Abstract
The reverse genetics system for influenza A viruses described by Hoffmann et al. (Virology 267(2):310–317, 2000, Proc Natl Acad Sci USA 97(11):6108–6113, 2000, ArchVirol 146(12):2275–2289, 2001) is one of the most commonly used. However, this cloning strategy is rather time-consuming and lacks a selection marker to identify positive clones carrying viral genes. We report here the optimization of the cloning protocol of viral genes into pHW2000 by (i) introducing a selection marker and (ii) simplifying the cloning strategy: now the cloning reaction takes only a few minutes and, in addition, is independent of internal restriction sites for BsmBI/Esp3I, BsaI or AarI. In order to accelerate the whole cloning protocol for the generation of recombinant viruses, we first introduced a lacP/Z-element (lac-promoter/lacZα-fragment) between the two BsmBI sites of pHW2000 to allow selection of positive clones by blue/white screening. Then we optimized the digestion/ligation-protocol: In our system, enzymatic digestion and ligation of PCR products into the vector is performed in a single “one-tube” reaction. Due to this strategy, time and material consumption is reduced by a great amount, as vector and cDNA do not have to be digested and purified prior to the ligation. Therefore, this one-tube reaction yields positive clones with high efficiency and fidelity, again saving time and material, which were formerly required for screening and analyzing clones. Finally, to add more versatility to the system, we also created an entry vector based on TA-cloning. This entry vector provides several advantages: inserted genes can easily be modified, e.g., by site-directed mutagenesis or tag attachment, and then subcloned into pHW2000 or other plasmids containing a similar cloning site (e.g., our modified pCAGGS-Esp-blue) by the same rapid and reliable one-tube reaction protocol described here. In fact, the presented protocol is suitable to be adapted to other reverse genetics systems (e.g., those for members of the order Mononegavirales or the family Bunyaviridae) or cloning of genes in general.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The generation of recombinant influenza A viruses from cloned cDNA provides a powerful tool to study the effects of reassortment events or single point mutations in viral genes. One of the most commonly used cloning systems is the bidirectional approach first described by Hoffmann and colleagues [1, 2]. In the described plasmid (pHW2000), the viral gene is flanked by both a pol II promoter element (truncated CMV promoter, bovine growth hormone poly-A signal [BGHpA]) and a pol I promoter element (truncated human pol I promoter, truncated murine pol I terminator) in positive and negative orientation, respectively. Therefore, from this plasmid, both viral mRNA (pol II) and vRNA (pol I) can be synthesized without the need of additional helper viruses or further transfection with other polymerase genes, e.g., influenza polymerase subunits [4, 5] or T7 RNA polymerase [6].
Due to the nature of the pol I promoter [7], viral genes have to be cloned into pHW2000 via a defined protocol in order to maintain the sequence integrity of the 3′ and 5′ ends of cRNA and vRNA. To achieve this specificity, a set of “outside cutting” endonucleases can be utilized: BsmBI (or its isoschizomer Esp3I), BsaI or AarI, as described by Hoffmann et al. [3]. These enzymes cut within a defined distance of the recognition site (e.g., for Esp3I, the recognition site is CGTCTCN1/N5) and thereby create overhangs with sequences that can be different from that of the recognition site itself. This property thus allows the ligation of viral genes into pHW2000 without adding, removing or changing the 3′ and 5′ ends of the viral RNA. There are, however, several circumstances that make cloning of a complete viral genome by the standard cloning technique difficult. Typically, viral cDNA is generated by reverse transcription PCR (RT-PCR) after RNA extraction. This cDNA has to be purified to remove small by-products and components of the reaction mix. Afterwards, the purified cDNA is digested by one of the three enzymes mentioned above to generate sticky ends for ligation. Here, the first difficulty arises as some viral genes contain recognition sites in their sequence for one or more of these three endonucleases. Representative examples are given in the publication of Hoffmann et al. [3], where WSN-PB2 contains two BsmBI sites while WSN-PB1 contains three BsaI sites and NP contains one AarI site. A mix of cloning primers, as described by Hoffmann and colleagues [3], is therefore required to create the full recombinant WSN set of plasmids. Next, the digested viral cDNA has to be purified again from the reaction mix prior to the ligation. After ligation and transformation of competent bacteria, clones have to be selected for analysis. Here, another drawback of the original pHW2000 plasmid is the lack of a selection marker helping to identify positive clones. This means that in the case of difficult-to-clone genes such as HA or PB2, a large number of clones have to be analyzed. This again is another time-consuming and expensive step.
