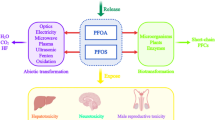
Abstract
Investigations on diverse aspects of fluoro-organic compounds have rapidly increased during the past decades. Because natural sources of fluoro-organic compounds are extremely rare, the industrial synthesis of fluorinated organic compounds and production of fluorinated natural product derivatives have greatly expanded in recent years because of their increasing importance in the agrochemical and pharmaceutical industries. Due to structural complexity or instability, synthetic modification is often not possible, and various biofluorination strategies have been developed in recent years for applications in the anti-cancer, anti-viral and anti-infection fields. Despite the industrial importance of fluorinated compounds, there have been serious concerns worldwide over the levels and synthetic routes of certain fluorinated organic compounds, in particular perfluorinated chemicals (PFCs). PFCs are emerging and recalcitrant pollutants which are widely distributed in the environment and have been detected in humans and wildlife globally. PFCs have been demonstrated to be potentially carcinogenic, adversely affect the neuroendocrine and immune systems, and produce neurotoxicity, heptatotoxicity and endocrine disrupting effects in vertebrate animals. Here, we provide an overview of recent advances in our understanding of the biology of various fluoro-organic compounds and perspectives for new enzymes and metabolic pathways for bioremediation of these chemicals.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Biodegradation
- Defluorogenase
- Environmental toxicity
- Fluorinase
- Human health
- Perfluorinated compounds
- Polyfluorinated compounds
1 Introduction
1.1 Natural Sources of Fluorinated Compounds
Fluorine exists naturally in the Earth’s crust and is the most abundant halogen and the 13th most abundant element. Compared with other halogens, fluorine shows very low levels in surface water and exists mainly in an insoluble form (CaF2) in nature, and thus has very little effects on the environment and biota. More than 4,000 natural products that contain chlorine, bromine, and even iodine have been reported in living organism, whereas only about a dozen fluorinated natural products have been isolated to date [1]. Fluoroacetate was the first natural organofluorinated compound to be identified in 1943 as a metabolite from Dichapetalum cymosum [2]. The low bioavailability of natural fluorinated compounds and fluorine’s very low concentration in surface water may be due to its largely insoluble form (CaF2). The fluoride ion has a high heat of hydration in aqueous solution, which thus limits its participation in displacement reactions. Fluorine cannot be transformed into organic substrates by haloperoxidases (which is a family of peroxidase enzymes that mediate the oxidation of halides by hydrogen peroxide) [3].
1.2 Biofluorination and Fluorinase
Fluorine substitution is widely used in pharmaceutical and agricultural applications because of the effects of fluorine on membrane permeability, metabolic stability, and receptor-binding properties [4, 5]. Because fluorinated products are extremely rare in nature, a number of methods have been developed for synthesis of fluorinated compounds [6, 7]. However, the greatest progress has been in the generation of nonselective fluorinated products, which often cause toxicity and are difficult to handle. Selective incorporation of fluorine is challenging; therefore, development of biologically-based methods for fluorochemical production is needed.
Some enzymatic systems have been reported to utilize fluoride ions. For example, pyruvate kinase is known to catalyze the generation of fluorophosphate from ATP fluoride [8], and more recently, mutant glycosyl transferases were reported to fluorinate 2,4-dinitrophenyl-activated sugars to form α-fluoroglycosides [9, 10]. However, these reactions are adventitious or the intermediates are unstable. In 2002, the first fluorinase was reported in Streptomyces cattleya (O’Hagan et al. 2002), which uses S-adenosyl-l-methionine (SAM) and a fluoride ion as substrates to catalyze the formation of 5-fluoro-5-deoxyadenosine (5-FDA) and l-methionine (l-Met), which is the first step in the biosynthetic pathway of the fluorometabolites, fluoroacetate and 4-fluorothreonine (Fig. 1; Hagan et al. 2002). As the only native fluorination enzyme that has been identified so far, fluorinase was used to explore the syntheses of diverse fluorinated derivatives. For example, an engineered organofluorine biosynthetic metabolite that is a potent anticancer agent, fluorosalinosporamide, was produced by introducing a fluorinase gene (flA) into Salinispora tropica using recombinant DNA technology [12]. This study showed that selective fluorination of drugs and drug candidates could be expanded by inserting the flA gene into a variety of microorganisms to initiate the biosynthesis of novel organo-fluorine compounds.
The fluorinase enzyme of S. cattleya is the first committed step in the biosynthetic pathway to produce fluoroacetate and 4-fluorothreonine [11]
In 2009, a chemo-enzymatic approach for selective fluorination was established whereby fluorine substitutions were used to produce a set of organic molecules including some prodrugs via a two-step regio- or stereo-selective procedure. The initial reaction is catalyzed by cytochrome P450 monooxygenases to insert oxygen selectively into non-reactive C–H bonds with deoxofluorination. The generated hydroxyl group was substituted by a nucleophilic fluorinating reagent, leading to selective fluorine substitution [13].
1.3 (Bio) Synthesis and Pharmaceutical Applications of Fluorinated Compounds
Burgeoning after the 1970s, the industrial synthesis of fluorinated organic compounds expanded because of their applications in pharmaceutical, agricultural, and other industrial areas. In medical applications, fluorine substitution often increases the hydrophobicity, metabolic stability, bioactivity, and bioavailability of molecules, thus improving their therapeutic indices. Medicinal production has focused on fluorinated drugs and drug candidates based on natural product analogs. While fluorinated natural products are very rare, the production of fluorinated natural product derivatives is increasingly common. Due to structural complexity or instability, synthetic modification is often not possible, and alternative strategies have been sought. In the past 20 years, synthetic methodologies in organic fluorine chemistry have focused on the biosynthesis of fluorinated analogs of natural products. Precursor-directed biosynthesis and mutasynthesis are two of the main industrial approaches for biosynthesis of fluorinated natural products. For example, fluorinated diazepinomicin analogs with modest anti-bacterial activity against Staphylococcus aureus have been generated through precursor-directed biosynthesis by supplementing Micromonospora cultures with various indole-related derivatives [14]. Using the mutasynthesis approach, auxotrophic strains of bacteria (which are unable to produce specific amino acids) have been successfully exploited to produce a number of fluorinated natural products [15]. For example, several new calcium-dependent antibiotics were produced by feeding 5-fluorotryptophan to a Streptomyces coelicolor tryptophan-auxotrophic strain [16].
Derivatives of anti-cancer drugs and other compounds such as the anti-inflammatory drugs fluorouracil and fluorocorticoids have been successfully biosynthesized. Other recent efforts have led to the development of fluorinated natural product derivatives, such as fluorine-substituted nucleosides, alkaloids, macrolides, steroids, amino acids, and prostaglandins, for applications in the anti-cancer, anti-viral, and anti-infection fields [15, 17]. Almost 20% of all pharmaceutical drugs on the market contain at least one fluorine atom, including the two best selling compounds, Lipitor (Atorvastatin; Fig. 2a), an inhibitor of cholesterol biosynthesis, and Advair Discus (a mixture of fluticasone (Fig. 2b) and salmeterol), a steroidal anti-inflammatory [18].
Structures of the fluorine containing market leading pharmaceuticals. (a) Lipitor (Atorvastatin, (3R,5R)-7-[2-(4-fluorophenyl)-3-phenyl-4-(phenylcarbamoyl)-5-(propan-2-yl)-1H-pyrrol-1-yl]-3,5-dihydroxyheptanoic acid). (b) Advair Discus (a combination of fluticasone [S-(fluoromethyl) (6 S,8 S,9R,10 S,11 S,13 S,14 S,16R,17R)-6,9-difluoro-11,17-dihydroxy-10, 13,16-trimethyl-3-oxo-6,7,8,11,12,14,15,16-octahydrocyclopenta [a] phenan-threne-17-carbothioate] and salmeterol-2-(hydroxymethyl)-4-{1-hydroxy-2-[6-(4-phenylbutoxy) hexylamino] ethyl} phenol) (O’Hagan 2010).
1.4 Perfluorinated Compounds
In industrial applications, fluorinated compounds, especially perfluorinated compounds perfluorooctanesulfonate (PFOS) and perfluorooctanoic acid (PFOA), play important roles in material science, including fluoropolymers, liquid crystals, and fire extinguishing products, due to their thermal and oxidative stability [19]. The phase-partitioning behavior of perfluoroalkanes makes them a prominent class of surfactants widely used in fire-fighting applications, herbicide and insecticide formulations, cosmetics, greases and lubricants, paints, polishes, and adhesives. In addition, poly/perfluorine derivatives are applied as oxygen carriers in blood substitutes [20]. Although production of many perfluorinated compounds such as PFOA and PFOS has ended in the USA and EU, these compounds are still produced in China and other developing countries.
1.4.1 Environmental Fate and Toxicity
Thousands of tons of fluorinated organic compounds have been emitted into the environment [19]. In recent years, concerns over the levels and synthetic routes of fluorinated organic compounds, especially perfluorinated compounds, have increased. Perfluorinated compounds show thermal, chemical, and biological stability, lipophilicity, worldwide distribution and accumulation in the atmosphere [21], river water [22], wildlife [22, 23], and in humans [24], which may lead to serious problems. The detection of organofluorines in wildlife and humans has been increasingly reported since 1968 [25, 26]. In 2003–2004, >99% of individuals sampled in one study in the US showed detectable PFOA in their serum [27]. In 2009, PFOS was included in Annex B of the Stockholm Convention on Persistent Organic Pollutants.
