Abstract
Ectopic ossification of the spinal ligament and its progression compresses the spinal cord, resulting in serious neurological deficiencies. Surgery is an established therapy that has had considerable success, but it is often associated with a higher risk of re-progression of ossification followed by neurological complications. Because the detailed mechanism of the onset and development of ectopic ossification have not been clarified, pharmaceutical treatment for this disease has yet to be established. However, a safe and effective treatment is required to improve the quality of life of patients. The disease is thought to be multifactorial, being influenced by both genetic factors and several environmental factors such as mechanical stress. In this review, we discuss recent progress in disease research, with particular focus on what causes initial ossification and which type of abnormality occurs. Mesenchymal stem cells of patients with ossification are thought to misdifferentiate into osteoblasts instead of target ligament cells because they have high osteogenic ability. Elucidation of the genetic and epigenetic transformation of these cells in spinal ligament tissue is crucial to understanding the disease mechanism.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Spinal ligament
- Ectopic ossification
- Mesenchymal stem cells
- Mechanical stress
- Genetic change
- Genome-wide association study
- Epigenetic modification
- DNA methylation
1 Introduction
The spine is formed by a series of vertebrae. Multiple longitudinal ligaments connect individual vertebrae longitudinally to stabilize the spine and play an important role in assuring coordinated movement. Two of the spinal ligaments, the posterior longitudinal ligament (PLL) and the ligamentum flavum (LF) located on the anterior and posterior surfaces of the vertebral canal, respectively, are in contact with the spinal cord running longitudinally in the vertebral foramen. Ossification of the spinal ligament (OPLL and OLF) is a disorder in which the spinal ligament undergoes ectopic ossification. It is possible for ligament tissue to ossify to a small extent because of aging. However, when ossification exceeds a certain amount it compresses the spinal cord, which may lead to numbness of hands and feet and even paralysis [1,2,3,4].
There are no established effective preventive methods for the development of spinal ligament ossification, and no treatment option other than spinal surgery which is extremely invasive and burdens the patient [5]. There is also often the possibility of recurrence after surgery. Therefore, to improve the quality of life of patients, the establishment of safe and effective drug treatments is required. For this reason, the determination of disease etiology and pathogenesis is essential.
Because the disorder was found to be hereditary, its cause is presumed to involve genetic factors and the search for causative genes is ongoing [6,7,8,9,10]. Epidemiologically, endocrine and metabolic diseases are considered to be associated with the etiology of spinal ligament ossification [11]. Clinically, the onset and progress of ossification are likely to occur at the ligament site where mechanical stress is applied [11]. Therefore, the ossification of spinal ligaments is considered to be a multifactorial disease caused by a combination of various factors [10, 11]. To clarify the etiology of this disease, we returned to its origin and asked what entity causes the initial ossification and what type of abnormality occurs there. Once the mechanism of onset is clear, the treatment target should also become apparent.
2 What Type of Cells Ossify During Ectopic Ossification of the Spinal Ligament?
2.1 Ossification of Cells in Ligament Tissue
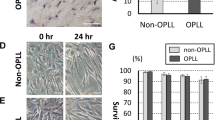
Advances in cell culture techniques enabled us to isolate cells from spinal ligament tissues and maintain them in vitro [13, 14]. Moreover, with informed patient consent, we also obtained spinal ligament tissue for research purposes that was assigned to be discarded after surgery [15]. From this tissue, ligaments can be digested with enzymes such as collagenase to isolate cells which are cultured to obtain fibroblast-like spinal ligament cells. Long-term culture of these cells, even in normal medium without passaging, results in their gradual calcification from the time point when confluency (uniform covering of cells over the entire surface of the culture dish) has passed. This can be used as an in vitro model system for the ossification of ligament tissue [13,14,15]. In ligament cells from patients with ossification, calcification progresses more extensively in response to culture medium used to induce ossification compared with cells from patients with no ossification such as those with cervical myelopathy (Fig. 7.1).
