Abstract
Antimicrobial peptides (AMPs) attack bacterial membranes selectively, killing microbes at concentrations that cause no toxicity to the host cells. This selectivity is not due to interaction with specific receptors but is determined by the different lipid compositions of the membranes of the two cell types and by the peculiar physicochemical properties of AMPs, particularly their cationic and amphipathic character. However, the available data, including recent studies of peptide-cell association, indicate that this picture is excessively simplistic, because selectivity is modulated by a complex interplay of several interconnected phenomena. For instance, conformational transitions and self-assembly equilibria modulate the effective peptide hydrophobicity, the electrostatic and hydrophobic contributions to the membrane-binding driving force are nonadditive, and kinetic processes can play an important role in selective bacterial killing in the presence of host cells. All these phenomena and their bearing on the final activity and toxicity of AMPs must be considered in the definition of design principles to optimize peptide selectivity.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Antimicrobial peptides
- Host defense peptides
- Selectivity
- Toxicity
- Peptide-membrane association
- Aggregation
- Hydrophobicity
- Amphipathicity
1 Introduction
The scientific and medical interest for antimicrobial peptides (AMPs), short peptides produced by most organisms as part of their innate immune defenses, derives from their wide-spectrum bactericidal properties and their possible application to fight drug-resistant bacteria. However, in view of clinical applications, the absence of significant toxicity is almost as important as a good activity. In this respect, one of the appealing properties of many AMPs is their cell selectivity, i.e., the ability to kill bacterial cells at concentrations significantly lower than those causing damage to cells of the host organism, at least in in vitro tests. Still, potential toxicity is commonly listed as one of the challenges limiting the clinical application of AMPs as systemic drugs (Hancock and Sahl 2006; Eckert 2011; Yeung et al. 2011; Seo et al. 2012; Carneiro et al. 2015; Pachón-Ibáñez et al. 2017), and therefore, several research efforts are devoted to understand and further improve AMP selectivity.
This chapter discusses AMP selectivity, the origin and the structural determinants of this property, the design strategies available to improve it, and the results of recent studies on the quantitative determination of peptide-cell association. Overall, the available data indicate that selectivity is the result of a complex interplay of several interconnected phenomena, including peptide association to target and host cells, peptide conformational equilibria, and AMP aggregation. Any modification to the peptide sequence and structure necessarily affects all of these processes, which must therefore be fully understood and considered in the rational design of new peptide or peptidomimetic molecules with improved selectivity properties. Our attempt is to give a critical overview of the available evidences, in order to provide a rationale for future efforts in this area. To this end, we have strived to derive, whenever possible, generalizations of the findings reported in the literature, but we have to stress from the beginning that the presence of exceptions to every rule is the norm, in such a diverse set as AMPs, also as a consequence of the complications mentioned above.
The different aspects of AMP selectivity have last been reviewed by Matsuzaki in 2009 (Matsuzaki 2009). Selectivity or toxicity has often been considered in general review articles on AMPs (Alba et al. 2012; Teixeira et al. 2012; Oddo and Hansen 2017; Hollmann et al. 2018). Some reviews have summarized our current knowledge on the structural determinants of AMP activity and selectivity (Takahashi et al. 2010; Huang et al. 2010; Strömstedt et al. 2010; Tossi 2011; Ruiz et al. 2014; Ebenhan et al. 2014a). Finally, for a recent discussion on how the interaction of AMPs with target and host cells determines their selectivity, see Savini et al. (2018).
As illustrated in other chapters of this book, AMPs have multiple functions, including anticancer, antifungal, and antiviral activities. For the sake of brevity and simplicity, in this chapter, we will essentially limit ourselves to discuss selectivity for bacterial versus host cells. Selectivity of anticancer peptides has been reviewed by Phoenix and coworkers (2012) and Harris et al. (2013).
2 AMPs Are Selective for Microbial Cells
AMPs have been isolated from natural sources based on their antimicrobial activity. The minimum inhibitory concentration (MIC, i.e., the lowest concentration of antimicrobial agent that inhibits the visible growth of a microorganism) (Wiegand et al. 2008) or the minimum bactericidal concentration (MBC, i.e., the minimal drug dosage killing at least 99.9% of the bacterial cells) (Lorian 2005) for AMPs is usually in the low μM range (Giacometti et al. 1998). AMPs are typically bactericidal, and therefore, the MIC and MBC values are usually similar (Giacometti et al. 1998).
The active concentration of a bioactive, therapeutically useful molecule must be much lower than the concentration causing toxic effects to the host cells. This property is quantified by the therapeutic index (TI), i.e., the ratio of the active concentration to the toxic concentration (see the legend to Table 11.1 for a detailed definition of these parameters). In the case of AMPs, whose main mechanism of bactericidal action is membranolytic (as discussed in Sect. 11.3), toxicity is most commonly assessed by measuring the lysis of erythrocytes (Fig. 11.1). Table 11.1 summarizes some TI values of natural and artificial AMPs, which are typically in the range 10–1000. However, it should be considered that unfortunately a strong variability is present in the literature regarding the definition of the toxic concentration, because different thresholds of lysed red blood cells (RBCs) are utilized to define the minimum hemolytic concentration (MHC), ranging from barely detectable to full hemolysis (see references cited in Table 11.1) (Bacalum and Radu 2015). In addition, MIC values depend on the specific strains tested in the assay.
Selective cytotoxicity in in vitro assays. Minimal inhibitory concentrations against different Gram- and Gram+ bacterial strains and minimal hemolytic concentration for the designed artificial AMP P5. The asterisk indicates that no hemolysis was observed for P5 in the peptide concentration range investigated (up to 100 μM). Adapted, with permission, from research originally published in Bobone et al. 2013, published by the European Peptide Society and John Wiley & Sons, Ltd.
When toxicity is assayed on other human cells, the results are generally not very different from those obtained using hemolysis (Table 11.2), but combining the two toxicity tests obviously provides a clearer picture of the selectivity of AMPs (Bacalum and Radu 2015).
AMP activity and toxicity are usually measured in separate assays performed under rather different conditions (for instance, regarding cell density, see Sect. 11.9) (Matsuzaki 2009). We have argued that experiments on bacterial and human cells in co-culture would provide a more stringent test of peptide selectivity (Savini et al. 2017, 2018). However, this approach has been employed only in a few cases. These studies, discussed in detail in Sect. 9.3, demonstrated that AMPs are selective even when acting on bacteria co-cultured with mammalian cells (Fig. 11.2a).