1.4.2 Fluorinated Compounds and Human Health
While fluorine is regarded as an essential element and is beneficial to human health at low concentrations, the environmental distribution of fluorinated organic compounds is dangerous to humans due to their bioaccumulation and potential impacts on metabolism. During the last two decades, concerns about the toxicity of fluorinated organic compounds, especially perfluorinated compounds, have increased. Most toxicological studies on PFCs have been conducted on rats or monkeys. In animal research, common PFCs such as PFOA and PFOS have been demonstrated to be potentially carcinogenic, to affect the neuroendocrine and immune systems, to cause neurotoxicity and hepatotoxicity, and to reduce serum cholesterol and triglycerides [28–30]. Effects on gestational and developmental toxicity were also confirmed [31]. In vitro studies on human cells also demonstrated the toxicity of PFCs on DNA integrity, intracellular organelles, and hormones ([32]; Vanden Heuvel et al. 2006; [33]). In population studies, some PFCs were reported to act as hormone disruptors and thus to affect human fecundity [34]. Human fetal birth weight was also reported to be impaired by background exposure to PFOA [35]. Additionally, exposure to PFCs causes altered hepatic function, immune function, thyroid function, and cholesterol metabolism, and has carcinogenic potential in humans [36].
2 Biodegradation of Organofluorinated Compounds
Biodegradation is the chemical dissolution of materials by bacteria or through other biological means. Over the years, scientists and engineers have developed a number of bioremediation and biotransformation methods to degrade, transform, or accumulate a huge range of man-made contaminants. A great variety of microbes such as Burkholderia, Rhodococcus, Pseudomonads, Aspergillus, and Beauveria have shown an extraordinary capability to degrade artificial pollutants such as hydrocarbons (e.g., oil), polychlorinated biphenyls (PCBs), polyaromatic hydrocarbons (PAHs), heterocyclic compounds (such as pyridine or quinoline), and pharmaceutical substances [37]. Biodegradation by microorganisms is perhaps one of the most effective methods to remove organic pollutants from the environment and has attracted considerable interest in bioremediation of organofluorinated compounds.
Although fluoroorganic compounds are well known for their inertness and contain the strong C–F bond, some organisms such as bacteria, fungi, algae, and even vertebrates can still biotransform and biodegrade fluoroorganic compounds because of the steric size similarity between fluorine, hydrogen, and hydroxyl groups. To date, little is known about the bacterial metabolism of fluoroorganic compounds, even though several reports have been published on the degradation of monofluorinated compounds. In 1954, the first report on biological defluorination described fluoride elimination of p-fluoroaniline using a horseradish peroxidase. Fluoroaliphatics such as fluoroacetate can be degraded with monofluoroacetate dehalogenase (Pseudomonas indoloxidans, Pseudomonas cepacia, Moraxella sp., Burkholderia sp., etc.) and biodegradation of trifluoroacetic acid has also been reported [38, 39]. Fluoroaromatic compounds can be biodegraded aerobically and anaerobically. However, the biodegradation pathways of perfluorinated chemicals are still not known.
2.1 Fluoroaliphatics
2.1.1 Fluoroacetate
Fluoroacetate is one of the most highly toxic compounds for mammals [40]. The dissociation energy of its C–F bond is among the highest found in natural products [41]. The presence of fluoroacetate in the environment and biota results from its industrial use as a vertebrate pest control agent as well as from metabolites of other compounds such as fluoroacetamide, which is used to control rodents, the anticancer drugs 5-fluorouracil and fluoroethyl nitrosourea, and the industrial chemical fluoroethanol [42].
Microbial defluorination of fluoroacetate was first reported in 1961 [43], followed by reports of the first enzymatic release of fluoride ion from fluoroacetate in both vertebrates and bacteria [44]. A wide variety of microorganisms such as Moraxella, Pseudomonas, and Burkholderia were isolated and shown to be capable of defluorinating fluoroacetate [39, 45]. Fluoroacetate dehalogenases have been characterized in Pseudomonas strains as well as other bacteria for decades (Fig. 3) [46–48]. Microbial degradation of fluoroacetate is now well understood at the mechanistic level. Two possible mechanisms were delineated from the enzyme reaction [49]. The ester intermediate pathway has been examined for fluoroacetate dehalogenases and other enzymes such as rat liver microsomal epoxide hydrolase [45, 50–52]. The carboxylate group of the aspartate residue at the active site acts as a nucleophile and first attacks the α-carbon atom of fluoroacetate to displace the fluorine atom, leading to the release of a fluoride ion. An ester intermediate is formed, which is subsequently hydrolyzed by a water molecule activated by a histidine residue, thereby regenerating the carboxylate group of the aspartate molecule [53].
Hydrolytic defluorination of fluoroacetate [40]
2.1.2 Fluoropyruvate
Fluoropyruvate is often used in the laboratory as an inhibitor to inactivate pyruvate carboxylase, lactate dehydrogenase, and the pyruvate dehydrogenase complex [54]. In recent years, there has been increasing focus on the use of 3-halopyruvate as an anti-cancer agent because it acts as an irreversible inhibitor of metabolic enzyme(s) associated with glycolysis. For example, it has been demonstrated that 3-bromo pyruvate shows high in vivo toxicity on tumors but has no adverse effect on healthy tissue [55]. In 1978, a pyruvate dehydrogenase component of Escherichia coli that catalyzes the conversion of 3-fluoropyruvate to acetate and fluoride ions was reported [56]. Fluoride is eliminated by β-elimination, which is the classical mechanism for dehydrogenases (Fig. 4). Recently, 19F NMR spectroscopy studies demonstrated the conversion of fluoropyruvate to fluoroacetate by D. cymosum, where fluoroacetate is mineralized followed by the release of fluoride [57].
Proposed enzymatic defluorination of 3-fluoropyruvate [38]
2.1.3 Maleylacetate
Fluorinated maleylacetates have been investigated as substrates of maleylacetate reductase for a number of years [58–61]. A maleylacetate reductase enzyme was first isolated in 1995 from Pseudomonas sp. strain B13 that catalyzes the halo-elimination of 2-fluoromaleylacetate as well as other halomaleylacetates (Fig. 5). This enzyme consumes two moles of NADH per mole of maleylacetate that contains a fluorine substituent in the 2-position, while only one mole of NADH is required for halide elimination in substrates without a fluorine substituent in the 2-position [58].
Proposed mechanism for the elimination of halogen substituents from the 2-position of maleylacetate [58]
2.1.4 Fluorinated Cycloalkyl N-Phenylcarbamates
Fluorine substitution of a hydrocarbon position in fluorinated cycloalkyl N-phenylcarbamates occurs in hydroxylation reactions by Beauveria bassiana, a soil-borne filamentous fungus. The hydroxylation of 4-cis-fluorocycloalkyl N-phenylcarbamates probably produces terminal fluorohydrins, which are not stable and thus are subsequently dehydrofluorinated to give the corresponding ketones [62]. Recently, the defluorination of trans-2-fluorocycloalkyl N-phenylcarbamate by B. bassiana was also reported, in which fluorine elimination could occur either via hydroxylation of the six member ring at C-4 or p-hydroxylation of the aromatic ring (Fig. 6).
Defluorination of trans-2-fluorocycloalkyl N-phenylcarbamate by Beauveria bassiana [63]
2.1.5 Fluorinated Carbohydrates
Fluorinated carbohydrates have a broad range of pharmaceutical and biomedical applications ranging from metabolic and biochemical studies to disease diagnoses. Replacement of a hydroxyl group with a fluorine atom in carbohydrates can affect their metabolic and biochemical behavior, including enzyme-carbohydrate interactions, lectin-carbohydrate affinities, antibody-carbohydrate binding [64, 65], and application in positron emission tomography for cancer diagnosis [66]. Therefore, fluorinated compounds are important reagents in metabolic studies and for disease diagnoses. The microbial catalytic defluorination of fluoromonosaccharides has been reported [67, 68]. Expression of a 65.5 kDa membrane protein is induced by 4-deoxy-4-fluoro-d-glucose (4-FG) or glucose and is associated with the active d-glucose transporter system in Pseudomonas putida [68]. P. putida defluorination of fluoro-d-glucose is stereospecific. In addition, 4-FG is converted to 2,3-dideoxy-d-glycero-pentonic acid with fluoride elimination while 3-deoxy-3-fluoro-d-glucose (3-FG) is metabolized without defluorination. Electron donors such as l-malate are required in these defluorination metabolic pathways [69].
2.2 Fluoroaromatics
Fluoroaromatics are widely used in industry as intermediates or end-products in the synthesis of pharmaceuticals, insecticides, plastics, and molecules related to liquid crystal technology [15, 70]. The broad applications of fluoroaromatics have led to their accumulation in the environment. Their widespread occurrence and potential toxicity have led to increasing interest in biodegradation and treatment processes for fluoroaromatics.
2.2.1 Fluorobenzoates
As model compounds of other fluoro-substituted aromatic compounds, fluorobenzoates have been widely used to study bacterial metabolism of fluorinated aromatics. For example, bacteria such as Pseudomonas [71, 72], Xanthobacter [73], and Sphingomonas [74] have been reported to exhibit fluorobenzoate degradation. In addition, the metabolism of 2-, 3-, and 4-fluorobenzoic acid has been well studied [71, 74, 75]. Using 18O2, Pseudomonas sp. was shown to form catechol from 2-fluorobenzoic acid by incorporation of two oxygen atoms from a single dioxygen molecule. This defluorination proceeds through a cyclic peroxide intermediate. In the major pathway, 1,2-dioxygenation of 2-fluorobenzoic acid leads to an unstable fluorohydrin, which is then defluorinated to catechol. Muconate is finally formed, which subsequently goes in the TCA cycle to produce energy (Fig. 7, pathway a). The minor pathway, 1,6-dioxygenation, also takes place, leading to the formation of 3-fluorocatechol and then 2-fluoro-cis-cis-muconate (Fig. 7, pathway b) [75]. 3-Fluorobenzoate is degraded by 1,2-dioxygenation to yield fluorocatechol, which is metabolized to 2-fluorobenzoic acid in the minor pathway (Fig. 7, pathway c) [74, 75]. The predominant pathway of 3-fluorobenzoate includes a 1,6-dioxygenation reaction to yield fluoromuconic acids. Defluorination then occurs to yield muconate [74] (Fig. 7, pathway d). 4-Fluorobenzoate is degraded by Pseudomonas sp. in similar pathways to 3-fluorobenzoate (Fig. 7, pathway e) [75, 76].
