Abstract
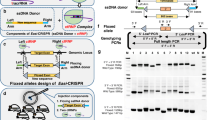
Recent development of Easi-CRISPR (Efficient additions with ssDNA inserts-CRISPR) that utilizes long single-stranded DNA (lssDNA) of 0.2–2 kbases in length as donor templates to insert large segments of novel DNA sequences or to replace endogenous genes at precise locations in the genome has enabled CRISPR-assisted genome editing to make strides toward a more simple and rapid workflow. By leveraging the notion that short single-stranded DNA oligo (<200 bases) serves as efficient donor in mouse zygotes for facilitating HDR-mediated genome editing, Easi-CRISPR expands to use lssDNA as the donor which accelerates the timeline to as little as 2 months for creating most types of genetically engineered mouse models (F0). Our lab (CGERC) has adopted Easi-CRISPR for multiple loci to generate mouse models over the past three plus years since its introduction. Here, we use two genes as examples to illustrate a step-by-step protocol for generating two commonly used models, including a knock-in (insertion of a reporter gene plus GOI) as well as a conditional knock-out model (via exon floxing). This protocol will focus more on molecular biology aspect, particularly we demonstrate two recently developed methods for lssDNA procuration: (1) PCR-based Takara Bio kit with modifications; (2) plasmid-retrieval-based CRISPR-CLIP (CRISPR-Clipped LssDNA via Incising Plasmid). Both methods are devised to retain sequence fidelity in lssDNA generated. In addition, CRISPR-CLIP directly retrieves lssDNA from DNA plasmid without using restriction enzymes through a PCR-free system hence carries virtually no restriction on sequence complexity, further mitigating limitations discussed in the original Easi-CRISPR protocol. We have alternated the use between both methods when suitable and successfully generated lssDNA templates via CRISPR-CLIP up to 3.5 kbases patched with multiple highly repetitive sequences, which is otherwise challenging to maneuver. Along with certain other modified workflow presented herein, Easi-CRISPR can be adapted to be more straightforward while applicable to generate mouse models in broader scope. (Certain figures and text passages presented in this chapter are reproduced from Shola et al. (The CRISPR J 3(2):109–122, 2020), published by Mary Ann Libert, Inc)
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
To maintain confidentiality of the research, partial sequences of the genes have been altered and some data shown herein are schematic for illustration purpose.
References
Cong L, Ran FA, Cox D et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Mali P, Yang L, Esvelt KM et al (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826
Suzuki K, Tsunekawa Y, Hernandez-Benitez R et al (2016) In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 540:144–149
Yang H, Wang H, Shivalila CS et al (2013) One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154:1370–1379
Gu B, Posfai E, Rossant J (2018) Efficient generation of targeted large insertions by microinjection into two-cell-stage mouse embryos. Nat Biotechnol 36:632–637
Quadros RM, Miura H, Harms DW et al (2017) Easi-CRISPR: a robust method for one-step generation of mice carrying conditional and insertion alleles using long ssDNA donors and CRISPR ribonucleoproteins. Genome Biol 18:92
Roth TL, Puig-Saus C, Yu R et al (2018) Reprogramming human T cell function and specificity with non-viral genome targeting. Nature 559:405–409
Yoshimi K, Kunihiro Y, Kaneko T et al (2016) ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes. Nat Commun 7:10431
Miura H, Quadros RM, Gurumurthy CB et al (2018) Easi-CRISPR for creating knock-in and conditional knockout mouse models using long ssDNA donors. Nat Protoc 13:195–215
Shola DTN, Yang C, Kewaldar V-S et al (2020) New additions to the CRISPR toolbox: CRISPR-CLONInG and CRISPR-CLIP for donor construction in genome editing. The CRISPR J 3(2):109–122
Gurumurthy CB, O’brien AR, Quadros RM et al (2019) Reproducibility of CRISPR-Cas9 methods for generation of conditional mouse alleles: a multi-center evaluation. Genome Biol 20:171
Terada R, Johzuka-Hisatomi Y, Saitoh M et al (2007) Gene targeting by homologous recombination as a biotechnological tool for rice functional genomics. Plant Physiol 144:846–856
Aida T, Chiyo K, Usami T et al (2015) Cloning-free CRISPR/Cas system facilitates functional cassette knock-in in mice. Genome Biol 16:87
Mashiko D, Young SA, Muto M et al (2014) Feasibility for a large scale mouse mutagenesis by injecting CRISPR/Cas plasmid into zygotes. Develop Growth Differ 56:122–129
Chen S, Lee B, Lee AY et al (2016) Highly efficient mouse genome editing by CRISPR ribonucleoprotein electroporation of zygotes. J Biol Chem 291:14457–14467
Bell CC, Magor GW, Gillinder KR et al (2014) A high-throughput screening strategy for detecting CRISPR-Cas9 induced mutations using next-generation sequencing. BMC Genomics 15:1002
Sentmanat MF, Peters ST, Florian CP et al (2018) A survey of validation strategies for CRISPR-Cas9 editing. Sci Rep 8:888
Li H, Beckman KA, Pessino V et al (2017) Design and specificity of long ssDNA donors for CRISPR-based knock-in. bioRxiv:178905
Richardson CD, Ray GJ, Dewitt MA et al (2016) Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat Biotechnol 34:339–344
Behringer RGM, Nagy KV, Nagy A (2014) Manipulating the mouse embryo: a laboratory manual, 4th edn. Cold Spring Harbor Laboratory Press, New York
Codner GF, Mianne J, Caulder A et al (2018) Application of long single-stranded DNA donors in genome editing: generation and validation of mouse mutants. BMC Biol 16:70
Acknowledgments
We thank our lab (CGERC) members Vhy-Shelta Kewaldar, Pradip Kar, and Jing Gao for their help and assistance with the experiments; Transgenic (TRTC) members Jahnney Torres, Roxana Cubias, William Ramirez, and Eunyoung Kim for zygote microinjection and mouse husbandry; Patrick Smith and David Knorr (LMGI) for collaborative efforts in trying out the Easi-CRISPR system while we were at early stage establishing relevant methods and protocols. We gratefully acknowledge the support and help of Ravi Tolwani, Associate Vice President, Comparative Bioscience Center of the Rockefeller University, for providing resources to make this manuscript possible. We also give our thanks to Betty Shih, Enlightagen Consulting for the editorial contribution and helpful discussions on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Shola, D.T.N., Yang, C., Han, C., Norinsky, R., Peraza, R.D. (2021). Generation of Mouse Model (KI and CKO) via Easi-CRISPR. In: Singh, S.R., Hoffman, R.M., Singh, A. (eds) Mouse Genetics . Methods in Molecular Biology, vol 2224. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1008-4_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1008-4_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1007-7
Online ISBN: 978-1-0716-1008-4
eBook Packages: Springer Protocols