Abstract
T cells recognize peptides bound to major histocompatibility complex (MHC) class I and class II molecules at the cell surface. This recognition is accomplished by the expression of T cell receptors (TCR) which are required to be diverse and adaptable in order to accommodate the various and vast number of antigens presented on the MHCs. Thus, determining TCR repertoires of effector T cells is necessary to understand the immunological process in responding to cancer progression, infection, and autoimmune development. Furthermore, understanding the TCR repertoires will provide a solid framework to predict and test the antigen which is more critical in autoimmunity. However, it has been a technical challenge to sequence the TCRs and provide a conceptual context in correlation to the vast number of TCR repertoires in the immunological system. The exploding field of single-cell sequencing has changed how the repertoires are being investigated and analyzed. In this review, we focus on the biology of TCRs, TCR signaling and its implication in autoimmunity. We discuss important methods in bulk sequencing of many cells. Lastly, we explore the most pertinent platforms in single-cell sequencing and its application in autoimmunity.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- T cells
- T cell receptors
- T cell receptor signaling
- Single cell sequencing
- Bulk sequencing
- Sjogren’s syndrome
3.1 Introduction
T cells are a critical component of the adaptive immune system capable of recognizing a nearly infinite variety of antigens. T cells recognize antigens through unique antigen receptors called T cell receptors (TCRs). Unlike the closely related B cell receptors (BCRs), which can secrete as an antibody, TCRs are strictly membrane bound and are not secreted [1, 2]. As a result, TCRs lack the opsonization and neutralization abilities of antibodies and are committed to participating in antigen recognition followed by intracellular signaling and subsequent T cell activation [3]. TCRs are further distinguished from BCRs by their mechanisms for antigen recognition. While BCRs can directly bind to a wide variety of molecules, TCRs are restricted to recognizing short peptides of protein antigens processed and presented by major histocompatibility complexes (MHCs) on the body’s own antigen presenting cells (APCs) [4]. Fundamentally, TCRs function as an antigen recognition receptor complexed to T cell activating signaling machinery and are activated in response to short continuous amino acid sequences presented on MHC. TCRs are composed of two heterodimeric polypeptide chains linked by a disulfide bond [1]. Each chain of the TCR consists of two extracellular immunoglobulin domains, a transmembrane region and a short cytoplasmic tail [1]. The two extracellular domains are made up of the variable (V) region and constant (C) region [4]. The heterodimeric structure of the TCR is analogous to the heavy and light chain heterodimers of BCRs [5]. However, the forked structure of the BCR consists of two antigen binding sites, whereas each TCR possesses a single antigen binding site. The majority of TCRs possess an α chain and a β chain and are referred to as αβ TCRs. A subset of T cells possesses a γ chain and δ chain and are referred to as γδ TCRs. γδ TCRs are capable of directly recognizing antigens outside the context of MHC and are even capable of recognizing non-peptide antigens [6,7,8].
T cells possess the ability to bind to a vast array of peptide antigens through their TCRs; it has been estimated that humans can produce between 1015 and 1020 possible unique TCR chains [9]. This enormous variety is imparted by an unusual genetic mechanism, largely shared with BCR generation, that provides diversity concentrated in the antigen binding regions of the TCR [10]. The V region is the portion of the TCR that participates in antigen binding. The V region is not encoded by a single segment of DNA, but rather is composed of multiple gene segments that are rearranged through somatic DNA recombination. Combinatorial diversity afforded through recombination of the gene segments is further augmented by junctional diversity through the random addition of nucleotides at the interface between segments, thus allowing for the generation of a nearly limitless array of TCRs [11]. The DNA encoding the α chain of the TCR possesses multiple variable (V) and joining (J) segments, whereas the β chain possesses multiple V, diversity (D), and segments [11, 12] as represented in Fig. 3.1. A functional TCR will consist of an α chain with a single V and J segment and a β chain with a single V, D, and J segment. The act of rearranging these segments into functional TCRs is referred to as VDJ recombination. During VDJ recombination, the gene segments for the α and β chains of an individual T cell are rearranged in order to produce a new functional TCR. VDJ recombination employs both the lymphoid-specific RAG1-RAG2 complex, so named for recombination activating genes (RAG), and enzymes of the more broadly utilized double stranded break repair (DSBR) system [12, 13]. Antigen binding within the TCR V region involves the three complementarity determining regions (CDRs) that contact the antigen MHC complex. CDR1 and CDR2 are primarily encoded in the V germline segments and therefore experience less diversity. CDR3 however includes the junctional regions and is the primary region in contact with the antigen [5, 14]. The diversity of the TCR repertoire can change or evolve at different stages of the disease. If one can grasp the repertoire diversity, one can understand the immune response under various disease conditions, specifically infectious diseases, cancer, and autoimmune diseases.
The genetic basis of TCR formation. The figure represents the mRNAs somatic VDJ recombination to form the alpha and beta chains of TCRs. As indicated, there is an arrangement step that recombines the VDJ segment for TCR β and V and J segments for the TCR α chain. The mRNAs formed have addition and deletion of nucleotides at the junctions of these segments leading to junctional diversity that leads to variability for assessing specific antigens. There may be different combinations of genes leading to the final formation of the TCR that consists of the TCR α and β subunit organized in a constant and variable region wherein the variable region is responsible for antigen recognition
In the past decades, in an attempt to examine the TCR repertoires, we relied on quantitative polymerase chain reaction (PCR) and spectratyping techniques to capture the TCR sequences. The limitations of these processes are the low resolution, favoring of the most dominant sequences, and the inability to pair the receptor expression. These constraints limit our ability to resolve the heterogeneity of TCR expression at high resolution. Currently, with the advancement of single-cell isolation processes and sequencing technologies, we are able to decipher the paired expression of TCRs at single-cell resolution. Understanding TCRs at the individual cell level will revolutionize how to profile immune cell repertoires, antigen discovery, disease prognosis, and treatment. The body of this review will focus on both the specific techniques and strategies used in the past and current single-cell technologies used to define and sequence TCR sequences, and how these techniques provide a better understanding of the immune cascade following antigen presentation in autoimmune diseases.
3.2 TCR Repertoires in Autoimmunity
3.2.1 TCR Signaling
TCRs recognize processed peptide antigen presented on MHC on the surface of the body’s own cells. The two conventional MHCs, MHC I and MHC II are both polygenic and polymorphic noncovalent protein complexes composed of two polypeptide chains [4, 15]. TCRs are specific to both peptide antigen and the MHC to which it is bound, a phenomenon known as MHC restriction [4, 16]. MHC I is on the surface of virtually all nucleated cells in the body. Peptides presented on MHC I are generally 8–10 amino acids in length and result from the processing of foreign intracellular proteins [17, 18]. For this reason, MHC I is frequently used to signal viral infection to cytotoxic CD8 T cells. MHC II is only present on the surface of antigen presenting cells of the immune system including B cells, macrophages, and dendritic cells. MHC II presents peptides of 13–17, amino acids in length that have been collected from the extracellular environment [15]. MHC II is recognized by CD4 helper T cells which upon activation stimulate the effector arms of the immune system [19]. The T cell co-receptors CD4 and CD8 increase the T cell sensitivity to antigens and serve to jumpstart TCR signaling [1, 2]. T cell activation is considered to require three discrete signals delivered to T cells [3] (Fig. 3.2). Signal 1 occurs through the interaction of TCR and antigenic peptide presented on MHC. Association of T cell co-receptors CD4 or CD8 and MHC II or MHC I respectively are also considered part of signal 1. Signal 2 consists of a co-stimulatory signal commonly transmitted by B7.1 and B7.2 interacting with CD28 on the T cell surface [19, 20]. Signal 1 and signal 2 together drive T cell survival and clonal expansion. Signal 3 is delivered through the cytokine environment acting on the T cell, and determines differentiation of CD4 T cells into T helper Th1, Th2, or Th17 subsets [19] which further in an immune response lead to different effector functions (Fig. 3.3).
Three discrete signals for T cell activation. Signal 1 occurs through the interaction of TCR and antigenic peptide presented on MHC. Signal 2 is indicated that consists of a co-stimulatory signal that is commonly transmitted by the B7.1 and B7.2 receptors of the antigen presenting cell that interact with CD28 on the T cell surface. Signal 3 that is delivered as a result of the cytokine environment acting that determines the differentiation of the T cell to the Th1, Th2, or Th17 subset
3.2.2 Importance of Shaping the T cell Repertoire in Autoimmunity
While VDJ recombination allows for the development of an arsenal of TCRs capable of recognizing a nearly limitless array of foreign peptides and mounting an immune response, some of the receptors generated will be reactive against self-peptides. These self-reactive TCRs can erroneously engage the machinery of the immune system against the body’s own tissues, resulting in tissue destruction and potentially an autoimmune disease [21]. T cells expressing TCRs that are strongly self-reactive are typically culled or inactivated through the mechanisms of central tolerance during T cell development in the thymus and peripheral tolerance after the T cells have exited the thymus and entered the secondary lymphoid tissues [21, 22]. However, the systems maintaining self-tolerance are imperfect and some self-reactive T cells will escape selection [23]. Additionally, some TCRs are specific for foreign peptides but also reactive against self-peptides [24]. Therefore, the immune system must balance the elimination of self-reactive T cells with the maintenance of a TCR repertoire capable of providing protection. During infection, when latently autoreactive T cells are exposed to self-peptide in an inflammatory environment, the mechanisms of self-tolerance can break down and permit the activation of autoreactive T cells [25, 26].