Metabolism of 2-, 3-, and 4-fluorobenzoic acid [38]
The anaerobic degradation of monofluorobenzoates under various electron-accepting conditions including denitrifying, sulfate-reducing, iron-reducing, and methanogenic conditions has also been studied [77–80]. After long-term incubation, 2-fluoro- and 4-fluorobenzoates are degraded by Pseudomonas with fluoride elimination [79]. Recently, dehalogenated 3-fluorobenzoate was investigated in Syntrophus aciditrophicus culture. Two hydrogen atoms are added to 3-fluorobenzoate to form a 3-fluorocyclohexadiene metabolite, leading to stoichiometric accumulation of benzoate and fluorine [80].
2.2.2 Fluorophenols
Fluorophenolic compounds are widely used in agricultural industries as herbicides, insecticides, and fungicides [81]. Fluorophenols are transferred to fluorocatechols and fluoromuconates via microbial degradation [82]. The fluorophenol metabolites of Exophiala jeanselmei, a yeast-like fungus, which are converted by the phenol hydroxylase and catechol 1,2-dioxygenase enzymes, have been characterized by 19F NMR spectroscopy. The conversion of fluorophenols (i.e., 3-fluoro-, 4-fluoro-, and 3,4-difluorophenol) via catechol 1,2-dioxygenase involves two common steps [81, 83]: (1) the introduction of ortho-hydroxyl groups and (2) ring cleavage by catechol dioxygenase. The resulting muconates and accumulation of stoichiometric amounts of fluoride anions have been detected (Fig. 8).
Pathways for defluorination of fluorinated phenols by P. benzenivorans. Dashed arrows show the known monofluorophenols pathway for comparison. 1: phenol hydroxylase, 2: catechol 1,2-dioxygenase [83]
2.2.3 Fluorotoluene
3-Fluorotoluene was reported to be accumulated and co-metabolized by Cladosporium sphaerospermum, a fungi culture grown on toluene [84]. 19F NMR was used to determine the catabolic pathway. A methyl group is first oxidized by the toluene monooxygenase enzyme followed by ring hydroxylation to form fluoroprotocatechuate. The remaining steps include decarboxylation of the fluoroprotocatechuate followed by ortho-cleavage (Fig. 9).
Proposed fungal catabolism of fluorotoluene [38]
2.2.4 Fluorobiphenyls
Fluorobiphenyls can be co-metabolized via the classical aromatic degradation pathways by fungi and bacteria [85–87]. Recently, the degradation pathway of 4,4-difluorobiphenyl was proposed. The hydrolase BphD catalyzes the transformation from 3-fluoro-2-hydroxy-6-oxo-6-(4-fluorophenyl)-hexa-2,4-dienoate to 3-fluoro-2-hydroxypenta-2,4-dienoate. Then, (Z)-3-fluoro-2-oxo-pent-3-enoate is formed and further catabolized, eventually yielding acetaldehyde and fluoropyruvate (Fig. 10) [88].
Proposed catabolism of 4,4-difluorobiphenyl along the upper and lower pathways. BphA biphenyl 2,3-dioxygenase; BphB dehydrogenase; BphC 2,3-dihydroxybiphenyl 1,2-dioxygenase; BphD 2-hydroxy-6-oxo-6-phenylhexa-2,4-dienoate hydrolase; BphX1 2-hydroxypenta-2,4-dienoate hydratase; BphX3 4-hydroxy-2-oxovalerate hydrolase [88]
2.2.5 Fluorophenylacetic Acid
The defluorination of p-fluorophenylacetic acid by Pseudomonas has been studied [76]. First, the aromatic ring is cleaved between C-1 and C-2. Then, C-2 is further modified by two alternative pathways. Hydrolyzation occurs to give 3-hydroxy-3-fluoroadipic acid. Fluorine elimination occurs and yields β-ketoadipic acid (Fig. 11, pathway a). Alternatively, after lactonization and formation of 4-carboxymethyl-3-fluoro-butanolide, hydrolyzation and cleavage of C–C bonds yield acetate and monofluorosuccinic acid (Fig. 11, pathway b). The latter compound is converted to oxaloacetate and hydrogen fluoride.
Degradative pathways for 3-fluoro-3-hexenedioic acid [76]
2.2.6 Fluorobenzene
A microbial consortium containing Sphingobacterium, Flavobacterium, and β-Proteobacterium was shown by Carvalho et al. in 2002 [89] to be capable of defluorinating fluorobenzene. In addition, a bacterial strain from the Labrys portucalensis group that uses fluorobenzene as a sole carbon and energy source has been purified [90]. The degradation of fluorobenzene via ortho cleavage of 4-fluorocatechol and catechol by Rhizobiales strain F11 has been investigated by Carvalho et al. in 2006 [91]. It was found that the initial attack on fluorobenzene by a dioxygenase enzyme could lead to two different pathways. In one pathway, a dihydrodiol dehydrogenase enzyme (step 1) transforms 4-fluoro- cis-benzene-1,2-dihydrodiol to 4-fluorocatechol. In the second pathway (step 10), 1-fluoro-cis-benzene-1,2-dihydrodiol is converted to catechol (Fig. 12).
Proposed pathway for fluorobenzene metabolism. The enzyme activities are denoted as follows: 1: fluorobenzene dioxygenase; 2: fluorobenzene dihydrodiol dehydrogenase; 3: fluoro-catechol 1,2-dioxygenase; 4: fluoromuconate cycloisomerase; 5 and 6: possible side reactions to cis-dienelactone by fluoromuconate cycloisomerase (activity 5) or by slow spontaneous conversion (activity 6); 7: trans-dienelactone hydrolase; 8: maleylacetate reductase; 9: fluorobenzene dioxygenase; 10: nonenzymatic defluorination; 11: catechol-1,2-dioxygenase; 12: muconate cycloisomerase; 13: muconolactone isomerase; 14: 3-oxoadipate enol-lactone hydrolase [91]
2.2.7 Fluoroquinolones
Fluoroquinolones are some of the most widely used antimicrobial agents for treating both Gram-negative and Gram-positive infections. Their widespread presence has been detected at multiple locations around the world [92]. Other reports have suggested their potential toxicity to plants and aquatic organisms [93, 94]. Many clinically relevant bacterial species including S. aureus and Pseudomonas aeruginosa are capable of developing resistance to quinolones [95]. Degradation of the fluoroquinolone, enrofloxacin, was observed in Gloeophyllum striatum, a brown rot fungus where a hydroxyl radical attacks fluorine at the C-6 position to form 6-hydroxyen-rofloxacin which is further hydroxylated to 5,6- and 6,8-dihydrox-yenrofloxacin [96]. The metabolism of enrofloxacin by seven basidiomycetous fungi from agricultural sites was recently reported by Wetzstein et al. [97]. Oxidative decarboxylation of enrofloxacin first occurs, then defluorination takes place in multiply hydroxylation and acetylation steps (Fig 13) [97].
Degradation of enrofloxacin by basidiomycetous fungi [97]
2.2.8 Fluorinated Anilines
Microsomal NADPH-dependent reaction pathways for biodehalogenation of fluorinated anilines have been investigated [98]. Three possible pathways for dehalogenation of fluorinated anilines, such as 2-fluoro-4-hydroxyaniline and pentafluoroaniline, in the presence of xanthine glutathione and NADPH were proposed. A study of the metabolism of 3,4-difluoroaniline with Pseudomonas fluorescens 26-K showed the formation of 3-fluoro-4-hydroxyaniline and the release of a fluoride ion [99]. Recently, biotransformation of 4-fluoroaniline was observed in the earthworm Eisenia veneta. The catabolic products were analyzed using 19-F NMR, but no fluoride ion was detected (Fig. 14) [100, 101].
Metabolism of 4-fluoroaniline in Eisenia veneta [100]
2.3 Biodegradation of Polyfluorinated Compounds
The degradation of polyfluorinated compounds, such as fluorotelomer alcohols (FTOHs), fluorotelomer ethoxylates, and polyfluoroalkyl phosphates, in atmospheric and aqueous systems has been established and has been reported to be a source of perfluorinated carboxylic acids (PFCAs). However, published information on the biodegradation of PFCAs is very limited. The aerobic and anaerobic biodegradability of three fluorinated surfactants have been described [102]. However, no release of fluoride has been found.
2.3.1 Fluorotelomer Alcohols
FTOH is the generic name of fluorinated compounds that contain even-numbered fluorocarbon chains and an ethanol moiety [103]. FTOHs are used in fire-fighting foams, grease-resistant food packaging, leather protectants, and stain-resistant carpeting and textiles. In addition, FTOHs are used industrially to generate acrylate polymers and as intermediates in the production of fluorinated surfactants. Consequently, FTOHs are widely detected in air. Furthermore, estrogen-like properties have been reported for these compounds [104].
8–2 FTOH degradation was first reported in detail in reactions catalyzed by a mixed microbial consortium [105–109]. Based on 14C analysis, 8–2 FTOH biodegradation in aerobic soils was proposed (Fig. 15). 8–2 FTOH is converted rapidly to 8–2 fluorotelomer aldehyde (FTAL) by an alcohol dehydrogenase and to 8–2 fluorotelomer acid (8–2 FTA) by an aldehyde dehydrogenase. The conversion of 8–2 FTA to 8–2 fluorotelomer unsaturated acid (8–2 FTUA) in soils is so rapid that no 8–2 FTA above the limit of quantification was observed.