In order to improve cloning efficiency and reduce time and material cost, we decided to modify pHW2000 and the whole cloning protocol in several ways. First of all, we introduced the lac operon (lacP/Z), containing the lac promoter and lacZα gene, into pHW2000 as a selection marker for blue/white screening to identify clones carrying a viral gene. Next, we adapted the one-tube cloning protocol that has been described for the commercially available StarGate® cloning system (IBA GmbH Göttingen, Germany [8]) and by Engler and colleagues [9] for cloning influenza gene segments into our modified plasmids. Since the general idea of our protocol is not restricted to pHW2000 alone, we also created pCAGGS-blue variants of this high-level expression plasmid [10] for Esp3I or BsaI cloning. In addition to that, we created an entry vector based on the well-characterized TA cloning system [11], which allows easy modification of viral genes and transfer of such genes by a one-tube reaction into any plasmid providing the matching four-nucleotide overhangs.
Materials and methods
Generation of modified pHW2000- and pCAGGS plasmids
All kits and enzymes were obtained from Fermentas/Thermo Scientific if not indicated otherwise. I) pHW2000-Esp-blue was generated from the pHW2000 [2] vector. First, lacP/Z (accession no. FB506781.1) was amplified from the pASG vector (IBA) by PCR using the indicated primers (forward, 5′ AAGGGAAGAGACGCTGCAGCCCAATACGCAAAC 3′, and reverse, 5′ TTTATTTGAGACGCTCGAGGTGAACCATCACCC 3′; the Esp3I recognition motif is underlined) and the proofreading Phusion Polymerase in HF buffer. Then, pHW2000 and the resulting PCR product were digested with Esp3I, separated by gel electrophoresis and extracted using a gel extraction kit. Ligation using T4 DNA ligase was performed for 1 h at 37 °C. Competent DH5α cells were transformed with the mixture [12] and plated onto LB-ampicillin plates containing X-Gal and IPTG. Blue colonies were picked, and clones were analyzed by restriction mapping. II) In pHW2000Δ-Esp-blue, we deleted all vector-based BsaI sites (Fig. 1, highlighted in bold; for this and all other vectors, “Δ” indicates BsaI-free). For this, we first exchanged the ApaLI fragment (containing part of the ampicillin cassette and part of the ori region) (Fig. 1, framed) as follows: pHW2000-Esp-blue and pESG5 (IBA) were digested with ApaLI (NEB), and fragments were purified using a gel extraction kit and ligated. The resulting plasmid (pHW2000i-Esp-blue) thus contained only one remaining BsaI site. To remove this second BsaI site, pHW2000i-Esp-blue was digested with NheI and NcoI, and pESG5 was digested with NcoI and XbaI to exchange part of the CMV enhancer/promoter (Fig. 1, underlined). Again, both fragments were purified by gel extraction and ligated. The resultant vector, pHW2000Δ-Esp-blue, was now free of any internal BsaI sites. III) Vector pHW2000Δ-Bsa-blue was generated by exchanging the lacP/Z insert. For this, pHW2000Δ-Esp-blue was digested with Esp3I. The new lacP/Z was amplified from pHW2000-Esp-blue by PCR with the indicated primers (forward, 5′ TATAGGGAAGAGACCCTGCAGCCCAATACGCAAAC 3′, and reverse, 5′ TTAATATTTGAGACCCTCGAGGTGAACCATCACCC 3′; the BsaI recognition motif is underlined) and digested with BsaI (NEB). The vector and PCR product were extracted from a gel and ligated, and clones were analyzed by restriction mapping. IV) pCAGGS-Esp-blue, pCAGGSΔ-Esp-blue and pCAGGSΔ-Bsa-blue were generated by exchanging the ApaLI fragment and inserting lacP/Z fragments as described above.