Autoreactive T cells are critically involved in many autoimmune diseases either as effector cells or through the promotion of autoantibody responses by providing T cell help to B cells [21]. Type I diabetes is characterized by the infiltration of leukocytes, particularly autoreactive CD8 T cells, into the pancreas [27] where the cytotoxic CD8 T cells kill the insulin producing beta cells, resulting in the development of disease [28]. Multiple sclerosis is another T cell mediated autoimmune disease where Th17 cells, subsets of CD4 T cells, propel an inflammatory response that results in the destruction of the myelin sheath around nerve axons [29,30,31]. During the pathogenesis of psoriasis, multiple autoreactive T cell subtypes including Th1 and Th17 cells are activated and contribute towards the development of inflammatory skin disease [32,33,34]. For many autoimmune diseases, the antigens capable of activating autoreactive T cells have yet to be defined. Discovery of the sequences of TCRs of clonally activated T cells in autoimmune diseases presents an opportunity to elucidate the antigen-specific response of these clones and gain a better understanding of the determinants of autoimmune disease.
3.3 Bulk Sequencing Technologies
Multiple techniques of immune repertoire analysis have been developed over the last decade. Pioneering studies on TCR repertoires were initially based on the cloning and sequencing of PCR-amplified cDNA collections [35,36,37,38,39,40]. The mRNA gene expression level of TCR variable gene segments in bulk is analyzed by techniques such as semi-quantitative PCR using TCR Vβ family specific primers [41,42,43,44]. Bulk sequencing and analysis do not capture the expression of single cells, but rather capture bulk samples or many cells within the samples. Despite being comprehensive, this qualitative and quantitative technique does not provide any information about CDR3 region of TCRs which forms an integral component of the receptor [4]. The following discussion will detail some of the major techniques for bulk TCR analysis.
3.3.1 CDR3 Sequence Analysis by Immunoscope
Immunoscope, also known as CDR3 spectratyping, was the first qualitative method at the molecular genetic level for analyzing CDR3 polymorphisms and sequence length diversity [9, 45,46,47]. This method based on electrophoretic analysis of CDR3-PCR fragment length has been the standard technique of TCR evaluation since the 1990s [39, 48]. Musette et al. found an expansion of oligoclonal Vβ5.3 + T cells population in HLA-DR2 multiple sclerosis (MS) patients [49]. Ercolini and Miller characterized the cross-reactive anti-self response induced by a peptide mimic on a model for MS [50]. Immunoscope analysis of TCRVβ repertoire diversity in patients with autoimmune lymphoproliferative syndrome showed an expansion of TCRα/β+CD4−CD8− double negative T cells derived extra-thymically from CD8+ T cells [51]. Immunoscope analysis has technical limitations and lacks the key parameters of capturing TCR diversity such as specific nucleotide sequences, codon usage, and amino acid composition [49]. In parallel to spectratyping, development of real-time PCRs opened the possibility to evaluate TCR repertoires with higher precision [39, 50,51,52,53,54]. Pairing immunoscope with other techniques allowed capture of quantitative and qualitative data for the repertoire of V domain and C region combinations [55,56,57]. Current approaches aim to quantify genomic DNA (gDNA) or mRNA sequences corresponding to CDR3 region of TCR β chain, the most variable TCR region [58, 59]. CDR3 sequences are usually unique markers and the key determinants of antigenic recognition, quantitative T cell responses, and clonal composition of the T cell repertoire [60]. However, CDR1 and CDR2 are also associated with MHC restriction [61, 62] and therefore sequencing of all regions is important to provide the complete information about TCR structure and its binding characteristics [63]. Selecting between gDNA and mRNA is an important consideration for sequencing. The gDNA presents high long-term stability, ease of isolation, and presence of a single DNA template per T cell. The disadvantages include possible errors of sequencing due to introns, cells with residues of VDJ rearrangements and interfering priming sites of the sample. Contrary to the presence of multiple TCR transcripts in gDNA, using mRNA provides the sequence of the entire V and J gene [51, 59, 63] which is the basis of specific antigen recognition. Lim et al. combined immunoscope with MHC peptides multimer-based T cell sorting; however, specific T cells could not be detected due to their low frequency and/or an insufficient amount of TCR β chain transcript [57, 64]. For the first time at the nucleotide sequence level, the TCR repertoire was analyzed using molecular cloning and Sanger sequencing [9, 65, 66]. Nonetheless these low-throughput approaches were limited due to their failure to individually sequence a large number of T cell clones. Over the years, there have been several methodologies taken into consideration for successful and correct sequencing analyses. With technological interventions, it has become possible to amplify both TCR α and β chains simultaneously. Fundamentally, the β chain is more preferred for analysis due to the unique presence of the D gene component [63].
3.3.2 High-Throughput Sequencing Techniques
3.3.2.1 Multiple-Based PCR Methods
All previous techniques presented several limitations to create an extensive analysis of TCR repertoire profiling until the development of high-throughput sequencing (HTS) techniques. These allowed for massive parallel sequencing of millions of TCR sequences. The application of HTS presented promising results in clinical needs pertaining to infectious diseases, cancer, and specially autoimmunity [60, 67, 68]. There are three principal PCR-based enrichment techniques that are used: the multiplex PCR, target enrichment PCR, and 5′RACE-switch-oligo nested PCR. The multiplex technique allows working with both starting materials: gDNA and RNA and contains a mix of primers complementary to all known V segments and a pool of reverse primers for J segment or constant region exons [69, 70]. There are several kits that are available on the market, for example: BGI, Adaptive Biotechnologies, iRepertoire [71,72,73] that facilitate the multiplex PCR technique. Target enrichment-based PCRs also work with both gDNA and RNA. TCR-specific baits complementary to α and β transcripts hybridize with molecules in the library, are conjugated to magnetic beads and then undergo amplification before sequencing. Kits such as SMARTer ThruPLEX have been developed by Takara Bio. Lastly the nested PCR approach is an alternative technology that has been introduced into the bulk sequencing and is based on the 5′RACE switch-oligo approach designed only for RNA. It involves presenting an insert of an adaptor molecule at the 5′ end of the cDNA during cDNA synthesis. This technique was developed by Clontech and follows the principle of a PCR where the forward primer binds to 5′ adaptor molecule and reverse primer binds to the C region of transcript improving specificity [74]. Hendriksen et al. analyzed T cell repertoires of paired liver and gut samples from patients suffering from primary sclerosing cholangitis with inflammatory bowel disease, and demonstrated their common origin [78]. Spreafico et al. discovered for the first time circulating pathogenic-like lymphocytes in patients with juvenile and rheumatoid arthritis TCR. These cells are enriched in synovial clonotypes, circulate through the site of autoimmune reaction, and correlated with progression of juvenile and adult autoimmune arthritis [79]. Expanded findings are summarized in Table 3.1.
3.3.2.2 Commercial Platforms for Next Gen Sequencing
Next Gen Sequencing (NGS) consists of current technologies that have higher accuracy as compared to Sanger High-Throughput Sequencing (HTS) platforms. The NGS platforms have refined technologies over time and as a result there are several available currently. Technologies such as GS FLX by 454 Life Sciences/Roche diagnostics, Genome Analyzer, HiSeq, MiSeq and NextSeq by Illumina, Inc., SOLiD by ABI, Ion Torrent by Life Technologies, Oxford Nanopore, Complete Genomics by Beijing Genomics Institute, and GnuBIO are a few platforms that have different characteristics of sequencing accuracy. The advantages they provide over HTS differ based on sequencing depth, read length, error type and their frequency but include individual TCR clonotype quantification [58, 75]. Since error rate is a major consideration, there is a trade-off in choosing a sequencing platform needed to correctly manage it. Library preparation for companies such as Illumina sequencing, Roche, Life Technologies, and Pacific Biosciences are almost the same with differences in parameters such as sequencing adapters, cluster generation and application of technologies such as paired end sequencing. The basic process includes RNA extraction and fragmentation from the sample and conversion into cDNA by reverse transcription. DNA fragments are ligated to sequencing adapters with specific sequences [76]. These sequences are designed to interact with a specific platform either the surface of an individual flow cell in case of Illumina or the microemulsion-based platform for Ion Torrent. Library construction involves three primary steps: (1) fragmentation of starting material DNA or RNA, (2) adaptor ligation, and (3) amplification [81, 82]. Initial DNA can be sheared into short fragments of 200–400 bp (depending on the sequencing platform) by physical methods (acoustic sonication and hydrodynamic shearing), enzymatic (endonuclease and transposase) or chemical methods (heat digestion with divalent metal cations) [81, 83]. Random/unbiased fragmentation allowing to produce overlapping segments of optimal length is one of successful factors for NGS library construction. After fragmentation end repair is required, single stranded segment ends are blunted and 5′ phosphorylated, following A-tailing by Taq or a Klenow fragment. The sequencing adaptors are then ligated to the DNA fragments forming the fragment library. Several cycles of PCR are performed to enrich for the library product.. Finally, the DNA library can be evaluated for quality control and followed by sequencing through NGS [81, 84].