Proposed biodegradation pathways of 8–2 FTOH [109]
Recently, the first study to investigate aerobic biodegradation of 6–2 FTOH [F(CF2)6CH2CH2OH] was described by Liu et al. [110]. Based on this investigation and previous studies on the mechanism of 8–2 FTOH biodegradation [107–109, 111, 112], several pathways for 6–2 FTOH degradation have been proposed. 6–2 FTOH is first converted to 6–2 FTAL through oxidation by alcohol dehydrogenase or cytochrome P450, and then to 6–2 FTA by aldehyde dehydrogenase. Using the 2,4-dinitrophenylhydrazine (DNPH) derivatization method previously described for the detection of 8–2 FTAL from 8–2 FTOH degradation in soil and mammals [108, 109], 6–2 FTAL was not detected in the soil extracts. Hydrogen fluoride (HF) is removed from 6–2 FTA to form 6–2 FTUA either because α-oxidation is not operable or because rapid HF elimination to 6–2 FTUA supersedes 6–2 FTA α-hydroxylation, which is necessary for α-oxidation (Martin et al. 2005). 6–2 FTUA degradation proceeds by two pathways (Fig. 16).
Proposed 6–2 FTOH aerobic biodegradation pathways. The double arrows indicate multiple transformation steps [110]
Biodegradation of a novel fluorotelomer alcohol, 1H,1H,2H,2H,8H,8H-perfluorododecanol (degradable telomer fluoroalcohol, DTFA), was investigated in a mixed bacterial culture obtained from activated sludge and the pathway was also proposed (Fig. 17) [103], First, through the catalytic activity of alcohol dehydrogenase and aldehyde dehydrogenase, DTFA is oxidized to 2H,2H,8H,8H-perfluorododecanoic acid (2H,2H,8H,8H-PFDoA) which is then defluorinated to 2H,8H,8H-2-perfluorododecenoic acid (2H,8H,8H-2-PFUDoA). Double bonds are formed between the internal –CH2– and –CF2– groups in 2H,8H,8H-2-PFUDoA which is then further degraded via two different β-oxidation pathways. In pathway a, through the removal of –CF2–, 2H,8H-2,8-PFUDoA is transformed into three different long-chain carboxylic acids which are further degraded into perfluorobutanoic acid (PFBA) with dicarboxylic acids containing different fluorocarbon lengths (C4–C6 compounds), whereas in pathway b, 2H,8H-2,8-PFUDoA is transformed into perfluoropentanoic acid (PFPeA) with three different fluorinated dicarboxylic acids (Fig. 17).
Proposed biodegradation pathway for DTFA. The double arrows indicate multiple transformation steps [103]
2.3.2 Fluorotelomer Ethoxylates
Fluorotelomer ethoxylates [F–(CF2–CF2–)x–(CH2–CH2–O)y–H] are an important class of non-ionic fluorinated surfactants and are regarded as a potential source of per- and polyfluorinated organic pollutants. Aerobic biotransformation of FTEOs was recently demonstrated by Frömel and Knepper [113]. Distinct from the biodegradation of FTOHs, ω-oxidation occurs and is responsible for the transformation of FTEOs to FTEO carboxylates (FTEOCs). After oxidation of the terminal hydroxyl group to a carboxylic acid, the carbon chain is subsequently shortened whereby the short-chained FTEOCs are not further degraded. In this chapter, no PFCA formation attributable to FTEO degradation was observed.
2.3.3 Fluorotelomer-Based Urethanes
Fluorotelomer-based urethanes are urethane polymers consisting of a series of fluorotelomer side-chains attached to a hydrocarbon backbone and are commercially used as stains and soil repellents for textiles. Russell et al. [114] tested the potential for microbial activities in four different soil samples to degrade a fluorotelomer-based urethane polymer under aerobic conditions over a 24-month period and demonstrated that fluorotelomer side-chains were released and transformed to perfluorocarboxylic acids including PFOA.
2.3.4 Polyfluoroalkyl Phosphates
Polyfluoroalkyl phosphates (PAPs) are used as commercial surfactants for oil repelling applications, and have been shown to be degraded to PFCAs in a rat model and waste water treatment plant system [115]. The pathway in Fig. 18 describes the aerobic degradation routes of 4:2, 6:2, 8:2, and 10:2 monosubstituted PAPs (monoPAPs) and 6:2 disubstituted PAP (diPAP) by a microbial mixture collected from sewage of a wastewater treatment plant. In the microbial system, 6:2 FTOH was initially oxidized into a series of acid metabolites. The intermediate metabolite, 6:2 saturated fluorotelomer carboxylic acid (6:2 FTCA), is converted via β-oxidation to 6:2 unsaturated fluorotelomer carboxylic acid (6:2 FTUCA) and perfluorohexanoic acid (PFHxA) (Fig. 18, pathway b), while 5:3 FTCA is transformed to perfluoropentanoic acid (PFPeA) (Fig. 18, pathway c). However, the production of PFPeA may also be attributed to other precursors. For example, 6:2 FTUCA may degrade into 5:2 fluorotelomer ketone (F(CF2)5CH(OH)CH3) which is further reduced to 5:2 secondary fluorotelomer alcohol (sFTOH, F(CF2)5CH(OH)CH3), and subsequently transformed to PFPeA (Fig. 18, pathway d). 6:2 FTCA and PFHpA production have been observed, supporting the possibility of oxidation of the α-carbon in FTCA to form odd-chain PFCAs (Fig. 18, pathway a).
Proposed degradation pathway of 6:2 diPAP and 6:2 monoPAP [115]
2.3.5 ω-(Bis(trifluoromethyl)amino)alkane-1-Sulfonates
Biodegradation of ω-(bis(trifluoromethyl)amino)alkane-1-sulfonates was detected in a fixed-bed bioreactor. Its incomplete mineralization revealed that degradation mostly takes place via desulfonation, oxidation, and further β-oxidation [116]. The C–F and C–N bonds in the bis(tri-fluoromethyl)amino (BTFMA) group cannot be accessed by microbes for biodegradation; therefore no defluorination was observed (Fig. 19).
Biotransformation pathways of ω-(bis(trifluoromethyl)amino)alkane-1-sulfonates [116]
2.3.6 N-Ethyl Perfluorooctane Sulfonamide Ethanol
N-Ethyl perfluorooctane sulfonamide ethanol (N-EtFOSE) is present in protective paper coatings. Although the only producer in the USA, 3M, has stopped production since 2002, N-EtFOSE can still be detected in the North American atmosphere [117]. Aerobic biotransformation of N-EtFOSE in activated sludge has been studied [118]. Fast oxidation of N-EtFOSE forms N-ethyl perfluorooctane sulfonamido acetic acid (N-EtFOSAA) through an aldehyde intermediate. N-Ethyl perfluorooctane sulfonamide (N-EtFOSA) undergoes direct dealkylation to perfluorooctane sulfonamide (FOSA), while perfluorooctane sulfonamido acetic acid (FOSAA) production proceeds at a slower rate. The extremely stable compound PFOS was observed as the final product (Fig. 20) [118].
Proposed transformation pathway of N-EtFOSE in activated sludge [118]
2.3.7 10-(Trifluoromethoxy) Decane-1-Sulfonate
10-(Trifluoromethoxy) decane-1-sulfonate is a fluorinated surfactant that has been globally distributed, thus leading to increasing concern on its environmental fate and toxicity. Biomineralization of 10-(trifluoromethoxy) decane-1-sulfonate was reported by Peschka et al. in 2008. Two proposed pathways, major and minor, have been described (Fig. 21). In the major degradation pathway, the carbon chain of the fluorinated alkylsulfonate derivative is shortened by β-oxidation after desulfonation and oxidation. The formed trifluoromethanol is unstable and mineralizes immediately (Fig. 21, pathway a). In the minor degradation pathway, insertion of oxygen occurs, and then, the molecule is subsequently cleaved and degraded (Fig. 21, pathway b).
Degradation pathways of 10-(trifluoromethoxy)decane-1-sulfonate [119]
2.4 Perspectives for the Biodegradation of Perfluorinated Compounds
As previously mentioned, biodegradation and biotransformation of several polyfluorinated compounds such as FTOHs, FTEOs, and PAPs have been reported. But to date there are still no reliable reports on the biodegradation or biotransformation of perfluoroalkyl compounds such as PFOS and PFOA. To date, the studies examining biodegradation and transformation of PFCs is very limited. PFCAs, including PFOA, are common transformation products from fluorotelomer chemicals [105, 106, 109, 120, 121]. No evidence about the biodegradation and biotransformation of PFCAs has been found. A recent study about the biodegradability of PFOA using five different microbial communities incubated for up to 259 days showed that PFOA is still microbiologically inert and thus is environmentally persistent [122]. Because of the high stability of the strong C–F covalent bonds, the rigidity of the perfluoroalkyl chain, and the lack of reactive substituent, PFOS is highly recalcitrant to biodegradation or chemical degradation under ambient conditions. Only two reports about the chemical degradation of PFOS have been published [123, 124]. No studies on biodegradation or biotransformation have been reported. Recently, the first report of reductive dehalogenation of PFOS catalyzed by vitamin B12 was published, in which PFOA was reduced and dehalogenated by Ti(III)-citrate [125]. These results suggested the potential for reductive dehalogenation of PFCs.
2.4.1 Thermodynamics of Organofluorine Biodegradation
Thermodynamics can be used to evaluate whether organisms can obtain energy for growth by catalyzing certain reactions. This approach has been applied to the study of reductive biodechlorination of chlorinated compounds such as 3-chlorobenzoate and chloromethanes. The amount of energy available from reductive dechlorination was reported to be between 100 and 180 kJ/mol [126], which is enough to support microorganism growth by using halogenated compounds as electron acceptors. The Gibbs free energy values of fluorinated compounds showed that the amount of energy obtained from defluorination is similar to the amount available from dechlorination and could support microorganism growth. Therefore, organisms may be able to obtain energy by catalyzing certain defluorination reactions for growth.
Although the thermodynamic properties of perfluoroalkylated compounds are not available, the Gibbs free energy values for the reductive removal of one fluorine atom from fluoropropane molecules (Table 1) showed that the energy yields from the hydrogenolysis of perfluorinated compounds and from less fluorinated compounds are similar. These results reveal the thermodynamic basis for reductive biodefluorination of perfluoroalkylated compounds, especially under anaerobic conditions.
