Abstract
The collection of microorganisms that inhabits the human gastrointestinal tract is usually called the gut microbiota. The gut microbiota is an important component for the development and function of the immune system, operating as a complex of microorganisms that produce substances that interact with the immune cells and respond to internal and external stimuli in the body. The gut microbiota has a different composition in healthy individuals and those who have a disease suggesting that it can be a disease marker. It is also suggested to educate the host immune response and keep homeostasis through sophisticated microbial cross-talk with the mucosal immune system that includes huge integrated signaling pathways and gene regulatory circuits. The imbalance of these delicate interactions between microbiota and immune cells is associated with the development not only of inflammatory diseases but of also several diseases such as neurological, autoimmune disease, and metabolic syndrome. Therefore, a better understanding becomes vital for comprehending the factors linked with the development and/or occurrence of these disorders. This chapter focuses on the current findings of the role of gut microbiota in the activation and function of immune cells and how this relation modulates homeostasis and health disorders associated with microbiome dysbiosis. Moreover, we point up new nanotechnology therapies concerning manipulating the microbiome for the management of microbiota alterations-related human disease, giving and discussing future challenges and the perspective for this emerging area.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
8.1 Introduction
Our body is endowed with the ability to respond to external environmental signals while maintaining host’s homeostasis. This task is particularly relevant in sites with closer contact with the external world. The gut microbiome is a reunion of all microorganisms inhabiting the intestine, such as fungi, viruses, and bacteria. The gut microbiota, in this case, is related to all the different bacteria that are somehow living in the intestine. It is appreciated now that gut microbiota composition as well as its product produced by different pathways are important to maintaining tissue homeostasis. Any perturbation either in its composition or in the substances released by the microbiota directly impacts gut-residing immune cells, triggers inflammation, and, thus, these microbiota factors have been associated with different inflammatory diseases, such as metabolic syndrome, autoimmune diseases, infectious diseases, and cancer. In this chapter, we summarize the findings reporting the cross-talk between gut microbiota and the activation and stimulation of immune cells in the gut and the potential association with inflammatory diseases. Moreover, we highlight the recent nanotechnology approaches focused on gut microbiota manipulation to treat intra- and extraintestinal diseases.
8.2 Gut Microbiome
The human gut microbiome, or gut microbiota, refers to the assembly of microorganisms including, archaea, fungi, bacteria, and even viruses that coexist in the digestive tract (Arumugam et al. 2011). A growing number of studies demonstrates that the microbiota is an important manager of physiological, metabolic, and protective functions (Schluter et al. 2020).
In healthy conditions, the gut microbiota composition is predominantly composed of six main phyla; Firmicutes (Lactobacillus, Bacillus, Clostridium, Enterococcus, and Ruminicoccus genera) and Bacteroidetes (Bacteroides and Prevotella genera) making up about 90% of gut microbiota and the remainders are constituted by Actinobacteria, Proteobacteria, Fusobacteria, and Verrucomicrobia (Krajmalnik-Brown et al. 2012). Diversity in the microbiota composition seems to be important for health since recent studies have shown that a high diversity rate and high microbial gene richness indicate a healthy gut microbiota (Lloyd-Price et al. 2016). Moreover, by responding to several environmental stimuli such as diet, lifestyle, and medication, the gut microbiota composition is constantly challenged and susceptible to rapid change in its composition (Fig. 8.1) (David et al. 2014; Radjabzadeh et al. 2020) which may persist and impact over generations (Sonnenburg et al. 2016).
Factors that influence the development, diversity, and composition of gut microbiota. The influences of exogenous (exercise, nutrition, geography, drugs therapy, disease) and endogenous (aging, genetics, pregnancy) factors upon gut microbiota. Arrows indicate interactions that might occur between the gut microbiota and a particular factor
The type and quantities of food consumed affect the composition and function of gut microbiota, indicating that a diverse diet modulates the composition and functions of bacteria in the intestine (Maurice et al. 2013). For instance, recent studies have shown that a diet high in fat and sugar causes shifts in gut microbiota that may be correlated to the increase in diseases such as type 2 diabetes, obesity, and inflammatory bowel diseases (IBD) (Charbonneau et al. 2016; Zhu and Goodarzi 2020). As well with the increase in Western diet consumption, modern dietary pattern characterized by high intakes of ultra-processed foods, for instance pre-packaged food, can trigger major distresses of the gut microbiota, inducing an abnormal gut microbiota profile with consequences for well-being that are not always well comprehended (Wu et al. 2017). Therefore, the obvious interrelationship between the diet and how it modulates microbiota and affects the host still demand studies that in the future may lead to dietary therapeutic applications.
Besides, drug therapy such as antibiotics, painkillers, and diabetes medication also impacts gut microbiota. It is well established now that antibiotics use act not only on pathogenic bacteria that cause infections but also affect the resident commensals, diminishing levels of bacterial diversity and changing relative abundances, and in some cases, leading to gut microbiota dysbiosis-associated diseases (Elvers et al. 2020). For instance, metformin changes microbiota composition both in vitro and in vivo, increasing short-chain fatty acid (SCFA)-producing bacteria (Uranga et al. 2016). As well, proton pump inhibitor drugs increased the number of bacteria typically found in the oral tract in the gut (Imhann et al. 2017). Besides that, the indiscriminate use of antibiotics can lead to antibiotic-resistant bacteria present in the intestinal lumen, and, thus, can enter into the bloodstream of vulnerable patients, resulting in disseminated infection. Antibiotic therapy results in changes in intestinal microbiota composition that diminishes the resistance to colonization by antibiotic-resistant bacteria in the intestinal lumen (Keith and Pamer 2019).
8.2.1 Dialog Between Gut Microbiota and the Immune System in Homeostasis
One of the known benefits of the gut microbiome to the host is the contribution to the development of the immune system (Fig. 8.2). This concept comes from germ-free (GF) studies that revealed that these animals do not present a fully functional immune system (Round and Mazmanian 2009). By being generated in a sterile environment, and, thus, having never ever been exposed to any bug, GF mice are a valuable tool to study the impact of gut microbiota in different host conditions. The immune system is also involved in shaping and preserving the microbiota community (Round and Mazmanian 2009). Antibiotic-treated mice that eliminate bacteria in the gut exhibited an impaired immune response (Lazar et al. 2018). Importantly, dysbiosis in gut microbiota may trigger an exacerbated immune response, culminating with the development of allergies, inflammatory, or autoimmune diseases (Lazar et al. 2018).
Gut microbiota shape innate and adaptive immune cells. Some of the ways that the intestinal microbiome modulates host immunity are illustrated, including effects on innate and adaptive immune cells. IECS intestinal epithelial cells, IgA immunoglobulin A, IL-10 interleukin 10, NK cells natural killer cells, NKp46 natural cytotoxicity receptor, NOD1 nucleotide-binding oligomerization domain containing 1, SFB segmented filamentous bacteria, T reg cells regulatory T cells
8.2.1.1 Interactions Between the Innate Immune System and the Microbiota
Numerous reports provide direct evidence of relevant protagonist for the gut microbiota in regulating the development of macrophages, neutrophils, conventional natural killers (NK) cells (Khosravi et al. 2014; Luo et al. 2015; Smythies et al. 2005) and hematopoiesis (Shi et al. 2011). For instance, antibiotic therapy decreases bone marrow granulocyte-macrophage colony formation in animal models, and also GF animals present a deficiency in innate immune cells (Goris et al. 1985; Maslowski et al. 2009). Thus, the gut microbiota has been shown to modulate the innate immune response.
A critical feature for innate immune cells in the intestine, mainly antigen-presenting cells (APCs), is their capacity to protect the body against possible infections and, at the same time, to maintain a tolerogenic state to the normal gut microbiota (Imaoka et al. 1996). Indeed, the gut microbiota modulates the development of APCs. In GF animals were observed a reduction in the number of intestinal but not in the systemic DCs. Notably, monocolonization of GF mice with Escherichia coli led to the recruitment of DC to the intestine (Iwasaki and Kelsall 1999; Smythies et al. 2005). Furthermore, peritoneal macrophages of GF mice present an impairment of chemotaxis, phagocytosis, and microbial activities (Mikkelsen et al. 2004) besides displaying a lack of activation markers, such as MHC II (Wu and Wu 2012), once again demonstrating the impact of the gut microbiota on immune cell function at distant organs. Interestingly, CX3CR1hi mononuclear phagocytes, an intestinal cell population, are capable of trafficking antigen bacteria from the intestinal lumen to mesenteric lymph nodes modulating gut barrier repair and immune response, such as T cell responses and IgA production. Inhibition of this capture pathway through a MyD88-dependent mechanism restricts immune priming against intestinal antigens and can be a mechanism by which pathogens bacteria evade the immune response (Kim et al. 2018).