Selective targeting of bacterial cells by AMPs in vitro and in vivo. (a) Optical and fluorescence microscopy image of a labeled ubiquicidin analogue (visible by the green fluorescence) selectively binding to S. aureus bacteria, in co-culture with isolated human neutrophils (blue arrows) Adapted, with permission, from research originally published in Akram et al. 2015, published by The Royal Society of Chemistry. (b) Positron emission tomography image of a patient with an infection in the left hand (indicated by the arrow), traced with a radiolabeled ubiquicidin analogue. No significant peptide uptake in the contralateral hand was noted. The image was obtained 30 min after tracer administration (Reproduced, with permission, from research originally published in Akhtar et al. 2012). (c) Visualization of in vivo targeting of human α-defensin 5 (HD5) toward E. coli cells. The mesenteric vein was imaged intravitally in mice by two-photon laser scanning microscopy, 30 min after injection of E.coli cells expressing a green fluorescent protein (visualized in the left panel), and treatment with HD5 labeled with a red fluorescent probe (imaged in the center panel). Colocalization is demonstrated by the overlapped images (right panel). Scale bars, 50 μm. Adapted, with permission, from Lei et al. 2018. Copyright (2018) American Chemical Society
Several evidences indicate that AMPs are selective also in vivo. For instance, a large body of studies starting in 1999 (Welling) has shown that radiolabeled (Lupetti et al. 2003; Brouwer et al. 2008; Akhtar et al. 2012; Ebenhan 2014a) or fluorescent (Akram et al. 2015) AMPs can be used to image infections in vivo and can even discriminate between infection and inflammation, thanks to their specific binding to bacterial cells (Welling et al. 2000) (Fig. 11.2b, c). Among the peptides used for this purpose, there are defensins, cathelicidins, lactoferricins, histatins, artificial peptoids, and particularly sequences derived from ubiquicidin (Lupetti et al. 2003; Brouwer et al. 2008; Akhtar et al. 2012; Ebenhan 2014a, b; Dutta et al. 2017; Lei et al. 2018). Several imaging studies employed ubiquicidin 29–41 (Meléndez-Alafort et al. 2004; Akhtar et al. 2005; Vallejo et al. 2008; Gandomkar et al. 2009; de Murphy et al. 2010; Assadi et al. 2011; Ostovar et al. 2013; Saeed et al. 2013; Kahrom et al. 2014; Ebenhan et al. 2018; Bhatt et al. 2018), which has moderate activity and selectivity in the standard assays (MIC 40 μM, TI > 5) (Brouwer et al. 2006; Lupetti et al. 2008) but accumulates at the site of infection. For instance, one study reported overall values of sensitivity, specificity, and accuracy for infection detection of 100%, 80%, and 94% (Akhtar et al. 2005). Analogues of the ubiquicidin peptide have been used also for targeted delivery of traditional antibiotics to the infection site (Chen et al. 2015).
Although in vivo studies of the activity of AMPs abound, similar investigations characterizing their toxicity are more sparse (Mahlapuu et al. 2016). Some TIs derived from animal studies are summarized in Table 11.3, and the range of values is similar to that obtained in vitro.
The selectivity of AMPs for bacterial cells is demonstrated also by the fact that these peptides have been exploited in sensing elements that can detect infection (Mannoor et al. 2010; Shriver-Lake et al. 2012; Silva et al. 2014; Hoyos-Nogués et al. 2018), even in whole blood (Shi et al. 2017) or other complex biological samples (Qiao et al. 2017).
Overall, the results collected in the literature support an interesting selectivity of AMPs for target versus host cells. The origin of this property is necessarily related to the mechanism of action of AMPs.
3 Cellular Membranes Are the Main Target of AMPs
Selectivity is not surprising when a biomolecule associates to a specific receptor or protein (Le Joncour and Laakkonen 2018). However, this is not the case for most AMPs. In general, natural AMPs and their enantiomers comprising all D-amino acids have a comparable antimicrobial activity (Fig. 11.3), while any interaction of a peptide with a protein, due to the chirality of both systems, would be favored for one enantiomer over the other. As is often the case for AMPs, exceptions to this rule have been reported (Otvos et al. 2000; Bulet and Stocklin 2005; de la Fuente-Núñez et al. 2015), showing that the mechanism of action of a minority of AMPs could be receptor mediated. On the other hand, microbiological assays of membrane permeability and microscopic imaging of bacteria treated with AMPs clearly show that cell membranes are damaged and that, as a consequence, transmembrane gradients are dissipated (Tiozzo et al. 1998; Arcidiacono et al. 2009; Hartmann et al. 2010; Agrawal and Weisshaar 2018) (Fig. 11.4). Usually, membrane perturbation and bacterial killing are correlated, further supporting membrane disruption as the main bactericidal mechanism. However, also in this case, exceptions exist, indicating that a subclass of AMPs might act through different killing mechanisms (He et al. 2014; Friedrich et al. 2000). Finally, it is worth mentioning that AMPs usually are able to perturb the permeability of artificial membranes, comprising only phospholipids (Fig. 11.4c) (Orioni et al. 2009; Bocchinfuso et al. 2011; Braun et al. 2017; Savini et al. 2018). This observation demonstrates that the membrane-perturbing activity is purely the result of physicochemical interaction between the peptide and the lipid bilayer, and not the consequence of some biological process.
Activity of natural AMPs and their enantiomeric analogues. Comparison of the antibacterial activities (circles for MBC, squares for MIC) of enantiomeric peptides. Data refer to magainin 2 (Bessalle et al. 1990) (violet), cecropin A (Wade et al. 1990) (blue), melittin (Juvvadi et al. 1996) (dark green), LL-37 (Dean et al. 2011) (light green), KSLK (Hong et al. 1999) (yellow), temporin A (Wade et al. 2000) (orange; data refer to IC50 values), camel 48 (Oh et al. 2000) (red), V681 and analogues (Chen et al. 2006) (dark red; data refer to LC50 values), lactoferricin B analogues (Wakabayashi et al. 1999) (silver), and cecropin B (Bland et al. 2001) (black). The blue line is the diagonal of the plot (corresponding to identical activity for D and L enantiomers) and not a fit. A version of this figure with a more limited set of data has been published previously (Savini et al. 2018)
Fluorescence and electron microscopy images of the effects of AMPs on bacterial and artificial membranes. (a) Scanning electron micrographs of E. coli (top) and S. aureus (bottom) before (images on the left) and after (images on the right) treatment with the synthetic AMP PGYa (30 min, 10 μM). The images show a considerable roughening of the bacterial membranes and formation of blebs on the cell surface, in contrast to the smooth surfaces of untreated bacteria, providing a strong indication that the membrane is being considerably altered by the peptide. Adapted, with permission, from research originally published in Tiozzo et al. 1998 © Elsevier. (b) Fluorescence microscopy images of an E. coli cell attacked by the AMP cecropin A (0.5 μM). The leakage of periplasmic green fluorescent protein (GFP), shown by the green fluorescence, indicates perturbation of the outer membrane, while uptake of the DNA stain Sytox Orange (red fluorescence) demonstrates pore formation in the plasma membrane (Adapted, with permission, from research originally published in Agrawal 2018 © Elsevier). (c) Fluorescence microscopy images of perturbation of a giant unilamellar vesicle by the AMP PMAP-23. The top panels report the green fluorescence emission from carboxyfluorescein molecules entrapped inside the GUV, which were completely released after peptide addition (right). By contrast, the vesicle was still present after peptide addition, as indicated by the red fluorescence of rhodamine-labeled phospholipids located in the GUV bilayer (bottom panels). Taken together, these images demonstrate pore formation by the AMP. The vesicle diameter is about 20 μm. Adapted, with permission, from research originally published in Orioni et al. 2009 © Elsevier
Overall, literature data clearly demonstrate that membrane perturbation is the main mechanism of direct bacterial killing for most AMPs. Even for those AMPs that act through a different antibacterial mechanism (Nicolas 2009; Otvos 2017), the cell envelope is the first cell component that the peptides encounter, and they have to cross the extracellular membrane (when present) and the cell wall to reach the plasma membrane and eventually the cell interior.
