Abstract
Autophagy has a direct and indirect role in health and disease. By definition, it is an exceedingly complex process which degrades and modifies damaged and surplus macromolecules in the cell. Autophagy has also been defined as a process which plays a role in dynamics of organelles using enzymes in lysosomes. Malaria parasite, i.e. Plasmodium, is a deadly parasite which encounters various conditions during its life cycle, ranging from temperature fluctuations to drug pressures. Role of autophagy in coping with these changes in Plasmodium is not well defined; however, there is growing evidence for role of this mechanism in life cycle of the parasite. This chapter highlights the link between different homologues of ATG protein present in Plasmodium and explores the mechanisms underlying these connections and their implications for cell physiology and survival of the parasite.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Last decade has seen tremendous gains in knowledge regarding autophagy pathways. Autophagy plays a role in a variety of processes, which range from its cellular role of degrading unwanted material to more sophisticated phenomenon of organellar homoeostasis. Autophagy now represents an emerging role in various human diseases including infectious disease and cancer. Autophagy also plays crosstalk and overlapping function with a number of cellular processes such as apoptosis and senescence. A lot of information is available about autophagy events; however, a constant effort to understand process of autophagy is ongoing. Cellular homoeostasis in malaria pathogen, i.e. Plasmodium, is not a much explored area, but it gives a challenging opportunity for researchers to understand this complex phenomenon in this highly evolved parasite. In this chapter, we review three main aspects: autophagy in the Plasmodium; autophagy in cellular homoeostasis in Plasmodium; and autophagy of host affecting the parasite life cycle. Plasmodium autophagy is an upcoming area of research; thus, we have tried to compile all the information regarding homologues of autophagy pathways and proteins in Plasmodium. Finally, we describe the role of host autophagy in limiting the spread of Plasmodium infection and crosstalk between host and pathogen autophagy mechanisms.
Autophagy and Signalling in Plasmodium
Plasmodium falciparum is a deadly pathogen which causes malaria in millions of people globally, leading to death of thousands annually. It also causes a socio-economic impact on the well-being of the human society as it leads to poverty and neurological sequelae for an undetermined number [1]. There have been extensive studies defining cellular signalling during the life cycle of the parasite, which clearly defines its mechanism for survival and pathogenesis under normal conditions. However, little is been studied to understand the mechanism parasite follows to cope with the stress conditions it faces during the life cycle, which can include “Crisis form” morphology of the parasite. In crisis form morphology, parasite appears as punctuated and condensed parasite [2, 3]. This crisis form is attributed to the immunological response of human system against the parasite. Later the crisis form morphology was also observed in chloroquine-treated parasites, which subsequently died after the treatment [4]. Similarly, different cellular stress on the parasite led to induction of specific sequence of cellular events which ultimately caused apoptosis-like cell death [5,6,7]. This apoptosis-like death of the parasite opened a big debate about regulated cell death mechanism of the parasite. Regulated cell death is a balance between different mechanisms involving survival strategies, e.g. autophagy, and those involved in cell death process. Autophagy-like events have been observed both during the cell adaptation to stress and during the regulated cell death process, and it could be playing role in cell adaptation to stress [8]. Interestingly, the cellular stress-induced effects on parasite are reversible for a particular period till the apoptosis pathways are induced, suggesting an anti-apoptotic survival mechanism in the parasite during initial phase of stress [5,6,7]. It will be crucial to understand the metabolic pathways induced for survival mechanism which may guide towards key proteins involved in cellular homoeostasis in the parasite.
Another important aspect which needs investigation is about cellular homoeostasis in the parasite during various stages of its life cycle. Plasmodium enters a human when an Anopheles mosquito bites and releases sporozoite in the bloodstream from where it travels to the hepatocyte. After rounds of inter-hepatic cycle, the merozoites get released in the blood and invade erythrocytes. During this life cycle, the parasites transit between various stages, ring, trophozoites and schizonts, which also involve changes at the organellar level. Mechanism involved in this transition is not known, but autophagy can be speculated as one of the possible pathways which can govern this phenomenon. One such example comes from the observation that post-invasion of hepatocytes, P. berghei ATG8-decorated micronemes (an invasion-related organelle) are expelled from the parasite and degraded by enzymes present in the parasitophorous vacuole (PV) lumen [9]. Therefore, formation of the “crisis form” and degradation of specific organelles during parasite differentiation need exploration of a mechanism which can explain these phenomena. Autophagy is one of such mechanism which is suggested to play similar roles in other organisms [8, 10]. Autophagy is a catabolic pathway conserved among eukaryotes that allows cells to rapidly eliminate large unwanted structures such as aberrant protein aggregates, superfluous or damaged organelles and invading pathogens. Autophagy is considered to be a survival mechanism and an important pathway in the maintenance of the cell homoeostasis. Disruption in the normal autophagy machinery leads cells towards non-apoptotic pathways. Autophagy can be both specific and non-specific towards elimination of damaged organelles or aggregates of unfolded proteins.
Autophagy was described for the first time almost five decades back. Various studies based on lysosomal, biochemical and morphological changes provided the role of such a mechanism. Further research in the field of yeast biology allowed a better understanding of the phenomenon at molecular and genetic level. Identification, isolation and characterization of 27 genes related to the autophagy mechanism, which now have been termed as autophagy-related genes (ATGs), have paved the path for better understanding of the phenomenon [11]. Identification of autophagy started with acid phosphatase-stained electron microscopy images showing presence of lysosomes in rat liver [12]. Purification of these lysosomes leads to identification of other structures termed as dense bodies which have almost the same properties of lysosomes including presence of hydrolases [13]. Later these dense bodies were named as autophagosome vacuoles or AVs, which are distinct from lysosomes. The presence of whole organelles like mitochondria in these autophagosome vacuoles from newborn mouse kidney cells highlighted the organelle degradative role of these structures [14]. It is becoming highly evident that autophagy is highly selective quality control mechanism whose basal levels are important to maintain cellular homoeostasis [15]. A number of organelles have been found to be selectively turned over by autophagy, and cargo-specific names have been given to distinguish the various selective pathways including the ER (reticulophagy or ER-phagy), peroxisomes (pexophagy), mitochondria (mitophagy), lipid droplets (lipophagy), secretory granules (zymophagy) and even parts of the nucleus (nucleophagy) [16,17,18]. In addition, it has been proposed for a long time that, although generally a cytoprotective mechanism, autophagy can initiate or execute cell death under certain conditions such as cellular stress or starvation [19].
The basic feature of autophagy is generation of autophagosome, which is the site where degradation of macromolecules to their native constituents takes place. Different stages for the autophagosome generation have been listed as induction, nucleation, expansion, fusion and cargo recycling. All of these stages involve different autophagy proteins along with different signalling molecules. Out of ~30 autophagy-related genes (ATGs) which were identified in yeast, the Plasmodium genome contains 15 homologues of autophagy-related genes (ATGs). Here we have discussed role of different ATGs in various steps of the autophagy process, their homologue in Plasmodium and their possible role in cellular homoeostasis of parasite life cycle.