The gut microbiota also influences the regulation of neutrophils function. For instance, a gut microbiota bacteria cell wall component, peptidoglycan, is recognized by the cytosolic receptor-nucleotide oligomerization domain 1 (NOD1), and this interaction intensifies the killing capacity of bone marrow neutrophils (Clarke et al. 2010). Conversely, GF rats are neutropenic and have diminished nitric oxide and superoxide anion generation and decreased phagocytosis in blood neutrophils (Ohkubo et al. 1990). When those GF rats were transferred to a conventional environment it has not observed a recovery in superoxide anion production, indicating an impairment of neutrophils functions perhaps due to a lack of bacteria antigenic stimulation by gut microbiota.
NK cells, which are innate lymphoid subsets responsible for antitumor and antiviral responses, are also found in the gut mucosa. Recent studies identified two different subsets of intestinal NK cells that expressed the natural cytotoxicity receptor NKp46. One subset of gut NKp46+ cells is very similar to conventional NK cells while the other subset diverges from classical NK which displays restricted IFN-γ translation and lacks perforin production (Satoh-Takayama et al. 2008). In contrast, these NKp46+ cells expressed the nuclear hormone receptor retinoic acid receptor-related orphan receptor gamma t (RORγt) and interleukin-22 (IL-22) in the presence of Citrobacter rodentium. In GF mice there is an absence of IL-22-producing NKp46+ cells indicating that signals from the gut microbiota contribute to the differentiation of IL-22-producing NKp46+ cells (Sanos et al. 2009).
Also, mast cells in the lamina propria (LP) represents 2–3% of cells in the GI tract (Boeckxstaens 2018) and in the GF mice there is a reduction in the proportion of the intestinal mast cell and an increase in mast cells circulating in the blood compared to normal mice (Kunii et al. 2011). These results suggest that intestinal bacteria may regulate the migration of blood mast cells to the intestine. Mechanistically, this migration is promoted by the releasing of the CXCR2 ligands from intestinal epithelial cells (IECs) in a TLR-dependent fashion and MyD88−/− mice had lower densities of intestinal mast cells than raised mice (Kunii et al. 2011).
8.2.1.2 Interactions Between the Adaptive Immune System and the Microbiota
Gut microbiota also has an essential role in B cells. Generally, B cells are found in the gut-associated lymphoid tissues, like Peyer’s patches and mesenteric lymph nodes, differentiated in immunoglobulin (Ig)A-secreting plasma cells (Hapfelmeier et al. 2010). IgA is an important form of secretory antibody found in gut mucosa composing a physical barrier and regulating the expression of genes by microbes in the intestine thus maintaining gut homeostasis (Peterson et al. 2007). Mechanistically, secretory IgA binds and obstructs the uptake of microbial antigens in the lumen, leads to bacterial disturbance and agglutination, and besides neutralizes pathogenic bacterial toxins (Cong et al. 2009). In addition, secretory IgA modulates and downregulates the expression of pro-inflammatory surface epitopes in the commensal bacterium Bacteroides thetaiotaomicron (Lindner et al. 2012). The absence of intestinal microbiota leads to a reduction in the number of plasma cells IgA+ in the gut, mostly in PP and in the LP, and a lower level of IgA (Wei et al. 2011). Likewise, deficient mice for Toll-like receptor 5 (TLR5), known to recognize bacterial flagellin, exhibit reduced levels of IgA leading to an abnormal expression of genes related to flagellum structure commensal bacteria (Cullender et al. 2013). In addition, people with IgA deficiency present a higher rate of bacteria with potentially inflammatory properties (Friman et al. 2002). Although gut IgA diversity is individual-specific, specific pathogen-free (SPF) mice have a much greater richness of gut IgA repertoires when compared to mono-colonized mice with bacteria or GF mice (Hapfelmeier et al. 2010). During mice aging, the IgA repertoire gets more complex while new B cell clones are persistently generated against new possible microbiota antigens. Interestingly, B cell clones acquired during the beginning of life are also kept, revealing a long-lasting memory B cell response (Lindner et al. 2015).
T cells are a key component of the adaptive immune system involved in killing infected host cells, activation of other immune cells, production of cytokines, and regulation of the immune response. T cells that express CD4 can be found in every organ of the body, and include a high amount of the T cells of the LP of the intestine (Lindner et al. 2012). Once activated, naive T CD4+ cells can differentiate in four subsets: T helper 1 (Th1), Th2, Th17, or regulatory T cell (Treg), which differ by distinct expression of various transcription factors and cytokines. The correct adjustment and balance of T cell subtypes is an essential element in shaping homeostasis state. Unrestrained Th responses are related to pathological disorders, for example, the Th1 and Th17 responses have been related to autoimmune diseases whereas the Th2 response has been related to allergic response (Geuking et al. 2011).
Likewise, gut microbiota modulates TCD4+ cell development, inside the LP and in other tissues. Thereby, GF mice exhibit a noticeable reduction in the number of T CD4+ cells in LP (Macpherson et al. 2002). In addition, observed defects in spleens and mesenteric lymph nodes of GF animals, with the absence of lymphocyte zones in these animals (Mazmanian et al. 2005). Also, GF animals have been shown a Th1/Th2 imbalance, their immune response going toward a Th2 response (Mazmanian et al. 2005). Some recent findings revealed that some specific bacterial species play a key role in the development of distinct T cell subtypes. For example, Bacteroides fragilis, through its polysaccharide A (PSA), induces the proliferation of CD4+ T cells into Foxp3+ Treg cells directing to a proper Th1/Th2 balance in the host (Round and Mazmanian 2010).
In addition, gut microbiota induces the development of Th17 cells. For instance, segmented filamentous bacteria (SFB) were revealed to be strong inducers of Th17 in the LP. Contrary to that, in GF mice the number of Th17 cells is reduced in the LP in their gut (Ivanov et al. 2009). Recent reports have shown that the physical interaction of SFB with intestinal epithelial cells (IECs) and Th17 that express T cell receptors (TCRs) specific for adhesive forms of these bacteria is essential for Th17 differentiation (Atarashi et al. 2015), postulating that SFB colonization must trigger distinctive signaling pathways in the intestine to induce Th17 response.
Intestinal Tregs are essential for the preservation of the immune tolerance to dietary antigens and the gut microbiota and the suppression of tissue damage suffered by an immune response against pathogenic bacteria. It was demonstrated that the number of peripheral Treg is diminished in GF mice (Bilate and Lafaille 2012). Some endogenous bacteria, such as Clostridium (cluster IV, XIVa, and XVIII), and bacterial products (PSA of B. fragilis) or SCFA can induce functional colonic Treg in LP (Atarashi et al. 2011) and modulate the pathogenesis of inflammatory diseases (Atarashi et al. 2013). Indeed, PSA of B. fragilis can interact with TLR2 on Treg cells and consequently suppress Th17 response (Round et al. 2011). In addition, DNA from gut microbiota activates TLR9 signaling and maintains immunity by controlling Treg cell conversion in the LP (Hall et al. 2008). Furthermore, colonic Tregs induced by microbiota colonization express low levels of Helios, a key transcription related to thymus-derived Treg (Yang et al. 2016) indicating that these cells are a consequence of induction of peripheral Treg not thymic Treg cells.
CD8+ T cells are cytotoxic cells able to kill infected cells, as well as cancer cells (van der Leun et al. 2020) and intestinal T CD8+ are found, mainly, in the intraepithelial layer of the gut (Imaoka et al. 1996). GF mice exhibit a reduction in the number and cytotoxicity of intestinal CD8+ T cells, suggesting that microbiota provide essential signals required for the maintenance of CD8+ T cell population (Wu and Wu 2012). The gut microbiota takes part in shaping CD8+ T cells perhaps due to the modulation of other peripheral immune cells, such as invariant natural killer T cells, plasmacytoid DCs, and marginal zone B cells (Wei et al. 2010). Interestingly, during the dysregulation of the gut barrier type I IFN signaling is needed leading to CD8 T cell accumulation and effector functions within the small intestine. In fact, blocking type I IFN receptor signaling or depleting CD8 T cell prevented barrier leakage caused by a viral infection, indicating that CD8 T cells response can be a crucial factor of intestinal leakage in the pathogenesis of chronic infections (Labarta-Bajo et al. 2020). Besides, recent findings indicate a novel role for butyrate on CD8+ T cells modulating the gene expression of effector molecules in CD8+ T lymphocytes (Luu et al. 2018).
