Abstract
Control of gene expression is crucial to optimize metabolic pathways and synthetic gene networks. Promoters and terminators are stretches of DNA upstream and downstream (respectively) of genes that control both the rate at which the gene is transcribed and the rate at which mRNA is degraded. As a result, both of these elements control net protein expression from a synthetic construct. Thus, it is highly important to discover and engineer promoters and terminators with desired characteristics. This chapter highlights various approaches taken to catalogue these important synthetic elements. Specifically, early strategies have focused largely on semi-rational techniques such as saturation mutagenesis to diversify native promoters and terminators. Next, in an effort to reduce the length of the synthetic biology design cycle, efforts in the field have turned towards the rational design of synthetic promoters and terminators. In this vein, we cover recently developed methods such as hybrid engineering, high throughput characterization, and thermodynamic modeling which allow finer control in the rational design of novel promoters and terminators. Emphasis is placed on the methodologies used and this chapter showcases the utility of these methods across multiple host organisms.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Promoters and terminators play an indispensable role in metabolic engineering and synthetic biology applications for controlling gene expression. These critical elements play a part in regulating both the strength of transcription and the longevity of the transcript. Together, these two forces dictate the overall abundance of mRNA within the cell and ultimately play a significant role in determining protein contents within cells. At the same time, optimizing microorganisms for chemical production via metabolic engineering often requires the use of these elements to create highly regulated intracellular flux [1], often through high-strength promoters [2]. Fine-level control, inducibility, and expression range are all quite important in these endeavors, as has been seen with large strain engineering efforts such as rewiring the yeast Saccharomyces cerevisiae for industrial-level heterologous artemisinin production [3]. Fortunately, our understanding and cataloging of synthetic control elements such as promoters and terminators is continuously improving. In this chapter we consider the selection and engineering of both promoters and terminators for a variety of possible host organisms. Initially, we describe early strategies which mainly relied on genome mining and semi-rational mutagenesis techniques to improve sequence diversity and function. Next, we describe recent advances in the design of these parts using techniques such as hybrid engineering, high-throughput characterization, thermodynamic modeling, synthetic part development, and rational design. In each of these cases, both our understanding and the utility of these parts are enhanced, thus increasing the rate of design cycles within cells.
2 Early Efforts of Promoter Identification and Diversification
2.1 Native Promoter Mining
The initial set of catalogued promoters for synthetic use was derived from the genome of the host organism or a phage that targets the host organism [4–8]. These promoters were often uncovered as a result of genomic dissections. The advent of genome sequencing and annotation (especially of hosts such as Escherichia coli and S. cerevisiae) allowed for the rapid discovery of endogenous promoters, especially when coupled with mRNA quantification methods. In a similar fashion, promoters for more complex systems such as mammalian hosts have largely been discovered via high-throughput screening methods such as “promoter trapping [9–11].” This approach typically involves random integration of a promoter-less vector containing GFP followed by fluorescence-based selection to determine adjacent, upstream regions of the genome that enable transcription. In similar fashion to other hosts, the sequencing of genomes (such as the CHO genome [12]) allowed for the discovery of novel, dynamic promoters such as pTXnip, which expresses proportionally to cell density [13].
Libraries of native promoters serve an important role as major synthetic parts and are among the most highly characterized [14, 15]; however, they remain limited in their ability to sample complete gene expression ranges. Although multiple gene overexpression techniques have been used in E. coli [16–18] and S. cerevisiae [19–22], among other organisms, this approach can be limited and leads to the build-up of toxic intermediates that reduce productivity [23]. In some cases – including commonly-used native promoters in S. cerevisiae – dependencies such as carbon-source metabolism [24] can impact part performance. Such a conditional function is exacerbated in mammalian hosts, as commonly-used viral promoters vary widely in performance between cell lines and are often unstable after many cell generations [25–27]. As a result, further engineering of promoters is necessary to obtain desired fine-tuned expression, stability, and conditional performance.
2.2 Mutagenesis Techniques to Diversify Promoter Strength
Random mutagenesis is a powerful approach to augment promoter function without explicitly requiring extensive knowledge of sequence-to-function mapping. Specifically, because mutagenesis techniques such as error-prone PCR (Ep-PCR) indiscriminately target both consensus and non-consensus promoter regions, libraries with a large dynamic range of promoter function can be easily obtained. For instance, error-prone PCR was used to generate a mutant library of the prokaryotic PL-λ bacteriophage-derived promoter, enabling a 196-fold dynamic range of expression in E. coli [28]. The utility of this library was demonstrated by optimizing the expression of phosphoenolpyruvate carboxylase (ppc) for biomass yield and deoxy-xylulose-P-synthase (dxs) for maximal lycopene production. The importance of an expression continuum was highlighted by the fact that optimal dxs expression was dependent on strain genetic background. Similar mutagenesis of the strong constitutive S. cerevisiae TEF1 promoter yielded a library exhibiting a 15-fold dynamic range [28, 29]. Likewise, this library was used to optimize glycerol 3-phosphate dehydrogenase (GPD1) expression for glycerol overproduction in yeast.
As an alternative to Ep-PCR, serial deletion of promoter regions has been used to modulate expression, especially for mammalian hosts. Initially, serial deletion was used as a genetic tool to systematically remove portions of a promoter sequence to better understand function [30–32]. As these deletions often tend to dampen promoter activity, this approach has recently been used to generate libraries of weaker promoters [33, 34]. In this regard, serial deletion has been used to create knockdown libraries of glutamine synthetase (GS) expression for the GS-CHO expression system [35]. Moreover, serial deletion can also identify promoter variants that are cell-line specific. For example, the human cytomegalovirus (hCMV) promoter was optimized for transgene expression in both CHO-K1 and HEK-293 cells [36]. This study found that the full-length promoter gave the highest stable expression in CHO-K1 cells whereas the addition of the first exon to the minimal enhancer and core promoters was optimal for expression in HEK293 cells.
Although Ep-PCR and serial deletion are effective at creating a large dynamic range of promoter strength, these approaches suffer from two major deficiencies: (1) higher level expression is hard to achieve and (2) large pools of inactive mutants are generated because of aberrant mutagenesis of elements critical for transcription [2]. Newer techniques (described in the sections below) are required to gain higher expression consistently. To address the second limitation of large inactive pools, more targeted approaches that make use of molecular understanding of promoter function can be employed. As an example, a saturation mutagenesis approach (Fig. 1a) was used to specifically modulate the sequence between consensus −35 “TTGACA” and −10 “TATAAT” motifs [37]. As these two motifs are both necessary and sufficient for the recruitment of the σ70 factor of RNA polymerase II (RNAP II) to initiate transcription [38], a randomized linker region was generated that resulted in a promoter library with a 400-fold dynamic range in Lactococcus lactis [39]. To improve the dynamic range further, a library including mutations of the −35 and −10 motifs exhibited another three orders of magnitude in range, thus demonstrating the importance of the entire promoter sequence [39].