Induction and Nucleation of Autophagosome Vacuoles
The induction of autophagy vacuoles and the source of membranes for their formation has remained a phenomenon which has been investigated and has remained complex for long time. Endoplasmic reticulum and Golgi bodies were pointed out as the source for the generation of membranes for these autophagosome vacuoles, although no confirmatory source of AVs are suggested till now [20,21,22]. Once these AVs are formed, irrespective of their source of generations, morphologic studies showed that these structures acquire degradative enzymes through fusion with mature lysosomes [23]. Different steps and various ATG proteins are involved in the generation of mature autophagolysosome [24, 25].
Change in the environment, i.e. nutritional starvation or temperature changes, are sensed by the cells using different receptors on their surfaces, which is further relayed to the downstream effector molecules using different signalling mechanisms. Autophagy initiation occurs with development of single site called the pre-autophagosomal structure (PAS; also called phagophore assembly site). Under a fluorescence microscope, the PAS is detected as a single, dot-like accumulation of Atg proteins next to the vacuole. In Mammal’s PAS is missing, but small sites where assembly starts are found to be associated initially with ER membrane [26, 27], Golgi membranes [28, 29] and nuclear membranes [30]. Nutrition starvation is one of the triggers which can throw a cell into the mechanism of autophagy. In deprivation of some amino acids or proteins, autophagy has been induced in the cell through target of rapamycin complex 1 (TORC1) which can sense changes in the extracellular environment of cell. TORC1 induction further leads to activation of ATG1 kinase complex. Plasmodium lacks homologue of TORC1 complex; PfTOR has very poor homology or even extensive variations that set it apart from other conserved sequences, which may suggest that PfTOR is regulated differently or its activity may be variable. Interestingly, phosphorylation of PfeIF2α by PfeIKI in response to starvation is described by Fennell et al. [31], which suggest that this system (downstream of TOR) regulates response to amino acid starvation. Similarly, Williams et al. [32] described an autophagy pathway independent of mTOR that is induced in a Ca2+-dependent fashion in neuronal precursor cells. This precedent suggests that it is possible for autophagy to operate independently of TOR. In Plasmodium, generation of autophagosomes has remained a very intriguing mechanism. As in other systems, source of phagophore membranes is not very clear and has remained a question of investigation.
Atg1 Kinase Complex
In yeast, which is a metazoan, autophagy has been inhibited by the TOR kinase, which phosphorylates components of Atg1 kinase complex. The ATG1 complex is composed of ATG1 and ATG13, along with a scaffold made up of ATG17-ATG31 and ATG29. This whole complex acts a nucleation site to bring all the ATG proteins required for complex functioning. ATG1 kinase activity in complex with ATG13-Atg17 is required for the phagophore formation. Further the activity of ATG1 has been regulated by TOR kinase which prevents interaction of ATG13 with ATG1 by phosphorylating the ATG13 [33]. TOR kinase is energy-sensing kinase and thus acts as a regulator for the initiation of autophagy sensitive to nutrient and growth factor availability. Similar function is carried out by ULK1 which is a homologue of Atg1 in reticulocytes [34]. In mammals, the complex comprises of ULK1, ATG13, ATG101 and FIP200. Out of the described Atg1 kinase complex defined in mammals, Atg1 and its mammalian binding partner FIP200 and Atg101 are present in the Plasmodium genome. ULK1 is the mammalian orthologue of Atg1, while FLIP200 is the mammalian counterpart of Atg17. Atg13 is an important partner of Atg1 complex; however it is not present in the Plasmodium genome [35].
PfAtg1 also differs from Atg1 from other organisms; it lacks C-terminal domain, which is required for interaction with ATG13. Another interesting observation about PfAtg1 is presence of Atg8-interacting domains, which points towards presence of Atg13-independent pathway of autophagy in Plasmodium, as has been shown in yeast [36].
ATG9 and Its Cycling System
Apart from the start point of phagosome formation, another important question about autophagy is the source of lipid used for formation of autophagosomes and their cycling to the site of initiation. Atg9 is an integral membrane protein which acts as a carrier of membrane during the assembly process [37]. Atg9 is little different from most of other Atg proteins, as most of Atg proteins has single localization on PAS, whereas ATG9 localizes to multiple puncta structures [38]. Plasmodium genome shows absence of PfAtg9, which points towards presence of other important protein compensating for the role of this important protein in the generation of autophagosomes.
Cycling of ATG9 between PAS and structure other than PAS is thought to be potential source of lipid for the formation of PAS. Anterograde movement of Atg9 to PAS involves several other Atg proteins in mammals including PfATG23 and PfAtg27. Plasmodium genome also lacks presence of PfAtg23 and pfAtg27 homologues. Retrieval of Atg9 from PAS to peripheral locations depends on Atg1-Atg13 kinase complex and Atg2, Atg18 and PtdIns3K complex I [38]. Atg2 and Atg8 is a set of important interacting peripheral membrane proteins which can interact with Atg9 [39]. Interaction of Atg18 requires both Atg2 and Atg8 [40, 41]. Atg18 has been shown to interact with two phosphoinositides, PtdIns(3)P and PtdIns[3, 5]P2; however Atg18 interacting with PtdIns(3)P is required for autophagy [42]. In yeast, Atg1-Atg13 complex promotes interaction of Atg9 with Atg2 and Atg18, and formation of the ternary complex leads to release of Atg9 [43].
PfAtg18 also shows interaction with PtdIns(3)P and gets localized to vesicles near apicoplast and food vacuole [44]. PfAtg18 disruption leads to increase in the number of PfATG18-positive vesicles with P. falciparum maturation, which corroborated well with increases in PfATG18 expression in mature trophozoites and schizonts. The PfATG18 vesicles were observed in close vicinity of the food vacuole and there was no apparent colocalization of PfATG18 with the apicoplast; some PfATG18-labeled vesicles were detected near the branching apicoplast and in proximity of vesicles containing downstream autophagy protein PfATG8. PfATG18 interact with 3′-PIPs via the FRRG motif, and mutation of this motif resulted in a significant loss of its vesicular localization. PfATG18 having FRRG domain and binding potential towards PI3PK have speculated role in binding of Atg2 and lead to attachment of PfAtg18-PfAtg2 to PAS.
In other systems ATG18 interaction with PtdIns(3)P leads to attachment of ATG8 to phagophore and its elongation [45]. In Plasmodium, no direct interaction has been shown between PfATG8 and PfATG18. However, downregulation of PfATG18 leads to decrease in the number of PfATG8-positive vesicles under normal conditions [44]. A putative homologue of PfAtg2 has also been identified in the genome.