8.3 Dysregulation of Gut Microbiota and the Association with Inflammation-Mediated Disease
The microbiota can modulate several cells of the immune system, through signaling molecules, as well as by its metabolites, such as SCFAs. Immune cells are responsible for the secretion of cytokines, which are signaling molecules that can, for example, recruit other cell types for the inflammatory site (O'Shea and Murray 2008). One of the ways in which the cross-talk between microbiota and the immune system occurs is through cytokines, such as IL-22 which is produced mainly by CD4+ Th17 T cells (Leung et al. 2014) and innate lymphoid cells (ILCs) (Zeng et al. 2019) and in GF mice, the production of IL-22 is impaired (Sanos et al. 2009; Satoh-Takayama et al. 2008). A recent study demonstrated that the microbiota regulates the production of IL-22 through SCFAs in T cells and ILCs inhibiting the inhibition of histone deacetylase (HDAC) and GPR41 and promoting the expression of aryl hydrocarbon receptor (AhR) and hypoxia-inducible factor (HIF-1α) (Yang et al. 2020). In addition, SCFAs propionate and butyrate facilitate the generation of Foxp3+ Tregs (Arpaia et al. 2013; Smith et al. 2013).
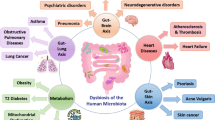
Thus, demonstrating that an intestinal dysbiosis influences the population of immune cells, being able to trigger inflammatory, infectious diseases, dysplasia, and others related to dysregulation of the immune system (Fig. 8.3).
Gut dysbiosis and diseases. Intestinal dysbiosis changes the immune cells and increases the risk of some diseases, such as rheumatoid arthritis, type 1 diabetes, multiple sclerosis, metabolic diseases, hypertension, atherosclerosis, melanoma, colorectal cancer, ulcerative colitis, and Crohn’s disease. CVD cardiovascular diseases, IBD inflammatory bowel diseases
8.3.1 Inflammatory Bowel Disease (IBD)
IBD comprises a range of diseases including Crohn’s disease (CD) and ulcerative colitis (UC) affecting the GI tract (Khor et al. 2011). Several strong evidence reveals that alterations in gut microbiota influence the pathogenesis of IBD (Comito et al. 2014). Indeed, most IBD patients show significant changes in the gut microbiota when compared with normal adults (Manichanh et al. 2006). Antibiotic therapy ameliorates the clinical condition both in patients with IBD and animal IBD models (Khan et al. 2011). Using IBD animal models it was demonstrated that GF rederivation leads to a milder form of the disease in the IL-2 knockout (KO) IBD model or protects against disease (in the T cell receptor α/β KO or IL-10 KO IBD models), suggesting that habitual gut microbiota modulates the inflammatory state of IBD (Schultz et al. 1999; Sellon et al. 1998). Lately, it has been observed a reduction in the relative abundance of Firmicutes and Bacteroides species and an accentuated growth of proteobacteria in feces/mucosa-associated microbiota of IBD patients (Frank et al. 2007).
CD patients present discontinuous lesions that affect any parts of the GI and include chronic and relapsing transmural inflammation resulting in severe abdominal pain, diarrhea, obstruction, and/or perianal lesion. Currently, CD pathogenesis involves the complex balance between genetic, microbiological, immunological, and environmental factors (Neuman and Nanau 2012). In fact, there is evidence that changes in the microbiome are involved with the inflammatory response of the disease (Baker et al. 2009). The intestinal commensal populations are distinctive in CD patients compared to healthy individuals. Especially, some protective microbes and normal anaerobic bacteria, such as Bacteroides sp., Eubacterium sp., and Lactobacillus sp., are remarkably reduced in active CD patients (Alhagamhmad et al. 2016). Also, adherent–invasive E. coli has been related to a higher prevalence of CD, due to over colonization of epithelial cells, mostly in ileal regions (Palmela et al. 2018). Interestingly, a study that compared the cytokines production by intestinal intraepithelial lymphocytes (IELs) in health and CD patients, found that IELs from subjects with CD secreted significantly larger amounts of TNF-α, IFN-γ, and IL-17A (Regner et al. 2018).
UC is triggered by chronic inflammation in the colon, which has been increasing in the number of patients in recent years. A variety of environmental and genetic factors are associated with the development of UC (Feuerstein et al. 2019). Also, the intestinal microbiota in balance with the immune system is essential for maintaining the epithelial barrier and homeostasis. Therefore, an imbalance between the microbiota and the immune system can trigger excessive intestinal inflammation, consequently leading to the development of a UC (Shen et al. 2018). In patients with UC, the diversity and quality of the microbial population are altered, with an increase in Proteobacteria, mainly E. coli, and variable changes in Bacteroidetes (Hansen et al. 2010). In experimental models, these changes vary according to the model used, in general, the variations are similar to the human (Lupp et al. 2007). Another interesting factor that is changed in UC is the immune cell profile, a study was done on CD45+ blood cells of patients with UC showed differences in the expression of Treg, T cell, and CD8+ tissue-resident memory cells (Boland et al. 2020).
The modulation of the microbiota through antibiotic treatment for UC alters the microbial composition in different forms varying according to the experimental model, being an alternative therapy that needs to be better explored (Rooks et al. 2014). Genetic ablation of genes responsible for an encoded protein that recognizes bacteria components is also important to maintain microbiota composition. Animals with innate immunodeficiency in the TLR5 that promotes intestinal inflammation are likely to develop colitis (Vijay-Kumar et al. 2007). Likewise, NLRP6 inflammasome KO mice are more susceptible to develop severe colitis (Elinav et al. 2011). Thus, the gut microbiota seems to be a determinant for this development, in which there is an increase in Proteobacteria and consequently a greater susceptibility to infections by E. coli (Carvalho et al. 2012). Another experimental model of UC used is through IL-10 deficient mice that develop inflammation in the colon spontaneously, with a CD4+ Th1 cell response and excessive production of pro-inflammatory cytokines IL-12, IL-17, and IFN-γ (Keubler et al. 2015). According to Maharshak et al., the IL-10−/− germ-free animal colonized with microbiota from SPF loses its wealth 4 weeks after colonization with an increase in Proteobacteria and Tenericutes (Maharshak et al. 2013), strengthening the idea of the role of immune cells in controlling gut microbiota composition.
The IL-22 signaling molecule plays an important role in maintaining intestinal homeostasis (Shohan et al. 2020) and can be regulated by microbiota through metabolites derived from microbial tryptophan such as kynurenine (Kyn) and indole-3-acetic acid (IAA) that are Ahr ligands (Zelante et al. 2013). The signaling of IL-22 and Ahr ligands has been associated with the expression of the CARD9 gene (Lamas et al. 2016) which is related to susceptibility to IBD (Lanternier et al. 2015). CARD9 deficient mice are more susceptible to colitis, by decreasing antimicrobial peptides REGIII-γ and REGIII-β, change in fungal and bacterial microbiota (Lamas et al. 2016). Fecal microbial transfer experiments from CARD9−/− mice into GF are sufficient to increase susceptibility to DSS-induced colitis, potentially by a deficiency in the induction of IL-22 by T cells and ILCs and decreasing AhR ligands (Lamas et al. 2016). AhR signaling is an essential component of intestinal immune response and the microbiota is one of the main ones responsible to produce Ahr ligands (Lamas et al. 2018). Thus, AhR signaling is one of the pathways whereby the gut microbiota and immune cells communicate (Modoux et al. 2021). Animals treated with indole-3-aldehyde (IAId), a tryptophan metabolite synthesized primarily by Lactobacilli, promote IL-22 production and reduce DSS-induced colitis (Zelante et al. 2013). In addition, according to Qiu et al. Rorc GFP+ AhR−/−, SPF and aged mice have increased Th17 cells producing IFN-γ and IL-17 and the development of chronic spontaneous colitis (Qiu et al. 2013). In Rag−/−AhR−/− animals the inflammation worsened, and it was later reversed with a decrease in the inflammatory infiltrate in the colon, through treatment with antibiotics (Qiu et al. 2013). Also, animals treated with indole-3-carbinol (I3C) AhR ligand of plant origin have their microbiota altered, and in the colitis model have an attenuation of inflammation with lower intestinal permeability, increasing the production of IL-22 by ILC3, with an increase in bacteria producing butyrate and consequently an increase in regulatory T cells (Busbee et al. 2020). Innate immunity cells participate in the colitis development process (Geremia et al. 2014); however, studies focusing on microbial AhR ligands and their association with innate immune system cells in UC models are scarce, and it is important to explore the role of Ahr in innate cells in colitis. In fact, in vitro experiments with LPS-stimulated macrophages showed the potential anti-inflammatory function of indole-3-acetate (I3A), by decreasing cytokines at mRNA levels of the IL-1β, MCP-1 e TNF-α (Krishnan et al. 2018). In an in vivo UC model, treatment with the microbial metabolite ligand of AhR, Indole-3-pyruvic acid (IPA), had an anti-inflammatory effect with increased DCs CD103+ CD11b− (Aoki et al. 2018). Together these findings indicate an essential function of the microbiota in the development of UC through AhR ligands signaling in AhR-expressing immune cells.
