Abstract
Cancer is one of the leading causes of death worldwide. Recent report from the World Health Organization suggested that, globally, one in six deaths is owing to cancer. In 2018, it was accountable for nearly 9.6 million deaths, and it is expected to be 14.6 million by the year 2035. The worldwide burden of cancer increase is due to aging and growth of population. In addition, cancer-associated lifestyle choices like smoking, sedentary habits and westernized diets increases the risk. Metastasis is complex and multistep process that results in the spread of cancerous cells from the primary site of the tumor to the surrounding tissues and to distant organs. Metastatic cancer is the primary cause of cancer morbidity and mortality. Several studies suggest that tumor has heterogeneous cell population and have numerically less cancer stem cell (CSC) population with self-renewal characteristics. CSCs are shown to drive tumor initiation, progression, metastasis, recurrence, and resistance. In addition, acquisition of epithelial-mesenchymal transition, expression of aberrant RNA-binding proteins, dysregulated microRNA expression, and increase in intercellular transfer of molecules via exosome cargo have been correlated with tumor progression, invasion, metastasis, poor survival, and an increased risk of cancer recurrence. Given the tumor initiating capacity, resistance, migratory potential and invasiveness, CSCs are the seeds of metastasis. This review article attempts to provide the details of the critical importance of CSCs on metastatic process and to offer a basis for the investigation of novel targets to curtail this deadly disease.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
12.1 Introduction
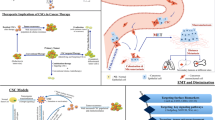
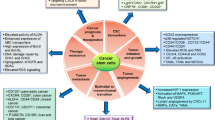
Cancer is one of the leading causes of death worldwide [1]. Recent report from the World Health Organization suggested that, globally, one in six deaths is owing to cancer. In 2015, it was accountable for nearly 8.8 million deaths, and it is expected to be 14.6 million by the year 2035. The worldwide burden of cancer increase is due to aging and growth of population. In addition, cancer-associated lifestyle choices like smoking, sedentary habits, and westernized diets increases the risk. Cancer is a complex disease with various cellular alterations that will result in self-sufficiency in growth signal leading to abnormal cell growth, evading apoptotic and growth suppressor signals, and increased angiogenesis, wherein, network of blood vessels develop and penetrates into the tumor to supply nutrients and oxygen for the cancerous cells. Some of the tumor cells invade surrounding tissues and distant organs through the blood circulation or lymph vessels. This spread of cancer cells from primary tumor to other sites is termed metastasis, which is shown to be responsible for more than 90% of cancer-related death. Cancer stem cells (CSCs) are cells within a tumor that exclusively have self-renewal capacity and can give rise to all cancer cell lineages within a tumor and are exclusively tumorigenic in vivo. They undergo asymmetric/symmetric cell division, can maintain and expand themselves, and also have a distinct profile of surface marker expression that has been linked to poor prognosis [2]. Intriguingly, it has been shown that CSCs drive tumor initiation, progression, metastasis, recurrence, and resistance. Identification of CSC-specific surface markers has provided opportunity to characterize CSCs and their role in tumor progression and metastasis. In addition, acquisition of epithelial-mesenchymal transition (EMT) features, expression of aberrant RNA-binding proteins, dysregulated microRNA expression, and increase in intercellular transfer of molecules via exosome cargo have been correlated with tumor progression, invasion, metastasis, poor survival, and an increased risk of cancer recurrence. Given the tumor initiating capacity, resistance, migratory potential and invasiveness, CSCs are the seeds of metastasis [3]. This review article attempts to provide the details of advances in the role of CSCs on metastatic process that will aid in better understanding of the involvement of cancer stem cells (CSCs) in the metastatic processes and to offer a basis for the investigation of novel targets to curtail this deadly disease (Fig. 12.1).
12.2 Epithelial-Mesenchymal Transition in Regulation of CSCs and Metastasis
Epithelial-mesenchymal transition (EMT) is a process where the cancer epithelial cells lose many of their epithelial characteristics and acquire various mesenchymal cell characteristics such as cell morphology, cytoskeletal organization, and cell junctions that will enable cell invasion and migration. Cancer cells have been observed in the circulation, whether as single or in clusters; these cells display signs of at least partial epithelial-mesenchymal transition [4]. Earlier reports have provided evidence that in the tumor, only CSC-enriched subpopulation exhibits aspects of EMT-related gene activation [5, 6]. In addition, induction of EMT-related gene expression in epithelial tumor cells has increased their capacity for tumor progression and metastasis [7]. Eliminating CSCs alone will not be sufficient to prevent tumor recurrence as the non-CSCs can undergo EMT and dedifferentiate into CSCs [8]. Therefore, for effective cancer therapeutic strategy, both CSCs and non-CSCs should be simultaneously targeted. We have listed the most important CSC markers for various cancer subtypes (Table 12.1).
The traits of EMT are the loss of epithelial cell surface marker, E-cadherin, and the gain of mesenchymal traits [28]. The initiating factors are seen to be mostly because of networks of transcriptional, translational, posttranscriptional, and posttranslational modifications seen in the cells [29]. The ALDH+ cells strongly displayed stem cell-like properties plus higher invasiveness, EMT, and antiapoptotic phenotypes [30]. In the case of human breast cancer cells, it was observed that a small population of cells, which exhibited EMT, also displayed stem cell-like phenotypes. Fascinatingly studies performed in transgenic cancer models in combination with S100A4 lineage tracing have stated that EMT of the breast cancer cells is not responsible for their metastasis to the lungs, but they had a significant role in promoting chemoresistance. The lowering the levels of E-cadherin in the mouse models showed that inhibition of the epithelial traits may promote migration but does not result in metastasis [31]. CD44 is a popular cell surface glycoprotein, which is strongly associated to the stemness of the cancer and its aggressiveness. In the case of ovarian cancer cells, the overexpression of CD44 resulted in population of cells with mesenchymal-like phenotypes (CD44S) and decreased the number of epithelial-like cells. The downregulation of ESRP1 and upregulation of TGFPs1 promoted EMT, invasiveness, and the gain of stem cell-like phenotypes and chemoresistance in CD44 cells [32].
In the colorectal cancer cells, abnormal expression of miR-26b induced EMT and stem cell-like characteristics. Lymphatic metastasis shows significantly upregulated levels of miR-26b. miR-26b directly targets many tumor suppressors along with phosphatase and tensin homolog (PTEN) and wingless-type MMTV integration site family member 5A (WWT5A) [33].
The role of IncRNAs was evaluated in two sets of cells: colorectal cancer with liver metastasis and colorectal cancer without liver metastasis. The expression levels of UICLM (upregulated in colorectal cancer liver metastasis) IncRNA was upregulated in the CRC with liver metastasis, and the knockdown of UICLM prevented cell proliferation, invasion, epithelial-mesenchymal transition, and CRC stem cell formation. Further experiments found that IncRNA UICLM regulated ZEB2 [34, 35].
MYC (c-Myc) is regarded as a very strong proto-oncogene observed to be highly expressed in many cancers. The (PARPI)-poly (adenosine diphosphate (ADP)) ribose polymerase inhibitor effect on triple-negative breast cancer can be chemically improved upon a blockade of MYC. Dinaciclib a cyclin-dependent kinase inhibitor downregulates Myc expression; this, administered along with PARPI-niraparib, downregulated EMT by reducing homologous recombination which resulted in reduced cancer stem cell-like phenotype. Also dinaciclib re-sensitized TBNC cells which displayed resistance toward niraparib. This combination of therapy also worked on ovarian, prostrate, pancreatic, lung, and colon cancer cells [36].
Claudin-6 (CLDN6) is a tight junction protein functioning as a tumor suppressor and also a stem cell marker. Triple-negative breast cancer (TNBC) cells show low levels of CLDN6. A study involving the overexpression of CLDN6 in TNBC cells (MDAMB231 cells) showed increase in epithelial marker E-cadherin and reduction in vimentin (mesenchymal marker); stem cell markers such as OCT4, SOX2, and Nanog were upregulated [37].
Another long noncoding RNA called the nuclear-enriched abundant transcripts (NEATs) plays a significant role in Non-small-cell lung carcinoma (NSCLC) stem cells. Experimental results suggest that NEAT1 was overexpressed and copper transporter 1 (CTR1) was downregulated in the NSCLC stem cells. NEAT1 knockdown reduced the cancer stem cell-like phenotype in these cells. NEAT1-expressing cells also exhibited Wnt pathways and EMT process [38].
In triple-negative breast cancer cell line SUMI159pr, low expression of miR-105 was recorded. Its targets were identified to be VEGFA, Erb33, Zab1, Fyn, and Lyn A/B, thus reducing cell proliferation and c-Myc with upregulated levels of participants.