Atg8 and Atg12 Ubiquitin Conjugation System
Another important pathway for autophagy mechanism in yeast is ATG5-ATG12 pathway, which involves ubiquitin-like enzymes for its implication. Plasmodium contains homologues for Atg8, Atg3 and Atg7 from this branch of autophagy [46,47,48]. PfAtg8 remains the choice of molecule that has been targeted in Plasmodium to understand the autophagy and its role in the life cycle of parasite. Atg8 orthologues from most organisms require proteolytic processing of one or several amino acids to expose a C-terminal glycine. PfAtg8 differs from its mammalian homologue, in having a C-terminal glycine. This exposed C-terminal glycine is required for attaching Atg8 to its E1 ubiquitin enzyme Atg7. In mammals ATG4-based processing is required for making the C-terminal glycine for conjugation of ATG8 towards phosphatidylethanolamine (PE). Lipidated form of Atg8 has been suggested to be a marker of autophagy in mammalian system, as ATG8-PE remains conjugated with autophagosome during fusion to the lysosome [49]. In Plasmodium various studies tried to decipher the localization of PfAtg8. PfAtg8 has been targeted in Plasmodium to understand the autophagy and its role in the life cycle of parasite [43,44,45,46,47,48, 50, 51]. C-terminal glycine of PfAtg8 has an important role in the association of PfAtg8 with these punctate structures and apicoplast as removal of this C-terminal glycine leads to diffuse localization of this protein in the cytoplasm in liver stage [47]. Recently structure of PfAtg8-PfAtg3 has been solved, which suggest that Plasmodium Atg8 contains an insertion of nine residues only conserved within Apicomplexa that comprise β3 and the turn in the loop [35].
In yeast, lipidation of Atg8 requires E1-type ligase Atg7, E2-type ligase Atg3 and a cysteine protease Atg4. Plasmodium genome has all of these three ATG proteins, Atg3, Atg4 and Atg7, which are shown to be transcribed in blood stages of the parasite life cycle [52]. Processing of C-terminus of Atg8 by Atg4, followed by activation using ATP, allows it to interact with Atg7. Atg8 is further transferred to Atg3, an E2-like conjugating enzyme, leading to formation of second thioester bond followed by conjugation to nitrogen of PE. This process also requires noncovalent interaction between Atg8 and Atg3 through a well-characterized Atg8-interacting motif (AIM) in Atg3 and two hydrophobic pockets, termed the W- and L-site, in Atg8. In Plasmodium Atg3 also contains Atg8-interacting motif along with the WLLP residues which are responsible for the interaction of Atg3-Atg8 [35]. Atg3-Atg8 interaction has been targeted well in Plasmodium as possible site for the action of small molecule inhibitors [53].
Deconjugation of PfAtg8 from its binding sites for recycling has been carried out by other cysteine proteases, i.e. PfOTU [55]. Depletion of PfOTU levels also affects PfAtg8 conjugated forms and thus affects transport of apicoplast proteins. PfAtg8 localization to the punctate structure during normal conditions and relocalization during nutritional starvation conditions or during drug treatment point towards important role in autophagy mechanism in the Plasmodium [56]. PfAtg8-associated double-membrane vesicular structures are also shown to contain PfRab7 [57]. During nutritional starvation conditions, these double-membrane vesicular structures are shown to be near to the food vacuole, which points towards food vacuole as the final site of their localization [57].
Atg7 Protein Complex
The PfAtg7 shows low identity (14.7%) and similarity (32.2%) to yeast Atg7; however, it shows conservation of key residues, including the catalytic cysteine and ATP-binding domain [54]. Atg7 is an E1-type activating enzyme, which is a ubiquitin-related modifier. It follows the mechanism of protein ubiquitination in order to lipidate Atg8. During the process, a thioester intermediate is formed between the E1 (Atg7) and ubiquitin (Atg8). Ubiquitin (Atg8) is then transferred to the catalytic cysteine residue of the ubiquitin-conjugating enzyme or E2 (Atg3). The final step includes transfer of ubiquitin (Atg8) to its target protein (PE) forming a covalent bond through an isopeptide linkage. This can occur directly by the E2 or through a third ubiquitin-protein ligase or E3 (Atg5-Atg12). Putative homologue of ATG12 and Atg5 has also been identified in the host.
PtdIns3K Complex
Another signalling mechanism which plays a role in the initiation process of autophagy is class III PI-kinase-based pathway. Vesicular protein sorting 34 or vps34 is a class III PI-kinase which carries out initiation of autophagy in yeast in complex with Atg6 and Beclin1 [58]. Interaction of Beclin1 and Vps34 promotes activity of Vps34, which leads to higher production of phosphatidylinositol-3-phosphate (PtdIns3p) required for phagophore formation and elongation. Beclin1-Vps34 complex has further been joined by various different regulatory molecules which leads to either promotion of autophagy or its inhibition. UVRAG, ATG14L, BIF-1 and AMBRA are shown to promote autophagy when they interact with beclin1-vps34 complex [22, 59], whereas Rubicon and Bcl-2 lead to inhibition of autophagy [60, 61]. Interaction of Bcl-2 with Beclin leads to disruption of beclin1-vps34 complex and leads to inhibition of autophagy [61, 62]. Class III PtdIns3K complex is comprised of five distinct proteins: Vps34, the regulatory kinase Vps15, Vps30/Atg6, Atg14 and Atg38 [63, 64]. This complex is responsible for the production of PtdIns3p, which plays a very important role for the correct localization of other ATG proteins including Atg18 and Atg2 enabling the recruitment of Atg8, Atg9 and Atg12 of the pre-autophagosomal site [65].
Plasmodium genome harbours homologue of vps34, which is also termed as PfPI3K (ref). PfPI3K has been identified and found to be localized at various places ranging from digestive vacuole (DV), the parasite membrane and vesicles in host erythrocytes [66]. PfPI3K is mainly shown to be involved in haemoglobin uptake and degradation pathways, as inhibition of pfPI3K with wortmannin causes accumulation of unutilized haemoglobin [66]. Pfvps34 do not contain pleckstrin homology domains and Ras binding sites that are usual features of class I and class II PI3Ks. PfVps34 is an essential protein for the survival of the parasite [67]. The Pfvps34, along with process of haemoglobin uptake and degradation, had been hypothesized to be a key regulatory checkpoint in the autophagy-like cascade of P. falciparum [35, 51]. As cellular material is likely to be digested within the digestive in parasite, Pfvps34 appears to be key component of Plasmodium falciparum autophagy cascade in absence of TOR kinase activity [35].
