Abstract
Cardiovascular Diseases (CVDs) as a leading cause of death worldwide inflict major stress on morbidity and societal costs. Though the studies pertaining to pathophysiology and genetics of CVDs have helped in prevention, diagnosis and treatment of diseases, there are still lacunas in our knowledge. So, novel tools that can define genomic regulation under different conditions are needed to bridge this gap. ‘Epigenetic’ mechanism helps the cells to quickly respond to ever changing environment by molecular mechanisms like methylation, histone modifications, nc-RNAs. These mechanisms act as a new layer of regulation in CVDs. The role of epigenetics as a key regulatory player in prevention, diagnosis and treatment of CVDs is emerging. Thus, the focus of present chapter is to decipher the role of epigenetics in CVDs and its potential to be used in risk assessment or as biomarkers in devising and deploying better diagnosis and treatment for different CVDs.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Background
Cardiovascular system responsible for transportation of nutrients as well as cellular waste products comprises of the heart, blood and blood vessels. The term cardiovascular diseases (CVDs) refer to any disease of cardiovascular system. CVDs inflict major burden on mortality worldwide making it global epidemic even after the advances made in their prevention and management. CVDs are not only public health issue but also accounts for massive societal costs as healthcare expenditure. They as a group of multifactorial disorders are associated with various genetic and acquired risk factors. But the known genetic and environmental influences cannot fully explain the variability in CVD risk among different populations contributing to the major road blocks in its treatment and prevention. Though thorough studies pertaining to pathophysiology, epidemiology, gene polymorphism, genetic linkage maps and various environmental stresses have paved a way for better diagnosis and treatment of cardiovascular medicine, but still there is lacuna in our understanding of interaction between environment and genome, in development of CVD. As inherited genome contribute to only a part of individual’s risk profile, ‘epi’genomics has emerged as a promising area of interest that can bridge gaps in our understanding pathophysiology to therapeutics of diseases. Epigenetic changes or modifications are the ‘heritable changes in the genome that does not include changes in DNA sequences’ which includes various mechanisms like DNA methylation, demethylation, histone modifications, RNA mediated alterations (microRNAs, circulatory RNAs, long non-coding RNAs) etc. Epigenetic mechanisms collectively empower the cell to quickly respond to environmental changes. Various established risk factors for CVD such as stress, pollution, smoking, nutrition, circadian rhythm, hypoxia etc. are associated with modifications in epigenetic markers. Therefore, the examination of these markers as regulators in CVD may emerge as early preventive and novel therapeutic approach. This book chapter therefore focuses on acquainting the vascular biology community with the advent filed of epigenetics and its role in cardiovascular medicine. The chapter consists of a brief introduction of cardiovascular disease, epigenetic mechanisms and evidences linking various epigenetic mechanisms with CVD. The main emphasize of this chapter would be on epigenetic regulator of CVDs to understand their translational potential and clinical reality in cardiovascular medicine.
2 Cardiovascular Diseases (CVDs)
CVDs are wide spectrum of disorders that comprises of abnormalities related to heart and blood vasculature. They include: Coronary artery diseases (CAD) like angina and myocardial infarction (heart attack), stroke, heart failure, coronary heart disease, cerebrovascular disease, peripheral arterial disease, rheumatic heart disease cardiomyopathy, heart arrhythmia, congenital heart disease, congenital heart disease, thromboembolic disorders and thrombosis. The underlying molecular mechanism pertaining to different CVDs vary considerably as majority of CVDs are regulated by multiple factors. For example, CAD and stroke involve atherosclerosis and rheumatic heart disease involves damage to heart muscle and valves caused by streptococcal bacteria. Even with different molecular mechanisms, nevertheless, most CVDs share common risk factors that are, high blood pressure, high blood cholesterol, diabetes mellitus, smoking, obesity, lack of exercise, alcohol consumption, hypoxia, to name a few.
CVDs are the leading cause of death globally accounting for most of annual mortality as compared to any other disease [1]. Over the period of last 100 years the prevalence of CVDs has increased tremendously and for the coming two decades it is likely to remain a major health issue even with sophisticated techniques and improved medical care [2,3,4]. The deaths caused by CVDs have increased over the course of time from 12.3 million (25.8%) in 1990 to 17.9 million (32.1%) in 2015 [5, 6]. In 2016, 31% of the global deaths that reached up to 17.9 million people were due to Different CVDs. Heart attack and stroke contributed to 85% of these deaths [7]. Studies estimate that 90% of the CVDs may be preventable [8, 9] by improving the known risk factors like hypertension, diabetes, smoking, alcohol consumption by healthy eating, avoidance of smoke and alcohol, exercise and appropriate medical counselling [10]. Thus, people who are at high cardiovascular risk need early detection and management in order to prevent the occurrence of disease.
The CVDs appeared as a huge public health hazard during 1940s, that triggered the research associated with the factors that influence occurrence of the disease. The research programmes like Framingham Heart Study were initiated to find out common genetic, biochemical, environmental and life style related factors that lead to occurrence or predisposition to various CVDs [11]. This resulted in identification of several conditions like obesity, hypertension and diabetes to detect individuals at risk of CVD and their early treatment. It in turn lead to significant decline in CVD mortality in USA and Western Europe over the past 3 decades. Therefore, it is important to understand the risk factors associated with CVDs.
2.1 Types of Cardiovascular Diseases
CVDs are group of disorders involving heart, blood vessels or both. Thus, they are characterised either as vascular diseases that majorly affects blood vasculature or heart diseases that majorly affects heart.