Overexpression of MiR205 inhibited the anchorage-independent growth, migratory and invasive nature of SUM159PT cell line with activated src kinases, and low levels of MMPs. The pathways and proteins associated with EMT like CD44, TAZ, E2AE12, twist, Snail A, and CK5 were also highly reduced with the expression of miR-205. The miR-205 also plays a critical role, and its co-expression along with anti-miR-205 reverted back all the reduced and the inhibited pathways of the triple-negative breast cancer cell line SUM159PT [39]. Ursodeoxycholic acid is an epimer of chenodeoxycholic acid found in the mammalian bile secretions, commonly abbreviated as UDCA. Reactive oxygen species (ROS) plays a critical role in cancer progression and advancement, and UDCA inhibited intracellular ROS. Pancreatic cancer cell lines treated with 0.2 mM UDCA showed elevated levels of E-cadherin and lower levels of N-cadherin and downregulation of sex-determining region Y-box 2 (SOX2). It reduced the sphere-forming abilities; thus it is evident as an effector inhibitor of cancer stem cell-like and EMT phenotypes [40].
Carnosol (CAR) is naturally found in our body that inhibits the MDM2/p53 complex. Its effects on U87MG, a glioblastoma-derived cancer stem cell line, showed that it reduced the CSC formation and promoted apoptosis of the cancer stem cells by functionally reactivating P53. Furthermore it also controlled the effects of TNF-alpha/TGF-beta and inhibited the effects of cytokines associated with EMT-regulating genes (slug, Snail, twist, ZEB1). It also promoted the activation of miR-200c, which is associated with EMT; adding on this it also increased the antiproliferative effects of temozolomide (TMZ) [41]. Lagunas et al. demonstrated that telomere DNA damage signaling regulates cancer stem cell evolution and metastasis. Telomeres are protected by the double-stranded DNA-binding protein TRF2 and maintained by telomerase or a recombination-based mechanism known as alternative lengthening of telomeres (ALT). Loss of TRF2 and Terc expression gives telomere DNA damage, severely decreases CD34+ and Lgr6+ cancer stem cells, and induces terminal differentiation of metastatic cancer cells [42]. The natural sphingolipid phytosphingosine (PHS) suppresses the stem cell-like phenotype and EMT-associated proteins and the highly malignant basal-type breast cancer cells (CD44+/CD24-) by downregulating EGFR/JAK1/STAT3 [43]. Slug and twist are important transcriptional factors that are highly associated with EMT; they are found to be regulated by two processes, namely, ubiquitination and degradation. Slug and twist are found to be in very stable conditions inside the cancer cells. It can be speculated that the stabilization of the slug and twist is because of the loss of ubiquitin by deubiquitinase (DUB). DUB3 was identified to be the deubiquitinase for both slug and twist. The upregulation of DUB3 amplified the expression levels of slug and twist in a dosage-dependent manner, also protecting the two genes from being degraded. IL-6, which plays a significant role in the metastasis of breast cancer cells, seemed to induce the expression of DUB3. Thus DUB3 is identified to play a critical role in stem cell-like phenotype, metastasis, and invasive and migratory traits in breast cancer cells [44].
Fusobacterium nucleatum has been identified to play a role in colorectal cancer. A study was conducted on stage 3 CRC patients. The Fusobacterium nucleatum levels were significantly high and were associated with the invasion, lymph node and metastasis and distant metastasis. Analysis showed the presence of Nanog, OCT4, and SOX2 (stem cell markers) and N-cadherin levels [45]. SOX8 was overexpressed in tongue squamous cell carcinoma (TSCC) resistant to cisplatin, which exhibited EMT and CSC-like (Wnt) phenotypes. It is found to be upgraded in chemoresistant patients affected by tongue squamous cell carcinoma (TSCC) and also correlated with normal lymph node metastasis [46].
12.3 Role of MicroRNAs on CSCs and Metastasis
The microRNAs are short noncoding segments of RNA with 21–25 nucleotides seen in plants, animals, and certain viruses. Their function is “RNA-mediated gene silencing” at the posttranscriptional stage by attacking the 3′-untranslated regions of the “target gene,” thereby degrading the specific mRNA [47]. miRNAs play a vital role in the human cancer progression and metastasis. The expression levels of the oncogenic miRNAs can be observed to be increased as cancer progresses. The improper regulation of the expression of the miRNAs influences the processes of the progression like antiapoptotic activity, drug resistance, tissue invasion, and metastasis [48] (Table 12.2).
12.3.1 miR-200 Family
miR-200a, miR-200b, miR-141, and miR-429 are the members of the miR family. Their regulations have a strong association with the cancer stem-like features and metastasis via EMT [63]. Experiments on human mammary epithelial cells show that these cells were able to transit from non-stem like to stem like upon the loss of miR family [64]. A strong connection between the levels of miR-200s and E-cadherin in the cancer cell lines and clinical samples showed that miR-200s maintained tumor epithelial traits and prevented EMT. This was achieved by the direct interaction of miR-200s with ZEB1 and ZEB2 transcription factors. Suppression of metastasis has been observed upon the upregulation of miR-200s, but the miRNAs are also known to promote metastasis by recent studies as higher concentration of miR-200s promoted the development of metastasis in breast cancer patients and promoted invasion of the lung by murine breast cancer cells. The direct downregulation of Sec23a resulted in loss of expression of metastasis-suppressive proteins IGFBP4 and Tinagl1 in the murine breast cancer cells [63].
Tumor suppressor p53 is shown to be associated with both EMT and breast CSCs associated with EMT by transcriptionally activating the miRNAs associated with stemness including miR-200C. The loss of P53 in mammary epithelial cells downregulated miR-200C expression and initiates EMT and increases CSCs. It also directly interacts with EMT and increases CSCs. It also directly interacts with the protein line ZEB1, ZEB2, and BMI1 [65].
12.3.2 miR-203
miR-203 inhibits the colony formation, migration, and invasion of many cancer cells. Enhanced regulation of Snail and downregulation of miR-203 in CD44+ human colorectal carcinoma cell lines showed higher metastasis; further miR-203 is suppressed by Snail [66].
miR-203 reduced the sphere-forming ability of the nearby cells by indirectly prompting DKK 1 (inhibitor of Wnt signaling) [67]. The effects of miR-203 are observed in CD44+/CD88− leukemia cancer stem cells by directly interacting with BMI1/survivin [68].
12.3.3 miR-34a
miR-34a has been called as a “star” miRNA in cancer research, acts as tumor suppressor, and is downregulated in many human cancers, and also studies have shown that the aberrant miR-34a expression has been linked to chemotherapy resistance in a variety of cancers [69].
miR-34a is a mediator of the p53 transcriptional network and has been identified as a tumor suppressor that contributes to the inhibition of the invasion and metastasis in various types of epithelial cancers [69]. miR-34a expression is significantly downregulated in primary tumors from head and neck cancer patients as well as in head and neck cancer cell lines. Ectopic expression of miR-34a in head and neck cell lines significantly inhibited tumor cell proliferation, migration, and colony formation by downregulating the expression of E2F3 and survivin [70]. Expression of miR-34a in bulk can inhibit prostate cancer cells (CD44+) through inhibition of clonogenic expansion, tumor regeneration, and metastasis, and expression of miR-34a antagomirs in CD44− prostate cancer cells promoted tumor development and metastasis [71], and miR-34a performs a key role in suppressing colorectal cancer metastasis by targeting and regulating Notch signaling [72].
12.3.4 miR-22
miR-22 epigenetically promoted stem-like traits and metastasis in breast cancer cells. Studies have shown miR-22 capable to directly inhibit TET expression and EMT induction in breast cancer. Ten eleven translocation (TET) enzyme has been linked to the demethylation of miRNA-200 promoter region. miR-200 is an anti-metastatic microRNA that inhibits stemness and EMT, and miR-22 is observed to be in association with TET family, thus promoting CSC-like properties and metastasis by repressing miR-200 family [48].
12.3.5 miR-17
Significant overexpression of miR-17 was seen in CD133+ cells of glioblastoma cell lines. The miR-17 is said to directly target calmodulin-binding transcriptional activator (CAMTA1), which is a transcription factor of antiproliferation cardiac hormone natriuretic peptide A. Downregulation of miR-17 in these cells reduced neurosphere formation and promotes cell differentiation. This shows that miR-17 is significantly correlated to stem (CD133+)-like traits in cells [48]. In osteosarcoma the levels of miR-17 were seen to be higher, and its inhibition resulted in reduced or suppressed cancer cell proliferation, migration effects, and invasion/metastasis. PTEN homolog was identified to be directly targeted by miR-17 [73]. PTEN levels are critical in maintaining stemness, and its suppression leads to promotion of cancer stem cells [74]. However, A549/DDP (cisplatin resistance) non-small cell lung cancer cells showed downregulated levels of miR-17, miR-20a, and miR-20b. The downregulated levels suppressed the TGF-beta signaling pathways and inhibited EMT pathways, thus affecting metastasis [75]. Cancer stem cells seem to have developed their stem-like properties via stem cell pathways like Wnt, TGF-beta, STAT, and Hippo-YAP/TAZ [76]. Colon cancer cells that overexpressed phosphatase of regenerating liver 3 (PRL-3) inducing the expression of miR-21, miR-17, and miR-19 by activating STAT3 [77]. Cancer stem cells are strongly associated with stemness-related STAT pathway [76]. Thus these miRNAs have increased the proliferation of primary colon cancer cells and the metastatic growth [77]. In ovarian cancer metastasis, the expression of miR-17 is inversely related to the levels of ITGA5 and ITGB1. Lower level of ITGA5 and ITGB1 suppressed peritoneal metastasis. The abnormal expression of miR-17 in ovarian cancer resulted in lowered expression of ILK phosphorylation and MMP-2. Thus higher levels of miR-17 suppress ovarian cancer cell peritoneal metastasis [78].