3.3.2.3 The Illumina Platform and Technologies
The Illumina platform presents the leading position in the NGS market and it provides sequencing by synthesis based on reversible dye-terminators. Illumina HiSeq and MiSeq use four-channel sequencing systems with individual image detection of each base. The NextSeq 500 presents a two-sequencing system wherein only two images are needed for four base calls that allow reduction in time, cost of sequencing, and number of cycles as there are two images simultaneously analyzed from both ends [77]. Illumina MiSeq is widely used for capturing the most common and expanded clonotypes, whereas Illumina HiSeq is mostly applied for increasing the depth in sequencing [59, 63]. The principal drawback of bulk sequencing is that it provides information about the frequency of single TCR chains, but no information regarding their pairing [63]. To remove this bottleneck, unique molecular identifiers (UMIs) have been introduced during cDNA synthesis that allow for reduction in errors during amplification during a PCR [74]. Using high-throughput deep TCRβ chain sequencing, Muraro et al. showed that autologous stem cell transplantation has distinctive effects on CD4+ and CD8+ T cell repertoires in multiple sclerosis patients [78]. Incorporating multiple-PCR and 5′ rapid-amplification of cDNA ends (RACE) to capture CDR3 regions and high-throughput sequencing platform HiSeq2000, the authors found no significant difference in TCR repertoires between pancreatic cancer patients versus healthy controls, and no differences were found between the samples of tumor tissue and the blood samples from patients. Applying a template-switch anchored RT-PCR and Illumina sequencing, diverse repertoire of clonally expanded tumor-reactive lymphocytes was identified on CD8+ melanoma tumor-infiltrating lymphocytes (TILs). Additionally, sequencing the TCRβ revealed that tumor-reactive and mutation-specific clonotypes were highly expanded in the CD8+ population and preferentially expanded in the PD-1+ population [79,80,81].
3.4 Single-Cell Sequencing Techniques
As opposed to the bulk cell analyses that provide coarse or broader resolution, the TCR repertoires can be analyzed at a second level that is more fine-tuned at higher resolution with single-cell sequencing analysis. This allows us to visualize the complex interplay between varied cell types of the immune system bringing about a systemic response. Bulk technologies fail mostly in understanding the nuances of the complex cell responses while single cells help in resolving ambiguity as analyzing the interactions microscopically between individual cells provides details giving insight to bigger reactions [82]. As indicated, bulk sequencing procedures are only able to catch the most common and expanded clonotypes which may or may not be ideally suggestive of the immune responses during an infection or autoimmune process [83]. Single-cell analyses on the other hand provide information indicative about rare TCR subset sequence repertoires that will yield a comprehensive understanding of the biological processes of individual T cells and their dynamic interaction within the biological system.
As discussed, initially platforms focused primarily on providing sequencing information on the dominant TCR α and β chains, but current technologies can sequence the rarer and less frequent cell populations such as TCR of γδ T cells [84]. In T cell repertoire formation, it is essential to understand the development and expansion of the cells based on TCR expression. The differentiation of a progenitor T cell into a specific subset can be identified by single-cell RNA (scRNA) sequencing as there is clonal expansion of a specific prototype [84, 85]. Since single-cell sequencing helps in identification of the TCR chains at a cellular level, depth becomes an important consideration as it provides vital information regarding repertoire formation [86, 87]. Major variability can be observed between two similar cells during the analysis of single-cell-based data. This variability may be attributed to either inherent biological variations or a result of the technique for repertoire analysis. Single-cell technologies for TCR repertoire analysis include PCR that involves the reverse transcription of the mRNA to amplify the cDNA from a single cell. The goal in single-cell receptor sequencing is to obtain maximum coverage of cell types and a low cost per cell, with comprehensive algorithms and platforms for analysis that provide extensive detail [88]. The challenges include the capture of single cells, isolation and amplification of sufficient materials, and generation of vast volumes of data that require faster methods of computation and algorithmic tools that allow efficient data management.
3.4.1 Microfluidic Technologies and Platforms
One of the challenges of single-cell sequencing is the capture of the single cells. Major technological improvement in the area of microfluidics has facilitated the capturing process. Microfluidic technology involves capturing of individual or population of cells in emulsion droplets in the picoliter or nanoliter range that is pumped in by different speeds using microfluidic devices [89]. The cell droplets constitution is compatible with mammalian cells and as a result it keeps cells functional for further investigation [90]. Cells are barcoded via specific primers and cDNAs are generated from thousands of cells in parallel [91, 92]. The cDNAs formed are pooled by breakage of the microfluid droplet, amplified by transcription and then sequencing libraries are prepared [93]. Microfluidic technologies are currently being used widely in the space of TCR enrichment sequencing [94]. The advantages that the process provides consist of providing T cell activation information, pathogens targeted by cytotoxic T cells and cytokine release [95,96,97]. Not only does it provide real-time information about the T cell repertoire, but it also provides the functional monitoring of the individual T cell types as opposed to the conventional methods [70, 98] such as identifying cell populations via FACS, carrying out PCR and then mass population sequencing that leads to loss of diversity [98, 99]. Chokkalingam et al. have used droplet technology combined with flow cytometry to identify T cell populations that secrete IL-2, IFNγ, and TNF-α [100]. Konry et al. have characterized T cells, natural killer cells, and antigen presenting complexes by multiplexing different parameters that are found in a single droplet. McDaniel et al. have proven that the pairing of both TCR chains (αβ or γδ) can be kept intact by single-cell sequencing by carrying out high-throughput sequencing on millions of T cell populations by obtaining individual chain sequences [101]. Further Segaliny et al. demonstrated the dynamics of TCR T cell interactions and downstream molecular analysis using single-cell sequences that confirmed their matching with target antigens with an exact specificity [102].
Droplet-based microfluidics has led to rapid discovery and has provided new tools for learning single-cell biology that include being a high-throughput technology [89, 91], providing insight to sequencing [103,104,105,106,107,108], secretome analysis [95, 96, 109], cell–cell interactions [97, 110], protein and antibody engineering [111], repertoire analysis and multiplex biological interactions [112]. Furthermore, microfluidics allows for analysis of drug interactions [113], antigen-specific T cell repertoires [97, 104, 114], CAR-T cell populations [115] and facilitates investigation of other protein secretion, e.g., antibodies [111, 116,117,118]. Microfluidics uses in-situ PCR and can display libraries in the same run simultaneously [119,120,121]. Platforms like the Chromium Single Cell Immune Profiling Solution by 10× Genomics can provide pertinent information on TCR, BCR, cell surface protein expression, and RNA expression in the same cells [122]. In addition, the small volume provides increased sensitivity, decreases time and it improves the signal to noise ratio [123]. As a result, it provides an insight into complex cellular heterogeneity instead of an average readout and provides information on molecular details that are essential for T cell activation. Not only can real-time sequencing be done with control of space time and environmental factors but it does not compromise the screening sensitivity [123]. There are multiple studies that have been conducted to understand multiplex biological responses, CDR3 CAR-T cell products, and adoptive T cells that can be used in varied therapeutic platforms, and provide real-time data for analysis by microfluidics [70].
3.4.2 Generation of TCR Sequences Using scRNA Sequencing
Single-cell TCR sequencing typically requires a distinct four step process. First, the isolation of single cells. This process is often performed using FACS sorting to isolate the target single cells. As discussed, microfluidic devices can be used to sequester single cells [86, 124]. Secondly, cells are isolated and the RNA is released into a fluid medium post lysis of the cell and Oligo dt primers recognize polyadenylated regions which are amplified as a result of selective reverse transcription. In the third step, the complimentary DNA molecule that is obtained from this process is amplified to produce multiple replicons, and then sequenced. Lastly, the output sequence reads from the reactions are analyzed and quantified [125]. The reference genome and transcriptome of interest can be traced back by comparing the sequence read subset to a reference genome or transcriptome [86, 124,125,126]. A shift has been observed in the single-cell sequencing landscape, as previously used capillary-based techniques are being replaced by emulsion-based PCR techniques. The methodology is improved as devices use technology where oil-in-water emulsions are pumped and high sequence read repeats are analyzed for individual cells. This provides a detailed insight on the αβ TCR chains as the mRNA coding them is released into the individual droplet that acts as an individual reaction platform. Within the droplet, the mRNA is amplified using a multiplex PCR-based approach [70] (Fig. 3.4).