2.1.1 Vascular Diseases
The common vascular diseases are coronary artery disease, peripheral arterial disease, cerebrovascular disease, renal artery stenosis, aortic aneurysm, stroke, thrombosis.
2.1.2 Heart Diseases
The common CVDs affecting heart are cardiomyopathy, hypertensive heart disease, heart failure, pulmonary heart disease, cardiac dysrhythmias, inflammatory heart disease, valvular heart disease, congenital heart disease, rheumatic heart disease.
2.2 The Risk Factors Associated with CVDs
The most contributing risk factors for stroke and heart disease are genetic predisposition, unhealthy diet, age, sex, excessive use of alcohol, obesity, hypertension, diabetes, hyperlipidemia, stress, air pollution etc. Though it is very difficult to access the individual contribution of each risk factor among different ethnic group or populations, but the overall impact of these risk factors is quite consistent [12].
2.2.1 Genetic Predisposition
Most CVDs are multifactorial in nature i.e. there are several genes or genetic factors that contribute to development and progression of the disease. There are many identified single nucleotide polymorphisms (SNPs) make individual susceptible to the disease [13, 14]. Even with the sophisticated techniques the contribution of all genetic factors and their influence of vascular diseases is poorly understood.
2.2.2 Age
Age is one of the most important contributing risk factors towards the development of heart diseases, with each passing decade it triples the risk of disease [15]. The case studies show that 82% coronary heart patients that die are either of 65 years or older [15]. Similarly, the risk of death by stroke doubles after every 10 passing years after the age of 55 [17]. There are multiple theories that explain the influence of age on CVDs, one relates to increased serum cholesterol level with age [18] other with the changes in vascular wall [15]. Age decreases the arterial elasticity and reduced arterial compliance that subsequently causes coronary artery disease.
2.2.3 Sex
Gender influences the prevalence of heart diseases. Men are more susceptible than pre-menopausal women [16]. Middle aged men are at 2–5 times higher risk of coronary heart diseases at compare to women [19]. A study by World Health Organisation (WHO) reveals that sex contributes to roughly 40% difference in the mortality rate caused by coronary heart diseases [20]. The influence of gender in CVDs may be due to hormonal differences between men and women [20].
2.2.4 Excessive Use of Alcohol
The effect of alcohol consumption on CVDs is complex and depends largely on the quantity of alcohol consumed. The high level of drinking is directly related to susceptibility to the disease [21].
2.2.5 Obesity
Another risk factor for developing CVDs is being overweight or obesity. An unhealthy diet and physical inactivity lead to high body mass index (BMI), that falls outside the normal range making a person obese. This may lead to diseases like coronary heart disease and congestive heart failure [22].
2.2.6 Hypertension
High blood pressure or hypertension cause stress on blood vasculature making them weak or cause them to clog. It is a potent risk factor that can lead to atherosclerosis as it causes narrowing of blood vessels which make them more prone to be blocked by blood clots or fatty deposits [23].
2.2.7 Diabetes
High blood glucose levels as experienced in diabetes make individual more vulnerable for developing CVDs. The high levels of glucose have damaging effect on arterial walls and more likely cause the build-up of fat deposits called atheroma. These fat depositions if occur in coronary arteries may lead to coronary heart disease and heart attacks [24].
2.2.8 Hyperlipidemia
Hyperlipidemia is presence of abnormal level of lipids (fats) in blood stream and is one of the major contributory risk factors for CVDs. The high levels of “bad cholesterol” i.e. low-density lipoprotein (LDL) cholesterol is associated with range of heart diseases as it results in fatty substance deposition on the vasculature leading to various complications [23].
2.2.9 Diet
Diet plays a crucial role in prevention and development of CVDs. A diet rich in saturated fatty acids increases the risk of stroke and heart diseases. It is estimated that 11% of stroke and 31% of coronary heart diseases worldwide are caused by high dietary saturated fatty acids [23].
2.2.10 Air Pollution
The effect of particulate matter present in the air for its short- and long-term exposure on CVDs has been accessed. It was found in recent studies that the long-term exposure of PM2.5 results in increased 8–18% CVD mortality risk [24].
The majority of above-mentioned CVD risk factors are manageable and can be reduced by opting alternative lifestyle. For example, reduction of alcohol consumption, fatty food, salt, regular physical exercise, consumption of vegetables and fruits reduce the risk of CVDs drastically. In addition, the treatment of hypertension, diabetes and hyperlipidemia is essential for reducing the risk of heart diseases. Thus, there are choices by which people can adopt and sustain healthy way of living, but we need health policies and conducive environments that can make these choices affordable and available to masses. The other major forces driving occurrence of CVDs are stress, poverty, urbanization, socio-economic and cultural changes. With all the knowledge on contributing factors and advancement of genomic era, we still fail to answer the variability in CVD risk, role of individual risk factors, their cumulative effect. Also, the clinical utility of identified genetic markers has been limited in both prevention and prediction of diseases [25].With the translational disappointment of genomics field, the vascular community is now exploiting epigenomics as new avenue for assessing risk stratification, preventive measures and treatment. Epigenetics is the heritable changes in the genome that are not coded by the DNA sequence [26]. There are three broad mechanisms that define epigenetic changes: DNA methylation, post- translational histone modifications and RNA based mechanisms (microRNAs, circular RNAs and non-coding RNAs) [27]. DNA methylation is the most common epigenetic modification in the mammalian genome [28, 29]. The recent studies though suggest that all the three epigenetic modifications contribute to the pathogenesis of CVDs [30]. The international projects like Human Epigenome Project (HEP) and International Human Epigenome Consortium (IHEC) have been initiated to study and catalogue human epigenome along with correlation of its pathophysiology with several diseases including CVDs [31].The focus of this chapter is epigenetics in CVDs, and will be discussed under the following sections.