12.3.6 miR-124
SNAI2 has been found to be upregulated in glioblastoma cells, and miR-124 has SNAI2 as its functional target. SNAI2 has also been associated with stemness. Experimental evidences showed that upregulation of miR-124 and SNAI2 knockdown reduced neurosphere formation, and the expression of stem cell markers like BMI1, Nanog, and Nestin was substantially reduced, and the effects can be reverted by the re-expression of SNAI2 in in vivo [79]. miR-124 was also seen to directly target STAT3 signaling. STAT3 is identified to have positive effects on T-cell-mediated suppression in tumor microenvironment. miR-124 is observed to be lost in all grades and in all pathological types of gliomas. The upregulation of miR-124 in glioma cancer stem cells (GCSCs) resulted in the inhibition of STAT3, and it reversed the GCSC-associated immunosuppression of T cells and the induction of FOXP3 and regulatory T cells (Tregs). T cells from immunosuppressed glioblastoma patients when treated with miR-124 resulted in upregulation of interleukin (IL-2), IFN-gamma, and TNF-alpha [80]. The abnormal expression of miR-124 in MDA-MB-231 cells (known for high invasiveness) suppressed spindle formation, invasive capacity, and adhesion to fibronectin and anoikis. These results show that miR-124 plays a critical role in the multistep process of metastasis in breast cancer cells [81].
12.3.7 miR-128
Patients with advanced glioma show downregulation of miR-128. miR-128 targets BMI1 [48]. miR-128 is reported to target VEGF-C and reduce the proliferation and the invasive properties of bladder cancer cells. The knockdown of miR-128 upregulates VEGF-C and induces proliferation, migration, and invasion of bladder cancer (BC) [82]. The metastatic and the stem cell-like properties of the hepatocellular carcinoma were inhibited by the upregulation of miR-128, and they were identified to target ITGA2 and ITGA5 [83]. The chemosensitivity is increased and invasive properties of prostate cancer cells were inhibited following upregulation of miR-128. Experimental results suggest that miR-128 directly targets zinc finger E-box-binding homeobox 1 (ZEB1) in prostate cancer cells and induces the sensitivity toward cisplatin and inhibits invasion [84].
12.3.8 miR-199b-5p
Studies show that miR-199b-5p is downregulated in medulloblastoma which results in its invasive properties. This happens by targeting HES1 transcription factor in Notch signaling pathways, thus inhibiting the self-renewal properties of Glioma Stem Cells (GSCs) by targeting the CD133+ cells [48]. But its expression plays an opposite role in the case of human osteosarcoma. The elevated levels of miR-199b-5p correlate to cell proliferation, invasion, and migration of these cells. Its expression levels are seen amplified in the higher grades of osteosarcoma [85]. In breast cancer cells, miR-199b-5p suppresses HER2 expression by negatively conferring with ERK1/2 and AKT pathways. This shows loss of migration, wound healing, and colony formation. This has also improved the sensitivity of HER2 cells towards trastuzumab, thus hampering cells’ migratory and clonogenicity properties [86]. miR-199b-5b targets N-cadherin and promotes cell aggregation and suppresses migration/invasion of hepatocellular carcinoma (HCC). This inhibits metastasis of tumor xenografts. It was also shown to reduce the effects of TGF-beta-induced AKT phosphorylation which results in EMT features [87].
12.3.9 miR-451
The lower expression of miR-451 shows enhanced levels of cyclooxygenase-2 (COX2) and macrophage migration inhibitory factor (MIF); this results in acquisition of stem cell-like properties. COX2 and MIF have been shown to be associated with Wnt pathway, which is a major regulator of cancer stem cells [48].
Papillary thyroid carcinoma with lymph node (PTCLN) metastasis shows amplified expression levels of four miRNAs: miR-2861, miR-451, miR-193b, and miR-1202. When compared with PTC without lymphoid metastasis, it was found that PTCLN had high levels of miR-2861 and miR-451 especially in lateral and lymph node (LLN) [88]. Zhang et al. reported that miR-144/451 re-expression markedly suppressed the migration and invasion of breast cancer and HNSCC cells through ADAM10 and ADAMTS5 modulation. PAX4 promoted migration and invasion in human epithelial cancers by decreasing miR-144 and miR-451 (miR-144/451) expression levels.
Paired box gene 4 (PAX4) has been promoting metastasis in human epithelial cancer by downregulating miR-144 and miR-451, while miR-144/451 has been observed to inhibit cancer migration even in PAX4-expressing cells by targeting a disintegrin and metalloproteinase (ADAM) protein family members in ADAMT55 and ADAM10 [89]. MiR-451 has also been shown to suppress cell proliferation and metastasis by targeting chemokine ligand 16 (CXCL16) in osteosarcoma patients [90]; it has been shown to promote significant metastasis in various cancer types like hepatocellular carcinoma by targeting c-Myc [91]. In neuroblastoma miR-451 has been shown to target macrophage migratory inhibitory factors [92], in A549 lung cancer cells miR-451 inhibits metastasis by targeting PSMB8 and NOS2 thereby reducing the expression of MMP-2, MMP-9, VEGF. miR-451 has also been associated with stemness and CXCR4 [93]. This miR-451 plays a vital role in inhibition of stem cell-like features by inhibiting stem cells and metastasis through numerous pathways.
12.3.10 miR-320
The miR-320 directly targets Wnt/beta-catenin expression in prostate cancer stem cells. Thus the expression of CD44+ PCa cell expressing Wnt is inversely proportional to miR-320 level [48]. Fatty acid synthase (FAS) was previously reported to be correlated with various clinicopathological features of cancer. Overexpression of FAS in NSCLC has been shown to be significantly associated with bone metastasis. Thus miR-320 contributes to cell proliferation, migration, and invasion by directly targeting FAS in NSCLC, and overexpression of miR-320 in NSCLC cell lines inhibits cell proliferation, migration, and invasion via downregulation of FAS. miR-320 may act as a tumor suppressor by inhibiting the oncogenic activity of FAS [94]. In addition, miR-320 inhibited migration by targeting FOXM1 in cervical cancer cells [95].
12.4 Functions of RBPs on CSCs and Metastasis
RNA-binding proteins (RBPs) act as epigenetic regulators of various RNA processing events, such as splicing, localization, stabilization, and translation, and can regulate various types of stem cells. Many RNA-binding proteins are overexpressed in cancers [96, 97]). Deregulation of RBPs affects every step of cancer development, such as sustained cell proliferation, inhibition of the apoptosis process, avoiding immunosurveillance, inducing angiogenesis, and activating metastasis. Some RBP proteins recognize cis-acting elements to translationally regulate proto-oncogenes, cytokines, and growth factors [98]. RNA-binding proteins that are abnormally expressed in cancers are the IMP-3, the CRD-BP/IMP-1, the p62, as well as members of the ELAV/Hu protein family, e.g., HuR. Upon binding to the AU-rich instability element (ARE) in the 3′Untranslated region (UTR) of rapidly degraded mRNAs of proto-oncogenes, cytokines, and growth factors, HuR regulates nucleocytoplasmic transport, stability, and translation. However, only very few of these RNA-binding proteins have been demonstrated to regulate tumor progression and metastasis and control the cancer stem cell self-renewal [97].
12.4.1 RBM3
Colorectal cancer mostly demonstrates the overexpression of Wnt/beta-catenin in the colon cancer stem cells. The RNA-binding protein RBM3 promotes cancer cell proliferation, angiogenesis, and resistance against apoptosis and even induces metastasis at higher levels by acting as a proto-oncogene [99]. Two cell lines HCT116 and DLD1 were taken to study the effects of RBM3 on colon cancer; upon the upregulation of RBM3, the number of sphere-forming cells increased in the HCT116 cells along with the increased expression of cancer stem cell marker DCLK1 in DLD1 cells. Further analysis have shown that RBM3 upregulated the levels of Wnt/beta-catenin but suppressed the expression of Notch [100].
12.4.2 PTBP3
The RNA polypyrimidine tract-binding protein (PTBP3) at upregulated state leads to the acquisition of cancer stem cell-like and EMT phenotype in breast cancer cells, thus enhancing metastasis. Mechanistically, the EMT regulatory transcription factor ZEB1 is upregulated due to PTBP3 binding to its mRNA at the 3′UTR [101].