Plasmid maps of pHW2000-Esp-blue and pCAGGS-Esp-blue. These plasmids where used for the generation of the ΔBsaI variants of pHW2000 and pCAGGS (pHW2000Δ-Esp-blue, pHW2000Δ-Bsa-blue, pCAGGSΔ-Esp-blue and pCAGGSΔ-Bsa-blue, respectively). For both pHW2000 and pCAGGS, the ApaLI fragment indicated (framed restriction sites) was exchanged with the corresponding ApaLI fragment of pESG-IBA5, which does not contain an internal BsaI restriction site. In pHW2000, the second BsaI site was removed by exchanging the corresponding fragment of the pESG-IBA5 vector (NcoI and NheI, underlined restriction sites). The vector-based BsaI restriction sites are shown in bold type
Construction of the entry vector pEntryT
The entry vector for TA cloning pEntryT was constructed by digestion of pBluescript (Stratagene) using LguI and AdeI to isolate lacP/Z. After digestion, ends of the excised lacP/Z were blunted using T4 DNA polymerase. The kanamycin backbone (including ori) was amplified by PCR from Entry-Vector (IBA) with the following primers: forward, 5′ ATCCAAAGGCGGTAATACGGTTATC 3′, and reverse, 5′ TGGCACTTTTCGGGGAAATGTGC 3′. The backbone and lacP/Z fragments were ligated, and chemically competent DH5α cells were transformed with the ligated fragments and plated onto LB-kanamycin plates with X-Gal and IPTG. Plasmid DNA from blue clones was analyzed. The vector that was obtained was then linearized with EcoRV and purified using a PCR purification kit. T-overhangs were generated using Taq polymerase [13] as follows: Linearized vector was incubated in Taq buffer with MgCl2 and dTTP. After 2 h at 72 °C, the vector was purified and ligated overnight at 16 °C. This ligation step eliminates vectors without or with only one T-tail by re-ligation, dimerisation or oligomerisation. The linear vector containing T-overhangs was then purified by gel extraction.
Generation of PCR fragments
To perform the following exchange reaction or for entry cloning, we generated viral cDNAs using the primers pairs described by Hoffmann and colleagues [3], except those for the NA segment. The following NA primers were used to generate overhangs with Esp3I instead of BsaI: forward Bs-NA-1, 5′ TATTCGTCTCAGGGAGCAAAAGCAGGAGT 3′, and reverse Bs-NA-1413R, 5′ ATATCGTCTCGTATTAGTAGAAACAAGGAGTTTTTT 3′ (the Esp3I recognition motif is underlined). As the template for our experiments, we used cloned segments of A/WSN/33 (H1N1) in pHW2000. The PCR conditions were as described [3], and the PCR products were isolated by gel extraction.
Transfer into pHW-blue and pCAGGS-blue vectors
We used 25 ng plasmid DNA and PCR products of PB1 (120 ng), PA (115 ng), HA (90 ng), NP (80 ng), NA (70 ng), M (55 ng), NS (45 ng) for transfer into pHW2000-Esp-blue, and 120 ng of PB2 PCR product for transfer into pHW2000Δ-Bsa-blue. For transfer of NA into pCAGGSΔ-Esp-blue we used 50 ng plasmid DNA instead. For transfer of PB2 into pHW2000Δ-Esp-blue and pCAGGSΔ-Esp-blue, we used 240 ng PCR product and a 1:2 mixture of Esp3I and BsaI. For transfer of PB2 into pCAGGSΔ-Bsa-blue, we used 120 ng of PCR product and BsaI only. The reaction mixture was incubated for 1 h at 37 °C and 1 h at RT. The incubation at RT is only necessary if the segment contains internal Esp3I or BsaI restriction sites (i.e., PB2 or NA), the addition of fresh T4 DNA ligase or ATP is not required, as 37 °C is within the temperature optimum of the enzyme of 16-37 °C according to the manufacturer (Fermentas/Thermo Scientific). In Table 1, the composition of each reaction mix is listed.
Transfer into pEntryT
To generate A-overhangs, the PCR product was incubated in Taq buffer with dNTPs and Taq polymerase for 25 minutes at 72 °C. After this step, the PCR product was purified using a PCR purification kit, and its concentration was measured. The ratio of vector to insert should be 1:2. The size of the vector is 2.5 kb, and the composition of the reaction mix is given in Table 2.