3 Leads from Genetics
Almost three decades ago the research on the genetic factors and genes that predispose an individual to CVDs has started. It was thought that the SNPs and mutations in the genome could be analogous to already known CVD risk factors. It was anticipated that they could be included in risk assessment model like Framingham score [30] in order to calculate and determine the risk of an individual to develop CVD and adopt apt preventive therapeutic approaches. The traditional approach was to study the role of candidate gene as risk factor, but now the genome-wide association studies grouped with availability of large patient cohorts are used to uncover the novel genes and genetic factors that act like risk factors for various CVDs [31]. The genetic research has enabled the cardiovascular community to better understand the disease origin and its pathophysiology. It has progressively also helped in the risk stratification, direct and indirect diagnosis and therapeutic interventions. The novel GWAS has widened the understanding of pathophysiology of various CVDs. The genetic studies have also led to discovery of new drug targets that may considerably improve the current therapeutic approaches for CVDs. The genotyping studies have been used for diagnostic and prognostic assessment by clinicians. But still genetic studies are unable to find the “holy grail” for diagnosis of CVDs. So, there is a need of more profound knowledge that can answer dynamics of disease. ‘Epi’genetics promises to deepen the present insights as it can well explain the influence of environment on the genome. It can unveil the interaction between the genetic variants and environment changes; thus, can elucidate the clinical drug responsiveness and patient outcome.
4 Epigenetic Revolutions – Genomics to Post-Genomic Era
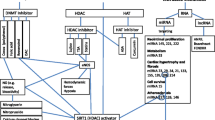
Epigenetics has emerged as a post genomic era that can reveal the lacuna in our understanding of inheritance of common traits. It provides the link between environment and its effect on genome, thus deciphering the mechanism by which cells are able to respond quickly to environmental stimuli. The phenotypic variations seen in humans that cannot be solely defined by genotype, can be better explained in the light of epigenetic modifications [32]. Epigenetic mechanisms are stable cellular memory, which let the propagation of gene activities generation after generation without the changes in the DNA sequence. There are three broad mechanisms that define epigenetic changes: DNA methylation, post- translational histone modifications and RNA based mechanisms (microRNAs, circular RNAs and long non-coding RNAs). The simple and static evaluation of the genetic code has limited the understanding of genetic influence on CVDs. In this respect, epigenetics as a dynamic mechanism can explain and modify the genome’s functionality under the influence of exogenous stimuli or environment condition. This would in turn help in identification of novel targets and mechanism of gene regulation with considerable acquisitions in CVD knowledge related to genetic risk and pathophysiology [33,34,35]. The environmental changes like pollution, diet, stress etc. leads to epigenetic modifications and impact the susceptibility to CVD (Fig. 6.1). There are reports that validate the role of epigenetic in various processes underlying CVDs like hypertension, inflammation and atherosclerosis [35].
Epigenetic modifications: The figure depicts the three most common epigenetic modifications that occur in mammalian genome namely, histone modification and chromatin remodelling, DNA methylation and modifications mediated by non-coding RNA (ncRNA) family like miRNA, circRNA and lncRNA. All these modifications are interlinked and enable cells to quickly respond to changing environment. The established risk factors of CVD like stress, pollution, age, diet, pollution etc. can act as environmental stimuli and triggers epigenetic modifications resulting in release of various regulatory molecules that can be used as biomarkers. The changes also influence the CVD outcomes
The following sections of the chapters will deal with different epigenetic modifications and their role in progression, prevention, treatment and diagnosis of CVDs.
4.1 DNA Methylation
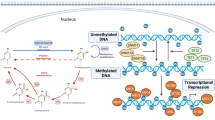
DNA methylation is the addition of the methyl group at the cytosine bases of the dinucleotide CpGs of eukaryotic DNA. As a result they are converted to 5-methylycytosine by de novo DNA methyltransferase (DNMT) enzymes such as DNMT1, DNMT3A and DNMT3B. In mammals, methylation is found sparsely but globally with the exception of CpG islands. Vertebrate CpG islands are short interspersed DNA sequences (>500 bp) generally found in the 5′ end of the gene that deviate significantly from the average genomic pattern by being GC-rich (G + C percentage greater than 55%), CpG rich (observed CpG/expected CpG of 0.65) and predominantly non-methylated [36].
Out of the several ways, the methylation of DNA is one of the most common ways of regulating gene expressions in eukaryotes [37]. It has emerged as an important process in numerous cellular processes like genomic imprinting, embryonic development, X-chromosome inactivation and many more. The first clue of the role of methylation in gene expression was provided by 5-azacytidine experiments in mouse studies [38]. The integration of 5-azacytidine in the growing strand of DNA severely inhibited the actions of DNMT enzymes to normally methylate DNA. Therefore, the comparisons of the cells before and after the treatment of 5-azacytidine allowed to see the impact of loss of methylation on gene expression [39]. The exact role of methylation in gene expression is unknown, perhaps it play a crucial role in repressing gene expression by blocking the promoters at which activating transcription factors bind. Approximately, 70% of annotated gene promoters are associated with CpG islands, making it the most common promoter type in the genome [36]. Not all CpG islands found are associated with the promoter. Recent works have found a large class of islands that are remote from the transcription start sites (TSSs) but still show evidence for the promoter function. The lack of CpG dinucleotides in the vertebrate genome except the CpG island is thought to be due to the loss of genomic CpGs due to deamination of methylated sequences [39].