12.4.3 Lin 28
Recent studies have shown that LIN 28A/B plays an important role in the formation of CSCs and is involved in tumor aggressiveness and metastasis. Higher expression rates of RNA-binding protein LIN 28 is correlated with the exhibition of malignant, cancer stem-like phenotypes in breast, colorectal, and esophageal cancer cells. Ovarian cancer cells with CSC-like trait express both LIN 28A and OCT4. LIN 28B is expressed in the colorectal stem cells with CSC markers like LGR5, KIT, and PROM1 (CD133) in colon cancer. LIN 28B might be correlated with intestinal CSCs, and LIN 28B regulated by IKKβ are able to maintain stemness by interacting with Wnt pathways as LIN 28 is expressed only by CSCs. A crosstalk between LIN 28 and let-7 is observed in CSCs; thus blocking this axis might be a solution to target the CSCs. In non-small cell lung cancer patients, higher levels of LIN 28 and lower levels of let-7 are associated with chemo- and radiotherapy resistance [102].
12.4.4 MUSASHI-1
MUSASHI-1 (MSI-1) is a stem cell marker in both normal and cancerous stem cells; in malignant colorectal cancer cells, MUSASHI1 is upregulated by the expression of Notch-3 [103]. In another experiment on gastric cancer cells, it was identified that MUSASHI1 played a significant role in the prognosis of the metastatic gastric cancer. The expression of MSI-1 can be associated with tumor node metastasis (TNM), Lauren’s classification, depth of invasion, vessel invasion, lymph node metastasis, and distant metastasis. The prognosis at each stages of TNM is worse than its previous one as there is an increase in MSI-1 expression [104].
12.4.5 MUSASHI-2
Elevated levels of MSI-2 is observed in metastatic non-small cell lung cancer cell lines; the suppression of MSI-2 led to decrease in metastatic potential and promoted the expression of claudin 3 (CLDN3), claudin 5 (CLDN5), and claudin 7 (CLDN7) and downregulated the expression of TGFβR1, SMAD3, and zinc finger proteins SNAI1 and SNAI2 (Slug) [105].
12.5 Effect of CSCs on Immunosurveillance and Metastasis
A perfect display of chemoresistance and immune resistance is seen during the escape phase when detectable tumor bodies of CSCs/TICs reappear after a long dormant phase. They have been reported to produce immunosuppressive molecules and recruit cells that suppress the immune system like Treg cells more tolerant toward the immune system via loss of expression of tumor antigen, loss of processing and presentation machinery, and downregulation of MHC1 and MHC2. All these factors along with aging related immune deficiency is utilized by CSCs to promote cancer. CSCs/TICs are able to escape the immune response because they are equipped with co-inhibiting molecules like cytotoxic T-lymphocyte antigen (CTLA4), B7-H2, B7-H3, and programmed death receptor 1 (PD-1). The lack of MHC2 leads to downregulation of low molecular weight proteins (LMP), transporter associated with antigen processing (TAP), and beta-macroglobulin, which assists immune escape. Cytokines like transforming growth factor-beta (TGF-beta), IL-10, and IL-13 were secreted in in vitro conditions. The CSCs/TICs from glioblastoma required expression of VEGF, macrophage chemoattractant protein-1 (MCP-1), macrophage inhibitory factor (MIF), and growth-related oncogene (GRO2) for their survival. Breast cancer stem cells and glioblastoma CSCs produce more TGFs than usual cancer cells; IL-4 produced by colon CSCs promotes drug resistance and stops antitumor immune responses. CSCs also produce CD200 that aids immune escape. The dissemination of CSCs of the melanoma is assisted by the expression of ATP-binding cassette subfamily member 5 (ABCB5) and shows low levels of lineage-related and Cancet-testis (CT) antigens. But the CSCs expressing CD133+ have upregulated levels of NY-ESO1 cancer testis antigen; and they produce specific T-cell response.
TAADDX3X expressed by CD133+ CSCs is susceptible to T cells in massive models, while the CD271+ CSCs do not express both lineage-related and CT antigens; hence their susceptibility toward T cells is not possible. Thus all these traits help the CSCs/TICs during tumor progression and metastasis.
Cancer stem cells of the lungs have developed ways to protect themselves from T cells and induce apoptosis in them. T cells express CTLA4/CD152 after exposure to antigen. The binding of CTLA4 to its ligand (CD80/CD86) on the antigen presenting cells (APC) induces T-cell apoptosis. Specifically lung cancer cells induce T cells to produce more CTLA4. Similarly PD4 is produced by the T cells, B cells, and some myeloid cells. Cytotoxic T cells with PD4 interacting with its ligand are marked for apoptosis. Upregulation of ALDH and B-cell lymphoma-2 (BCL-2) protein and its family seems to enhance the chemoresistant phenotype [106].
Cancer stem cells derived from histopathologically negative prostrate training lymph nodes (PDLN) in mice with prostrate intraepithelial neoplasia (mPIN) controlled by oncogene was similar to the CSCs from mPIN tumor bodies. CSCs from both PDLN and mPIN produced extracellular matrix protein tenascin-C (TNC) and CXCR4. TNC interacts with α5β1 integrin on the cell surface of the T cell and inhibits T-cells receptor dependent T cell activation proliferation and cytokine production [107].
Macrophages play a vital role during the stages of metastasis of the cancer cells. Macrophages are made tumor friendly by various cytokines and chemokines like colony-stimulating factor 1 (CSF1), vascular endothelial growth factor A (VEGF-A), semaphorin3A (SEMA3A), CC-chemokine ligand 2 (CCL2), and CXC-chemokine ligand 12. Tumor-associated macrophages (TAM) are known for their CD8+-suppressing properties by directly producing DDL1 and BT-H4 or indirectly Treg cells [108].
The vascular endothelial growth factor A (VEGF-A) is a vital oncogenic factor that also plays important role in cancer stem cell maintenance, proliferation, malignancy, immunosuppression and also EMT. Myc and SOX2 were upregulated in the breast and lung cancer stem cells through VEGF receptor 2 (VEGFR2)/STAT3 [109]. VEGF has been identified to target dendritic cells by disrupting its maturation from the progenitor cells. It also plays a role in the T cells and macrophages. The functional maturation of the DCs from its progenitor CD34+ was identified to have disturbed the VEGF produced by breast and colon cancer cells. The M2-polarized macrophages that are tumor friendly express VEGF, hence promoting angiogenesis. Initial experiments on mice with elevated levels of VEGF equivalent to the amount found in advanced cancer patients showed that the number of CD8/CD4 thymocytes was reduced. The VEGF affected the progenitor of the T cells rather than on the T cells themselves [110]. The human cancer cell A549 and breast cancer cells MDA-MB-231 recruit many tumor-associated dendritic cells (TADCs) overexpressing CCL2. CCL2 increases the stem cell-like features, migratory/invasive properties, malignancy, and also the immunosuppressive tumor-associated macrophages. The CCL2 amplifies the phosphorylation of STAT3 in the cells. 6-Shogaol can inhibit CCL2 and suppress the proliferative and the metastatic properties of the lung and the breast cancer cell lines [111]. In case of the hepatocellular carcinoma, increased expression of CXCR4 corresponds to lymph node metastasis [112], but the cytoplasmic expression of CXCR4 does not seem to play a role in the lymph node metastasis [113].
A cross talk between the CXCR4 pathway and TGF-beta pathways has been reported in the Hepatocellular carcinoma (HCC); CXCL12/CXCR4 has also been identified to promote the expression matrix metalloproteinase 10 (MMP10) that enhances the migration and metastatic properties of the HCC. A cross talk between CXCR4 and Sonic hedgehog (SHH) pathways has been reported in the human pancreatic cancer and medulloblastoma as well. SHH/CXCR4 interaction has been associated with promotion of stem cell properties and malignancy in these cells.
A cross-link between alpha-fetoprotein (AFP) and CXCR4 has been observed. AFP promotes migration by activating AKT/mTOR signaling through CXCR4 in HCC [114].
The regulatory T cells are utilized by the breast cancer cells with lung metastasis by expressing CCL22/CCL5, which is very immunosuppressive expressing alpha-chain/CD25 and forkhead boxp3 (FOXP3). Suppressing of metastasis in the breast cancer models has shown to deplete number of CD4 + CD25+ cells; overexpression of prostaglandin E2 by the breast cancer cells also recruits Treg cells there by inducing CD8+ T-cell apoptosis and enhances cancer cell bone metastasis. Melanoma utilized Treg cell to overexpress TNR to promote lung metastasis [108]. The galectin-1 produced by the breast cancer cells suppresses the immune system by regulating clonal expansion and via linker for activation of T cells resulting in the promotion of breast cancer metastasis. Galectin-1 has been reported to be highly expressed in CD133+ cells (stem cell marker) [115, 116]. Treg cells seem to target NK cells and induce apoptosis by expressing BETA-galoctoside-binary protein (BETA-GBP) in lung metastasis that Treg cells suppressed the cytotoxicity of the NK cells via cell to cell contact and expressing TGF-beta [108]. Neutrophils have an interesting role in cancer promotion. The human fibrosarcoma and prostate cancer cells associated neutrophil promote angiogenesis by secreting MMP9. In intrahepatic cholangiocarcinoma xenograft model, CXCL15 recruits neutrophils and enhances lung metastasis. But other studies show that reducing the neutrophil contact increases the lung metastatic loci in breast cancer models. The neutrophils extracted from tumor-bearing mice can kill cancer cells in other mice models by producing Hydrogen peroxide (H2O2). However TGF-beta has been reported to reverse the antitumor properties to tumor-promoting traits of neutrophils [117].