Incidentally, the fact that AMPs target microbial membranes determines their broad-spectrum activity, their bactericidal, rather than bacteriostatic, mechanism of action and also the higher difficulty for bacteria in developing resistance against them (compared to resistance against conventional antibiotics acting on a protein target) (Perron et al. 2006; Otvos 2017).
4 Bacterial and Host Cells Have Different Membrane Structure and Composition
If membranes are the target, then it is conceivable that selectivity arises from a difference in membrane composition of the various cell types. Indeed, bacterial and eukaryotic cells have very different cell envelopes (Wang 2017). Bacteria can be divided into Gram-positive and Gram-negative, depending on whether they are colored by the Gram stain or not. This assay reflects differences in the composition of the cell envelope. In both cases, the plasma membrane is surrounded by a cell wall. However, in Gram-positive bacteria, this is formed by a thick peptidoglycan and lipoteichoic acid layer (40–80 nm). By contrast, in Gram negatives, a thin peptidoglycan layer (8 nm thick) is contained in a second (outer) membrane, with asymmetric composition: phospholipids are the main components of the inner leaflet, while the outer layer is mainly formed by lipopolysaccharides (LPS). On the other hand, eukaryotic cells only have the plasma membrane, with asymmetric lipid composition in the two leaflets of the bilayer (Fig. 11.5).
Schematic depiction of the structure of the cellular envelope in different cell types. The three panels, from left to right, schematize the structure of the cellular envelope in Gram- and Gram+ bacteria and in human cells, respectively. Proteins, glycolipids, and lipoteichoic acids (in Gram+ bacteria) have been omitted, for the sake of clarity. LPS, light gray; peptidoglycan, dark gray; PC, light green; PE, dark green; SM, light brown; PI, light gray; PG, red; PS, dark red; CL, orange; L-lysyl PG, blue; cholesterol, beige
In addition to the different structures of the cell envelope, important differences are present in the lipid composition of the cellular membranes. Tables 11.4 and 11.5 summarize the lipid content of bacterial and RBC membranes.
Bacterial membranes contain a significant fraction of negatively charged lipids: in Gram negatives, both membranes contain phosphatidylglycerol (PG, ~20% overall) and cardiolipin (CL, ~5% overall); in Gram-positive bacteria, the content of anionic lipids is much higher, again with PG and CL being the most important components (Malanovic and Lohner 2016). However, the membranes of these cells can also contain the positively charged l-lysyl-PG (LPG). In both cases, the main zwitterionic component is phosphatidylethanolamine (PE), and no sterols are present. Of course, the values for the composition reported in Table 11.4 are only approximate, since they change with the specific strain and growth conditions. In addition, lipid composition is not homogeneous over the cell surface (Renner and Weibel 2011; Oliver et al. 2014).
Human cells contain cholesterol and have no anionic phospholipids in the outer leaflet of their cell membrane. Some negatively charged glycolipids, such as gangliosides, are present on the cell surface (Miyazaki et al. 2012), but they are minor components in most cell membranes (with the exception of nerve cells) (Storch and Kleinfeld 1985). These properties are exemplified by RBCs, which are commonly used to test toxicity and selectivity (Table 11.5). For eukaryotes, the main zwitterionic components are phosphatidylcholine (PC), sphingomyelin (SM), and PE.
Overall, we can translate these differences in lipid composition in distinct physicochemical properties. Bacterial membranes contain more anionic lipids in the outer surface of their bilayers than eukaryotic cells. This difference combines with the additional negative charges conferred to bacterial cells by teichoic and teichuronic acids and LPS. Furthermore, the transmembrane potential of bacterial cells is more inside-negative than that of normal mammalian cells (Yeaman and Yount 2003). For all these reasons, bacteria have stronger electrostatic interactions with positively charged molecules than eukaryotic cells. Another difference is that bacterial membranes are more disordered and less well packed than those of eukaryotes, due to the lack of cholesterol. In addition, they contain larger amounts of “non-bilayer” lipids, with negative or positive values for the “intrinsic curvature,” such as PE, CL, and PA, or LPG, respectively (McMahon and Boucrot 2015; Malanovic and Lohner 2016). This property depends on the relative sizes of the phospholipid head-groups and acyl chains. Lipids where the cross-sectional area occupied by head-groups and tails is similar (e.g., PC, PG, PS) are said to have a cylindrical shape and pack well in locally flat bilayer structures (zero intrinsic curvature). By contrast, lipids where the head-group is smaller than the tails (e.g., PE or PA) favor concave shapes of the monolayer (negative curvature). The opposite is true for lipids with comparatively larger polar heads (e.g., LPG) which have a positive curvature (Koller and Lohner 2014).
The differences in lipid composition and in physical properties between bacterial and human cell membranes are considered to be the origin of AMP selectivity. Similar considerations on membrane composition (particularly regarding the content of anionic lipids and sterols) have been proposed to explain the selectivity for cancer cells (Hoskin and Ramamoorthy 2008; Schweizer 2009; Phoenix et al. 2012; Gaspar et al. 2013), fungi (van der Weerden et al. 2013; Rautenbach et al. 2016), protozoa (Rivas et al. 2009), and enveloped viruses (Aloia et al. 1993; Findlay et al. 2013), since in all cases the lipid distribution is different from that of a normal eukaryotic cell.
5 Lipid Composition Determines the Affinity of AMPs for Lipid Bilayers
The hypothesis of a selectivity based on differences in lipid composition has been tested by studying the interaction of AMPs with model membranes mimicking the composition of the natural bilayers. With liposomes, it is possible to vary the lipid composition at will and to measure both peptide-membrane association and peptide-induced membrane permeability (Bocchinfuso et al. 2011; Savini et al. 2018). The role of various membrane properties in AMP selectivity is summarized in the following sections.
5.1 Membrane Charge
In model membranes, the presence of anionic lipids increases peptide association to the bilayer and, as a consequence, peptide-induced leakage (Matsuzaki et al. 1989, 1995; Gazit et al. 1995; Abraham et al. 2005; Sood et al. 2008; Russell et al. 2010; Bobone et al. 2013; Golbek et al. 2017; Maturana et al. 2017). On the other hand, the positively charged lipid lysyl-PG, present in Gram+ bacteria, inhibits AMP activity (Nishi et al. 2004; Andra et al. 2011). These findings are a straightforward consequence of electrostatic interaction of the membranes with the positively charged AMPs (see Sect. 11.6). Regarding the anionic gangliosides present in the outer leaflet of eukaryotic membranes, Matsuzaki and coworkers (2012) demonstrated that, although their acidic moieties favor the association of AMPs to model membranes, this interaction does not lead to strong membrane perturbation, since the peptides remain trapped in the sugar region.