Saturation mutagenesis strategies used to diversify promoters and improve understanding of promoter design rules. (a) Prokaryotic promoters have a highly constrained architecture with consensus −35 and −10 motifs spaced by exactly 17 base pairs for optimal function. (b, c) Eukaryotic promoters lack a rigidly-defined consensus architecture. (b) In yeast, promoters can be broken down into an upstream activating sequence (UAS) containing transcription factor binding sites (TFBSs), such as those for GCR1p (CT-Box) and Rap1p (RPG-box), and a core promoter which serves to recruit RNA Polymerase II. (c) In mammalian hosts, promoters follow a similar general architecture but contain additional consensus motifs such as the initiator element (INR, shown above), transcription factor IIB recognition sequence (BRE), motif ten element (MTE), and downstream promoter element (DPE)
Eukaryotic promoters, although more complex and less rigidly defined than prokaryotic counterparts, can be broken down into a core promoter [40, 41] and upstream enhancer element(s) [42, 43] located 5′ of the core promoter. Efforts to engineer these distinct elements have been successful. For example, Jeppsson et al. created an ENO1-based promoter scaffold (Fig. 1b) containing two GCR1p TFBSs, two Rap1p TFBSs, and a TATA box coupled by spacers whose length was based on the architecture of native promoters [44, 45]. Randomization of these spacer regions afforded 37 synthetic promoters that spanned 3 orders of magnitude in strength. The utility of this library was demonstrated for the controlled knockdown of ZWF1 expression, resulting in a 16% increase in yeast ethanol production from xylose fermentation. Finally, this same approach of creating synthetic promoter scaffolds followed by saturation mutagenesis has been applied to mammalian promoters (Fig. 1c) in which mutagenesis of regions between TFBSs in the JeT promoter afforded a weakened synthetic promoter library with a tenfold range [46].
Collectively, these early mutagenesis techniques demonstrate that utilizing native promoters (prokaryotes) or constructing synthetic promoters (eukaryotes) followed by randomization of spacer regions can provide a promoter library marked by downregulation. Although efforts continue to use these approaches, a greater understanding of promoter architecture and high-throughput characterization techniques have yielded new methods to design promoters rationally with highly specific expression characteristics as described in the following sections.
3 Rational Construction of Promoters with Desired Characteristics
3.1 Hybrid Promoter Engineering
Once essential components of promoter architecture are defined, it is possible to combine disparate elements in a “hybrid promoter engineering” scheme. Importantly, in contrast to Ep-PCR and saturation mutagenesis, the construction of hybrid promoters often yields synthetic promoters which are stronger than the core scaffold [2]. Thus, this technique serves as a potent way to amplify the expression of promoters – an important goal of many engineering endeavors. The first instance of hybrid promoter engineering involved the fusion of the trp and lac promoters to create the tacI and tacII promoters [47]. Notably, this resulted in promoters that were between 7 and 11 times stronger than the derepressed lac promoter although maintaining the same regulation. Similar approaches in E. coli have been utilized to generate regulated promoters. For instance, a strong binding site for the FadR transcription factor was placed upstream of the strong phage promoters PL and PT7 to create a dynamic biosensor-regulator for acyl-CoA conversion to fatty acids in E. coli [48]. A similar concept was used to produce a malonyl-CoA responsive hybrid promoter that controlled flux from acyl-CoA to malonyl-CoA [49]. However, prokaryotic promoters may also be limited by promoter escape after transcript initiation, meaning that the addition of redundant hybrid elements is not guaranteed to improve transcription and can reduce transcription in some cases [50].
Unlike prokaryotic promoters, eukaryotic promoters are largely enhancer-limited, meaning that the addition of enhancer elements (by including additional binding sites) can both regulate and amplify promoter activity (Fig. 2a) [51]. Combining previously isolated Upstream Activating Sequences (UASs) from CYC1 [52, 53], CLB2 (UASCLB) [54], CIT1 (UASCIT) [55], GAL1-10 (UASGAL) [56], and TEF1 (UASTEF) [51] with core promoters such as GPD (PGPD) [24], TEF1 (PTEF) [4], LEU2 (PLEUM) [52], and CYC1 (PCYC) [57] can result in a predictable increase in transcriptional activity [51]. Ultimately, the strongest constitutive promoter in yeast was generated which had mRNA levels 2.5-fold higher than the GPD promoter [24]. Hybrid yeast promoters can also be designed for altered regulation. For example, linking various elements of UASGAL to a constitutive core results in a functional, galactose inducible promoter [51]. A similar approach has been conducted with regulated regions of the ARO9 UAS [58]. Collectively, these approaches resulted in a library of galactose-inducible promoters with a 40-fold range in induced expression strength, and a tryptophan-inducible promoter with a 29-fold range in induced expression strength. This hybrid promoter approach has been extended to non-conventional yeasts such as the host Yarrowia lipolytica. For example, hybrid engineering on the LEU2 core promoter resulted in a constitutive promoter library with 400-fold range in expression [49]. Most importantly, this work demonstrated the generalizability of the hybrid promoter approach to multiple core promoters and alternative UAS elements [59]. Such strong promoters were used in the rewiring of Y. lipolytica, in which constitutive overexpression of DGA1 using the UAS1B16-TEF1 hybrid promoter (among other genetic changes) resulted in a 60-fold improvement in lipogenesis [60].
Promoter engineering strategies. (a) Hybrid promoter engineering uses combinations of sequence motifs to modify expression and regulation. (b) Synthetic promoter scaffolds may be constructed based on native promoters with desired characteristics. These scaffolds can then be diversified using saturation mutagenesis and modified via hybrid promoter engineering. (c) Minimal synthetic core and enhancer elements may be selected using randomization followed by FACS. (d) Promoter elements have been fully characterized via high-throughput oligo library synthesis followed by FACS sorting into different expression bins. (e) Expression can be tuned by altering nucleosome occupancy using a nucleosome prediction model or via addition of nucleosome-disfavoring poly (dA:dT) tracts. Chromatin regulators (CRs) can program a diverse range of transcriptional logic when targeted to synthetic promoters, thus creating more efficient synthetic circuits
Finally, the hybrid promoter approach has been further generalized to mammalian systems. For instance, the binding site of repressor PDX1 in the hCMV promoter was removed, enhancing expression fourfold in transient luciferase experiments [61]. The traditional additive hybrid approach has also been generalized to mammalian hosts to increase expression [62], improve transgene expression in specific hosts [63, 64], and impart novel regulation on promoters. As an example, a strong, cold-inducible promoter was created by combining a mild-cold responsive enhancer (MCRE) to the hCMV promoter [65]. Using this promoter and shifting temperature from 37°C to 32°C afforded sixfold higher erythropoietin production. Collectively, these results indicate that the hybrid promoter approaches are useful in both increasing net expression and imparting unique regulation.
3.2 Synthetic Promoter Scaffolds and Libraries
More recently, efforts have been made to establish synthetic and/or orthogonal [66, 67] promoters. Certainly bacterial systems can take advantage of the T7 RNA polymerase system [68] to generate short, synthetic, and orthogonal promoters for usage in logic gates [69–71]. However, the diversity of synthetic prokaryotic promoters is limited by the strict consensus promoter architecture not found in eukaryotes. To create a library of orthogonal core promoters in S. cerevisiae, native promoters were screened over a wide range of growth conditions to find a promoter scaffold that would exhibit the least amount of natural regulation [67]. The resulting candidate promoter, PFY1 (PPFY1), was then de-constructed to produce a minimal promoter scaffold (Fig. 2b) containing the ~100-bp core promoter, a Reb1p binding site, and a poly-dT element that maintained nucleosome depletion and constant DNA bending for constitutive RNA polymerase II access. By randomizing the spacer regions within this core promoter, a library of 36 minimally-regulated promoters with a 10-fold dynamic range in expression was created. This same methodology has been generalized to other organisms including Pichia pastoris, where four natively regulated promoters were sequence aligned to create a set of minimal core promoters from which sequence elements were transferred to modify the native AOX1 promoter [72]. This same approach has been applied to human liver cells where a synthetic promoter scaffold with enhanced TF binding was created via the alignment of the hCMV and HEF1α promoters [64].