12.6 Overview of CSC-Derived Exosomes on Metastasis
There are many evidences proving that exosomes from cancer cells cause organ-specific metastasis. Their specificity can be identified by the presence of certain ECM, membrane proteins, lipids, and adhesion molecules present within the exosomes. Tumor-derived exosomes (TDEs) initiate metastasis by three ways: firstly, autocrine and paracrine signaling which initiates EMT formation. Secondly, they help in the formation of pre-metastatic niche. Thirdly, they modulate the body’s immune system promoting metastasis.
Exosomes from various cancer cells have Notch-1 (MMPs), miR-100, HIFα, casein kinase Iiα, and annexin A2. Hypoxia is a popular condition associated with the development of metastasis particularly; EMT-inducing molecules like TGFβ, MMPs, TNF-α, IL6, AKT, ILK1, caveolin 1, PDGFs, and β-catenin are the contents of exosomes under hypoxic conditions. Nasopharyngeal carcinoma (NPC) (CNE2) cell line co-cultured with exosomes expressed more of N-cadherin and vimentin and downregulated expression of E-cadherin [118]. Primary tumors targeting the lungs express integrins like α6β4 and α6β1. The integrin αvβ5 promotes metastasis to the lungs [119]. In another experiment where exosomes from fluorescently labeled B16-FI0 melanoma cells were injected into a mice, the exosomes combined together with the regional lymph node nearest to the point of injection. Tumor and organ-specific metastasis is a characteristic behavior of cancer stem cells [120]. CSCs secrete and uptake exosomes; hence their metastatic phenotype can also be determined by studying the contents of their exosomes. The CD105+ exosomes from the renal cancer stem cells when injected into SCID mice developed only lung metastasis [118]. Breast cancer stem cells recruit Treg cells and promote lung metastasis [108]. Tumor-derived exosomes (TDEs) use Treg cells and induce CD8+ T-cell apoptosis and suppress NK cells. The exosomes from NPC recruit Treg cells and confer with T helper cell (Th1) and Th17 differentiation; also these cells recruit CD4 + CD25- T cells and convert them to CD4+CD25+ T cells. Tumor-tropic patient-derived adipose cancer stem cells when treated with exsosomes from prostate cells induced mesenchymal to epithelial transition (MET) and lead to the development of a more aggressive prostate like secondary tumor [118]. Malignant breast cancer stem cells are also associated with CD4+CD25+ T cells [108]. Thus it is possible for the Breast cancer stem cells to utilize exosomes in a similar manner.
The paracrine activity of adult stem cells and cancer stem cells is mediated by the release of exosomes [121]. Cancer stem cells communicate with nearby cancer cells and stromal cells by uptaking the exosomes by the cells. The exosomes derived from the fibroblast mediate Notch signaling, overexpress ALDH, and promote stemness in the breast cancer cells. The intake of exosomes from melanoma cancer cells by the bone marrow progenitor cells leads to the acquisition of malignant phenotype [122]. The exosomes from more aggressive TNBC cell lines Hs578Ts (i) transmitted their aggressive phenotype to secondary breast cancer cells (Hs578T, SKBR3, MDA-MB-231, and HCC1954). The noninvasive nature of the mammary epithelial cell line HMLE was reversed upon its exposure to miR-10b from MDA-MB-231 cells. miR-10b has been reported to be highly expressed by the TNBC cells. In an in vivo study mice were intravenously injected with exosomal miR-105 from MDA-MB-231 cells, and the MDA-MB-231 cells also were intracardially injected resulted in the development of lung and brain metastasis. Under hypoxic conditions the exosomes from breast cancer cells have been shown to promote invasiveness and malignant phenotypes. The expression of RAB22A by the breast cancer cell lines (MCF-7, MDA-MB-231, and MDA-MB-435) is seen. The knockdown of the RAB22A by shRNA showed suppressed invasion and long colonization. The fibroblast promotes metastasis by Wnt signaling. The CD81 secreted by the fibroblast L cells through exosomes was taken up by the breast cancer cells and induces metastasis of the MDA-MB-231 cells. The knockdown of CD81 in L cells suppressed the malignancy [123, 124].
12.7 Future Prospects
Understanding the roles of CSCs on tumor progression and metastasis will provide the strategies for targeting CSCs to prevent the seed for cancer metastasis. This chapter has highlighted the need for future research on the various factors that regulate the dissemination of cancer from its primary site. The RNA-binding proteins and their role in posttranscriptional regulation during cancer progression and metastasis have provided various targets for regulating cancer stem cells. In addition, the development of combination therapies for the above highlighted multiple targets will improve patient’s outcome. The latest development in the field has enabled us to understand the contents and role of CSC-derived exosomes on metastasis. There is a clear lack of information on content loading of exosomes and target or recipient cell identification. The epigenetic regulation of CSCs by microRNAs and RNA-binding proteins has highlighted the targets and identified the biomarkers for tumor progression and metastasis.
References
Lazer LM, Sadhasivam B, Palaniyandi K, Muthuswamy T, Ramachandran I, Balakrishnan A, Pathak S, Narayan S, Ramalingam S (2018) Chitosan-based nano-formulation enhances the anticancer efficacy of hesperetin. Int J Biol Macromol 107:1988–1998
Nguyen LV, Vanner R, Dirks P et al (2012) Cancer stem cells: an evolving concept. Nat Rev Cancer 12(2):133
Mashouri L, Yousefi H, Aref AR, Mohammad Ahadi A, Molaei F, Alahari SK (2019) Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer 18(1):75
Shibue T, Weinberg RA (2017) EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat Rev Clin Oncol 14(10):611–629
Pang R, Law WL, Chu AC et al (2010) A subpopulation of CD26+ cancer stem cells with metastatic capacity in human colorectal cancer. Cell Stem Cell 6(6):603–615
Chen C, Wei Y, Hummel M et al (2011) Evidence for epithelial-mesenchymal transition in cancer stem cells of head and neck squamous cell carcinoma. PLoS One 6(1):e16466
Morel AP, Lièvre M, Thomas C et al (2008) Generation of breast cancer stem cells through epithelialmesenchymal transition. PLoS One 3(8):e2888
Marjanovic ND, Weinberg RA, Chaffer CL (2013) Cell plasticity and heterogeneity in cancer. Clin Chem 59(1):168–179
Kim WT, Ryu CJ (2017 Jun) Cancer stem cell surface markers on normal stem cells. BMB reports. 50(6):285
Huang R, Rofstad EK (2017) Cancer stem cells (CSCs), cervical CSCs and targeted therapies. Oncotarget 8(21):35351
Li Y, Lin K, Yang Z et al (2017) Bladder cancer stem cells: clonal origin and therapeutic perspectives. Oncotarget 8(39):66668–66679
Lin W, Modiano JF, Ito D (2017) Stage-specific embryonic antigen: determining expression in canine glioblastoma, melanoma, and mammary cancer cells. J Vet Sci 18(1):101–104
Yuan ZX, Mo J, Zhao G, Shu G, Fu HL, Zhao W (2016) Targeting strategies for renal cell carcinoma: from renal cancer cells to renal cancer stem cells. Front Pharmacol 7:423
Govaere O, Wouters J, Petz M, Vandewynckel YP, Van den Eynde K, Verhulst S, Dollé L, Gremeaux L, Ceulemans A, Nevens F, van Grunsven LA (2016) Laminin-332 sustains chemoresistance and quiescence as part of the human hepatic cancer stem cell niche. J Hepatol 64(3):609–617
