Summary
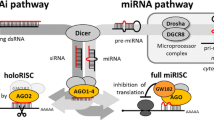
An efficient RNAi largely depends on optimal design of the siRNA. In recent studies, Dicer substrates were found to be more potent than classical synthetic 21-mer siRNAs, suggesting a coupling of the Dicer-mediated processing step to the efficient assembly of the silencing complex, RISC. We describe the fundamental principles and experimental results leading to optimal Dicer substrates.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Fire, A., Xu, S., Montgomery, M. K., Kostas, S. A., Driver, S. E., and Mello, C. C. (1998). Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811.
Hannon, G. J., and Rossi, J. J. (2004). Unlocking the potential of the human genome with RNA interference. Nature 431, 371–378.
Elbashir, S. M., Harborth, J., Lendeckel, W., Yalcin, A., Weber, K., and Tuschl, T. (2001). Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498.
Jackson, A. L., Bartz, S. R., Schelter, J., et al. (2003). Expression profiling reveals off-target gene regulation by RNAi. Nat. Biotechnol. 21, 635–637.
Kim, D. H., Behlke, M. A., Rose, S. D., Chang, M. S., Choi, S., and Rossi, J. J. (2005). Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nat. Biotechnol. 23, 222–226.
Siolas, D., Lerner, C., Burchard, J., et al. (2005). Synthetic shRNAs as potent RNAi triggers. Nat. Biotechnol. 23, 227–231.
Pham, J. W., Pellino, J. L., Lee, Y. S., Carthew, R. W., and Sontheimer, E. J. (2004). A Dicer-2-dependent 80s complex cleaves targeted mRNAs during RNAi in Drosophila. Cell 117, 83–94.
Gregory, R. I., Chendrimada, T. P., Cooch, N., and Shiekhattar, R. (2005). Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123, 631–640.
Maniataki, E., and Mourelatos, Z.(2005). A human, ATP-independent, RISC assembly machine fueled by pre-miRNA.Genes Dev.19, 2979–2990.
Krol, J., Sobczak, K., Wilczynska, U., et al.(2004). Structural features of microRNA(miRNA) precursors and their relevance to miRNA biogenesis and small interfering RNA/short hairpin RNA design.J. Biol. Chem.279, 42230–42239.
Rose, S. D., Kim, D. H., Amarzguioui, M., et al.(2005). Functional polarity is introduced by Dicer processing of short substrate RNAs. Nucleic Acids Res.33, 4140–4156.
Elbashir, S. M., Lendeckel, W., and Tuschl, T.(2001). RNA interference is mediated by 21- and 22-nucleotide RNAs.Genes Dev.15, 188–200.
Heale, B. S., Soifer, H. S., Bowers, C., and Rossi, J. J.(2005). siRNA target site secondary structure predictions using local stable substructures.Nucleic Acids Res.33, e30.
Amarzguioui, M., Holen, T., Babaie, E., and Prydz, H.(2003). Tolerance for mutations and chemical modifications in a siRNA.Nucleic Acids Res.31, 589–595.
Du, Q., Thonberg, H., Wang, J., Wahlestedt, C., and Liang, Z. (2005). A systematic analysis of the silencing effects of an active siRNA at all single-nucleotide mismatched target sites.Nucleic Acids Res.33, 1671–1677.
Elbashir, S. M., Martinez, J., Patkaniowska, A., Lendeckel, W., and Tuschl, T.(2001). Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate.EMBO J.20, 6877–6888.
Holen, T., Amarzguioui, M., Wiiger, M. T., Babaie, E., and Prydz, H. (2002). Positional effects of short interfering RNAs targeting the human coagulation trigger Tissue Factor.Nucleic Acids Res.30, 1757–1766.
Miller, V. M., Gouvion, C. M., Davidson, B. L., and Paulson, H. L. (2004). Targeting Alzheimer’s disease genes with RNA interference: An efficient strategy for silencing mutant alleles.Nucleic Acids Res.32, 661–668.
Birmingham, A., Anderson, E. M., Reynolds, A., et al.(2006). 3′ UTR seed matches, but not overall identity, are associated with RNAi off-targets.Nat. Meth.3, 199–204.
Hornung, V., Guenthner-Biller, M., Bourquin, C., et al.(2005). Sequence-specific potent induction of IFN-α by short interfering RNA in plasmacytoid dendritic cells through TLR7.Nat. Med.11, 263–270.
Judge, A. D., Sood, V., Shaw, J. R., Fang, D., McClintock, K., and MacLachlan, I.(2005). Sequence-dependent stimulation of the mammalian innate immune response by synthetic siRNA.Nat. Biotechnol.23, 457–562.
Kariko, K., Bhuyan, P., Capodici, J., and Weissman, D.(2004). Small interfering RNAsmediate sequence-independent gene suppression and induce immune activation by signaling through Toll-like receptor 3. J. Immunol.172, 6545–6549.
Robbins, M. A., and Rossi, J. J.(2005). Sensing the danger in RNA. Nat. Med.11, 250–251.
Sioud, M.(2005). Induction of inflammatory cytokines and interferon responses by double- stranded and single-stranded siRNAs is sequence-dependent and requires endosomal localization.J. Mol. Biol.348, 1079–1090.
Marques, J. T., Devosse, T., Wang, D., et al.(2006). A structural basis for discriminating between self and nonself double-stranded RNAs in mammalian cells.Nat. Biotechnol.24, 559–565.
Amarzguioui, M.(2004). Improved siRNA-mediated silencing in refractory adherent cell lines by detachment and transfection in suspension.Biotechniques 36, 766–768, 770.
Rossi, J. J.(2006). Partnering aptamer and RNAi technologies.Mol. Ther. 14, 461–462.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2008 Humana Press, a part of Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Amarzguioui, M., Rossi, J.J. (2008). Principles of Dicer Substrate (D-siRNA) Design and Function. In: Barik, S. (eds) RNAi. Methods in Molecular Biology™, vol 442. Humana Press. https://doi.org/10.1007/978-1-59745-191-8_1
Download citation
DOI: https://doi.org/10.1007/978-1-59745-191-8_1
Publisher Name: Humana Press
Print ISBN: 978-1-58829-874-4
Online ISBN: 978-1-59745-191-8
eBook Packages: Springer Protocols