The reaction was incubated at 4 °C overnight. As an example, we inserted the NA segment (70 ng) and PB2 (115 ng) into the pEntryT vector to obtain pEntry-NA and pEntry-PB2. Two microliters of each reaction mix was introduced into 50 μl of chemically competent DH5α cells, which were then plated onto LB-ampicillin plates with X-Gal and IPTG. DNA from white clones was analyzed. It has been published that the proportion of positive clones is around 80 % when using the Taq method, and 90 % when using terminal transferase and ddTTP [13].
Transfer from pEntry-NA and pEntry-PB2 into destination vectors (pHW2000-Esp-blue, pHW2000Δ-Bsa-blue, pCAGGS-Esp-blue or pCAGGSΔ-Bsa-blue)
The reaction conditions are given in Table 1. The amount of entry vector (pEntry-NA/pEntry-PB2) instead of the PCR product in this reaction was 100 ng.
Transfer of the M and HA segments into pHW2000-Esp-blue (kinetics)
A double standard reaction was prepared (50 μl, see Table 1) without adding Esp3I. Of this, 2 μl was taken as the 0 minute control. Next we added Esp3I and transferred 9 μl to a single tube for each time point. Incubation at 37 °C was stopped after 5, 10, 20, 30 and 60 minutes by transferring the tube to liquid nitrogen. After the last sample was collected, all tubes were thawed, and 2 μl of each reaction mixture was added to 50 μl of competent DH5α cells. Cells were incubated on ice for 30 minutes and 30 seconds at 42 °C. After an additional 2 minutes on ice, the cells were plated onto LB-ampicillin plates with X-Gal and IPTG.
The concentrations of plasmid DNA and PCR products were determined using a Nanodrop 1000, (PeqLab, Erlangen, Germany). Pictures of the reaction kinetic experiment were created using Molecular Imager® ChemiDoc XRS+ (BIO-RAD, Munich, Germany). Plasmid maps were generated with CloneManager 9 Basic Edition (Scientific & Educational Software).
Results
The lacP/Z gene region was cloned between the two BsmBI/Esp3I-sites, replacing the short nucleotide spacer between the restriction sites. If the ligation of a viral gene is successful, the whole lacP/Z is replaced by the viral gene, ensuring the correct expression of viral RNA from the plasmid. Next, we performed restriction digestion of both vector and purified cDNA, as well as ligation in a single reaction. To obtain higher restriction/ligation-rates, we decided to use the isoschizomer Esp3I (Fermentas/Thermo Scientific) instead of BsmBI (NEB). Esp3I has a temperature optimum of 37 °C instead of 55 °C for BsmBI, resulting in a higher activity of the T4 DNA ligase, which has a temperature optimum around 16-37 °C.
Direct cloning of viral cDNAs into pHW2000-Esp-blue, pHW2000Δ-Esp-blue, pHW2000Δ-Bsa-blue, pCAGGS-Esp-blue, pCAGGSΔ-Esp-blue and pCAGGSΔ-Bsa-blue
The mechanism of the exchange reaction is illustrated in Fig. 2. Both the PCR product (highlighted in dark gray throughout the figure) (Fig. 2A) and the vector (Fig. 2B) are digested by Esp3I or BsaI in the presence of T4 DNA ligase. Due to the parallel digestion/ligation reaction, the cleaved ends of the PCR product, as well as LacP/Z can be re-ligated and re-cut. Thus, reactions A and B are both equilibrium reactions. However, reaction C is the key step of this cloning system. Since the Esp3I recognition sequences are located at the far ends of the PCR product and LacP/Z, they are removed from both viral cDNA and the vector after digestion. Therefore, once the vector and the cDNA are ligated, this product cannot be digested with Esp3I or BsaI anymore, removing it from the reaction equilibrium of digestion/re-ligation (Fig. 2C). In the case of cDNAs where no internal Esp3I sites are present (PB1, HA, PA, NP, M and NS), an incubation of up to 1 h at 37 °C is sufficient, and competent bacteria can be transformed directly with the plasmids (in this experiment we used DH5α, but in principle, any strain with the ΔlacZM15 genotype would be suitable). If there are internal Esp3I recognition sites (NA has one internal Esp3I motif), the end product of the reaction is an equilibrium between closed and open vector. This equilibrium can be shifted towards the closed vector by further incubation at RT due to the different temperature optima of Esp3I (37 °C) and T4 DNA ligase (16-37 °C). The same applies for segments with two internal Esp3I sites (PB2).