5.2 Cholesterol Content
Several studies also reported an AMP inhibitory effect of cholesterol. For instance, the presence of cholesterol inhibits the membrane-perturbing activity of magainin, pardaxin, LL-37, temporin L, human defensin HNP1, and other AMPs (Matsuzaki et al. 1995; Tytler et al. 1995; Hallock et al. 2002; Sood et al. 2008; Sood and Kinnunen 2008; Gonçalves et al. 2012; Verly et al. 2008; Wu et al. 2010; McHenry et al. 2012). The membrane-ordering effects of cholesterol in fluid bilayers are well established: insertion of the rigid ring structure of the sterol limits the possibility for trans-gauche isomerization for adjacent phospholipid tails, leading to an increase in bilayer order, packing, thickness, and rigidity (Henriksen et al. 2006; Mouritsen and Zuckermann 2004). All these effects could contribute to reduce peptide binding and membrane perturbation (McIntosh et al. 2002). However, the relevance of cholesterol for AMP selectivity has been recently questioned. While all the investigations listed above were performed on simple lipid mixtures, a comprehensive study by Ramamoorthy and coworkers on more realistic lipid compositions showed that cholesterol’s protective effect against AMPs does not occur in lipid systems containing raft domains and presenting phase separation (McHenry et al. 2012; Brender et al. 2012). There are examples where the activity of AMPs is not affected by the presence of cholesterol even in simple lipid mixtures (Bobone et al. 2013). On the other hand, Matsuzaki et al. (1995) demonstrated an AMP-inhibiting effect of cholesterol in real cells, by artificially varying the cholesterol content of RBCs.
It is worth mentioning that the inhibitory effect is specific of cholesterol, while ergosterol, present in fungal membranes, does not appear to inhibit peptide binding and activity to the same extent, in agreement with the specific activity of antifungal peptides (Sood and Kinnunen 2008; Gonçalves et al. 2012) and with the comparatively smaller effects of ergosterol on membrane order (Henriksen et al. 2006).
5.3 Intrinsic Curvature
The situation is less clear regarding the effect of the presence of negative curvature lipids (PE) in bacterial membranes. PE has been shown to inhibit pore formation by magainin, melittin, alamethicin, PMAP-23, and mastoparan X (Matsuzaki et al. 1998; Allende et al. 2005; Lee et al. 2005; Bobone et al. 2012). On the other hand, the activity of some AMPs is favored by the presence of PE (Schröder-Borm et al. 2003; Epand et al. 2006; Leite et al. 2015). Inhibition of pore formation by PE can be understood by considering that the peptides act by inserting in the head-group region of the membrane, thus imposing a positive curvature strain, which is released after a threshold of membrane-bound peptide concentration is reached, through the formation of membrane defects or pores. The presence of lipids with negative intrinsic curvature would counteract this mechanism (Matsuzaki et al. 1998, Lee et al. 2005). Similar considerations, on the other hand, suggest that PE can favor membrane binding of AMPs, by reducing the intrinsic curvature strain needed for peptide insertion in the polar region of the bilayer; an increased binding to PE-containing membranes has been reported for some AMPs (Schröder-Borm et al. 2003; Phoenix et al. 2015). However, reasoning only in terms of intrinsic curvature might be misleading. For instance, the peculiar lipid-lipid interactions made possible by the structure of PE could inhibit peptide insertion: this phospholipid contains a primary amine (lacking in PC), which allows it to form strong hydrogen bonds with phosphate or CO groups in other lipids (Lewis and McElhaney 2005). This H-bond network is responsible for the melting temperatures of PE lipids being higher than those of their corresponding PC analogues (Lewis and McElhaney 2005). This difference might lead to an increase in the energy needed to insert a peptide in the bilayer, or to open a pore, in the presence of PE. Finally, an additional mechanism invoked to explain different activities on vesicles lacking or containing PE involves peptide-induced formation of lipid domains (Epand et al. 2006). Overall, these considerations can explain why different final effects are observed on the membrane-perturbing activity of AMPs, depending on which of the various phenomena predominates in each specific case. In any case, it is difficult to ascribe a well-defined role in AMP selectivity to the PE content.
6 Thermodynamics of Peptide-Membrane Association
In principle, selectivity for different membrane compositions could result from two effects. AMPs could have a higher affinity for bacterial membranes than for human bilayers, or they could be more effective in perturbing the former, once inserted (Wimley and Hristova 2011). The data on model membranes presented above clearly indicate that differential binding is an important aspect of AMP selectivity.
AMPs are usually short (about 10–50 residues in length), and their sequences and structures have no common features, except for the cationic charge (most AMPs fall in the range of +2 to +4 e), and amphipathic character, with an overall content of about 50% hydrophobic residues (Wang 2017). The role of these properties is easily rationalized: charge imparts selectivity toward bacterial versus eukaryotic membranes, and apolar residues provide a hydrophobic driving force for binding and insertion into membranes, leading to perturbation of bilayer integrity. Indeed, peptides interacting only electrostatically usually do not cause significant membrane leakage, because their depth of insertion is too shallow (Wimley 2010a).
6.1 Hydrophobic and Electrostatic Driving Forces Are Nonadditive
Different treatments are used in the literature to describe peptide-membrane interactions, and comprehensive reviews are available on this topic (White and Wimley 1999; Wieprecht and Seelig 2002; Simon and McIntosh 2002; Santos et al. 2003; Seelig 2004; Wimley 2010a). Here we will just briefly mention that, since peptide-membrane association does not have a specific stoichiometry, it is not correctly described by a binding equilibrium, and it is better treated as a partition equilibrium between the water and the membrane phase (White and Wimley 1999; Wieprecht and Seelig 2002; Santos et al. 2003; Wimley 2010a). In this view, the main effect of Coulombic interactions can be described as an increase in local peptide concentration in the vicinity of the bilayer, according to Gouy-Chapman theory (Beschiaschvili and Seelig 1990; Wieprecht and Seelig 2002; Seelig 2004). However, the thermodynamic contributions of electrostatic and hydrophobic effects to the driving force of peptide water-membrane partition are not simply additive (Ladokhin and White 2001). This finding is due mainly to the different depths of polar and aliphatic moieties of phospholipids in the bilayer: charged groups are located on the surface of the membrane, well separated from the hydrocarbon core, and the physicochemical properties of the bilayer vary steeply in the head-group region (Wimley 2010a). As a consequence, the depth of insertion of a peptide in the membrane is determined by the interplay between hydrophobic effect and Coulombic forces (Wimley 2010a). Highly charged, hydrophilic molecules sit on the membrane surface, and strongly hydrophobic peptides insert into the hydrocarbon core, while cationic, amphipathic peptides are located at an intermediate position, which depends on their specific properties (Bocchinfuso et al. 2009; Farrotti et al. 2015). In turn, the depth of insertion in the bilayer modulates the intensity of electrostatic and hydrophobic contributions: strongly simplifying, one could say that the position of a peptide in the membrane determines the average distance between the peptide and the charged lipid moieties and the degree of insertion of the peptide in the water-free hydrocarbon core. An increase in peptide hydrophobicity ultimately reduces the effect of electrostatic interactions; on the other hand, augmenting the Coulombic forces diminishes the hydrophobic contribution to the binding free energy (Ladokhin and White 2001).
6.2 Multiple Interconnected Equilibria Modulate Peptide Activity and Selectivity
Several artificial peptides were designed having the required characteristics of cationic charge and amphipathic character. However, in many cases, such peptides turned out to be highly toxic (Dathe et al. 1996; Cornut et al. 1994; Bobone et al. 2013). These findings demonstrated that a cationic charge is not sufficient for specificity and provided a first indication of the complexities of peptide-membrane interaction discussed above. Peptides in solution can assume different conformations and aggregation states, and once membrane-bound, they can change conformation, orientation, insertion depth, and aggregation state (Fig. 11.6). All these phenomena are regulated by interconnected equilibria, and therefore they contribute in determining the final membrane-perturbing activity (Stella et al. 2004; Mazzuca et al. 2005; Gatto et al. 2006; Bobone et al. 2013). Every modification in peptide properties can affect all these processes (Gatto et al. 2006). In our opinion, this is the reason why the rational design of peptides with improved selectivity has met with limited success, and it has progressed through a trial-and-error process. Even so, several useful principles for the optimization of AMP selectivity have been defined.