3.4.3 Single-Cell Sequencing Data
Multiplex PCR systems and high-throughput systems have been used in the past for analyzing the TCR sequences. The protocol as described above uses isolation, amplification, library preparation, and sequencing for TCR α and β chains [79, 127]. There are two fundamental strategies: barcoding and full-length complimentary DNA sequencing. PCR has been used in the process of single-cell barcoding, a method of identification of cell subtypes. A barcode is a unique nucleotide sequence obtained from a single cell that tags cell transcripts and allows tracing the RNA back to its origin. This once linked to the TCR can help in identifying individual T cell types and subsets with distinct functions [128]. Full-length sequencing on the other hand involves pooling the cDNA obtained from each cell and then making a single-cell library for comparison of the sequence. This process is time consuming and expensive but sensitive and can provide information about changes in individual base pairs, mutations in the CDR3 region, difference in isoforms, and differences in α and β chain pairing [98]. The SMART-seq protocol that was developed by Stubbington was one of the first tools developed to reconstruct paired TCR α and β chains. Named TraCer, it was used to validate the sequences of the TCR αβ chains. The same cDNA was used for sequencing libraries and this validated the experimental approach as the start was the same PCR-based approach and the result a comparison for the chain sequences. This procedure has given an impetus to the requirement of efficiency in the approach [98]. TRAPes was another tool that was created to read short-read single-cell RNA-sequence libraries [129]. Other methods such as the non-strand specific poly-A tailing described by Tang et al. showed a weak positional bias for the 3′ end of the RNA [105]. The CEL-seq and MARS-seq that are similar techniques are also based on the principle of in-vitro transcription and having the same strong 3′ positional bias [130]. Quartz-seq a platform based on Poly-A tailing is able to provide low depth sequencing accurate results as it is one of the few that use a non-specific 3′ weak positional bias [131]. Further, platforms like the VDJ puzzle allow the analysis of T cell gene expression and TCR diversity, which consist of a library formation and then comparison of the sequences to the antigen-specific circulating CD8 T cells for additional validation [132].
3.4.4 An Application of Single-Cell TCR Sequencing
It has always been the goal of TCR research to identify the T cells at the genetic and biological levels simultaneously; it is an ongoing quest to understand the individuality of pathogenic or protective T cells in diseases. The challenges often encountered include the insufficiently small size of the human patients’ biopsies and the near impossibility of simultaneously examining biological functions of individual live cells isolated from the glands and their genetic profiles. Numerous attempts have been made using techniques such as laser capture micro-dissection or flow cytometric sorting to obtain single-cell populations. However, both techniques are cumbersome and neither can satisfactorily ensure single-cell analysis. Furthermore, the integrity of such isolated cells is greatly compromised due to the vigorous isolation process, which can affect subsequent micromanipulation and even functions of infiltrating T cells. Due to the fact that the purified cells are often either not viable or damaged, important information regarding the biological function(s) is difficult or impossible to obtain. To circumvent these challenges, we utilize the single-cell microengraving technology which was developed originally by the Love group at MIT as a tool for identification, profiling, and selection of single cells, specifically based on their reactivity to or secretion of proteins [118, 133,134,135]. The nanochip is fabricated via soft lithography, where arrays of nanowells are manufactured onto a glass slide consisting of 24 × 72 blocks containing 7 × 7 arranged nanowells within each block, yielding 84,672 50 μm nanowells per array. Additionally, these can be manufactured to produce 30 μm nanowells, where the arrays consist of 248,832 nanowells. Sylgard 184 silicone elastomer base (polydimethyl-siloxane, PDMS) is cured in a custom-built aluminum mold to produce these microchips. After cells are stained for desired cell surface markers with fluorescently conjugated antibodies and/or calcein (a live/dead marker), the cells are distributed on the microchip. As shown in Fig. 3.5, they can be analyzed by fluorescent microscopy, where images are taken of the 7 × 7 grid of nanowells to determine the identity of the cells. Next, the microchip is hybridized with a capture slide. This is a poly-l-Lysine slide coated with a desired protein or an antibody against a desired protein. For example, for identification of T helper cells, one would coat the slide with anti-IFNγ and anti-IL-17. In combination with the data from the fluorescent microscopy, identification of Th1, Th2, and Th17 cells would be possible. Alternatively, to isolate reactive antibodies, one would coat the capture slide with proteins of interest in order to identify B cell reactivity. Notably, in the case of identifying cells based on secreted proteins or antibodies, single-cell microengraving is advantageous over comparable techniques, such as flow, because the cells remain active during the process and do not need to be permeabilized, such as in the case of flow cytometry. After hybridization, the capture slide is exposed to detection (fluorescent) antibodies and/or proteins. The capture slide can be analyzed on a microarray scanner and the array data is then aligned against the fluorescent microscopy data to give a complete profile of the cells.
Single-cell microengraving technology. The fabricated nanowells were loaded with lymphocytes obtained from biopsies and were imaged using an automated epifluorescence microscope to cell populations based on surface markers. Hybridized nanowells with capture slides containing anti-human immunoglobulins against IL-17 and IFNγ for Th17 and Th1 cells as an example were analyzed. Post incubation, nanowells containing intact live cells and capture slides were separated and antibody-specific fluorochrome-conjugated antibodies were used detection antibodies. Lastly, micrographs of microarrays were generated by scanning using a GenePix Autoloader 4200AL microarray scanner
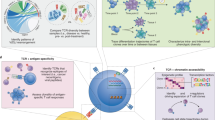
One notable application of this approach is the area of Sjogren’s syndrome (SjS) in which we sought to determine the TCR repertoires of pathogenic T cells that infiltrate the salivary glands, the main targeted tissue in SjS. Other groups have been trying to sequence pathogenic T cells for well over two decades utilizing a variety of techniques [136,137,138,139,140,141]. Commonly, the final technique involves nested PCRs to identify the TCR sequences. Single-cell sorting by flow is the most common technique to isolate T cells prior to this [142,143,144,145]. As previously mentioned, the pitfall of this technique is the lack of specific subset identification of the T cells. The one measure to counteract this is to permeabilize the membrane, killing the cells. Currently, there is no way to identify intracellular antigens while keeping the cells alive. Recently, single-cell microengraving has been successfully utilized in this effort [146]. As outlined in Fig. 3.6, cells presenting the correct phenotypes were selected, e.g., live Th1, Th17 cells, from the biopsies of SjS patients and sicca controls (patients who present a similar disease phenotype, but who do not meet the criteria of SjS). After the cells were lysed to retrieve RNA, RT-PCR was performed, immediately followed by nested PCR for the alpha and beta TCR chains and those products were sequenced. These sequences were then aligned against databases for identification of VDJ regions and compared between the subject and control groups as well as the subset of cells to identify unique subsets presenting specific TCRs, implying a shared antigen. Our recent study has identified that glandular Th1 and Th17 cells of control and SjS patients expressed common TCRβ variable (TRBV)3-1 and TRBV20, whereas TCRα variable (TRAV)8-2 was uniquely expressed by Th1 of SjS patients [146]. Using the SjS animal model, we have shown that salivary Th1 cells of male mice selected for TRAV8 and TRBV16 in Th1 and Th17 cells, whereas female Th1 cells selected for TRAV8, TRAV13D-2, and TRBV23 [147]. Other studies attest to our findings by ascertaining unique glandular TCRs in the human and animal models of SjS [145, 148]. Our seminal studies clearly imply that the clonal expansion of the effector T cells with the conserved TCRs is driven by SG cell antigens, and autoimmune responses to SG cell autoantigens evidence a specific loss of immunological self-tolerance.