In an effort to generate more minimal, synthetic promoters using a library-based approach, Redden and Alper [73] developed an S. cerevisiae minimal core promoter scaffold (Fig. 2c) by dissecting both the core element and the UAS element and identifying functional, minimal units using a library-based approach involving FACS analysis and a series of robustness tests. Ultimately, a series of nine generic core elements were isolated which have limited homology to the genome. The same methodical workflow was used to isolate six synthetic 10-bp UAS sequences that activated these synthetic core promoters. Finally, these elements were combined to generate a minimal promoter with 70% the activity of GPD with an 80% reduction in size. Importantly, these promoters represent a minimal scaffold with highly defined consensus regions similar to those of prokaryotic promoters and thus these elements may be further rationally engineered for desired characteristics. Finally, in HeLa cells, synthetic 100-bp enhancers were created via construction of a library containing tandem repeats of random, micro-array printed 10-bp oligonucleotides [74]. This approach resulted in an enhancer with twice the strength of the hCMV enhancer. Thus, rationally constructing purely synthetic libraries can result in novel promoters with prescribed function across multiple hosts.
4 Sequence-Level Prediction and Specification of Promoters
Most of the methods described above rely heavily on repeated iterations of the synthetic biology design-build-test cycle [75, 76]. In contrast, the ability to specify promoter function at the DNA level would rapidly accelerate the field of synthetic biology by reducing the number of design cycles. This section describes many of the efforts that have been made toward this end.
4.1 Promoter Characterization and Standardization
Promoters, composed of a vast array of distinct regulatory elements, behave as a system that integrates an input from the host to produce an output: gene expression. As high-throughput oligo synthesis [77] and quantification of DNA, mRNA, and protein levels have improved, large combinatorial libraries may be generated to measure promoter performance across a wide range of contexts (Fig. 2d). For instance, in prokaryotes, the Ribosome Binding Site (RBS) controls the binding of the ribosome to the mRNA transcript, thus regulating gene expression at the translational level whereas the promoter regulates expression at the transcriptional level. The independent function of these two regulatory elements has been thoroughly characterized and modeled via the construction of a library containing combinations of 114 promoters and 111 RBSs [78]. Although the model could explain 96% of RNA levels, its prediction of 82% of protein levels demonstrates the complex regulation of prokaryotic gene expression at the translational level. Thus, it is important to consider RBS performance when designing expression cassettes in pathways.
Eukaryotic transcription is regulated by a complex “program” of TF binding and RNAP II recruitment, and thus underlying “design rules” can be extracted that determine how the orientation, copy number, and context of TFBSs affect transcription. To parse these design rules, Sharon et al. [79] created a combinatorial library varying these parameters for 75 transcription factors. Fluorescence-activated cell sorting (FACS) coupled with high-throughput sequencing of 6,500 barcoded promoters generated a large dataset that uncovered regulatory design rules for TFs. For instance, in promoters that contained a Gcn4p binding site, expression and binding site location were related via a periodic function. Using a similar high-throughput characterization technique in mouse liver cells, it was possible to rapidly screen thousands of rationally designed enhancer haplotype variants [80]. This study found that enhancers are highly robust to single nucleotide variation (SNV), but that combinations of SNVs have an additive negative effect on function. This study also determined novel expression-enhancing motifs and characterized predicted TFBSs, thus laying the foundation for future enhancer design rules. In mammalian hosts, a similar predictive model has been used to identify K-mers that denote enhancers recognized by certain TFs [81, 82]. This model can be trained on CHIP-seq data [83] to predict enhancers throughout the genome.
Whereas TFBSs with a well-characterized function may be added to tune expression rationally, sequence-function mapping for core promoters is less understood. The core promoter sequence determines how RNAP II binds in the TATA region, forms the pre-initiation complex to unwind the DNA directly downstream, scans for a TSS, and initiates transcription [84–86]. Moving towards rational design, 859 native S. cerevisiae promoters were characterized using flow cytometry to generate a model relating maximal expression to short oligo motifs (K-mers) which impact these steps [86]. Although this model only accounted for 25% of the variance in an aggregate test promoter set, it nonetheless mapped expression-enhancing and repressing characteristics to short motifs in the core promoter to allow prediction of novel synthetic promoters. These results were improved upon via construction and high-throughput characterization of 13,000 specifically designed synthetic core promoters [87], leading to a model relating expression to the presence and orientation of consensus core promoter regions. However, despite analysis of thousands of systematically designed core promoters, the design rules for sequence level specification of core promoter activity are much less understood than those for UAS manipulation.
4.2 Thermodynamic Modeling and Prediction of Promoters
To fully expedite the synthetic biology design cycle, it is desirable to develop methods to design entire promoters de novo for predictable expression. In prokaryotes, thermodynamic models of ribosome interaction with mRNA secondary structure have been constructed to calculate the proportion of bound RBS-mRNA complexes, and thus translation rate [88, 89]. A thermodynamics-based RBS calculator was able to predict expression levels within a factor of 2.3 over an expression range of five orders of magnitude. Most importantly, this RBS calculator takes into account variations in translation rate depending on the genetic context of the RBS, thus allowing a “forward engineering” approach for novel applications.
Although eukaryotic transcriptional regulation involves countless protein factor binding events prior to transcription initiation, it is nonetheless possible to thermodynamically model individual steps as a surrogate for transcription initiation rate. A thermodynamic model incorporating both TF-DNA and TF-TF interactions was trained upon a promoter library containing different TFBS combinations using “effective TF concentration” as a floating parameter to fit the data [90]. Overall, the model predicted 56% of the variance in expression across a wide variety of TFBS arrangements, thus laying a foundation for de novo design of regulatory logic at the DNA sequence level.
To generalize this model further, other events in transcription initiation have been considered. Thermodynamic modeling of the TATA–TATA-binding protein (TBP) complex formed as a first step in the recruitment of RNAP II [91] and re-design of promoters with different consensus TATA boxes created a promoter library which predictably scaled with the thermodynamic affinity of TBP to each TATA Box [92]. Incorporating the thermodynamic model for the TBP–TATA complex with the previously developed model for TF-RNA Polymerase II and TF-TF binding [90] explained 75% of variance in promoter expression across a wide variety of genetic contexts. These examples demonstrate the utility of thermodynamically modeling transcription initiation steps as a means to predict expression. Since discovering promoters is highly important for uncharacterized mammalian hosts, thermodynamic sequence-level approaches have been used to predict novel promoters based on DNA structural properties such as duplex stability and bendability [93, 94]. In addition, mammalian promoter regions have been modeled at the sequence level using an “alpha score,” which describes the likelihood that a genomic region contains a promoter based on its nucleotide composition. Remodeling the X-linked gene cancer/testis antigen 1A promoter to have twice the alpha score improved expression in a non-quantitative manner [95]. Although predictive of high expression, these techniques are limited as they cannot design promoters de novo with prescribed expression. Nevertheless, they demonstrate the potential to use heuristic models for the design and prediction of DNA function.
4.3 Prediction and Rational Modulation of Promoter Nucleosome Occupancy
In eukaryotes, the secondary structure of promoter DNA wound around nucleosomes controls access to the transcription machinery [96]. As a result, the rational design of novel promoters must consider how primary sequence contributes to DNA secondary structure. Nucleosome occupancy at promoters strongly regulates gene expression because nucleosome binding can occlude TFBSs and RNAP II recruitment to the core promoter [97]. Accordingly, rational addition of a tunable nucleosome-disfavoring poly(dA:dT) element [91, 98, 99] upstream of the natural Gcn4p binding site in a synthetic His3-based promoter library afforded predictable control over nucleosome occupancy and thus expression [100]. Similarly, mutation of CpG islands known to be prone to methylation and silencing by histones eliminated promoter silencing during long-term transgene expression in embryonic stem cells [101]. Thus, nucleosome-disfavoring sequences may be considered part of the rational eukaryotic promoter engineering toolbox along with the addition of hybrid enhancers (Fig. 2e).