7 Selectivity of AMPs Is Determined by Their Physicochemical Properties
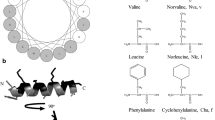
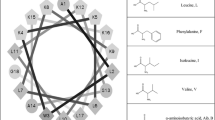
Helical peptides are the most abundant and best-characterized class of AMPs. Investigations on the structural determinants of AMP selectivity have mostly focused on this type of peptides. They are usually disordered in solution but attain a helical conformation when membrane-bound, with a spatially amphipathic distribution of the side chains, where most of the hydrophobic residues face toward the membrane center and the polar and charged residues are oriented toward the water phase. From a physicochemical point of view, they can be characterized by several parameters, such as charge, hydrophobicity, or amphipathicity. In addition to the considerations discussed in the previous section, on the multiple processes involved in peptide-membrane interactions, investigations on the role of each of these parameters are complicated by the fact that varying the sequence by a single amino acid substitution usually causes a variation in multiple physicochemical properties of the peptides (Wieprecht et al. 1997a; Dathe et al. 2001). For instance, inserting an additional cationic residue does not vary only the peptide charge but also its hydrophobicity and amphipathicity. However, some systematic studies have been performed where the authors tried to compare peptide sequences where multiple substitutions were inserted to cause the significant variation of one parameter only, while the others were kept as constant as possible (Dathe et al. 1997, 2001, 2002; Wieprecht et al. 1997a, b, c; Dathe and Wieprecht 1999; Giangaspero et al. 2001; Zelezetsky et al. 2005; Zelezetsky and Tossi 2006).
7.1 Cationic Charges Favor Selectivity
Based on the results on the importance of anionic lipids for the membrane activity of AMPs, it is not surprising that a positive correlation between peptide positive charge and antimicrobial activity and selectivity has often been described (Bessalle et al. 1992; Matsuzaki et al. 1997; Dathe et al. 2001; Giangaspero et al. 2001; Zelezetsky and Tossi 2006; Bobone et al. 2011). However, several studies reported that increasing cationicity above a certain level (+5, +8, or +9 depending on the specific case) is not beneficial and might even cause a decrease in activity or selectivity (Dathe et al. 2001; Giangaspero et al. 2001; Zelezetsky and Tossi 2006; Jiang et al. 2008). This last finding might be due to an overly shallow insertion of the peptide in the bilayer (Wimley 2010a) and to the nonadditivity of electrostatic and hydrophobic effects, discussed in Sect. 11.6.
One of the ways to increase the total positive charge of the peptide is C-terminal amidation, which is frequent in natural sequences and has the additional advantage of reducing susceptibility to proteolytic degradation (Huang et al. 2010; Mura et al. 2016). However, this approach to increase peptide selectivity is not generally valid, possibly because it also affects the stability of helical conformations in solution (Dennison et al. 2009) (see Sect. 11.8).
7.2 Hydrophobicity Is Necessary for Activity but Correlates with Toxicity: The Two Thresholds
The other main parameter influencing peptide affinity for membranes is hydrophobicity. Several studies concur to support the view that two hydrophobicity thresholds exist (Dathe et al. 1997; Kondejewski et al. 1999, 2002; Stark et al. 2002; Chen et al. 2007; Glukhov et al. 2008; Mojsoska et al. 2015; Uggerhøj et al. 2015). A first threshold hydrophobicity value must be reached to obtain peptides with significant membrane binding and insertion and thus endowed with antimicrobial activity. However, if hydrophobicity surpasses a second, higher threshold, toxicity is observed, because binding to neutral membranes becomes significant. The difference between these two thresholds is due to the electrostatic contributions to peptide binding to bacterial membranes. Therefore, an optimal range of hydrophobicity values exists, in which peptides exhibit antimicrobial activity, but no significant toxicity. Above a third, even higher threshold, activity decreases, because of peptide aggregation and lack of solubility (Gatto et al. 2006; Chen et al. 2007; Chu-Kung et al. 2010; Wimley 2010a). It is difficult to provide quantitative values for these thresholds, since different hydrophobicity scales are used in the literature. Just as an example, Deber and coworkers identified values of 0.4 and approximately 2 in the Liu-Deber scale for the activity and toxicity thresholds, respectively, for the hydrophobicity of the core segment of a series of model peptides (Glukhov et al. 2008).
It is interesting to note that hydrophobicity affects binding to neutral membranes more than to charged bilayers and hemolysis more than bactericidal activity (Wieprecht et al. 1997a; Dathe et al. 2002). The rationale underlying this finding is not immediately obvious, since the hydrophobic driving force is present for both membrane types, and therefore any variation in hydrophobicity should affect both antimicrobial activity and toxicity to the same extent. The experimental observations can be explained based on the nonadditivity of electrostatic and hydrophobic effects (Sect. 11.6).
In the case of highly hydrophilic peptides, modifications that increase hydrophobicity can enhance the antimicrobial activity (first threshold), without inducing strong toxicity (second threshold). Malmsten and coworkers have reported addition of hydrophobic oligopeptide stretches to the N- or C-terminus of the sequence as a way to improve peptide activity and selectivity (Pasupuleti et al. 2009; Schmidtchen et al. 2009, 2011, 2014). Comparison of different hydrophobic modifications indicated that tagging by oligo-Trp sequences at the C-terminus is the most effective one, leading to a substantial increase in activity, without significant enhancement of toxicity. Trp residues have peculiar properties, since they are known to have an affinity for membrane interfaces, thanks to their ability to interact both with hydrophobic moieties and with charged groups (through cation-aromatic interactions) (Yau et al. 1998). It has been speculated that the specificity-enhancing effect of Trp might be linked to the difficulty of inserting such a bulky residue in the tightly packed, cholesterol-containing membranes of eukaryotes (Pasupuleti et al. 2009; Schmidtchen et al. 2009, 2011, 2014). However, preferential interaction of Trp with cholesterol has also been hypothesized (de Kruijff 1990), although it is disputed (Holt et al. 2008), and in some cases, introduction of Trp residues has been linked to enhanced peptide toxicity (Oddo and Hansen 2017; Matsuzaki et al. 1997). Another common approach to increase the hydrophobicity of highly hydrophilic peptides is lipidation (Gatto et al. 2006). Shai’s group demonstrated that highly polar peptides, originally devoid of antimicrobial activity, can become antimicrobial, but not toxic, after this modification (Avrahami and Shai 2004; Malina and Shai 2005; Makovitzki et al. 2006, 2008). However, in other cases, lipidation led to strong toxicity (Chu-Kung et al. 2004; Laverty et al. 2010), or even to loss of activity, when it compromised peptide solubility (Toniolo et al. 1996; Gatto et al. 2006; Chu-Kung et al. 2010). These findings highlight the fine-tuning of AMP hydrophobicity needed for optimal activity and selectivity properties.