An example of T helper cell profiling using single-cell microengraving technology. Representation of cells (bright field [BF]) as observed in the process of microengraving in nanowells labeled with calcein (indicating live cells). Micrographs of matching microarray by microengraving show detection signals for CD3, CD4, CD8, IFNγ, and IL-17A responsive cells as represented in the demonstration
3.4.5 Advantages and Limitations of Single-Cell Sequencing
A variety of bulk sequencing measurements give insight about large, mixed populations of cells. ELISA, PCR, and transcriptional profiling provide information about rapidly multiplying cells [149]. The bulk measurements stated above do not provide input on unique subsets of cells such as clonally expanded B cells and T cells. The parameters tested in bulk measurements average out phenotypic states of cells and give a generic outlook of the basic interactions among cells [150]. TCR profiling includes analyzing the transcriptome that is generated by the RNA. While offering many advantages, there are gaps in information which are unaccounted for in single-cell sequencing techniques. Some include accountability for stress conditions in the surrounding environment that affect individual cells. Cells may lie in varied cycle phases and each individual unit may display a variation in gene expression, size, and RNA content. Furthermore, every run in the single-cell sequencing process requires a large cell population harvested from fresh material, and it is expensive and covers limited cell populations [151]. Hence, despite the accuracy the variation that arises is mostly a consequence of all these factors. This aspect is not addressed clearly in most commercially available platforms available for TCR analysis [88, 92]. To combat these issues, there is a requirement of unprejudiced and reproducible gene expression patterns in single cells [152]. Seeing the varied factors that influence TCR analysis, there are different computational approaches that use variable models to account for variability in results and provide a better insight to the data obtained [92].
3.5 Conclusion
Significant technological advancement has catapulted the field of single-cell TCR sequencing. The remaining unanswered quests are how fast the field of bioinformatics will evolve to analyze and manage the large data generated. Recent interests in systemic biology of single T cells involve identification of the T cell subsets based on surface markers, gene expression at the single-cell transcriptome, and single-cell TCR expression. Our ability to comprehend and organize this set of new data will be limited if we are not able to organize the data in a biologically meaningful way. The second aspect that we must grapple with is its usefulness in antigen discovery. The current approach in examining and testing the antigen specificity of individual T cell clone is labor intensive using vast peptide-MHC libraries under culture conditions mimicking the in-vivo environment. In order to improve diagnosis and define therapy, specifically personalized medicine, it is imperative that we can identify and test the antigen specificity based on the single-cell TCRs. Emerging single-cell technologies bring promising discoveries but there are challenges that need to be addressed in order to fully comprehend the TCR repertoires and its application.
References
Chothia C, Boswell DR, Lesk AM (1988) The outline structure of the T-cell alpha beta receptor. EMBO J 7:3745–3755
Davis MM, Bjorkman PJ (1988) T-cell antigen receptor genes and T-cell recognition. Nature 334:395
Smith-Garvin JE, Koretzky GA, Jordan MS (2009) T cell activation. Annu Rev Immunol 27:591–619. https://doi.org/10.1146/annurev.immunol.021908.132706
Rudolph MG, Stanfield RL, Wilson IA (2006) How TCRs bind MHCs, peptides, and coreceptors. Annu Rev Immunol 24:419–466. https://doi.org/10.1146/annurev.immunol.23.021704.115658
Claverie J, Prochnickachalufour A, Bougueleret L (1989) Implications of a Fab-like structure for the T-cell receptor. Immunol Today 10:10–14. https://doi.org/10.1016/0167-5699(89)90058-3
Kaufmann SH (1996) gamma/delta and other unconventional T lymphocytes: what do they see and what do they do? Proc Natl Acad Sci U S A 93:2272–2279. https://doi.org/10.1073/pnas.93.6.2272
Hayday AC (2000) [gamma][delta] cells: a right time and a right place for a conserved third way of protection. Annu Rev Immunol 18:975–1026. https://doi.org/10.1146/annurev.immunol.18.1.975
Holtmeier W, Kabelitz D (2005) gammadelta T cells link innate and adaptive immune responses. Chem Immunol Allerg 86:151–183. https://doi.org/10.1159/000086659
Arstila TP et al (1999) A direct estimate of the human alphabeta T cell receptor diversity. Science 286:958–961. https://doi.org/10.1126/science.286.5441.958
van Dongen JJ, Wolvers-Tettero IL (1991) Analysis of immunoglobulin and T cell receptor genes. Part I: Basic and technical aspects. Clin Chim Acta 198:1–91. https://doi.org/10.1016/0009-8981(91)90246-9
Pannetier C et al (1993) The sizes of the CDR3 hypervariable regions of the murine T-cell receptor beta chains vary as a function of the recombined germ-line segments. Proc Natl Acad Sci U S A 90:4319–4323. https://doi.org/10.1073/pnas.90.9.4319
Cabaniols JP, Fazilleau N, Casrouge A, Kourilsky P, Kanellopoulos JM (2001) Most alpha/beta T cell receptor diversity is due to terminal deoxynucleotidyl transferase. J Exp Med 194:1385–1390. https://doi.org/10.1084/jem.194.9.1385
Sadofsky MJ (2001) The RAG proteins in V(D)J recombination: more than just a nuclease. Nucleic Acids Res 29:1399–1409. https://doi.org/10.1093/nar/29.7.1399
Borg NA et al (2005) The CDR3 regions of an immunodominant T cell receptor dictate the ‘energetic landscape’ of peptide-MHC recognition. Nat Immunol 6:171–180. https://doi.org/10.1038/ni1155
Stern LJ, Wiley DC (1994) Antigenic peptide binding by class I and class II histocompatibility proteins. Structure 2:245–251
Sim BC, Zerva L, Greene MI, Gascoigne NR (1996) Control of MHC restriction by TCR Valpha CDR1 and CDR2. Science 273:963–966. https://doi.org/10.1126/science.273.5277.963
Fremont DH, Matsumura M, Stura EA, Peterson PA, Wilson IA (1992) Crystal structures of two viral peptides in complex with murine MHC class I H-2Kb. Science 257:919–927. https://doi.org/10.1126/science.1323877
Madden DR, Garboczi DN, Wiley DC (1993) The antigenic identity of peptide-MHC complexes: a comparison of the conformations of five viral peptides presented by HLA-A2. Cell 75:693–708. https://doi.org/10.1016/0092-8674(93)90490-h
Janeway CA, Bottomly K (1994) Signals and signs for lymphocyte responses. Cell 76:275–285. https://doi.org/10.1016/0092-8674(94)90335-2
June CH, Bluestone JA, Nadler LM, Thompson CB (1994) The B7 and CD28 receptor families. Immunol Today 15:321–331. https://doi.org/10.1016/0167-5699(94)90080-9
Skapenko A, Leipe J, Lipsky PE, Schulze-Koops H (2005) The role of the T cell in autoimmune inflammation. Arthritis Res Ther 7:S4. https://doi.org/10.1186/ar1703
Sakaguchi S, Sakaguchi N, Asano M, Itoh M, Toda M (1995) Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains (CD25). Breakdown of a single mechanism of self-tolerance causes various autoimmune diseases. J Immunol 155:1151–1164
Bouneaud C, Kourilsky P, Bousso P (2000) Impact of negative selection on the T cell repertoire reactive to a self-peptide: a large fraction of T cell clones escapes clonal deletion. Immunity 13:829–840. https://doi.org/10.1016/S1074-7613(00)00080-7
Liblau R et al (1991) T cell response to myelin basic protein epitopes in multiple sclerosis patients and healthy subjects. Eur J Immunol 21:1391–1395. https://doi.org/10.1002/eji.1830210610
Rosenblum MD, Remedios KA, Abbas AK (2015) Mechanisms of human autoimmunity. J Clin Invest 125:2228–2233. https://doi.org/10.1172/JCI78088
Mills KH (2011) TLR-dependent T cell activation in autoimmunity. Nat Rev Immunol 11:807–822. https://doi.org/10.1038/nri3095
Itoh N et al (1993) Mononuclear cell infiltration and its relation to the expression of major histocompatibility complex antigens and adhesion molecules in pancreas biopsy specimens from newly diagnosed insulin-dependent diabetes mellitus patients. J Clin Invest 92:2313–2322. https://doi.org/10.1172/JCI116835