Phagophore Expansion
Ubiquitin-like systems also play key role in the autophagy phagophore expansion process. Two major key ubiquitin pathways are Ag5-Atg12 complex and the LC3 processing pathway. ATG12 and ATG8 are the most important ubiquitin-like proteins in the process of autophagy. They have high similarity to the ubiquitin at the structural level, but they are not the real homologues. ATG5-ATG12 pathway starts with ATG7 binding to the carboxyl terminal of ATG12 activating the later one in an ATP-dependent manner. This activated ATG12 then has been transferred to ATG10. ATG10 acts like an E2-ubiquitin-like enzyme which carries out attachment of ATG12 to residue 130 lysine of ATG5 leading to formation of ATG12-Atg5 complex. This conjugated ATG5-ATG12 pairs with ATG16L dimers to form bigger multimers of ATG5-ATG12-ATG16L which leads to the extension of phagophore membrane. This growing multimer complex then further has been joined with LC3B-II, the product of second important ubiquitin system playing a role in autophagy in mammals. Microtubule-associated protein light chain 3 (LC3B) is full-length cytosolic protein, which has been acted upon by ATG4 which is a cysteine protease, leading to formation of LC3B-I, which contains a C-terminally exposed glycine residue. This exposed glycine residue has then been activated by ATG7 in an ATP-dependent manner. Activated LC3B-I is then transferred to Atg3; a different E2-like carrier protein before phosphatidylethanolamine (PE) is conjugated to the carboxyl glycine to generate processed LC3B-II.
PfAtg5 is a recently characterized ATG protein in Plasmodium [52]; it has been found to be associated with punctuate structures throughout the parasite cytosol, which may point towards presence of autophagosome-like structures near the mitochondria and ER as observed in mammalian systems [52]. The putative PfAtg5 is a much longer protein due to insertions as compared to its counterparts. The lysine amino acid residues present at position 479 are involved in conjugation with PfAtg12; in addition, a lysine residue is present at position 480.
Roles of Autophagy in Plasmodium
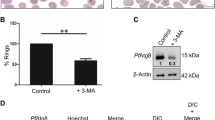
After being released in the blood after mosquito bites, sporozoite moves to the hepatic cells and tries to establish liver stage development of the parasite. During this process, sporozoite gets converted into metabolically active trophozoites. This whole metamorphosis involves lot of activity at the cellular level and involves changes in the shape of the parasite. Along with this shape changes, they start losing some of the organelles [68], and at the end of this metamorphosis, parasite is only left with organelles required for replication. The mechanism that underlies this process was speculated to be autophagy at the time, although much of ATGs identified in yeast have not been recognized in the parasite. Plasmodium cellular homoeostasis is a very important but complex mechanism in the life cycle of the parasite. Plasmodium sporozoites treated with 3-methyladenine (3-MA), which targets Vps34, needed for PI(3)P production and phagophore formation, delay conversion into trophozoites [69].
Autophagy in Protein Secretion and Trafficking
Protein secretion in Plasmodium is an important process which is required for its survival and pathogenesis. Parasite remodels the host and exports a number of proteins to the host cytoplasm and to the surface [70]. Plasmodium stays in the host encapsulated in the parasitophorous vacuole, from which parasite proteins have to travel across the PV and plasma membrane, which require novel pathways for secretion. Exophagy is one of the mechanisms through which these kinds of protein secretion are been observed in secretion of Acb1 protein in yeast, which utilizes autophagy mechanism involving GRASP65, t-SNARE and sso1 [71]. Plasmodium genome contains homologue of GRASP65 and t-SNARE [72], which points towards possibility of exophagy as part of transport mechanism in the parasite.
Another example regarding the role of autophagy in protein secretion comes from type III PI3K, e.g. vps34, a lipid kinase at the ER that produce PI(3)P on the outer leaflet, and recruit proteins to the phagophore and formation of autophagosome. Plasmodium homologue of the Vps34, PfVps34, from lysate has also been shown to have PI3K activity and can be inhibited by wortmannin class III PI3K. PfVps34 localizes to the food vacuole and different vesicular structures near the food vacuole and the plasma membrane in the blood stage [73]. Further, Vps34 inhibition in sporozoites delays its development to trophozoite [73]. The electron microscopy studies of liver stage parasite showed presence of double-membrane structures resembling autophagosomes having microneme point towards role of exophagy in protein secretion in the parasite.
PfAtg8 role in protein trafficking has also been highlighted in early-mid trophozoite, where it is suggested to be linked to haemoglobin uptake [57]. Chloroquine treatment to the early-mid trophozoite leads to increase in vesicles, which are assumed to be autophagosomic vesicles. Prolonged starvation or high cytocidal concentration of chloroquine (CQ) leads to relocalization of PfAtg8-containing vesicles from the parasite to host cytoplasm [51]. However, Cytostatic concentration of chloroquine (100 μm and 300 μM) and shorter treatment duration doesn’t show any relocalization of PfAtg8 to host cytoplasm [74]. PfAtg8-containing vesicles are also shown to harbour Rab7 molecule on their surfaces, which also points towards the role of PfAtg8 in protein trafficking [57].
Autophagy in Programmed Cell Death
In the early 2000s, studies about effect of drugs on the development on the parasite pointed out towards probable role of programmed cell death in the Plasmodium. Treatment of the parasite with chloroquine, S-nitroso-N-acetylpenicillamine (SNAP) or staurosporine showed presence of apoptosis-like features during the death of the parasite [75]. In absence of canonical apoptosis pathway in the parasite, these features of death lead to investigate the role of other mechanisms in the parasite life cycle. Programmed cell death in unicellular organisms has not been supported earlier as the apoptotic pathway machinery is lacking as has been shown in yeast [76, 77]. Although presence of metacaspases in some of the unicellular eukaryotes including Plasmodium suggested a primordial form of apoptosis to exist, the cannonical pathway of apoptosis remains missing [78].
It is suggested that Plasmodium berghei invasion of hepatocytes involves death of a lot of parasites through autophagic mechanism [79]. This phenomenon was observed in mice model and thus seems to be a physiological mechanism of host defence. This phenomenon is seen to be associated with development of later stages of parasite in which double-membrane structures have been observed which are termed as “membranous whorls” [80]. PbAtg8 does not localize with these structures and remains associated with apicoplast with no changes in localization [9]. Another interesting observation is that treatment with rapamycin, a known TOR kinase inhibitor which has been used to induce autophagy in yeast, leads to decrease in parasite size and morphological changes in apicoplast in Plasmodium.