Sun JH, Luo Q, Liu LL, Song GB (2016) Liver cancer stem cell markers: progression and therapeutic implications. World J Gastroenterol 22(13):3547
Ming XY, Fu L, Zhang LY, Qin YR, Cao TT, Chan KW, Ma S, Xie D, Guan XY (2016) Integrin α7 is a functional cancer stem cell surface marker in oesophageal squamous cell carcinoma. Nat Commun 7(1):1–4
Sahlberg SH, Spiegelberg D, Glimelius B, Stenerlöw B, Nestor M (2014) Evaluation of cancer stem cell markers CD133, CD44, CD24: association with AKT isoforms and radiation resistance in colon cancer cells. PLoS One 9(4):e94621
Dawood S, Austin L, Cristofanilli M (2014) Cancer stem cells: implications for cancer therapy. Oncology 28(12):1101–1107, 1110
Guo Z, Hardin H, Lloyd RV (2014) Cancer stem-like cells and thyroid cancer. Endocr Relat Cancer 21(5):T285–T300
Bao B, Ahmad A, Azmi AS, Ali S, Sarkar FH (2013) Overview of cancer stem cells (CSCs) and mechanisms of their regulation: implications for cancer therapy. Curr Protoc Pharmacol 61(1):14–25
Gao MQ, Choi YP, Kang S, Youn JH, Cho NH (2010) CD24+ cells from hierarchically organized ovarian cancer are enriched in cancer stem cells. Oncogene 29(18):2672–2680
Saikawa Y, Fukuda K, Takahashi T, Nakamura R, Takeuchi H, Kitagawa Y (2010) Gastric carcinogenesis and the cancer stem cell hypothesis. Gastric Cancer 13(1):11–24
Li C, Heidt DG, Dalerba P, Burant CF, Zhang L, Adsay V, Wicha M, Clarke MF, Simeone DM (2007) Identification of pancreatic cancer stem cells. Cancer Res 67(3):1030–1037
Bao S, Wu Q, Sathornsumetee S, Hao Y, Li Z, Hjelmeland AB, Shi Q, McLendon RE, Bigner DD, Rich JN (2006) Stem cell–like glioma cells promote tumor angiogenesis through vascular endothelial growth factor. Cancer Res 66(16):7843–7848
Fang D, Leishear K, Nguyen TK, Finko R, Cai K, Fukunaga M, Li L, Brafford PA, Kulp AN, Xu X, Smalley KS (2006) Defining the conditions for the generation of melanocytes from human embryonic stem cells. Stem Cells 24(7):1668–1677
Yun EJ, Lo UG, Hsieh JT (2016) The evolving landscape of prostate cancer stem cell: therapeutic implications and future challenges. Asian J Urol 3(4):203–210
Singh SK, Hawkins C, Clarke ID, Squire JA, Bayani J, Hide T, Henkelman RM, Cusimano MD, Dirks PB (2004) Identification of human brain tumour initiating cells. Nature 432(7015):396–401
García de Herreros A (2014) Epithelial to mesenchymal transition in tumor cells as consequence of phenotypic instability. Front Cell Dev Biol 12(2):71
Kim DH, Xing T, Yang Z, Dudek R, Lu Q, Chen YH (2018) Epithelial mesenchymal transition in embryonic development, tissue repair and cancer: a comprehensive overview. J Clin Med 7(1):1
Clark DW, Palle K (2016) Aldehyde dehydrogenases in cancer stem cells: potential as therapeutic targets. Ann Transl Med 4(24):518
Sikandar SS, Kuo AH, Kalisky T, Cai S, Zabala M, Hsieh RW, Lobo NA, Scheeren FA, Sim S, Qian D, Dirbas FM (2017) Role of epithelial to mesenchymal transition associated genes in mammary gland regeneration and breast tumorigenesis. Nat Commun 8(1):1–9
Bhattacharya R, Mitra T, Ray Chaudhuri S, Roy SS (2018) Mesenchymal splice isoform of CD44 (CD44s) promotes EMT/invasion and imparts stem‐like properties to ovarian cancer cells. J Cell Biochem 119(4):3373–3383
Fan D, Lin X, Zhang F, Zhong W, Hu J, Chen Y, Cai Z, Zou Y, He X, Chen X, Lan P (2018) Micro RNA 26b promotes colorectal cancer metastasis by downregulating phosphatase and tensin homolog and wingless‐type MMTV integration site family member 5A. Cancer Sci 109(2):354–362
Chen DL, Chen LZ, Lu YX, Zhang DS, Zeng ZL, Pan ZZ, Huang P, Wang FH, Li YH, Ju HQ, Xu RH (2017) Long noncoding RNA XIST expedites metastasis and modulates epithelial–mesenchymal transition in colorectal cancer. Cell Death Dis 8(8):e3011
Hudis CA, Gianni L (2011) Triple-negative breast cancer: an unmet medical need. Oncologist 16(Suppl 1):1–11
Carey JP, Karakas C, Bui T, Chen X, Vijayaraghavan S, Zhao Y, Wang J, Mikule K, Litton JK, Hunt KK, Keyomarsi K (2018) Synthetic lethality of PARP inhibitors in combination with MYC blockade is independent of BRCA status in triple-negative breast cancer. Cancer Res 78(3):742–757
Yang M, Li Y, Shen X, Ruan Y, Lu Y, Jin X, Song P, Guo Y, Zhang X, Qu H, Shao Y (2017) CLDN6 promotes chemoresistance through GSTP1 in human breast cancer. J Exp Clin Cancer Res 36(1):1–5
Jiang P, Chen A, Wu X, Zhou M, ul Haq I, Mariyam Z, Feng Q (2018) NEAT1 acts as an inducer of cancer stem cell‐like phenotypes in NSCLC by inhibiting EGCG‐upregulated CTR1. J Cell Physiol 233(6):4852–4863
Mayoral-Varo V, Calcabrini A, Sánchez-Bailón MP, Martín-Pérez J (2017) miR205 inhibits stem cell renewal in SUM159PT breast cancer cells. PLoS One 12(11):e0188637
Kim YJ, Jeong SH, Kim EK, Kim EJ, Cho JH (2017) Ursodeoxycholic acid suppresses epithelial-mesenchymal transition and cancer stem cell formation by reducing the levels of peroxiredoxin II and reactive oxygen species in pancreatic cancer cells. Oncol Rep 38(6):3632–3638
Giacomelli C, Daniele S, Natali L, Iofrida C, Flamini G, Braca A, Trincavelli ML, Martini C (2017) Carnosol controls the human glioblastoma stemness features through the epithelial-mesenchymal transition modulation and the induction of cancer stem cell apoptosis. Sci Rep 7(1):1–7
Lagunas AM, Wu J, Crowe DL (2017) Telomere DNA damage signaling regulates cancer stem cell evolution, epithelial mesenchymal transition, and metastasis. Oncotarget 8(46):80139
Kang HM, Son HS, Cui YH, Youn B, Son B, Kaushik NK, Uddin N, Lee JS, Song JY, Kaushik N, Lee SJ (2017) Phytosphingosine exhibits an anti-epithelial–mesenchymal transition function by the inhibition of EGFR signaling in human breast cancer cells. Oncotarget 8(44):77794
Lin Y, Wang Y, Shi Q, Yu Q, Liu C, Feng J, Deng J, Evers BM, Zhou BP, Wu Y (2017) Stabilization of the transcription factors slug and twist by the deubiquitinase dub3 is a key requirement for tumor metastasis. Oncotarget 8(43):75127
Yan X, Liu L, Li H, Qin H, Sun Z (2017) Clinical significance of Fusobacterium nucleatum, epithelial–mesenchymal transition, and cancer stem cell markers in stage III/IV colorectal cancer patients. Onco Targets Ther 10:5031
Xie SL, Fan S, Zhang SY, Chen WX, Li QX, Pan GK, Zhang HQ, Wang WW, Weng B, Zhang Z, Li JS (2018) SOX8 regulates cancer stem‐like properties and cisplatin‐induced EMT in tongue squamous cell carcinoma by acting on the Wnt/β‐catenin pathway. Int J Cancer 142(6):1252–1265
Garg M (2015) Emerging role of microRNAs in cancer stem cells: implications in cancer therapy. World J Stem Cells 7(8):1078
Takahashi RU, Miyazaki H, Ochiya T (2014) The role of microRNAs in the regulation of cancer stem cells. Front Genet 4:295
Zhou L, Liu F, Wang X, Ouyang G (2015) The roles of microRNAs in the regulation of tumor metastasis. Cell Biosci 5(1):32
El Helou R, Pinna G, Cabaud O, Wicinski J, Bhajun R, Guyon L, Rioualen C, Finetti P, Gros A, Mari B, Barbry P (2017) miR-600 acts as a bimodal switch that regulates breast cancer stem cell fate through WNT signaling. Cell Rep 18(9):2256–2268
Wang ZM, Du WJ, Piazza GA, Xi Y (2013) MicroRNAs are involved in the self-renewal and differentiation of cancer stem cells. Acta Pharmacol Sin 34(11):1374–1380
Shimono Y, Mukohyama J, Nakamura SI, Minami H (2016) MicroRNA regulation of human breast cancer stem cells. J Clin Med 5(1):2
