Abstract
Crop yield is adversely being influenced very frequently due to biotic and abiotic stresses, globally. The insufficient yield and low nutritional value of the crops are due to the numerous ambient stresses. Which has challenged the nutritional security of people in developing and underdeveloped nations that are already been malnourished. The tremendously changing global climate and the ever-increasing world population are the principal apprehensions guiding towards the adaptation of a neoteric technique that can aid in achieving the sustainable development of agriculture with enriched nutritional value, plant resilience, and improved yield potential. The clustered regularly interspaced short palindromic repeat (CRISPR) and CRISPR-associated (Cas) protein-based genome-editing (CRISPR-Cas) tool is the most valuable technique to boon the modern world and offers an edge over to meganucleases, zinc finger nucleases (ZFNs), and transcription activator-like effector nucleases (TALENs) by being its tremendous potency, accuracy, ease of use, and versatility. Here, we have highlighted the neoteric advancements of the CRISPR-Cas-based approaches that have revolutionized the way of food production in the agriculture industry. It has paved the way for food security by modifying crucial crop attributes by introducing desirable characteristics that employ knockout and/or knockin of targeted genes to generate resistant crop plants with enriched nutritional quality, yield enhancement, and stress resilience. In addition, we have also shed light on different mechanisms, challenges, approaches for the minimization of off-target effects, and future possibilities of these neoteric genome-editing tools. Furthermore, with the advent of the CRISPR-based platform, the numerous emerging biotechnologies have broadened the basic crop research toolbox and synthetic biotechnology via the incorporation of artificial intelligence (AI) and various bioinformatics frameworks. Eventually, the current global regulatory stratagems and social approval of CRISPR-Cas-based crop trait enhancement have been explored.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
9.1 Introduction
9.1.1 Genome Editing: An Introduction to the Plethora of Tools in the Arsenal of Gene Editing
Genetic modification for the development of desired traits in plants utilized for food began before the end of the Pleistocene era about 12,000–11,000 years ago (Larson et al. 2014). Since then, numerous changes have happened due to natural evolutionary processes, which resulted in new crop species that are now genetically different from their ancestors. After establishing Mendel’s ‘principle of genetics’ in 1865, actual genetic modification was started. Plant genetic engineering has designed to generate plants with neoteric attributes that could conquer sustainability goals. Hence, it necessitates introducing advanced genetic engineering strategies, for instance, mutagenesis, transgenic approach, RNAi approaches, genome editing (GEd) via ZFNs, TALENS, CRISPR-Cas approaches. GEd was promoted by introducing double-strand breaks (DSBs) at the targeted locus, which relies on sequence-specific nucleases (SSNs). Recently, the toolkit of GEd comprises four classes of SSNs: meganucleases, ZFNs, TALEN, and CRISPR-Cas systems.
Meganucleases are the naturally available endonucleases, also familiar as homing endonuclease. It is the first-generation SSN and came into the limelight as a self-splicing component of mitochondrial large ribosomal DNA (mtLrDNA) introns of Saccharomyces cerevisiae (Colleaux et al. 1988). This can identify a wide range of DNA (14–40 bp) (Orlowski et al. 2007). Due to variation in target recognition and cleavage site, these SSNs can be grouped into six major families, for instance, His-Cys Box, LAGLIDADG, HNH, EDxHD, GIY-YIG, and PD-(D/E)xK (Belfort et al. 2014). I-SecI is the most frequently utilized meganucleases and was first utilized in tobacco plant (Puchta 1999), since then it was being used by plant biologists for GEd. D’Halluin et al. (2007) reported the utilization of meganucleases in maize. Nevertheless, the lack of editing capability of broad target sequences via protein redesign mightily narrows this SSN’s applications (Rosen et al. 2006).
The re-programmability lacking of meganucleases was solved with ZFNs. ZFNs comprise multiple zinc finger domain harbouring proteins. Those protein domains are generated from the typical Cys-2-His 2-zinc finger domain (Gaj et al. 2013); after recognition of specific sequences, those protein motifs were fastened to DNA in a sequence-specific manner (Weeks et al. 2016). The C-terminal part of each ZFN motif is responsible for targeted sequence recognition. The composition of each ZFN motif binds a 3-bp DNA sequence and is made up of almost 30 amino acids (Maeder et al. 2008). Therefore, unlike meganucleases, these separate domain arrangements made ZFN simpler. Nevertheless, due to lack of endonuclease activity, they require to be fused with Fok I endonuclease domain for cleavage of DNA at the target site (Kaul et al. 2019). The efficiency of ZFNs-mediated gene editing was first successfully employed in Arabidopsis (Lloyd et al. 2005). Similarly, Dicer-like DCL4a and DCL4b gene in soybean was successfully edited utilizing ZFNs (Curtin et al. 2011). ZFN is also used for HDR-mediated gene editing, for instance, the amino acid substitution of SuRa and SuRb gene in tobacco, which conferred resistance to sulphonylurea herbicide (Townsend et al. 2009). However, two separate ZFN motifs target two proximal sites and double the construct size, which may complicate the design of this SSN.
TALENs are specific DNA-binding proteins, large sequences (>30 bp) targets make them more precise (Miller et al. 2011). The TALEs proteins were identified from plant pathogen Xanthomonas sp. Unlike ZFNs, these proteins have DNA-binding modular domains, specifically recognizing one single base instead of three (Moscou and Bogdanove 2009). Additionally, like ZFNs, this single DNA identification modules must be fused with Fok I endonuclease domain (Mahfouz et al. 2014). The possibility of TALEN-based editing was realized in 2009, wherein successful gene editing was first reported in yeast (Christian et al. 2013). Over the past few years, TALENs have emerged as a choice of GEd in plants. It has been successfully employed in a variety of crops, for instance, tobacco (Moore et al. 2014), barley (Budhagatapalli et al. 2015), tomato Čermák et al. 2015), Arabidopsis (Forner et al. 2015). Genome modification via TALENs is handy in comparison to ZFNs, due to its simplicity in using TALEs repeats for each of the DNA nucleotide recognition.
Amongst all the approaches, recently discovered CRISPR-Cas9-based GEd tools have replaced the ZFNs and TALENs and opened the way to modify plant’s genomes with unprecedented precision. This GEd system is revolutionizing the field of plant biology due to its efficiency, specificity, unparalleled flexibility, and target design simplicity. Apart from these, CRISPR-Cas9 has additional advantages over ZFNs and TALENs, including target specificity design, efficiency in incorporating the guide RNA (gRNA), and the RNAs guided Cas9 protein and the ability of multiplexing in a single event. Compared to the previously available techniques, designing a CRISPR-Cas9 vector is easy and efficient with the availability and accessibility of enhanced bioinformatics tools, which can be utilized to find the most selective sequences for designing gRNAs, eliminating the potential for screening libraries to find the most effective target. This technology has been rapidly and widely adopted for a range of applications for instance, multiplex gene knockout, targeted sequence insertion, base editing, prime editing, and so on. A variety of strategies have been developed for optimizing the CRISPR-Cas9 reagents and their delivery systems. This chapter tries to compile a detailed review of the existing GEd approaches, emphasizing the CRISPR-Cas9 technique. We also shed light on a glimpse of information about novel breakthrough and milestone achievements of CRISPR-Cas9 systems and the impact of this system as the next gene tool for crop improvement.
9.2 Era of CRISPR-Cas-Based Genome Editing
The invention of the CRISPR-Cas microbial self-defense mechanism and its ongoing achievement as a genome-editing tool represents the findings of numerous researchers all over the world. Our concise historical era will represent the contributions of different scientists who pushed this GEd field forward from the initial discovery. The clusters of repeats which are separated by spacers were first observed in 1987 during the study of E. coli harbouring jap gene (Ishino et al. 1987). In 1989, the structure of the CRISPR array was defined but without its functional mechanism (Nakata et al. 1989). Interestingly, similar structures were identified later in numerous bacteria and archaea (Hermans et al. 1991; Mojica et al. 1995; Bult et al. 1996). Francisco Mojica characterizes those sequences for the first time in 1993, what is now known as CRISPR locus, and the potentiality of this locus was shown in 2000 (Mojica et al. 2000). Simultaneously, 45 protein families were identified with clusters of CRISPR-associated genes (Haft et al. 2005). After increasing the volume of prokaryotic sequence data, the crucial breakthrough happened in 2005. It was reported that identified CRISPR sequences showed similarity with some bacteriophage and led to immunity against those infectious bacteriophages (Mojica et al. 2005; Pourcel et al. 2005). Bolotin revealed some anomaly in the CRISPR locus and found a large protein with nuclease activity, which is now known as Cas9 (Bolotin et al. 2005). Although found some viral genes resemble sequences at one end, those are the PAM sequence. In the same year 2005, Jennifer Doudna and Jillian Banfield started their investigation on CRISPR and their functions. In 2006, the hypothetical scheme of the CRISPR-Cas adaptive immunity mechanism was proposed by Koonim (Makarova et al. 2006). Later on, this hypothesis was confirmed by Barrangou et al. (2007). After that, scientists started to report the CRISPR-Cas action that how this RNA-mediated system interferes with the invading phage DNA (Brouns et al. 2008). It was also demonstrated that this system could target both DNA (Marraffini and Sontheimer 2008) and RNA (Hale et al. 2009). Alongside, PAM sequences are also essential for some systems described simultaneously (Mojica et al. 2009). Details about the transcription mechanisms of crRNAs were also revealed in 2010 (Haurwitz et al. 2010). The classification of the CRISPR-Cas systems was demonstrated in 2011 (Makarova et al. 2011). In the same year, 2011, Emmanuelle Charpentier and Jenifer Dounda conjointly started to study the CRISPR-Cas mechanism, and they discovered the function of tracrRNA for the Cas9 system. Moreover, the role of RNase III in pre-crRNA and tracrRNA processing was characterized in 2011 (Deltcheva et al. 2011). In 2012, Siksnys and his team mechanically characterized the mode of adaptation of Cas9 via understanding the cell infection kinetics with the CRISPR-Cas system (Datsenko et al. 2012; Gasiunas et al. 2012). At the same time, in 2012, similar findings were reported by Jennifer Doudna in collaboration with Emmanuelle Charpentier. They demonstrated that synthetic gRNAs could be generated via fusion of the crRNA and the tracrRNA (Jinek et al. 2012; Qi et al. 2021). Finally, the newly invented CRISPR-Cas system was led to use for targeted genome modification in bacteria (Gasiunas et al. 2012), yeast (DiCarlo et al. 2013), human (Cong et al. 2013a, b; Jinek et al. 2013; Mali et al. 2013a, b). In 2013, CRISPR-Cas machinery was successfully employed to engineer plant genomes (Shan et al. 2013). In 2014, CRISPR-Cas9 was demonstrated in primates (Cas9 mRNA and sgRNAs were coinjected in monkey embryo); therefore, Cas9/sgRNA screens were established as a tool for genetic analysis in mammalian cells (Shen 2014). In 2015, US and UK research scientist, Medical Research Council (MRC) declared their support for using GEd strategy for human cells (Charo 2015). After that, the International Summit on human gene editing was met to discuss about the medical and ethical issues (Charo 2016). New protein-Cpf1 was invented in the same year, which made gene editing become simpler (Koonin et al. 2017). Moreover, US scientists reported about the modified CRISPR-Cas9 technique with fewer off-target effects. The first clinical trial of the genetically modified human embryo was approved in 2016 (Cyranoski 2016; Reardon 2016). A new base editing technique was discovered in 2016 by US scientists, offered a new approach where any gene can modify without cleavage of double-stranded DNA as well as without donor DNA template (Rees and Liu 2018; Porto et al. 2020; Bharat et al. 2020). For RNA editing, a new CRISPR approach was identified in 2017 (Cox et al. 2017; Adli 2018). In 2018, Weissman’s lab created a new GEd strategy called CRISPRa (for ‘activation’), which activate the gene expression, and they also made CRISPRi (for ‘interference’) technology (Kampmann 2018). In 2018, a group of scientists identified pre-existing Cas9 antibodies in cells, leading to immune issues during gene therapy employing CRISPR-Cas9 (Crudele and Chamberlain 2018; Wagner et al. 2021). Same year another Cas variant Cas14 (a-c) was identified (Harrington et al. 2018) In 2019, Chinese researchers declared their first gene-edited human baby (Wang et al. 2020). Newly developed ‘search and replace’ tool for GEd known as prime editing was discovered in 2020 (Hampton 2020). In the same year 2020, a Chinese researcher was convicted for employing CRISPR-Cas9 in a human baby (Cyranoski 2020). Moreover, in 2020 for the first time, one patient received gene editing therapy employing the CRISPR-Cas9 approach (He 2020; Ledford 2020). In early 2020, a novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) outbreaks rapidly evolved into a global pandemic. For the detection and quantification of SARS-CoV-2 RNA, a CRISPR-Cas13-based approach was employed (Konwarh 2020; Kumar et al. 2020). In 2020, Jennifer Doudna and Emmanuelle Charpentier were jointly awarded the Nobel Prize in Chemistry for the identification of an efficient method in GEd known as the CRISPR-Cas9 technique (Ledford and Callaway 2020). The CRISPR ‘on-off switch’- a new genome-editing approach was discovered by MIT and UCSF researchers in 2021, successfully implicated in Alzheimer disease (James et al. 2021). Any part of the targeted genome can be silent via controlling the gene’s expression without altering DNA sequences. Therefore, unlike first-generation GEd tool, the CRISPR-Cas9 technology has empowered researchers with an unprecedented toolbox via breakthrough discoveries and methodological advancements in science.
9.3 CRISPR-Cas: New-Fangled Dawn in Genome Editing
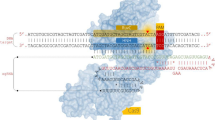
CRISPR-Cas is the most efficacious and ease-to-design editing tool, which generate a buzz in the field of research in current times. This is one of the crucial tools in an endless arms race between bacterial and archaeal hosts and viruses (Newsom et al. 2021). The CRISPR immunity gets triggered when a virus’ foreign genetic material (DNA/RNA) is introduced into bacterial cells. The bacterial cell effectively produced specialized molecules (Cas protein) that can recognize the past similarity of foreign DNA and destroy them as antibodies work. The defense mechanism of this system comprises into three-stage process, i.e. (i) Adaptation: small DNA sequences (protospacers) of foreign plasmid are chosen and incorporated into the particular CRISPR locus of the host genome; (ii) Biogenesis of crRNA: multiple gRNA spacers and their repeats are transcribed into a precursor RNA and processed into mature gRNAs. Targeting complexes are produced via fastening of gRNAs with the Cas enzyme, which contain a distinctive spacer sequence resembling foreign target DNA; and (iii) Interference: Cas nuclease starts searching the unique sequences complementary to the gRNA. Cas nucleases fasten up to the gRNA resemble target foreign DNA site via complementary base pairing and cleave the targeted DNA sequences. By utilizing this machinery, bacteria generated the ability to avoid transcribing the matching targeted viral DNA, making its genome resistant to viral invasion. As research gains grounded, numerous CRISPR-Cas systems have been identified for GEd. All these systems have their own attributes, for instance, variation at PAM regions, varying sizes of Cas protein, and different cleavage sites. Amongst all, the type II CRISPR-Cas9 system provided the most simple, versatile precision editing in crop plants. This system required only two key molecules, i.e. Cas9 endonuclease and gRNA: fusion of CRISPR RNA (crRNA, a 20-nucleotide sequence complementary to the target DNA) and trans-activating crRNA (tracrRNA, acts as a binding scaffold for the Cas9 endonuclease). It also should be noted that the gRNA can be expressed as synthetic sgRNA, where the crRNA and tracrRNA are fused into one molecule for ease of expression (Fig. 9.1). Widely accepted SpCas9 (Streptococcus pyogenes) comprises a conserved core with two major big globular recognition lobe, for instance, REC (recognition) and a NUC (nuclease) lobe for nucleic acid binding. Wherein, the REC (functional domain of Cas9) contains bi-partite domain, for instance, REC1 & REC2 and bridge helix cd domain. It was revealed that base pairing between the ligand DNA strand and the seed region of gRNA (up to 8–12 bp) triggers the development of RNA–DNA heteroduplex, which occupied by both NUC and REC lobe (Anders et al. 2014). The small NUC nuclease lobe comprises a highly conserved RuvC- & HNH- and PI- domain (arginine-rich alpha-helical bridge helix) (Hsu et al. 2014). Simultaneously, RuvC and HNH nick the complimentary and non-complimentary strand in the target sequence, introducing double-strand breaks (DSBs) (Nishimasu et al. 2014). According to previous studies, the PI domain plays a crucial role in the PAM site (5′-NGG-3′) recognition because of having a tryptophan-rich flexible loop (Jinek et al. 2014). At 3 bp prior to PAM sites, the assembled CRISPR-Cas complex created DSBs. DSBs can be repaired at defined positions by integrating numerous alterations utilizing DNA repairing machinery, i.e. HDR and NHEJ (Fig. 9.2).
CRISPR mechanism in action: Natural vs Engineered CRISPR system. This system required only two key molecules, i.e. Cas9 endonuclease and gRNA. gRNA is fusion of CRISPR RNA (crRNA, a 20-nucleotide sequence complementary to the target DNA) and trans-activating crRNA (tracrRNA, acts as a binding scaffold for the Cas9 endonuclease). Inactive Cas9 is become active when bind with gRNA. In synthetic sgRNA, the crRNA and tracrRNA are fused into one molecule for ease of expression
Schematic representation of CRISPR/Cas9-based DSBs repair mechanism, including NHEJ and HDR-mediated repair pathways. The CRISPR-associated endonuclease Cas9 generated DSBs in the target DNA. NHEJ pathway results in random indels via gene disruption at the target site. HDR pathway uses homologous donor DNA sequences for accurate insertions or base substitutions between DSB sites. DSB double-strand break; NHEJ non-homologous end joining, HDR homology donor repair, Indels insertions and deletions
NHEJ is the primary DSB fixing pathway in plant cells and is comparatively effortless to exploit for GEd (Lieber 2010; Pannunzio et al. 2017). This error-prone pathway generally introduces indel mutations (insertions and/or deletions) by disrupting the targeted DNA, resulting in gene knockout (KO). The CRISPR-Cas9-based KO is utilized in gene function study, and modifying a variety of beneficial traits, for instance, stress resistance (Singh et al. 2020); disease resistance (Schenke and Cai 2020); higher yield (Huang et al. 2018; Ma et al. 2019; Liu et al. 2021; Tabassum et al. 2021); nutritional enhancement (Zhang et al. 2018a; Sanchez-Leon et al. 2018; Ku and Ha 2020; Dong et al. 2020; Huang et al. 2020; Dong et al. 2019; Xu et al. 2020; Yang et al. 2020; Zeng et al. 2020; Kaul et al. 2020a, b; Sashidhar et al. 2020; Tiwari et al. 2020); and male sterility (Chen et al. 2021). For the achievement of successful KO, it is recommended to target early exon because functional activities of a gene will be less if indel mutation is generated in either 3′ end of exon sequences or intron region. Nevertheless, due to alternative splicing if target gene enciphers various proteins, then frameshift mutation or stop codon introduction in early exon may not reveal gene KO. In this situation, complete gene deletion can be possible by utilizing the multiplex CRISPR-Cas9 KO strategy by targeting the gene’s 3′ and 5′ end. For example, (115–245) kb in size chromosomal deletions were generated via gene cluster deletion in rice (Zhou et al. 2014) employing multiplex CRISPR-based KO strategy. Recently, multiplex CRISPR-Cas9 system utilized for simultaneous KO of multiple genes and revealed de novo domestication of wild tomato (Zsögön et al. 2018; Xie and Liu 2021). Gene KO is extremely difficult in polyploidy species due to its gene functional redundancy. It was successfully utilized in hexaploid wheat to develop fungal-resistant wheat by KO of disease susceptible S-gene (Wang et al. 2014; Wang et al. 2018a). Corteva Agriscience generated amylopectin rich (waxy) corn via KO of Wx1 gene (DuPont Pioneer 2016). Similarly, two japonica rice varieties (glutinous sticky) were achieved through Waxy (OsWx) gene KO (Yunyan et al. 2019). Moreover, amylose-rich rice grain was revealed via targeted modification of the SBEIIb gene (Sun et al. 2017). Gaoneng et al. (2017) developed fragrance enriched rice via targeted KO of the BADH2 gene (negative regulator for aroma production). Additionally, KO of OsERF922 gene generated blast-resistant rice lines was reported by Wang et al. (2016). Edited rice lines with pale green colour in leaf were generated via KO of chlorophyll biosynthesis regulated gene OsCAO1 gene (Jung et al. 2021). Interestingly, OsHOL1 plays a major role in the production of methyl iodide, and KO of this gene abolished methyl iodide emissions from rice plants (Carlessi et al. 2021). Targeted KO of TERMINAL FLOWER 1 (TF1) gene in Brassica napus altered the flowering time and plant architecture (Sriboon et al. 2020). Targeted KO mutations of HvHPT and HvHGGT gene rendered a high level of vitamin E (tocopherol) in barley (Zeng et al. 2020). According to Li et al. (2019), KO of numerous genes, i.e. SVP, AP1, and TFL elicited floral features advancement in Arabidopsis. High-oleic acid content was generated in allotetraploid cotton (Gossypium hirsutum L.) (Chen et al. 2021) and tobacco (Tian et al. 2020) via KO of GhFAD2 and NtFAD2–2 genes, respectively, as well as Monounsaturated Fatty Acid (MUFAs) contents enhancement in Hexaploid Camelina sativa seed oil was generated through FAD2 Gene KO using CRISPR-Cas9 (Lee et al. 2021). Functional KO of StDND1, StCHL1, and StDMR6–1 generated potatoes highly resistance against late blight disease (Kieu et al. 2021).