Host Autophagy Machinery During Plasmodium Life Cycle
During the life cycle, parasite transits between two hosts, mosquito and human. In human host the erythrocyte doesn’t play any role in the control of the parasite infection in blood stage; however, liver cells employ its host cell autophagy machinery to stop the infection [87]. During its stay in the liver cell, Plasmodium parasite is protected from its host by parasite vacuole membrane (PVM). PVM has been generated from host cell membrane during invasion but has been modified by parasite using its own protein [81,82,83]. Some of these proteins, e.g. UIS3 and UIS4, have been shown to be important in the survival of the parasite in the liver cell. Disruption of UIS3 and UIS4 showed to cause growth arrest of the parasite [84]. For a long time, liver stage infection of the malaria parasite has been considered as “silent stage”. It is becoming clear now that host cell can sense the parasite and take care of it very effectively using autophagy [79]. Induction of canonical nonselective autophagy with rapamycin or starvation enhances the development of parasite in hepatocytes and leads to increase in the number of liver stage parasite [85]. In addition, if host autophagy machinery is disrupted genetically, the parasite development is affected in liver stage [85]; however, there are contradictory reports as well, which may have arisen due to different cell types used in the experiment [79]. These different opinions on the role of host autophagy in Plasmodium led to intense discussion whether it is an advantage or disadvantage regarding the malaria infection [87]. Recent views on liver stage development could represent a non-canonical form of autophagy, termed as Plasmodium Associated Autophagic-like Response (PAAR) [85,86,87].
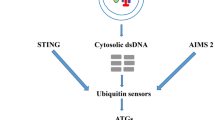
The process and the proteins involved in host autophagy during the hepatic stages of Plasmodium are well elucidated. One of the signature processes of parasite liver stage development is the rapid acquisition of host LC3 and other interacting proteins such as P62, NBR1, NDP52 and ubiquitin on the PVM [79]. The process of LC3 acquisition is so rapid that it suggests that either the parasite hijacks the host autophagy machinery or it is readily sensed by the host. Association of LC3 with the PVM itself has some striking aspects, such as LC3 decoration on the PVM does not involve the formation of new canonical double-membrane autophagosome, as it has been observed in other systems, rather LC3 associates with the existing PVM. This is also different from LC3-associated phagocytosis phenomenon as sporozoite invasion to the liver cells is an active process which is different from conventional phagocytosis. Another aspect which is different from canonical xerophagy pathway is the association of the LC3-binding proteins, including ubiquitin, to the PVM that is to a large extent directly mediated by LC3 [79]. Recruitment of LC3-binding proteins to the PVM appears to be in reverse order leading to the idea of an “inverted” recruitment of LC3-associated proteins on the PVM [76]. Further LC3 association with the PVM is temporary [85], and it gets dissociated in later stages, which is shown to be important for the proper development of the parasite [85]. LC3 recruitment to the PVM is governed by lipidation suggesting the role of conjugation machinery involving upstream ATGs such as ATG5. However, initiation complexes of autophagy such as FIP200 are not required [83].
The pathway and the factors that initiate the conjugation of LC3 to the PVM due to Plasmodium infection are not known in detail. UIS3 binds directly to and retains LC3 on the PVM as evident by uis3(−) parasite, which are arrested in development in wild-type hepatocytes. These uis3(−) parasite develop normally in ATG5−/-MEFs, pointing towards central role of UIS3 in interaction with host autophagy machinery (Table 1).
References
White, N. J. (2014). Malaria: A molecular marker of artemisinin resistance. Lancet (London, England) [Internet]., 383(9927), 1439–1440. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24766952.
Ockenhouse, C. F., Schulman, S., Shear, H. L., et al. (1984). Journal of Immunology [Internet], 133(3), 1601–1608. Available from: http://www.ncbi.nlm.nih.gov/pubmed/6431003.
Jensen, J. B., Boland, M. T., & Akood, M. (1982). Induction of crisis forms in cultured Plasmodium falciparum with human immune serum from Sudan. Science [Internet], 216(4551), 1230–1233. Available from: http://www.ncbi.nlm.nih.gov/pubmed/7043736.
Picot, S., Burnod, J., Bracchi, V., Chumpitazi, B. F., & Ambroise-Thomas, P. Apoptosis related to chloroquine sensitivity of the human malaria parasite Plasmodium falciparum. Transactions of The Royal Soceity Tropical Medicine Hygiene [Internet], 91(5), 590–591. Available from: http://www.ncbi.nlm.nih.gov/pubmed/9463676.
Rathore, S., Datta, G., Kaur, I., Malhotra, P., Mohmmed, A., et al. (2015). Cell Death and Disease [Internet], 6, e1803. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26136076.
Rathore, S., Jain, S., Sinha, D., Gupta, M., Asad, M., Srivastava, A., et al. (2011). Disruption of a mitochondrial protease machinery in Plasmodium falciparum is an intrinsic signal for parasite cell death. Cell Death and Disease [Internet], 2, e231. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22113196.
Jain, S., Rathore, S., Asad, M., Hossain, M. E., Sinha, D., Datta, G., et al. (2013). The prokaryotic ClpQ protease plays a key role in growth and development of mitochondria in Plasmodium falciparum. Cellular Microbiology [Internet], 15(10), 1660–1673. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23521916.
Denton, D., Aung-Htut, M. T., Lorensuhewa, N., Nicolson, S., Zhu, W., Mills, K., et al. (2013). UTX coordinates steroid hormone-mediated autophagy and cell death. Nature Communications [Internet], 4, 2916. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24336022.
Voss, C., Ehrenman, K., Mlambo, G., Mishra, S., Kumar, K. A., Sacci, J. B., et al. (2016). Overexpression of Plasmodium berghei ATG8 by liver forms leads to cumulative defects in organelle dynamics and to generation of noninfectious Merozoites. MBio [Internet], 7(3). Available from http://www.ncbi.nlm.nih.gov/pubmed/27353755.
Betin, V. M. S., Singleton, B. K., Parsons, S. F., Anstee, D. J., & Lane, J. D. (2013). Autophagy facilitates organelle clearance during differentiation of human erythroblasts: Evidence for a role for ATG4 paralogs during autophagosome maturation. Autophagy [Internet], 9(6), 881–893. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23508006.
Klionsky, D. J., Cregg, J. M., Dunn, W. A., Emr, S. D., Sakai, Y., Sandoval, I. V., et al. (2003). A unified nomenclature for yeast autophagy-related genes. Developmental Cell [Internet], 5(4), 539–545. Available from: http://www.ncbi.nlm.nih.gov/pubmed/14536056.
De Duve, C., Pressman, B. C., Gianetto, R., Wattiaux, R., & Appelmans, F. (1955). Tissue fractionation studies. 6. Intracellular distribution patterns of enzymes in rat-liver tissue. The Biochemical Journal [Internet], 60(4), 604–617. Available from: http://www.ncbi.nlm.nih.gov/pubmed/13249955.
Novikoff, A. B. (1955). Some aspects of hepatoma NK. Journal of the National Cancer Institute [Internet], 15(5, Suppl), 1533–1534. Available from: http://www.ncbi.nlm.nih.gov/pubmed/13243091.