Another group of microbial metabolites important in the modulation of immune cells and inflammatory diseases is SCFA. The three most studied SCFA are the butyrate, produced mainly by phylum Firmicutes, and the acetate and the propionate, produced by the phylum Bacteroidetes (Bilotta and Cong 2019). SCFA receptors are G-protein coupled receptors or GPCR or GPR expressed in several immune and intestinal epithelial cells and the stimulation of GPRs by SCFA in these cells contributes to the maintenance of gut homeostasis (Bilotta and Cong 2019). The SCFAs themselves induce Treg in the colon GPR43-dependent manner (Smith et al. 2013), and butyrate and propionate, but not acetate, are involved in the increase in peripheral Treg, through the inhibition of HDAC (Arpaia et al. 2013; Furusawa et al. 2013). Corroborating with data showing that in an IL-10−/− colitis model, mice treated with antibiotics have a reduction in CD4+ Treg and Th1 cells, a decrease in bacteria belonging to the Firmicutes and Bacteroidetes groups, and a reduction in the total levels of SCFA (Shen et al. 2019), known Treg cell regulators (Arpaia et al. 2013). In addition, animals deficient in HIF-1α in epithelial cells with DSS colitis induction reduce the amount of butyrate-producing bacteria and after treatment with sodium butyrate there was a decrease in F4/80+ cells, cytokines IL-6, TNF-α, and IL-1β (Zhou et al. 2020). Also, acetate is another SCFA with anti-inflammatory potential, reducing migration (Kamp et al. 2016) and infiltration/activation of neutrophils in UC, GPR43-dependent (Maslowski et al. 2009), indicating that bacterial metabolites SCFAs have an anti-inflammatory effect.
Intestinal dysbiosis and alteration of the immune system are present in patients and experimental models of UC, requiring a balance between these factors for a better prognosis of the disease. Indicating an immunomodulatory capacity of microbiota and its metabolites, which are possible therapeutic targets for UC.
8.3.2 Cancer
One of the triggers to the development of cancers is due to an accumulation of genetic changes within the cell, which can be a consequence of genetic predisposition, environmental factors, and individual habits such as smoking and food, among others (Lewandowska et al. 2019). There is a necessary immune balance for cell types to behave in an antitumor manner, as the abundance of both Treg cells and effector T cells (CD8+) can cause pro- and antitumor effects, respectively (Farhood et al. 2019).
The intestinal microbiota is an important factor when we talk about the development of cancer, in which the change in its composition can impact patient prognosis (Gopalakrishnan et al. 2018a). This is exemplified by a study demonstrating that GF animals with colorectal cancer (CRC) induction by azoxymethane (AOM), when conventionalized with feces from patients with CRC, has an increase in Th17 and Th1 cells, directly involved in CRC development process, in addition to positive regulation of proliferation, metastasis and angiogenesis genes (Wong et al. 2017). Consequently, these animals develop high-grade dysplasia when compared to controls (Wong et al. 2017). In a CRC model using AOM+ sodium dextran sulfate (DSS), the bacterial composition proved to be essential, in which the colonization of GF animals with Bacteroides fragilis decreased the infiltration of granulocytes and the formation of tumors (Lee et al. 2019), showing the potential of modulation of the microbiota in the development of the CRC.
The impact of the microbiota during treatment with checkpoint inhibitors, which target immunomodulatory T cell molecules, has been gaining emphasis in recent years. Oral administration of the bacterial genus Bifidobacterium improved antitumor immunity, with the recruitment of CD8+ T cells for TME and the combination with anti-PDL1 checkpoint inhibitor led to tumor elimination in an experimental model of melanoma (Sivan et al. 2015). According to Routy et al. animals and patients treated with antibiotics had a response to anti-PDL1 therapy compromised (Routy et al. 2018). In addition, patients who received anti-PDL1 therapy and with a high abundance of Faecalibacterium, Clostridiales, and Ruminococcaceae had high frequencies of CD8+ and TCD4+ T cells as well as a better response to therapy, while the abundance of Bacteroidales is related to increased Tregs and a lower response to therapy (Chaput et al. 2017; Gopalakrishnan et al. 2018a). Another checkpoint inhibitor used in cancer therapies is anti-CTLA-4, and it has been shown that microbiota modulation is essential for its efficiency and affects Th1-type responses in melanoma and CRC models (Vetizou et al. 2015). Taken together, the efficacy of checkpoint blockade treatment should consider gut microbiota composition to increase the extraordinary potential of this therapy to treat cancer patients.
8.3.3 Hypertension and Cardiovascular Diseases
In addition to influencing diseases related to the gastrointestinal system, a change in the microbiota can favor cardiovascular diseases (CVD), through the modulation of the immune system by the intestinal microbiota causing systemic immune effects (Kitai and Tang 2018). Among the CVD that are altered or alter the intestinal microbiota are hypertension, heart failure, and atherosclerosis (Roth et al. 2017), and research with a focus on alternative therapies to combat these diseases is extremely necessary.
Blood pressure (BP) is regulated by several factors, such as genes, environment, hormones, and it is currently suggested that the intestinal microbiota is also a regulatory factor (Kitai and Tang 2018). It is observed in animals with hypertension, a reduction in the production of SCFAs and changes in the composition of the microbiota, with an increase in Firmicutes and a decrease in Bacteroidetes, as well as in patients who have a lower microbiome richness and diversity (Yang et al. 2015). In addition, SCFAs receptors that are present in various cardiac tissues can modulate blood pressure through SCFAs signaling (Pluznick 2014). Propionate produced by the intestinal microbiota, the intravenous infusion of propionate resulted in a drop in BP; however, GPR41-deficient mice this effect is attenuated, suggesting that Gpr41 mediates the hypotensive effects of propionate (Pluznick et al. 2013). Also, to the direct influence of the microbiota on hypertension, it can act by modulating the immune system, Toral et al. performed fecal transplantation of hypertensive in normotensive animals and observed an increase in BP and inflammatory markers in the aortic infiltrate, TNF-α, IFN-γ, Rorγ, and IL-17 as well as a decrease in FoxP3 and IL-10, after administration of neutralizing IL-17 antibody there was a decrease in BP and pro-inflammatory markers (Toral et al. 2019).
Arteriosclerosis is considered another chronic inflammatory disease, involving the entire immune system modulating the onset and progression of lesions, being characterized mainly by the accumulation of fat in the arterial walls (Gui et al. 2012). Intestinal dysbiosis can also contribute to the development of atherosclerosis through systemic inflammation (Duttaroy 2021) indicating an immunomodulatory role of the microbiota through its bacterial products. A study using GF animals showed a decrease in arteriosclerosis, as well as a decrease in plasma levels of LPS and inflammatory markers, IL-6, IL-1β, and TNF-α (Kasahara et al. 2017). In addition, trimethylamine and trimethylamine N-oxide (TMAO) are metabolites derived from the intestinal microbiota and oral supplementation with TMAO has a pro-atherogenic effect linked to cardiovascular risks, dependent on the microbiota (Koeth et al. 2013) with enrichment of specific microbial group, increase in Coriobacteriaceae, Erysipelotrichaceae, and Allabaculum, and a decrease in Candidatus arthromitus, Lachnospiraceae, Oscillospira, and Ruminococcus (Zhu et al. 2016). The effects of TMAO in atherosclerosis have recently been associated, in endothelial cells, with increased oxidative stress, the activation of NLRP3 with the release of inflammatory cytokines, IL1β and IL18, an increase in protein kinase C (PKC) activation and Nuclear factor κB (NF-κB) phosphorylation, consequently inducing positive regulation of Vascular cell adhesion protein 1 (VCAM-1) and an increase in monocyte adhesion (Ma et al. 2017; Sun et al. 2016).