On the other hand, HDR mechanism introduces specific base pair substitution point mutations via target DNA recombination with complementary HDR template (Reis et al. 2014; Sander and Joung 2014). Knocking-in of targeted and precise sequences has been more challenging. Repairing Cas9-induced DSBs or nicks using HDR-mediated pathway makes GEd more accurate. Thus, unlike NHEJ, in case of knockin, the incision must be embedded precisely, without extra insertions/deletions (indel) mutation. Unfortunately, GEd frequency employing HDR mechanism is relatively low in plants in comparison to NHEJ. Amongst numerous approaches to recurrence, the HDR efficiency in plants, the utilization of mastrevirus (Geminiviridae) vectors for delivery of donor template is the most successful generated so far. This method was first demonstrated in tobacco to develop bean yellow dwarf virus-resistant (Baltes et al. 2014). The HDR template frequency was increased dramatically in nucleus due to replication of donor template, revealing a high editing frequency. Later on, it was employed in tomato ANT1 gene to precisely insert a promoter upstream of the gene, resulted in pigment accumulation in foliage, flowers, and fruits via controlling anthocyanin biosynthesis (Čermák et al. 2015). Moreover, point mutation was introduced in the potato ALS1 gene, conferring herbicide resistance (Butler et al. 2016). A viable alternative method is delivering a large copy of the donor template into plant genome employing biolistic approach. This approach was successfully utilized in rice and maize (Baltes et al. 2015; Gil-Humanes et al. 2017; Wang et al. 2017) resulted in higher precise edits. In line with this, numerous advancements had been developed, for instance, in Arabidopsis the absence of a repair protein, KU70/80 may lead to a 5–16 fold enhancement in HDR editing frequency via suppressing the NHEJ repair pathway (Endo et al. 2016). Moreover, Lu et al. (2020) discovered a tandem repeat-HDR (TR-HDR) approach for high frequency targeted sequence replacement, wherein the precise editing frequencies ranged from 3.4 to 11.4%. According to Shi et al. (2017), the promoter swapping, for instance, GOS2 promoter by the native ARGOS8 promoter employing CRISPR-Cas based GEd via HDR approach generated drought tolerance in maize. Tomato lines with higher self-life were generated via T317A substitution in the ALC gene (Yu et al. 2017). Newly developed RNA-mediated CRISPR/Cpf1-based approach also rendered efficient, targeted gene insertion in tomatoes (Vu et al. 2020). Therefore conjointly, CRISPR-Cas9 and CRISPR/Cpf1 may overcome all difficulties for precise gene knockin via HDR mechanism for crop plant enhancement.
9.4 Novel Technical Breakthrough of Genome Editing in Plants
The necessity to genetically improve crop varieties became the reason for the discovery of target-specific endonucleases (TSENs) and since 2005 there has been a significant improvement and addition of new tools/techniques in the GEd toolkit. The genetic engineering field experienced another boost with the discovery of the CRISPR-Cas9 system which back in 1987 was recognized as the bacterial immune system (Ishino et al. 1987). The last decade has witnessed the evolution of this technique to reduce the bottleneck in terms of efficacy, efficiency, applicability, and other already discussed shortcomings. Substantial diverseness in genes, loci configuration, and action mechanisms of CRISPR-Cas approach made their classification a formidable task. An updated classification was reported by MaKarova et al. (2020), which include 2 classes, 6 types, and 33 subtypes; they identified novel class 2 CRISPR-Cas systems including 3 types and 17 subtypes (Table 9.1). Class 1 systems contain ~90% of all discovered CRISPR-Cas loci constituting type I, III, and IV (Makarova et al. 2015). The Class 2 system contains 10% of CRISPR-Cas loci (Makarova et al. 2015) and clearly differentiating into type II, V, and VI (Makarova et al. 2020). Numerous Cas9 variants are identified in recent years to broaden the opportunity of genome alteration. Thus to greatly expand the range of targets, different orthologs of Cas9 were reviewed, and VQR (5′-NGA-3′) and VRER (5′- NGCG-3′) variants of Cas9 were developed for plants (Hua et al. 2016). In the same line of study, an ortholog from Francisella novicida (Fncas9) was engineered to recognize 5′-YG-3′ PAM. FnCas9 is also known to target the RNA substrate, consequently it can be utilized to gain viral resistance in plants (Zhang et al. 2018b). Later to increase the penetrability of the Fn Cas9, proximal CRISPR (proxy-CRISPR) was developed (Chen et al. 2017). To increase the target specificity and reduce the off-target effect, Cas9 nickases mutants (nCas9) came into the picture by introducing point mutations like D10A in RuvC (Jinek et al. 2012) and N863A or H840A in HNH domains (Nishimasu et al. 2014). Along the same line of work, Satomura et al. (2017) designed a ‘CRISPR Nickase system’ (CNS) to target sequences that were non-editable with the conventional CRISPR-Cas9 tool. Furthermore, the inducible Cas9 or split Cas9 can be used for temporally and spatially restricted Cas9 expression (Zhou et al. 2018; Carlson-Stevermer et al. 2020).
The continuous endeavour led to the discovery of class II type V CRISPR from Prevotella and Francisella 1- Cpf1/Cas12a was a potential alternative to Cas9 primarily because it could target AT-rich (5′-TTTN-3′) PAM instead of GC rich PAM (Doudna and Charpentier 2014). Besides cis-cleavage of the target double strand, it can also cleave non-specific ssDNA in trans (Swarts and Jinek 2019) which contributed to the invention of a sensitive nucleic acid detection technique, i.e. DETECTOR (Li et al. 2018a). Recently, Zhang et al. (2021) have contributed exceptionally with the discovery of six highly efficient orthologs (ErCas12a, Lb5Cas12a, BsCas12a, Mb2Cas12a, TsCas12a, and MbCas12a) of Cas12a. Similarly, a related enzymatic activity harbouring Cas12b (C2c1) from Alicyclobacillus acidiphilus, i.e. AaCas12b was prospected as a potential add-on to the tool kit. Another class II type VI-A Cas protein, i.e. Cas13a (C2c2) is an effective tool that possesses RNA-guided RNase activity (Abudayyeh et al. 2016). Single strand RNA (ssRNA) targeting LshCas13a (Leptotrichia shahii) and other orthologs (b,c,d) have two HEPN (Higher Eukaryotes and Prokaryotes Nucleotide-binding) domain with no requirement of PAM (Bandaru et al. 2020). Cas13a has been utilized for RNA/transcript knockdown and RNA editing (REPAIR and RESCUE; Cox et al. 2017; Abudayyeh et al. 2019). Moreover, Cas13a has been utilized for SHERLOCK, PAC-MAN, and SARS/Covid 19 detection kits (Gootenberg et al. 2018; Joung et al. 2020; Zhang et al. 2020). In plants, it can be utilized to gain viral resistance against specific viral pathogens (Abudayyeh et al. 2017). Another class II type V effector, i.e. the Cas14 family (Cas14a-c: 400–700 amino acids) present in archaea came as a significant discovery (Harrington et al. 2018; Savage 2019). Cas14s can target both ssDNA and dsDNA with no PAM or AT-rich PAM (5′-TTAT-3′) requirement (Karvelis et al. 2019). Due to sensitivity towards mismatching, it can be used for high precision SNP genotyping and because of trans cleavage activity it can be used as a Cas14-DETECTOR and to gain viral resistance in plants (Aquino-Jarquin 2019; Khan et al. 2019a). With continued hustle to discover better alternatives for GEd, the database mining led to the discovery of smaller Cas proteins such as Cas12f and the features closely related to the previously known Cas 14s (Karvelis et al. 2020). The recent classification thus unifies these proteins together Cas12f1 (Cas14a and type V-U3), Cas12f2 (Cas14b), and Cas12f3 (Cas14c, type V-U2 and U4) and expands the utility tools in the GEd artillery (Makarova et al. 2020).
Further expanding the smaller type V effectors family, DpbCasX (Deltaproteobacteria) is one such mini (~980 aa) novel protein (Liu et al. 2019a). CasX (alias Cas12e) is a dual RNA (crRNA and tracrRNA) guided protein (naturally combined into single-guide RNA; sgRNA) targeting dsDNA adjacent to 5′-TTCN-3′ PAM to generate 10 nt staggered break (Yang and Patel 2019). However, it shows the nominal trans activity as compared to other type V effectors which highlight structural differences between Cas X and other enzymes. Recently, in Doudna’s lab a supercompact CRISPR-CasΦ system encoded by bacteriophage genome has been discovered, where a bacteriophage uses the system to target other competing phages. CasΦ (Cas12j) also has a C-terminal RuvC domain but shares no similarity (<7% amino acid identity) with type V effectors, rather it is remotely related to the TnpB enzymes. The CasΦ locus lacks the spacer acquisition enzymes such as Cas1, 2, and 4 which results in a really compact CRISPR array and the locus also lacks the presence of tracrRNA. CasΦ represents the consolidated form of the CRISPR-Cas system and thus can be utilized to its full potential for genome manipulation (Pausch et al. 2020). With the discovery of such versatile, flexible, and miniature (400–1093 amino acids) effectors, the Cas12 family is expanding and till date, there are 11 subtypes of type V which has been reported, namely Cas12a to k (Li et al. 2021) and a subtype V-U which is more closely related to transposon TnpB. Cas12a, Cas12b, Cas12e, Cas12h, and Cas12i specifically target dsDNA with PAM assisted unwinding (Yan et al. 2019). However, an ortholog Cas12g (thermostable) was reported to initially target ssRNA and then indiscriminately degrade both ssDNA and ssRNA (Chen et al. 2018). Till now, class 2 effectors have dominated the terrain of GEd primarily because it utilizes single subunit protein effectors, whereas class 1 CRISPR-Cas system utilizes multiple subunit protein effectors (Makarova et al. 2018). Type III effectors of class 1 are known to target RNA substrates and hold the potential to be developed into diagnostic tools or to attain tolerance against viruses or mobile genetic elements (MGE) (Samai et al. 2015; Staals et al. 2014) in any system. Type III effectors are divided into III-A (Csm), III-B (Cmr), III-C, III-D subtypes, and the common feature between these subtypes is the presence of Cas10 (Csm1 or Cmr2) (Burmistrz et al. 2020) in the complex. Cas10 predominantly has two domains (Makarova et al. 2018) notably the palm domain (the cyclase activity of palm domain is absent in type III-C effectors) and a nuclease HD-type (unavailable in type III-D effectors) domain (Zhu et al. 2018). Typically, this multi-subunit complex protein effector is composed of two parallel filaments from which the first filament is generally made of six subunits of Cas7 protein and the other is made up of three subunits of Cas11 homolog (Csm2 or Cmr5) protein. The crRNA is stretched in between these filaments (Lintner et al. 2011) and the 5′- end having the handle derived from repeat is capped by Cas10 and Cas5 (Csm4 or Csm3) proteins (Staals et al. 2014), whereas the maturation of crRNA from pre-crRNA is catalysed by the Cas6 (Nickel et al. 2018) protein. Cas7 as a family of protein (members like Thermofilum pendens Csc2 protein) has members in both type I-D (commonly present in Archaea and Cyanobacteria) and type III branches of classification (Staals and Brouns 2013; Cai et al. 2013). Moreover, the protein organization in CASCADE (CRISPR-associated interference complex type I) and type III complex is similar, and proteins of type I-D have HD domain fused to the Cas10 (type III protein) and thus Cas7 is considered as the evolutionary link between type I and type III CRISPR-Cas system (Hrle et al. 2014). Exceptionally, the type III system has three different nuclease activities, and primarily it possesses sequence-specific RNase activity where acidic residues of the RNA-recognition motif (RRM) of Cas7 targets a specific RNA sequence (Estrella et al. 2016). The CRISPR-associated Rossman fold (CARF) located at the N-terminal of Csm6 sense the presence of the cyclic oligoadenylate while the C-terminally located higher eukaryotes and prokaryotes nucleotide-binding (HEPN) domain non-specifically cleaves the ssRNA (Niewoehner and Jinek 2016) molecule. Considering the potency of Csm6 protein, it has been included in the SHERLOCKv2 and this resulted in the three fold increase in the efficiency of the technique by improving the reporter signal (Gootenberg et al. 2017; Kellner et al. 2019). With continued exploration and screening over 11 billion protein sequences revealed the existence of a single-protein effector under type III-D2 CRISPR-Cas system, referred as Cas7x3 which have three Cas7 protein fused into a single protein (Özcan et al. 2021). In consonance, a novel breakthrough has resulted in the discovery of a programmable type III RNA targeting single-protein effector termed as Cas7–11, structurally having four Cas7 proteins fused to a putative Cas11 protein (Makarova et al. 2020). DiCas7–11 from Desulfonema ishimotonii is a programmable RNase with no reported collateral activity. The discovery of this protein further expands the classification nomenclature by adding a type III-E subtype to the previously known subtypes. Both Cas7x3 and Cas7–11 process their own pre-crRNA into mature crRNA for targeting specific sequence template and not even display any toxic effect in mammalian cells (Özcan et al. 2021). However, they still need to be developed into programmable CRISPR-Cas tools to utilize their full potential for GEd across different systems including plants.
Introducing foreign DNA and generating of DSBs in any system for GEd raised some regulatory concerns which led to the evolution of the DNA-free GEd strategy. Under this, the ribonucleoproteins (RNP) which are pre-assembled Cas nucleases with the target-specific gRNA are delivered into the target system to achieve the desired GEd in plant and animal systems (Woo et al. 2015; Wu et al. 2020). The use of CRISPR-Cas tool in prokaryotic (Qi et al. 2013) and eukaryotic (Gilbert et al. 2013) systems introducing DSB leads to unexpected changes and toxicity. Precision transcriptional regulation without the introduction of any DSBs, i.e. without changing the underlying DNA sequence, with the strategies like CRISPRi and CRISPRa has revolutionized the field of genetic engineering (Liu et al. 2019b). In CRISPRi and CRISPRa, a dCas9 is fused with transcriptional effector to either repress (repressor like Kruppel associated box, or KRAB) or activate (activators like VP64 and p65) the gene expression (Lawhorn et al. 2014; Mali et al. 2013a, b). The newly developed customizable epigenome memory writer ‘CRIPSR on-off’ technique can alter gene expression by generating heritable epigenome modification. CRISPRoff is a fusion protein with dCas9 with DNA methyltransferase (DNMT1) and KRAB domains to silence the gene expression. However, the modifications are specific, tunable, and reversible as the methylation can be removed (inhibitor of DNMT1, i.e. 5-aza-2′-deoxycytidine (5-aza-dC)) by CRISPRon, and gene expression can be activated via recruitment of the transcriptional machinery. Genome-wide screen helped to find the targetable genes and showed that genes lacking the CpG islands can also be silenced with the CRISPR-off technique (Nunez et al. 2021). ‘CRISPR on-off’ is a complementary technique to the already existing CRIPSRi, CRISPRa, and CRISPR nuclear approaches.
Base editors (BEs) in conjunction with the CRISPR-Cas tool have been used for precise, specific single base modification with no induction of DSB and as an alternative to HDR-based GEd (Komor et al. 2016). dCas9 or any inactive RNA-guided Cas protein with cytidine base editor (CBE; cytidine deaminase) can catalyse target specific C-to-U (Uracil recognized as T) base substitution which results in C-G to T-A base pair conversion (Rees and Liu 2018) and with adenine base editor (ABE; deoxyadenosine deaminase), it can catalyse the A-to-I (Inosine; recognized as G) base substitution which results in A-T to G-C base pair conversion (Gaudelli et al. 2017), respectively. CRISPR-BE has gone under severe optimization and development and in a recent generation a D10A nCas9 (to induce nick in unedited strand) fused with cytidine deaminase enzyme, i.e. rAPOBEC1 (rat apolipoprotein B mRNA editing enzyme) or to Lamprey cytidine deaminase (pmCDA1) for activation-induced cytidine deaminase (AID) at N-terminal, and two copies of uracil DNA glycosylase inhibitor (UGI) at C-terminal are used for base editing. The fusion of UGI increases the efficiency of editing in the case of C-to-U conversion as it helps in retaining the U in the target sequence till the next cycle of replication by inhibiting the inherent conversion of U-to-C again by uracil DNA glycosylase (UDG) (Abdullaha et al. 2020). Whereas, nCas9 (D10A) was also utilized in conjunction with the TadA (tRNA adenosine deaminase) and TadA* (modified at K157N, I156F, E155V, R152P, D147Y, S146C, H123Y, D108 N, A106V, L84F, R51L, P48A, H36L, W23R) domains connected via varying linker length, for ABE optimization and development (Bharat et al. 2020). Along with DNA base editing, RNA base editing can be achieved with the RNA directed RNA targeting dCas13. RNA editing comprises REPAIR (RNA editing for programmable A-to-I (G) replacement; catalysed by ADARs) and RESCUE (RNA editing for specific C-to-U exchange; catalysed by cytidine deaminase) techniques in plants (Abudayyeh et al. 2019). Although the base editing approach has faced few challenges in terms of off-target, range of editing, and bystander editing (Jeong et al. 2020). These shortcomings have led to the revolutionary discovery of prime editing (PE) which is based on the search and replace ideology and is a template free strategy (Anzalone et al. 2019). PE2 system is dependent on an amalgamation of the nCas9 (H840A), reverse transcriptase (RT; M-MLV from mouse-murine leukaemia virus), and the prime guide RNA (pegRNA). pegRNA have a primer binding site (PBS) sharing sequence complementarity to the sequence of the nicked DNA strand upstream of PAM and a reverse transcriptase template strand (RT strand). The 3′ flap is utilized as the primer to transcribe the desired sequence (written in the RT template), whereas the 5′ flap is cleaved via structure-specific host endogenous flap endonuclease (FEN1; Flap endonuclease Homo sapiens). Later the edited strand is ligated after 5′ flap digestion forming a heteroduplex of edited and unedited strands co-exist (Anzalone et al. 2019). The induction of a second nick on the unedited strand 10–12 nt away from the original pegRNA cut on the edited strand resulted in the development of the PE3 system. In PE3 when the unedited strand is repaired after induction of the second nick, it leads to the formation of the homoduplex of the edited dsDNA (Anzalone et al. 2020). In order to avoid incorporation of indel mutation by PE3 while repairing, in PE3b the second nick was introduced after successful completion of the flap resolution and editing (Kantor et al. 2020). PE till now displays really low events of off-target in any system (Scholefield and Harrison 2021). In a study by Lin et al. (2020), the efficiency of the plant PE system increased at some locus by using the PPE-Ribozyme (PPE-R) system where the PE protein transcript is expressed by Polymerase II (Pol II) and pegRNA is processed by the ribozyme. Prime editing has come as a boon in the field of GEd and has expanded the toolbox for deep genome modification with enhanced efficiency, specificity, and tenacity even in polyploidy genomes such as wheat, as well (Lin et al. 2020).