To map nucleosome occupancy to primary sequence for predictive engineering of promoters, a Hidden Markov Model (HMM) was trained on a genome-wide nucleosome map [102]. This model was utilized to investigate nucleosome occupancy of the previously mentioned TEF1 promoter library, demonstrating that expression correlated inversely with predicted cumulative nucleosome occupancy in a very robust manner. To create a predictive model, a greedy algorithm was developed which allowed re-design of native promoters for up to 16-fold greater strength [103]. Furthermore, this approach was used for the successful de novo design of synthetic yeast promoters. Importantly, sequence-level prediction of nucleosome occupancy affords a predictive method to optimize native promoters fully regardless of genetic context. As a result, future efforts in this area must consider the precise control of nucleosome occupancy to modulate expression.
4.4 Design of Synthetic Promoters with Controlled Chromatin Environment
Moving forward from nucleosome models, the context of eukaryotic DNA is important in considering promoter function. Specifically, eukaryotic DNA is wound around histone octamers in 147 base pair increments and packaged together tightly to create the “bead-on-a-string” backbone of the chromatin [104]. This structure is not composed randomly; in fact, the structure of chromatin surrounding genes has a direct impact on their regulation [105–111]. Thus, any endeavor to engineer promoters rationally as synthetic biology “parts” that exhibit defined functions in any genetic context must take into account the chromatin environment of the promoter.
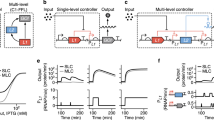
The first step towards any rational bottom-up synthetic biology engineering approach is to parse design rules from the native system. To create design rules for chromatin-based control, a combinatorial library of zinc finger-based synthetic transcription factors was created with specific yeast chromatin regulators (CRs) tethered as the activation domain [112]. These CRs impact gene expression by regulating PIC formation, remodeling and assembly of nucleosomes, chromatin accessibility via histone modification, and transcriptional elongation. From this library screening approach, many different classes of CRs were delineated: activators and repressors, synergistic regulators, spatially encoded regulators that could repress transcription from a non-canonical position downstream of genes, and CRs that could activate or repress multiple genes simultaneously over a long range of genomic space. These minimal chromatin-based components can thus act as synthetic “parts” to create a diverse array of transcriptional logic and predictably tune expression by altering chromatin state. These initial efforts demonstrate the first work towards considering greater genetic context for promoters.
In closing, promoter discovery and characterization has progressed from genome mining to random mutagenesis to combinatorial and rational design. In some of these later cases, the use of computational models has been able to speed the design-build-test cycle. Although limitations still exist with respect to inducible promoters, pure synthetic design, and maximal expression levels, the field has progressed rapidly in recent years.
5 Terminator Discovery and Characterization
In addition to promoters, terminators serve as an important control point when tuning expression in circuits and pathways [113, 114]. Unlike promoters, terminator cataloguing has not been as extensive until recently. In fact, most commonly used terminators have been relics from past experiments and are not often the most efficient. As an example, commonly used terminators such as the native bacteriophage T7 terminator exhibit low termination efficiencies, meaning that transcriptional flux continues through the expression cassette and affects the regulation of downstream genes and limits polymerase recycling [113–115]. Furthermore, the collection of terminators available to researchers has traditionally been much smaller in breadth than promoters [116], thus limiting large-scale pathways and circuits because of the fear of genetic instability via homologous recombination [117, 118]. Terminators also serve as a control point to tune expression in eukaryotes via the stability of the 3′ end of the mRNA transcript [119–121]. Thus, the base of commonly used terminators must be diversified to meet pathway specifications via both discovery and engineering techniques. We highlight various approaches from terminator mining to synthetic design and models in the following sections.
5.1 Native Terminator Mining
To diversify initially from the commonly used terminator library in E. coli, an extensive library of 582 natural and synthetic terminators [122, 123] was constructed and analyzed for its termination efficiency [124]. To enable further terminator engineering, the study also delineated terminator design rules based on a mechanism where RNAP stalls at the U:A tract, allowing an RNA hairpin to form within the RNA exit channel and terminating transcription. It was shown that the composition of the terminator U-tract effectively controls polymerase dissociation and can thus be rationally designed to impact terminator strength. This work served as one of the more exhaustive studies for bacteria to determine alternative terminators for synthetic constructs.
In contrast to prokaryotic intrinsic termination, eukaryotic mRNA transcript stability is regulated by recruited protein factors such as the cleavage and polyadenylation specificity factor (CPSF) and cleavage stimulation factor (CstF) [125]. Thus, terminators must be characterized not only by their termination efficiency but also by their impact on mRNA and protein levels. Yamanishi et al. undertook the first genome-scale flow cytometry characterization of yeast terminators, determining that the majority of terminators enabling higher expression from a synthetic construct came from ribosomal protein genes [120]. A separate, high-capacity terminator library was constructed by selecting a subset of terminators originating from genes shown to have higher mRNA half-lives [121]. Characterization of this library established a direct relationship between terminator strength and mRNA half-life, thus laying the groundwork for terminator design rules. In addition, the utility of these alternative terminators was proven by improved pathway flux with similar or lower promoter strength as those originally paired with a “traditional” terminator. Thus, terminators clearly serve as an important synthetic part that must be rationally specified to tune expression for metabolic engineering applications.
6 Rational Construction of Terminators with Desired Characteristics
6.1 Hybrid Terminator Engineering
Similar to promoters, the hybrid engineering approach has yielded synthetic terminators with enhanced efficiencies. Multiple combinations of both native and synthetic termination signals were used to enhance the termination efficiency of the T7 terminator while retaining its orthogonality [126]. However, this hybrid approach faces limitations in eukaryotes because termination is a highly concerted process regulated by multiple disparate elements (Fig. 3a).
Evolution of terminators in S. cerevisiae. (a) Unlike promoters, native terminators consist of many defined consensus motifs. (b) A minimal terminator scaffold was created by spacing these terminator motifs 10 bp apart. (c) This scaffold was engineered by rationally modifying the linkers between consensus motifs, adding upstream and downstream elements, and changing the length and sequence of consensus motifs. Licensing: Reprinted with permission from Curran KA, Morse NJ, Markham KA et al. (2015) Short, Synthetic Terminators for Improved Heterologous Gene Expression in Yeast. ACS Synth. Biol. 2015, 4, 824–832. doi:10.1021/sb5003357. Copyright © 2015 American Chemical Society
6.2 Synthetic Terminator Scaffolds and Libraries
To overcome the limitations of hybrid terminator engineering in yeast, a synthetic minimal terminator scaffold (TGuo) was constructed by stringing together defined consensus efficiency, positioning, and poly-adenylation elements which cooperate in the cleavage and 3′ polyadenylation of the mRNA transcript (Fig. 3b) [127]. This minimal scaffold was both diversified and enhanced using modified consensus termination elements and mRNA stability elements [128] to produce a library of rationally designed synthetic terminators (Fig. 3c) which were functional in multiple hosts and improved CAD1 expression for itaconic acid production [129]. Importantly, this technique allowed delineation of design rules based on consensus element identity and spacing, enabling potential rational design of synthetic terminators. These resulting terminators were much shorter in size than native terminators with the additional benefit of enhanced mRNA stability and increased protein production. Thus, in a similar fashion as described with promoters above, once a fundamental understanding of molecular function is obtained, synthetic part design can proceed.
7 Sequence-Level Prediction and Specification of Terminators
Although the previously described methods of synthetic terminator design allow rational diversification of the terminator library, they are nonetheless limited by the natural sequence space. Pure de novo design of terminators requires a fundamental understanding of the constraints underlying terminator function. Very early studies have begun to elucidate underlying design principles for terminators; however, this area is lagging behind the progress made with promoters as described above.
7.1 Terminator Characterization and Standardization
To this end, high-throughput studies have been carried out to measure quantitatively the performance of terminators and determine predictive sequence features for design in both prokaryotes and eukaryotes. For instance, systematic variation of terminator U-tract and hairpin stem-loop sequences in the aforementioned E. coli terminator library [124] afforded optimal expression-enhancing consensus sequences for rational construction of synthetic terminators.