2.4.2 Perspectives for the Biodegradation of Perfluorinated Compounds
Thermodynamically, perfluorinated compounds should be potentially biodegradable, especially under anaerobic conditions [127]. To date, the reductive biodefluorination of perfluorinated compounds has not yet been observed. Moreover, the rate at which microorganisms can evolve the capability to grow on this potential source of energy and the function of the enzymatic machinery that catalyzes this reaction are largely unknown. However, co-metabolic degradation of several polyfluorinated compounds under aerobic conditions and without thermodynamic facilitation has been studied in detail [109, 110, 115]. This information formed the basis for technology that has been applied in the field for the degradation of other polyhalogenated compounds such as trichloroethene. The search for co-metabolic degradation of poly- and perfluorinated compounds, and studies to understand its mechanisms better will continue.
3 Defluorination Pathways and Defluorogenases
3.1 Enzymatic Metabolic Pathways
3.1.1 Aerobic Metabolism
Limited reports are available about the biodegradation of fluorinated organic compounds, and therefore little is known about the enzyme-catalyzed defluorination pathway. Under aerobic conditions, fluorinated organic compounds are usually degraded via the electron donor or co-metabolic pathways. It has been reported that 4-fluorophenol can be utilized as the sole source of carbon and energy for Arthrobacter sp. strain IF1, and that two gene clusters are involved [128]. Cluster A harbors fpdA1DE and includes an FADH2-dependent monooxygenase, a putative maleylacetate reductase, and a hydrolase gene. In Cluster B, fpdA2 encodes a 4-FP monooxygenase, fpdB encodes a flavin reductase, and fpdC encodes a putative hydroxyquinol dioxygenase (Fig. 22). The proposed catabolic pathway is shown in Fig. 23.
Organization of the open reading frames (ORFs) in the fpd gene regions of Arthrobacter sp. strain IF1. Open arrows indicate the size and direction of each ORF [128]
Proposed catabolic pathway for 4-FP degradation [128]
To date, the well-known enzymes involved in fluoro-degradation are normally responsible for the catabolism of non-fluorinated compounds. Evidence suggests that enzymes are specifically employed for the catabolism of these substrates. Enzymes for degrading aromatic compounds such as monooxygenases, cleavage dioxygenases, and maleylacetate reductase have exhibited biodefluorination activity. As shown in Fig. 24, a number of enzymes that do not show specific activity for fluoroaromatic compounds are involved in the catabolism of 3-fluorobenzoic acid [86].
Catabolism of 3-fluorobenzoic acid in aerobic bacteria [40]
Because of the similar steric sizes of hydrogen and fluorine, substitution of hydrogen for fluorine is considered to have an important role in defluorination. Many oxygenases and (de)hydroxylases make up a group of defluorinating enzymes that act on both fluorinated aromatics and fluoroaliphatics. In 1978, a pyruvate dehydrogenase component of E. coli was found to catalyze the conversion of 3-fluoropyruvate to acetate and fluoride ions and to eliminate the fluorine [56]. Later, the proposed mechanism of fluorine elimination by dehydrogenases was proposed. p-Hydroxybenzoate hydroxylase, a NADPH-dependent flavin-containing monooxygenase from P. fluorescens and Candida parapsilosis, was reported to degrade several fluorine-substituted p-hydroxybenzoates such as fluorohydroxybenzoate [129, 130]. Fluorobiphenyl metabolism is catalyzed by a series of dioxygenases dehydrogenases and hydrolases to yield fluoropyruvate and 4-fluorobenzoate [86, 87].
3.1.2 Anaerobic Metabolism
Less is known about the degradation of fluoroaromatic compounds under anaerobic conditions. Defluorination of 2-hydroxybenzoate and 3-fluorobenzoate was observed in S. aciditrophicus cultures under anaerobic conditions (Mouttaki et al. 2009). Recently, co-metabolism in a bacterial culture was found to catalyze 4-fluorobiphenyl to a carboxylic acid derivative [131]. Both methanogenic and sulfate-reducing defluorination generated trifluoroacetic acid (TFA) via a co-metabolism pathway [132, 133]. Denitrifying bacteria have also been reported to mineralize 2- and 4-fluorobenzoate [79].
3.2 Defluorinases
Among all the dehalogenations, defluorination is most difficult because the C–F bond is one of the most stable bonds in nature. Partly because of limited studies on defluorinases, most man-made organofluorine compounds are degraded via co-metabolism pathways. Enzymes with alterable substrates play an important role, although few fluorine-specific enzymes have been identified.
3.2.1 Fluoroacetate Dehalogenase
As the most common fluorinated natural product, fluoroacetate was reported in 1965 to be degraded by Pseudomonas fluoroacetate dehalogenase, which catalyzes the hydrolytic cleavage of the C–F bond to yield glycolate and a fluoride ion [46]. Other fluoroacetate dehalogenases have been isolated from microorganisms such as Moraxella, Delftia, and Burkholderia [45, 48, 134]. Fluoroacetate dehalogenase belongs to the α/β hydrolase superfamily protein. The mechanism of C–F bond cleavage by fluoroacetate dehalogenase has been extensively investigated (Keuning et al. 1985; [45]). The three-dimensional structure of FAc-DEX FA1, a fluoroacetate dehalogenase from Burkholderia sp. strain FA1 [53], suggested a mechanism whereby fluoroacetate is degraded by an initial nucleophilic attack on the α-carbon atom by the carboxylate group of Asp104 which displaces the fluoride ion to form an ester intermediate. The ester intermediate is then hydrolyzed by a His271-activated water molecule, which yields glycolate and regenerates the carboxylate group of Asp104 (Fig. 25).
Proposed reaction mechanism of FAc-DEX FA1 [53]
3.2.2 Fluoroacetate-Specific Defluorinases
Detoxification of fluoroacetate in mammals is catalyzed by fluoroacetate-specific defluorinases (FSDs) such as the glutathione-S-transferase isozyme GSTZ which is distinct from bacterial fluoroacetate dehalogenases (Fig 26) [135]. Two distinct FSD activities have been identified in rat liver: one is glutathione-S-transferase-like and the other more predominant activity is apparently a new type of dehalogenase, which is considered to be an FSD. Interestingly, the amino acid sequence of the latter FSD is similar to the sorbitol dehydrogenase protein, which does not show defluorination activity on fluoroacetate.
Enzymatic C–F bond cleavage. (a) Fluoroacetate dehalogenase. (b) Fluoroacetate-specific defluorinase [40]
3.2.3 4-Fluorobenzoate Dehalogenase
4-Fluorophenol (4-FP) monooxygenase (FpdA2) was first cloned and purified from Arthrobacter sp. strain IF1. In combination with FpdB, which uses NADH to reduce either flavine-adenine dinucleotide (FAD) or flavin Mononucleotide (FMN), FpdA2 transforms various halogenated phenols via para-substitution, leading to halide release and hydroquinone formation (Fig. 27) [128].
4-FP monooxygenase [128]
3.3 Perspectives for New Enzymes and Metabolic Pathways
As mentioned above, co-metabolism is the main degradation pathway for mono- and polyfluorinated organic compounds. However, due to very limited research, only a few co-metabolism pathways have been found in the laboratory. Further studies in this area will help to investigate the pathways and enzymes involved in defluorination. Compared with a series of dechlorinases that catalyze various kinds of reactions, only three kinds of defluorinases have been identified to date. In 2007, biodegradation of 4-fluoroglutamate was reported via an unusual pathway, yielding equimolar concentrations of fluoride ions and ammonia, indicating that an enzyme such as glutamate dehydrogenase is not responsible for the biotransformation (Fig. 28) [136]. In addition, the defluorinating/deaminating activity was found in the soluble fraction of the cell and was not related to the dechlorinating/deaminating activity, which was located in the cell membrane. These results suggest the existence of a potential new fluoroglutamate dehalogenase. Under anaerobic conditions, defluorination was detected in methanogenic, sulfate-reducing, and denitrifying bacteria, indicating that extensive defluorination occurs under anaerobic conditions [79, 132]. Reductive defluorination is a thermodynamically feasible mechanism to derive cellular energy under anaerobic conditions. However, microbes that are able to obtain energy for growth by reductive defluorination have yet to be isolated [127]. And there is much be done to elucidate the defluorination mechanisms and properties of the enzymes involved.
4-Fluoroglutamate dehalogenase/deaminase. Arrows indicate that more than one reaction might be occurring. GSH=glutathione [40]
4 Summary and Perspectives
Brominated and chlorinated compounds have been investigated in previous research on the biodegradation of halogenated compounds. However, fluorinated chemicals have thus far received much less attention [127]. The inertness of fluorine results in persistence and leads to accumulation in the environment, making it necessary to explore microbial degradation of fluoroorganic compounds. Until recently, only a few microbes including bacteria, fungi, and algae have been found to be capable of fluoro-degradation. For most fluorinated substrates, the mechanism of fluoro-degradation is still not clear. Several monofluorinated compounds, including fluoroaliphatics [57, 136, 137], fluoroaromatics [71, 74, 81, 88], and a few other polyfluorinated compounds [105, 106, 109, 110], can be degraded. However, the mechanisms of these degradation reactions are largely unknown. No biodegradation of perfluorinated compounds has been observed [25, 122]. Perfluorinated and polyfluorinated compounds are widely used as surfactants, catalysts, and insecticides [18, 19]. These compounds are highly recalcitrant and have been detected throughout the global environment [26, 27]. Biodegradation of perfluorinated compounds is thermodynamically possible under reductive conditions, but has not been measured [127]. Despite a great increase in knowledge over the last few decades, we are still far from being able to predict the biodegradation of fluorinated organic compounds as well as the mechanism of defluorination. Although the dehalogenation of both fluorinated and chlorinated organic compounds is largely mediated by soil microflora, limited knowledge of the factors influencing these microorganisms is available. Development of systematic biological and molecular genetics studies will help in the study of soil microbial species and communities, thus facilitating the discovery of new microbes capable of defluorination.