9.5 Revisiting Challenges and Impediments of CRISPR-Based Approach for Precise Genome Editing
CRISPR is regularly portrayed as ‘cut and paste’ approach for genes, but the actual procedure is not that easy. However, further research is needed to gain a deeper understanding of the CRISPR-Cas process and its neoteric uses in plants. To date, researchers face umpteen obstacles related to utilizing the CRISPR approach in plant research, including hurdles in GMO regulation. Recently, researchers have achieved huge achievements utilizing CRISPR in its native and closely related organisms. But, employing CRISPR into bigger genomes containing complex organisms has accompanied its own set of difficulties. Some plants have multiple copies of each chromosome, for instance, hexaploid wheat (6 copies), strawberries (up to 10 copies), which is become strenuous to engineer compared to humans and animals. Subsequently, the probability of getting target gene editing in each copy decreases as the quantity of chromosome copies increases (Yang et al. 2020). Scientists are improving traditional CRISPR-Cas workflow by employing varying modifications so that multiple copies of the identical gene can be altered at once (Wilson et al. 2019; Lin et al. 2020; Jouanin et al. 2020; Smedley et al. 2021). Lamentably, this type of alteration sometimes create off-target mutation/s. Screening of accurate mutation and potential off-target sites is a very sensitive and significant challenge in the field of gene editing.
Earlier PCR/RE strategy was utilized to screen mutation in edited plants (Shan et al. 2014). The T7 endonuclease I (T7E1) assay was employed to detect off-target mutations; however, it is neither feasible nor cost-effective for large-scale screening due to its deprived sensitivity. Therefore, RNA-guided endonucleases, i.e. SpCas9- or FnCpf1- based PCR/RNP method for identifying indel/s, overcome the PCR/RE strategy (Liang et al. 2018). Unlike T7EI, this PCR/RNP-based technique can differentiate the mutant types, i.e. homozygous, heterozygous, bi-allelic, and mosaic mutants. It is also a SNPs independent mutation detection method essential for polyploidy plants like wheat (Liang et al. 2018). Numerous web-based approaches, for instance, deep sequencing (mutation detection range: 0.01–0.1%), genome-wide, unbiased identification of DSBs facilitated by sequencing (GUIDE-seq), RNA-guided endonucleases (RGEN), had been widely adapted (Wu et al. 2014; Zhang et al. 2015; Tsai and Joung 2016; Kosicki et al. 2018). Consequently, different bioinformatics-based programs (TALE-NT, CAS-OFF Finder, PROGNOS) have been developed to profile off-target mutations via CRISPR-Cas nucleases (Fine et al. 2013; Listgarten et al. 2018; Minkenberg et al. 2019). Recently, genome-wide off-target edit frequencies were identified using the whole-genome resequencing (WGRS) approach in rice, maize, cotton (Tang et al. 2018; Lee et al. 2019; Li et al. 2019).
In addition, the plant regeneration and transformation approach is quintessential for delivering the editing reagents into plant cells for genome editing. Wherein, genotype-dependency is one of the major bottlenecks in completely appearing the incredible capability of genome altering in plant species (Alpeter et al. 2016). The development regulator (DR) genes of maize: Baby Boom (Bbm) and Wuschel2 (Wus2) in combination with phytohormones lead to enhance the transformation efficiency in plants (Lowe et al. 2016; Maher et al. 2020). Moreover, Agrobacterium-mediated transformation is frequently restricted due to the narrow range of genotypes within a species. As well as plant growth conditions, co-incubation time & temperature, pre-treatment with phytohormones, variability of Agrobacterium tumefaciens are well-known factors to affect transformation efficiency (Zambre et al. 2003; Gelvin 2006). However, these shortcomings can be overcome by utilizing the biolistic transformation approach due to its efficient and potent high transformation efficiency (Wu et al. 2015; Li et al. 2019; Kaul et al. 2021). The CRISPR-Cas-based genome editing in crop plants can only be manifested by fine-tuning the targeted gene or genetic elements (Kwon et al. 2019; Oliva et al. 2019).
Identifying the targets (quantity) due to an inadequate understanding of biological networks and their interactions with environmental factors is another critical obstacle for CRISPR-Cas-based plant genome editing. Applications of multidisciplinary strategy, for instance, genome-wide, and high-throughput functional genomics strategy for identification of beneficial agronomic traits harbouring targets in both the model and non-model crop plants are crucial for genome editing (Lu et al. 2017; Meng et al. 2017; Araus et al. 2018). Alongside, the achievement of high base substitution efficiency via fragment knockout and knockin of homology donor repair (HDR) is an important implication for crop enhancement. However, precision editing in plants employing an HDR-based approach is a significant challenge due to its lower editing potential. Optimizing the optimal quantity and the effective delivery methods of the donor DNA template might ease the base substitution editing approach (Kaul et al. 2020a, b).
The presence of protein inhibitors of CRISPR-Cas systems, known as anti-CRISPR (Acr) proteins, enables the generation of more precision in CRISPR-Cas-based GEd. More than 50 Acr proteins are currently shown to interact with CRISPR-Cas variants, for instance, Cascade-Cas3, Cas9, Cas12, and Cas13 (Dolgin 2019; Marino et al. 2020). The functional mechanism of ACr proteins is one of three ways: firstly, prevention of DNA binding: Acr either blocks or reduced Cas9’s interaction with the PAM recognition site; secondly, prevention of crRNA loading: the interaction of Cas9 may disrupt or prevents the proper integration of the crRNA-Cas complex; and thirdly, and blocking of DNA extraction: Acr binds with HNH endonuclease domain of Cas9 and inhibits its activity (Dong et al. 2017; Zhu et al. 2019). However, Acrs can be used to eliminate allergies in unidentified areas (Aschenbrenner et al. 2020; Shin et al. 2017), unwanted mutations in unintentional cell types or tissues (Hirosawa et al. 2019; Hoffmann et al. 2020). In addition, Acrs (AcrII4s) can be employed as a ligand biosensor to detect and measure CRISPR-Cas9 RNP affinity reagents (Johnston et al. 2019). Similarly, other alternative approaches also being developed to prevent Cas9 activity, for instance, nucleic acid-base inhibitors and (Barkau et al. 2021) and smaller molecules of inhibitors (Maji et al. 2019). Despite precision genome alteration, Acrs provide a prospect to exploit their ability to inhibit Cas9 and to address other engineering limitations of the Cas9 genome.
Comparative genomic analysis revealed that CRISPR and its associated proteins, especially Cas9, were present in umpteen bacterial phylogenetic groups (Lillestøl et al. 2006; Makarova et al. 2006). Cas9 from S. pyogenes showed 23 to 58% and 35% similarity to Cas9 proteins from Streptococcus thermophilus and Lactobacillus plantarum, respectively. Those organisms were utilized for various human edible food processing, for instance, yoghurt, cheese, kefir, fermented drinks, and so many (Settachaimongkon et al. 2014; Sidira et al. 2017; Behera et al. 2018). Thus, humans were exposed to Cas9 protein in their diet long before the development of CRISPR-Cas9 genome editing. Additionally, Cas9 from S. pyogenes showed 80% sequence similarities with a variety of gram-positive and negative bacteria that present in human body (Qin et al. 2010; Louwen et al. 2014). The above-mentioned findings do not imply that human exposure to Cas9 used in genome-editing planning is insignificant (Pineda et al. 2019). However, the biosafety risk assessment regarding human exposure to Cas9 after consuming GEd plants product requires further testing.
Another hindrance is the adoption of edited crop plants success in natural field conditions. An enormous number of researches on genome editing reported so far, but the majority is only about proof of concepts in the greenhouse environment. The performances uncertainties of the edited plants are still existed due to the lack of field trials. Despite all these challenges and impediments, the CRISPR-Cas9 approach is considered the most promising tool due to its precision editing. This approach incorporates numerous heritable traits in plants, which may produce modified plants similar to those developed through conventional breeding. CRISPR-Cas9 strategy leads towards a progressive change via high yielding crop plant production to meet food security globally.
9.6 Overcoming Challenges for ‘Off-Target’ Mutations
Alteration of plant genome employing the CRISPR-Cas approach sometimes resulted in off-target effects (alteration of the additional region beyond the target region of the genome), which is a pivotal impediment of this application. However, numerous strategies can be employed to minimize off-target mutations. Till date, above than 30 plant varieties (~100 attribute traits) have been edited successfully employing the CRISPR-Cas9system. The precession binding of Cas9 depends on the 7–8 nucleotides seed sequence and the existence of the PAM close to the target sequence, but unwanted insertions/deletions could happen in the genome (Hajiahmadi et al. 2019). To improve genome-editing efficiency, scientists devised in vivo/vitro biological analysis and algorithm-based computational methods to uncover and increase gene editing efficiency. Promoters and target genes are essential elements involved in the regulation of gene expression by modifying transcription factors (TFs) via RNA polymerase recognition. The specificity of a promoter is essential for controlling transgenic expression in target tissues or throughout the plant. Over the last few years, constitutive promoters like the cauliflower mosaic virus (CaMV35S) promoter (Paparini and Romano-Spica 2006; McCaw et al. 2021) and the maize ubiquitin (pZmUbi) promoter (Xu et al. 2018; Samalov and Moore 2021) have already been used. In dicot plants, the CaMV35S driven promoter showed a high level of expression, in contrast, monocot plants employ pZmUbi promoter more effectively. In Arabidopsis, the promoters of (rd29A and rd29B) genes showed well performance to a variety of stress stimuli, such as salinity and drought (Bihmidine et al. 2013). Salt induces activity in the BADH promoter from Suaeda liaotungensis (Zhang et al. 2008). The Rab16A promoter might up-regulate GUS expression in transgenic rice under salt stress (Rai et al. 2009). The TsVP1 promoter from Thellungiella halophila is effective in almost all tissues except the seeds, and salt stress in leaves and roots, particularly root tips (Sun et al. 2010). DREB2 coordinated expression of transcription factors will generate successful regulatory activity; thus, monocotyledonous plant promoters’ operations are higher in monocots as in dicots. Heat-shock protein 17.5E (Hsp17.5E) gene promoter from soybean (Glycine max) has been utilized to direct Cas9 expression in rice for genome editing. Several methods have been described to reduce off-target mutations; primarily, the effect can be minimized by using a highly specific Cas nuclease or a stringent sgRNA design that differs from the other genomic regions by three mismatches, in addition to one mismatch in the PAM proximal region. The designed sgRNAs determine the occurrence of a ‘off-target’ effect; sgRNAs with more than 50% GC content are competent enough to promote on-target mutagenesis due to strong binding to the target sites (Kim et al. 2015; Ren et al. 2014). Precisely designed sgRNAs enable specific targeting, even if so many homologous loci are present in the studied genome (Baysal et al. 2016). Many recently introduced computer-based innovations, i.e. Cas-OFF Finder that identifies the unique target sequences and possible off-target sites in the genomes of various species minimizes the off-target sites (Cong et al. 2013a, b; Hsu et al. 2013). CRISPR-P enables gRNA design for substantially all plant species with accessible genome sequences, as well as off-target site and restriction enzyme sequence analyses (Lei et al. 2014). Subsequently, CRISPR-PLANT has used a genome-wide platform of highly comprehensive RNAs in more than eight plant species and favours restriction endonuclease analysis of target sites. Various guidelines for sgRNAs design to lower the potential off-target effects which can be beneficial for various crop species have been documented in recent articles. It is critical to avoid using sgRNAs with seeds that are homologous to various other genome loci in order to minimize off-target mutations. Indispensable components of CRISPR-Cas9, the PAM and seed sequence, need to be carefully designed.
The sites cleaved with the genome-editing tool CRISPR-Cas9 system can be both on-target and off-target sites and that need to appropriately balance according to the different experimental purposes. To avoid these events, bioinformatics tools, for instance, E-CRISPR and Cas OT, can promote sgRNA design concerning whole-genome sequence information. A vector performs as a vehicle for delivering an element of interest. The vector only needs two components: the single-guide RNA (sgRNA) sequence and the Cas9 gene, both of which may be expressed from a single vector system. A variable crRNA (approximately 20 bp) and a constant tracrRNA make up the sgRNA. To boost performance and eliminate off-target impacts, various target sequences of the same gene might be introduced. The Cas9 gene encompasses multiple nuclear localization signals (NLS) for nuclear targeting, and Cas9 can be defined in a variety of ways (Heintze et al. 2013). In addition to this, various delivery methods such as agrobacterium-mediated, bombardment or biolistic approach, PEG-mediated protoplast, and floral-dip are widely used in plants to regulate genes properly (Table 9.2). There are widely used transformation mechanisms, but the agrobacterium-mediated method is extensively used for the delivery of various Cas enzymes (Ali et al. 2015). The RNP strategy is another important way to reduce the intended effect when sgRNA and RNP nuclease processes are introduced by biolistic and electroporation into plant protoplasts, showing a small frequency of target changes and successfully reported on various plants such as maize (Zea mays), rice (Oryza sativa), tomato (Solanum lycopersicum), and many others (Woo et al. 2015). Nanoparticle-mediated RNP delivery systems have been successfully adopted in plant species due to the reduction of unwanted changes via the potentiality of RNP. The recommended system is time-effective, affordable, species-independent, and equipment-independent CRISPR-Cas9 vector or ribonucleoprotein complexes. Consequently, the specificity of CRISPR-Cas9 is influenced by several parameters, including the aggregation of the Cas9/sgRNA complex and the characteristics of the off-target sites.
Off-target mutation is a major apprehension in the emergence of the CRISPR-Cas9 system in plants used whole genome sequencing WGS and deep sequencing, respectively, to investigate CRISPR-Cas9 specificity in Arabidopsis thaliana. According to their findings, CRISPR-Cas9 is highly specific in plants owing to low Cas9 protein expression levels, which resulted in undetectable levels of off-target alterations. Most CRISPR-Cas9 investigations in plants have reported a low frequency of off-target mutation, which could be attributed to its occurrence in non-coding areas and, as a result, the inability to detect off-target implications (Zhang et al. 2018a). CRISPR-PLANT v2 is a popular tool for predicting off-target mutations in plants. This software has the highest sensitivity of among all off-target prediction tool and can be utilized in the genomes of seven plants, including Sorghum bicolor, Arabidopsis thaliana, Oryza sativa, Medicago truncatula, Solanum lycopersicum, Glycine max, and Brachypodium distachyon. However, in eukaryotes, several strategies for off-target recognition have been introduced, including deep sequencing and online prediction software. Although in vitro approaches for investigating potential off-target sites have been established, exact prophecies of the prevalence of undesired mutations in vivo are difficult to acquire. Digenome-seq, SITE-seq, and CIRCLE-seq are the most used in vitro genome-wide detection systems and quantifying off-target effects (Cameron et al. 2017). Digested genome sequencing (Digenome-seq) is a reliable, delicate (~ 0.1%), and frequently used for detecting Cas9 and other nucleases for off-target effects in genome-wide. The most prominent strategies established to solve the Digenome-seq difficulties are selective enrichment and identification of tagged genomic DNA ends by sequencing (SITE-Seq), followed by circularization for in vitro reporting of cleavage effects by sequencing (CIRCLE–seq). The SITE-Seq approach could map all of the Cas9 cleavage sites in a genome (Naeem et al. 2020). This study employed sgRNA and Cas9 RNPs in a cell-free environment to cleave purified genomic DNA. Afterwards, both (on- and off-target) cleavage fragments are tagged, and off-target sites are detected using next-generation sequencing (NGS). The total amount of off-target sites has a considerable impact on nuclease concentration. RNPs (low to high) were employed as variable concentrations to recover off-target locations with low and high cleavage sensitivity. When low doses of RNPs are subjected to cell identification, they exhibit a significant proclivity for off-target alterations. SITE-Seq also requires less NGS read depth than Digenome-seq, with some procedural modifications; CIRCLE-Seq has a similar concept. In CIRCLE-Seq, the DNA is first trimmed, then circularized, and finally destroyed. Prior to treatment with (Cas9–sgRNA) RNPs, the degradation phase practically eliminates high background DNA to boost sensitivity, condensing NGS read space that would otherwise be squandered on random reads. Following that, DNA is linearized using Cas9 and then exposed to NGS for off-target detection. CIRCLE-Seq, like SITE-Seq, could be employed in a reference-independent manner to discover off-target cleavage sites, for organisms whose genome sequences are less well-characterized and/or show considerable genetic variability. Several approaches were proposed, including bioinformatics tools for in silico detection of off-target mutations and increased on-target efficiency to mitigate off-target impacts. However, off-target effects might have happened, yet the alterations will be lower than those developed via conventional breeding. Thus, GEd employing the CRISPR-Cas approach produces a far less off-target effect in comparison to the traditional crop enhancement strategy.
9.7 CRISPR Implementation in Sustainable Agriculture: Climate-Smart and Nutritionally Secure Crops
The global population is assumed to increase 9.2 billion in 2050, and so agronomic production needs to rise by about 70% from existing levels to encounter the increased demand of food, as predicted by Food and Agriculture Organization (accessed on 1 February 2021). Cereal crops such as rice, wheat, and maize are the world’s most important sources of energies intended for humans, livestock feed for animals, and raw material for biofuel. Therefore, improving cereal-crop-grain production is critical to meet further demand. For most cereal crops, the annual yield relates to grain production. Until the last decade, the core crop improvement strategies banked upon chemical mutations, hybrids, and expression of trans gene/s (Chari et al. 2017). The shift from the conventional breeding approach, which relied on the occurrence of the naturally relevant variations to the molecular breeding approach, has alleviated some barriers attached with the conventional methods. Now, the targeted traits can be swiftly incorporated into the plant system to generate a new plant variety for food as well as nutritional security. The gradual increase in human population, deteriorating arable land conditions, the drastic climatic changes through uplifted temperature, and escalated pollutants by excessive emission of greenhouse gases (GHG) causes threat to agriculture and food security (Asseng et al. 2014). Therefore, to develop climate-smart crops via sustainable agriculture, the need of the hour is to achieve a ‘triple win’ by targeting enhanced productivity, improved adaptivity, and GHG mitigation. Targeted GEd made the revolution in molecular biology by discovering programmable SSNs (Chandrasegaran and Carroll 2016). CRISPR-Cas-based GEd has become an essential tool that has effectively caused enormous ripple effects in plant research. Throughout the last decade, we have seen fast development in numerous fields, including plant functional genomics and crop enhancement (>45 genera of plants) in a manner that straightforwardly benefits consumers (Shan et al. 2020). In plant species, the practice of CRISPR-Cas9-mediated genome alteration in diverse crops was successful, for instance, in maize, rice, wheat, maize, and cotton. In 2015, the fourth quarter experienced the employment of DNA-free, pre-assembled RNP complex of CRISPR-Cas9 for genome alteration in model plants such as Arabidopsis, rice, lettuce, tobacco, wheat, maize, and so on (Woo et al. 2015). An extremely systematic transgene integration free GEd and most importantly callus-based methodology were introduced for wheat pertaining transitory expression of CRISPR-Cas9 in the form of DNA or RNA (dubbed TECCDNA or TECCRNA, respectively), the technique had the potential to be applied in different crops (Zhang et al. 2016). The crops developed via RNP complex mediated and TECCRNA-based editing techniques are foreign gene integration free, thus they could be spared from GMO regulatory concerns. In a recent study, by altering the sequence of a S gene, namely SIAGAMOUS-LIKE 6 (SIAGL6) which is linked to enhanced fruit setting even under heat stress, tolerance towards high temperature was attained in tomato (Klap et al. 2017). Further, optimization of method for targeting multiple genes via CRISPR-Cas9 in a single organism was done for numerous crops which include rice, cotton, maize, and wheat (Miao et al. 2013; Gao et al. 2017; Char et al. 2017; Wang et al. 2018b). Thus, CRISPR-Cas9 is a remarkable technique, which is potent enough to develop crop with multiple stress tolerance by choosing concurrently different S genes as a target in exclusively high productive but sensitive cultivars.