Both native and synthetic terminator libraries have been constructed and characterized to tease apart the functions of different terminator motifs [130] in regulating mRNA abundance in yeast [131, 132]. Characterization of these libraries showed that the AU-rich efficiency element upstream of the poly(A) site plays a major role in 3′ end processing and transcription termination. In addition, terminators were broken down into mono- and di-nucleotide K-mers, leading to identification of dA:dT elements as a major determinant in terminator strength. From these studies, it appears that terminators can be broken down into a collection of tunable elements for rational design.
7.2 Thermodynamic Modeling and Prediction of Terminators
To generate a finer continuum of terminator function, it has become necessary to engineer entirely synthetic terminator sequences based on known design rules and thermodynamic prediction. In prokaryotes, multiple biophysical models have been developed to predict terminator strength based on elementary steps in termination, including U:A hybrid formation, hairpin formation, and mRNA transcript dissociation [122, 133, 134]. Training one of these models on a set of natural and synthetic terminators over a large dynamic range in termination efficiencies afforded a linear sequence-function model with a high coefficient of determination (R 2 = 0.81) [134].
In S. cerevisiae, however, terminator function is much less predictable based simply on distinct sequence elements whose function is determined by biophysical models. In fact, characterization of the aforementioned rationally designed synthetic library [129] demonstrated that consensus termination motifs were not entirely additive. This suggests there is a fundamental code underlying termination in yeast which remains to be uncovered before thermodynamic prediction becomes feasible. However, with a more rigidly defined architecture than promoters, yeast terminators are highly amenable to rational engineering for desired characteristics. Thus, creating fundamental models to describe eukaryotic termination and half-life stabilization are required to advance the field of terminator engineering.
8 Future Directions in Promoter and Terminator Engineering
Improved promoters and terminators help minimize the length of the design cycle. Optimal design of these elements must meet three criteria: robustness, orthogonality, and predictable tunability. Promoters and terminators must be robust in that they function consistently regardless of genetic background, genetic context, and cellular environment [135]. In this regard, unexpected deviation from desired promoter or terminator function is a severe hindrance to the rapid development of circuits and pathways leading to multiple iterations of the design cycle. To improve robustness, efforts have been made to create synthetic promoter scaffolds based on highly constitutive promoters which function consistently across many different cellular environments. However, to date, few significant efforts have been made to engineer eukaryotic promoters that are robust to differing genetic contexts. These efforts are also complicated by the fact that eukaryotic promoters are highly regulated by the chromatin environment in which they are placed. It is thus imperative to develop design rules that govern promoter and terminator chromatin environment to predict and control these factors for optimal gene expression. The promise of purely orthogonal elements can bypass some of the robustness issues as these promoters and terminators seem to function more ubiquitously. Overall, many strides have been made in the past 5 years to provide novel expression capabilities to promoters and terminators. However, because of the regulatory complexity of microorganism hosts, new techniques must be developed to predict and design promoters and terminators for desired function. Nevertheless, these new synthetic parts have greatly improved the ability to engineer strains for metabolic engineering and synthetic biology applications.
References
Blazeck J, Alper H (2010) Systems metabolic engineering: genome-scale models and beyond. Biotechnol J 5:647–659. doi:10.1002/biot.200900247
Blazeck J, Alper HS (2013) Promoter engineering: recent advances in controlling transcription at the most fundamental level. Biotechnol J 8:46–58. doi:10.1002/biot.201200120
Paddon CJ, Westfall PJ, Pitera DJ et al (2013) High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496:528–532. doi:10.1038/nature12051
Gatignol A, Dassain M, Tiraby G (1990) Cloning of Saccharomyces cerevisiae promoters using a probe vector based on phleomycin resistance. Gene 91:35–41
Hauf J, Zimmermann F, Müller S (2000) Simultaneous genomic overexpression of seven glycolytic enzymes in the yeast Saccharomyces cerevisiae. Enzyme Microb Technol 26:688–698
Hawley DK, McClure WR (1983) Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res 11:2237–2255
Reifenberger E, Boles E, Ciriacy M (1997) Kinetic characterization of individual hexose transporters of Saccharomyces cerevisiae and their relation to the triggering mechanisms of glucose repression. Eur J Biochem 245:324–333
Diderich JA, Schepper M, van Hoek P et al (1999) Glucose uptake kinetics and transcription of HXT genes in chemostat cultures of Saccharomyces cerevisiae. J Biol Chem 274:15350–15359. doi:10.1074/jbc.274.22.15350
Pontiller J, Gross S, Thaisuchat H et al (2008) Identification of CHO endogenous promoter elements based on a genomic library approach. Mol Biotechnol 39:135–139. doi:10.1007/s12033-008-9044-9
Pontiller J, Maccani A, Baumann M et al (2010) Identification of CHO endogenous gene regulatory elements. Mol Biotechnol 45:235–240. doi:10.1007/s12033-010-9278-1
Chen J, Haverty J, Deng L et al (2013) Identification of a novel endogenous regulatory element in Chinese hamster ovary cells by promoter trap. J Biotechnol 167:255–261. doi:10.1016/j.jbiotec.2013.07.001
Xu X, Nagarajan H, Lewis NE et al (2011) The genomic sequence of the Chinese hamster ovary (CHO)-K1 cell line. Nat Biotechnol 29:735–741. doi:10.1038/nbt.1932
Le H, Vishwanathan N, Kantardjieff A et al (2013) Dynamic gene expression for metabolic engineering of mammalian cells in culture. Metab Eng 20:212–220. doi:10.1016/j.ymben.2013.09.004
Partow S, Siewers V, Bjørn S et al (2010) Characterization of different promoters for designing a new expression vector in Saccharomyces cerevisiae. Yeast 27:955–964. doi:10.1002/yea.1806
Sun J, Shao Z, Zhao H et al (2012) Cloning and characterization of a panel of constitutive promoters for applications in pathway engineering in Saccharomyces cerevisiae. Biotechnol Bioeng 109:2082–2092. doi:10.1002/bit.24481
Terpe K (2006) Overview of bacterial expression systems for heterologous protein production: from molecular and biochemical fundamentals to commercial systems. Appl Microbiol Biotechnol 72:211–222. doi:10.1007/s00253-006-0465-8
Studier FW, Moffatt BA (1986) Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol 189:113–130