Direct cloning of cDNAs into pHW2000-/pCAGGS-blue vectors. A: Both ends of the PCR product (highlighted in gray) are cleaved by Esp3I/BsaI and can be re-ligated or re-cut during the reaction. B: Esp3I/BsaI digestion of the vector removes the selection marker, creating compatible ends for ligation of the PCR product. The vector and selection marker can be re-ligated or re-cut as well. C: Since Esp3I/BsaI digestion removes the recognition sites from both the PCR product and the vector, the PCR product cannot be re-cut from the final ligation product once ligation has occurred
Plasmids pHW2000-Esp-blue, pHW2000Δ-Esp-blue, pHW2000Δ-Bsa-blue, pCAGGS-Esp-blue, pCAGGSΔ-Esp-blue and pCAGGSΔ-Bsa-blue were generated and prepared as described in Materials and methods. Viral cDNAs for all eight gene segments were generated and purified by gel extraction. The specific one-tube reaction conditions for each gene segment are listed in Table 1 and explained in detail in Materials and methods. Table 3 gives an overview of the results obtained for all viral cDNAs and the restriction enzymes used for analysis. As indicated, direct cloning of cDNAs into the modified pHW2000 vector yields a very large number of positive clones (12 of 12 for all genes), which can be easily discriminated from negative clones, which turn blue.
For direct cloning of cDNAs into our modified pCAGGS plasmids, we selected PB2 and NA as examples, because they both contain internal Esp3I sites (two in PB2 and one in NA) and therefore represent difficult-to-clone genes. As listed in Table 4, our system again resulted in a very large number of positive clones (pCAGGSΔ-Esp-blue (PB2 12/12) pCAGGSΔ-Bsa-blue (PB2 11/12) and pCAGGSΔ-Esp-blue (NA 12/12), as determined by restriction analysis.
Cloning into pEntryT by TA cloning and subcloning of cDNAs into pHW- and pCAGGS-related vectors
Subcloning of segments from one plasmid into another is a common technique. It is often easier and faster to use commercially available systems such as TA cloning to quickly clone a given gene in order to obtain sequence information or to simply amplify enough DNA for traditional cloning methods. In the case of influenza virus genes, however, TA cloning requires some further consideration. Subcloning of a TA-cloned cDNA into pHW2000 using any other restriction enzyme other than the three described outside cutters, BsmBI/Esp3I, BsaI and AarI, is impossible, since the restrictive requirements of the pol I promoter cassette do not allow any modification of the 3′ and 5′ ends of the viral cDNA/RNA. Excision of the viral cDNA by one of the aforementioned restriction enzymes and successive ligation into pHW2000 is possible, but again, this step faces the already described problems of time and material consumption as well as the consideration of internal restriction sites. Especially for enzymes like BsaI, which was used previously as a restriction enzyme for cloning of PB2 and NA, internal TA-vector BsaI sites can make the cloning procedure even more complicated. Since digestion and ligation reactions in our new cloning system are performed in a single tube, it is important to separate the template vector from the resulting vector after the reaction. Therefore, we designed our TA cloning vector to harbor an antibiotic resistance cassette that is different from pHW2000 and pCAGGS. Both plasmids carry an ampicillin selection cassette. For this reason, we constructed a small vector as an entry vector for TA cloning that lacks internal BsaI or BsmBI/Esp3I sites and provides kanamycin resistance.
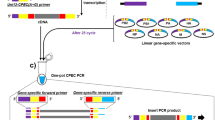
The cloning procedure for the TA entry vector is presented in Fig. 3A. First, we prepared the entry vector pEntryT and the purified cDNAs of two gene segments as described in Materials and methods. For our experiments, we again selected PB2 and NA for the same reasons as described above. As listed in Table 5, ligation of both cDNAs was successful and yielded a large number of positive clones (pEntry-PB2 7/10 and pEntry-NA 9/12) as determined by digestion and gel electrophoresis. We next selected one of the positive clones of each gene for subcloning (Fig. 3B) into pHW2000Δ-Bsa-blue and pCAGGSΔ-Bsa-blue (PB2) or pHW2000-Esp-blue and pCAGGS-Esp-blue (NA). The results of this experiment are listed in Table 6. The transfer reaction yielded a large number of white colonies, and restriction analysis of white colonies resulted in 12 out of 12 clones being positive for all reactions except pHW2000-NA (11/12).