The growth of plants is linked to diverse developmental and environmental cues. Plants receive and respond to those cues via cellular signaling cascades, which regulate gene expression at the pre-mRNA level by tuning splicing patterns and controlling the transcript abundance at mature-mRNA level. Alternatively, spliced pre-mRNA represents the genome’s coding potential for multi-exon genes and synchronizes gene expression by different mechanisms. In Solanum tuberosum (potato), vegetative reproduction (tuberization) is regulated via photoperiod, for example, flowering controlling transcription factor- StCDF1 (CYCLING DOF FACTOR 1), which regulate the antisense transcript of StFLORE to gain drought tolerance. Loss of function mutation in promoter of this StFLORE via CRISPR-Cas9 revealed drought tolerance via stomatal size and number regulation (Gonzales et al. 2020).
In agriculture, weed control is critical for a high yield of crop production, which can reduce the phytotoxicity of herbicides to crops, cut off the cost of the weeding, and upgrade the efficiency of the chemical weeding. Consequently, substantial attempts to develop herbicide -resistant crop varieties have been undertaken to contribute the frugal and economic tools to serve farmers for clean and effortless weed management. To develop robust herbicide-resistant crop plants, endogenous genes like cellulose synthase A catalytic subunit 3 (CESA3), splicing factor 3B subunit 1 (SF3B1) and more commonly acetolactate synthase (ALS), 5-enolpyruvylshikimate-3-phosphate synthase (EPSPS) are targeted for CRISPR-Cas9-mediated gene editing. The crucial amino acid substitution in EPSPS and ALS genes in rice employing CRISPR-Cas9 HDR-mediated machinery conferred resistance to glyphosate and sulfonylurea herbicide, respectively (Li et al. 2015; Sun et al. 2016). Similarly, T102I/P106S and T102I/P106A substitution were introduced in EPSPS gene of flax (Sauer et al. 2016) and cassava plant (Hummel et al. 2017). To acquire effective gene replacement, CRISPR-Cas9 is employed with target sequence-specific sgRNAs directing the CRISPR-associated RNA endoribonuclease csy4 from Pseudomonas aeruginosa, for sequence-specific induction of DSBs (Wang et al. 2021). Till now among the developed crop germplasm specifically resistant to herbicides, crops only resistance towards ALS-inhibiting herbicides, ACCase-inhibiting herbicides, and glyphosate has been successfully established. One of the greatest important applications intended for gene editing in agriculture is biotic stress resistance. The genetic mechanisms of the agents that cause biotic stressors in plants can be examined in order to overcome these stresses by GEd (Yin and Qiu 2019; Zafar et al. 2020; Pak et al. 2020). In addition to some crop species like rice, a CRISPR-Cas9 targeted mutation in the ethylene responsive factor, OsERF922, has been effectively established to improve resistance to Magnaporthe oryzae blast disease (Wang et al. 2016). Similarly, OsMPK5, a negative regulator of biotic and abiotic stressors in rice, was identified for targeted mutagenesis in rice protoplasts utilizing three gRNAs by using a more precise gRNA design strategy with a low level of off-targets (Xie and Yang 2013). By producing genetically modified resistant crop varieties, which have proven to be a significant effort to fight against biotic stressors. Despite CRISPR-Cas9 inimitable accomplishment, there are substantial trials in incorporating this technology into agricultural research, especially with transformation-resistant crops reproduced asexually. Several projects are presently in progress to fine-tune CRISPR-Cas9-based technologies for precise editing in the plant genome of the target locus.
Consequently, crop improvement now targets not only improving quantity (yield), but quality (nutrition) of the crop product as well. Great quality food grains have a critical and direct impact on human health and well-being, as plants produce numerous molecules with anti-inflammatory, anti-cancerous, and anti-oxidation properties (Liu et al. 2021) that have beneficial effects on human health. Thus, plants are the major source of nutrients and natural dietary products and are considered as ‘dietary doctors’ as they can cure the prevalent undernourishment (FAO 2020). Thus, crops biofortified with micronutrients and minerals such as iron, zinc, selenium, and iodine can curb the nutrient deficiency in addition making anti-nutrient, such as heavy metals, phytate, and gluten, devoid crops can make the unavailable nutrient available for absorption in human body and protect humans from developing allergies, metabolic disorders, and chronic ailments. Conventional breeding accompanied by technology has saved humanity from the food crisis in the past but now these approaches culminate into no added benefit in enhancing the productivity, whereas new techniques like CRISPR-Cas hold the potential to drive the way towards sustainable food security. Recent breakthroughs (Table 9.3) have paved the way to introduce or manipulate the inherent genes to improve the quality of the majorly consumed crops. Alteration of genes for crop biofortification as well as for removing anti-nutrients have the potential to provide macro and micronutrients and alleviate the ‘hidden hunger’ (Majumder et al. 2019) condition as well as to cure and prevent the non-infectious, lifestyle related chronic ailments in humans. Although the CRISPR-Cas system is still developing and evolving, the latent potential of this technique has resulted in some benchmark studies, and it will continue to bestow the field of genetic engineering with more novel breakthroughs.
9.8 Amalgamation of MI and CRISPR-Based Genome Editing
Despite being one of the common genetic engineering techniques, CRISPR-Cas9 GEd relies on the accuracy of well-designed guide RNAs as it is an essential aspect of successful target gene editing (Cox et al. 2015). In recent years, various algorithms have been generated for assessing CRISPR activity (on-target) and specificity (off-target) as well as web-based tools for in silico gRNA designing (Henry et al. 2014; Zhu 2015). Machine learning (ML) and Artificial Intelligence (AI) offer revolutionary approaches for utilizing the CRISPR-Cas9 technology to analyse edited crop lines with better features, for example, higher nutrient value, palatability, modified root, flower architectures, stress tolerance, and so on. Some examples of CRISPR-based design tools are described in Table 9.4. All of these gRNA design tools, off- and on-target prediction tools have contributed to the success and application of CRISPR genome technology.
Several functions have been shown to be important for target gRNA activity, including secondary structure, sequence composition, thermodynamics, and physicochemical characteristic, but for off-target predictions, this is the size, composition, and combination of discrepancies. Many machines and deep learning methods have been established to represent the activity of CRISPR, which can be broadly divided into two types. (1) Machine learning based, which includes CRISPRscan, sgRNA Scorer, SSC, sgRNA Designer, and CRISPRater. CRISPRScan, CRISPRater, and SSC are trained using simple linear models, and Azimuth2.0 and TUSCAN are trained using general linear models that are logistic regression and random forests, respectively (Listgarten et al. 2018). (2) Deep learning based. CNN_std, DeepCas9, DeepCRISPR, CRISPRpred, and DeepCpf1 predict sgRNA activity builds on automatic recognition of sequence characters using a Convolutional Nuclear Network (CNN). MIT server estimates off-targets based on the distance and number between unpaired nucleotides (Hsu et al. 2013). Subsequently, a cutting frequency determination (CFD) score was developed that predicts off-target scores by reproducing the frequency of bases in gRNA spacer sequence (Doench et al. 2016). Synergizing CRISPR combines the projection results of five different models (CCTop, CFD, CROPIT, MIT, and MIT website) into an input function based on hypothetical and statistical methods (Dobson et al. 2015; Singh et al. 2015). There are currently numerous procedures available to generate accurate sgRNAs using basic rules. Here, a new algorithm called CRISPR target estimation (CRISTA) was introduced as part of ML, which performed the important task of identifying specific genomic regions to be accurately removed via given sgRNAs. The CRISTA predictions have been proven to be more accurate than previously predicted thresholds (Abadi et al. 2017). However, identifying prospective off-target sites required the recognition of short sequence motifs up to 20 bp, besides the PAM with frequent mismatches. In most cases, the aligners first match the seed sequence and extend the seed sequence in a specific direction and then check for a match. Therefore, ML and AI analysed possible regression points that may converge or deviate from on-target and off-target specificity charts.
The precision of these tools for predicting gRNA activity in different species and cell types remains unclear (Chuai et al. 2017). Large variations between species have led to the development of species-specific software (e.g. CRISPR-P for plants, flyCRISPR for fruit flies, CRISPRscan for zebrafish, and EuPaGDT for pathogens). Of these, only CRISPRscan was generated based on ML, and the rest were theoretical software. Since organisms cannot rapidly limit the previous off-target scoring process, researchers wanted to create a new procedure for assessing off-target action called CASPER (Mendoza and Trinh 2018). Although these tools can be selected for prior study when performing experiments by editing them in corresponding species, their prediction of sgRNA efficiency and target in various cell types is debatable. However, these tools have been proved in the laboratory using mouse cell lines, and human or both, major and cross-species variations have not yet been testified. Therefore, ML-based learning approaches can effectively predict lethal sgRNA interactions and characterize target regions in specific gene combinations. However, there is a large amount of work to be tested and optimized for utilizing CRISPR-Cas9 gene editing in plant systems. In the future, genome-wide engineering crops will include trained data sets, including variants and orthologs.
9.9 Regulatory Aspects of Genome Edited Crops
GEd technology has proved its potential uses in a broad array of industries, notably human and animal health, food, agriculture, and others, in a relatively short period of time. GEd innovations, the same as any other new technology, have dual-use prospective and so raise both safety and security concerns. Novel GEd techniques, particularly CRISPR-Cas9, have a unified mechanism for the insertion of elite attributes in crops plants, allowing unconstrained base substitutions, additions, deletions, and gene introduction or replacement. The offspring produced are similar to those produced by random mutagenesis, natural genetic variants, and traditional breeding. The Cartagena Protocol governs the regulation of genetically modified organisms (GMOs), which is part of the worldwide regulatory framework for living modified organisms (LMOs). LMOs, according to their definition, are living organisms with a unique combination of genetic material that has been improved via the use of contemporary technological methods. In contrast to GMO, the integration site is pre-decided, precise and without an insertion of foreign DNA in GEd organisms. The Cartagena Protocol is based on international terms and conditions that each state and its government must adhere to when enacting biosafety legislation. In addition, the lack of clear perception mentioned in the protocol has been a subject of an argument to date.
Universally, there are differing perspectives on how to harmonize genome edited product/process-based policy in every region (Fig. 9.3). One argument is that GEd species do not need to be regulated because there is no trace of genetic engineering in particular categories, and they resemble organisms that have evolved naturally. The opposing point of view is that GEd organisms must be regulated, but they do not have to go through the same stringent biosafety regulatory process as all GMOs/LMOs. Such divergent viewpoints reflect the rules and regulations that govern the regulation of GE organisms and products in each country. The insertion of considerable modifications to the genomes of GE crop plants generated using gene editing or Site-Directed Nuclease (SDN) technologies showed genetic differences. There are three types of SDN technology: SDN-1: These were made by cleaving double-stranded DNA in the existing genome without involving of foreign DNA particles, as a result the end products characteristics are almost similar to what arose from natural plant mechanisms and or artificial mutation . SDN-2: involves a short homologous DNA fragment that contains few base pair different from the targeted DNA template. Double strand cut is recognized by the host repair system and simultaneously repaired with the help of donor DNA fragment and introduces predetermined mutations. Lastly, SDN-3: requires a DNA repair donor template longer than 20 bp for incorporation into the target area, which is accomplished by a DSBs nick in the gene that is accomplished by a fragment carrying a gene or other genetic material template. The first and second SDN approaches lack foreign DNA insertions or recombinant DNA because they do not produce new plant varieties. SDN-3, on the other hand, would be subject to GMO regulation if newly created plant types comprised more than 20 bp foreign DNA insertions, showing the same outcome as the classic recombinant DNA technique (Pauwels et al. 2014). Mutation breeding (induced random mutagenesis) or CRISPR-Cas9 (gene editing technology) can be used to create crop features with similar phenotypes, and they will fall into the same category. The change to genetic modifications is appealing due to the possibility for developers to use SDN technology to build superior crops that could bypass the cumbersome regulatory assessments associated with GE crop adoption (Arora and Narula 2017; Yin et al. 2017; Pacher and Puchta 2017; Kumlehn et al. 2018; Sedeek et al. 2019). Policymaker laws that facilitate the commercialization of gene-edited crops could reduce the time between the lab and the farmer even more. Globally, the countries that have welcomed GM crop production and export policy have a planned structure that is quick, simple to comprehend and follow, and enforced (Levin 1994). Notwithstanding their various process or product-based techniques, Argentina, Brazil, Chile, Costa Rica, Honduras, Mexico, and Uruguay were among the first Latin American countries to give GM agricultural permits (Ishii and Araki 2017; Rosado and Craig 2017). This day, these nations are fast forward in cultivating biotech crops and thus, their economic success could be explained by something other than the GMO framework (Table 9.5) (Rosado and Craig 2017). SDN-1 products are almost universally regarded as non-GMO, and the final product would go through the same legislative framework as classically produced plant species (Schmidt et al. 2020).
Divergence re-emerges, however, when it comes to SDN-2 techniques: Australia and Japan have taken a cautious approach, determining that organisms modified with the SDN-2 technology will be classified as GMOs (Thygesen 2019; Tsuda et al. 2019). Plants that have undergone a genetic modification requiring an initial assessment on the basis of their creation using NBTs are characterized as gene-edited organisms, with the exception of those that have been modified without a template or with a modest template. This is not always a negative attitude; in fact, it is one of the key causes driving the formation of biosafety regulation in the first place: it upholds societal ideals of risk assessment and risk management with the ultimate goal of safeguarding human, animal, and environmental health.
9.10 Conclusion
Implementation of Noble Prize winner CRISPR-Cas GEd technique for plant GEd and regulation has revolutionized the field of genetic engineering and advanced the plant molecular breeding aspect for crop improvement. Recent advances in genome sequencing (reading) and DNA editing or engineering (writing) techniques have led to an era where we can read and write or even re-write the complex genome of plants. With novel breakthroughs of CRISPR-Cas system, we have witnessed the rise of genetic engineering 2.0 which has contributed enormously to the development of practical, valuable, applicable, and multifaceted tools. These tools are the arsenal for future gene editing, genome modification, metabolic engineering avenues via gene knockout, knockin, replacement, point mutations, fine-tuning of gene regulation, and other modifications at any gene locus. This comprehensive review highlights the successful implementation of the CRISPR-Cas system in plant GEd as well as aids in the documentation of some novel events in the field of plant genetic engineering. In addition, it addresses the technical limitations & shortcomings of the tools, and how to overcome those challenges. Additionally, grieve regulatory concerns and applicability of the machine learning approach i achieve the next-generation engineering or breeding technique. However, it encourages the utilization of the new addition of the CRISPR tool kit for their development into programmable nucleases for efficient, precise, and easy to achieve plant GEd tools. While public acceptance will always be a great concern but there has been a shift in the general notion of disapproval of the genome edited crops however not completely. But even a pitch positive turn with the subsisting endeavours of the scientific community and government ministries of INDIA will be a great achievement, and a way forward to the release of the CRISPR generated new robust variety in the global market.