Given the critical role of DNA methylation in gene expression and cell differentiation, it seems obvious that the errors in methylation could give rise to a number of devastating consequences, including various diseases. As a result, a growing number of human diseases have been found to be associated with aberrant DNA methylation [40]. The methylation of the promoter region bearing transcriptional start sites of many genes encoding tumor suppressors such as tumor protein p53, retinoblastoma-associated protein 1, tumor protein p16, breast cancer 1 and many more resulting in the reduced expression of these genes have been found in a large number of cancers like retinoblastoma, colon, lung and ovarian [40, 41]. 5-methylcytosine (5-mC) is spontaneously converted to thymine by deamination and is thought to be responsible for about one-third of all disease-causing mutations in the germline [39]. The mechanisms for establishing, maintaining and removing the methyl group are dependent on nucleosomal DNA and the histone modifications within the nucleosome [39]. Global methylation studies are the first ever epigenetics studies in the area of CVD research. The outcome of these studies though were conflicting as some studies highlighted decreased global DNA methylations with CVDs [40] while some associated increased global methylations with these disorders [41, 42]. Coronary heart disease have been associated with elevated homocystein and increased methylations [43]. DNA methylation studies include investigation of repetitive sequences across the genome such as Alu elements and long interspersed nucleotide element-1 (LINE)-1. Hypomethylation of LINE-1 was associated with cardiovascular risk factors such as higher serum vascular cell adhesion molecule [44] and higher low-density lipoprotein (LDL) and lower high-density lipoprotein (HDL) cholesterol [47]. LINE-1 hypomethylation has also been associated with higher prevalence of metabolic syndrome, and therefore elevated risk for CVD [48]. There are several genes responsible for DNA methylation and investigation of these candidate genes demonstrated some interesting results. Fat mass and obesity-associated protein (FTO), a prominent obesity-associated gene has been linked to levels of DNA methylation [49] and CVD risk independently of its effect on body mass index [50].Similarly, other candidate genes such as F2R Like Thrombin Or Trypsin Receptor 3, Insulin, GNAS Antisense RNA 1, Phospholipase A2 Group VII, Insulin Like Growth Factor 2, to name a few has been associated with various CVDs [51,52,53,54]. With the advent of whole epigenome array several markers and novel distinct patterns of DNA methylations have been identified in the context of CVDs [55,56,57,58]. The outcome of all these studies could result in the novel therapeutic interventions in cardiovascular diseases.
4.2 Post-translation Histone Modifications and Chromatin Remodelling
Histone modification is a covalent post-translational modification, which includes methylation, phosphorylation, acetylation, sumoylation and ubiquitylation, to histone proteins. These modifications are known to play an important role in replication, transcription, heterochromatin formation, chromatin compaction, and DNA damage repair. Investigation of histone modifications in CVDs reveals the crucial role of histone deacetylases (HDACs). It is the most extensively studied family of histone-modifying enzymes in the cardiovascular system [59]. Studies have demonstrated the effect of its inhibition in attenuating hypertension [60] and in prevention of proliferation of vascular smooth muscle cells [61, 62]. Class III HDACs also known as sirtuins have been shown to participate in cardiac hypertrophy and myocardial ischemia [63]. The other class of enzyme, histone acetyl transferase has been associated in the settings of atherosclerosis through regulation of genes that inhibit endothelial cell inflammation [64]. Similarly, the loss-of-function studies of histone methyltransferase in the adult heart showed hypertrophy, dilation, and derepression of some cardiac disease genes [65]. The polycomb repressive complex, one of the best-studied gene silencing complexes, has been implicated in a wide variety of phenotypes in the cardiovascular system. They have been shown to be differentially involved in cardiac development and regeneration [66].
4.3 RNA Based Modifications (miRNAs, circRNAs, lncRNAs)
The RNA based modifications include microRNAs (miRNAs), circular RNAs (circRNAs) and recently discovered long non-coding RNAs (lncRNAs). They all are non-coding endogenous RNAs that regulate the genome without being translated into proteins. The recent studies show the implication of these non-coding RNAs in CVD pathophysiology. There is large number of growing evidences that show role of miRNAs and lncRNAs in both animal and cellular models of CVD. In the following sections, we will be studying the role of these non-coding RNAs in CVDs and how they can be used as either biomarker or for treatment in CVDs.