Calcification of spinal ligament cells from ossified and nonossified patients. Ligament cells prepared from ligament tissue of nonossified and ossified patients were cultured for more than 4 weeks, and mineral deposition (calcification) was detected by alizarin red S staining. Marked calcification occurred in cells derived from ossified patients 4 weeks after the initiation of culture
When cultured spinal ligament cells are exposed to various stresses such as mechanical stress in the form of repetitive stretch stimuli, which is presumed to be a trigger for ossification, the expression of various ossification-related genes is induced, promoting calcification of the cells [14,15,16,17,18] (Fig. 7.2).
Mechanical stress loading on cultured ligament cells. Ligament cells were cultured in an extensible chamber (upper panel) and subjected to uniaxial cyclic stretch at a constant cycle (120%, 0.5 Hz) to apply mechanical stress for indicated time periods. The production of the osteogenic marker BMP2 was analyzed by western blot (lower panel). β-actin was used as an internal standard of cell proteins. The level of BMP2 expression was enhanced in a time-dependent manner in ligament cells from patients with ossification but not in those from nonossified patients. The figure is reproduced with permission from Bone [13]
Furthermore, in cells from patients with ossification, the sensitivity of increased expression of ossification-related genes in response to mechanical stress was shown to be much higher than in cells derived from non-ossifying tissues [13,14,15,16,17,18]. This demonstrated that ligament cells derived from patients with ossification are more likely to be ossified at the isolated cell level, strongly suggesting the following: (1) the main body undergoing ossification is the cell itself which comprises ligament tissue, (2) patient-derived ligament cells undergo some transformation, and (3) the addition of various stresses and environmental factors to a pre-existing abnormality is likely to cause ossification. Based on these findings, we suggest that ligament cells are ossified in the process of repairing the damaged part of the ligament. This is supported by clinical findings that ossification is likely to occur at sites susceptible to mechanical stress, and that it often progresses after spinal surgery [12].
2.2 Mesenchymal Stem Cells as Ossifying Entities and their Localization in Ligament Tissue
Previous research used cells obtained by the digestion of ligament tissue with collagenase that were then cultured. Therefore, it is possible that a mixture of cell types had been obtained. Indeed, the calcification of cells in culture vessels is varied, and differences in calcification abilities can be observed among cells. Because the identification of cells with high ossification potentials would likely lead to an understanding of the mechanism of ossification, we hypothesized that cells responsible for the repair of tissue damage are present in ligament tissue but that they carry a mutation enabling them to ossify. Therefore, when tissue repair occurs they behave like osteoblasts rather than ligament cells.
First, we tried to prove that mesenchymal stem cells (MSCs) exist in ligament tissue. Using flow cytometry, we separated cells bearing MSC surface markers (CD73-, CD90-, and CD105-positive, and CD11b-, CD19-, CD34-, CD45-, and human leukocyte antigen-DR-negative) from a cell population obtained by enzymatic digestion of ligament tissue with collagenase [19, 20]. Next, MSCs in ligament tissue were purified by a cell sorter, and shown to have no significant difference in their proliferative potential between ossified and non-ossified patient tissues. Furthermore, MSCs from both tissue types had multilineage (osteogenic, chondrogenic, and adipogenic) differentiation potential. However, the osteogenic potential of MSCs from patients with ossification was significantly higher than those from patients without ossification [21].