References
Abadi S, Yan WX, Amar D, Mayrose I (2017) A machine learning approach for predicting CRISPR-Cas9 cleavage efficiencies and patterns underlying its mechanism of action. PLoS Comput Biol 13:1–24. https://doi.org/10.1371/journal.pcbi.1005807
Abdullaha JZ, Honga X, Zhanga S, Yao R, Xiao Y (2020) CRISPR base editing and prime editing: DSB and template-free editing systems for bacteria and plants. Synth Syst Biotechnol 5:277–292. https://doi.org/10.1016/jsynbio.2020.08.003
Abe K, Araki E, Suzuki Y, Toki S, Saika H (2018) Production of high oleic/low linoleic rice by genome editing. Plant Physiol Biochem 131:58–62
Abudayyeh OO, Gootenberg JS, Konermann S, Joung J, Slaymaker IM, Cox DB, Shmakov S, Makarova KS, Semenova E, Minakhin L, Severinov K, Regev A, Lander ES, Koonin EV, Zhang F (2016) C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353(6299):5573. https://doi.org/10.1126/science.aaf5573
Abudayyeh OO, Gootenberg JS, Essletzbichler P, Han S, Joung J, Belanto JJ, Verdine V, Cox DBT, Kellner MJ, Regev A, Lander ES, Voytas DF, Ting AY, Zhang F (2017) RNA targeting with CRISPR-Cas13. Nature 550(7675):280–284
Abudayyeh G, Gootenberg JS, Franklin B, Koob J, Kellner MJ, Ladha A, Joung J, Kirchgatterer P, Cox DBT, Zhang F (2019) A cytosine deaminase for programmable single-base RNA editing. Science 365(6451):382–386
Adli M (2018) The CRISPR tool kit for genome editing and beyond. Nat Commun 9:1911. https://doi.org/10.1038/s41467-018-04252-2
Akama K, Akter N, Endo H, Kanesaki M, Endo M, Toki S (2020) An in vivo targeted deletion of the calmodulin-binding domain from rice glutamate decarboxylase 3 (OsGAD3) increases γ-aminobutyric acid content in grains. Rice 13:20
Ali Z, Abulfaraj A, Idris A, Ali S, Tashkandi M, Mahfouz MM (2015) CRISPR/Cas9-mediated viral interference in plants. Genome Biol 16:238
Alkan F, Wenzel A, Anthon C, Havgaard JH, Gorodkin J (2018) CRISPR-Cas9 off-targeting assessment with nucleic acid duplex energy parameters. Genome Biol 19:1–13. https://doi.org/10.1186/s13059-018-1534-x
Anders C, Niewoehner O, Duerst A, Jinek M (2014) Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513:569–573. https://doi.org/10.1038/nature13579
Anzalone AV, Randolph PB, Davis JR, Sousa AA, Koblan LW, Levy JM, Chen PJ, Wilson C, Newby GA, Raguram A, Liu DR (2019) Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576:149–157
Anzalone AV, Koblan LW, Liu DR (2020) Genome editing with CRISPR–Cas nucleases base editors transposases and prime editors. Nat Biotechnol 38:824–844. https://doi.org/10.1038/s41587-020-0561-9
Aquino-Jarquin G (2019) CRISPR-Cas14 is now part of the artillery for gene editing and molecular diagnostic. Nanomedicine 18:428–431
Arndell T, Sharma N, Langridge P, Baumann U, Watson-Haigh NS, Whitford R (2019) gRNA validation for wheat genome editing with the CRISPR-Cas9 system. BMC Biotechnol 19(1):1–12
Arora L, Narula A (2017) Gene editing and crop improvement using CRISPR-Cas9 system. Front Plant Sci 8:1932. https://doi.org/10.3389/fpls.2017.01932
Aschenbrenner S, Kallenberger SM, Hoffmann MD, Huck A, Eils R, Niopek D (2020) Coupling Cas9 to artificial inhibitory domains enhances CRISPR- Cas9 target specificity. Sci Adv 6(6):0187
Asseng S, Ewert F, Martre P, Rötter RP, Lobell DB, Cammarano D, Kimball BA, Ottman MJ, Wall GW, White JW, Reynolds MP, Alderman PD, Prasad PVV, Aggarwal PK, Anothai J, Basso B, Biernath C, Challinor AJ, De Sanctis G, Doltra J, Fereres E, Garcia-Vila M, Gayler S, Hoogenboom G, Hunt LA, Izaurralde RC, Jabloun M, Jones CD, Kersebaum KC, Koehler AK, Müller C, Naresh KS, Nendel C, O’Leary G, Olesen JE, Palosuo T, Priesack E, Eyshi Rezaei E, Ruane AC, Semenov MA, Shcherbak I, Stöckle C, Stratonovitch P, Streck T, Supit I, Tao F, Thorburn PJ, Waha K, Wang E, Wallach D, Wolf J, Zhao Z, Zhu Y (2014) Rising temperatures reduce global wheat production. Nat Clim Change 5(2):143–147. https://doi.org/10.1038/nclimate2470
Bae S, Park J, Kim JS (2014) Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30:1473–1475. https://doi.org/10.1093/bioinformatics/btu048
Baltes NJ, Gil-Humanes J, Cermak T, Atkins PA, Voytas DF (2014) DNA replicons for plant genome engineering. Plant Cell 26:151–163. https://doi.org/10.1105/tpc.113.119792
Baltes NJ, Hummel AW, Konecna E, Cegan R, Bruns AN, Bisaro DM, Voytas DF (2015) Conferring resistance to geminiviruses with the CRISPR–Cas prokaryotic immune system. Nat Plants 1:15145. https://doi.org/10.1038/nplants.2015.145
Banakar R, Eggenberger AL, Lee K, Wright DA, Murugan K, Zarecor S, Lawrence-Dill CJ, Sashital DG, Wang K (2019) High-frequency random DNA insertions upon co-delivery of CRISPR-Cas9 ribonucleoprotein and selectable marker plasmid in rice. Sci Rep 9(1):1–13
Bari VK, Nassar JA, Kheredin SM, Gal-On A, Ron M, Britt A, Steele D, Yoder J, Aly R (2019) CRISPR/Cas9-mediated mutagenesis of CAROTENOID CLEAVAGE DIOXYGENASE 8 in tomato provides resistance against the parasitic weed Phelipanche aegyptiaca. Sci Rep 9:11438
Barkau CL, Reilly D, Eddington SB, Damha MJ, Gagnon KT (2021) Small nucleic acids and the path to the clinic for anti-CRISPR. Biochem Pharmacol 189:114492
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712. https://doi.org/10.1126/science.1138140
Baysal C, Bortesi L, Zhu C, Farré G, Schillberg S, Christou P (2016) CRISPR/Cas9 activity in the rice OsBEIIb. Mol Breed 36:1–11
Behera SS, Ray RC, Zdolec N (2018) Lactobacillus plantarum with functional properties: an approach to increase safety and shelf-life of fermented foods. Biomed Res Int 2018:9361614. https://doi.org/10.1155/2018/9361614
Belfort M, Roberts RJ (1997) Homing endonucleases: keeping the house in order. Nucleic Acids Res 25(17):3379–3388. https://doi.org/10.1093/nar/25.17.3379
Bharat SS, Li S, Li J, Yan L, Xia L (2020) Base editing in plants: current status and challenges. Crop J 8(3):384–395. https://doi.org/10.1016/j.cj.2019.10.002
Bihmidine S, Lin J, Stone JM, Awada T, Specht JE, Clemente TE (2013) Activity of the Arabidopsis RD29A and RD29B promoter elements in soybean under water stress. Planta 237(1):55–64. https://doi.org/10.1007/s00425-012-1740-9
Bolotin A, Quinquis B, Sorokin A, Ehrlich SD (2005) Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151:2551–2561. https://doi.org/10.1099/mic028048-0
Brouns SJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJ, Snijders AP, Dickman MJ, Makarova KS, Koonin EV, Van der Oost J (2008) Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321:960–964. https://doi.org/10.1126/science.1159689
Budhagatapalli N, Rutten T, Gurushidze M, Kumlehn J, Hensel G (2015) Targeted modification of gene function exploiting homology-directed repair of TALEN-mediated double-strand breaks in Barley. G3 (Bethesda) 5(9):1857–1863. https://doi.org/10.1534/g3.115.018762
Bult CJ, White O, Olsen GJ, Zhou L, Fleischmann RD, Sutton GG, Blake JA, FitzGerald LM, Clayton RA, Gocayne JD, Kerlavage AR, Dougherty BA, Tomb JF, Adams MD, Reich CI, Overbeek R, Kirkness EF, Weinstock KG, Merrick JM, Glodek A, Scott JL, Geoghagen NS, Venter JC (1996) Complete genome sequence of the methanogenic archaeon Methanococcus jannaschii. Science 273:1058–1073. https://doi.org/10.1126/science.273.5278.1058
Burmistrz M, Krakowski K, Krawczyk-Balska A (2020) RNA-targeting CRISPR–Cas systems and their applications. Int J Mol Sci 21:1122. https://doi.org/10.3390/ijms21031122
Butler NM, Baltes NJ, Voytas DF, Douches DS (2016) Geminivirus-mediated genome editing in potato (Solanum tuberosum L) using sequence-specific nucleases. Front Plant Sci 7:1045. https://doi.org/10.3389/fpls.2016.01045
Cai F, Axen SD, Kerfeld CA (2013) Evidence for the widespread distribution of CRISPR-Cas system in the Phylum Cyanobacteria. RNA Biol 10:687–693
Cameron P, Fuller CK, Donohoue PD, Jones BN, Thompson MS, Carter MM, Gradia S, Vidal B, Garner E, Slorach EM, Lau E, Banh LM, Lied AM, Edwards LS, Settle AH, Capurso D, Llaca V, Deschamps S, Cigan M, Young JK, May AP (2017) Mapping the genomic landscape of CRISPR-Cas9 cleavage. Nat Methods 14(6):600–606. https://doi.org/10.1038/nmeth.4284
Carlessi M, Mariotti L, Giaume F, Fornara F, Perata P, Gonzali S (2021) Targeted knockout of the gene OsHOL1 removes methyl iodide emissions from rice plants. Sci Rep 11:17010. https://doi.org/10.1038/s41598-021-95198-x
Carlson-Stevermer J, Kelso R, Kadina A, Joshi S, Rossi N, Walker J, Stoner R, Maures T (2020) CRISPRoff enables spatio-temporal control of CRISPR editing. Nat Commun 11:5041. https://doi.org/10.1038/s41467-020-18853-3
Čermák T, Baltes NJ, Čegan R, Zhang Y, Voytas DF (2015) High-frequency precise modification of the tomato genome. Genome Biol 16:232. https://doi.org/10.1186/s13059-015-0796-9
Chandrasegaran S, Carroll D (2016) Origins of programmable nucleases for genome engineering. J Mol Biol 428:963–989. https://doi.org/10.1016/j.jmb.2015.10.014
Char SN, Neelakandan AK, Nahampun H, Frame B, Main M, Spalding MH, Becraft PW, Meyers BC, Walbot V, Wang K, Yang B (2017) An agrobacterium-delivered CRISPR/Cas9 system for high-frequency targeted mutagenesis in maize. Plant Biotechnol 15:257–268
Chari R, Mali P, Moosburner M, Church GM (2015) Unraveling CRISPR-Cas9 genome engineering parameters via a library-on-library approach. Nat Methods 12:823–826. https://doi.org/10.1038/nmeth.3473
Charo RA (2015) Yellow lights for emerging technologies. Science 349(6246):384–385. https://doi.org/10.1126/science.aab3885
Charo RA (2016) The legal and regulatory context for human gene editing. Issues Sci Technol 32(3):39–45
Che J, Yamaji N, Ma JF (2021) Role of a vacuolar iron transporter OsVIT2 in the distribution of iron to rice grains. New Phytol 230:1049–1062
Chen F, Ding X, Feng Y, Seebeck T, Jiang Y, Davis GD (2017) Targeted activation of diverse CRISPR-Cas systems for mammalian genome editing via proximal CRISPR targeting. Nat Commun 8:14958. https://doi.org/10.1038/ncomms14958
Chen JS, Ma E, Harrington LB, Da Costa M, Tian X, Palefsky JM, Doudna JA (2018) CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360(6387):436–439. https://doi.org/10.1126/science.aar6245
Chen Y, Fu M, Li H et al (2021) High-oleic acid content nontransgenic allotetraploid cotton (Gossypium hirsutum L) generated by knockout of GhFAD2 genes with CRISPR/Cas9 system. Plant Biotechnol J 19(3):424–426. https://doi.org/10.1111/pbi.13507
Christian M, Qi Y, Zhang Y, Voytas DF (2013) Targeted mutagenesis of Arabidopsis thaliana using engineered TAL effector nucleases. G3(Bethesda) 3:1697–1705
Chuai G, Hui WQL, Liu Q (2017) In silico meets in vivo: towards computational CRISPR-based sgRNA design. Trends Biotechnol 35:12–21. https://doi.org/10.1016/jtibtech201606008
Chuai G, Ma H, Yan J, Chen M, Hong N, Xue D et al (2018) DeepCRISPR: optimized CRISPR guide RNA design by deep learning. Genome Biol 19:1–18. https://doi.org/10.1186/s13059-018-1459-4
Colleaux L, D’Auriol L, Galibert F, Dujon B (1988) Recognition and cleavage site of the intron-encoded omega transposase. Proc Natl Acad Sci U S A 85(16):6022–6026. https://doi.org/10.1073/pnas.85.16.6022
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013a) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823. https://doi.org/10.1126/science1231143
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Zhang F (2013b) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Cox DBT, Platt RJ, Zhang F (2015) Therapeutic genome editing: prospects and challenges. Nat Med 21:121–131. https://doi.org/10.1038/nm.3793
Cox DBT, Gootenberg JS, Abudayyeh OO, Franklin B, Kellner MJ, Joung J, Zhang F (2017) RNA editing with CRISPR-Cas13. Science 358(6366):1019–1027. https://doi.org/10.1126/science.aaq0180
Crudele JM, Chamberlain JS (2018) Cas9 immunity creates challenges for CRISPR gene editing therapies. Nat Commun 9:3497. https://doi.org/10.1038/s41467-018-05843-9
Curtin SJ, Zhang F, Sander JD, Haun WJ, Starker C, Baltes NJ, Reyon D, Dahlborg EJ, Goodwin MJ, Coffman AP, Dobbs D, Joung JK, Voytas DF, Stupar RM (2011) Targeted mutagenesis of duplicated genes in soybean with zinc-finger nucleases. Plant Physiol 156(2):466–473. https://doi.org/10.1104/pp.111.172981
Cyranoski D (2016) CRISPR gene-editing tested in a person for the first time. Nature 539:479. https://doi.org/10.1038/nature.2016.20988
Cyranoski D (2020) What CRISPR-baby prison sentences mean for research. Nature 577:154–155. https://doi.org/10.1038/d41586-020-00001-y
D’Halluin K, Vanderstraeten C, Stals E, Cornelissen M, Ruiter R (2007) Homologous recombination: a basis for targeted genome optimization in crop species such as maize. Plant Biotechnol 6(1):93–102. https://doi.org/10.1111/j1467-7652200700305x
Datsenko K, Pougach K, Tikhonov A, Wanner BL, Severinov K, Semenova E (2012) Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system. Nat Commun 3:945. https://doi.org/10.1038/ncomms1937
DiCarlo JE, Norville JE, Mali P, Rios X, Aach J, Church GM (2013) Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems. Nucleic Acids Res 41(7):4336–4343. https://doi.org/10.1093/nar/gkt135
Dobson L, Reményi I, Tusnády GE (2015) CCTOP: a consensus constrained topology prediction web server. Nucleic Acids Res 43:408–412. https://doi.org/10.1093/nar/gkv451
Doench JG, Hartenian E, Graham DB, Tothova Z, Hegde M, Smith I, Sullender M, Ebert BL, Xavier RJ, Root DE (2014) Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat Biotechnol 32(12):1262–1267. https://doi.org/10.1038/nbt.3026
Doench JG, Fusi N, Sullender M, Hegde M, Vaimberg EW, Donovan KF, Smith I, Tothova Z, Wilen C, Orchard R, Virgin HW, Listgarten J, Root DE (2016) Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol 34:184–191. https://doi.org/10.1038/nbt.3437
Dolgin E (2019) Finding the CRISPR off-switch. Nature 577:309
Dong D, Guo M, Wang S, Zhu Y, Wang S, Xiong Z, Yang J, Xu Z, Huang Z (2017) Structural basis of CRISPR–SpyCas9 inhibition by an anti-CRISPR protein. Nature 546(7658):436–439
Dong L, Qi X, Zhu J, Liu C, Zhang X, Cheng B, Mao L, Xie C (2019) Supersweet and waxy: meeting the diverse demands for specialty maize by genome editing. Plant Biotechnol 17:1853–1855. https://doi.org/10.1111/pbi.13144
Dong OX, Yu S, Jain R, Zhang N, Duong PQ, Butler C, Li Y, Lipzen A, Martin JA, Barry KW, Schmutz J, Tian L, Ronald PC (2020) Marker-free carotenoid-enriched rice generated through targeted gene insertion using CRISPR-Cas9. Nat Commun 11:1178. https://doi.org/10.1038/s41467-020-14981-y
Doudna JA, Charpentier E (2014) The new frontier of genome engineering with CRISPR-Cas9. Science 346:1258096
DuPont Pioneer (2016) DuPont Announces Intentions to Commercialize First CRISPR/Cas Product Press Release. https://www.pioneercom/home/site/about/news-media/newreleases/template
Endo A, Masafumi M, Kaya H, Toki S (2016) Efficient targeted mutagenesis of rice and tobacco genomes using Cpf1 from Francisella novicida. Sci Rep 6:38169
Estrella MA, Kuo FT, Bailey S (2016) RNA-activated DNA cleavage by the Type III-B CRISPR-Cas effector complex. Genes Dev 30:460–470
Feng Z, Zhang B, Ding W, Liu X, Yang DL, Wei P, Cao F, Zhu S, Zhang F, Mao Y, Zhu JK (2013) Efficient genome editing in plants using a CRISPR/Cas system. Cell Res 23(10):1229
Fine EJ, Cradick TJ, Zhao CL, Lin Y, Bao G (2013) An online bioinformatics tool predicts zinc finger and TALE nuclease off-target cleavage. Nucleic Acids Res 42(6):2014
Forner J, Pfeiffer A, Langenecker T, Manavella PA, Lohmann JU (2015) Correction: germline-transmitted genome editing in Arabidopsis thaliana using TAL-effector-nucleases. PLoS One 10(7):0133945
Gaj T, Gersbach CA, Barbas CF (2013) ZFN TALEN and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31(7):397–405. https://doi.org/10.1016/j.tibtech.2013.04.004
Gao W, Long L, Tian X, Xu F, Liu J, Singh PK, Botella JR, Song C (2017) Genome editing in cotton with the CRISPR/Cas9 system. Front Plant Sci 8:1364
Gaoneng S, Lihong GJ, Xiangjin W, Zhonghua S, Shaoqing T, Peisong HU (2017) CRISPR/CAS9-mediated editing of the fragrant gene Badh2 in rice. Chin J Rice Sci 31:216–222
Gasiunas G, Barrangou R, Horvath P, Siksnys V (2012) Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci U S A 109:2579–2586. https://doi.org/10.1073/pnas.1208507109
Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, Liu DR (2017) Programmable base editing of A• T to G• C in genomic DNA without DNA cleavage. Nature 551:464–471
Gelvin SB (2006) Agrobacterium virulence gene induction. Methods Mol Biol 343:77–84. https://doi.org/10.1385/1-59745-130-4:77
Gilbert LA, Larson MH, Morsut L, Liu Z, Brar GA, Torres SE, Stern-Ginossar N, Brandman O, Whitehead EH, Doudna JA, Lim WA, Weissman JS, Qi LS (2013) CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154(2):442–451
Gil-Humanes JY, Wang Z, Liang Q, Shan CV, Ozuna S, Sánchez-León NJ, Baltes C, Starker F, Barro C, Gao DF, Voytas DF (2017) High-efficiency gene targeting in hexaploid wheat using DNA replicons and CRISPR/Cas9. Plant J 89(6):1251–1262
Gonzales LR, Li S, Bergonzi S, Oortwijn M, Bachem C (2020) Potato cycling Dof Factor1 and its lncRNA counterpart StFLORE link tuber development and drought response. Plant J 105:855–869. https://doi.org/10.1111/tpj.15093
Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, Verdine V, Donghia N, Daringer NM, Freije CA, Myhrvold C, Bhattacharyya RP, Livny J, Regev A, Koonin EV, Hung DT, Sabeti PC, Collins JJ, Zhang F (2017) Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356(6336):438–442. https://doi.org/10.1126/science.aam9321
Gratz SJ, Ukken FP, Rubinstein CD, Thiede G, Donohue LK, Cummings AM, O'Connor-Giles KM (2014) Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in Drosophila. Genetics 196:961–971. https://doi.org/10.1534/genetics.113.160713
Haeussler M, Schönig K, Eckert H, Eschstruth A, Mianné J, Renaud JB, Schneider-Maunoury S, Shkumatava A, Teboul L, Kent J, Joly JS, Concordet JP (2016) Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol 17:1–12. https://doi.org/10.1186/s13059-016-1012-2
Haft DH, Selengut J, Mongodin EF, Nelson KE (2005) A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/Cas subtypes exist in prokaryotic genomes. PLoS Comput Biol 1:60