7.3 Excessive Amphipathicity Causes Toxicity
The total quantities of charged and hydrophobic residues provide only a very rough measure of peptide properties, since also their position in the sequence and structure are obviously important. Amphipathicity measures the degree of asymmetry in the distribution of polar and hydrophobic residues. This property can be quantified by the hydrophobic moment. This quantity is usually defined assuming an ideal helical structure and summing the vectors indicating the position of each residue with respect to the helix axis, multiplied by their respective hydrophobicity values (in analogy with the definition of an electric dipole). To compare sequences of different lengths, the mean hydrophobic moment can be obtained by normalizing for the number of amino acids (Eisenberg et al. 1982; Phoenix and Harris 2002). Peptide amphipathicity is a very important parameter for determining the free energy of membrane binding (Fernández-Vidal et al. 2007). As early as 1981, De Grado demonstrated that amphipathicity is sufficient to induce lytic activity in a helical peptide (De Grado et al. 1981). The specific value of the hydrophobic moment becomes particularly important for selectivity in an intermediate range of hydrophobicity values, when the hydrophilic or hydrophobic components of the peptide do not predominate in determining its behavior (Dathe and Wieprecht 1999; Dathe et al. 2002). Similar to what has been reported for hydrophobicity, an increased hydrophobic moment affects the activity on neutral membranes more than that on charged bilayers (Wieprecht et al. 1997b; Dathe et al. 2002). Increasing amphipathicity above a critical threshold results in strong interaction with neutral membranes, leading to toxicity (Wieprecht et al. 1997b; Dathe and Wieprecht 1999; Fernández-Vidal et al. 2007; Kindrachuk and Napper 2010). As discussed above for hydrophobicity, also in this case, it is difficult to provide a quantitative, generally valid value for this threshold.
Another measure of the distribution of polar and hydrophobic residues is the angle subtended by the polar face of the amphipathic helix, again assuming an ideal conformation and looking along the helix axis (Uematsu and Matsuzaki 2000). The available data on the role of this property in selectivity are limited, but a comprehensive study by Dathe and coworkers (2002) provided some indications. As discussed above, hydrophobicity and hydrophobic moment mostly affect the affinity for neutral membranes and thus the toxic activity. By contrast, in model membranes, the polar angle affects AMP ability to perturb the bilayer, after membrane binding: in charged bilayers, peptide-induced membrane leakage decreases with increasing polar angle, while it is essentially unaffected in neutral membranes (Dathe et al. 2002). However, the effects of the polar angle in cellular assays of activity and toxicity are more limited (Dathe et al. 2002).
8 Conformational and Aggregation Equilibria Play an Important Role in Membrane Selectivity: The Concept of Effective Hydrophobicity
All the considerations reported in the previous section are based on hydrophobicity values determined from the peptide amino acid composition and on amphipathicity calculated assuming an ideal helical conformation. In addition, a monomeric peptide state is always considered. However, as discussed in Sect. 11.6, peptides in solution and in the membrane attain specific ensembles of conformations, which can deviate significantly from an ideal alpha-helix. In addition, amphipathic peptides have a strong tendency to aggregate (Fig. 11.6). Conformational equilibria and self-assembly affect the degree to which the hydrophobic moieties of AMPs are exposed to the water phase and therefore modulate the hydrophobic driving force for membrane binding (Bobone et al. 2013). Similarly, water-membrane partition is affected by the peptide conformation, orientation, and depth of insertion in the bilayer. Based on these considerations, in our opinion, AMP selectivity is not determined by the “ideal” peptide hydrophobicity or amphipathicity but by what we call “effective” hydrophobicity and amphipathicity, i.e., the value these parameters assume in the actual conformation and aggregation state attained by the peptide in solution and in the bilayer (Bobone et al. 2013; Uggerhøj et al. 2015).
If peptide conformation and aggregation influence peptide hydrophobicity, the opposite is also true: high hydrophobicity and amphipathicity values favor a stable secondary structure by allowing the formation of intramolecular interactions between apolar residues (Fernández-Vidal et al. 2007). Peptide structure is influenced by self-assembly processes, too (Sal-Man et al. 2002). Therefore, in order to fully understand the determinants of peptide selectivity, conformational and self-assembly equilibria should be considered.
8.1 Helicity Correlates with Toxicity
A correlation between peptide helicity and toxicity has been reported in many studies (Tossi et al. 2000; Giangaspero et al. 2001; Zelezetsky et al. 2005; Chen et al. 2005; Khandelia and Kaznessis 2006; Zhang et al. 2011; Mangoni et al. 2011; Chapuis et al. 2012; Bobone et al. 2013; Cherry et al. 2014). In addition, helix-destabilizing Gly or Pro residues are often present close to the center of the sequence of natural, selective AMPs that attain a helical conformation in membranes (Tossi et al. 2000, Bobone et al. 2013). These amino acids are important for peptide selectivity, since their deletion, substitution, or insertion significantly affects toxicity, through the perturbation of the secondary structure (Thennarasu and Nagaraj 1996; Zhang et al. 1999; Shin et al. 2001; Yang et al. 2002, 2006a, b; Lee et al. 2004, 2007; Song et al. 2004; Carotenuto et al. 2008; Bobone et al. 2013; Wang et al. 2015). An increase in selectivity with a reduction in helical structure has been reported for other helix-breaking strategies, such as the insertion of d-amino acids (Shai and Oren 1996, 2001; Oren and Shai 1997; Papo et al. 2002; Chen et al. 2005; Zhu et al. 2007c; Kaminski and Feix 2011; Nan et al. 2012; Huang et al. 2014) or peptoid residues (N-substituted glycines, which lack a H-bonding proton on the N backbone atom and comprise a flexible main-chain methylene group) (Song et al. 2005; Zhu et al. 2007a, b; Kim et al. 2010). Incidentally, these non-proteinogenic residues have the added advantage of reducing peptide susceptibility to proteolysis (Papo et al. 2002; Kim et al. 2010).
The correlation between helicity and toxicity was tentatively explained by proposing that a stable helical conformation enhances the peptide propensity to aggregate (Kindrachuk and Napper 2010; Vermeer et al. 2012): in helical amphipathic peptides, the hydrophobic face of the helix is totally exposed to the aqueous phase, and therefore aggregation is hydrophobically favored. Aggregation, in turn, would inhibit crossing of the LPS layer and cell wall and thus access to the plasma membrane of bacteria (see below). However, helicity normally affects toxicity, rather than antibacterial activity. In addition, in the peptides we investigated, aggregation was significant only at concentrations higher than the membrane-perturbing values, and therefore it was not relevant for activity (Bobone et al. 2013). Probably a higher tendency to aggregate and an enhanced toxicity are just two independent consequences of the hydrophobicity induced by a stable helical structure, but the lack of selectivity in helical peptides is not caused by peptide aggregation (see also below). Based on a systematic study in which a central proline residue was moved along the sequence or deleted, we obtained data supporting an alternative explanation (Bobone et al. 2013). In a perfectly helical, amphipathic structure, the apolar residues are completely exposed on the hydrophobic face of the helix. Even though short peptides are often unstructured in water, helical conformations can be at least partially populated, also thanks to the stabilization due to the interaction between hydrophobic side chains aligned along the helix (a motif often called “leucine zipper” (Asthana et al. 2004)). White and coworkers reported a strong correlation between the amphiphilicity of a peptide sequence in an ideal helical conformation and both the degree of helicity in solution and the affinity for neutral membranes (Fernández-Vidal et al. 2007). Destabilization of the helical conformation allows the peptide to fold onto itself, hiding the apolar side chains from the water phase, and reducing the effective hydrophobicity of the peptide, and thus the driving force for binding neutral membranes (Bobone et al. 2013; Büttner et al. 1992). Destabilization of the helix also increases the entropic cost of membrane binding, since association to the bilayer is normally followed by peptide structuring (Zelezetsky et al. 2005). Interestingly, once membrane-bound, the helix-destabilizing modifications do not preclude the attainment of an amphipathic helical conformation (Bobone et al. 2013; Orioni et al. 2009; Oren and Shai 2000). Therefore, variations in the free energy of membrane binding are determined essentially by changes in the solution conformation. Unstructured conformations would inhibit binding to neutral membranes but would affect only marginally the affinity for charged bilayers, leading to enhanced selectivity. Related to this interpretation is the concept of position-dependent hydrophobicity: hydrophobic residues in unstructured regions of the peptide contribute to the effective hydrophobicity and to toxicity less than those in helical segments (Tachi et al. 2002).