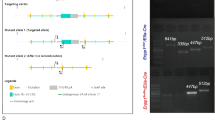
Subsequently, when the localization of MSCs was examined by immunohistochemical staining, pericytes around the blood vessels were found to express MSC markers. Additionally, while the tissue of non-ossified patients contained few blood vessels, that of ossified patients contained vascular tissue that extended around the ossification site to the ligament parenchyma. Moreover, MSCs were widely present around blood vessels in the ossification area [22], while hypertrophic chondrocyte-like cells, often found at the ossification junction, expressed MSC markers [19, 22]. This strongly suggested that MSCs participate in ectopic ossification of the ligament by endochondral ossification pathways including angiogenesis, in which changes in MSC localization play an important role. Such changes may involve the MSC regulatory factor stromal cell-derived factor (SDF)-1 and its receptor C-X-C motif chemokine receptor (CXCR)-4 [23]. Osteogenic ligament-derived MSCs showed high expression of SDF-1 and CXCR4 compared with healthy ligaments, together with enhanced motility of MSCs and chemotaxis to SDF-1. This motility and chemotaxis were suppressed by AMD3100, a specific inhibitor of CXCR4 [24], suggesting that MSCs migrating by the SDF-1/CXCR4 system cause ossification (Fig. 7.3).
SDF-1 and CXCR4 are expressed in spinal ligament tissue and control the migration activity of MSCs. (a) CXCR4 mRNA levels were analyzed by RT-PCR and its protein expression was evaluated by western blotting in MSCs isolated from ossified (OLF) and nonossified (nonOLF) patients. CXCR4 expression was normalized against β-actin as an internal standard. (b) The migration activity of MSCs was analyzed by a wound healing assay in the presence of AMD3100, a potent inhibitor of CXCR4. (c) MSC migration toward SDF-1 was measured in a chemotaxis chamber containing various concentrations of SDF-1. The figure is reproduced with permission from Journal of Pharmacology and Experimental Therapeutics [21]
2.3 Endochondral Ossification and MSCs
In the parenchyma of ossified ligament tissue, hypertrophic chondrocytes have been observed near the ossification front [19]. These cells express the chondrocyte-specific connective tissue growth factor [25]. Furthermore, those cells near the ossification front were observed to express MSC markers, indicating that they derive from MSCs [22].
An abundance of blood vessels was reported in the ossified ligament of ossified patients and the amount of blood loss during their spinal surgery was high [26]. This suggests that ligament ossification also follows the pathway of endochondral ossification, which is a normal mechanism of bone metabolism involving conversion to bone via cartilage through blood vessel invasion. We observed that chondromodulin-1, a factor expressed in cartilage tissue that suppresses osteogenesis and maintains cartilage tissue [27], is also expressed in MSCs of ligament tissue. Moreover, its expression was significantly reduced in MSCs from patients with ossification compared with those without ossification.
Considering that ligament ossification follows the process of endochondral ossification [27], it is conceivable that decreased expression of chondromodulin-1 permits vascular invasion and promotes ectopic ossification. This mechanism could be used as a target for drug treatment.
3 What Determines the Difference Between Healthy and Ossifying MSCs?
3.1 Genetic Mechanism
How did MSCs in the spinal ligament tissue of ossified patients acquire (transform) their high ossification capacity? Progressive ossifying fibroplasia (fibrodysplasia ossificans progressiva, FOP) is an example of a genetic disease of ectopic ossification caused by abnormal MSCs [28, 29]. This disease caused by a mutation in the activin receptor-like kinase-2 gene results in the transformation of cells derived from vascular endothelium into mesenchymal cells, leading to ossification. To identify candidate genes responsible for the genetic variation of spinal ligament ossification disease, genome-wide association studies have been carried out by Nakajima et al. [9, 30] and Takegami et al. [31]. They focused on R-spondin 2 as a susceptible gene for ectopic ossification of spinal ligaments. The R-spondin 2 protein is a key player in endochondral ossification that facilitates the differentiation of proliferating chondrocytes into hypertrophic chondrocytes. It is conceivable that its function is altered in patients with ossification, promoting ectopic ossification. However, it is unclear whether only one gene is responsible for disease, so the involvement of mutations in other genes will also need to be considered.