Clark, S. L. (1957). Cellular differentiation in the kidneys of newborn mice studies with the electron microscope. The Journal of Biophysical and Biochemical Cytology [Internet], 3(3), 349–362. Available from: http://www.ncbi.nlm.nih.gov/pubmed/13438920.
He, C., & Klionsky, D. J. (2009). Regulation mechanisms and signaling pathways of autophagy. Annual Review of Genetics [Internet], 43, 67–93. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19653858.
Beau, I., Esclatine, A., & Codogno, P. (2008). Lost to translation: When autophagy targets mature ribosomes. Trends in Cell Biology [Internet], 18(7), 311–314. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18508269.
Kraft, C., Reggiori, F., & Peter, M. (2009). Selective types of autophagy in yeast. Biochimica et Biophysica Acta [Internet], 1793(9), 1404–1412. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19264099.
van der Vaart, A., Mari, M., & Reggiori, F. (2008). A picky eater: Exploring the mechanisms of selective autophagy in human pathologies. Traffic [Internet], 9(3), 281–289. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17988219.
Gordy, C., & He, Y.-W. (2012). The crosstalk between autophagy and apoptosis: Where does this lead? Protein and Cell [Internet], 3(1), 17–27. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22314807.
De Duve, C., & Wattiaux, R. (1966). Functions of lysosomes. Annual Review of Physiology [Internet], 28, 435–492. Available from: http://www.ncbi.nlm.nih.gov/pubmed/5322983.
Novikoff, A. B., & Shin, W. Y. (1978). Endoplasmic reticulum and autophagy in rat hepatocytes. Proceedings of the National Academy of Sciences of United States America [Internet], 75(10), 5039–5042. Available from: http://www.ncbi.nlm.nih.gov/pubmed/283412.
Liang, C., Feng, P., Ku, B., Dotan, I., Canaani, D., Oh, B.-H., et al. (2006). Autophagic and tumour suppressor activity of a novel Beclin1-binding protein UVRAG. Nature Cell Biology [Internet], 8(7), 688–699. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16799551.
Deter, R. L., Baudhuin, P., De Duve, C., et al. (1967). The Journal of Cell Biology [Internet], 35(2), C11–C16. Available from: http://www.ncbi.nlm.nih.gov/pubmed/6055998.
Dunn, W. A. (1994). Autophagy and related mechanisms of lysosome-mediated protein degradation. Trends Cell Biol [Internet], 4(4), 139–143. Available from: http://www.ncbi.nlm.nih.gov/pubmed/14731737.
Levine, B. (2005). Eating oneself and uninvited guests: Autophagy-related pathways in cellular defense. Cell [Internet], 120(2), 159–162. Available from: http://www.ncbi.nlm.nih.gov/pubmed/15680321.
Hayashi-Nishino, M., Fujita, N., Noda, T., Yamaguchi, A., Yoshimori, T., & Yamamoto, A. (2009). A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nature Cell Biology [Internet], 11(12), 1433–1437. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19898463.
Ylä-Anttila, P., Vihinen, H., Jokitalo, E., & Eskelinen, E.-L. (2009). 3D tomography reveals connections between the phagophore and endoplasmic reticulum. Autophagy [Internet], 5(8), 1180–1185. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19855179.
Axe, E. L., Walker, S. A., Manifava, M., Chandra, P., Roderick, H. L., Habermann, A., et al. (2008). Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. Journal of Cell Biology [Internet], 182(4), 685–701. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18725538.
Mizushima, N. (2007). The role of mammalian autophagy in protein metabolism. Proceedings of the Japan Academy Series B Physical and Biology Sciences [Internet], 83(2), 39–46. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24019583.
English, L., Chemali, M., Duron, J., Rondeau, C., Laplante, A., Gingras, D., et al. (2009). Autophagy enhances the presentation of endogenous viral antigens on MHC class I molecules during HSV-1 infection. Nature Immunology [Internet], 10(5), 480–487. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19305394.
Fennell, B. J., Al-shatr, Z. A., & Bell, A. (2008). Isotype expression, post-translational modification and stage-dependent production of tubulins in erythrocytic Plasmodium falciparum. International Journal Parasitology [Internet], 38(5), 527–539. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17977543.
Williams, A., Sarkar, S., Cuddon, P., Ttofi, E. K., Saiki, S., Siddiqi, F. H., et al. (2008). Novel targets for Huntington’s disease in an mTOR-independent autophagy pathway. Nature Chemical Biology [Internet], 4(5), 295–305. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18391949.
Díaz-Troya, S., Pérez-Pérez, M. E., Florencio, F. J., & Crespo, J. L. (2008). The role of TOR in autophagy regulation from yeast to plants and mammals. Autophagy [Internet], 4(7), 851–865. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18670193.
Kundu, M., Lindsten, T., Yang, C.-Y., Wu, J., Zhao, F., Zhang, J., et al. (2008). Ulk1 plays a critical role in the autophagic clearance of mitochondria and ribosomes during reticulocyte maturation. Blood [Internet], 112(4), 1493–1502. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18539900.
Hain, A. U. P., & Bosch, J. (2013). Autophagy in Plasmodium, a multifunctional pathway? Computational and Structural Biotechnology Journal [Internet], 8, e201308002. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24688742.
Kraft, C., Kijanska, M., Kalie, E., Siergiejuk, E., Lee, S. S., Semplicio, G., et al. (2012). Binding of the Atg1/ULK1 kinase to the ubiquitin-like protein Atg8 regulates autophagy. EMBO Journal [Internet], 31(18), 3691–3703. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22885598.
Noda, T., Kim, J., Huang, W. P., Baba, M., Tokunaga, C., Ohsumi, Y., et al. (2000). Apg9p/Cvt7p is an integral membrane protein required for transport vesicle formation in the Cvt and autophagy pathways. Journal of Cell Biology [Internet], 148(3), 465–480. Available from: http://www.ncbi.nlm.nih.gov/pubmed/10662773.
Reggiori, F., Shintani, T., Nair, U., & Klionsky, D. J. (2005). Atg9 cycles between mitochondria and the pre-autophagosomal structure in yeasts. Autophagy [Internet], 1(2), 101–109. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16874040.
Suzuki, K., Kubota, Y., Sekito, T., & Ohsumi, Y. (2007). Hierarchy of Atg proteins in pre-autophagosomal structure organization. Genes to Cells [Internet], 12(2), 209–218. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17295840.
Wang, C. W., Kim, J., Huang, W. P., Abeliovich, H., Stromhaug, P. E., Dunn, W. A., et al. (2001). Apg2 is a novel protein required for the cytoplasm to vacuole targeting, autophagy, and pexophagy pathways. Journal of Biology Chemistry [Internet], 276(32), 30442–30451. Available from: http://www.ncbi.nlm.nih.gov/pubmed/11382760.