Hajiahmadi Z, Shirzadian-Khorramabad R, Kazemzad M, Sohani MM (2019) Enhancement of tomato resistance to Tuta absoluta using a new efficient mesoporous silica nanoparticle-mediated plant transient gene expression approach. Sci Hortic 243:367–375
Hale CR, Zhao P, Olson S, Duff MO, Graveley BR, Wells L, Terns RM, Terns MP (2009) RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 139:945–956
Hampton T (2020) DNA prime editing: a new CRISPR-based method to correct most disease-causing mutations. JAMA 323(5):405–406. https://doi.org/10.1001/jama.2019.21827
Harrington LB, Burstein D, Chen JS, Paez-Espino D, Ma E, Witte IP, Cofsky JC, Kyrpides NC, Banfield JF, Doudna JA (2018) Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362:839–842
Haurwitz RE, Jinek M, Wiedenheft B, Zhou K, Doudna JA (2010) Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 329(5997):1355–1358. https://doi.org/10.1126/science.1192272
He S (2020) The first human trial of CRISPR-based cell therapy clears safety concerns as new treatment for late-stage lung cancer. Sig Transduct Target Ther 5:168. https://doi.org/10.1038/s41392-020-00283-8
Heigwer F, Kerr G, Boutros M (2014) E-CRISP: fast CRISPR target site identification. Nat Methods 11:122–123. https://doi.org/10.1038/nmeth.2812
Henry VJ, Bandrowski AE, Pepin AS, Gonzalez BJ, Desfeux A (2014) OMICtools: an informative directory for multi-omic data analysis. Database (Oxford) 2014:bau069. https://doi.org/10.1093/database/bau069
Hermans PW, Van Soolingen D, Bik EM, De Haas PE, Dale JW, Van Embden JD (1991) Insertion element IS987 from Mycobacterium bovis BCG is located in a hot-spot integration region for insertion elements in Mycobacterium tuberculosis complex strains. Infect Immun 59:2695–2705
Hirosawa M, Saito H (2021) Cell-type-specific CRISPR-Cas9 system with miRNAs In CRISPR-Cas methods. Humana, New York, NY, pp 265–279
Hodgkins A, Farne A, Perera S, Grego T, Parry-Smith DJ, Skarnes WC, Iyer V (2015) WGE: a CRISPR database for genome engineering. Bioinformatics 31:3078–3080. https://doi.org/10.1093/bioinformatics/btv308
Hoffmann M, Kleine-Weber H, Schroeder S et al (2020) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181(2):271–280. https://doi.org/10.1016/j.cell.2020.02.052
Hrle A, Maier L, Sharma K, Ebert J, Basquin C, Urlaub H, Marchfelder A, Conti E (2014) Structural analyses of the CRISPR protein Csc2 reveal the RNA-binding interface of the type I-D Cas7 family. RNA Biol 11(8):1072–1082. https://doi.org/10.4161/rna.29893
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O, Cradick TJ, Marraffini LA, Bao G, Zhang F (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31:827–832. https://doi.org/10.1038/nbt.2647
Hua K, Tao X, Han P, Wang R, Zhu JK (2019) Genome engineering in rice using Cas9 variants that recognize NG PAM sequences. Mol Plant 12:1003–1014
Huang J, Li J, Zhou J, Wang L, Yang S, Hurst LD, Li WH, Tian D (2018) Identifying a large number of high-yield genes in rice by pedigree analysis whole-genome sequencing and CRISPR-Cas9 gene knockout. Proc Natl Acad Sci USA 115:7559–7567. https://doi.org/10.1073/pnas.18061101
Huang L, Li Q, Zhang C, Chu R, Gu Z, Tan H, Zhao D, Fan X, Liu Q (2020) Creating novel Wx alleles with fine-tuned amylose levels and improved grain quality in rice by promoter editing using CRISPR/Cas9 system. Plant Biotechnol 18(11):2164–2166. https://doi.org/10.1111/pbi.13391
Hummel AW, Chauhan RD, Cermak T, Mutka AM, Vijayaraghavan A, Boyher A, Starker CG, Bart R, Voytas DF, Taylor NJ (2017) Allele exchange at the EPSPS locus confers glyphosate tolerance in cassava. Plant Biotechnol 16:1275–1282
Ibrahim S, Saleem B, Rehman N, Zafar SA, Naeem MK, Khan MR (2021) CRISPR/Cas9 mediated disruption of Inositol pentakisphosphate 2-kinase 1(TaIPK1) reduces phytic acid and improves iron and zinc accumulation in wheat grains. J Adv Res 37:33–41. https://doi.org/10.1016/j.jare.2021.07.006
Ishii T, Araki M (2017) A future scenario of the global regulatory landscape regarding genome-edited crops GM. Crop Food 8:44–56. https://doi.org/10.1080/21645698.2016.1261787
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A (1987) Nucleotide sequence of the iap gene responsible for alkaline phosphatase isozyme conversion in Escherichia coli and identification of the gene product. J Bacteriol 169:5429–5433. https://doi.org/10.1128/jb.169.12.5429-5433.1987
Jacob H, Christin L, Robin K (2013) A CRISPR CASe for high-throughput silencing. Front Plant Sci 4:193. https://doi.org/10.3389/fgene.2013.00193
Jacobs TB, LaFayette PR, Schmitz RJ, Parrott WA (2015) Targeted genome modifications in soybean with CRISPR/Cas9. BMC Biotechnol 15:16
Jacquin ALS, Odom DT, Lukk M (2019) Crisflash: open-source software to generate CRISPR guide RNAs against genomes annotated with individual variation. Bioinformatics 35:3146–3147. https://doi.org/10.1093/bioinformatics/btz019
James K, Nuñez JC, Greg C, Pommier JZC, Joseph MR, Carmen A, Gokul NR, Quanming S, King L, Avi JS, Angela NP, James YSK, Amanda C, Manuel DL, Howard YC, Martin K, Bradley EB, Hovestadt V, Luke AG, Jonathan SW (2021) Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184(9):2503–2519.e17. https://doi.org/10.1016/j.cell.2021.03.025
Jeong YK, Song B, Bae S (2020) Current status and challenges of DNA base editing tools. Mol Ther 28:91938–91952
Jiang W, Zhou H, Bi H, Fromm M, Yang B, Weeks DP (2013) Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis tobacco sorghum and rice. Nucleic Acids Res 41(20):188–188
Jiang WZ, Henry IM, Lynagh PG, Comai L, Cahoon EB, Weeks DP (2017) Significant enhancement of fatty acid composition in seeds of the allohexaploid Camelina sativa using CRISPR/Cas9 gene editing. Plant Biotechnol J 15:648–646
Jiang M, Liu Y, Liu Y, Tan Y, Huang J, Shu Q (2019) Mutation of inositol 134-trisphosphate 5/6-kinase6 impairs plant growth and phytic acid synthesis in rice. Plan Theory 8:114. https://doi.org/10.3390/plants8050114
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821. https://doi.org/10.1126/science.1225829
Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J (2013) RNA-programmed genome editing in human cells. Elife 2:00471
Jinek M, Jiang F, Taylor DW, Sternberg SH, Kaya E, Ma E, Kaplan M (2014) Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343:1247997
Johnston RK, Seamon KJ, Saada EA, Podlevsky JD, Branda SS, Timlin JA, Harper JC (2019) Use of anti-CRISPR protein AcrIIA4 as a capture ligand for CRISPR/Cas9 detection. Biosens Bioelectron 141:111361. https://doi.org/10.1016/jbios.2019.111361
Joung J, Ladha A, Saito M, Segel M, Bruneau R, Huang MW, Kim NG, Yu X, Li J, Walker BD, Greninger AL, Jerome KR, Gootenberg JS, Abudayyeh OO, Zhang F (2020) Point-of-care testing for COVID-19 using SHERLOCK diagnostics. medRxiv. https://doi.org/10.1101/2020.05.04.20091231
Jung YJ, Lee HJ, Yu J, Bae S, Cho YG, Kang KK (2021) Transcriptomic and physiological analysis of OsCAO1 knockout lines using the CRISPR/Cas9 system in rice. Plant Cell Rep 40:1013–1024. https://doi.org/10.1007/s00299-020-02607-y
Kampmann M (2018) CRISPRi and CRISPRa screens in mammalian cells for precision biology and medicine. ACS Chem Biol 13(2):406–416. https://doi.org/10.1021/acschembio.7b00657
Kantor A, McClements ME, MacLaren RE (2020) CRISPR-Cas9 DNA base-editing and prime-editing. Int J Mol Sci 21:6240. https://doi.org/10.3390/ijms21176240
Kapusi E, Corcuera-Gomez M, Melnik S, Stoger E (2017) Heritable genomic fragment deletions and small indels in the putative ENgase gene induced by CRISPR/Cas9 in barley. Front Plant Sci 8:540. https://doi.org/10.3389/fpls.2017.00540
Karvelis T, Bigelyte G, Young JK, Hou Z, Zedaveinyte R, Pociute K, Silanskas A, Venclovas C, Siksnys V (2019) PAM recognition by miniature CRISPR-Cas14 triggers programmable double-stranded DNA cleavage. BioRxiv Preprint. https://doi.org/10.1101/654897
Karvelis T, Bigelyte G, Young JK, Hou Z, Zedaveinyte R, Budre K, Paulraj S, Djukanovic V, Gasior S, Silanskas A, Venclovas Č, Siksnys V (2020) PAM recognition by miniature CRISPR-Cas12f nucleases triggers programmable double-stranded DNA target cleavage. Nucleic Acids Res 48(9):5016–5023. https://doi.org/10.1093/nar/gkaa208
Kaul T, Raman NM, Eswaran M, Thangaraj A, Verma R, Sony SK, Sathelly KM, Kaul R, Yadava P, Agrawal PK (2019) Data mining by pluralistic approach on CRISPR gene editing in plants. Front Plant Sci 10:801. https://doi.org/10.3389/fpls.2019.00801
Kaul T, Sony SK, Raman NM, Eswaran M, Verma R, Thangaraj A, Bharti J, Motelb KFA, Kaul R (2020a) How crisp is CRISPR? CRISPR/Cas mediated crop improvement with special focus on nutritional traits. In: Tuteja N, Tuteja R, Passricha N, Saifi S (eds) Advancement in crop improvement techniques, vol 20. Woodhead Printing, New Delhi, pp 159–197
Kaul T, Sony SK, Verma R, Motelb KFA, Thangaraj A, Eswaran M, Bharti J, Nehra M, Kaul R (2020b) Revisiting CRISPR–Cas mediated crop improvement:special focus on nutrition. J Biosci 45:137
Kaul T, Sony SK, Raman NM, Motelb KFA, Bharti J (2021) Genotype–independent regeneration and transformation protocol for rice cultivars. In: Bandyopadhyay A, Thilmony R (eds) Rice genome engineering and gene editing. Methods in Molecular Biology. Humana, New York, NY, p 2238
Kaur K, Gupta AK, Rajput A, Kumar M (2016) Ge-CRISPR - An integrated pipeline for the prediction and analysis of sgRNAs genome editing efficiency for CRISPR/Cas system. Sci Rep 6:1–12. https://doi.org/10.1038/srep30870
Kaur N, Alok A, Shivani KP, Kaur N, Awasthi P, Chaturvedi S, Pandey P, Pandey A, Pandey AK, Tiwari S (2020) CRISPR/Cas9 directed editing of lycopene epsilon-cyclase modulates metabolic flux for beta-carotene biosynthesis in banana fruit. Metab Eng 59:76–86. https://doi.org/10.1016/j.ymben.2020.01.008
Kellner MJ, Koob J, Gootenberg JS, Abudayyeh OO, Zhang F (2019) SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat Protoc 14(10):2986–3012. https://doi.org/10.1038/s41596-019-0210-2
Khan MZ, Haider S, Mansoor S, Amin I (2019a) Targeting plant ssDNA viruses with engineered miniature CRISPR-Cas14a. Trends Biotechnol 37(8):800–804
Khan MSS, Basnet R, Islam SA, Shu Q (2019b) Mutational analysis of OsPLD1 reveals its involvement in phytic acid biosynthesis in rice grains. J Agric Food Chem 67:11436–11443
Kieu NP, Lenman M, Wang ES, Petersen BL, Andreasson E (2021) Mutations introduced in susceptibility genes through CRISPR/Cas9 genome editing confer increased late blight resistance in potatoes. Sci Rep 11:4487. https://doi.org/10.1038/s41598-021-83972-w
Kim D, Bae S, Park J, Kim E, Kim S, Yu HR, Kim JS (2015) Digenome-seq: genome-wide profiling of CRISPR/Cas9 off-target effects in human cells. Nat Methods 12:237
Kim HK, Song M, Lee J, Menon AV, Jung S, Kang YM (2017) In vivo high-throughput profiling of CRISPR-Cpf1 activity. Nat Methods 14:153–159. https://doi.org/10.1038/nmeth4104
Klap C, Yeshayahou E, Bolger AM, Arazi T, Gupta SK, Shabtai S, Usadel B, Salts Y, Barg R (2017) Tomato facultative parthenocarpy results from SlAGAMOUS-LIKE 6 loss of function. Plant Biotechnol J 15:634–647
Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR (2016) Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533:420–424. https://doi.org/10.1038/nature17946
Konwarh R (2020) Can CRISPR/Cas technology be a felicitous stratagem against the COVID-19 fiasco? Prospects and hitches. Front Mol Biosci 7:557377. https://doi.org/10.3389/fmolb2020557377
Koonin EV, Makarova KS, Zhang F (2017) Diversity classification and evolution of CRISPR-Cas systems. Curr Opin Microbiol 37:67–78. https://doi.org/10.1016/j.mib.2017.05.008
Kosicki M, Tomberg K, Bradley A (2018) Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nat Biotechnol 36:765–771
Ku HK, Ha SH (2020) Improving nutritional and functional quality by genome editing of crops: status and perspectives. Front Plant Sci 11:577313. https://doi.org/10.3389/fpls.2020.577313
Kumar P, Malik YS, Ganesh B, Rahangdale S, Saurabh S, Natesan S, Srivastava A, Sharun K, Yatoo MI, Tiwari R, Singh RK, Dhama K (2020) CRISPR-Cas system: an approach with potentials for COVID-19 diagnosis and therapeutics. Front Cell Infect Microbiol 10:576875. https://doi.org/10.3389/fcimb.2020.576875
Kumlehn J, Pietralla J, Hensel G, Pacher M, Puchta H (2018) The CRISPR/Cas revolution continues: From efficient gene editing for crop breeding to plant synthetic biology. J Integr Plant Biol 60:1127–1153. https://doi.org/10.1111/jipb.12734
Kurtz S (2003) The Vmatch large scale sequence analysis software Ref Type. Computer Program:4–12
Labuhn M, Adams FF, Ng M, Knoess S, Schambach A, Charpentier EM, Schwarzer A, Mateo JL, Klusmann JH, Heckl D (2018) Refined sgRNA efficacy prediction improves largeand small-scale CRISPR-Cas9 applications. Nucleic Acids Res 46:1375–1385. https://doi.org/10.1093/nar/gkx1268
Labun K, Montague TG, Gagnon JA, Thyme SB, Valen E (2016) CHOPCHOP v2: a web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res 44:272–276. https://doi.org/10.1093/nar/gkw398
Labun K, Montague TG, Krause M, Torres CYN, Tjeldnes H, Valen E (2019) CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res 47:171–174. https://doi.org/10.1093/nar/gkz365
Larson G, Piperno DR, Allaby RG, Purugganan MD, Andersson L, Arroyo-Kalin M, Barton L, Climer VC, Denham T, Dobney K, Doust AN, Gepts P, Gilbert MTP, Gremillion KJ, Lucas L, Lukens L, Marshall FB, Olsen KM, Pires JC, Richerson PJ, Rubio CR, Sanjur OI, Thomas MG, Fuller DQ (2014) Current perspectives and the future of domestication studies. Proc Natl Acad Sci 111(17):6139–6146. https://doi.org/10.1073/pnas1323964111
Lawhorn IE, Ferreira JP, Wang CL (2014) Evaluation of sgRNA target sites for CRISPR-mediated repression of TP53. PLoS One 9:113232
Ledford H (2020) CRISPR treatment inserted directly into the body for first time. Nature 579:185. https://doi.org/10.1038/d41586-020-00655-8
Ledford H, Callaway E (2020) Pioneers of revolutionary CRISPR gene editing win chemistry Nobel. Nature 586:346–347. https://doi.org/10.1038/d41586-020-02765-9
Lee K, Zhang Y, Kleinstiver BP et al (2019) Activities and specificities of CRISPR/Cas9 and Cas12a nucleases for targeted mutagenesis in maize. Plant Biotechnol J 17(2):362–372. https://doi.org/10.1111/pbi.12982
Lei Y, Lu L, Liu HY, Li S, Xing F, Chen LL (2014) CRISPR-P: a web tool for synthetic single-guide RNA design of CRISPR-system in plants. Mol Plant 7:1494–1496. https://doi.org/10.1093/mp/ssu044
Levin M (1994) The role of risk assessment in developing statutes and regulations. In: Krattiger AF, Rosemarin A (eds) Biosafety for sustainable agriculture: sharing biotechnology regulatory experiences of the western hemisphere. ISAAA and Stockholm Environment Institute, Ithaca/Stockholm, pp 127–137
Li Z, Liu ZB, Xing A, Moon BP, Koellhoffer JP, Huang L, Ward RT, Clifton E, Falco SC, Cigan AM (2015) Cas9- guide RNA directed genome editing in Soybean. Plant Physiol 169:960–970
Li C, Liu C, Qi X, Wu Y, Fei X, Mao L, Cheng B, Li X, Xie C (2017) RNA-guided Cas9 as an in vivo desired-target mutator in maize. Plant Biotechnol J 15:1566–1576
Li S, Cheng Q, Liu J, Nie X, Zhao G, Wang J (2018a) CRISPR-Cas12a has both cis- and trans-cleavage activities on single-stranded DNA. Cell Res 28:1–3. https://doi.org/10.1038/s41422-018-0022-x
Li X, Wang Y, Chen S, Tian H, Fu D, Zhu B, Luo Y, Zhu H (2018b) Lycopene is enriched in tomato fruit by CRISPR/Cas9-mediated multiplex genome editing. Front Plant Sci 9:559
Li S, Li J, He Y, Xu M, Zhang J, Du W, Zhao Y, Xia L (2019) Precise gene replacement in rice by RNA transcript templated homologous recombination. Nat Biotechnol 37:445–450
Li Z, Zhang H, Xiao R, Han R, Chang L (2021) Cryo-EM structure of the RNA-guided ribonuclease Cas12g. Nat Chem Biol 17(4):387–393. https://doi.org/10.1038/s41589-020-00721-2
Liang Z, Zhang K, Chen K, Gao C (2014) Targeted mutagenesis in Zea mays using TALENs and the CRISPR/Cas system. Genet Genomics 41:63–68
Liang Z, Chen K, Yan Y, Zhang Y, Gao C (2018) Genotyping genome-edited mutations in plants using CRISPR ribonucleoprotein complexes. Plant Biotechnol J 16(12):2053–2062. https://doi.org/10.1111/pbi.12938
Lieber MR (2010) The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem 79:181–211. https://doi.org/10.1146/annurev.biochem.052308.093131
Lillestøl RK, Redder P, Garrett RA, Brügger KA (2006) A putative viral defence mechanism in archaeal cells. Archaea 2(1):59–72. https://doi.org/10.1155/2006/542818
Lin J, Wong KC (2018) Off-target predictions in CRISPR-Cas9 gene editing using deep learning. Bioinformatics 34:656–663. https://doi.org/10.1093/bioinformatics/bty.554
Lin Q, Zong Y, Xue C, Wang S, Jin S, Zhu Z, Wang Y, Anzalone AV, Raguram A, Doman JL, Liu DR, Gao C (2020) Prime genome editing in rice and wheat. Nat Biotechnol 38:582–585. https://doi.org/10.1038/s41587-020-0455-x
Lintner NG, Kerou M, Brumfield SK, Graham S, Liu H, Naismith JH, Sdano M, Peng N, She Q, Copie V et al (2011) Structural and functional characterization of an archaeal clustered regularly interspaced short palindromic repeat (CRISPR)-associated complex for antiviral defense (CASCADE). J Biol Chem 286:21643–21656. https://doi.org/10.1074/jbcM111238485
Listgarten J, Weinstein M, Kleinstiver BP, Sousa AA, Joung JK, Crawford J, Gao K, Hoang L, Elibol M, Doench JG, Fusi N (2018) Prediction of off-target activities for the end-to-end design of CRISPR guide RNAs. Nat Biomed Eng 2:38–47. https://doi.org/10.1038/s41551-017-0178-6
Liu H, Wei Z, Dominguez A, Li Y, Wang X, Qi LS (2015) CRISPR-ERA: a comprehensive design tool for CRISPR-mediated gene editing repression and activation. Bioinformatics 31:3676–3678. https://doi.org/10.1093/bioinformatics/btv423
Liu H, Ding Y, Zhou Y, Jin W, Xie K, Chen LL (2017) CRISPR-P 20: an improved CRISPR-Cas9 tool for genome editing in plants. Mol Plant 10:530–532. https://doi.org/10.1016/jmolp.2017.01.003
Liu JJ, Orlova N, Oakes BL, Ma E, Spinner HB, Baney KLM, Chuck J, Tan D, Knott GJ, Harrington LB, Al-Shayeb B, Wagner A, Brötzmann J, Staahl BT, Taylor KL, Desmarais J, Nogales E, Doudna JA (2019a) CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566(7743):218–223. https://doi.org/10.1038/s41586-019-0908-x