4.3.1 miRNAs in CVD Scenario
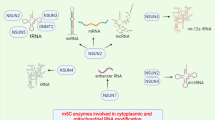
miRNAs are class of endogenous interfering RNAs that are coded by human genome. Mature miRNAs are 20–25 nucleotide long RNA sequence that are synthesised in canonical pathway from a large RNA precursor, pri-miRNA. In the nucleus pri-miRNA is transcribed and matured by RNA polymerase II. It is then subsequently cleaved by RNase III Drosha and associated protein, giving a 60–70 nucleotide long pre-miRNA, which is released into the cytoplasm. In cytoplasm it is cleaved by RNaseIII Dicer resulting in 21–25 nucleotide long double stranded miRNA/miRNA∗ duplex. This duplex is unwounded by RNA induced-silencing complex (RISC), which carries mature single stranded miRNA to target messenger RNA causing subsequent gene silencing. Thus, miRNAs decrease the expression of gene by binding to them and causing translational repression [67]. Non-coding RNAs like miRNAs have emerged as prime candidates for discovering novel biomarkers for CVDs. As there is correlation between the pathologies and blood miRNA levels, miRNAscan be great tool for being non-invasive biomarkers. The general hypothesis is that miRNAs would be differentially expressed in people having CVD and are at risk of CVD and in normal population. So, plasma miRNA levels can be promising avenue for evaluating early CVD risk and patient outcome, along with assessing individual patient response towards various surgical procedures or treatment. The pioneering work in this field is done by Dimmeler et al., they highlighted the change in serum miRNA levels of CAD patients Vs. control [42]. There are other studies that have shown miRNAs as potential biomarkers to predict CVD outcome. For instance, the work done by Seronde et al. has shown that low serum levels of miR-423-5p are associated with a poor long-term outcome in acute heart failure patients, emphasising the role of miR-423-5p as a prognostic biomarker for predicting acute heart failure [43]. The cardiovascular deaths caused by acute coronary syndrome were precisely predicted by the sericlevels of miR-132, miR-140-3p, and miR-210 in the study consisting of 1114 patients [44]. Several miRNAs are very sensitive to even small exogenous stimuli like changes in blood pressure that can be predictive of circulatory stress or hypertension (established risk factor for CVD) [45]. miR-22 is a potential biomarker candidate for predicting CVD in elderly patients asmiR-22 regulates cardiac autophagy specially in myocardium of elderly patients thus circulating levels this miRNA gives prognostic clue on eventual progression of disease ultimately leading to heart failure in elderly [46]. Regular exercise can reduce the outcomes of CVDs and several miRNAs have been proposed as biomarkers to monitor physiological effect of regular exercise in different populations. For example, decreased level of miR-146a andmiR-221, and increased level of miR-149 are seen after acute exercise [47]. Whereas the altered level of expression of miR-1, miR-133, and miR-206 is seen after the endurance exercise [48]. A recent study has highlighted the role of miR-145 in impeding the thrombus formation in vivo by targeting tissue factor in the case of venous thrombosis [49].
4.3.2 circRNAs in CVD Scenario
circular RNAs are single stranded RNAs that form a covalently closed continuous loop. They are expressed in mammalian tissues as transcriptional and translational regulators [50]. Just like miRNAs, circRNAs are stable and detectable in blood stream, making it possible to use them as non-invasive biomarker. It is seen that in aged human hearts the level of circAmotl1decreases dramatically [51]. The risk of atherosclerotic vascular diseases is correlated with the expression of the increased expression circANRILin CAD patients [52]. The diagnostic biomarkers for various CVDs identified in peripheral blood mononuclearcells (PBMCs) are Hsa_circ_0005836, hsa_circRNA_025016, andhsa_circ_0124644 [53, 54].
4.3.3 Long Non-coding (lnc)-RNAs in CVD Scenario
lncRNAs are the RNA sequences that exceed more than 200 nucleotides and don’t code for any functional transcript. They differ from other non-coding RNAs like miRNAs as they are poorly conserved across the species also, they regulate gene expression at both transcription and post-transcriptional level. They allow microRNAs’ target mRNA escape degradation as they act as decoy for microRNA [55]. The advancement in sequencing techniques like deep sequencing has made it possible to study the lncRNA profiles in different CVDs as well as normal healthy population. And it is found that lncRNAs regulate multiple biological pathways in aging and cardiac development [56]. The 9 lncRNAs namely CDKN2BAS1/ANRIL, RMRP, RNY5, SOX2-OT, SRA1 EGOT, H19, HOTAIR, and LOC285194/TUSC7 were significantly modulated in non-ischemic myocardial biopsies of patients suffering from heart failure and dilated ischemic cardiomyopathy. Also, in mouse model the expression level of RMRP, H19, and HOTAIRlncRNAs were upregulated when measured in hypertrophic heart sample [57]. MIAT (myocardial infarction-associated transcript) and SENCR (smooth muscle and endothelial cell enriched migration/differentiation-associated lncRNA), are other lncRNA markers that are associated with dysfunction and myocardial infarction [58, 59]. Nodal lncRNAs act as key regulators of cardiomyocte cell cycle as revealed by the data of single cardiomyocyte nuclear transcriptome analysis of normal and failing heart [60]. Interestingly, the severity of fibrosis in human diseased heart is directly correlated with a cardiac fibroblast enriched lncRNA- Wisper (Wisp2 super enhancer-associated RNA) [61]. Growing evidences show that majority of lncRNAs are localised in nucleus and are capable of triggering chromatin remodelling by recruitment of various epigenetic factors, thereby causing activation or repression of genes. The genes involved in cardiac hypertrophy are regulated by a cardiac-enriched lncRNA via its direct interaction with catalytic subunit of PRC2 causing inhibition of methylation of histone H3 lysine 27 [62].
At last we can conclude that non-coding RNAs play a role in controlling gene expression and shaping genome organisation. With increasingly more functions and roles of non-coding RNAs being discovered, we need to use better biochemical approaches along with deep sequencing analysis coupled with novel bioinformatics strategies so that a comprehensive understanding of their role in CVDs can be provided.