Reggiori, F., Wang, C.-W., Nair, U., Shintani, T., Abeliovich, H., & Klionsky, D. J. (2004). Early stages of the secretory pathway, but not endosomes, are required for Cvt vesicle and autophagosome assembly in Saccharomyces cerevisiae. Molecular Biology of the Cell [Internet], 15(5), 2189–2204. Available from: http://www.ncbi.nlm.nih.gov/pubmed/15004240.
Strømhaug, P. E., Reggiori, F., Guan, J., Wang, C.-W., & Klionsky, D. J. (2004). Atg21 is a phosphoinositide binding protein required for efficient lipidation and localization of Atg8 during uptake of aminopeptidase I by selective autophagy. Molecular Biology of the Cell [Internet], 15(8), 3553–3566. Available from: http://www.ncbi.nlm.nih.gov/pubmed/15155809.
Reggiori, F., Tucker, K. A., Stromhaug, P. E., & Klionsky, D. J. (2004). The Atg1-Atg13 complex regulates Atg9 and Atg23 retrieval transport from the pre-autophagosomal structure. Developmental Cell [Internet], 6(1), 79–90. Available from: http://www.ncbi.nlm.nih.gov/pubmed/14723849.
Bansal, P., Tripathi, A., Thakur, V., Mohmmed, A., & Sharma, P. (2017). Autophagy-related protein ATG18 regulates Apicoplast biogenesis in apicomplexan parasites. MBio [Internet], 8(5). Available from: http://www.ncbi.nlm.nih.gov/pubmed/29089429.
Papinski, D., & Kraft, C. (2014). Atg1 kinase organizes autophagosome formation by phosphorylating Atg9. Autophagy [Internet], 10(7), 1338–1340. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24905091.
Kitamura, K., Kishi-Itakura, C., Tsuboi, T., Sato, S., Kita, K., Ohta, N., et al. (2012). Autophagy-related Atg8 localizes to the apicoplast of the human malaria parasite Plasmodium falciparum. PLoS One [Internet], 7(8), e42977. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22900071.
Eickel, N., Kaiser, G., Prado, M., Burda, P.-C., Roelli, M., Stanway, R. R., et al. (2013). Features of autophagic cell death in Plasmodium liver-stage parasites. Autophagy [Internet], 9(4), 568–580. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23388496.
Jayabalasingham, B., Voss, C., Ehrenman, K., Romano, J. D., Smith, M. E., Fidock, D. A., et al. (2014). Characterization of the ATG8-conjugation system in 2 Plasmodium species with special focus on the liver stage: Possible linkage between the apicoplastic and autophagic systems? Autophagy [Internet], 10(2), 269–284. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24342964.
Xie, Z., Nair, U., & Klionsky, D. J. (2008). Dissecting autophagosome formation: The missing pieces. Autophagy [Internet], 4(7), 920–922. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18719358.
Cervantes, S., Bunnik, E. M., Saraf, A., Conner, C. M., Escalante, A., Sardiu, M. E., et al. (2014). The multifunctional autophagy pathway in the human malaria parasite, Plasmodium falciparum. Autophagy [Internet], 10(1), 80–92. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24275162.
Gaviria, D., Paguio, M. F., Turnbull, L. B., Tan, A., Siriwardana, A., Ghosh, D., et al. (2013). A process similar to autophagy is associated with cytocidal chloroquine resistance in Plasmodium falciparum. PLoS One [Internet], 8(11), e79059. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24278114.
Joy, S., Thirunavukkarasu, L., Agrawal, P., Singh, A., Sagar, B. K. C., Manjithaya, R., et al. (2018). Basal and starvation-induced autophagy mediates parasite survival during intraerythrocytic stages of Plasmodium falciparum. Cell Death Discovery [Internet], 4, 43. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30302277.
Hain, A. U. P., Bartee, D., Sanders, N. G., Miller, A. S., Sullivan, D. J., Levitskaya, J., et al. (2014). Identification of an Atg8-Atg3 protein-protein interaction inhibitor from the medicines for malaria venture malaria box active in blood and liver stage Plasmodium falciparum parasites. Journal of Medicinal Chemistry [Internet]., 57(11), 4521–4531. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24786226.
Walker, D. M., Mahfooz, N., Kemme, K. A., Patel, V. C., Spangler, M., & Drew, M. E. (2013). Plasmodium falciparum erythrocytic stage parasites require the putative autophagy protein PfAtg7 for normal growth. PLoS One [Internet], 8(6), e67047. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23825614.
Datta, G., Hossain, M. E., Asad, M., Rathore, S., & Mohmmed, A. (2017). Plasmodium falciparum OTU-like cysteine protease (PfOTU) is essential for apicoplast homeostasis and associates with noncanonical role of Atg8. Cell Microbiology [Internet], 19(9), e12748. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28423214.
Sinai, A. P., & Roepe, P. D. (2012). Autophagy in Apicomplexa: A life sustaining death mechanism? Trends in Parasitology [Internet], 28(9), 358–364. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22819059.
Tomlins, A. M., Ben-Rached, F., Williams, R. A., Proto, W. R., Coppens, I., Ruch, U., et al. (2013). Plasmodium falciparum ATG8 implicated in both autophagy and apicoplast formation. Autophagy [Internet], 9(10), 1540–1552. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24025672.
Backer, J. M. (2008). The regulation and function of class III PI3Ks: Novel roles for Vps34. Biochemical Journal [Internet], 410, 1):1–1)17. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18215151.
Fimia, G. M., Stoykova, A., Romagnoli, A., Giunta, L., Di Bartolomeo, S., Nardacci, R., et al. (2007). Ambra1 regulates autophagy and development of the nervous system. Nature [Internet], 447(7148), 1121–1125. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17589504.
Matsunaga, K., Saitoh, T., Tabata, K., Omori, H., Satoh, T., Kurotori, N., et al. (2009). Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nature Cell Biology [Internet], 11(4), 385–396. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19270696.
Pattingre, S., Tassa, A., Qu, X., Garuti, R., Liang, X. H., Mizushima, N., et al. (2005). Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell [Internet], 122(6), 927–939. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16179260.
Maiuri, M. C., Zalckvar, E., Kimchi, A., & Kroemer, G. (2007). Self-eating and self-killing: Crosstalk between autophagy and apoptosis. Nature Reviews Molecular Cell Biology [Internet], 8(9), 741–752. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17717517.
Schu, P. V., Takegawa, K., Fry, M. J., Stack, J. H., Waterfield, M. D., & Emr, S. D. (1993). Phosphatidylinositol 3-kinase encoded by yeast VPS34 gene essential for protein sorting. Science [Internet], 260(5104), 88–91. Available from: http://www.ncbi.nlm.nih.gov/pubmed/8385367.
Kihara, A., Noda, T., Ishihara, N., & Ohsumi, Y. (2001). Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. Journal of Cell Biology [Internet], 152(3), 519–530. Available from: http://www.ncbi.nlm.nih.gov/pubmed/11157979.