3.2 Epigenetic Mechanism
We are tackling this problem using an alternative approach of an epigenetic mechanism, in which transformation occurs by the modification of genomic DNA that is not caused by variation in the nucleic acid sequence. Changes in DNA methylation have been shown to be involved in the pathogenesis of several diseases [32]. For example, Montes et al. reported on the calcification of vascular smooth muscle as an example of ectopic calcification involving an epigenetic mechanism [33]. The calcification of blood vessels in chronic renal failure is presumed to be caused by decreased expression of the vascular smooth muscle specific protein SM22α, resulting in the transformation of vascular smooth muscle cells into osteoblasts. This study found that the decrease was caused by increased methylation of the SM22α transcriptional regulatory site. Several epigenetic mechanisms are known, but we focused on the regulation of expression by genomic DNA methylation. This involves a high density of cytosine–guanine nucleotide sequences (CpG) centered on the transcriptional regulatory region of the gene, and the suppression of gene expression by methylation of a CpG island.
The treatment of MSCs with an inhibitor of DNA methylation (5-aza-2′-deoxycytidine, 5AdC) to demethylate DNA resulted in marked changes in the expression of some osteogenic genes [34]. We found that MSCs derived from tissues of both ossified and non-ossified patients were demethylated by 5AdC treatment. Analysis of expression changes in the transcriptome by microarray enabled us to screen genes that are highly expressed through low DNA methylation in MSCs from patients with ossification, and those that are suppressed through high methylation in MSCs from non-ossified patients. This approach identified two candidate genes involved in controlling ossification-related genes: Wnt family member 5A (WNT5A) and glial cell line-derived neurotrophic factor (GDNF). Treatment of MSCs with small interfering RNA targeting each of these genes significantly suppressed their expression as well as that of osteogenic genes such as alkaline phosphatase, bone morphogenetic protein 2, and Runt-related transcription factor 2. From this, we presumed that WNT5A and GDNF are involved in the upstream regulation of these osteogenic genes. It also suggested that WNT5A and GDNF are not suppressed by methylation in MSCs derived from ossified patients, so they are more likely to be ossified. This provided evidence that DNA methylation is involved in ligament ossification. Furthermore, because their expression was suppressed by strong methylation, MSCs deriving from patients with ossification can be used to identify genes contributing to ossification. It has been shown that certain genes involved in angiogenesis are regulated by DNA methylation, and that MSCs deriving from ossified patients have different methylation profiles from those of non-ossified patients (our unpublished observations). In future work, we plan to investigate the abnormal transformation of MSCs by methylation using next-generation sequencing to reveal the profile of genome-wide DNA methylation and its changes related to ossification.
4 Summary
The cells responsible for ectopic ossification of the spinal ligament are considered to be MSCs present in the ligament tissue. Their primary role is the repair of damaged tissue which is achieved by differentiation and proliferation through their self-replication ability and multipotency into ligament cells as required. However, the MSCs of patients with ossification are thought to misdifferentiate into osteoblasts instead of target ligament cells because they have high osteogenic ability. We consider epigenetics to be one of the mechanisms responsible for the transformation of patient MSCs (Fig. 7.4). Currently, we are focusing on DNA methylation as a mechanism of epigenetics, but we also intend to analyze other mechanisms such as histone acetylation and regulation by non-coding RNA. The importance of basic research in clinical medicine lies in how the results can be connected to the therapeutics. Therefore, if the mechanism of MSC transformation is elucidated, it will be possible to develop targeted therapies and therapeutic agents.
Role of mesenchymal stem cells (MSCs) in the ossification of spinal ligaments. The function of normal MSCs present in ligament tissue is to differentiate and proliferate into ligament cells to repair damage to ligament tissue. In ossified patients, MSCs in the ligament tissue transform in response to a genetic mutation or an epigenetic change caused by several environmental factors or mechanical stresses. Therefore, during tissue repair, they erroneously differentiate into osteoblasts, undergo endochondral ossification and angiogenesis, and ossify ligament tissue
Abbreviations
- CXCR4:
-
C-X-C motif chemokine receptor 4
- LF:
-
Ligamentum flavum
- MSCs:
-
Mesenchymal stem cells
- OLF:
-
Ossification of the ligamentum flavum
- OPLL:
-
Ossification of the posterior longitudinal ligament
- PLL:
-
Posterior longitudinal ligament
- SDF-1:
-
Stromal cell-derived factor
References
Resnick D, Shaul SR, Robins JM. Diffuse idiopathic skeletal hyperostosis (DISH). Radiography. 1975;115:513–24.