New technologies for chemical analysis have made highly sophisticated studies practical in the laboratory. Fluorine-19 nuclear magnetic resonance spectroscopy (19F NMR) and isotopic labeling techniques have helped to contribute to a deeper understanding of several key processes in the catalyzed reactions of fluorinated substances [74, 109, 110]. The rapid growth of bioinformatics has led to the development of databases that search for organic persistence information. Furthermore, scientists have created computer programs such as MultiCASE based on general quantitative structure-degradation relationships (QSDRs) to predict the degradation/persistence of organic chemicals in the environment that have not been characterized [138]. One of these popular computer programs, BIOWIN, contains a series of models collectively referred to as biodegradability probability. Based on QSDR models as well as six aerobic biodegradation models and one anaerobic model, BIOWIN can predict the biodegradability probability under aerobic and anaerobic conditions. If a metabolic pathway is available for a chemical, it is assumed to be biodegradable [139]. Similar programs, including the UM-BBD Pathway Prediction and System CATABOL program, have also been developed for determining the biodegradability probability [140, 141]. These tools provide unique approaches to studying biodifluorination.
In general, biodegradation studies need interdisciplinary collaborations between microbiology, ecology, genetics, biochemistry, and analytical chemistry to resolve complex problems. As more research attention is given to this field and more technologies are developed and applied, further mechanisms of the biodegradation of fluorine-containing organic compounds will be elucidated.
References
Gribble GW (2004) Natural organohalogens: a new frontier for medical agents? J Chem Educ 81:1441–1449
Deng H, O’Hagan D, Schaffrath C (2004) Fluorometabolite biosynthesis and the fluorinase from Streptomyces cattleya. Nat Prod Rep 21:773–784
Murphy CD, Schaffrath C, O’Hagan D (2003) Fluorinated natural products: the biosynthesis of fluoroacetate and 4-fluorothreonine in Streptomyces cattleya. Chemosphere 52:455–461
Park BK, Kitteringham NR, O’Neill PM (2001) Metabolism of fluorine-containing drugs. Annu Rev Pharmacol Toxicol 41:443–470
Muller K, Faeh C, Diederich F et al (2007) Fluorine in pharmaceuticals: looking beyond intuition. Science 317:1881–1886
Kirsch P (2004) Modern fluoroorganic chemistry: synthesis, reactivity, applications. Wiley-VCH, Weinheim
Shimizu M, Hiyama T (2005) Modern synthetic methods for fluorine-substituted target molecules. Angew Chem Int Ed Engl 44:214–231
Flavin M, Castro-Mendoza H, Ochoa S (1957) Metabolism of propionic acid in animal tissues. J Biol Chem 229:981–996
Nashiru O, Zechel DL, Stoll D et al (2001) β–Mannosynthase:synthesis of β-mannosides with a mutant β-mannosidase. Angew Chem Int Ed Engl 113:431–434
Zechel DL, Reid SP, Nashiru O et al (2001) Enzymatic synthesis of carbon-fluorine bonds. J Am Chem Soc 123:4350–4351
Cobb SL, Deng H, McEwan AR et al (2006) Substrate specificity in enzymatic fluorination. The fluorinase fromStreptomyces cattleya accepts 2-deoxyadenosine substrates. Org Biomol Chem 4:1458–1460
Eustáquio AS, O’Hagan D, Moore BS (2010) Engineering fluorometabolite production fluorinase expression in Salinispora tropica yields fluorosalinosporamide. J Nat Prod 73:378–382
Rentmeister A, Arnold FH, Fasan R (2009) Chemo-enzymatic fluorination of unactivated organic compounds. Nat Chem Biol 5:26–28
Ratnayake AS, Janso JE, Feng X, Schlingmann G, Goljer I, Carter GT (2009) Evaluating indole-related derivatives as precursors in the directed biosynthesis of diazepinomicin analogues. J Nat Prod 72:496–499
Murphy CD, Clark BR, Amadio J (2009) Metabolism of fluoroorganic compounds in microorganisms: impacts for the environment and the production of fine chemicals. Appl Microbiol Biotechnol 84:617–629
Amir-Heidari B, Thirlway J, Micklefield J (2008) Auxotrophic-precursor directed biosynthesis of nonribosomal lipopeptides with modified tryptophan residues. Org Biomol Chem 6:975–978
Begue JP, Bonnet-Delpon D (2006) Recent advances (1995–2005) in fluorinated pharmaceuticals based on natural products. J Fluor Chem 127:8992–1012
Ojima (2009) Fluorinein medicinal chemistry and chemical biology. Blackwell, Chichester
Prevedouros K, Cousins IT, Buck RC et al (2006) Sources, fate and transport of perfluorocarboxylates. Environ Sci Technol 40:32–44
Lewandowski G, Meissner E, Milchert E et al (2006) Special applications of fluorinated organic compounds. J Hazard Mater 136:385–391
Barber JL, Berger U, Chaemfa C et al (2007) Analysis of per- and polyfluorinated alkyl substances in air samples from Northwest Europe. J Environ Monit 9:530–541
Senthilkumar K, Ohi E, Sajwan K et al (2007) Perfluorinated compounds in river water, river sediment, market fish, and wildlife samples from Japan. Bull Environ Contam Toxicol 79:427–431
Houde M, Martin JW, Letcher RJ et al (2006) Biological monitoring of polyfluoroalkyl substances: a review. Environ Sci Technol 40:3463–3473
Olsen GW, Burris JM, Ehresman DJ et al (2007) Half-life of serum elimination of perfluorooctanesulfonate, perfluorohexanesulfonate, and perfluorooctanoate in retired fluorochemical production workers. Environ Health Perspect 115:1298–1305
Kennedy GL, Butenhoff JL, Olsen GW et al (2004) The toxicology of perfluorooctanoate. Crit Rev Toxicol 34:351–84
Lau C, Butenhoff JL, Rogers JM et al (2004) The developmental toxicity of perfluoroalkyl acids and their derivatives. Toxicol Appl Pharmacol 198:231–41
Calafat AM, Wong LY, Kuklenyik Z et al (2007) Polyfluoroalkyl chemicals in the U.S. population: data from the National Health and Nutrition Examination Survey (NHANES) 2003–2004 and comparisons with NHANES 1999–2000. Environ Health Perspect 115:1596–602
Yang Q, Xie Y, Eriksson AM et al (2001) Further evidence for the involvement of inhibition of cell proliferation and development in thymic and splenic atrophy induced by the peroxisome proliferator perfluoroctanoic acid in mice. Biochem Pharmacol 62:1133–40
Nakayama S, Harada K, Inoue K et al (2005) Distributions of perfluorooctanoic acid (PFOA) and perfluorooctane sulfonate (PFOS) in Japan and their toxicities. Environ Sci 12:293–313
Tilton SC, Orner GA, Benninghoff AD et al (2008) Genomic profiling reveals an alternate mechanism for hepatic tumor promotion by perfluorooctanoic acid in rainbow trout. Environ Health Perspect 116:1047–55
Asakawa A, Toyoshima M, Fujimiya M et al (2007) Perfluorooctane sulfonate influence feeding behavior and gut motility via the hypothalamus. Int J Mol Med 19:733–739
Yao X, Zhong L (2005) Genotoxic risk and oxidative DNA damage in HepG2 cells exposed to perfluorooctanoic acid. Mutat Res 587:38–44
Ishibashi H, Ishida H, Matsuoka M et al (2007) Estrogenic effects of fluorotelomer alcohols for human estrogen receptor isoforms alpha and beta in vitro. Biol Pharm Bull 30:1358–1359
Fei C, McLaughlin JK, Lipworth L et al (2009) Maternal levels of perfluorinated chemicals and subfecundity. Hum Reprod 24:1200–1205
Fei C, McLaughlin JK, Tarone RE et al (2008) Fetal growth indicators and perfluorinated chemicals:a study in the Danish National Birth Cohort. Am J Epidemiol 168:66–72
Frisbee SJ (2008) The C8 health project:how a class action lawsuit caninteract with public health – history of events. Robert C. Byrd Health Science Center. Available at: http://www.hsc.wvu.edu/som/cmed/c8/index.asp. Accessed 14 Feb 09
Díaz E (2008) Microbial biodegradation: genomics and molecular biology. Caister Academic Press, Norfolk
Natarajana R, Azerada R, Badet B et al (2005) Microbial cleavage of C–F bond. J Fluor Chem 126:425–436
Yu M, Faan Y-W, Chung WYK et al (2007) Isolation and characterization of a novel haloacid permease from Burkholderia cepacia MBA4. Appl Environ Microbiol 73:4874–4880
Murphy CD (2010) Biodegradation and biotransformation of organofluorine compounds. Biotechnol Lett 32:351–359
Hiyama T (2000) Organofluorine compounds: chemistry and applications. Springer, Berlin
Goncalves LPB, Antunes OAC, Pinto GF et al (2003) Kinetic aspects involved in the simultaneous enzymatic synthesis of (S)-3-fluoroalanine and (R)-3-fluorolactic acid. J Fluor Chem 124:219–227
Horiuchi N, Agric J (1961) The CF bond rupture of monofluoroacetate by soil microbes. Chem Soc Jpn 35:870–873
Horiuchi N (1962) The CF bond rupture in monofluoroacetate by soil microbes. II. Some properties of the bacteria and the enzyme. Jpn J Biochem Soc 34:92–98
Liu JQ, Kurihara T, Ichiyama S et al (1998) Reaction mechanism of fluoroacetate dehalogenase from Moraxella sp.B. J Biol Chem 273:30897–30902
