Abstract
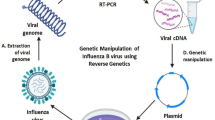
Reverse genetics is the creation of a virus from a full-length cDNA copy of the viral genome, referred to as an “infectious clone,” and is one of the most powerful genetic tools in modern virology. Since its development in 1999, plasmid-based reverse genetics has been effectively applied to numerous aspects of influenza studies which include revolutionizing the production of seasonal and pandemic influenza vaccine seed strains. Although continual improvement in reverse genetics system is being made in different laboratories for the efficient rescue of the influenza virus, the basic concept of synthesizing viral RNA using RNA polymerase I remains the same. Coupled with in vitro mutagenesis, reverse genetics can be applied widely to accelerate progress in understanding the influenza virus life cycle, the generation of customized vaccine seed strains, development of live-attenuated vaccines, and the use of influenza virus as vaccine and gene delivery vectors.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Engelhardt OG (2013) Many ways to make an influenza virus: review of influenza virus reverse genetics methods. Influenza Other Respir Viruses 7:249–256
Boyer J-C, Haenni A-L (1994) Infectious transcripts and cDNA clones of RNA viruses. Virology 198:415–426
Palese P, Zheng H, Engelhardt OG, Pleschka S, Garcia-Sastre A (1996) Negative-strand RNA viruses: genetic engineering and applications. Proc Natl Acad Sci U S A 93:11354–11358
Lamb RA, Krug RM (2001) Orthomyxoviridae: the viruses and their replication. In: Knipe DM, Howle PM (eds) Fields virology. Lippincott Williams & Wilkins, Philadelphia, PA, pp 1487–1532
Neumann G, Watanabe T, Ito H, Watanabe S, Goto H, Gao P, Hughes M, Perez DR, Donis R, Hoffmann E, Hobom G, Kawaoka Y (1999) Generation of influenza A viruses entirely from cloned cDNAs. Proc Natl Acad Sci U S A 96:9345–9350
Fodor E, Devenish L, Engelhardt OG, Palese P, Brownlee GG, Garcia-Sastre A (1999) Rescue of influenza A virus from recombinant DNA. J Virol 73:9679–9682
Neumann G, Kawaoka Y (2002) Synthesis of influenza virus: new impetus from an old enzyme, RNA polymerase I. Virus Res 82:153–158
Hoffmann E, Nuemann G, Hobom G, Webster RG (2000) “Ambisense” approach for the generation of influenza A virus: vRNA and mRNA synthesis from one template. Virology 267:310–317
Neumann G, Fujii K, Kino Y, Kawaoka Y (2005) An improved reverse genetics system for influenza A virus generation and its implications for vaccine production. Proc Natl Acad Sci U S A 102:16825–16829
Chen H, Ye J, Xu K, Angel M, Shao H, Ferrero A, Sutton T, Perez DR (2012) Partial and full PCR-based reverse genetics strategy for influenza viruses. PLoS One 7:e46378
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Plainview, NY
Hoffmann E, Krauss S, Perez D, Webby R, Webster RG (2002) Eight-plasmid system for rapid generation of influenza virus vaccines. Vaccine 20:3165–3170
Wang L, Lee CW (2009) Sequencing and mutational analysis of the non-coding regions of influenza A virus. Vet Microbiol 135:239–247
Lee CW, Senne DA, Suarez DL (2004) Generation of reassortant influenza vaccines by reverse genetics that allows utilization of a DIVA (Differentiating Infected from Vaccinated Animals) strategy for the control of avian influenza. Vaccine 22:3175–3181
Lee CW, Lee YJ, Senne DA, Suarez DL (2006) Pathogenic potential of North American H7N2 avian influenza virus: a mutagenesis study using reverse genetics. Virology 353:388–395
Lee CW, Senne DA, Suarez DL (2006) Production of reference antisera against 15 hemagglutinin subtypes of influenza virus by DNA vaccination of chickens. Clin Vaccine Immunol 13:395–402
Czudai-Matwich V, Schnare M, Pinkenburg O (2013) A simple and fast system for cloning influenza A virus gene segments into pHW2000- and pCAGGS-based vectors. Arch Virol 158:2049–2058
Mostafa A, Kanrai P, Ziebuhr J, Pleschka S (2013) Improved dual promotor-driven reverse genetics system for influenza viruses. J Virol Methods 193:603–610
Massin P, Rodrigues P, Marasescu M, van der Werf S, Naffakh N (2005) Cloning of the chicken RNA polymerase I promoter and use for reverse genetics of influenza A viruses in avian cells. J Virol 79:13811–13816
Wang Z, Duke GM (2007) Cloning of the canine RNA polymerase I promoter and establishment of reverse genetics for influenza A and B in MDCK cells. Virol J 4:102
Song MS, Baek YH, Pascua PN, Kwon HI, Park SJ, Kim EH, Lim GJ, Choi YK (2013) Establishment of Vero cell RNA polymerase I-driven reverse genetics for Influenza A virus and its application for pandemic (H1N1) 2009 influenza virus vaccine production. J Gen Virol 94:1230–1235
Tang Y, Lee CW, Zhang Y, Senne DA, Dearth R, Byrum B, Perez D, Suarez DL, Saif YM (2005) Isolation and characterization of H3N2 influenza A virus from Turkeys. Avian Dis 49:207–213
Acknowledgments
The authors would like to thank Dr. Gerd Hobom (Institut fur Mikro- und Molekularbiologie, Giessen, Germany) for kindly providing pHH21 vector and Dr. Yoshihiro Kawaoka (University of Wisconsin) for providing plasmids for A/WSN/33 (H1N1) reverse genetics.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this protocol
Cite this protocol
Lee, CW. (2014). Reverse Genetics of Influenza Virus. In: Spackman, E. (eds) Animal Influenza Virus. Methods in Molecular Biology, vol 1161. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-0758-8_4
Download citation
DOI: https://doi.org/10.1007/978-1-4939-0758-8_4
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-0757-1
Online ISBN: 978-1-4939-0758-8
eBook Packages: Springer Protocols