Elvin CM, Thompson PR, Argall ME et al (1990) Modified bacteriophage lambda promoter vectors for overproduction of proteins in Escherichia coli. Gene 87:123–126
Walfridsson M, Hallborn J, Penttilä M et al (1995) Xylose-metabolizing Saccharomyces cerevisiae strains overexpressing the TKL1 and TAL1 genes encoding the pentose phosphate pathway enzymes transketolase and transaldolase. Appl Environ Microbiol 61:4184–4190
Lu C, Jeffries T (2007) Shuffling of promoters for multiple genes to optimize xylose fermentation in an engineered Saccharomyces cerevisiae strain. Appl Environ Microbiol 73:6072–6077. doi:10.1128/AEM.00955-07
Wisselink HW, Toirkens MJ, del Rosario Franco Berriel M et al (2007) Engineering of Saccharomyces cerevisiae for efficient anaerobic alcoholic fermentation of L-arabinose. Appl Environ Microbiol 73:4881–4891. doi:10.1128/AEM.00177-07
Alper H, Stephanopoulos G (2009) Engineering for biofuels: exploiting innate microbial capacity or importing biosynthetic potential? Nat Rev Microbiol 7:715–723. doi:10.1038/nrmicro2186
Keasling JD (2010) Manufacturing molecules through metabolic engineering. Science 330:1355–1358. doi:10.1126/science.1193990
Da Silva NA, Srikrishnan S (2012) Introduction and expression of genes for metabolic engineering applications in Saccharomyces cerevisiae. FEMS Yeast Res 12:197–214. doi:10.1111/j.1567-1364.2011.00769.x
Addison CL, Hitt M, Kunsken D, Graham FL (1997) Comparison of the human versus murine cytomegalovirus immediate early gene promoters for transgene expression by adenoviral vectors. J Gen Virol 78(Pt 7):1653–1661
Xia W, Bringmann P, McClary J et al (2006) High levels of protein expression using different mammalian CMV promoters in several cell lines. Protein Expr Purif 45:115–124. doi:10.1016/j.pep.2005.07.008
Kim M, O’Callaghan PM, Droms KA, James DC (2011) A mechanistic understanding of production instability in CHO cell lines expressing recombinant monoclonal antibodies. Biotechnol Bioeng 108:2434–2446. doi:10.1002/bit.23189
Alper H, Fischer C, Nevoigt E, Stephanopoulos G (2005) Tuning genetic control through promoter engineering. Proc Natl Acad Sci U S A 102:12678–12683
Nevoigt E, Kohnke J, Fischer CR et al (2006) Engineering of promoter replacement cassettes for fine-tuning of gene expression in Saccharomyces cerevisiae. Appl Environ Microbiol 72:5266–5273. doi:10.1128/AEM.00530-06
Boshart M, Weber F, Jahn G et al (1985) A very strong enhancer is located upstream of an immediate early gene of human cytomegalovirus. Cell 41:521–530
Dorsch-Häsler K, Keil GM, Weber F et al (1985) A long and complex enhancer activates transcription of the gene coding for the highly abundant immediate early mRNA in murine cytomegalovirus. Proc Natl Acad Sci U S A 82:8325–8329
Nelson JA, Reynolds-Kohler C, Smith BA (1987) Negative and positive regulation by a short segment in the 5′-flanking region of the human cytomegalovirus major immediate-early gene. Mol Cell Biol 7:4125–4129
Prentice HL, Tonkin CJD, Caamano L, Sisk WP (2007) High level expression of proteins using sequences from the ferritin heavy chain gene locus. J Biotechnol 128:50–60. doi:10.1016/j.jbiotec.2006.09.021
Thaisuchat H, Baumann M, Pontiller J et al (2011) Identification of a novel temperature sensitive promoter in CHO cells. BMC Biotechnol 11:51. doi:10.1186/1472-6750-11-51
Fan L, Kadura I, Krebs LE et al (2013) Development of a highly-efficient CHO cell line generation system with engineered SV40E promoter. J Biotechnol 168:652–658. doi:10.1016/j.jbiotec.2013.08.021
Mariati NYK, Chao S-H et al (2010) Evaluating regulatory elements of human cytomegalovirus major immediate early gene for enhancing transgene expression levels in CHO K1 and HEK293 cells. J Biotechnol 147:160–163. doi:10.1016/j.jbiotec.2010.02.022
Nair TM, Kulkarni BD (1994) On the consensus structure within the E. coli promoters. Biophys Chem 48:383–393
Gruber BTM, Gross CA (2003) Assay of Escherichia coli RNA polymerase: sigma–core interactions. Methods Enzymol 370:206–212
Jensen PR, Hammer K (1998) The sequence of spacers between the consensus sequences. Appl Environ Microbiol 64:82–87
Juven-Gershon T, Hsu J-Y, Kadonaga JT (2006) Perspectives on the RNA polymerase II core promoter. Biochem Soc Trans 34:1047–1050. doi:10.1042/BST0341047
Juven-Gershon T, Hsu J-Y, Theisen JW, Kadonaga JT (2008) The RNA polymerase II core promoter – the gateway to transcription. Curr Opin Cell Biol 20:253–259. doi:10.1016/j.ceb.2008.03.003
Struhl K (1984) Genetic properties and chromatin structure of the yeast gal regulatory element: an enhancer-like sequence. Proc Natl Acad Sci U S A 81:7865–7869
Struhl K (1995) Yeast transcriptional regulatory mechanisms. Annu Rev Genet 29:651–674. doi:10.1146/annurev.ge.29.120195.003251
Jeppsson M, Johansson B, Jensen PR et al (2003) The level of glucose-6-phosphate dehydrogenase activity strongly influences xylose fermentation and inhibitor sensitivity in recombinant Saccharomyces cerevisiae strains. Yeast 20:1263–1272. doi:10.1002/yea.1043
Drazinic CM, Smerage JB, López MC, Baker HV (1996) Activation mechanism of the multifunctional transcription factor repressor-activator protein 1 (Rap1p). Mol Cell Biol 16:3187–3196
Tornøe J, Kusk P, Johansen TE, Jensen PR (2002) Generation of a synthetic mammalian promoter library by modification of sequences spacing transcription factor binding sites. Gene 297:21–32. doi:10.1016/S0378-1119(02)00878-8
De Boer HA, Comstock LJ, Vasser M (1983) The tac promoter: a functional hybrid derived from the trp and lac promoters. Proc Natl Acad Sci U S A 80:21–25
Zhang F, Carothers JM, Keasling JD (2012) Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol 30:354–359. doi:10.1038/nbt.2149
Blazeck J, Liu L, Redden H, Alper H (2011) Tuning gene expression in Yarrowia lipolytica by a hybrid promoter approach. Appl Environ Microbiol 77:7905–7914. doi:10.1128/AEM.05763-11
Hsu LM (2002) Promoter clearance and escape in prokaryotes. Biochim Biophys Acta 1577:191–207. doi:10.1016/S0167-4781(02)00452-9
Blazeck J, Garg R, Reed B, Alper HS (2012) Controlling promoter strength and regulation in Saccharomyces cerevisiae using synthetic hybrid promoters. Biotechnol Bioeng 109:2884–2895. doi:10.1002/bit.24552
Guarente L, Hoar E (1984) Upstream activation sites of the CYC1 gene of Saccharomyces cerevisiae are active when inverted but not when placed downstream of the “TATA box”. Proc Natl Acad Sci U S A 81:7860–7864
Guarente L, Lalonde B, Gifford P, Alani E (1984) Distinctly regulated tandem upstream activation sites mediate catabolite repression of the CYC1 gene of S. cerevisiae. Cell 36:503–511
Van Slyke C, Grayhack EJ (2003) The essential transcription factor Reb1p interacts with the CLB2 UAS outside of the G2/M control region. Nucleic Acids Res 31:4597–4607
Rosenkrantz M, Kell CS, Pennell EA et al (1994) Distinct upstream activation regions for glucose-repressed and derepressed expression of the yeast citrate synthase gene CIT1. Curr Genet 25:185–195
West RW, Yocum RR, Ptashne M (1984) Saccharomyces cerevisiae GAL1-GAL10 divergent promoter region: location and function of the upstream activating sequence UASG. Mol Cell Biol 4:2467–2478