5 Perspective
The role of epigenetic alterations in the capacity of DNA methylation, chromatin remodelling and non coding-RNAs has been emerging in the development of CVDs. However, the results of epigenetic studies are inconsistent and contradictory. Epigenetic phenomenon, as we know, is not a stable or heritable event. It is dynamic in the sense that it gets altered by environmental factors and can vary with time. These all features have to be taken into considerations while designing the studies for any type of epigenetics research. Not just the nature of samples and the conditions they are exposed to are important but the numbers of times the samples are taken are equally critical. There are various upcoming combinatorial technologies through which researchers can explore epigenetic landscapes. Bisulfite-sequencing combined to ChIP, de novo methylation simultaneously examining nucleosome occupancy and CpG methylation can be utilized for studying multiple epigenetic marks simultaneously. Research in epigenetics is a relatively new approach but it has a remarkable potential to identify new biomarkers in the CVD field. These biomarkers would not be just helpful in CVD diagnosis, outcome, prognosis and treatment but could lead us to new avenues for novel targeted CVD therapies. However, more epigenomic studies are warranted that will help to decipher the complex link between genetics, epigenetics, and CVDs.
References
Murray C, Lopez A. Alternative projections of mortality and disability by cause1990–2020: global burden of disease study. Lancet. 1997;349:1498–504.
Mathers CD, Loncar D. Projections of global mortality and burden of disease from 2002 to 2030. PLoS Med. 2006;3:e442.
MacKinnon AU. The origin of the modern epidemic of coronary artery disease in England. J R Coll Gen Pract. 1987;37:174–6.
Azambuja MI, Levins R. Coronary heart disease (CHD)—one or several diseases? Changes in the prevalence and features of CHD. Perspect Biol Med. 2007;50:228–42.
GBD. Mortality and causes of death collaborators (2014) Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990–2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet. 2013;385(9963):117–71.
GBD. Mortality and Causes of Death Collaborators (2017) Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: a systematic analysis for the Global Burden of Disease Study 2015. Lancet. 2015;388(10053):1459–544.
McGill HC, McMahan CA, Gidding SS. Preventing heart disease in the 21st century: implications of the Pathobiological Determinants of Atherosclerosis in Youth (PDAY) study. Circulation. 2008;117(9):1216–27.
O’Donnell MJ, Chin SL. Global and regional effects of potentially modifiable risk factors associated with acute stroke in 32 countries (INTERSTROKE): a case-control study. Lancet. 2016;388(10046):761–75.
Mendis S, Puska P, Norrving B, editors. Global atlas on cardiovascular disease prevention and control. Geneva: World Health Organization/World Heart Federation/World Stroke Organization; 2011.
GBD. Mortality and causes of death collaborators (2016) Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: a systematic analysis for the Global Burden of Disease Study 2015. Lancet. 2015;388(10053):1459–544.
Yusuf S, Hawken S, Ounpuu S, Dans T, Avezum A, Lanas F, Mc Queen M, Budaj A, Pais P, Varigos J, Lisheng L, INTERHEART Study Investigators. Effect of potentially modifiable risk factors associated with myocardial infarction in 52 countries (the INTERHEART study): case-control study. Lancet. 2004;364(9438):937–52.
Nikpay M, Goel A, Won H, Hall LM, Willenborg C, Kanoni S, Saleheen D, Kyriakou T, Nelson CP. A comprehensive 1000 Genomes–based genome-wide association meta-analysis of coronary artery disease. Nat Genet. 2015;47(10):1121–30.
MacRae CA, Vasan RS. The future of genetics and genomics: closing the phenotype gap in precision medicine. Circulation. 2016;133(25):2634–9.
Finegold JA, Asaria P, Francis DP. Mortality from ischaemic heart disease by country, region, and age: statistics from World Health Organisation and United Nations. Int J Cardiol. 2012;168(2):934–45.
World Health Organization. The atlas of heart disease and stroke/Judith Mackay and George Mensah with Shanthi Mendis and Kurt Greenland. World Health Organization. 2004.
Jousilahti P, Vartiainen E, Tuomilehto J, Puska P. Sex, age, cardiovascular risk factors, and coronary heart disease. Circulation. 1999;99(9):1165–72.
Jani B, Rajkumar C. Ageing and vascular ageing. Postgrad Med J. 2006;82(968):357–62.
Mukamal KJ, Chen CM, Rao SR, Breslow RA. Alcohol consumption and cardiovascular mortality among U.S. Adults, 1987 to 2002. J Am Coll Cardiol. 2010;55(13):1328–35.
Hubert HB, Feinleib M, McNamara PM, Castelli WP. Obesity as an independent risk factor for cardiovascular disease: a 26-year follow-up of participants in the Framingham Heart Study. Circulation. 1983;67(5):968–77.
Anderson KM, Odell PM, Wilson PW, Kannel WB. Cardiovascular disease risk profiles. Am Heart J. 1991;121:293–8.
Kannel WB, McGee DL. Diabetes and cardiovascular disease. the Framingham study. J Am Med Assoc. 1979;241(19):2035–8.
Finks SW, Airee A, Chow SL, Macaulay TE, Moranville MP, Rogers KC, Trujillo TC. Key articles of dietary interventions that influence cardiovascular mortality. Pharmacotherapy. 2012;32(4):e54–87.
Khallaf M. The impact of air pollution on health, economy, environment and agricultural sources. Rijeka: InTech; 2011. p. 69–92. ISBN 978-953-307-528-0
Di Angelantonio E, Butterworth AS. Clinical utility of genetic variants for cardiovascular risk prediction: a futile exercise or insufficient data? Circ Cardiovasc Genet. 2012;5:387–90.