Xiao Y, Humphries B, Yang C, Wang Z (2019) MiR-205 dysregulations in breast cancer: the complexity and opportunities. Non-coding RNA 5(4):53
Bimonte S, Barbieri A, Leongito M, Palma G, Del Vecchio V, Falco M, Palaia R, Albino V, Piccirillo M, Amore A, Petrillo A (2016) The role of miRNAs in the regulation of pancreatic cancer stem cells. Stem Cells Int 2016:8352684
Hu J, Qiu M, Jiang F, Zhang S, Yang X, Wang J, Xu L, Yin R (2014) MiR-145 regulates cancer stem-like properties and epithelial-to-mesenchymal transition in lung adenocarcinoma-initiating cells. Tumor Biol 35(9):8953–8961
Fan X, Chen X, Deng W, Zhong G, Cai Q, Lin T (2013) Up-regulated microRNA-143 in cancer stem cells differentiation promotes prostate cancer cells metastasis by modulating FNDC3B expression. BMC Cancer 13(1):61
Liu C, Liu R, Zhang D, Deng Q, Liu B, Chao HP, Rycaj K, Takata Y, Lin K, Lu Y, Zhong Y (2017) MicroRNA-141 suppresses prostate cancer stem cells and metastasis by targeting a cohort of pro-metastasis genes. Nat Commun 8(1):1–4
Bao B, Li Y, Ahmad A, Azmi AS, Bao G, Ali S, Banerjee S, Kong D, H Sarkar F (2012) Targeting CSC-related miRNAs for cancer therapy by natural agents. Curr Drug Targets 13(14):1858–1868
Yu CC, Lo WL, Chen YW, Huang PI, Hsu HS, Tseng LM, Hung SC, Kao SY, Chang CJ, Chiou SH (2011) Bmi-1 regulates snail expression and promotes metastasis ability in head and neck squamous cancer-derived ALDH1 positive cells. J Oncol 2011:609259
Li XJ, Ren ZJ, Tang JH (2014) MicroRNA-34a: a potential therapeutic target in human cancer. Cell Death Dis 5(7):e1327
Wang Y, Kim S, Kim IM (2014) Regulation of metastasis by microRNAs in ovarian cancer. Front Oncol 4:143
Liu C, Tang DG (2011) MicroRNA regulation of cancer stem cells. Cancer Res 71(18):5950–5954
Pencheva N, Tavazoie SF (2013) Control of metastatic progression by microRNA regulatory networks. Nat Cell Biol 15(6):546–554
Lim YY, Wright JA, Attema JL, Gregory PA, Bert AG, Smith E, Thomas D, Lopez AF, Drew PA, Khew-Goodall Y, Goodall GJ (2013) Epigenetic modulation of the miR-200 family is associated with transition to a breast cancer stem-cell-like state. J Cell Sci 126(10):2256–2266
Wu M-J, Chen Y-S, Kim MR, Chang C-J (2016) Regulation of epithelial plasticity and cancer stemness via microRNAs. J Mol Genet Med 10:2. ISSN: 1747-0862
Ju SY, Chiou SH, Su Y (2014) Maintenance of the stemness in CD44+ HCT-15 and HCT-116 human colon cancer cells requires miR-203 suppression. Stem Cell Res 12(1):86–100
Taube JH, Malouf GG, Lu E, Sphyris N, Vijay V, Ramachandran PP, Ueno KR, Gaur S, Nicoloso MS, Rossi S, Herschkowitz JI (2013) Epigenetic silencing of microRNA-203 is required for EMT and cancer stem cell properties. Sci Rep 3:2687
Zhang Y, Zhou SY, Yan HZ, Xu DD, Chen HX, Wang XY, Wang X, Liu YT, Zhang L, Wang S, Zhou PJ (2016) miR-203 inhibits proliferation and self-renewal of leukemia stem cells by targeting survivin and Bmi-1. Sci Rep 6(1):1–2
Yu G, Yao W, Xiao W, Li H, Xu H, Lang B (2014) MicroRNA-34a functions as an anti-metastatic microRNA and suppresses angiogenesis in bladder cancer by directly targeting CD44. J Exp Clin Cancer Res 33(1):779
Kumar B, Yadav A, Lang J, Teknos TN, Kumar P (2012) Dysregulation of microRNA-34a expression in head and neck squamous cell carcinoma promotes tumor growth and tumor angiogenesis. PLoS One 7(5):e37601
Liu C, Kelnar K, Liu B, Chen X, Calhoun-Davis T, Li H, Patrawala L, Yan H, Jeter C, Honorio S, Wiggins JF, Bader AG, Fagin R, Brown D, Tang DG (2011) The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nat Med 17:211–215
Zhang X, Ai F, Li X, Tian L, Wang X, Shen S, Liu F (2017) MicroRNA‑34a suppresses colorectal cancer metastasis by regulating Notch signaling. Oncol Lett 14(2):2325–2333
Gao Y, Luo LH, Li S, Yang C (2014) miR-17 inhibitor suppressed osteosarcoma tumor growth and metastasis via increasing PTEN expression. Biochem Biophys Res Commun 444(2):230–234
Schubbert S, Jiao J, Ruscetti M, Nakashima J, Wu S, Lei H, Xu Q, Yi W, Zhu H, Wu H (2016) Methods for PTEN in stem cells and cancer stem cells. In: PTEN. Humana Press, New York, pp 233–285
Jiang Z, Yin J, Fu W, Mo Y, Pan Y, Dai L, Huang H, Li S, Zhao J (2014) MiRNA 17 family regulates cisplatin-resistant and metastasis by targeting TGFbetaR2 in NSCLC. PLoS One 9(4):e94639
Ajani JA, Song S, Hochster HS, Steinberg IB (2015) Cancer stem cells: the promise and the potential. In: Seminars in oncology, vol 42. WB Saunders, pp S3–S17
Zhang J, Xiao Z, Lai D, Sun J, He C, Chu Z, Ye H, Chen S, Wang J (2012) miR-21, miR-17 and miR-19a induced by phosphatase of regenerating liver-3 promote the proliferation and metastasis of colon cancer. Br J Cancer 107(2):352–359
Gong C, Yang Z, Wu F, Han L, Liu Y, Gong W (2016) miR-17 inhibits ovarian cancer cell peritoneal metastasis by targeting ITGA5 and ITGB1. Oncol Rep 36(4):2177–2183
Xia H, Cheung WK, Ng SS, Jiang X, Jiang S, Sze J, Leung GK, Lu G, Chan DT, Bian XW, Kung HF (2012) Loss of brain-enriched miR-124 microRNA enhances stem-like traits and invasiveness of glioma cells. J Biol Chem 287(13):9962–9971
Wei J, Wang F, Kong LY, Xu S, Doucette T, Ferguson SD, Yang Y, McEnery K, Jethwa K, Gjyshi O, Qiao W (2013) MiR-124 inhibits STAT3 signaling to enhance T cell-mediated immune clearance of glioma. Cancer Res 73(13):3913–3926
Lv XB, Jiao Y, Qing Y, Hu H, Cui X, Lin T, Song E, Yu F (2011) miR-124 suppresses multiple steps of breast cancer metastasis by targeting a cohort of pro-metastatic genes in vitro. Chin J Cancer 30(12):821
Zhou XU, Qi L, Tong S, Cui YU, Chen J, Huang T, Chen Z, Zu XB (2015) miR-128 downregulation promotes growth and metastasis of bladder cancer cells and involves VEGF-C upregulation. Oncol Lett 10(5):3183–3190
Zhao X, Wu Y, Lv Z (2015) miR-128 modulates hepatocellular carcinoma by inhibition of ITGA2 and ITGA5 expression. Am J Transl Res 7(9):1564
Sun X, Li Y, Yu J, Pei H, Luo P, Zhang J (2015) miR-128 modulates chemosensitivity and invasion of prostate cancer cells through targeting ZEB1. Jpn J Clin Oncol 45(5):474–482
Zeng H, Zhang Z, Dai X, Chen Y, Ye J, Jin Z (2016) Increased expression of microRNA-199b-5p associates with poor prognosis through promoting cell proliferation, invasion and migration abilities of human osteosarcoma. Pathol Oncol Res 22(2):253–260
Fang C, Zhao Y, Guo B (2013) MiR‐199b‐5p targets HER2 in breast cancer cells. J Cell Biochem 114(7):1457–1463
Zhou SJ, Liu FY, Zhang AH, Liang HF, Wang Y, Ma R, Jiang YH, Sun NF (2017) MicroRNA-199b-5p attenuates TGF-β1-induced epithelial–mesenchymal transition in hepatocellular carcinoma. Br J Cancer 117(2):233–244
Wang Z, Zhang H, Zhang P, Li J, Shan Z, Teng W (2013) Upregulation of miR-2861 and miR-451 expression in papillary thyroid carcinoma with lymph node metastasis. Med Oncol 30(2):577
Zhang J, Qin X, Sun Q, Guo H, Wu X, Xie F, Xu Q, Yan M, Liu J, Han Z, Chen W (2015) Transcriptional control of PAX4-regulated miR-144/451 modulates metastasis by suppressing ADAMs expression. Oncogene 34(25):3283–3295
Zhang F, Huang W, Sheng M, Liu T (2015) MiR-451 inhibits cell growth and invasion by targeting CXCL16 and is associated with prognosis of osteosarcoma patients. Tumor Biol 36(3):2041–2048
Huang JY, Zhang K, Chen DQ, Chen J, Feng B, Song H, Chen Y, Zhu Z, Lu L, De W, Wang R (2015) MicroRNA-451: epithelial-mesenchymal transition inhibitor and prognostic biomarker of hepatocelluar carcinoma. Oncotarget 6(21):18613