Guarente L, Ptashne M (1981) Fusion of Escherichia coli lacZ to the cytochrome c gene of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 78:2199–2203
Kim D, Kim JD, Baek K et al. (2003) Improved mammalian expression systems by manipulating transcriptional termination regions. Biotechnol Prog 19:1620–1622. doi:10.1021/bp0341186
Blazeck J, Reed B, Garg R et al (2013) Generalizing a hybrid synthetic promoter approach in Yarrowia lipolytica. Appl Microbiol Biotechnol 97:3037–3052. doi:10.1007/s00253-012-4421-5
Blazeck J, Hill A, Liu L et al (2014) Harnessing Yarrowia lipolytica lipogenesis to create a platform for lipid and biofuel production. Nat Commun 5:3131. doi:10.1038/ncomms4131
Chao S-H, Harada JN, Hyndman F et al (2004) PDX1, a cellular homeoprotein, binds to and regulates the activity of human cytomegalovirus immediate early promoter. J Biol Chem 279:16111–16120. doi:10.1074/jbc.M312304200
Berg DT, Mooney PQ, Baez M, Grinnell BW (1988) Tandem promoter/enhancer units create a versatile regulatory element for the expression of genes in mammalian cells. Nucleic Acids Res 16:1635
Gehrke S, Jérôme V, Müller R (2003) Chimeric transcriptional control units for improved liver-specific transgene expression. Gene 322:137–143
Magnusson T, Haase R, Schleef M et al (2011) Sustained, high transgene expression in liver with plasmid vectors using optimized promoter-enhancer combinations. J Gene Med 13:382–391. doi:10.1002/jgm.1585
Sumitomo Y, Higashitsuji H, Higashitsuji H et al (2012) Identification of a novel enhancer that binds Sp1 and contributes to induction of cold-inducible RNA-binding protein (cirp) expression in mammalian cells. BMC Biotechnol 12:72. doi:10.1186/1472-6750-12-72
Garg A, Lohmueller JJ, Silver PA, Armel TZ (2012) Engineering synthetic TAL effectors with orthogonal target sites. Nucleic Acids Res 40:7584–7595. doi:10.1093/nar/gks404
Blount BA, Weenink T, Vasylechko S, Ellis T (2012) Rational diversification of a promoter providing fine-tuned expression and orthogonal regulation for synthetic biology. PLoS One 7:1–11. doi:10.1371/journal.pone.0033279
Rong M, He B, McAllister WT, Durbin RK (1998) Promoter specificity determinants of T7 RNA polymerase. Proc Natl Acad Sci U S A 95:515–519
Temme K, Hill R, Segall-Shapiro TH et al (2012) Modular control of multiple pathways using engineered orthogonal T7 polymerases. Nucleic Acids Res 40:8773–8781. doi:10.1093/nar/gks597
Shis DL, Bennett MR (2013) Library of synthetic transcriptional AND gates built with split T7 RNA polymerase mutants. Proc Natl Acad Sci U S A 110:5028–5033. doi:10.1073/pnas.1220157110
Segall-Shapiro TH, Meyer AJ, Ellington AD et al (2014) A “resource allocator” for transcription based on a highly fragmented T7 RNA polymerase. Mol Syst Biol 10:742
Vogl T, Ruth C, Pitzer J et al (2014) Synthetic core promoters for Pichia pastoris. ACS Synth Biol 3:188–191. doi:10.1021/sb400091p
Redden H, Alper HS (2015) The development and characterization of synthetic minimal yeast promoters. Nat Commun 6:7810. doi:10.1038/ncomms8810
Schlabach MR, Hu JK, Li M, Elledge SJ (2010) Synthetic design of strong promoters. Proc Natl Acad Sci U S A 107:2538–2543. doi:10.1073/pnas.0914803107
Khalil AS, Collins JJ (2010) Synthetic biology: applications come of age. Nat Rev Genet 11:367–379. doi:10.1038/nrg2775
Way JC, Collins JJ, Keasling JD, Silver PA (2014) Integrating biological redesign: where synthetic biology came from and where it needs to go. Cell 157:151–161. doi:10.1016/j.cell.2014.02.039
Kosuri S, Eroshenko N, LeProust EM et al (2010) Scalable gene synthesis by selective amplification of DNA pools from high-fidelity microchips. Nat Biotechnol 28:1295–1299. doi:10.1038/nbt.1716
Kosuri S, Goodman DB, Cambray G et al (2013) Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc Natl Acad Sci U S A 110:14024–14029. doi:10.1073/pnas.1301301110
Sharon E, Kalma Y, Sharp A et al (2012) Inferring gene regulatory logic from high-throughput measurements of thousands of systematically designed promoters. Nat Biotechnol 30:521–530. doi:10.1038/nbt.2205
Patwardhan RP, Hiatt JB, Witten DM et al (2012) Massively parallel functional dissection of mammalian enhancers in vivo. Nat Biotechnol 30:265–270. doi:10.1038/nbt.2136
Lee D, Karchin R, Beer MA (2011) Discriminative prediction of mammalian enhancers from DNA sequence. Genome Res 21:2167–2180. doi:10.1101/gr.121905.111
Fletez-Brant C, Lee D, McCallion AS, Beer MA (2013) kmer-SVM: a web server for identifying predictive regulatory sequence features in genomic data sets. Nucleic Acids Res 41:W544–W556. doi:10.1093/nar/gkt519
Johnson DS, Mortazavi A, Myers RM, Wold B (2007) Genome-wide mapping of in vivo protein-DNA interactions. Science 316:1497–1502. doi:10.1126/science.1141319
Giardina C, Lis JT (1993) DNA melting on yeast RNA polymerase II promoters. Science 261:759–762
Sugihara F, Kasahara K, Kokubo T (2011) Highly redundant function of multiple AT-rich sequences as core promoter elements in the TATA-less RPS5 promoter of Saccharomyces cerevisiae. Nucleic Acids Res 39:59–75. doi:10.1093/nar/gkq741
Lubliner S, Keren L, Segal E (2013) Sequence features of yeast and human core promoters that are predictive of maximal promoter activity. Nucleic Acids Res 41:5569–5581. doi:10.1093/nar/gkt256
Lubliner S, Regev I, Lotan-pompan M et al (2015) Core promoter sequence in yeast is a major determinant of expression level. 1008–1017. doi:10.1101/gr.188193.114.1008
Salis HM, Mirsky EA, Voigt CA (2010) Automated design of synthetic ribosome binding sites to precisely control protein expression. Nat Biotechnol 27:946–950. doi:10.1038/nbt.1568.Automated
Salis HM (2011) The ribosome binding site calculator. Methods Enzymol 498:19–42. doi:10.1016/B978-0-12-385120-8.00002-4
Gertz J, Cohen BA (2009) Environment-specific combinatorial cis-regulation in synthetic promoters. 1–9. doi:10.1038/msb2009.1
Iyer V, Struhl K (1995) Poly(dA:dT), a ubiquitous promoter element that stimulates transcription via its intrinsic DNA structure. EMBO J 14:2570–2579
Mogno I, Vallania F, Mitra RD, Cohen BA (2010) TATA is a modular component of synthetic promoters. Genome Res 20:1391–1397. doi:10.1101/gr.106732.110
Abeel T, Saeys Y, Bonnet E et al (2008) Generic eukaryotic core promoter prediction using structural features of DNA. Genome Res 18:310–323. doi:10.1101/gr.6991408
Gan Y, Guan J, Zhou S (2012) A comparison study on feature selection of DNA structural properties for promoter prediction. BMC Bioinformatics 13:4. doi:10.1186/1471-2105-13-4
Grabherr MG, Pontiller J, Mauceli E et al (2011) Exploiting nucleotide composition to engineer promoters. PLoS One 6, e20136. doi:10.1371/journal.pone.0020136
Kornberg RD, Lorch Y (1999) Twenty-five years of the nucleosome, fundamental particle of the eukaryote chromosome. Cell 98:285–294. doi:10.1016/S0092-8674(00)81958-3
Basehoar AD, Zanton SJ, Pugh BF (2004) Identification and distinct regulation of yeast TATA box-containing genes. Cell 116:699–709. doi:10.1016/S0092-8674(04)00205-3