The balance between the immune system and microbiota is altered in experimental models and patients with CVD and its modulation has been shown to be effective in reducing pathology, highlighting the therapeutic potential targeting the immunity-microbiota axis in CVD (Adnan et al. 2017).
8.3.4 Autoimmune Diseases
Changes in gut microbial populations have been connected to autoimmune diseases modulating the immune sensing that recognizes between self and nonself, impacting autoimmune diseases (Leipe et al. 2010).
Rheumatoid arthritis (RA) is also an inflammatory and systemic disease that causes the destruction of bone and cartilage and in consequence, evolving into functional disability. Recent studies demonstrated that RA is related to the Th1- and Th17-mediated inflammatory response and it seems that the disproportion between Th17 and Tregs has been linked to the etiology and progression of RA (Xu et al. 2019). Using experimental collagen-induced arthritis (CIA) animals Mui et al. identified the influence of gut microbiome on arthritis susceptibility (Wu et al. 2010). There were observed changes in the gut microbiota composition between CIA-susceptible and CIA-resistant or healthy mice. During RA, in CIA-susceptible, the relative abundance of families Bacteroidaceae, Lachnospiraceae overlap significantly the Lactobacillus genus found before arthritis onset (Wu et al. 2010). Using the model of RA study (IL-1 receptor antagonist deficient (IL-1Rn−/−) it was observed that the gut microbiota was involved in RA development in mice while GF IL-1Rn−/− mice did not exhibit the disease (Abdollahi-Roodsaz et al. 2008). Furthermore, monocolonizated with Lactobacillus bifidus of the GF IL-1Rn−/− mice restored the disease. Besides, the decrease of Th17 and Tregs cells observed in the lymph nodes and spleens were correlated with disease intensification in non-GF TLR2−/− IL-1Rn−/− mice and disease improvement in non-GF TLR4−/− IL-1Rn−/− mice (Abdollahi-Roodsaz et al. 2008). In the K/BxN mouse, a reduction in RA symptoms in GF-K/Bx was observed (Wu et al. 2010). Mechanistically, the authors demonstrated that the gut microbiota-induced LP small intestine Th17 cell migrated into the peripheral lymphoid tissue, then, stimulated B cells differentiation and autoantibody production in an IL-17-dependent fashion that can lead to the progression of the disease (Wu et al. 2010).
In humans, RA patients show less diverse gut microbiota composition compared with controls, indicating a relationship between the RA duration and autoantibody levels in these patients (Picchianti-Diamanti et al. 2018). A taxon-level analysis revealed that control samples have higher Actinobacteria levels when compared to RA patients (Chen et al. 2016). A study using forest algorithms indicated that Collinsella, Eggerthella, and Faecalibacterium are correlated to RA. The abundance of Collinsella was related to increased levels of alpha-aminoadipic acid and asparagine as long as the production of IL-17A, and Collinsella is involved in the process of disrupting gut permeability and RA severity in the experimental arthritis model (Wang and Xu 2019).
Type 1 diabetes (T1D) is one more disorder framed in autoimmune disease in which β cells are abolished by T cell-mediated response and in consequence very little or no insulin is produced by the islets of Langerhans in the pancreas (Katsarou et al. 2017). The observation that intestinal Tregs were reduced in T1D patients, indicated the likely implication of the gut microbiota in T1D pathogenesis (Badami et al. 2011). Using non-obese diabetic (NOD) mice have been observed that diabetic incidence is markedly higher in GF NOD mice when compared with their SPF controls (Alam et al. 2011). In harmony with these findings, SPF MyD88−/− NOD, lacking MyD88 protein, did not develop T1D, whereas GF MyD88−/− NOD mice readily developed TD1. Interestingly, colonization of GF MyD88-negative NOD mice with a microbial consortium, likely to the phyla normally present in the human gut attenuates T1D, the same result was observed when GF NOD mice were exposed to the microbiota of SPF MyD88-negative NOD donors ameliorating T1D in the GF mice (Wen et al. 2008).
Multiple sclerosis (MS) is an autoimmune disease featured by an abnormal immune system response directed against the central nervous system, leading to demyelination of this system (Oh et al. 2018). Since there is no specific murine model for human MS, the researchers have focused on the use of experimental autoimmune encephalomyelitis (EAE), the most common and accepted experimental model for the human inflammatory demyelinating disease (Constantinescu et al. 2011). Using GF induced for EAE models, Lee et al. noted an attenuation of disease phenotype in these mice, associated with reduced production of pro-inflammatory cytokines, such as IL-17 (Lee et al. 2011). In addition, intestinal colonization with SFB, a known stimulator of IL-17 production in the gut, induces IL-17A-producing CD4(+) T cells (Th17) in the CNS and restoring the phenotype of EAE and worsening the progression of the disease (Lee et al. 2011), indicating a role for SFB in EAE pathogenesis. On the other hand, some commensals may be beneficial in lessening EAE development. The colonization with human commensal B. fragilis can attenuate disease, due to the expression of PSA enhancing the number of Treg cells and CD5+ B cells in the animals treated with B. fragilis (Ochoa-Reparaz et al. 2010). Interestingly, the treatment with Lactobacillus strains (L. paracasei DSM 13434, L. plantarum DSM 15312, and DSM 15313) lead to suppression and reversion suppressed of the clinical symptoms of EAE, and these therapeutic effect was due to IL-10-producing Tregs stimulated by Lactobacillus presence (Lavasani et al. 2010).
8.4 The Gut Microbiota Manipulation as a Treatment in Diseases
As mentioned on the topics above, the gut microbiota composition and/or molecules produced by the microbiota are important to prevent or attenuated intestinal and extraintestinal diseases as well as to maintain the health states of the body thus demonstrating that regulation of the microbiota composition and microbiota-producing product are extremely regulated where different pathways, cell, and molecules take place. In this sense, targeting the microbiome as a strategy to prevent or treat disease may be an interesting approach. A review discusses the importance of alteration in the gut microbiota in pancreatic ductal adenocarcinoma patients, point out that these patients had a decreased microbial diversity, decreased beneficial bacteria, and abundance of potential pathogens (Yu et al. 2021).
There is a strategy already approved by the Food and Drug Administration (FDA) in the USA as a treatment for severe, recurrent Clostridium difficile infections (Napolitano and Covasa 2020) called fecal microbiota transplantation (FMT). This technique appears to be a potential therapy that can modify the human gut microbiome, once the transfer of living microorganisms from a donor to an afflicted person, presents an improvement of the response of the body related to not only Clostridium difficile infections (Napolitano and Covasa 2020) but also obesity and metabolic syndrome (Marotz and Zarrinpar 2016) cancer (Gopalakrishnan et al. 2018b) and liver disease as hepatic encephalopathy (Hassouneh and Bajaj 2021).
In the field of innovation, nanotechnology brings nanoparticles (NPs) and their application in drugs, and medication delivery (Wang et al. 2021) facilitating the treatment of several diseases.
One of the options for the treatment of IBD is Cyclosporine A (CYA) which some patients present resistance to the steroid and adverse effects like toxicity and infections after the treatment (Kornbluth 1999). Targeted delivery of CYA by polymeric nanoparticles shows to improve the therapy of IBD on DSS-induced acute colitis in mice model and shows that the effect of the drug is focused on the intestinal mucosa and not in the systemic absorption by oral administration (Melero et al. 2017). This work also demonstrated that nanoparticles and microparticles develop the same effect on IBD.
Not only CYA polymeric nanoparticles for IBD but a lot of other options of nanoparticle-based drugs have been studied as treatment strategies. Deliverable targets include intestinal epithelium, mucus, immune cells, LP, and the extracellular matrix are targets for NP and routes of NP administration such as oral, injection, and rectal administration (Yang and Merlin 2019). The oral administration of platinum nanoparticles (PtNPs) shows to improve and attenuate colonic and systemic inflammation in DSS-induced colitis mice model, protect their gut barrier from acute colitis, and in macrophage RAW264.7 cell murine culture, PtNPs attenuate inflammation from LPS, suppression Toll-like receptor 4/Nf-κB signaling, although this administration results in dysbiosis (Zhu et al. 2019). Gold nanoparticles (AuNPs) display a potential therapeutic action, and like PtNPs, AuNPs protect against colitis, suppressing TLR4, impacting negatively in mice microbiota, and inducing gut dysbiosis (Zhu et al. 2018).