The effective, conformation-dependent hydrophobicity can be calculated from peptide structures (Gaillard et al. 1994), but it can also be determined experimentally. Reversed-phase chromatography retention times have resulted to be an accurate measure of the effective hydrophobicity of peptides (Krause et al. 1995; Zhou et al. 1990; Kim et al. 2005). Interestingly, strong correlation between RP-HPLC retention times and the hemolytic activity of AMPs has been reported (Blondelle and Houghten 1991; Kondejewski et al. 1999; Tachi et al. 2002).
8.2 Imperfect Amphipathicity Optimizes Selectivity
Another idea related to effective hydrophobicity is imperfect amphipathicity. Insertion of a polar/charged residue in the hydrophobic face of an amphipathic helix has proven to be a reliable method to increase AMP selectivity (Asthana et al. 2004; Chen et al. 2005; Ahmad et al. 2006, 2009a, b; Hawrani et al. 2008; Pandey et al. 2010, 2011; Jiang et al. 2011, 2014, 2018; Son et al. 2013; Dalzini et al. 2016; Zhang et al. 2016). Hodges and coworkers even termed these misplaced polar residues “specificity determinants” (Jiang et al. 2011, 2014, 2018), even though exceptions to the selectivity-improving effect of this approach have been reported (Wang et al. 2018). Interestingly, an imperfectly amphipathic structure is a common property of many natural, selective AMPs (Wimley 2010b; Orioni et al. 2009). Again, this approach reduces the hydrophobic driving force for binding neutral membranes. On the other hand, antimicrobial activity is usually not affected significantly by these changes: in charged bilayers, binding takes place all the same, thanks to the electrostatic attraction; once membrane-bound, the peptide is able to attain a (possibly distorted) helical conformation, as demonstrated by spectroscopic and simulative studies (Hawrani et al. 2008; Orioni et al. 2009). In the bilayer, imperfect amphipathicity will contribute to membrane disruption, by driving some polar head-groups in the hydrophobic core of the membrane, as we observed for PMAP-23 (Orioni et al. 2009) (Fig. 11.7). This type of membrane activity has been termed “interfacial activity” by Wimley (2010b). Finally, it is worth mentioning that we recently observed an effect of imperfect amphipathicity on toxicity also in the case of peptidomimetic antimicrobial molecules (Konai et al. 2018). Two small amphipathic, cationic molecules were characterized by the same compositional hydrophobicity but had very different selectivity. By combining molecular dynamics simulations and RP-HPLC retention times, we demonstrated that this was due to imperfect amphipathicity and lower effective hydrophobicity of the selective analogue compared to the toxic compound.
Interfacial activity of an imperfectly amphipathic AMP. Effects of the imperfectly amphipathic AMP PMAP-23 on the structure of a lipid bilayer, as observed in MD simulations. Two charged residues located on the hydrophobic side of the helix drive three phospholipid head-groups and some water molecules into the hydrophobic core of the membrane. Water is represented in cyan, phospholipids in gray, and phospholipids’ phosphorus atoms as yellow spheres. The peptide backbone is shown in gray, charged side chains in red, polar amino acids in orange, apolar residues in blue, and prolines in green. The lipid composition was POPG/POPC (1:3 mol/mol).The bottom panel reports the density map of the lipid phosphorus atoms (red) and of the peptide backbone atoms (blue). Adapted, with permission, from research originally published in Orioni et al. 2009 © Elsevier
8.3 Effects of Peptide Aggregation in the Aqueous Phase on Activity and Toxicity Are System Dependent
AMPs, due to their amphipathic nature, are susceptible to aggregation in water (Tian et al. 2015). Some peptides oligomerize through the interaction of the apolar sides of their amphipathic helices (Oren et al. 1999; Asthana et al. 2004; Raimondo et al. 2005; Ahmad et al. 2006), while others form micellar structures (Liu et al. 2009; Wang et al. 2010; Joshi et al. 2015; Lin and Grossfield 2015; Haney et al. 2017; Lei et al. 2018), fibrils (Tu et al. 2007; Chen et al. 2010; Chen and Liang 2013; Shankar et al. 2013; Chairatana and Nolan 2014; Ravi et al. 2015), or even hydrogels (Veiga et al. 2012; McCloskey et al. 2014; Haney et al. 2017). Often the aggregates disassemble into monomers once membrane-bound (Ghosh et al. 1997). The critical concentration for self-assembly can vary significantly from one specific case to the other, also depending on the experimental conditions and particularly on salt concentration.
The results of experimental and theoretical studies on the effects of aggregation on peptide activity and selectivity are extremely contradictory. Both negative (Feder et al. 2000; Kustanovich et al. 2002; Chen et al. 2006, 2007; Daschbach et al. 2012; Lin and Grossfield 2015; Farrotti et al. 2017; Haney et al. 2017; Bagheri et al. 2018; Zou et al. 2018) and positive (Sal-Man et al. 2002; Avrahami and Shai 2002; Liu et al. 2009; Chen et al. 2010; Joshi et al. 2015; Ravi et al. 2015; Lei et al. 2018) correlations between aggregation and activity have been reported, as well as lack of activity changes following aggregation (Chen and Liang 2013). Similarly, some studies found that toxicity was not significantly affected by aggregation (Lei et al. 2018), while others reported an increase (Chen and Liang 2013; Lin and Grossfield 2015) or a decrease (Kustanovich et al. 2002; Chen et al. 2006; Chen et al. 2007) in selectivity upon self-assembly. Shankar et al. (2013) suggested that toxicity of self-assembled lipopeptides depends on the specific structure of the fibrillar aggregates.
The reported discrepancies are most likely due to the fact that peptide aggregation is usually controlled by varying the peptide properties or by modulating electrostatic interactions by changing the ionic strength of the solution. It is therefore difficult to discriminate between the direct effects of these changes (e.g., an increase in hydrophobicity) and the consequence of the variations they induce in aggregation. One approach to solve this problem is covalent linking of the monomers (Sal-Man et al. 2002; Dempsey et al. 2003), but it does not exactly mimic self-assembly driven by hydrophobic interactions.