Liu Y, Wan X, Wang B (2019b) Engineered CRISPRa enables programmable eukaryote-like gene activation in bacteria. Nat Commun 10:3693
Liu L, Gallagher J, Arevalo ED, Chen R, Skopelitis T, Wu Q, Bartlett M, Jackson D (2021) Enhancing grain-yield-related traits by CRISPR-Cas9 promoter editing of maize CLE genes. Nat Plants 7(3):287–294. https://doi.org/10.1038/s41477-021-00858-5
Lloyd A, Plaisier CL, Carroll D, Drews GN (2005) Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc Natl Acad Sci U S A 102:232–2237
Louwen R, Staals RH, Endtz HP, Van Baarlen P, van der Oost J (2014) The role of CRISPR-Cas systems in virulence of pathogenic bacteria. Microbiol Mol Biol Rev 78(1):74–88. https://doi.org/10.1128/MMBR00039-13
Lu Y, Tian Y, Shen R, Yao Q, Wang M, Chen M, Dong J, Zhang T, Li F, Lei M, Zhu JK (2020) Targeted efficient sequence insertion and replacement in rice. Nat Biotechnol 38(12):1402–1407. https://doi.org/10.1038/s41587-020-0581-5
Luo J, Chen W, Xue L, Tang B (2019) Prediction of activity and specificity of CRISPR-Cpf1 using convolutional deep learning neural networks. BMC Bioinformatics 20:1–10. https://doi.org/10.1186/s12859-019-2939-6
Ma X, Feng F, Zhang Y, Elesawi IE, Xu K, Li T, Mei H, Liu H, Gao N, Chen C, Luo L, Yu S (2019) A novel rice grain size gene OsSNB was identified by genome-wide association study in natural population. PLoS Genet 15:1008191. https://doi.org/10.1371/journal.pgen.1008191
MacPherson CR, Scherf A (2015) Flexible guide-RNA design for CRISPR applications using Protospacer Workbench. Nat Biotechnol 33:805–806. https://doi.org/10.1038/nbt.3291
Maeder ML, Thibodeau-Beganny S, Osiak A, Wright DA, Anthony RM, Eichtinger M, Jiang T, Foley JE, Winfrey RJ, Townsend JA, Unger-Wallace E, Sander JD, Müller-Lerch F, Fu F, Pearlberg J, Göbel C, Dassie JP, Pruett-Miller SM, Porteus MH, Sgroi DC, Iafrate AJ, Dobbs D, McCray PB, Cathomen T, Voytas DF, Joung JK (2008) Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol Cell 31(2):294–301. https://doi.org/10.1016/j.molcel.2008.06.016
Mahfouz MM, Piatek A, Stewart CN (2014) Genome engineering via TALENs and CRISPR/Cas9 systems: challenges and perspectives. Plant Biotechnol 12(8):1006–1014. https://doi.org/10.1111/pbi.12256
Maji B, Gangopadhyay SA, Lee M, Shi M, Wu P, Heler R, Mok B, Lim D, Siriwardena SU, Paul B, Dančík V (2019) A high-throughput platform to identify small-molecule inhibitors of CRISPR-Cas9. Cell 177(4):1067–1079
Makarova KS, Grishin NV, Shabalina SA, Wolf YI, Koonin EV (2006) A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery functional analogies with eukaryotic RNAi and hypothetical mechanisms of action. Biol Direct 1:7. https://doi.org/10.1186/1745-6150-1-7
Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P, Moineau S, Mojica FJ, Terns RM, Terns MP, White MF, Yakunin AF, Garrett RA, Van der Oost J, Backofen R, Koonin EV (2011) Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol 9:467–477. https://doi.org/10.1038/nrmicro2577
Makarova KS, Wolf YI, Alkhnbashi OS, Costa F, Shah SA, Saunders SJ (2015) An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol 13:722–736. https://doi.org/10.1038/nrmicro3569
Makarova KS, Wolf YI, Koonin EV (2018) Classification and nomenclature of CRISPR-Cas systems: where from here? Cris J 1:325–336
Makarova KS, Wolf YI, Iranzo J, Shmakov SA, Alkhnbashi OS, Brouns SJJ, Charpentier E, Cheng D, Haft DH, Horvath P, Moineau S, Mojica FJM, Scott D, Shah SA, Siksnys V, Terns MP, Venclovas Č, White MF, Yakunin AF, Yan W, Zhang F, Garrett RA, Backofen R, van der Oost J, Barrangou R, Koonin EV (2020) Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat Rev Microbiol 18:67–83. https://doi.org/10.1038/s41579-019-0299-x
Mali P, Aach J, Stranges PB, Esvelt KM, Moosburner M, Kosuri S, Yang L, Church GM (2013a) CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat Biotechnol 31:833–838
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM (2013b) RNA-guided human genome engineering via Cas9. Science 339:823–826. https://doi.org/10.1126/science.1232033
Malnoy M, Viola R, Jung MH, Koo OJ, Kim S, Kim JS, Velasco R, Nagamangala KC (2016) DNA-free genetically edited grapevine and apple protoplast using Crispr/Cas9 ribonucleoproteins. Front Plant Sci 7:1904
Marino ND, Pinilla-Redondo R, Csörgő B, Bondy-Denomy J (2020) Anti- CRISPR protein applications: natural brakes for CRISPR-Cas technologies. Nat Methods 17(5):471–479
Marraffini LA, Sontheimer EJ (2008) CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322:1843–1845. https://doi.org/10.1126/science.1165771
McCaw ME, Lee K, Kang M, Zobrist JD, Azanu MK, Birchler JA, Wang K (2021) Development of a transformable fast-flowering mini-maize as a tool for maize gene editing. Front Genome Ed 2:622227. https://doi.org/10.3389/fgeed.2020.622227
McKenna A, Shendure J (2018) FlashFry: a fast and flexible tool for large-scale CRISPR target design. BMC Biol 16:4–9. https://doi.org/10.1186/s12915-018-0545-0
Mendoza BJ, Trinh CT (2018) Enhanced guide-RNA design and targeting analysis for precise CRISPR genome editing of single and consortia of industrially relevant and non-model organisms. Bioinformatics 34:16–23. https://doi.org/10.1093/bioinformatics/btx564
Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X, Wan J, Gu H, Qu LJ (2013) Targeted mutagenesis in rice using CRISPR–Cas system. Cell Res 23:1233–1236
Miller JC, Tan S, Qiao G, Barlow KA, Wang J, Xia DF, Meng X, Paschon DE, Leung E, Hinkley SJ, Dulay GP, Hua KL, Ankoudinova I, Cost GJ, Urnov FD, Zhang HS, Holmes MC, Zhang L, Gregory PD, Rebar EJ (2011) A TALE nuclease architecture for efficient genome editing. Nat Biotechnol 29(2):143–148
Minkenberg B, Zhang J, Xie K, Yang Y (2019) CRISPR-PLANT v2: an online resource for highly specific guide RNA spacers based on improved off-target analysis. Plant Biotechnol J 17:5–8. https://doi.org/10.1111/pbi.13025
Mojica FJ, Ferrer C, Juez G, Rodríguez-Valera F (1995) Long stretches of short tandem repeats are present in the largest replicons of the archaea Haloferax mediterranei and Haloferax volcanii and could be involved in replicon partitioning. Mol Microbiol 17:85–93. https://doi.org/10.1111/j1365-29581995mmi_17010085x
Mojica FJ, Díez-Villaseñor C, Soria E, Juez G (2000) Biological significance of a family of regularly spaced repeats in the genomes of archaea bacteria and mitochondria. Mol Microbiol 36:244–246. https://doi.org/10.1046/j1365-2958200001838x
Mojica FJM, Díez-Villaseñor C, García-Martínez J, Soria E (2005) Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 60:174–182. https://doi.org/10.1007/s00239-004-0046-3
Mojica FJ, Diez-Villasenor C, Garcia-Martinez J, Almendros C (2009) Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 155:733–740
Montague TG, Cruz JM, Gagnon JA, Church GM, Valen E (2014) CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res 42:401–407. https://doi.org/10.1093/nar/gku410
Moore R, Chandrahas A, Bleris L (2014) Transcription activator-like effectors: a toolkit for synthetic biology. ACS Synth Biol 3(10):708–716. https://doi.org/10.1021/sb400137b
Moreno-Mateos MA, Vejnar CE, Beaudoin JD, Fernandez JP, Mis EK, Khokha MK, Giraldez AJ (2015) CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat Methods 12:982–988. https://doi.org/10.1038/nmeth3543
Moscou MJ, Bogdanove AJ (2009) A simple cipher governs DNA recognition by TAL effectors. Science 326(5959):1501
Naeem M, Majeed S, Hoque MZ, Ahmad I (2020) Latest developed strategies to minimize the off-target effects in CRISPR-Cas-mediated genome editing. Cell 9(7):1608. https://doi.org/10.3390/cells9071608
Naito Y, Hino K, Bono H, Ui-Tei K (2015) CRISPRdirect: Software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31:1120–1123. https://doi.org/10.1093/bioinformatics/btu743
Nakata A, Amemura M, Makino K (1989) Unusual nucleotide arrangement with repeated sequences in the Escherichia coli K-12 chromosome. J Bacteriol 171:3553–3556. https://doi.org/10.1128/jb.171.6.3553-3556.1989
Newsom S, Parameshwaran HP, Martin L, Rajan R (2021) The CRISPR-Cas mechanism for adaptive immunity and alternate bacterial functions fuels diverse biotechnologies. Front Cell Infect Microbiol 10:619763. https://doi.org/10.3389/fcimb.2020.619763
Nickel L, Ulbricht A, Alkhnbashi OS, Förstner KU, Cassidy L, Weidenbach K, Backofen R, Schmitz RA (2018) Cross-cleavage activity of Cas6b in crRNA processing of two different CRISPR-Cas systems in Methanosarcina mazei Gö1. RNA Biol 16:492–503
Niewoehner O, Jinek M (2016) Structural basis for the endoribonuclease activity of the type III-A CRISPR-associated protein Csm6. RNA 22:318–329
Nishimasu H, Ran FA, Hsu PD, Konermann S, Shehata SI, Dohmae N, Ishitani R, Zhang F, Nureki O (2014) Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156:935–949. https://doi.org/10.1016/jcell201402001
Nonaka S, Arai C, Takayama M, Matsukura C, Ezura H (2017) Efficient increase of ɣ-aminobutyric acid (GABA) content in tomato fruits by targeted mutagenesis. Sci Rep 7:7057
Nunez JK, Chen J, Pommier GC, Cogan JZ, Replogle JM, Adriaens C, Ramadoss GN, Shi Q, Hung KL, Samelson AJ, Pogson AN, Kim JYS, Chung A, Leonetti MD, Chang HY, Kampmann M, Bernstein BE, Hovestadt V, Gilbert LA, Weissman JS (2021) Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184(9):2503–2519. https://doi.org/10.1016/jcell202103025
O’Brien A, Bailey TL (2014) GT-Scan: Identifying unique genomic targets. Bioinformatics 30:2673–2675. https://doi.org/10.1093/bioinformatics/btu354
Okuzaki A, Ogawa T, Koizuka C, Kaneko K, Inaba M, Imamura J, Koizuka N (2018) CRISPR/Cas9-mediated genome editing of the fatty acid desaturase 2 gene in Brassica napus. Plant Physiol Biochem 131:63–69
Orlowski J, Boniecki M, Bujnicki JM (2007) I-Ssp6803I: the first homing endonuclease from the PD-(D/E)XK superfamily exhibits an unusual mode of DNA recognition. Bioinformatics 23(5):527–530
Özcan A, Krajeski R, Ioannidi E, Lee B, Gardner A, Makarova KS, Koonin EV, Abudayyeh OO, Gootenberg JS (2021) Programmable RNA targeting with the single-protein CRISPR effector Cas7-11. Nature 597(7878):720–725. https://doi.org/10.1038/s41586-021-03886-5
Pacher M, Puchta H (2017) From classical mutagenesis to nuclease-based breeding –directing natural DNA repair for a natural end-product. Plant J 90:819–833. https://doi.org/10.1111/tpj.13469
Pak H, Wang H, Kim Y, Song U, Tu M, Wu D, Jiang L (2020) Creation of male-sterile lines that can be restored to fertility by exogenous methyl jasmonate for the establishment of a two-line system for the hybrid production of rice (Oryza sativa L). Plant Biotechnol J 19:365–374. https://doi.org/10.1111/pbi.13471
Pannunzio NR, Watanabe G, Lieber MR (2017) Nonhomologous DNA end joining for repair of DNA double-strand breaks. J Biol Chem:117000374. https://doi.org/10.1074/jbcTM117000374
Paparini A, Romano-Spica V (2006) Gene transfer and cauliflower mosaic virus promoter 35S activity in mammalian cells. J Environ Sci Health B41(4):437–449. https://doi.org/10.1080/03601230600616957
Park J, Bae S, Kim JS (2015) Cas-designer: a web-based tool for choice of CRISPR-Cas9 target sites. Bioinformatics 31:4014–4016. https://doi.org/10.1093/bioinformatics/btv537
Pausch P, Al-Shayeb B, Bisom-Rapp E, Tsuchida CA, Li Z, Cress BF, Knott GJ, Jacobsen SE, Banfield JF, Doudna JA (2020) CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 369(6501):333–337. 10.1126/science.abb1400
Pauwels K, Podevin N, Breyer D, Carroll D, Herman P. 2014 Engineering nucleases for gene targeting: safety and regulatory considerations. N Biotechnol 31(1):18-27. https://doi.org/10.1016/j.nbt.2013.07.001
Peng D, Tarleton R (2015) EuPaGDT: a web tool tailored to design CRISPR guide RNAs for eukaryotic pathogens. Microb Genomics 1:1–7. https://doi.org/10.1099/mgen.0.000033
Peng H, Zheng Y, Blumenstein M, Tao D, Li J (2018) CRISPR/Cas9 cleavage efficiency regression through boosting algorithms and Markov sequence profiling. Bioinformatics 34:3069–3077. https://doi.org/10.1093/bioinformatics/bty298
Pineda M, Lear A, Collins JP, Kiani S (2019) Safe CRISPR: challenges and possible solutions. Trends Biotechnol 37:389–401. https://doi.org/10.1016/jtibtech.2018.09.010
Pliatsika V, Rigoutsos I (2015) “Off-Spotter”: very fast and exhaustive enumeration of genomic lookalikes for designing CRISPR/Cas guide RNAs. Biol Direct 10:1–10. https://doi.org/10.1186/s13062-015-0035-z
Porto EM, Komor AC, Slaymaker IM, Yeo GW (2020) Base editing: advances and therapeutic opportunities. Nat Rev Drug Discov 19:839–859. https://doi.org/10.1038/s41573-020-0084-6
Pourcel C, Salvignol G, Vergnaud G (2005) CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA and provide additional tools for evolutionary studies. Microbiology 151:653–663. https://doi.org/10.1099/mic.0.27437-0
Prykhozhij SV, Rajan V, Gaston D, Berman JN (2015) CRISPR multitargeter: a web tool to find common and unique CRISPR single guide RNA targets in a set of similar sequences. PLoS One 10:1–18. https://doi.org/10.1371/journal.pone.0119372
Puchta H (1999) Double-strand break-induced recombination between ectopic homologous sequences in somatic plant cells. Genetics 152(3):1173–1181
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA (2021) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 184(3):844
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Mende DR, Li J, Xu J, Li S, Li D, Cao J, Wang B, Liang H, Zheng H, Xie Y, Wang J (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464(7285):59–65. https://doi.org/10.1038/nature.08821
Rahman MK, Rahman MS (2017) CRISPRpred: a flexible and efficient tool for sgRNAs on-target activity prediction in CRISPR/Cas9 systems. PLoS One 12:1–14. https://doi.org/10.1371/journal.pone.0181943
Rai M, He C, Wu R (2009) Comparative functional analysis of three abiotic stress-inducible promoters in transgenic rice. Transgenic Res 18:787–799
Reardon S (2016) First CRISPR clinical trial gets green light from US panel. Nature. https://doi.org/10.1038/nature.2016.20137
Rees HA, Liu DR (2018) Base editing: precision chemistry on the genome and transcriptome of living cells. Nat Rev Genet 19(12):770–788. https://doi.org/10.1038/s41576-018-0059-1
Reis A, Hornblower B, Robb B, Tzertzinis G (2014) CRISPR/Cas9 & targeted genome editing: new era in molecular biology. Biolabs
Ren X, Yang Z, Xu J, Sun J, Mao D, Hu Y, Yang SJ, Qiao HH, Wang X, Hu Q, Deng P, Liu LP, Ji JY, Li JB, Ni JQ (2014) Enhanced specificity and efficiency of the CRISPR/Cas9 system with optimized sgRNA parameters in Drosophila. Cell Rep 9:1151–1162
Ren C, Liu X, Zhang Z, Wang Y, Duan W, Li S, Liang Z (2016) CRISPR/Cas9-mediated efficient targeted mutagenesis in Chardonnay (Vitis vinifera L). Sci Rep 6:1–9
Roginsky J (2018) Analyzing CRISPR editing results. Genet Eng Biotechnol News 38:24–26. https://doi.org/10.1089/gen.381113
Rosado A, Craig W (2017) Biosafety regulatory systems overseeing the use of genetically modified organisms in the Latin America and Caribbean region. AgBioForum 20:120–132
Rosen LE, Morrison HA, Masri S, Brown MJ, Springstubb B, Sussman D, Stoddard BL (2006) Seligman LM: homing endonuclease I-CreI derivatives with novel DNA target specificities. Nucleic Acids Res 34(17):4791–4800
Samai P, Pyenson N, Jiang W, Goldberg GW, Hatoum-Aslan A, Marraffini LA (2015) Co-transcriptional DNA and RNA cleavage during type III CRISPR–Cas immunity. Cell 161:1164–1174
Samalov M, Moore I (2021) The steroid-inducible pOp6/LhGR gene expression system is fast, sensitive and does not cause plant growth defects in rice (Oryza sativa). BMC Plant Biol 21:461. https://doi.org/10.1186/s12870-021-03241-w
Sanchez-Leon S, Gil-Humanes J, Ozuna CV, Gimenez MJ, Sousa C, Voytas DF, Barro F (2018) Low-gluten nontransgenic wheat engineered with CRISPR/Cas9. Plant Biotechnol 16:902–910. https://doi.org/10.1111/pbi.12837
Sander JD, Joung JK (2014) CRISPR-Cas systems for editing regulating and targeting genomes. Nat Biotechnol 32:347. https://doi.org/10.1038/nbt.2842
Sander JD, Maeder ML, Reyon D, Voytas DF, Joung JK, Dobbs D (2010) ZiFiT (Zinc Finger Targeter): an updated zinc finger engineering tool. Nucleic Acids Res 38:462–468. https://doi.org/10.1093/nar/gkq.319
Sashidhar N, Harloff HJ, Potgieter L, Jung C (2020) Gene editing of three BnITPK genes in tetraploid oilseed rape leads to significant reduction of phytic acid in seeds. Plant Biotechnol 18(11):2241–2250. https://doi.org/10.1111/pbi.13380
Satomura A, Nishioka R, Mori H, Sato K, Kuroda K, Ueda M (2017) Precise genome-wide base editing by the CRISPR Nickase system in yeast. Sci Rep 7:2095. https://doi.org/10.1038/s41598-017-02013-7
Sauer NJ, Narvaez-Vasquez J, Mozoruk J, Miller RB, Warburg ZJ, Woodward MJ, Mihiret YA, Lincoln TA, Segami RE, Sanders SL, Walker KA, Beetham PR, Schöpke CR, Gocal GF (2016) Oligonucleotide-mediated genome editing provides precision and function to engineered nucleases and antibiotics in plants. Plant Physiol 170:1917–1928
Savage DF (2019) Cas14: big advances from small CRISPR proteins. Biochem 58:1024–1025
Schenke D, Cai D (2020) Applications of CRISPR/Cas to improve crop disease resistance: beyond inactivation of susceptibility factors. iScience 23(9):101478. https://doi.org/10.1016/j.isci.2020.101478
Schmidt SM, Belisle M, Frommer WB (2020) The evolving landscape around genome editing in agriculture. EMBO Rep 21:19–22. https://doi.org/10.15252/embr.202050680
Scholefield J, Harrison PT (2021) Prime editing – an update on the field. Gene Ther 28:396–401. https://doi.org/10.1038/s41434-021-00263-9
Sedeek KEM, Mahas A, Mahfouz M (2019) Plant genome engineering for targeted improvement of crop traits. Front Plant Sci 10:114. https://doi.org/10.3389/fpls.2019.00114
Settachaimongkon S, Nout MJR, Antunes FEC, Van Hooijdonk Toon CM, Zwietering Marcel H, Smid Eddy J, Van Valenberg Hein JF (2014) The impact of selected strains of probiotic bacteria on metabolite formation in set yoghurt. Int Dairy J 38(1):1–10. https://doi.org/10.1016/j.idairyj.2014.04.002
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu JL, Gao C (2013) Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31(8):686–688