Varela-Calvino R, Calviño-Sampedro C, Gómez-Touriño I, Cordero OJ (2017) Apportioning blame: autoreactive CD4. Arch Immunol Ther Exp 65:275–284. https://doi.org/10.1007/s00005-016-0452-4
Tzartos JS et al (2008) Interleukin-17 production in central nervous system-infiltrating T cells and glial cells is associated with active disease in multiple sclerosis. Am J Pathol 172:146–155. https://doi.org/10.2353/ajpath.2008.070690
Kebir H et al (2007) Human TH17 lymphocytes promote blood-brain barrier disruption and central nervous system inflammation. Nat Med 13:1173–1175. https://doi.org/10.1038/nm1651
Compston A, Coles A (2008) Multiple sclerosis. Lancet 372:1502–1517. https://doi.org/10.1016/S0140-6736(08)61620-7
Kryczek I et al (2008) Induction of IL-17+ T cell trafficking and development by IFN-gamma: mechanism and pathological relevance in psoriasis. J Immunol 181:4733–4741. https://doi.org/10.4049/jimmunol.181.7.4733
Cai Y, Fleming C, Yan J (2012) New insights of T cells in the pathogenesis of psoriasis. Cell Mol Immunol 9:302–309. https://doi.org/10.1038/cmi.2012.15
Karczewski J, Dobrowolska A, Rychlewska-Hańczewska A, Adamski Z (2016) New insights into the role of T cells in pathogenesis of psoriasis and psoriatic arthritis. Autoimmunity 49:435–450. https://doi.org/10.3109/08916934.2016.1166214
Rosenberg WM, Moss PA, Bell JI (1992) Variation in human T cell receptor V beta and J beta repertoire: analysis using anchor polymerase chain reaction. Eur J Immunol 22:541–549. https://doi.org/10.1002/eji.1830220237
Moss PA, Rosenberg WM, Zintzaras E, Bell JI (1993) Characterization of the human T cell receptor alpha-chain repertoire and demonstration of a genetic influence on V alpha usage. Eur J Immunol 23:1153–1159. https://doi.org/10.1002/eji.1830230526
Moss PA, Bell JI (1995) Sequence analysis of the human alpha beta T-cell receptor CDR3 region. Immunogenetics 42:10–18
Moss PA, Bell JI (1996) Comparative sequence analysis of the human T cell receptor TCRA and TCRB CDR3 regions. Hum Immunol 48:32–38
Boudinot P et al (2008) New perspectives for large-scale repertoire analysis of immune receptors. Mol Immunol 45:2437–2445. https://doi.org/10.1016/j.molimm.2007.12.018
Pannetier C, Delassus S, Darche S, Saucier C, Kourilsky P (1993) Quantitative titration of nucleic acids by enzymatic amplification reactions run to saturation. Nucleic Acids Res 21:577–583. https://doi.org/10.1093/nar/21.3.577
Choi YW et al (1989) Interaction of Staphylococcus aureus toxin “superantigens” with human T cells. Proc Natl Acad Sci U S A 86:8941–8945. https://doi.org/10.1073/pnas.86.22.8941
Genevée C et al (1992) An experimentally validated panel of subfamily-specific oligonucleotide primers (V alpha 1-w29/V beta 1-w24) for the study of human T cell receptor variable V gene segment usage by polymerase chain reaction. Eur J Immunol 22:1261–1269. https://doi.org/10.1002/eji.1830220522
Lehner PJ et al (1995) Human HLA-A0201-restricted cytotoxic T lymphocyte recognition of influenza A is dominated by T cells bearing the V beta 17 gene segment. J Exp Med 181:79–91. https://doi.org/10.1084/jem.181.1.79
Pantaleo G et al (1994) Major expansion of CD8+ T cells with a predominant V beta usage during the primary immune response to HIV. Nature 370:463–467. https://doi.org/10.1038/370463a0
Pannetier C, Even J, Kourilsky P (1995) T-cell repertoire diversity and clonal expansions in normal and clinical samples. Immunol Today 16:176–181
Babel N, Stervbo U, Reinke P, Volk HD (2019) The identity card of T cells-clinical utility of T-cell receptor repertoire analysis in transplantation. Transplantation 103:1544–1555. https://doi.org/10.1097/TP.0000000000002776
Cochet M et al (1992) Molecular detection and in vivo analysis of the specific T cell response to a protein antigen. Eur J Immunol 22:2639–2647. https://doi.org/10.1002/eji.1830221025
Currier JR, Robinson MA (2001) Spectratype/immunoscope analysis of the expressed TCR repertoire. Curr Protoc Immunol Chapter 10:Unit 10.28. https://doi.org/10.1002/0471142735.im1028s38
Chaara W et al (2018) RepSeq data representativeness and robustness assessment by Shannon entropy. Front Immunol 9:1038. https://doi.org/10.3389/fimmu.2018.01038
Gallard A, Foucras G, Coureau C, Guéry JC (2002) Tracking T cell clonotypes in complex T lymphocyte populations by real-time quantitative PCR using fluorogenic complementarity-determining region-3-specific probes. J Immunol Methods 270:269–280. https://doi.org/10.1016/s0022-1759(02)00336-8
Six A et al (2013) The past, present, and future of immune repertoire biology - the rise of next-generation repertoire analysis. Front Immunol 4:413. https://doi.org/10.3389/fimmu.2013.00413
Casrouge A et al (2000) Size estimate of the alpha beta TCR repertoire of naive mouse splenocytes. J Immunol 164:5782–5787. https://doi.org/10.4049/jimmunol.164.11.5782
Seo KS, Park JY, Terman DS, Bohach GA (2010) A quantitative real time PCR method to analyze T cell receptor Vbeta subgroup expansion by staphylococcal superantigens. J Transl Med 8:2. https://doi.org/10.1186/1479-5876-8-2
Wettstein P, Strausbauch M, Therneau T, Borson N (2008) The application of real-time PCR to the analysis of T cell repertoires. Nucleic Acids Res 36:e140. https://doi.org/10.1093/nar/gkn634
Manfras BJ, Rudert WA, Trucco M, Boehm BO (1997) Analysis of the alpha/beta T-cell receptor repertoire by competitive and quantitative family-specific PCR with exogenous standards and high resolution fluorescence based CDR3 size imaging. J Immunol Methods 210:235–249. https://doi.org/10.1016/s0022-1759(97)00197-x
VanderBorght A et al (1999) Identification of overrepresented T cell receptor genes in blood and tissue biopsies by PCR-ELISA. J Immunol Methods 223:47–61. https://doi.org/10.1016/s0022-1759(98)00201-4
Lim A et al (2002) Combination of MHC-peptide multimer-based T cell sorting with the Immunoscope permits sensitive ex vivo quantitation and follow-up of human CD8+ T cell immune responses. J Immunol Methods 261:177–194. https://doi.org/10.1016/s0022-1759(02)00004-2
Calis JJ, Rosenberg BR (2014) Characterizing immune repertoires by high throughput sequencing: strategies and applications. Trends Immunol 35:581–590. https://doi.org/10.1016/j.it.2014.09.004
Woodsworth DJ, Castellarin M, Holt RA (2013) Sequence analysis of T-cell repertoires in health and disease. Genome Med 5:98. https://doi.org/10.1186/gm502
Clemente MJ et al (2013) Deep sequencing of the T-cell receptor repertoire in CD8+ T-large granular lymphocyte leukemia identifies signature landscapes. Blood 122:4077–4085. https://doi.org/10.1182/blood-2013-05-506386
Lynch JN, Donermeyer DL, Weber KS, Kranz DM, Allen PM (2013) Subtle changes in TCRα CDR1 profoundly increase the sensitivity of CD4 T cells. Mol Immunol 53:283–294. https://doi.org/10.1016/j.molimm.2012.08.020
Chlewicki LK, Holler PD, Monti BC, Clutter MR, Kranz DM (2005) High-affinity, peptide-specific T cell receptors can be generated by mutations in CDR1, CDR2 or CDR3. J Mol Biol 346:223–239. https://doi.org/10.1016/j.jmb.2004.11.057
Rosati E et al (2017) Overview of methodologies for T-cell receptor repertoire analysis. BMC Biotechnol 17:61. https://doi.org/10.1186/s12896-017-0379-9
Lin C (2016) Analysis of TCR repertoires for tracking and evaluating distinct T cell subsets. Stem Cell Investig 3:12. https://doi.org/10.21037/sci.2016.04.01
Correia-Neves M, Waltzinger C, Mathis D, Benoist C (2001) The shaping of the T cell repertoire. Immunity 14:21–32
Sant’Angelo DB et al (1998) A molecular map of T cell development. Immunity 9:179–186
Kirsch I, Vignali M, Robins H (2015) T-cell receptor profiling in cancer. Mol Oncol 9:2063–2070. https://doi.org/10.1016/j.molonc.2015.09.003
Ma Y, Shi N, Li M, Chen F, Niu H (2015) Applications of next-generation sequencing in systemic autoimmune diseases. Genom Proteom Bioinformatics 13:242–249. https://doi.org/10.1016/j.gpb.2015.09.004
Okino ST, Kong M, Sarras H, Wang Y (2016) Evaluation of bias associated with high-multiplex, target-specific pre-amplification. Biomol Detect Quantif 6:13–21. https://doi.org/10.1016/j.bdq.2015.12.001