Another option is the hyaluronic acid-bilirubin nanomedicine which accumulates in the colonic epithelium, restoring the epithelium barriers in an experimental model of acute colitis, which can modulate gut microbiota, associate with pro-inflammatory macrophages, regulating innate immune response (Lee et al. 2020).
Nanobiotechnology proved to be a promising action in the treatment of diseases, mainly for IBD. For a more efficient action of these NPs, it is necessary to interact with the immune system so that both help each other and manage to fight the disease.
8.5 Conclusion and Prospects
As discussed in this chapter, the gut microbiota is capable of modulating, directly or indirectly, the aspects and functions of innate and adaptive immunity directly impacting inflammatory, autoimmune, metabolic, and neurologic diseases. Although promising, there is still a lot to understand about how gut bacteria mechanistically affect local and systemic immunity. Beyond the genetic and immunological factors, environmental factors are a fundamental key to shape the gut microbiota. These features should be considered with precaution as inapt practices such as indiscriminate use of antibiotics is related to higher risks of inflammatory diseases mediated by the microbiota immunomodulation. The impact of intestinal commensals on health state and disease due to the regulation of immune system function has become a new field of science with potential clinical importance for disease therapy. For instance, nanotechnology approaches have emerged as a powerful strategy for manipulating gut microbiota to prevent intestinal and extraintestinal diseases, such as IBD, obesity, and metabolic syndromes. Such therapies when considered for human application must take into consideration the wide variation in gut microbiome diversity and innate immune responses that occur between each individual.
A better understanding of the mutual interactions of the microbiota and host immune system, as well as gut microbiota role, modulate the immune system will contribute to many strategies for manipulating the intestinal microbiome for therapeutic benefit, especially in inflammatory and autoimmune diseases.
References
Abdollahi-Roodsaz S et al (2008) Stimulation of TLR2 and TLR4 differentially skews the balance of T cells in a mouse model of arthritis. J Clin Invest 118(1):205–216
Adnan S et al (2017) Alterations in the gut microbiota can elicit hypertension in rats. Physiol Genomics 49(2):96–104
Alam C et al (2011) Effects of a germ-free environment on gut immune regulation and diabetes progression in non-obese diabetic (NOD) mice. Diabetologia 54(6):1398–1406
Alhagamhmad MH et al (2016) An overview of the bacterial contribution to Crohn disease pathogenesis. J Med Microbiol 65(10):1049–1059
Aoki R et al (2018) Indole-3-pyruvic acid, an aryl hydrocarbon receptor activator, suppresses experimental colitis in mice. J Immunol 201(12):3683–3693
Arpaia N et al (2013) Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504(7480):451–455
Arumugam M et al (2011) Enterotypes of the human gut microbiome. Nature 473(7346):174–180
Atarashi K et al (2011) Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331(6015):337–341
Atarashi K et al (2013) Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500(7461):232–236
Atarashi K et al (2015) Th17 cell induction by adhesion of microbes to intestinal epithelial cells. Cell 163(2):367–380
Badami E et al (2011) Defective differentiation of regulatory FoxP3+ T cells by small-intestinal dendritic cells in patients with type 1 diabetes. Diabetes 60(8):2120–2124
Baker PI, Love DR, Ferguson LR (2009) Role of gut microbiota in Crohn’s disease. Expert Rev Gastroenterol Hepatol 3(5):535–546
Bilate AM, Lafaille JJ (2012) Induced CD4+Foxp3+ regulatory T cells in immune tolerance. Annu Rev Immunol 30:733–758
Bilotta AJ, Cong Y (2019) Gut microbiota metabolite regulation of host defenses at mucosal surfaces: implication in precision medicine. Precis Clin Med 2(2):110–119
Boeckxstaens GE (2018) The emerging role of mast cells in irritable bowel syndrome. Gastroenterol Hepatol (N Y) 14(4):250–252
Boland BS et al (2020) Heterogeneity and clonal relationships of adaptive immune cells in ulcerative colitis revealed by single-cell analyses. Sci Immunol 5(50):eabb4432
Busbee PB et al (2020) Indole-3-carbinol prevents colitis and associated microbial dysbiosis in an IL-22-dependent manner. JCI Insight 5(1):e127551
Carvalho FA et al (2012) Transient inability to manage proteobacteria promotes chronic gut inflammation in TLR5-deficient mice. Cell Host Microbe 12(2):139–152
Chaput N et al (2017) Baseline gut microbiota predicts clinical response and colitis in metastatic melanoma patients treated with ipilimumab. Ann Oncol 28(6):1368–1379
Charbonneau MR et al (2016) Sialylated milk oligosaccharides promote microbiota-dependent growth in models of infant undernutrition. Cell 164(5):859–871
Chen J et al (2016) An expansion of rare lineage intestinal microbes characterizes rheumatoid arthritis. Genome Med 8(1):43
Clarke TB et al (2010) Recognition of peptidoglycan from the microbiota by Nod1 enhances systemic innate immunity. Nat Med 16(2):228–231
Comito D, Cascio A, Romano C (2014) Microbiota biodiversity in inflammatory bowel disease. Ital J Pediatr 40:32
Cong Y et al (2009) A dominant, coordinated T regulatory cell-IgA response to the intestinal microbiota. Proc Natl Acad Sci U S A 106(46):19256–19261
Constantinescu CS et al (2011) Experimental autoimmune encephalomyelitis (EAE) as a model for multiple sclerosis (MS). Br J Pharmacol 164(4):1079–1106
Cullender TC et al (2013) Innate and adaptive immunity interact to quench microbiome flagellar motility in the gut. Cell Host Microbe 14(5):571–581
David LA et al (2014) Diet rapidly and reproducibly alters the human gut microbiome. Nature 505(7484):559–563
Duttaroy AK (2021) Role of gut microbiota and their metabolites on atherosclerosis, hypertension and human blood platelet function: a review. Nutrients 13(1):144
Elinav E et al (2011) NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145(5):745–757
Elvers KT et al (2020) Antibiotic-induced changes in the human gut microbiota for the most commonly prescribed antibiotics in primary care in the UK: a systematic review. BMJ Open 10(9):e035677
Farhood B, Najafi M, Mortezaee K (2019) CD8(+) cytotoxic T lymphocytes in cancer immunotherapy: a review. J Cell Physiol 234(6):8509–8521
Feuerstein JD, Moss AC, Farraye FA (2019) Ulcerative colitis. Mayo Clin Proc 94(7):1357–1373
Frank DN et al (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci U S A 104(34):13780–13785
Friman V et al (2002) Increased frequency of intestinal Escherichia coli carrying genes for S fimbriae and haemolysin in IgA-deficient individuals. Microb Pathog 32(1):35–42
Furusawa Y et al (2013) Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504(7480):446–450
Geremia A et al (2014) Innate and adaptive immunity in inflammatory bowel disease. Autoimmun Rev 13(1):3–10
Geuking MB et al (2011) Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity 34(5):794–806
Gopalakrishnan V et al (2018a) The influence of the gut microbiome on cancer, immunity, and cancer immunotherapy. Cancer Cell 33(4):570–580
Gopalakrishnan V et al (2018b) Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359(6371):97–103
Goris H, de Boer F, van der Waaij D (1985) Myelopoiesis in experimentally contaminated specific-pathogen-free and germfree mice during oral administration of polymyxin. Infect Immun 50(2):437–441
Gui T et al (2012) Diverse roles of macrophages in atherosclerosis: from inflammatory biology to biomarker discovery. Mediators Inflamm 2012:693083
Hall JA et al (2008) Commensal DNA limits regulatory T cell conversion and is a natural adjuvant of intestinal immune responses. Immunity 29(4):637–649
Hansen J, Gulati A, Sartor RB (2010) The role of mucosal immunity and host genetics in defining intestinal commensal bacteria. Curr Opin Gastroenterol 26(6):564–571
Hapfelmeier S et al (2010) Reversible microbial colonization of germ-free mice reveals the dynamics of IgA immune responses. Science 328(5986):1705–1709
Hassouneh R, Bajaj JS (2021) Gut microbiota modulation and fecal transplantation: an overview on innovative strategies for hepatic encephalopathy treatment. J Clin Med 10(2):330