Goldman P (1965) The enzymatic cleavage of the carbon-fluorine bond in fluoroacetate. J Biol Chem 240:3434–3438
Tonomura K, Futai F, Tanabe O et al (1965) Defluorination of monofluoroacetate by bacteria. Agric Biol Chem 29:124–128
Kawasaki H, Yahara H, Tonomura K (1984) Cloning and expression in Escherichia coli of the haloacetate dehalogenase genes from Moraxella plasmid pUOl. Agric Biol Chem 48:2627–2632
Goldman P, Milne GWA, Keister DB et al (1968) Carbon-halogen bond cleavage. J Biol Chem 243:428–434
Lacourciere GM, Armstrong RN (1993) The catalytic mechanism of microsomal epoxide hydrolase involves an ester intermediate. J Am Chem Soc 115:10466–10467
Verschueren KHG, Seljee F, Rozeboom HJ et al (1993) Crystallographic analysis of the catalytic mechanism of haloalkane dehalogenase. Nature 363:693–698
Tzeng H-F, Laughlin LT, Armstrong RN et al (1998) Semifunctional site-specific mutants affecting the hydrolytic half-reaction of microsomal epoxide hydrolase. Biochemistry 37:2905–2911
Jitsumori K, Omi R, Kurihara T et al (2009) X-Ray crystallographic and mutational studies of fluoroacetate dehalogenase from Burkholderia sp. strain FA1. J Bacteriol 191:2630–2637
Zeczycki TN, Maurice MS, Attwood PV (2010) Inhibitors of pyruvate carboxylase. Open Enzym Inhib J 3:8–26
Vali M, Vossen JA, Buijs M et al (2008) Targeting of VX2 rabbit liver tumor by selective delivery of 3-bromopyruvate: a biodistribution and survival study. J Pharmacol Exp Ther 327:32–37
Leung LS, Frey PA (1978) Fluoropyruvate: an unusual substrate for pyruvate dehydrogenase. Biochem Biophys Res Commun 81:274–279
Meyer JJM, O’Hagan D (1992) Conversion of fluoropyruvate to fluoroacetate by Dichapetalum cymosum. Phytochemistry 31:499–501
Kaschabek SR, Reineke W (1995) Maleylacetate reductase of Pseudomonas sp strain B13: specificity of substrate conversion and halide elimination. J Bacteriol 177:320–325
Muller D, Schlomann M, Reineke W et al (1996) Maleylacetate reductases in chloroaromatic-degrading bacteria using the modified ortho pathway: comparison of catalytic properties. J Bacteriol 178:298–300
Bertau M (2001) Novel unusual microbial dehalogenation during enantioselective reduction of ethyl 4,4,4-trifluoro acetoacetate with baker’s yeast. Tetrahedron Lett 42:1267–1268
Perez-Pantoja D, Donoso RA, Sanchez MA et al (2009) Genuine genetic redundancy in maleylacetate-reductase-encoding genes involved in degradation of haloaromatic compounds by Cupriavidus necator JMP134. Microbiology 155:3641–3651
Haufe G, Pietz S, Wölker D et al (2003) Synthesis of fluorinated cycloalkyl N‐phenyl-carbamates and their microbial defluorination/oxygenation by Beauveria bassiana. J Org Chem 21:2166–2175
Zhan J, Gunatilaka AAL (2009) Microbial transformation by Beauveria bassiana. In: Rai M (ed) Advances in fungal biotechnology. I. K, International, New Delhi
Akiyama Y, Hiramatsu C, Fukuhara T et al (2006) Selective introduction of a fluorine atom into carbohydrates and a nucleoside by ring-opening fluorination reaction of epoxides. J Fluor Chem 127:920–923
Li X, Turánek J, Knötigová P et al (2009) Hydrophobic tail length, degree of fluorination and headgroup stereochemistry are determinants of the biocompatibility of (fluorinated) carbohydrate surfactants. Colloids Surf B Biointerfaces 73:65–74
Moller AKH, Loft A, Berthelsen AK et al (2011) 18F-FDG PET/CT as a diagnostic tool in patients with extracervical carcinoma of unknown primary site: a literature review. Oncologist 16:445–451
D’Amore TD, Taylor NF (1982) The reaction of 4-deoxy-4-fluoro-D-glucose with an outer membrane protein of Pseudomonas putida. FEBS Lett 143:247–251
Sbrissa D, McIntosh JM, Taylor NF (1990) The metabolism of 4-deoxy-4-fluoro-D-glucose in Pseudomonas putida. Carbohydr Res 203:271–280
Tejada ML, Green JR, Taylor NF (1993) The defluorination of 4-deoxy-4-fluoro-d-glucose in the cytoplasmic membrane of Pseudomonas putida. Carbohydr Res 249:207–219
Zhang C, Zhou Q, Chen L et al (2007) Biodegradation of meta-fluorophenol by an acclimated activated sludge. J Hazard Mater 141:295–300
Engesser KH, Auling G, Busse J et al (1990) 3-Fluorobenzoate enriched bacterial strain FLB-300 degrades benzoate and all 3 isomeric monofluorobenzoates. Arch Microbiol 153:193–199
Misiak K, Casey E, Murphy CD (2011) Factors influencing 4-fluorobenzoate degradation in biofilm cultures of Pseudomonas knackmussii B13. Water Res 45:3512–3520
Emanuelsson MAE, Osuna ME, Jorge RMF (2009) Isolation of a Xanthobacter sp. degrading dichloromethane and characterization of the gene involved in the degradation. Biodegradation 20:235–244
Boersma FGH, McRoberts WC, Cobb SL et al (2004) A F-19 NMR study of fluorobenzoate biodegradation by Sphingomonas sp HB-1. FEMS Microbiol Lett 237:355–361
Schreiber A, Hellwig M, Dorn E et al (1980) Critical reactions in fluorobenzoic acid degradation by Pseudomonas sp. B13. Appl Environ Microbiol 39:58–67
Harper DB, Blakley ER (1971) The metabolism of p-fluorobenzoic acid by a Pseudomonas sp. Can J Microbiol 17:1015–1023
Schennen U, Braun K, Knackmuss H-J (1985) Anaerobic degradation of 2-fluorobenzoate by benzoate–degrading, denitrifying bacteria. J Bacteriol 161:321–325
Drzyzga O, Jannsen S, Blotevogel KH (1994) Mineralization of monofluorobenzoate by a diculture under sulfate-reducing conditions. FEMS Microbiol Lett 116:215–219
Vargas C, Song B, Camps M et al (2000) Anaerobic degradation of fluorinated aromatic compounds. Appl Microbiol Biotechnol 53:342–347
Mouttaki H, Nanny MA, McInerney MJ et al (2009) Metabolism of hydroxylated and fluorinated benzoates by Syntrophus aciditrophicus and detection of a fluorodiene metabolite. Appl Environ Microbiol 75:998–1004
Shimoda K, Hamada H (2010) Bioremediation of fluorophenols by glycosylation with immobilized marine microalga Amphidinium Crassum. Environ Health Insights 4:87–91
Bondar VS, Boersma MG, Vervoort J et al (1998) 19F NMR study on the biodegradation of fluorophenols by various Rhodococcus species. Biodegradation 9:475–486
Kim EJ, Jeon JR, Kim YM et al (2010) Mineralization and transformation of monofluorophenols by Pseudonocardia benzenivorans. Appl Microbiol Biotechnol 87:1569–1577
Prenafeta-Boldu FX, Luykx D, Vervoort J et al (2001) Fungal metabolism of toluene: monitoring of fluorinated analogs by F-19 nuclear magnetic resonance spectroscopy. Appl Environ Microbiol 67:1030–1034
Green NA, Meharg AA, Till C et al (1999) Degradation of 4-fluorobiphenyl by mycorrhizal fungi as determined by 19 F nuclear magnetic resonance spectroscopy and 14 C radiolabelling analysis. Appl Environ Microbiol 65:4021–4027
Murphy CD, Quirke S, Balogun O (2008) Degradation of fluorobiphenyl by Pseudomonas pseudoalcaligenes KF707. FEMS Microbiol Lett 286:45–49
Amadio J, Murphy CD (2010) Biotransformation of fluorobiphenyl by Cunninghamella elegans. Appl Microbiol Biotechnol 86:345–351
Hughes D, Clark BR, Murphy CD (2011) Biodegradation of polyfluorinated biphenyl in bacteria. Biodegradation 22:741–749
Carvalho MF, Alves CCT, Ferreira MIM et al (2002) Isolation and initial characterization of a bacterial consortium able to mineralize fluorobenzene. Appl Environ Microbiol 68:102–105
Carvalho MF, Marco PDe, Duque AF et al (2008) Labrys portucalensis sp. nov., a fluorobenzene-degrading bacterium isolated from an industrially contaminated sediment in northern Portugal. Int J Syst Evol Microbiol 58:692–698
Carvalho MF, Ferreira MIM, Moreira IS et al (2006) Degradation of fluorobenzene by Rhizobiales strain F11 via ortho cleavage of 4-fluorocatechol and catechol. Appl Environ Microbiol 72:7413–7417
Gros M, Petrovic M, Barcelo D (2007) Wastewater treatment plants as a pathway for aquatic contamination by pharmaceuticals in the Ebro river basin (Northeast Spain). Environ Toxicol Chem 26:1553–1562
Brain RA, Johnson DJ, Richards SM et al (2004) Effects of 25 pharmaceutical compounds to Lemna gibba using a seven-day static-renewal test. Environ Toxicol Chem 23:371–382
Robinson AA, Belden JB, Lydy MJ et al (2005) Toxicity of fluoroquinolone antibiotics to aquatic organisms. Environ Toxicol Chem 24:423–430
Hernández A, Sánchez MB, Martínez JL (2011) Quinolone resistance: much more than predicted. Front Microbio 2:22
Wetzstein HG, Schmeer N, Karl W (1997) Degradation of the fluoroquinolone enrofloxacin by the brown rot fungus Gloeophyllum striatum: identification of metabolites. Appl Environ Microbiol 63:4272–4281
Wetzstein H-G, Schneider J, Karl W et al (2006) Patterns of metabolites produced from the fluoroquinolone enrofloxacin by Basidiomycetes indigenous to agricultural sites. Appl Microbiol Biotechnol 71:90–100
Rietjens IM, Tyrakowska B, Veeger C et al (1990) Reaction pathways for biodehalogenation of fluorinated anilines. Eur J Biochem 194:945–954