De Simone M, Rossetti G, Pagani M (2018) Single cell T cell receptor sequencing: techniques and future challenges. Front Immunol 9:1638. https://doi.org/10.3389/fimmu.2018.01638
Wu SG et al (2019) High throughput sequencing of T-cell receptor repertoire using dry blood spots. J Transl Med 17:47. https://doi.org/10.1186/s12967-019-1796-4
Hou X et al (2016) High throughput sequencing of T cell antigen receptors reveals a conserved TCR repertoire. Medicine 95:e2839. https://doi.org/10.1097/MD.0000000000002839
Rytlewski J et al (2019) Model to improve specificity for identification of clinically-relevant expanded T cells in peripheral blood. PLoS One 14:e0213684. https://doi.org/10.1371/journal.pone.0213684
Kivioja T et al (2011) Counting absolute numbers of molecules using unique molecular identifiers. Nat Methods 9:72–74. https://doi.org/10.1038/nmeth.1778
Bolotin DA et al (2012) Next generation sequencing for TCR repertoire profiling: platform-specific features and correction algorithms. Eur J Immunol 42:3073–3083. https://doi.org/10.1002/eji.201242517
Head SR et al (2014) Library construction for next-generation sequencing: overviews and challenges. Biotechniques 56:61–64., 66, 68, passim. https://doi.org/10.2144/000114133
Reuter JA, Spacek DV, Snyder MP (2015) High-throughput sequencing technologies. Mol Cell 58:586–597. https://doi.org/10.1016/j.molcel.2015.05.004
Muraro PA et al (2014) T cell repertoire following autologous stem cell transplantation for multiple sclerosis. J Clin Invest 124:1168–1172. https://doi.org/10.1172/JCI71691
Han A, Glanville J, Hansmann L, Davis MM (2014) Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat Biotechnol 32:684–692. https://doi.org/10.1038/nbt.2938
Bai X et al (2015) Characteristics of tumor infiltrating lymphocyte and circulating lymphocyte repertoires in pancreatic cancer by the sequencing of T cell receptors. Sci Rep 5:13664. https://doi.org/10.1038/srep13664
Gros A et al (2014) PD-1 identifies the patient-specific CD8+ tumor-reactive repertoire infiltrating human tumors. J Clin Invest 124:2246–2259. https://doi.org/10.1172/JCI73639
Chattopadhyay PK, Gierahn TM, Roederer M, Love JC (2014) Single-cell technologies for monitoring immune systems. Nat Immunol 15:128–135. https://doi.org/10.1038/ni.2796
Nagafuchi Y, Shoda H, Fujio K (2019) Immune profiling and precision medicine in systemic lupus erythematosus. Cell 8:140. https://doi.org/10.3390/cells8020140
Miconnet I (2012) Probing the T-cell receptor repertoire with deep sequencing. Curr Opin HIV AIDS 7:64–70. https://doi.org/10.1097/COH.0b013e32834ddcae
Newell EW, Davis MM (2014) Beyond model antigens: high-dimensional methods for the analysis of antigen-specific T cells. Nat Biotechnol 32:149–157. https://doi.org/10.1038/nbt.2783
Kiselev VY, Andrews TS, Hemberg M (2019) Challenges in unsupervised clustering of single-cell RNA-seq data. Nat Rev Genet 20:273–282. https://doi.org/10.1038/s41576-018-0088-9
Byrne A et al (2017) Nanopore long-read RNAseq reveals widespread transcriptional variation among the surface receptors of individual B cells. Nat Commun 8:16027. https://doi.org/10.1038/ncomms16027
Stegle O, Teichmann SA, Marioni JC (2015) Computational and analytical challenges in single-cell transcriptomics. Nat Rev Genet 16:133–145. https://doi.org/10.1038/nrg3833
Shembekar N, Chaipan C, Utharala R, Merten CA (2016) Droplet-based microfluidics in drug discovery, transcriptomics and high-throughput molecular genetics. Lab Chip 16:1314–1331. https://doi.org/10.1039/c6lc00249h
Zykwinska A, Marquis M, Sinquin C, Cuenot S, Colliec-Jouault S (2016) Assembly of HE800 exopolysaccharide produced by a deep-sea hydrothermal bacterium into microgels for protein delivery applications. Carbohydr Polym 142:213–221. https://doi.org/10.1016/j.carbpol.2016.01.056
Mazutis L et al (2013) Single-cell analysis and sorting using droplet-based microfluidics. Nat Protoc 8:870–891. https://doi.org/10.1038/nprot.2013.046
Buettner F et al (2015) Computational analysis of cell-to-cell heterogeneity in single-cell RNA-sequencing data reveals hidden subpopulations of cells. Nat Biotechnol 33:155–160. https://doi.org/10.1038/nbt.3102
Thomsen ER et al (2016) Fixed single-cell transcriptomic characterization of human radial glial diversity. Nat Methods 13:87–93. https://doi.org/10.1038/nmeth.3629
Gee MH et al (2018) Antigen identification for orphan T cell receptors expressed on tumor-infiltrating lymphocytes. Cell 172:549–563.e516. https://doi.org/10.1016/j.cell.2017.11.043
Konry T, Dominguez-Villar M, Baecher-Allan C, Hafler DA, Yarmush ML (2011) Droplet-based microfluidic platforms for single T cell secretion analysis of IL-10 cytokine. Biosens Bioelectron 26:2707–2710. https://doi.org/10.1016/j.bios.2010.09.006
Diercks AH et al (2009) A microfluidic device for multiplexed protein detection in nano-liter volumes. Anal Biochem 386:30–35. https://doi.org/10.1016/j.ab.2008.12.012
Sarkar S, Motwani V, Sabhachandani P, Cohen N, Konry T (2015) T cell dynamic activation and functional analysis in nanoliter droplet microarray. J Clin Cell Immunol 6:334. https://doi.org/10.4172/2155-9899.1000334
Stubbington MJT et al (2016) T cell fate and clonality inference from single-cell transcriptomes. Nat Methods 13:329–332. https://doi.org/10.1038/nmeth.3800
Staudt N, Müller-Sienerth N, Wright GJ (2014) Development of an antigen microarray for high throughput monoclonal antibody selection. Biochem Biophys Res Commun 445:785–790. https://doi.org/10.1016/j.bbrc.2013.12.033
Chokkalingam V et al (2013) Probing cellular heterogeneity in cytokine-secreting immune cells using droplet-based microfluidics. Lab Chip 13:4740–4744. https://doi.org/10.1039/c3lc50945a
McDaniel JR et al (2018) Identification of tumor-reactive B cells and systemic IgG in breast cancer based on clonal frequency in the sentinel lymph node. Cancer Immunol Immunother 67:729–738. https://doi.org/10.1007/s00262-018-2123-2
Segaliny AI et al (2018) Functional TCR T cell screening using single-cell droplet microfluidics. Lab Chip 18:3733–3749. https://doi.org/10.1039/c8lc00818c
Klein AM et al (2015) Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161:1187–1201. https://doi.org/10.1016/j.cell.2015.04.044
DeKosky BJ et al (2013) High-throughput sequencing of the paired human immunoglobulin heavy and light chain repertoire. Nat Biotechnol 31:166–169. https://doi.org/10.1038/nbt.2492
Tang F et al (2009) mRNA-seq whole-transcriptome analysis of a single cell. Nat Methods 6:377–382. https://doi.org/10.1038/nmeth.1315
Linnemann C et al (2013) High-throughput identification of antigen-specific TCRs by TCR gene capture. Nat Med 19:1534–1541. https://doi.org/10.1038/nm.3359
Chen B et al (2016) High-throughput analysis and protein engineering using microcapillary arrays. Nat Chem Biol 12:76–81. https://doi.org/10.1038/nchembio.1978
Macosko EZ et al (2015) Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161:1202–1214. https://doi.org/10.1016/j.cell.2015.05.002
Han Q et al (2012) Polyfunctional responses by human T cells result from sequential release of cytokines. Proc Natl Acad Sci U S A 109:1607–1612. https://doi.org/10.1073/pnas.1117194109
Yamanaka YJ et al (2012) Single-cell analysis of the dynamics and functional outcomes of interactions between human natural killer cells and target cells. Integr Biol 4:1175–1184. https://doi.org/10.1039/c2ib20167d
Fitzgerald V et al (2015) Exploiting highly ordered subnanoliter volume microcapillaries as microtools for the analysis of antibody producing cells. Anal Chem 87:997–1003. https://doi.org/10.1021/ac503547j
Junkin M et al (2016) High-content quantification of single-cell immune dynamics. Cell Rep 15:411–422. https://doi.org/10.1016/j.celrep.2016.03.033
Brouzes E et al (2009) Droplet microfluidic technology for single-cell high-throughput screening. Proc Natl Acad Sci U S A 106:14195–14200. https://doi.org/10.1073/pnas.0903542106
Pavesi A et al (2017) A 3D microfluidic model for preclinical evaluation of TCR-engineered T cells against solid tumors. JCI Insight 2:89762. https://doi.org/10.1172/jci.insight.89762