Obara, K., & Ohsumi, Y. (2008). Dynamics and function of PtdIns(3)P in autophagy. Autophagy [Internet], 4(7), 952–954. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18769109.
Vaid, A., Thomas, D. C., & Sharma, P. (2008). Role of Ca2+/calmodulin-PfPKB signaling pathway in erythrocyte invasion by Plasmodium falciparum. Journal of Biological Chemistry [Internet], 283(9), 5589–5597. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18165240.
Tawk, L., Chicanne, G., Dubremetz, J.-F., Richard, V., Payrastre, B., Vial, H. J., et al. (2010). Phosphatidylinositol 3-phosphate, an essential lipid in Plasmodium, localizes to the food vacuole membrane and the apicoplast. Eukaryotic Cell [Internet], 9(10), 1519–1530. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20709789.
Jayabalasingham, B., Bano, N., & Coppens, I. (2010). Metamorphosis of the malaria parasite in the liver is associated with organelle clearance. Cell Research [Internet], 20(9), 1043–1059. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20567259.
Coppens, I. (2011). Metamorphoses of malaria: The role of autophagy in parasite differentiation. Essays in Biochemistry [Internet], 51, 127–136. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22023446.
Rug, M., Cyrklaff, M., Mikkonen, A., Lemgruber, L., Kuelzer, S., Sanchez, C. P., et al. (2014). Export of virulence proteins by malaria-infected erythrocytes involves remodeling of host actin cytoskeleton. Blood [Internet], 124(23), 3459–3468. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25139348.
Manjithaya, R., Anjard, C., Loomis, W. F., & Subramani, S. (2010). Unconventional secretion of Pichia pastoris Acb1 is dependent on GRASP protein, peroxisomal functions, and autophagosome formation. Journal of Cell Biology [Internet], 188(4), 537–546. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20156962.
Ayong, L., Pagnotti, G., Tobon, A. B., & Chakrabarti, D. (2007). Identification of Plasmodium falciparum family of SNAREs. Molecular and Biochemical Parasitology [Internet], 152(2), 113–122. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17240462.
Hassett, M. R., & Roepe, P. D. (2018). PIK-ing new malaria chemotherapy. Trends in Parasitology [Internet], 34(11), 925–927. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29934102.
Navale, R., Atul, A. A. D., & Sijwali, P. S. (2014). Characterization of the autophagy marker protein Atg8 reveals atypical features of autophagy in Plasmodium falciparum. PLoS One [Internet], 9(11), e113220. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25426852.
Totino, P. R. R., Daniel-Ribeiro, C. T., Corte-Real, S., de Fátima Ferreira-da-Cruz, M., et al. (2008). Experimental Parasitology [Internet], 118(4), 478–486. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18226811.
Carmona-Gutierrez, D., Ruckenstuhl, C., Bauer, M. A., Eisenberg, T., Büttner, S., & Madeo, F. (2010). Cell death in yeast: Growing applications of a dying buddy. Cell Death and Differentiation [Internet], 17(5), 733–734. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20383156.
Madeo, F., Eisenberg, T., & Kroemer, G. (2009). Autophagy for the avoidance of neurodegeneration. Genes and Development [Internet], 23(19), 2253–2259. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19797764.
Meslin, B., Beavogui, A. H., Fasel, N., & Picot, S. (2011). Plasmodium falciparum metacaspase PfMCA-1 triggers a z-VAD-fmk inhibitable protease to promote cell death. PLoS One [Internet], 6(8), e23867. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21858231.
Schmuckli-Maurer, J., Reber, V., Wacker, R., Bindschedler, A., Zakher, A., & Heussler, V. T. (2017). Inverted recruitment of autophagy proteins to the Plasmodium berghei parasitophorous vacuole membrane. PLoS One [Internet], 12(8), e0183797. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28841718.
Stewart, M. J., Schulman, S., & Vanderberg, J. P. (1985). Rhoptry secretion of membranous whorls by Plasmodium berghei sporozoites. Journal of Protozoology [Internet], 32(2), 280–283. Available from: http://www.ncbi.nlm.nih.gov/pubmed/3925131.
Meis, J. F., Verhave, J. P., Jap, P. H., Sinden, R. E., Meuwissen, J. H., et al. (1983). Journal of Protozoology [Internet], 30(2), 361–366. Available from: http://www.ncbi.nlm.nih.gov/pubmed/6355454.
Lingelbach, K., & Joiner, K. A. (1998). The parasitophorous vacuole membrane surrounding Plasmodium and toxoplasma: An unusual compartment in infected cells. Journal of Cell Science [Internet], 111(Pt 1), 1467–1475. Available from: http://www.ncbi.nlm.nih.gov/pubmed/9580555.
Nyboer, B., Heiss, K., Mueller, A.-K., & Ingmundson, A. (2018). The Plasmodium liver-stage parasitophorous vacuole: A front-line of communication between parasite and host. International Journal of Medical Microbiology [Internet], 308(1), 107–117. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28964681.
Real, E., Rodrigues, L., Cabal, G. G., Enguita, F. J., Mancio-Silva, L., Mello-Vieira, J., et al. (2018). Plasmodium UIS3 sequesters host LC3 to avoid elimination by autophagy in hepatocytes. Nature Microbiology [Internet], 3(1), 17–25. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29109477.
Prado, M., Eickel, N., De Niz, M., Heitmann, A., Agop-Nersesian, C., Wacker, R., et al. (2015). Long-term live imaging reveals cytosolic immune responses of host hepatocytes against Plasmodium infection and parasite escape mechanisms. Autophagy [Internet], 11(9), 1561–1579. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26208778.
Wacker, R., Eickel, N., Schmuckli-Maurer, J., Annoura, T., Niklaus, L., Khan, S. M., et al. (2017). LC3-association with the parasitophorous vacuole membrane of Plasmodium berghei liver stages follows a noncanonical autophagy pathway. Cellular Microbiology [Internet], 19(10). Available from: http://www.ncbi.nlm.nih.gov/pubmed/28573684.
Coppens, I. (2017). How toxoplasma and malaria parasites defy first, then exploit host autophagic and endocytic pathways for growth. Current Opinionin Microbiology [Internet], 40, 32–39. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29102900.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Muneer, A., Singh, S., Narwal, M., Malhotra, P., Mohmmed, A., Rathore, S. (2019). Cellular Homoeostasis and Cell Signalling in Malaria Parasite: Role of Autophagy. In: Hameed, S., Fatima, Z. (eds) Pathogenicity and Drug Resistance of Human Pathogens. Springer, Singapore. https://doi.org/10.1007/978-981-32-9449-3_11
Download citation
DOI: https://doi.org/10.1007/978-981-32-9449-3_11
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-32-9448-6
Online ISBN: 978-981-32-9449-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)