Egger G, Liang G, Aparicio A, Jones PA. Epigenetics in human disease and prospects for epigenetic therapy. Nature. 2004;429:457–63.
Kaikkonen MU, Lam MT, Glass CK. Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc Res. 2011;90:430–40.
Udali S, Guarini P, Moruzzi S, Choi SW, Friso S. Cardiovascular epigenetics: from DNA methylation to microRNAs. Mol Asp Med. 2013;34:883–901.
Abbott A. Project set to map marks ongenome. Nature. 2010;463:596–7.
Wilson PW, D’Agostino RB, Levy D, Belanger AM, Silbershatz H, Kannel WB. Prediction of coronary heart disease using risk factor categories. Circulation. 1998;97:1837–47.
Evans A, Salomaa V, Kulathinal S, Asplund K, Cambien F, Ferrario M, Perola M, Peltonen L, Shields D, Tunstall-Pedoe H, Kuulasmaa K. MORGAM (an international pooling of cardiovascular cohorts). Int J Epidemiol. 2005;34:21–7.
Turan N, Katari S, Coutifaris C, Sapienza C. Explaining inter-individual variability in phenotype:is epigenetics up to the challenge? Epigenetics. 2010;5:16–9.
Khalil CA. The emerging role of epigenetics in cardiovascular disease. Ther Adv Chronic Dis. 2014;5(4):178–87.
Webster AL, Yan MS, Marsden PA. Epigenetics andcardiovascular disease. Can J Cardiol. 2013;29(1):46–57.
Muka T, Koromani F, Portilla E, O'Connor A, Bramer WM, Troup J, Chowdhury R, Dehghan A, Franco OH. The role of epigeneticmodifications in cardiovascular disease: a systematic review. Int J Cardiol. 2016;212:174–83.
Bird A. CpG-rich islands and the function of DNA methylation. Nature. 1986;321:209–13.
Deaton AM, Bird A. CpG islands and the regulation of transcription. Genes Dev. 2011;25(10):1010–22.
McGhee JD, Ginder GD. Specific DNA methylation sites in the vicinity of the chicken beta-globin genes. Nature. 1979;280:419–20.
Han L, Su B, Li WH, Zhao Z. CpG island density and its correlations with genomic features in mammalian genomes. Genome Biol. 2008;9(5):R79.
Robertson KD. DNA methylation and human disease. Nat Rev Genet. 2005;6(8):597–610.
Gopalakrishnan S, Van Emburgh BO, Robertson KD. DNA methylation in development and human disease. Mutat Res. 2008;647(1–2):30–8.
Handy DE, Castro R, Loscalzo J. Epigenetic modifications: basic mechanisms and role in cardiovascular disease. Circulation. 2011;123:2145–56.
Huh I, Zeng J, Park T, Yi SV. DNA methylation and transcriptional noise. Epigenetics Chromatin. 2013;6(1):9.
Hsiung DT, Marsit CJ, Houseman EA, Eddy K, Furniss CS, McClean MD, Kelsey KT. Global DNA methylation level in whole blood as a biomarker in head and neck squamous cell carcinoma. Cancer Epidemiol Biomark Prev. 2007;16:108–14.
Sharma P, Kumar J, Garg G, Kumar A, Patowary A, Karthikeyan G, Ramakrishnan L, Brahmachari V, Sengupta S. Detection of altered global DNA methylation in coronary artery disease patients. DNA Cell Biol. 2008;27:357–65.
Baccarelli A, Tarantini L, Wright RO, Bollati V, Litonjua AA, Zanobetti A, Sparrow D, Vokonas PS, Schwartz J. Repetitive element DNA methylation and circulating endothelial and inflammation markers in the VA normative aging study. Epigenetics. 2010;5(3):222–8.
Cash HL, McGarvey ST, Houseman EA, Marsit CJ, Hawley NL, Lambert-Messerlian GM, Viali S, Tuitele J, Kelsey KT. Cardiovascular disease risk factors and DNA methylation at the LINE-1 repeat region in peripheral blood from Samoan Islanders. Epigenetics. 2011;6:1257–64.
Turcot V, Tchernof A, Deshaies Y, Pérusse L, Bélisle A, Marceau S, Biron S, Lescelleur O, Biertho L, Vohl MC. LINE-1 methylation in visceral adipose tissue of severely obese individuals is associated with metabolic syndrome status and related phenotypes. Clin Epigenetics. 2012;4:10.
Bell CG, Finer S, Lindgren CM, Wilson GA, Rakyan VK, Teschendorff AE, Akan P, Stupka E, Down TA, Prokopenko I, Morison IM, Mill J, Pidsley R, International Type 2 Diabetes 1q Consortium, Deloukas P, Frayling TM, Hattersley AT, MI MC, Beck S, Hitman GA. Integrated genetic and epigenetic analysis identifies haplotype-specific methylation in the FTO type 2 diabetes and obesity susceptibility locus. PLoS One. 2010;5:e14040.
Liu C, Mou S, Pan C. The FTO gene rs9939609 polymorphism predicts risk of cardiovascular disease: a systematic review and meta-analysis. PLoS One. 2013;8:e71901.
Breitling LP, Salzmann K, Rothenbacher D, Burwinkel B, Brenner H. Smoking, F2RL3 methylation, and prognosis in stable coronary heart disease. Eur Heart J. 2012;33:2841–8.