Xue Q et al (2017) Single-cell multiplexed cytokine profiling of CD19 CAR-T cells reveals a diverse landscape of polyfunctional antigen-specific response. J Immunother Cancer 5:85. https://doi.org/10.1186/s40425-017-0293-7
El Debs B, Utharala R, Balyasnikova IV, Griffiths AD, Merten CA (2012) Functional single-cell hybridoma screening using droplet-based microfluidics. Proc Natl Acad Sci U S A 109:11570–11575. https://doi.org/10.1073/pnas.1204514109
Sista RS et al (2008) Heterogeneous immunoassays using magnetic beads on a digital microfluidic platform. Lab Chip 8:2188–2196. https://doi.org/10.1039/b807855f
Love JC, Ronan JL, Grotenbreg GM, van der Veen AG, Ploegh HL (2006) A microengraving method for rapid selection of single cells producing antigen-specific antibodies. Nat Biotechnol 24:703–707. https://doi.org/10.1038/nbt1210
Adler AS et al (2017) Rare, high-affinity anti-pathogen antibodies from human repertoires, discovered using microfluidics and molecular genomics. MAbs 9:1282–1296. https://doi.org/10.1080/19420862.2017.1371383
Hatch AC et al (2011) 1-Million droplet array with wide-field fluorescence imaging for digital PCR. Lab Chip 11:3838–3845. https://doi.org/10.1039/c1lc20561g
Williams R et al (2006) Amplification of complex gene libraries by emulsion PCR. Nat Methods 3:545–550. https://doi.org/10.1038/nmeth896
Demaree B, Weisgerber D, Lan F, Abate AR (2018) An ultrahigh-throughput microfluidic platform for single-cell genome sequencing. J Vis Exp. https://doi.org/10.3791/57598
Eyer K et al (2017) Single-cell deep phenotyping of IgG-secreting cells for high-resolution immune monitoring. Nat Biotechnol 35:977–982. https://doi.org/10.1038/nbt.3964
Menon V (2018) Clustering single cells: a review of approaches on high-and low-depth single-cell RNA-seq data. Brief Funct Genom 17:240–245. https://doi.org/10.1093/bfgp/elx044
Poulin JF, Tasic B, Hjerling-Leffler J, Trimarchi JM, Awatramani R (2016) Disentangling neural cell diversity using single-cell transcriptomics. Nat Neurosci 19:1131–1141. https://doi.org/10.1038/nn.4366
Singh M et al (2019) High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes. Nat Commun 10:3120. https://doi.org/10.1038/s41467-019-11049-4
Kim SM et al (2012) Analysis of the paired TCR α- and β-chains of single human T cells. PLoS One 7:e37338. https://doi.org/10.1371/journal.pone.0037338
Howie B et al (2015) High-throughput pairing of T cell receptor α and β sequences. Sci Transl Med 7:301ra131. https://doi.org/10.1126/scitranslmed.aac5624
Afik S et al (2017) Targeted reconstruction of T cell receptor sequence from single cell RNA-seq links CDR3 length to T cell differentiation state. Nucleic Acids Res 45:e148. https://doi.org/10.1093/nar/gkx615
Hashimshony T, Wagner F, Sher N, Yanai I (2012) CEL-seq: single-cell RNA-seq by multiplexed linear amplification. Cell Rep 2:666–673. https://doi.org/10.1016/j.celrep.2012.08.003
Sasagawa Y et al (2017) Erratum to: Quartz-seq: a highly reproducible and sensitive single-cell RNA sequencing method, reveals non-genetic gene-expression heterogeneity. Genome Biol 18:9. https://doi.org/10.1186/s13059-017-1154-x
Eltahla AA et al (2016) Linking the T cell receptor to the single cell transcriptome in antigen-specific human T cells. Immunol Cell Biol 94:604–611. https://doi.org/10.1038/icb.2016.16
Love KR, Bagh S, Choi J, Love JC (2013) Microtools for single-cell analysis in biopharmaceutical development and manufacturing. Trends Biotechnol 31:280–286. https://doi.org/10.1016/j.tibtech.2013.03.001
Torres AJ, Hill AS, Love JC (2014) Nanowell-based immunoassays for measuring single-cell secretion: characterization of transport and surface binding. Anal Chem 86:11562–11569. https://doi.org/10.1021/ac4030297
Esfandiary L et al (2016) Single-cell antibody nanowells: a novel technology in detecting anti-SSA/Ro60- and anti-SSB/La autoantibody-producing cells in peripheral blood of rheumatic disease patients. Arthritis Res Ther 18:107. https://doi.org/10.1186/s13075-016-1010-5
Yonaha F et al (1992) Restricted junctional usage of T cell receptor V beta 2 and V beta 13 genes, which are overrepresented on infiltrating T cells in the lips of patients with Sjogren’s syndrome. Arthritis Rheum 35:1362–1367. https://doi.org/10.1002/art.1780351118
Sumida T et al (1992) T cell receptor repertoire of infiltrating T cells in lips of Sjogren’s syndrome patients. J Clin Invest 89:681–685. https://doi.org/10.1172/JCI115635
Freimark B, Pickering L, Concannon P, Fox R (1989) Nucleotide sequence of a uniquely expressed human T cell receptor beta chain variable region gene (V beta) in Sjogren’s syndrome. Nucleic Acids Res 17:455. https://doi.org/10.1093/nar/17.1.455
Matsumoto I et al (1996) Common T cell receptor clonotype in lacrimal glands and labial salivary glands from patients with Sjogren’s syndrome. J Clin Invest 97:1969–1977. https://doi.org/10.1172/JCI118629
Smith MD et al (1994) Selective expression of V beta families by T cells in the blood and salivary gland infiltrate of patients with primary Sjogren’s syndrome. J Rheumatol 21:1832–1837
Sumida T, Matsumoto I, Maeda T, Nishioka K (1997) T-cell receptor in Sjogren’s syndrome. Br J Rheumatol 36:622–629. https://doi.org/10.1093/rheumatology/36.6.622
Dash P et al (2011) Paired analysis of TCRalpha and TCRbeta chains at the single-cell level in mice. J Clin Invest 121:288–295. https://doi.org/10.1172/JCI44752
Kim SM et al (2012) Analysis of the paired TCR alpha- and beta-chains of single human T cells. PLoS One 7:e37338. https://doi.org/10.1371/journal.pone.0037338
Han A et al (2013) Dietary gluten triggers concomitant activation of CD4+ and CD8+ alphabeta T cells and gammadelta T cells in celiac disease. Proc Natl Acad Sci U S A 110:13073–13078. https://doi.org/10.1073/pnas.1311861110
Joachims ML et al (2016) Single-cell analysis of glandular T cell receptors in Sjogren’s syndrome. JCI Insight 1:e85609. https://doi.org/10.1172/jci.insight.85609
Voigt A et al (2018) Unique glandular ex-vivo Th1 and Th17 receptor motifs in Sjogren’s syndrome patients using single-cell analysis. Clin Immunol 192:58–67. https://doi.org/10.1016/j.clim.2018.04.009
Wanchoo A et al (2017) Single-cell analysis reveals sexually dimorphic repertoires of Interferon-gamma and IL-17A producing T cells in salivary glands of Sjogren’s syndrome mice. Sci Rep 7:12512. https://doi.org/10.1038/s41598-017-12627-6
Karabiyik A, Peck AB, Nguyen CQ (2013) The important role of T cells and receptor expression in Sjogren’s syndrome. Scand J Immunol 78:157–166. https://doi.org/10.1111/sji.12079
Mahony J et al (2007) Development of a respiratory virus panel test for detection of twenty human respiratory viruses by use of multiplex PCR and a fluid microbead-based assay. J Clin Microbiol 45:2965–2970. https://doi.org/10.1128/JCM.02436-06
Perfetto SP, Chattopadhyay PK, Roederer M (2004) Seventeen-colour flow cytometry: unravelling the immune system. Nat Rev Immunol 4:648–655. https://doi.org/10.1038/nri1416
Proserpio V, Mahata B (2016) Single-cell technologies to study the immune system. Immunology 147:133–140. https://doi.org/10.1111/imm.12553
Svensson V, Vento-Tormo R, Teichmann SA (2018) Exponential scaling of single-cell RNA-seq in the past decade. Nat Protoc 13:599–604. https://doi.org/10.1038/nprot.2017.149
Acknowledgements
CQN was supported financially in part by PHS grants DE023433, DE018958, DE028544 from the National Institutes of Health. RW is supported by the T90/R90 Comprehensive Training Program in Oral Biology (NIDCR, T90 DE21990). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Gupta, S., Witas, R., Voigt, A., Semenova, T., Nguyen, C.Q. (2020). Single-Cell Sequencing of T cell Receptors: A Perspective on the Technological Development and Translational Application. In: Yu, B., Zhang, J., Zeng, Y., Li, L., Wang, X. (eds) Single-cell Sequencing and Methylation. Advances in Experimental Medicine and Biology, vol 1255. Springer, Singapore. https://doi.org/10.1007/978-981-15-4494-1_3
Download citation
DOI: https://doi.org/10.1007/978-981-15-4494-1_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-4493-4
Online ISBN: 978-981-15-4494-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)