Matsunaga S, Sakou T. In: Yonenobu K, Sakou T, Ono K, editors. Ossification of the posterior longitudinal ligament. Tokyo: Springer; 1997. p. 11–7.
Kawaguchi Y, Nakano M, Yasuda T, Seki S, Hori T, Suzuki K, Makino H, Kimura T. Characteristics of ossification of the spinal ligament; incidence of ossification of the ligamentum flavum in patients with cervical ossification of the posterior longitudinal ligament—analysis of the whole spine using multidetector CT. J Orthop Sci. 2016;21:439–45.
Tetreault L, Nakashima H, Kato S, Kryshtalskyj M, Nagoshi N, Nouri A, Singh A, Fehlings MG. A systematic review of classification systems for cervical ossification of the posterior longitudinal ligament. Global Spine J. 2019;9:85–103.
Joaquim AF, Makhni MC, Riew KD. Post-operative nerve injuries after cervical spine surgery. Int Orthop. 2019;43:791–5.
Taketomi E, Sakou T, Matsunaga S, Yamaguchi M. Family study of a twin with ossification of the posterior longitudinal ligament in the cervical spine. Spine. 1992;17:S55–6.
Okawa A, Nakamura I, Goto S, Moriya H, Nakamura Y, Ikegawa S. Mutation in Npps in a mouse model of ossification of the posterior longitudinal ligament of the spine. Nat Genet. 1998;19:271–3.
Tanaka T, Ikari K, Furushima K, Okada A, Tanaka H, Furukawa K, Yoshida K, Ikeda T, Ikegawa S, Hunt SC, Takeda J, Toh S, Harata S, Nakajima T, Inoue I. Genomewide linkage and linkage disequilibrium analyses identify COL6A1, on chromosome 21, as the locus for ossification of the posterior longitudinal ligament of the spine. Am J Hum Genet. 2003;73:812–22.
Nakajima M, Takahashi A, Tsuji T, Karasugi T, Baba H, Uchida K, Kawabata S, Okawa A, Shindo S, Takeuchi K, Taniguchi Y, Maeda S, Kashii M, Seichi A, Nakajima H, Kawaguchi Y, Fujibayashi S, Takahata M, Tanaka T, Watanabe K, Kida K, Kanchiku T, Ito Z, Mori K, Kaito T, Kobayashi S, Yamada K, Takahashi M, Chiba K, Matsumoto M, Furukawa K, Kubo M, Toyama Y, Genetic Study Group of Investigation Committee on Ossification of the Spinal Ligaments, Ikegawa S. A genomewide association study identifies susceptibility loci for ossification of the posterior longitudinal ligament of the spine. Nat Genet. 2014;46:1012–7.
Inamasu J, Guiot BH, Sachs DC. Ossification of the posterior longitudinal ligament: an update on its biology, epidemiology, and natural history. Neurosurgery. 2006;58:1027–39.
Furukawa KI. Pharmacological aspect of ectopic ossification in spinal ligament tissues. Pharmacol Ther. 2008;118:352–8.
Matsunaga S, Sakou T, Taketomi E, Nakanisi K. Effects of strain distribution in the intervertebral discs on the progression of ossification of the posterior longitudinal ligaments. Spine. 1996;21:184–9.
Kon T, Yamazaki M, Tagawa M, Goto S, Terakado A, Moriya H, Fujimura S. Bone morphogenetic protein-2 stimulates differentiation of cultured spinal ligament cells from patients with ossification of the posterior longitudinal ligament. Calcif Tissue Int. 1997;60:291–6.