Imaoka A et al (1996) Proliferative recruitment of intestinal intraepithelial lymphocytes after microbial colonization of germ-free mice. Eur J Immunol 26(4):945–948
Imhann F et al (2017) The influence of proton pump inhibitors and other commonly used medication on the gut microbiota. Gut Microbes 8(4):351–358
Ivanov II et al (2009) Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139(3):485–498
Iwasaki A, Kelsall BL (1999) Freshly isolated Peyer’s patch, but not spleen, dendritic cells produce interleukin 10 and induce the differentiation of T helper type 2 cells. J Exp Med 190(2):229–239
Kamp ME et al (2016) G protein-coupled receptor 43 modulates neutrophil recruitment during acute inflammation. PLoS One 11(9):e0163750
Kasahara K et al (2017) Commensal bacteria at the crossroad between cholesterol homeostasis and chronic inflammation in atherosclerosis. J Lipid Res 58(3):519–528
Katsarou A et al (2017) Type 1 diabetes mellitus. Nat Rev Dis Primers 3:17016
Keith JW, Pamer EG (2019) Enlisting commensal microbes to resist antibiotic-resistant pathogens. J Exp Med 216(1):10–19
Keubler LM et al (2015) A multihit model: colitis lessons from the interleukin-10-deficient mouse. Inflamm Bowel Dis 21(8):1967–1975
Khan KJ et al (2011) Antibiotic therapy in inflammatory bowel disease: a systematic review and meta-analysis. Am J Gastroenterol 106(4):661–673
Khor B, Gardet A, Xavier RJ (2011) Genetics and pathogenesis of inflammatory bowel disease. Nature 474(7351):307–317
Khosravi A et al (2014) Gut microbiota promote hematopoiesis to control bacterial infection. Cell Host Microbe 15(3):374–381
Kim M et al (2018) Critical role for the microbiota in CX3CR1(+) intestinal mononuclear phagocyte regulation of intestinal T cell responses. Immunity 49(1):151–163
Kitai T, Tang WHW (2018) Gut microbiota in cardiovascular disease and heart failure. Clin Sci (Lond) 132(1):85–91
Koeth RA et al (2013) Intestinal microbiota metabolism of l-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat Med 19(5):576–585
Kornbluth A (1999) Cyclosporine in inflammatory bowel disease. Curr Gastroenterol Rep 1(6):486–490
Krajmalnik-Brown R et al (2012) Effects of gut microbes on nutrient absorption and energy regulation. Nutr Clin Pract 27(2):201–214
Krishnan S et al (2018) Gut microbiota-derived tryptophan metabolites modulate inflammatory response in hepatocytes and macrophages. Cell Rep 23(4):1099–1111
Kunii J et al (2011) Commensal bacteria promote migration of mast cells into the intestine. Immunobiology 216(6):692–697
Labarta-Bajo L et al (2020) Type I IFNs and CD8 T cells increase intestinal barrier permeability after chronic viral infection. J Exp Med 217(12):e20192276
Lamas B et al (2016) CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat Med 22(6):598–605
Lamas B, Natividad JM, Sokol H (2018) Aryl hydrocarbon receptor and intestinal immunity. Mucosal Immunol 11(4):1024–1038
Lanternier F et al (2015) Inherited CARD9 deficiency in otherwise healthy children and adults with Candida species-induced meningoencephalitis, colitis, or both. J Allergy Clin Immunol 135(6):1558–1568
Lavasani S et al (2010) A novel probiotic mixture exerts a therapeutic effect on experimental autoimmune encephalomyelitis mediated by IL-10 producing regulatory T cells. PLoS One 5(2):e9009
Lazar V et al (2018) Aspects of gut microbiota and immune system interactions in infectious diseases, immunopathology, and cancer. Front Immunol 9:1830
Lee YK et al (2011) Proinflammatory T-cell responses to gut microbiota promote experimental autoimmune encephalomyelitis. Proc Natl Acad Sci U S A 108(Suppl 1):4615–4622
Lee YP et al (2019) The germ-free mice monocolonization with Bacteroides fragilis improves azoxymethane/dextran sulfate sodium induced colitis-associated colorectal cancer. Immunopharmacol Immunotoxicol 41(2):207–213
Lee Y et al (2020) Hyaluronic acid-bilirubin nanomedicine for targeted modulation of dysregulated intestinal barrier, microbiome and immune responses in colitis. Nat Mater 19(1):118–126
Leipe J et al (2010) Role of Th17 cells in human autoimmune arthritis. Arthritis Rheum 62(10):2876–2885
Leung JM et al (2014) IL-22-producing CD4+ cells are depleted in actively inflamed colitis tissue. Mucosal Immunol 7(1):124–133
Lewandowska AM et al (2019) Environmental risk factors for cancer—review paper. Ann Agric Environ Med 26(1):1–7
Lindner C et al (2012) Age, microbiota, and T cells shape diverse individual IgA repertoires in the intestine. J Exp Med 209(2):365–377
Lindner C et al (2015) Diversification of memory B cells drives the continuous adaptation of secretory antibodies to gut microbiota. Nat Immunol 16(8):880–888
Lloyd-Price J, Abu-Ali G, Huttenhower C (2016) The healthy human microbiome. Genome Med 8(1):51
Luo Y et al (2015) Microbiota from obese mice regulate hematopoietic stem cell differentiation by altering the bone niche. Cell Metab 22(5):886–894
Lupp C et al (2007) Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2(2):119–129
Luu M et al (2018) Regulation of the effector function of CD8(+) T cells by gut microbiota-derived metabolite butyrate. Sci Rep 8(1):14430
Ma G et al (2017) Trimethylamine N-oxide in atherogenesis: impairing endothelial self-repair capacity and enhancing monocyte adhesion. Biosci Rep 37(2):BSR20160244
Macpherson AJ, Martinic MM, Harris N (2002) The functions of mucosal T cells in containing the indigenous commensal flora of the intestine. Cell Mol Life Sci 59(12):2088–2096
Maharshak N et al (2013) Altered enteric microbiota ecology in interleukin 10-deficient mice during development and progression of intestinal inflammation. Gut Microbes 4(4):316–324
Manichanh C et al (2006) Reduced diversity of faecal microbiota in Crohn’s disease revealed by a metagenomic approach. Gut 55(2):205–211
Marotz CA, Zarrinpar A (2016) Treating obesity and metabolic syndrome with fecal microbiota transplantation. Yale J Biol Med 89(3):383–388
Maslowski KM et al (2009) Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461(7268):1282–1286
Maurice CF, Haiser HJ, Turnbaugh PJ (2013) Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 152(1–2):39–50
Mazmanian SK et al (2005) An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122(1):107–118
Melero A et al (2017) Targeted delivery of cyclosporine A by polymeric nanocarriers improves the therapy of inflammatory bowel disease in a relevant mouse model. Eur J Pharm Biopharm 119:361–371
Mikkelsen HB et al (2004) Macrophages in the small intestinal muscularis externa of embryos, newborn and adult germ-free mice. J Mol Histol 35(4):377–387
Modoux M et al (2021) Tryptophan metabolism as a pharmacological target. Trends Pharmacol Sci 42(1):60–73
Napolitano M, Covasa M (2020) Microbiota transplant in the treatment of obesity and diabetes: current and future perspectives. Front Microbiol 11:590370
Neuman MG, Nanau RM (2012) Inflammatory bowel disease: role of diet, microbiota, life style. Transl Res 160(1):29–44
Ochoa-Reparaz J et al (2010) Central nervous system demyelinating disease protection by the human commensal Bacteroides fragilis depends on polysaccharide a expression. J Immunol 185(7):4101–4108
Oh J, Vidal-Jordana A, Montalban X (2018) Multiple sclerosis: clinical aspects. Curr Opin Neurol 31(6):752–759
Ohkubo T et al (1990) Impaired superoxide production in peripheral blood neutrophils of germ-free rats. Scand J Immunol 32(6):727–729
O'Shea JJ, Murray PJ (2008) Cytokine signaling modules in inflammatory responses. Immunity 28(4):477–487
Palmela C et al (2018) Adherent-invasive Escherichia coli in inflammatory bowel disease. Gut 67(3):574–587
Peterson DA et al (2007) IgA response to symbiotic bacteria as a mediator of gut homeostasis. Cell Host Microbe 2(5):328–339
Picchianti-Diamanti A et al (2018) Analysis of gut microbiota in rheumatoid arthritis patients: disease-related dysbiosis and modifications induced by etanercept. Int J Mol Sci 19(10):2938
Pluznick J (2014) A novel SCFA receptor, the microbiota, and blood pressure regulation. Gut Microbes 5(2):202–207
Pluznick JL et al (2013) Olfactory receptor responding to gut microbiota-derived signals plays a role in renin secretion and blood pressure regulation. Proc Natl Acad Sci U S A 110(11):4410–4415