Anderson JD, Widom J (2001) Poly(dA-dT) promoter elements increase the equilibrium accessibility of nucleosomal DNA target sites. Mol Cell Biol 21:3830–3839. doi:10.1128/MCB.21.11.3830-3839.2001
De Boer CG, Hughes TR (2014) Poly-dA:dT tracts form an in vivo nucleosomal turnstile. PLoS One 9, e110479. doi:10.1371/journal.pone.0110479
Raveh-Sadka T, Levo M, Shabi U et al (2012) Manipulating nucleosome disfavoring sequences allows fine-tune regulation of gene expression in yeast. Nat Genet 44:743–750. doi:10.1038/ng.2305
Swindle CS, Kim HG, Klug CA (2004) Mutation of CpGs in the murine stem cell virus retroviral vector long terminal repeat represses silencing in embryonic stem cells. J Biol Chem 279:34–41. doi:10.1074/jbc.M309128200
Xi L, Fondufe-Mittendorf Y, Xia L et al (2010) Predicting nucleosome positioning using a duration hidden Markov model. BMC Bioinformatics 11:346. doi:10.1186/1471-2105-11-346
Curran KA, Crook NC, Karim AS et al (2014) Design of synthetic yeast promoters via tuning of nucleosome architecture. Nat Commun 5:4002. doi:10.1038/ncomms5002
Keung AJ, Joung JK, Khalil AS, Collins JJ (2015) Chromatin regulation at the frontier of synthetic biology. Nat Rev Genet. doi:10.1038/nrg3900
Ellis L, Atadja PW, Johnstone RW (2009) Epigenetics in cancer: targeting chromatin modifications. Mol Cancer Ther 8:1409–1420. doi:10.1158/1535-7163.MCT-08-0860
Ernst J, Kheradpour P, Mikkelsen TS et al (2011) Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473:43–49. doi:10.1038/nature09906
Gaspar-Maia A, Alajem A, Meshorer E, Ramalho-Santos M (2011) Open chromatin in pluripotency and reprogramming. Nat Rev Mol Cell Biol 12:36–47. doi:10.1038/nrm3036
Onder TT, Kara N, Cherry A et al (2012) Chromatin-modifying enzymes as modulators of reprogramming. Nature 483:598–602. doi:10.1038/nature10953
Rheinbay E, Louis DN, Bernstein BE, Suvà ML (2012) A tell-tail sign of chromatin: histone mutations drive pediatric glioblastoma. Cancer Cell 21:329–331. doi:10.1016/j.ccr.2012.03.001
Schuster-Böckler B, Lehner B (2012) Chromatin organization is a major influence on regional mutation rates in human cancer cells. Nature 488:504–507. doi:10.1038/nature11273
Wang X, Chen J, Quinn P (2012) Reprogramming microbial metabolic pathways. Zhurnal Eksp i Teor Fiz 181–201. doi:10.1007/978-94-007-5055-5
Keung AJ, Bashor CJ, Kiriakov S et al (2014) Using targeted chromatin regulators to engineer combinatorial and spatial transcriptional regulation. Cell 158:110–120. doi:10.1016/j.cell.2014.04.047
Du L, Gao R, Forster AC (2009) Engineering multigene expression in vitro and in vivo with small terminators for T7 RNA polymerase. Biotechnol Bioeng 104:1189–1196. doi:10.1002/bit.22491
Du L, Villarreal S, Forster AC (2012) Multigene expression in vivo: supremacy of large versus small terminators for T7 RNA polymerase. Biotechnol Bioeng 109:1043–1050. doi:10.1002/bit.24379
Carter AD, Morris CE, McAllister WT (1981) Revised transcription map of the late region of bacteriophage T7 DNA. J Virol 37:636–642
Redden H, Morse N, Alper HS (2014) The synthetic biology toolbox for tuning gene expression in yeast. FEMS Yeast Res 1–12. doi:10.1111/1567-1364.12188
Sleight SC, Bartley BA, Lieviant JA, Sauro HM (2010) Designing and engineering evolutionary robust genetic circuits. J Biol Eng 4:12. doi:10.1186/1754-1611-4-12
Renda BA, Hammerling MJ, Barrick JE (2014) Engineering reduced evolutionary potential for synthetic biology. Mol Biosyst 10:1668–1678. doi:10.1039/c3mb70606k
Abe H, Aiba H (1996) Differential contributions of two elements of rho-independent terminator to transcription termination and mRNA stabilization. Biochimie 78:1035–1042
Yamanishi M, Ito Y, Kintaka R et al (2013) A genome-wide activity assessment of terminator regions in Saccharomyces cerevisiae provides a “Terminatome” toolbox BT. ACS Synth Biol 2:337–347
Curran KA, Karim AS, Gupta A, Alper HS (2013) Use of expression-enhancing terminators in Saccharomyces cerevisiae to increase mRNA half-life and improve gene expression control for metabolic engineering applications. Metab Eng 19:88–97. doi:10.1016/j.ymben.2013.07.001
Von Hippel PH, Yager TD (1991) Transcript elongation and termination are competitive kinetic processes. Proc Natl Acad Sci U S A 88:2307–2311
Gama-Castro S, Jiménez-Jacinto V, Peralta-Gil M et al (2008) RegulonDB (version 6.0): gene regulation model of Escherichia coli K-12 beyond transcription, active (experimental) annotated promoters and Textpresso navigation. Nucleic Acids Res 36:D120–D124. doi:10.1093/nar/gkm994
Chen Y-J, Liu P, Nielsen AAK et al (2013) Characterization of 582 natural and synthetic terminators and quantification of their design constraints. Nat Methods 10:659–664. doi:10.1038/nmeth.2515
Nag A, Narsinh K, Martinson HG (2007) The poly(A)-dependent transcriptional pause is mediated by CPSF acting on the body of the polymerase. Nat Struct Mol Biol 14:662–669. doi:10.1038/nsmb1253
Mairhofer J, Wittwer A, Cserjan-puschmann M, Striedner G (2014) Synthetic termination signal capable of improving bioprocess. ACS Synth Biol. doi:10.1021/sb5000115
Guo Z, Sherman F (1996) Signals sufficient for 3′-end formation of yeast mRNA. Mol Cell Biol 16:2772–2776
Geisberg JV, Moqtaderi Z, Fan X et al (2014) Global analysis of mRNA isoform half-lives reveals stabilizing and destabilizing elements in yeast. Cell 156:812–824. doi:10.1016/j.cell.2013.12.026
Curran KA, Morse NJ, Markham KA et al (2015) Short, synthetic terminators for improved heterologous gene expression in yeast. ACS Synth Biol 4(7):824–832. doi:10.1021/sb5003357
Mischo HE, Proudfoot NJ (2013) Disengaging polymerase: terminating RNA polymerase II transcription in budding yeast. Biochim Biophys Acta 1829:174–185. doi:10.1016/j.bbagrm.2012.10.003
Shalem O, Carey L, Zeevi D et al (2013) Measurements of the impact of 3′ end sequences on gene expression reveal wide range and sequence dependent effects. PLoS Comput Biol. doi:10.1371/journal.pcbi.1002934
Shalem O, Sharon E, Lubliner S et al (2015) Systematic dissection of the sequence determinants of gene 3′ end mediated expression control. PLoS Genet 11:e1005147. doi:10.1371/journal.pgen.1005147
Yager TD, von Hippel PH (1991) A thermodynamic analysis of RNA transcript elongation and termination in Escherichia coli. Biochemistry 30:1097–1118
Cambray G, Guimaraes JC, Mutalik VK et al (2013) Measurement and modeling of intrinsic transcription terminators. Nucleic Acids Res 41:5139–5148. doi:10.1093/nar/gkt163
Leavitt JM, Alper HS (2015) Advances and current limitations in transcript-level control of gene expression. Curr Opin Biotechnol 34:98–104. doi:10.1016/j.copbio.2014.12.015
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Deaner, M., Alper, H.S. (2016). Promoter and Terminator Discovery and Engineering. In: Zhao, H., Zeng, AP. (eds) Synthetic Biology – Metabolic Engineering. Advances in Biochemical Engineering/Biotechnology, vol 162. Springer, Cham. https://doi.org/10.1007/10_2016_8
Download citation
DOI: https://doi.org/10.1007/10_2016_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-55317-7
Online ISBN: 978-3-319-55318-4
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)