Tanaka H, Nagai E, Murata H, Tsubone T, Shirakura Y, Sugiyama T, Taguchi T, Kawai S. Involvement of bone morphogenic protein-2 (BMP-2) in the pathological ossification process of the spinal ligament. Rheumatology (Oxford, England). 2001;40:1163–8.
Tanno M, Furukawa KI, Ueyama K, Harata S, Motomura S. Uniaxial cyclic stretch induces osteogenic differentiation and synthesis of bone morphogenetic proteins of spinal ligament cells derived from patients with ossification of the posterior longitudinal ligaments. Bone. 2003;33:475–84.
Iwasaki K, Furukawa KI, Tanno M, Kusumi T, Ueyama K, Tanaka M, Kudo H, Toh S, Harata S, Motomura S. Uni-axial cyclic stretch induces Cbfa1 expression in spinal ligament cells derived from patients with ossification of the posterior longitudinal ligament. Calcif Tissue Int. 2004;74:448–57.
Iwasawa T, Iwasaki K, Sawada T, Okada A, Ueyama K, Motomura S, Harata S, Inoue I, Toh S, Furukawa KI. Pathophysiological role of endothelin in ectopic ossification of human spinal ligaments induced by mechanical stress. Calcif Tissue Int. 2006;79:422–30.
Li JM, Zhang Y, Ren Y, Liu BG, Lin X, Yang J, Zhao HC, Wang YJ, Song L. Uniaxial cyclic stretch promotes osteogenic differentiation and synthesis of BMP2 in the C3H10T1/2 cells with BMP2 gene variant of rs2273073 (T / G). PLoS One. 2014;9:e106598.
Asari T, Tanaka S, Kudo H, Mizukami H, Ono A, Numasawa T, Kumagai G, Motomura S, Yagihashi S, Toh S. Mesenchymal stem cell isolation and characterization from human spinal ligaments. Biochem Biophys Res Commun. 2012;417:1193–9.
Nancarrow-Lei R, Mafi P, Mafi R, Khan W. A systemic review of adult mesenchymal stem cell sources and their multilineage differentiation potential relevant to musculoskeletal tissue repair and regeneration. Curr Stem Cell Res Ther. 2017;12:601–10.
Harada Y, Furukawa KI, Asari T, Chin S, Ono A, Tanaka T, Mizukami H, Yagihashi S, Motomura S, Ishibashi Y. Osteogenic lineage commitment of mesenchymal stem cells from patients with ossification of the posterior longitudinal ligament. Biochem Biophys Res Commun. 2014;443:1014–20.
Chin S, Furukawa KI, Ono A, Asari T, Harada Y, Wada K, Tanaka T, Inaba W, Mizukami H, Motomura S, Yagihashi S, Ishibashi Y. Immunohistochemical localization of mesenchymal stem cells in ossified human spinal ligaments. Biochem Biophys Res Commun. 2013;436:698–704.
Kucia M, Reca R, Miekus K, Wanzeck J, Wojakowski W, Janowska-Wieczorek A, Ratajczak J, Ratajczak MZ. Trafficking of normal stem cells and metastasis of cancer stem cells involve similar mechanisms pivotal role of the SDF-1-CXCR4 axis. Stem Cells. 2005;23:879–94.
Chin S, Furukawa KI, Kurotaki K, Nagasaki S, Wada K, Kumagai G, Motomura S, Ishibashi Y. Facilitation of chemotaxis activity of mesenchymal stem cells via stromal cell-derived factor-1 and its receptor may promote ectopic ossification of human spinal ligaments. J Pharmacol Exp Ther. 2019;369:1–8.
Yamamoto Y, Furukawa KI, Ueyama K, Nakanishi T, Takigawa M, Harata S. Possible roles of CTGF/Hcs24 in the initiation and development of ossification of the posterior longitudinal ligament. Spine. 2002;27:1852–7.
Kishiya M, Furukawa KI, Yokoyama T, Kudo H, Ono A, Numasawa T, Wada K, Toh S. Comparison of cardiovascular parameters between patients with ossification of posterior longitudinal ligament and patients with cervical spondylotic myelopathy. J Spin Disord Tech. 2009;22:361–6.