Shan Q, Wang Y, Li J, Gao C (2014) Genome editing in rice and wheat using the CRISPR/Cas system. Nat Protoc 9:2395–2410
Shan S, Soltis PS, Soltis DE, Yang B (2020) Considerations in adapting CRISPR/Cas9 in nongenetic model plant. Appl Plant Sci 8:11314. https://doi.org/10.1002/aps3.11314
Shen H (2014) First monkeys with customized mutations born. Nature. https://doi.org/10.1038/nature.2014.14611
Shi J, Gao H, Wang H, Lafitte HR, Archibald RL, Yang M, Hakimi SM, Mo H, Habben JE (2017) ARGOS 8 variants generated by CRISPR/Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol J 15:207–216
Shin J, Jiang F, Liu JJ, Bray NL, Rauch BJ, Baik SH, Nogales E, Bondy-Denomy J, Corn JE, Doudna JA (2017) Disabling Cas9 by an anti-CRISPR DNA mimic. Sci Adv 3:1701620
Shufen C, Yicong C, Baobing F, Guiai J, Zhonghua S, Ju L, Shaoqing T, Jianlong W, Peisong H, Xiangjin W (2019) Editing of rice isoamylase gene ISA1 provides insights into its function in starch formation. Ric Sci 26(2):77–87. https://doi.org/10.1016/j.rsci.2018.07.001
Sidira MV, Santarmaki M, Kiourtzidis AA, Argyri OS, Papadopoulou N, Chorianopoulos C, Tassou S, Kaloutsas A, Galanis KY (2017) Evaluation of immobilized Lactobacillus plantarum 2035 on whey protein as adjunct probiotic culture in yoghurt production. Lebensm Wiss Technol 75:137–146
Singh R, Kuscu C, Quinlan A, Qi Y, Adli M (2015) Cas9-chromatin binding information enables more accurate CRISPR off-target prediction. Nucleic Acids Res 43:1–8. https://doi.org/10.1093/nar/gkv575
Singh A, Roychowdhury R, Singh T, Wang W, Yadav D, Kumar A, Modi A, Rai AC, Ghughe S, Kumar A, Kumar Singh P (2020) Improvement of crop’s stress tolerance by gene editing CRISPR/CAS9 system. In: Choudhury RR, Hasanuzzaman S, Srivastava M (eds) Sustainable agriculture in the era of climate change. Springer, Cham
Sriboon S, Li H, Guo C et al (2020) Knock-out of TERMINAL FLOWER 1 genes altered flowering time and plant architecture in Brassica napus. BMC Genet 21:52. https://doi.org/10.1186/s12863-020-00857-z
Staals RH, Brouns SJ (2013) In: Barrangou R, van de Oost J (eds) Distribution and mechanism of the type I CRISPR-Cas systems. Springer, Berlin, Heidelberg, pp 115–144
Staals RH, Zhu Y, Taylor DW, Kornfeld JE, Sharma K, Barendregt A, Koehorst JJ, Vlot M, Neupane N, Varossieau K, Sakamoto K, Suzuki T, Dohmae N, Yokoyama S, Schaap PJ, Urlaub H, Heck AJ, Nogales E, Doudna JA, Shinkai A, van der Oost J (2014) RNA targeting by the type III-A CRISPR–Cas Csm complex of Thermus thermophiles. Mol Cell 56:518–530
Stemmer M, Thumberger T, Del SKM, Wittbrodt J, Mateo JL (2015) CCTop: an intuitive flexible and reliable CRISPR/Cas9 target prediction tool. PLoS One 10:1–11. https://doi.org/10.1371/journal.pone.0124633
Sun Q, Gao F, Zhao L, Li K, Zhang J (2010) Identification of a new 130 bp cis-acting element in the TsVP1 promoter involved in the salt stress response from Thellungiella halophile. BMC Plant Biol 10:90
Sun X, Hu Z, Chen R, Jiang Q, Song G, Zhang H, Xi Y (2015) Targeted mutagenesis in soybean using the CRISPR-Cas9 system. Sci Rep 5:10342
Sun Y, Zhang X, Wu C, He Y, Ma Y, Hou H, Guo X, Du W, Zhao Y, Xia L (2016) Engineering herbicide-resistant rice plants through CRISPR/Cas9-mediated homologous recombination of acetolactate synthase. Mol Plant 9:628–631
Sun Y, Jiao G, Liu Z, Zhang X, Li J, Guo X, Du WDJ, Francis F, Zhao Y, Xia L (2017) Generation of high amylose rice through CRISPR/Cas9-mediated targeted mutagenesis of starch branching enzymes. Front Plant Sci 8:298
Sun SK, Xu XJ, Tang Z, Tang Z, Huang XY, Wirtz M, Hell R, Zhao FJ (2021) A molecular switch in sulfur metabolism to reduce arsenic and enrich selenium in rice grain. Nat Commun 12:1392
Svitashev S, Young JK, Schwartz C, Gao H, Falco SC, Cigan AM (2015) Targeted mutagenesis precise gene editing and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol 169:931–945
Swarts DC, Jinek M (2019) Mechanistic insights into the Cis- and Trans-acting deoxyribonuclease activities of Cas12a. Mol Cell 73:589–600. https://doi.org/10.1016/j.molcel.2018.11.021
Tabassum J, Ahmad S, Hussain B, Mawia AM, Zeb A, Ju L (2021) Applications and potential of genome-editing systems in rice improvement: current and future perspectives. Agronomy 11:1359. https://doi.org/10.3390/agronomy11071359
Tang L, Mao B, Li Y, Lv Q, Zhang L, Chen C, He H, Wang W, Zeng X, Shao Y, Pan Y, Hu Y, Peng Y, Fu X, Li H, Xia S, Zhao B (2017) Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield. Sci Rep 7:14438
Tang T, Yu X, Yang H, Gao Q, Ji H, Wang Y, Yan G, Peng Y, Luo H, Liu K, Li X, Ma C, Kang C, Dai C (2018) Development and validation of an effective CRISPR/Cas9 vector for efficiently isolating positive transformants and transgene-free mutants in a wide range of plant species. Front Plant Sci 9:1533. https://doi.org/10.3389/fpls.2018.01533
Thygesen P (2019) Clarifying the regulation of genome editing in Australia: situation for genetically modified organisms. Transgenic Res 28:151–159. https://doi.org/10.1007/s11248-019-00151-4
Tian Y, Chen K, Li X, Zheng Y, Chen F (2020) Design of high-oleic tobacco (Nicotiana tabacum L) seed oil by CRISPR-Cas9-mediated knockout of NtFAD2–2. BMC Plant Biol 20:233. https://doi.org/10.1186/s12870-020-02441-0
Tiwari M, Trivedi P, Pandey A (2020) Emerging tools and paradigm shift of gene editing in cereals fruits and horticultural crops for enhancing nutritional value and food security. Food Ener Secur 10(1):e258. https://doi.org/10.1002/fes.3258
Townsend JA, Wright DA, Winfrey RJ, Fu F, Maeder ML, Joung JK, Voytas DF (2009) High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 459:442–445
Tsai SQ, Joung JK (2016) Defining and improving the genome-wide specificities of CRISPR-Cas9 nucleases. Nat Rev Genet 17(5):300–312. https://doi.org/10.1038/nrg.2016.28
Tsuda M, Watanabe KN, Ohsawa R (2019) Regulatory status of genome edited organisms under the Japanese Cartagena act. Front Bioeng Biotechnol 7:387. https://doi.org/10.3389/fbioe.2019.00387
Tuncel A, Corbin KR, Ahn-Jarvis J, Harris S, Hawkins E, Smedley MA, HarwoodW WFJ, Patron NJ, Smith AM (2019) Cas9-mediated mutagenesis of potato starch-branching enzymes generates a range of tuber starch phenotypes. Plant Biotechnol J 17:2259–2271
Upadhyay SK, Sharma S (2014) SSFinder: high throughput CRISPR-Cas target sites prediction tool. Biomed Res Int. https://doi.org/10.1155/2014/742482
Upadhyay SK, Kumar J, Alok A, Tuli R (2013) RNA guided genome editing for target gene mutations in wheat. G3 (Bethesda) 3:2233–2238
Vu TV, Sivankalyani V, Kim EJ, Doan DTH, Tran MT, Kim J, Kim JY (2020) Highly efficient homology-directed repair using CRISPR/Cpf1-geminiviral replicon in tomato. Plant Biotechnol J 18(10):2133–2143. https://doi.org/10.1111/pbi.13373
Wagner DL, Peter L, Schmueck-Henneresse M (2021) Cas9-directed immune tolerance in humans—a model to evaluate regulatory T cells in gene therapy? Gene Ther 28(9):549–559. https://doi.org/10.1038/s41434-021-00232-2
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32:947–951
Wang F, Wang C, Liu P, Lei C, Hao W, Gao Y, Liu YG, Zhao K (2016) Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS One 11(4):0154027. https://doi.org/10.1371/journal.pone.0154027
Wang M, Lu Y, Botella JR, Mao Y, Hua K, Zhu JK (2017) Gene targeting by homology-directed repair in rice using a geminivirus-based CRISPR/Cas9 system. Mol Plant 10(7):1007–1010. https://doi.org/10.1016/jmolp.2017.03.002
Wang W, Simmonds J, Pan Q, Davidson D, He F, Battal A, Akhunova A, Trick HN, Uauy C, Akhunov E (2018a) Gene editing and mutagenesis reveal inter-cultivar differences and additivity in the contribution of TaGW2 homoeologues to grain size and weight in wheat. Theor Appl Genet 131:2463–2475
Wang M, Wang S, Liang Z, Shi W, Gao C, Xia G (2018b) From genetic stock to genome editing: gene exploitation in wheat. Trends Biotechnol 36:160–172
Wang H, Wu Y, Zhang Y, Yang J, Fan W, Zhang H, Zhao S, Yuan L, Zhang P (2019) CRISPR/Cas9-based mutagenesis of starch biosynthetic genes in sweet potato (Ipomoea Batatas) for the improvement of starch quality. Int J Mol Sci 20:4702
Wang D, Wang K, Cai Y (2020) An overview of development in gene therapeutics in China. Gene Ther 27:338–348
Weeks DP, Spalding MH, Yang B (2016) Use of designer nucleases for targeted gene and genome editing in plants. Plant Biotechnol 14:483–495
Wong N, Liu W, Wang X (2015) WU-CRISPR: characteristics of functional guide RNAs for the CRISPR/Cas9 system. Genome Biol 16:1–8. https://doi.org/10.1186/s13059-015-0784-0
Woo JW, Kim J, Kwon SI, Corvalán C, Cho SW, Kim H, Kim S, Kim S, Choe S, Kim J (2015) DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat Biotechnol 33(11):1162–1164. https://doi.org/10.1038/nbt.3389
Wu X, Scott DA, Kriz AJ, Chiu AC, Hsu PD, Dadon DB (2014) Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat Biotechnol 32:670–676. https://doi.org/10.1038/nbt.2889
Wu H, Awan FS, Vilarinho A, Zeng Q, Kannan B, Phipps T (2015) Transgene integration complexity and expression stability following biolistic or Agrobacterium-mediated transformation of sugarcane. In Vitro Cell Devel Biol Plant 51:603–611
Wu S, Zhu H, Liu J, Yang Q, Shao X, Bi F, Hu C, Huo H, Chen K, Yi G (2020) Establishment of a PEG-mediated protoplast transformation system based on DNA and CRISPR/Cas9 ribonucleoprotein complexes for banana. BMC Plant Biol 20:425. https://doi.org/10.1186/s12870-020-02609-8
Xiao A, Cheng Z, Kong L, Zhu Z, Lin S, Gao G et al (2014) CasOT: a genome-wide Cas9/gRNA off-target searching tool. Bioinformatics 30:1180–1182. https://doi.org/10.1093/bioinformatics/btt.764
Xie X, Liu YG (2021) De novo domestication towards new crops. Nat Sci Rev 8(4):033. https://doi.org/10.1093/nsr/nwab.033
Xie K, Yang Y (2013) RNA-guided genome editing in plants using a CRISPR-Cas system. Mol Plant 6:1975–1983. https://doi.org/10.1093/mp/sst.119
Xie S, Shen B, Zhang C, Huang X, Zhang Y (2014) SgRNAcas9: a software package for designing CRISPR sgRNA and evaluating potential off-target cleavage sites. PLoS One 9:1–9. https://doi.org/10.1371/journal.pone.0100448
Xie X, Ma X, Zhu Q, Zeng D, Li G, Liu YG (2017) CRISPR-GE: a convenient software toolkit for CRISPR-based genome editing. Mol Plant 10:1246–1249. https://doi.org/10.1016/j.molp.2017.06.004
Xu H, Xiao T, Chen CH, Li W, Meyer CA, Wu Q, Wu D, Cong L, Zhang F, Liu JS, Brown M, Liu XS (2015) Sequence determinants of improved CRISPR sgRNA design. Genome Res 25:1147–1157. https://doi.org/10.1101/gr.191452.115
Xu R, Qin R, Li H, Li J, Yang J, Wei P (2018) Enhanced genome editing in rice using single transcript unit CRISPR-LbCpf1 systems. Plant Biotechnol J 17:553–555. https://doi.org/10.1111/pbi.13028
Xu J, Kang BC, Naing AH, Bae SJ, Kim JS, Kim H, Kim CK (2019) CRISPR/Cas9-mediated editing of 1-aminocyclopropane-1-carboxylate oxidase1 enhances Petunia flower longevity. Plant Biotechnol J 18(1):287–297. https://doi.org/10.1111/pbi.13197
Xu Y, Lin Q, Li X, Wang F, Chen Z, Wang J, Li W, Fan F, Tao Y, Jiang Y, Wei X, Zhang R, Zhu QH, Bu Q, Yang J, Gao C (2020) Fine-tuning the amylose content of rice by precise base editing of the Wx gene. Plant Biotechnol 19(1):11–13. https://doi.org/10.1111/pbi.13433
Xue L, Tang B, Chen W, Luo J (2019) Prediction of CRISPR sgRNA activity using a deep convolutional neural network. J Chem Inf Model 59:615–624. https://doi.org/10.1021/acs.jcim.8b00368
Yan L, Wei S, Wu Y, Hu R, Li H, Yang W, Xie Q (2015) High-efficiency genome editing in arabidopsis using YAO promoter-driven CRISPR/Cas9 system. Mol Plant 8:1820–1823. https://doi.org/10.1016/j.molp.2015.10.004
Yan WX, Hunnewell P, Alfonse LE, Carte JM, Keston-Smith E, Sothiselvam S, Garrity AJ, Chong S, Makarova KS, Koonin EV, Cheng DR, Scott DA (2019) Functionally diverse type V CRISPR-Cas systems. Science 363(6422):88–91. https://doi.org/10.1126/science.aav7271
Yang H, Patel DJ (2019) CasX: a new and small CRISPR gene-editing protein. Cell Res 29:345–346. https://doi.org/10.1038/s41422-019-0165-4
Yang Q, Zhong X, Li Q, Lan J, Tang H, Qi P, Ma J, Wang J, Chen G, Pu Z, Li W, Lan X, Deng M, Harwood W, Li Z, Wei Y, Zheng Y, Jiang Q (2020) Mutation of the d-hordein gene by RNA-guided Cas9 targeted editing reducing the grain size and changing grain compositions in barley. Food Chem 311:125892. https://doi.org/10.1016/j.foodchem.2019.125892
Yin K, Qiu JL (2019) Genome editing for plant disease resistance: applications and perspectives. Biol Sci 374(1767):20180322. https://doi.org/10.1098/rstb.2018.0322
Yin K, Gao C, Qiu JL (2017) Progress and prospects in plant genome editing. Nat Plants 3:17107. https://doi.org/10.1038/nplants.2017.107
Yu Q, Wang B, Li N, Tang Y, Yang S, Yang T, Xu J, Guo YP, Wang Q, Asmutola P (2017) CRISPR/Cas9- induced targeted mutagenesis and gene replacement to generate long-shelf life tomato lines. Sci Rep 7:11874
Yunyan F, Jie Y, Fangquan W, Fangjun F, Wenqi L, Jun W, Yang X, Jinyan Z, Weigong Z (2019) Production of two elite glutinous rice varieties by editing Wx Gene. Ric Sci 26:118–124
Zafar SA, Zaidi SS, Gaba Y, Singla-Pareek SL, Dhankher OP, Li X, Mansoor SP (2020) A engineering abiotic stress tolerance via CRISPR/Cas-mediated genome editing. J Exp Bot 71(2):470–479. https://doi.org/10.1093/jxb/erz476
Zambre M, Terryn N, De CJ, De BS, Dillen W, Van MM, Van DSD, Angenon G (2003) Light strongly promotes gene transfer from Agrobacterium tumefaciens to plant cells. Planta 216(4):580–586. https://doi.org/10.1007/s00425-002-0914-2
Zeng Z, Han N, Liu C, Buerte B, Zhou C, Chen J, Wang M, Zhang Y, Tang Y, Zhu M, Wang J, Yang Y, Bian H (2020) Functional dissection of HGGT and HPT in barley vitamin E biosynthesis via CRISPR/Cas9-enabled genome editing. Ann Bot 126(5):929–942. https://doi.org/10.1093/aob/mcaa115
Zhai Y, Yu K, Cai S, Hu L, Amoo O, Xu L, Yang Y, Ma B, Jiao C, Zhang C (2020) Targeted mutagenesis of BnTT8 homologs controls yellow seed coat development for effective oil production in Brassica napus L. Plant Biotechnol J 18:1153–1168
Zhang F (2019) Development of CRISPR-Cas systems for genome editing and beyond. Q Rev Biophys 52. https://doi.org/10.1017/S0033583519000052
Zhang Y, Yin H, Li D, Zhu W, Li Q (2008) Functional analysis of BADH gene promoter from Suaeda liaotungensis K. Plant Cell Rep 27(3):585–592. https://doi.org/10.1007/s00299-007-0459-8
Zhang XH, Tee LY, Wang XG, Huang QS, Yang SH (2015) Off-target effects in CRISPR/Cas9-mediated genome engineering. Mol Ther Nucl Acids 4(11):264. https://doi.org/10.1038/mtna.2015.37
Zhang Y, Liang Z, Zong Y, Wang Y, Liu J, Chen K, Qiu JL, Gao C (2016) Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat Commun 7:12617
Zhang J, Zhang H, Botella JR, Zhu JK (2018a) Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of the Waxy gene in elite rice varieties. J Integr Plant Biol 60:369–375. https://doi.org/10.1111/jipb.12620
Zhang T, Zheng Q, Yi X, An H, Zhao Y, Ma S, Zhou G (2018b) Establishing RNA virus resistance in plants by harnessing CRISPR immune system. Plant Biotechnol J 16(8):1415–1423. https://doi.org/10.1111/pbi.12881
Zhang S, Li X, Lin Q, Wong KC (2019) Synergizing CRISPR/Cas9 off-target predictions for ensemble insights and practical applications. Bioinformatics 35:1108–1115. https://doi.org/10.1093/bioinformatics/bty748
Zhang F, Abudayyeh OO, Gootenberg JS (2020) A protocol for detection of COVID-19 using CRISPR diagnostics Broad Institute. Bioarchive
Zhang Y, Ren Q, Tang X, Liu S, Malzahn AA, Zhou J, Wang J, Yin D, Pan C, Yuan M, Huang L, Yang H, Zhao Y, Fang Q, Zheng X, Tian L, Cheng Y, Le Y, McCoy B, Franklin L, Selengut JD, Mount SM, Que Q, Zhang Y, Qi Y (2021) Expanding the scope of plant genome engineering with Cas12a orthologs and highly multiplexable editing systems. Nat Commun 12:1944. https://doi.org/10.1038/s41467-021-22330-w
Zhou H, Liu B, Weeks DP, Spalding MH, Yang B (2014) Large chromosomal deletions and heritable small genetic changes induced by CRISPR/Cas9 in rice. Nucleic Acids Res 42(17):10903–10914. https://doi.org/10.1093/nar/gku806
Zhou XX, Zou X, Chung HK, Gao Y, Liu Y, Qi LS, Lin MZ (2018) A single-chain photoswitchable CRISPR-Cas9 architecture for light-inducible gene editing and transcription. ACS Chem Biol 13:443–448. https://doi.org/10.1021/acschembio.7b00603
Zhu LJ (2015) Overview of guide RNA design tools for CRISPR-Cas9 genome editing technology. Front Biol (Beijing) 10:289–296. https://doi.org/10.1007/s11515-015-1366-y
Zhu LJ, Holmes BR, Aronin N, Brodsky MH (2014) CRISPRseek: a bioconductor package to identify target-specific guide RNAs for CRISPR-Cas9 genome-editing systems. PLoS One 9(9):e108424. https://doi.org/10.1371/journal.pone.0108424
Zhu H, Misel L, Graham M, Robinson ML, Liang C (2016) CT-finder: a web service for CRISPR optimal target prediction and visualization. Sci Rep 6:1–8. https://doi.org/10.1038/srep25516
Zhu Y, Klompe SE, Vlot M, van der Oost J, Staals RHJ (2018) Shooting the messenger: RNA-targetting CRISPR-Cas systems. Biosci Rep 38(3):BSR20170788
Zhu Y, Gao A, Zhan Q, Wang Y, Feng H, Liu S, Gao G, Serganov A, Gao P (2019) Diverse mechanisms of CRISPR/Cas9 inhibition by type IIC anti-CRISPR proteins. Mol Cell 74:296–309
Zsögön A, Čermák T, Naves E et al (2018) De novo domestication of wild tomato using genome editing. Nat Biotechnol 36:1211–1121
Acknowledgements
The authors thank the ICAR scheme-National Agricultural Science Fund (NASF/GTR-7025/2018-19) for financially supporting this research. We also thank Department of Biotechnology (DBT) Funds (BT/PR38339/GET/119/334/2020) for additional support towards the completion of this work.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Kaul, T. et al. (2022). CRISPR Genome Editing Brings Global Food Security into the First Lane: Enhancing Nutrition and Stress Resilience in Crops. In: Gowdra Mallikarjuna, M., Nayaka, S.C., Kaul, T. (eds) Next-Generation Plant Breeding Approaches for Stress Resilience in Cereal Crops. Springer, Singapore. https://doi.org/10.1007/978-981-19-1445-4_9
Download citation
DOI: https://doi.org/10.1007/978-981-19-1445-4_9
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-1444-7
Online ISBN: 978-981-19-1445-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)