Talens RP, Jukema JW, Trompet S, Kremer D, Westendorp RG, Lumey LH, Sattar N, Putter H, Slagboom PE, Heijmans BT, PROSPER Group. Hypermethylation at loci sensitive to the prenatal environment is associated with increased incidence of myocardial infarction. Int J Epidemiol. 2012;41:106–15.
Jiang D, Zheng D, Wang L, Huang Y, Liu H, Xu L, Liao Q, Liu P, Shi X, Wang Z, Sun L, Zhou Q, Li N, Xu L, Le Y, Ye M, Shao G, Duan S. Elevated PLA2G7 gene promoter methylation as a gender-specific marker of aging increases the risk of coronary heart disease in females. PLoS One. 2013;8:e59752.
Perkins E, Murphy SK, Murtha AP, Schildkraut J, Jirtle RL, Demark-Wahnefried W, Forman MR, Kurtzberg J, Overcash F, Huang Z, Hoyo C. Insulin-like growth factor 2/H19 methylation at birth and risk of overweight and obesity in children. J Pediatr. 2012;161:31–9.
Irvin MR, Zhi D, Joehanes R, Mendelson M, Aslibekyan S, Claas SA, Thibeault KS, Patel N, Day K, Jones LW, Liang L, Chen BH, Yao C, Tiwari HK, Ordovas JM, Levy D, Absher D, Arnett DK. Epigenome-wide association study of fasting blood lipids in the genetics of lipid lowering drugs and diet network study. Circulation. 2014;130(7):565–72.
Guay SP, Voisin G, Brisson D, Munger J, Lamarche B, Gaudet D, Bouchard L. Epigenome-wide analysis in familial hypercholesterolemia identified new loci associated with high-density lipoprotein cholesterol concentration. Epigenomics. 2012;4:623–39.
Haas J, Frese KS, Park YJ, Keller A, Vogel B, Lindroth AM, Weichenhan D, Franke J, Fischer S, Bauer A, Marquart S, Sedaghat-Hamedani F, Kayvanpour E, Köhler D, Wolf NM, Hassel S, Nietsch R, Wieland T, Ehlermann P, Schultz JH, Dösch A, Mereles D, Hardt S, Backs J, Hoheisel JD, Plass C, Katus HA, Meder B. Alterations in cardiac DNA methylation in human dilated cardiomyopathy. EMBO Mol Med. 2013;5:413–29.
Movassagh M, Choy MK, Knowles DA, Cordeddu L, Haider S, Down T, Siggens L, Vujic A, Simeoni I, Penkett C, Goddard M, Lio P, Bennett MR, Foo RS. Distinct epigenomic features in end-stage failing human hearts. Circulation. 2011;124:2411–22.
Han P, Hang CT, Yang J, Chang CP, Bruneau B. Chromatin remodeling in cardiovascular development and physiology. Circ Res. 2011;108:378–96.
Cardinale JP, Sriramula S, Pariaut R, Guggilam A, Mariappan N, Elks CM, Francis J. HDAC inhibition attenuates inflammatory, hypertrophic, and hypertensive responses in spontaneously hypertensive rats. Hypertension. 2010;56:437–44.
Okamoto H, Fujioka Y, Takahashi A, Takahashi T, Taniguchi T, Ishikawa Y, Yokoyama M. Trichostatin A, an inhibitor of histone deacetylase, inhibits smooth muscle cell proliferation via induction of p21(WAF1). J Atheroscler Thromb. 2006;13:183–91.
Kong X, Fang M, Li P, Fang F, Xu Y. HDAC2 deacetylates class II transactivator and suppresses its activity in macrophages and smooth muscle cells. J Mol Cell Cardiol. 2009;46:292–9.
Matsushima S, Sadoshima J. The role of sirtuins in cardiac disease. Am J Physiol Heart Circ Physiol. 2015;309:H1375–89.
Zhang Y, Qiu J, Wang X, Zhang Y, Xia M. AMP-activated protein kinase suppresses endothelial cell inflammation through phosphorylation of transcriptional coactivator p300. Arterioscler Thromb Vasc Biol. 2011;31:2897–908.
Franklin S, Kimball T, Rasmussen TL, Rosa-Garrido M, Chen H, Tran T, Miller MR, Gray R, Jiang S, Ren S, Wang Y, Tucker HO, Vondriska TM. The chromatin-binding protein Smyd1 restricts adult mammalian heart growth. Am J Physiol Heart Circ Physiol. 2016;311:H1234–47.
Ai S, Yu X, Li Y, Peng Y, Li C, Yue Y, Tao G, Li C, Pu WT, He A. Divergent requirements for EZH1 in heart development versus regeneration. Circ Res. 2017;121:106–12.
Guo H, Ingolia NT, Weissman JS, Bartel DP. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature. 2010;466(7308):835–40.
Fichtlscherer S, Zeiher AM, Dimmeler S. Circulating microRNAs: biomarkers or mediators of cardiovascular diseases? Arterioscler Thromb Vasc Biol. 2011;31(11):2383–90.
Competing Financial Interests
The authors declare no competing financial interests.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Sharma, S., Mishra, A., Ashraf, M.Z. (2020). Involvement of Epigenetic Control and Non-coding RNAs in Cardiovascular System. In: Xiao, J. (eds) Non-coding RNAs in Cardiovascular Diseases. Advances in Experimental Medicine and Biology, vol 1229. Springer, Singapore. https://doi.org/10.1007/978-981-15-1671-9_6
Download citation
DOI: https://doi.org/10.1007/978-981-15-1671-9_6
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1670-2
Online ISBN: 978-981-15-1671-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)