Qiu J et al (2013) Group 3 innate lymphoid cells inhibit T-cell-mediated intestinal inflammation through aryl hydrocarbon receptor signaling and regulation of microflora. Immunity 39(2):386–399
Radjabzadeh D et al (2020) Diversity, compositional and functional differences between gut microbiota of children and adults. Sci Rep 10(1):1040
Regner EH et al (2018) Functional intraepithelial lymphocyte changes in inflammatory bowel disease and spondyloarthritis have disease specific correlations with intestinal microbiota. Arthritis Res Ther 20(1):149
Rooks MG et al (2014) Gut microbiome composition and function in experimental colitis during active disease and treatment-induced remission. ISME J 8(7):1403–1417
Roth GA et al (2017) Global, regional, and National Burden of Cardiovascular Diseases for 10 causes, 1990–2015. J Am Coll Cardiol 70(1):1–25
Round JL, Mazmanian SK (2009) The gut microbiota shapes intestinal immune responses during health and disease. Nat Rev Immunol 9(5):313–323
Round JL, Mazmanian SK (2010) Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc Natl Acad Sci U S A 107(27):12204–12209
Round JL et al (2011) The Toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science 332(6032):974–977
Routy B et al (2018) Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359(6371):91–97
Sanos SL et al (2009) RORgammat and commensal microflora are required for the differentiation of mucosal interleukin 22-producing NKp46+ cells. Nat Immunol 10(1):83–91
Satoh-Takayama N et al (2008) Microbial flora drives interleukin 22 production in intestinal NKp46+ cells that provide innate mucosal immune defense. Immunity 29(6):958–970
Schluter J et al (2020) The gut microbiota is associated with immune cell dynamics in humans. Nature 588(7837):303–307
Schultz M et al (1999) IL-2-deficient mice raised under germfree conditions develop delayed mild focal intestinal inflammation. Am J Physiol 276(6):G1461–G1472
Sellon RK et al (1998) Resident enteric bacteria are necessary for development of spontaneous colitis and immune system activation in interleukin-10-deficient mice. Infect Immun 66(11):5224–5231
Shen ZH et al (2018) Relationship between intestinal microbiota and ulcerative colitis: mechanisms and clinical application of probiotics and fecal microbiota transplantation. World J Gastroenterol 24(1):5–14
Shen B et al (2019) Antibiotics exacerbated colitis by affecting the microbiota, Treg cells and SCFAs in IL10-deficient mice. Biomed Pharmacother 114:108849
Shi C et al (2011) Bone marrow mesenchymal stem and progenitor cells induce monocyte emigration in response to circulating toll-like receptor ligands. Immunity 34(4):590–601
Shohan M et al (2020) Interleukin-22 and intestinal homeostasis: protective or destructive? IUBMB Life 72(8):1585–1602
Sivan A et al (2015) Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350(6264):1084–1089
Smith PM et al (2013) The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341(6145):569–573
Smythies LE et al (2005) Human intestinal macrophages display profound inflammatory anergy despite avid phagocytic and bactericidal activity. J Clin Invest 115(1):66–75
Sonnenburg ED et al (2016) Diet-induced extinctions in the gut microbiota compound over generations. Nature 529(7585):212–215
Sun X et al (2016) Trimethylamine N-oxide induces inflammation and endothelial dysfunction in human umbilical vein endothelial cells via activating ROS-TXNIP-NLRP3 inflammasome. Biochem Biophys Res Commun 481(1–2):63–70
Toral M et al (2019) Role of the immune system in vascular function and blood pressure control induced by faecal microbiota transplantation in rats. Acta Physiol (Oxf) 227(1):e13285
Uranga JA et al (2016) Food, nutrients and nutraceuticals affecting the course of inflammatory bowel disease. Pharmacol Rep 68(4):816–826
van der Leun AM, Thommen DS, Schumacher TN (2020) CD8(+) T cell states in human cancer: insights from single-cell analysis. Nat Rev Cancer 20(4):218–232
Vetizou M et al (2015) Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350(6264):1079–1084
Vijay-Kumar M et al (2007) Deletion of TLR5 results in spontaneous colitis in mice. J Clin Invest 117(12):3909–3921
Wang Q, Xu R (2019) Data-driven multiple-level analysis of gut-microbiome-immune-joint interactions in rheumatoid arthritis. BMC Genomics 20(1):124
Wang H et al (2021) Update on nanoparticle-based drug delivery system for anti-inflammatory treatment. Front Bioeng Biotechnol 9:630352
Wei B et al (2010) Commensal microbiota and CD8+ T cells shape the formation of invariant NKT cells. J Immunol 184(3):1218–1226
Wei M et al (2011) Mice carrying a knock-in mutation of Aicda resulting in a defect in somatic hypermutation have impaired gut homeostasis and compromised mucosal defense. Nat Immunol 12(3):264–270
Wen L et al (2008) Innate immunity and intestinal microbiota in the development of type 1 diabetes. Nature 455(7216):1109–1113
Wong SH et al (2017) Gavage of fecal samples from patients with colorectal cancer promotes intestinal carcinogenesis in germ-free and conventional mice. Gastroenterology 153(6):1621–1633
Wu HJ, Wu E (2012) The role of gut microbiota in immune homeostasis and autoimmunity. Gut Microbes 3(1):4–14
Wu HJ et al (2010) Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32(6):815–827
Wu H et al (2017) Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat Med 23(7):850–858
Xu H et al (2019) The dynamic interplay between the gut microbiota and autoimmune diseases. J Immunol Res 2019:7546047
Yang C, Merlin D (2019) Nanoparticle-mediated drug delivery systems for the treatment of IBD: current perspectives. Int J Nanomed 14:8875–8889
Yang T et al (2015) Gut dysbiosis is linked to hypertension. Hypertension 65(6):1331–1340
Yang BH et al (2016) Foxp3(+) T cells expressing RORgammat represent a stable regulatory T-cell effector lineage with enhanced suppressive capacity during intestinal inflammation. Mucosal Immunol 9(2):444–457
Yang W et al (2020) Intestinal microbiota-derived short-chain fatty acids regulation of immune cell IL-22 production and gut immunity. Nat Commun 11(1):4457
Yu Q, Jobin C, Thomas RM (2021) Implications of the microbiome in the development and treatment of pancreatic cancer: thinking outside of the box by looking inside the gut. Neoplasia 23(2):246–256
Zelante T et al (2013) Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 39(2):372–385
Zeng B et al (2019) ILC3 function as a double-edged sword in inflammatory bowel diseases. Cell Death Dis 10(4):315
Zhou C et al (2020) SCFAs induce autophagy in intestinal epithelial cells and relieve colitis by stabilizing HIF-1alpha. J Mol Med (Berl) 98(8):1189–1202
Zhu T, Goodarzi MO (2020) Metabolites linking the gut microbiome with risk for type 2 diabetes. Curr Nutr Rep 9(2):83–93
Zhu W et al (2016) Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165(1):111–124
Zhu S et al (2018) Orally administered gold nanoparticles protect against colitis by attenuating Toll-like receptor 4- and reactive oxygen/nitrogen species-mediated inflammatory responses but could induce gut dysbiosis in mice. J Nanobiotechnol 16(1):86
Zhu S et al (2019) Platinum nanoparticles as a therapeutic agent against dextran sodium sulfate-induced colitis in mice. Int J Nanomed 14:8361–8378
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
This study was supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP): Grant number: 2019/14755-0. Vinicius Andrade-Oliveira is also a fellow of the Pew Latin American Fellow program of the Pew Foundation. This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES)—Finance Code 001, where Renan Willian Alves and Lorena Doretto-Silva received a grant for their PhD degree. Eloisa Martins da Silva is recipient of PhD fellowship from FAPESP (2020/14388-4).
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
da Silva, E.M., Alves, R.W., Doretto-Silva, L., Andrade-Oliveira, V. (2023). Cross-Talk Between Gut Microbiota and Immune Cells and Its Impact on Inflammatory Diseases. In: Ribeiro de Araujo, D., Carneiro-Ramos, M. (eds) Biotechnology Applied to Inflammatory Diseases. Interdisciplinary Biotechnological Advances. Springer, Singapore. https://doi.org/10.1007/978-981-19-8342-9_8
Download citation
DOI: https://doi.org/10.1007/978-981-19-8342-9_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-8341-2
Online ISBN: 978-981-19-8342-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)