Liu G, Xu Z, Hao D (2016) MicroRNA‑451 inhibits neuroblastoma proliferation, invasion and migration by targeting macrophage migration inhibitory factor. Mol Med Rep 13(3):2253–2260
Yin P, Peng R, Peng H, Yao L, Sun Y, Wen L, Wu T, Zhou J, Zhang Z (2015) MiR-451 suppresses cell proliferation and metastasis in A549 lung cancer cells. Mol Biotechnol 57(1):1–11
Lei T, Zhu Y, Jiang C, Wang Y, Fu J, Fan Z, Qin H (2016) MicroRNA-320 was downregulated in non-small cell lung cancer and inhibited cell proliferation, migration and invasion by targeting fatty acid synthase. Mol Med Rep 14(2):1255–1262
Shi C, Zhang Z (2017) MicroRNA‑320 suppresses cervical cancer cell viability, migration and invasion via directly targeting FOXM1. Oncol Lett 14(3):3809–3816
Pereira B, Billaud M, Almeida R (2017) RNA-binding proteins in cancer: old players and new actors. Trends Cancer 3(7):506–528
Denkert C, Koch I, von Keyserlingk N, Noske A, Niesporek S, Dietel M, Weichert W (2006) Expression of the ELAV-like protein HuR in human colon cancer: association with tumor stage and cyclooxygenase-2. Modern Pathol 19(9):1261–1269
Hong S (2017) RNA binding protein as an emerging therapeutic target for cancer prevention and treatment. J Cancer Prev 22(4):203
Sureban SM, Ramalingam S, Natarajan G, May R, Subramaniam D, Bishnupuri KS, Morrison AR, Dieckgraefe BK, Brackett DJ, Postier RG, Houchen CW (2008) Translation regulatory factor RBM3 is a proto-oncogene that prevents mitotic catastrophe. Oncogene 27(33):4544–4556
Venugopal A, Subramaniam D, Balmaceda J, Roy B, Dixon DA, Umar S, Weir SJ, Anant S (2016) RNA binding protein RBM3 increases β‐catenin signaling to increase stem cell characteristics in colorectal cancer cells. Mol Carcinog 55(11):1503–1516
Hou P, Li L, Chen F, Chen Y, Liu H, Li J, Bai J, Zheng J (2018) PTBP3-mediated regulation of ZEB1 mRNA stability promotes epithelial–mesenchymal transition in breast cancer. Cancer Res 78(2):387–398
Mukohyama J, Shimono Y, Minami H, Kakeji Y, Suzuki A (2017) Roles of microRNAs and RNA-binding proteins in the regulation of colorectal cancer stem cells. Cancers 9(10):143
Pastò A, Serafin V, Pilotto G, Lago C, Bellio C, Trusolino L, Bertotti A, Hoey T, Plateroti M, Esposito G, Pinazza M (2014) NOTCH3 signaling regulates MUSASHI-1 expression in metastatic colorectal cancer cells. Cancer Res 74(7):2106–2118
Shou Z, Jin X, He X, Zhao Z, Chen Y, Ye M, Yao J (2017) Overexpression of Musashi-1 protein is associated with progression and poor prognosis of gastric cancer. Oncol Lett 13(5):3556–3566
Kudinov AE, Deneka A, Nikonova AS, Beck TN, Ahn YH, Liu X, Martinez CF, Schultz FA, Reynolds S, Yang DH, Cai KQ (2016) Musashi-2 (MSI2) supports TGF-β signaling and inhibits claudins to promote non-small cell lung cancer (NSCLC) metastasis. Proc Natl Acad Sci U S A 113(25):6955–6960
Codony-Servat J, Rosell R (2015) Cancer stem cells and immunoresistance: clinical implications and solutions. Transl Lung Cancer Res 4(6):689
Jachetti E, Caputo S, Mazzoleni S, Brambillasca CS, Parigi SM, Grioni M, Piras IS, Restuccia U, Calcinotto A, Freschi M, Bachi A (2015) Tenascin-C protects cancer stem–like cells from immune surveillance by arresting T-cell activation. Cancer Res 75(10):2095–2108
Kitamura T, Qian BZ, Soong D, Cassetta L, Noy R, Sugano G, Kato Y, Li J, Pollard JW (2015) CCL2-induced chemokine cascade promotes breast cancer metastasis by enhancing retention of metastasis-associated macrophages. J Exp Med 212(7):1043–1059
Zhao D, Pan C, Sun J, Gilbert C, Drews-Elger K, Azzam DJ, Picon-Ruiz M, Kim M, Ullmer W, El-Ashry D, Creighton CJ (2015) VEGF drives cancer-initiating stem cells through VEGFR-2/Stat3 signaling to upregulate Myc and Sox2. Oncogene 34(24):3107–3119
Li YL, Zhao H, Ren XB (2016) Relationship of VEGF/VEGFR with immune and cancer cells: staggering or forward? Cancer Biol Med 13(2):206
Hsu YL, Hung JY, Tsai YM, Tsai EM, Huang MS, Hou MF, Kuo PL (2015) 6-Shogaol, an active constituent of dietary ginger, impairs cancer development and lung metastasis by inhibiting the secretion of CC-chemokine ligand 2 (CCL2) in tumor-associated dendritic cells. J Agric Food Chem 63(6):1730–1738
Xiang ZL, Zeng ZC, Fan J, Wu WZ, He J, Zeng HY, Tang ZY (2011) A clinicopathological model to predict bone metastasis in hepatocellular carcinoma. J Cancer Res Clin Oncol 137(12):1791
Kim SW, Kim HY, Song IC, Jin SA, Lee HJ, Yun HJ, Kim S, Jo DY (2008) Cytoplasmic trapping of CXCR4 in hepatocellular carcinoma cell lines. Cancer Res Treat 40(2):53
Jeng KS, Jeng CJ, Jeng WJ, Chang CF, Sheen I (2017) Role of CXC chemokine ligand 12/CXC chemokine receptor 4 in the progression of hepatocellular carcinoma. Oncol Lett 14(2):1905–1910
Geiger P, Mayer B, Wiest I, Schulze S, Jeschke U, Weissenbacher T (2016) Binding of galectin-1 to breast cancer cells MCF7 induces apoptosis and inhibition of proliferation in vitro in a 2D-and 3D-cell culture model. BMC Cancer 16(1):1–9
Zhou X, Li D, Wang X, Zhang B, Zhu H, Zhao J (2015) Galectin-1 is overexpressed in CD133+ human lung adenocarcinoma cells and promotes their growth and invasiveness. Oncotarget 6(5):3111
Kitamura T, Qian B-Z, Pollard JW (2015) Immune cell promotion of metastasis. Nat Rev Immunol 15(2):73–86
Syn N, Wang L, Sethi G et al (2016) Exosome-mediated metastasis: from epithelial–mesenchymal transition to escape from immunosurveillance. Trends Pharmacol Sci 37(7):606–617
Desgrosellier JS, Cheresh DA (2010) Integrins in cancer: biological implications and therapeutic opportunities. Nat Rev Cancer 10(1):9–22
Peinado H, Alečković M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G, Hergueta-Redondo M, Williams C, García-Santos G, Ghajar CM, Nitadori-Hoshino A (2012) Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med 18(6):883–891
Parfejevs V, Sagini K, Buss A, Sobolevska K, Llorente A, Riekstina U, Abols A (2020) Adult stem cell-derived extracellular vesicles in cancer treatment: opportunities and challenges. Cells 9(5):1171
Hannafon BN, Ding WQ (2015) Cancer stem cells and exosome signaling. Stem Cell Invest 2:11
O’Brien K, Rani S, Corcoran C, Wallace R, Hughes L, Friel AM, McDonnell S, Crown J, Radomski MW, O’Driscoll L (2013) Exosomes from triple-negative breast cancer cells can transfer phenotypic traits representing their cells of origin to secondary cells. Eur J Cancer 49(8):1845–1859
Lowry MC, Gallagher WM, O’Driscoll L (2015) The role of exosomes in breast cancer. Clin Chem 61(12):1457–1465
Acknowledgments
The authors would like to thank the SRM Institute of Science and Technology for funding and providing the laboratory facility. We would also like to thank Science and Engineering Research Board (SERB)-EMR/2017/002874, Indian Council of Medical Research (ICMR)-2019-5526/CMB/BMS and Department of Biotechnology (DBT)-BT/PR26189/GET/119/226/2017 for the funding support provided.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Lizha Mary, L., Vasantha Kumar, M., Satish, R. (2020). Cancer Stem Cells as a Seed for Cancer Metastasis. In: Pathak, S., Banerjee, A. (eds) Cancer Stem Cells: New Horizons in Cancer Therapies. Springer, Singapore. https://doi.org/10.1007/978-981-15-5120-8_12
Download citation
DOI: https://doi.org/10.1007/978-981-15-5120-8_12
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-5119-2
Online ISBN: 978-981-15-5120-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)