Travkin VM, Solyanikova IP, Rietjens IMCM et al (2003) Degradation of 3,4-dichloro- and 3,4-difluoroaniline by Pseudomonas fluorescens 26-K. J Environ Sci Health B 38:121–132
Cobb SL, Murphy CD (2009) 19 F NMR applications in chemical biology. J Fluor Chem 130:132–143
Duckett CJ, Wilson ID, Douce DS et al (2007) Metabolism of 2-fluoro-4-iodoaniline in earthworm Eisenia veneta using super (19) F-NMR spectroscopy, HPLC-MS, and HPLC-ICPMS. Xenobiotica 37:1378–1393
Remde A, Debus R (1996) Biodegradability of fluorinated surfactants under aerobic and anaerobic conditions. Chemosphere 32:1563–1574
Arakaki A, Ishii Y, Tokuhisa T et al (2010) Microbial biodegradation of a novel fluorotelomer alcohol, 1H,1H,2H,2H,8H,8H-perfluorododecanol, yields short fluorinated acids. Appl Microbiol Biotechnol 88:1193–1203
Maras M, Vanparys C, Muylle F et al (2006) Estrogen-like properties of fluorotelomer alcohols as revealed by MCF-7 breast cancer cell proliferation. Environ Health Perspect 114:100–105
Wang N, Szostek B, Folsom PW et al (2005) Aerobic biotransformation of 14C-labeled 8–2 telomer B alcohol by activated sludge from a domestic sewage treatment plant. Environ Sci Technol 39:531–538
Wang N, Szostek B, Buck RC et al (2005) Fluorotelomer alcohol biodegradation:direct evidence that perfluorinated carbon chains breakdown. Environ Sci Technol 39:7516–7528
Fasano WJ, Carpenter SC, Gannon SA et al (2006) Absorption, distribution, metabolism, and elimination of 8–2 fluorotelomer alcohol in the rat. Toxicol Sci 91:341–355
Nabb DL, Szostek B, Himmelstein MW et al (2007) In vitro metabolism of 8–2 fluorotelomer alcohol: interspecies comparisons and metabolic pathway refinement. Toxicol Sci 100:333–344
Wang N, Szostek B, Buck RC et al (2009) 8–2 Fluorotelomer alcohol aerobic soil biodegradation: pathways, metabolites, and metabolite yields. Chemosphere 75:1089–1096
Liu J, Wang N, Szostek B et al (2010) 6–2 Fluorotelomer alcohol aerobic biodegradation in soil and mixed bacterial culture. Chemosphere 78:437–444
Fasano WJ, Sweeney LM, Mawn MP et al (2009) Kinetics of 8–2 fluorotelomer alcohol and its metabolites, and liver glutathione status following daily oral dosing for 45 days in male and female rats. Chem Biol Interact 180:281–295
Martin JW, Chan K, Mabury SA et al (2009) Bioactivation of fluorotelomer alcohols in isolated rat hepatocytes. Chem Biol Interact 177:196–203
Frömel T, Knepper TP (2010) Fluorotelomer ethoxylates:sources of highly fluorinated environmental contaminants part I: Biotransformation. Chemosphere 80:1387–1392
Russell MH, Berti WR, Szostek B et al (2010) Evaluation of PFO formation from the biodegradation of a fluorotelomer-based urethane polymer product in aerobic soils. Polym Degrad Stabil 95:79–85
Lee H, D’eon J, Mabury SA (2010) Biodegradation of polyfluoroalkyl phosphates as a source of perfluorinated acids to the environment. Environ Sci Technol 44:3305–3310
Frömel T, Peschka1 M, Fichtner N et al (2008) ω-(Bis(trifluoromethyl)amino)alkane-1-sulfonates:synthesis and mass spectrometric study of the biotransformation products. Rapid Commun Mass Spectrom 22:3957–3967
Stock NL, Lau FK, Ellis DA et al (2004) Polyfluorinated telomer alcohols and sulfonamides in the North American troposphere. Environ Sci Technol 38:991–996
Rhoads KR, Janssen EML, Luthy RG et al (2008) Aerobic biotransformation and fate of N-ethyl perfluorooctane sulfonamidoethanol (N-EtFOSE) in activated sludge. Environ Sci Technol 42:2873–2878
Peschka M, Fichtner N, Hierse W et al (2008) Synthesis and analytical follow-up of the mineralization of a new fluorosurfactant prototype. Chemosphere 72:1534–1540
Dinglasan MJA, Ye Y, Edwards EA et al (2004) Fluorotelomer alcohol biodegradation yields poly- and perfluorinated acids. Environ Sci Technol 38:2857–2864
Liu J, Lee LS, Nies LF et al (2007) Biotransformation of 8: 2 fluorotelomer alcohol in soil and by soil bacteria isolates. Environ Sci Technol 41:8024–8030
Liou JS-C, Szostek B, DeRito CM et al (2010) Investigating the biodegradability of perfluorooctanoic acid. Chemosphere 80:176–183
Moriwaki H, Takagi Y, Tanaka M et al (2005) Sonochemical decomposition of perfluorooctane sulfonate and perfluorooctanoic acid. Environ Sci Technol 39:3388–3392
Hori H, Nagaoka Y, Yamamoto A et al (2006) Efficient decomposition of environmentally persistent perfluorooctanesulfonate and related fluorochemicals using zerovalent iron in subcritical water. Environ Sci Technol 40:1049–1054
Ochoa-herrera V, Sierra-alvarez R, Somogy A et al (2008) Reductive defluorination of perfluorooctane sulfonate. Environ Sci Technol 42:3260–3264
Dolfing J (2003) Thermodynamic considerations for dehalogenation. In: Häggblom MM, Bossert ID (eds) Dehalogenation:microbial processes and environmental applications. Springer, Berlin
Parsons JR, Sáez M, Dolfing J et al (2009) Biodegradation of perfluorinated compounds. Rev Environ Contam Toxicol 196:53–71
Ferreira MIM, Iida T, Hasan SA et al (2009) Analysis of two gene clusters involved in the degradation of 4-fluorophenol by Arthrobacter sp. Strain IF1. Appl Environ Microbiol 75:7767–7773
Husain M, Entsch B, Ballou DP et al (1980) Fluoride elimination from substrates in hydroxylation reactions catalyzed by p-hydroxybenzoate hydroxylase. J Biol Chem 255:4189–4197
van der Bolt FJT, van den Heuvel RHH, Vervoort J et al (1997) 19F NMR study on the regiospecificity of hydroxylation of tetrafluoro-4-hydroxybenzoate by wild-type and Y385F p-hydroxybenzoate hydroxylase: evidence for a consecutive oxygenolytic dehalogenation mechanism. Biochemistry 36:14192–14201
Selesi D, Meckenstock RU (2009) Anaerobic degradation of the aromatic hydrocarbon biphenyl by a sulfate-reducing enrichment culture. FEMS Microbiol Ecol 68:86–93
Visscher PT, Culbertson CW, Oremland RS (1994) Degradation of trifluoroacetate in oxic and anoxic sediments. Nature 369:729–731
Kim BR, Suidan MT, Wallington TJ et al (2000) Biodegradability of trifluoroacetic acid. Environ Eng Sci 17:337–342
Ichiyama S, Kurihara T, Miyagi M et al (2002) Catalysis-linked inactivation of fluoroacetate dehalogenase by ammonia: a novel approach to probe the active-site environment. J Biochem 131:671–677
Tu LQ, Wright PFA, Rix CJ et al (2006) Is fluoroacetate-specific defluorinase a glutathione S-transferase? Comp Biochem Physiol C 143:59–66
Donnelly C, Murphy CD (2007) Bacterial defluorination of 4-fluoroglutamic acid. Appl Microbiol Biotechnol 77:699–703
Peters RA (1957) Mechanism of the toxicity of the active constituent of dichapetalum cymosum and related compounds. Adv Enzymol Relat Subj Biochem 18:113–159
Howard PH (2009) Howard predicting the persistence of organic compounds. Hdb Env Chem 2:17–41
Meylan WM, Boethling RS, Aronson D et al (2007) Chemical structure-based predictive model for methanogenic anaerobic biodegradation potential. Environ Toxicol Chem 26:1785–1792
Dimitrov S, Pavlov T, Nedelcheva D et al (2007) A kinetic model for predicting biodegradation. SAR QSAR Environ Res 18:443–457
Wicker J, Fenner K, Ellis L et al (2010) Predicting biodegradation products and pathways: a hybrid knowledge- and machine learning-based approach. Bioinformatics 26:814–821
O’Hagan D, Schaffrath C, Cobb S et al (2002) Biosynthesis of an organofluorine molecule. Nature 416:279
O’Hagan D (2010) Fluorine in health care: Organofluorine containing blockbuster drugs. J Fluorine Chem 131:1071–1081
Vanden Heuvel JP, Thompson JT, Frame SR et al (2006) Differential activation of nuclear receptors by perfluorinated fatty acid analogs and natural fatty acids: a comparison of human, mouse, and rat peroxisome proliferator-activated receptor-alpha, -beta, and -gamma, liver X receptor-beta, and retinoid X receptor-alpha. Toxicol Sci 92:476–89
Martin JW, Mabury SA, O’Brien PJ (2005) Metabolic products and pathways of fluorotelomer alcohols in isolated rat hepatocytes. Chem Biol Int 155:165-180
Mouttaki H, Nanny MA, McInerney MJ et al (2009) Metabolism of hydroxylated and fluorinated benzoates by Syntrophus aciditrophicus and detection of a fluorodiene metabolite. Appl Environ Microbiol 75:998–1004
Keuning S, Janssen DB, Witholt B (1985) Purification and characterization of hydrolytic haloalkane dehalogenase from Xanthobacter autotrophicus gj10. J Bacteriol 163:635–639
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Zhang, XJ., Lai, TB., Kong, R.YC. (2011). Biology of Fluoro-Organic Compounds. In: Horváth, I. (eds) Fluorous Chemistry. Topics in Current Chemistry, vol 308. Springer, Berlin, Heidelberg. https://doi.org/10.1007/128_2011_270
Download citation
DOI: https://doi.org/10.1007/128_2011_270
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-25233-4
Online ISBN: 978-3-642-25234-1
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)