Hiraki Y, Inoue H, Iyama K, Kamizono A, Ochiai M, Shukunami C, Iijima S, Suzuki F, Kondo J. Identification of chondromodulin I as a novel endothelial cell growth inhibitor. Purification and its localization in the avascular zone of epiphyseal cartilage. J Biol Chem. 1997;272:32419–26.
van Dinther M, Visser N, de Gorter DJ, Doorn J, Goumans MJ, de Boer J, ten Dijke P. ALK2 R206H mutation linked to fibrodysplasia ossificans progressiva confers constitutive activity to the BMP type I receptor and sensitizes mesenchymal cells to BMP-induced osteoblast differentiation and bone formation. J Bone Miner Res. 2010;25:1208–15.
Medici D, Shore EM, Lounev VY, Kaplan FS, Kalluri R, Olsen BR. Conversion of vascular endothelial cells into multipotent stem-like cells. Nat Med. 2010;16:1400–6.
Nakajima M, Kou I, Ohashi H, Genetic Study Group of the Investigation Committee on the Ossification of Spinal Ligaments, Ikegawa S. Identification and functional characterization of RSPO2 as a susceptibility gene for ossification of the posterior longitudinal ligament of the spine. Am J Hum Genet. 2016;99:202–7.
Takegami Y, Ohkawara B, Ito M, Masuda A, Nakashima H, Ishiguro N, Ohno K. R-spondin 2 facilitates differentiation of proliferating chondrocytes into hypertrophic chondrocytes by enhancing Wnt/β-catenin signaling in endochondral ossification. Biochem Biophys Res Commun. 2016;473:255–64.
Pérez-Campo FM, Riancho JA. Epigenetic mechanisms regulating mesenchymal stem cell differentiation. Curr Genomics. 2015;16:368–83.
Montes de Oca A, Madueño JA, Martinez-Moreno JM, Guerrero F, Muñoz-Castañeda J, Rodriguez-Ortiz ME, Mendoza FJ, Almaden Y, Lopez I, Rodriguez M, Aquilera-Tejero E. High-phosphate-induced calcification is related to SM22α promoter methylation in vascular smooth muscle cells. J Bone Miner Res. 2010;25:1996–2005.
Chiba N, Furukawa KI, Takayama S, Asari T, Chin S, Harada Y, Kumagai G, Wada K, Tanaka T, Ono A, Motomura S, Ishibashi Y. Decreased DNA methylation in the promoter region of the WNT5A and GDNF genes may promote the osteogenicity of mesenchymal stem cells from patients with ossified spinal ligaments. J Pharmacol Sci. 2015;127:467–73.
Acknowledgement
This research is the result of a joint project with Hirosaki University Graduate School of Medicine Orthopedic Surgery. I would like to express my heartfelt thanks to my collaborators. This work was supported by a Grant-in-Aid from the Practical Research Project for Rare/Intractable Diseases from the Japan Agency for Medical Research and Development (AMED) and a Grant-in-Aid from the Ministry of Education, Culture, Sports, Science and Technology of Japan [Grant 17K10916]. No potential conflicts of interest relevant to this article are reported. We thank Sarah Williams, PhD, from Edanz Group (www.edanzediting.com) for editing a draft of this chapter.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Furukawa, KI., Chin, S., Asari, T., Wada, K., Kumagai, G., Ishibashi, Y. (2020). Ectopic Ossification of Human Spinal Ligaments Caused by Mesenchymal Stem Cell Abnormalities. In: Okawa, A., Matsumoto, M., Iwasaki, M., Kawaguchi, Y. (eds) OPLL. Springer, Singapore. https://doi.org/10.1007/978-981-15-3855-1_7
Download citation
DOI: https://doi.org/10.1007/978-981-15-3855-1_7
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-3854-4
Online ISBN: 978-981-15-3855-1
eBook Packages: MedicineMedicine (R0)