Cloning of cDNA into TA-vector pEntryT and subcloning into pHW-/pCAGGS-plasmids. A: First, the A-tailed PCR product (highlighted in gray) is ligated into T-tailed pEntryT by TA cloning. pEntryT is suitable for mutagenesis or modification of the insert. B: To subclone the gene of interest from the entry vector into any target vector (e.g., pHW2000- and pCAGGS-Esp or Bsa-blue), the same one-tube reaction described for direct cloning can now be employed. In the case of BsaI digestion, the “Δ-plasmids” are recommended for a higher ligation efficiency, since all internal BsaI sites have been removed. Note that pEntryT and the target vectors confer kanamycin and ampicillin resistance, respectively, for positive selection of clones
Direct cloning in a single reaction is performed in a few minutes
To analyze whether a longer incubation time in our one-tube reaction would result in an even higher yield of positive clones, we performed a reaction kinetics experiment. We selected the M segment as well as the HA segment (M represents an easy-to-clone segment, whereas HA is difficult to clone), and incubated the reaction for one hour. We stopped the reaction after 5, 10, 20, 30 and 60 minutes by freezing and thereafter transformed competent DH5α bacteria. As depicted in Fig. 4, we obtained a very large number of white clones compared to blue colonies after only five minutes of incubation. The ratio of white to blue colonies can be further improved by a longer incubation time, with only a few colonies turning blue after one hour of incubation. For HA, we obtained fewer white colonies after five minutes of incubation (data not shown). However, of these, 12 of 12 colonies were positive for HA insertion, as determined by restriction digestion. With a prolonged reaction time, the number of blue colonies decreased, and after one hour of incubation, the ratio of blue to white colonies was comparable to that obtained with the M segment.
Reaction kinetics of the M segment. LB plates (Amp/IPTG/X-Gal) for the cloning reaction of the M segment were scanned with Molecular Imager® ChemiDoc XRS+. As little as 5 minutes of digestion/ligation yields a large number of white colonies. With increasing reaction time, the number of blue colonies decreases
Discussion
In order to generate recombinant influenza viruses, the cloning system published by Hoffmann et al. [2] has several advantages over other published systems (i.e., the reduced amount of plasmids needed, no need for another helper virus or complementary RNA polymerase). Another reason why this system is so easy to use is that it is compatible with the cloning method described in ref 3. To generate a full set of eight influenza virus genes for the generation of recombinant viruses, only a small number of cloning primers is needed. As we have already pointed out, however, there are some disadvantages of the protocol described in those two publications, namely the need to select the appropriate cloning primers based on internal restriction sites within the genes of interest and, subsequently, the time and materials needed to purify and screen the ligation components and the resulting clones.
While in the process of creating a new recombinant virus, we faced some difficulties with the cloning procedure, initiating our search for a way to clone viral genes and screen for positive clones. Here, we present a new protocol in which we have modified the previously described plasmids pHW2000 and pCAGGS to allow a fast and reliable screening of positive clones by blue/white screening and, furthermore, optimized the cloning protocol itself. It is now possible to generate plasmids that are ready for use in the generation of recombinant viruses or protein expression in a one-tube reaction with very high efficiency within a few minutes. We combined the cloning strategies from different sources (TA Cloning® from Invitrogen, StarGate® Cloning from IBA) and designed our own protocol, which was optimized for the cloning of influenza virus genes into pHW2000 and pCAGGS.
The one-tube restriction/ligation protocol we have presented here has several advantages compared to the standard cloning protocol:
-
1.
The amount of insert DNA and plasmid DNA required has been reduced from several micrograms to a few nanograms.
-
2.
Neither the insert nor the plasmid has to be pre-digested; the plasmid can be used directly from a plasmid preparation (mini or maxi scale), and the viral cDNA has to be gel-purified only once after RT-PCR.
-
3.
Since only very small amounts of DNA are required, the restriction/ligation reaction is performed with only a few units of restriction enzyme and DNA ligase, reducing the costs even further.
-
4.
The incubation time can be as short as five minutes and still yields almost 100 % positive clones. As we demonstrated for all eight influenza virus genes, there is a very high rate of white colonies resulting from successful cloning. This large proportion of positive clones is also a major step forward in reducing the time and material used to screen for positive cloning results.
-
5.
The one-tube reaction allows the successful cloning of genes in their full length and with the correct sequence, even if they contain internal restriction sites for the digestion enzyme used (i.e., BsmBI/Esp3I in PB2 or NA), as restriction digestion and ligation are at an equilibrium at 37 °C, which is shifted towards ligation of available ends when this reaction is further incubated at room temperature. This results in a large number of positive clones. Therefore, with our protocol, it is not necessary to consider internal restriction sites, as they are re-ligated as well, allowing the use of just one set of primers.
Over the last few years, three new cloning techniques were published that allow rapid cloning of viral genes and do not require digestion of cDNAs, thereby circumventing the problem of internal restriction sites. The first method, published by Stech et al. [14] describes the use of viral cDNA as megaprimers for target-primed plasmid amplification. The other two groups describing rapid cloning methods base their protocols on “ligation-independent cloning” instead, where Wang et al. [15] utilize homologous recombination of viral cDNA with modified pHH21 plasmids that harbour the conserved 3′ and 5′ UTRs of influenza A virus gene segments. Similarly, Zhou et al. [16] introduce a technique where apparently all eight gene segments are amplified in a single “multiplex” RT-PCR reaction and then inserted into a modified pHH21 vector by In-Fusion cloning (Clonetech). All of these protocols, despite accelerating the cloning process, have several drawbacks compared to our method: they either require larger amounts of cDNA and several successive PCR reactions, which may introduce unwanted mutations [14] or yield a lower ratio of positive clones in the case of larger gene segments like those of the polymerase subunits [15, 16]. Additionally, with the protocol of Zhou et al., a very large number of clones probably have to be screened in order to obtain plasmids for each of the eight gene segments. Furthermore, for the two protocols utilizing ligation-independent cloning, the vector has to be digested and purified prior to the cloning reaction, and the In-Fusion technique also requires the use of a commercial cloning kit. Therefore, our protocol is advantageous with respect to fidelity, efficiency and time and material consumption.
As cloning into pHW2000 is a one-way process, successive subcloning into other plasmids such as pCAGGS is also quite time-consuming. Previously, we cut viral genes from pHW2000 with SalI and SmaI and ligated them into SmaI/XhoI-digested pCAGGS. As is already apparent, this protocol requires time and a large amount of plasmid DNA, since a small amount of DNA is lost with each purification step. Another method, adapted to the one-tube cloning reaction using pHW2000, is to generate a PCR product of the viral gene (using pHW2000 as template and the cloning primers described by Hoffmann) for ligation into our pCAGGS-blue vectors. This protocol, just as for pHW2000 requires only a few nanograms of DNA and is quite fast. To minimize the risk of PCR-based errors, we created an entry vector from which any inserted gene can be transferred to all Esp- and Bsa vectors (pHW2000-Esp-blue or pCAGGS-Esp-blue, etc.) by a simple one-tube exchange reaction following the same protocol used for direct cloning. This new vector (pEntryT) has the advantage that any viral cDNA synthesized by RT-PCR can be ligated into it without the need for enzymatic digestion [13]. As the primers used for RT-PCR contain the required Esp3I or BsaI recognition sites, these are included in the vector as well. Therefore, any gene ligated into the entry vector can be excised by Esp3I or BsaI and provides the correct overhangs for ligation into our modified pHW2000 or pCAGGS plasmids. Additionally, the entry vector is designed to be as small as possible, making site-directed mutagenesis or other modifications (e.g., attachment of tags) very easy. In this way, a library of entry vectors harboring the viral genes can be generated very fast, modified as desired, and then transferred into any expression plasmid that provides the matching overhangs.
There is one commercially available TOPO TA vector that is suitable as an entry vector, pCR®8/GW/TOPO®, 2817 bp, that uses spectinomycin as resistance maker. With such a TOPO TA vector, cloning can be performed as described above within a few minutes and even blunt-end ligation of purified cDNAs with high efficiency is possible [17]. For any non-TOPO TA cloning vector like our pEntryT-vector, however, one has to consider the fact that most proof reading polymerases such as Pfu or Phusion® do not create the required A-overhangs to match the vector-based T-overhang. It is therefore necessary to include a short incubation of the PCR product with Taq polymerase and dNTPs prior to ligation as described above. As a final remark, all of the plasmids presented here, as well as the cloning protocol, have been used extensively in our lab to create either recombinant viruses of different subtypes or to perform protein studies, ensuring that the method and plasmids are working as intended.
In summary, we present here an optimized protocol for the cloning and subcloning of influenza virus gene segments into pHW2000- and pCAGGS-related vectors, ready for the generation of recombinant viruses or protein expression. In addition, a novel entry vector is presented that allows TA cloning of cDNAs directly from RT-PCR and further modifications, as well as the option to subclone the inserted gene segments into any other vector harbouring the corresponding ligation sequences.
References
Hoffmann E et al (2000) “Ambisense” approach for the generation of influenza A virus: vRNA and mRNA synthesis from one template. Virology 267(2):310–317
Hoffmann E et al (2000) A DNA transfection system for generation of influenza A virus from eight plasmids. Proc Natl Acad Sci USA 97(11):6108–6113
Hoffmann E et al (2001) Universal primer set for the full-length amplification of all influenza A viruses. Arch Virol 146(12):2275–2289
Neumann G et al (1999) Generation of influenza A viruses entirely from cloned cDNAs. Proc Natl Acad Sci USA 96(16):9345–9350
Fodor E et al (1999) Rescue of influenza A virus from recombinant DNA. J Virol 73(11):9679–9682
de Wit E et al (2007) A reverse-genetics system for Influenza A virus using T7 RNA polymerase. J Gen Virol 88(Pt 4):1281–1287
Zobel A, Neumann G, Hobom G (1993) RNA polymerase I catalysed transcription of insert viral cDNA. Nucleic Acids Res 21(16):3607–3614
Selmer T, Pinkenburg O (2008) WO2008095927 (A1) Method of cloning at least one nucleic acid molecule of interest using type IIs restriction endonucleases, and corresponding cloning vectors, kits and system using type IIs restriction endonucleases. Patent Assignee: Philipps-University Marburg; Inventors: Selmer T, Pinkenburg O
Engler C, Kandzia R, Marillonnet S (2008) A one pot, one step, precision cloning method with high throughput capability. PLoS One 3(11):e3647
Niwa H, Yamamura K, Miyazaki J (1991) Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 108(2):193–199
Holton TA, Graham MW (1991) A simple and efficient method for direct cloning of PCR products using ddT-tailed vectors. Nucleic Acids Res 19(5):1156
Inoue H, Nojima H, Okayama H (1990) High efficiency transformation of Escherichia coli with plasmids. Gene 96(1):23–28
Zhou MY, Gomez-Sanchez CE (2000) Universal TA cloning. Curr Issues Mol Biol 2(1):1–7
Stech J et al (2008) Rapid and reliable universal cloning of influenza A virus genes by target-primed plasmid amplification. Nucleic Acids Res 36(21):e139
Wang S et al (2008) Simplified recombinational approach for influenza A virus reverse genetics. J Virol Methods 151(1):74–78
Zhou B et al (2009) Single-reaction genomic amplification accelerates sequencing and vaccine production for classical and Swine origin human influenza a viruses. J Virol 83(19):10309–10313
Geng L et al (2006) A universal cloning vector using vaccinia topoisomerase I. Mol Biotechnol 33(1):23–28
Acknowledgments
We thank Folker Schwalm for providing primers for the PA, HA and PB2 segments, Erich Hoffmann and Robert G. Webster for the pHW2000 plasmid, and J. Miyazaki for the pCAGGS plasmid. This work was supported by the FluResearchNet, funded by the Federal Ministry of Education and Research.
Conflict of interest
The authors have declared that no conflict of interest exists.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Czudai-Matwich, V., Schnare, M. & Pinkenburg, O. A simple and fast system for cloning influenza A virus gene segments into pHW2000- and pCAGGS-based vectors. Arch Virol 158, 2049–2058 (2013). https://doi.org/10.1007/s00705-013-1697-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-013-1697-4