Thermodynamic considerations on the interconnected equilibria involved in AMP activity indicate a possible positive role of aggregation in enhancing peptide selectivity. Aggregation, which is hydrophobically driven, reduces the effective peptide hydrophobicity by hiding the apolar moieties in the molecule from the aqueous phase. As a consequence, the hydrophobic driving force for membrane binding is reduced in the aggregates (Stella et al. 2004; Mazzuca et al. 2005; Gatto et al. 2006; Chen et al. 2007; Chu-Kung et al. 2010; Farrotti et al. 2017). Considering the various hydrophobicity thresholds discussed above, aggregation could therefore lead to a reduced toxicity. At the same time, preassembly of AMPs causes a local release of a high concentration at a single site in the membrane, and this could cause higher activity (Ravi et al. 2015). In addition, computational studies suggested that self-assembly could lead to membrane selectivity also by affecting the kinetics of membrane binding (Lin and Grossfield 2015): binding to host mammalian membranes will be slow and inefficient as long as the lipopeptides are micellized in solution, while binding to the bacterial surface will still be efficient, thanks to electrostatic interactions and to the higher fluidity of the membrane. On the other hand, in cellular assays, the large size of the aggregates, compared to monomers, could impair selectivity: preassembled AMPs might be unable to cross the LPS layer or the cell wall and thus to reach the plasma membrane of bacteria. At the same time, they would still be able to interact with the “naked” membrane of host cells (Oren and Shai 2000; Kustanovich et al. 2002; Sal-Man et al. 2002; Mangoni and Shai 2009).
Discussing aggregation, it is important to note that this phenomenon reduces susceptibility to proteolytic degradation and affects the pharmacokinetics and pharmacodynamics in vivo (Raimondo et al. 2005; Tu et al. 2007; Chen and Liang 2013; Lei et al. 2018). It is also worth mentioning that human α-defensin 6 (HD6) has negligible direct killing activity but prevents infections by self-assembling into a network of fibrils that capture pathogens and thus contrast microbial invasion (Chairatana and Nolan 2014).
9 AMP Binding to Cells
As discussed in the previous sections, AMP selectivity is usually interpreted essentially on the basis of the different affinities observed in liposome studies for bilayers mimicking the membranes of bacteria or eukaryotes. However, quite surprisingly, peptide affinities toward the two types of cells are largely uncharacterized. In addition, if peptide activity is modulated by a cell-binding equilibrium, it should depend on the density of cells, but antimicrobial activity and toxicity assays are usually carried out using standardized, fixed cell densities, which are not necessarily representative of the cell concentrations present in a typical infection site (Savini et al. 2018). Finally, bactericidal and hemolytic activities are routinely determined in separate assays, but when the two cell populations are present at the same time, they compete for peptide association. All these aspects have received limited attention, until quite recently. Biophysical studies on model membranes allow the determination of both membrane-binding and bilayer-perturbing activities, while microbiological studies usually report activities only in terms of total peptide concentration. We recently reviewed the few studies that are trying to apply to cellular experiments the same quantitative approaches normally used with model systems (Savini et al. 2018). Here, only the aspects relevant to AMP selectivity are summarized.
9.1 AMPs Have a Higher Affinity for Bacterial Than for Eukaryotic Cells
Only a handful of studies reported data on AMP binding to bacterial and eukaryotic cells. As soon as 1988, Bruce Merrifield and his group (Steiner et al. 1988) measured binding of cecropin A and some of its analogues to Escherichia coli, B. megaterium, B. thuringiensis, and P. aeruginosa cells and to erythrocytes. While binding to the bacteria was significant (between 70% and 80% for the natural peptide, under the conditions studied), no detectable association was observed for RBCs, at a cell density corresponding to a membrane area similar to that present in the experiments with bacteria. Welling et al. (2000) measured the binding of defensins 1–3, ubiquicidin, and human lactoferrin to bacteria and activated murine peritoneal leucocytes. In the presence of the same cell density (2 · 107 cell/mL), the peptides bound 5–500 times more efficiently to bacteria than to mammalian cells, even though the latter are much bigger. Similarly, Ferro-Flores et al. (2003) reported that, in the presence of 2 · 107 cell/mL, an ubiquicidin analogue was 35% bound in the case of bacteria (S. aureus) while less than 4% in the case of human tumor cell lines LS174T and ACHN (which, again, are significantly bigger than bacteria). Comparable results have been reported for two ubiquicidin analogues (approximately 45–100% binding to S. aureus while only 10% to leukocytes, in the presence of 2 · 105 cell/mL), although in this case, selectivity was surprisingly observed also for an anionic peptide used as negative control (Ebenhan et al. 2014b). Wimley and coworkers (Starr et al. 2016) measured the binding of the artificial AMP ARVA to E. coli, S. aureus, and RBCs. In all cases, association to bacteria was more favorable than to RBCs. After accounting for the differences in cell size, the authors estimated that the affinity for bacterial membranes was more than two orders of magnitude higher than for erythrocytes. Finally, an analogue of LL37 was reported to bind E. coli, S. aureus, and M. smegmatis, but not to hepatic cells, under conditions of comparable cell numbers (Dutta et al. 2017). Overall, these data indicate that the differential affinity routinely observed with model bilayers is present also for the membranes of real cells.
9.2 Activity and Toxicity Are Cell Density Dependent
Another aspect that has remained essentially uncharacterized until very recently is whether the activities of AMPs depend on the density of cells present in the assays. Based on a partition equilibrium, the fraction of membrane-bound peptide obviously depends on the concentration of cells in the sample. Therefore, it is to be expected that MIC/MBC/MHC values depend on the concentration of cells used in the assays. In broth dilution assays of antimicrobial activity, the recommended value for the initial cell density (inoculum) is 5 · 105 cell/mL (Patel et al. 2012), which was selected for minimizing false-positive and false-negative results in the clinical practice (Wiegand et al. 2008). However, bacterial cell densities in clinically relevant infections range from 1 to 109 cell/mL. Similarly, hemolytic activity assays are normally performed with 5 · 108 cell/mL, which is 1/10 of the cell density in whole blood (Savini et al. 2018 and references therein). Matsuzaki (2009) pointed out that the cell densities in the two assays are very different, also considering that the membrane area of an erythrocyte is approximately ten times bigger than that of a typical bacterium. Therefore, he wondered if TI values such as those reported in Table 11.1 are an experimental artifact due simply to the fact that more peptide is probably needed to kill a higher number of bigger cells.
In the case of traditional antibiotics, it is well known that the MIC often depends on the size of the bacterial inoculum (“inoculum effect”). By contrast, in the case of AMPs, this possible dependence has been investigated only in very few studies (Savini et al. 2018). In the 1990s, Levison et al. (1993) reported that the bactericidal activity of magainins against P. aeruginosa was inoculum dependent above 3 · 105 cell/mL, but it did not vary if the inoculum was reduced below this value. Similarly, Jones et al. (1994) observed an inoculum effect for lactoferricin B against E. coli, with a plateau at inoculum densities below 106 cell/mL. Ulrich and coworkers measured MIC values for gramicidin S and PGLa at two cell density values and observed a cell density dependence (Hartmann et al. 2010). More recently, we measured the MBC values for a fluorescent analogue of PMAP-23 in the presence of different E. coli cell densities (Savini et al. 2017). Also in our case, the MBC increased with inoculum size but reached a plateau at densities of 5 · 106 cell/mL and below. A similar trend was reported by Poon and coworkers for pexiganan (Jepson et al. 2016). Finally, a recent study reported an inoculum effect also for LL-37 (Snoussi et al. 2018).
Overall, these studies show that AMP activity is strongly dependent on the density of cells in the assay, with a linear (Savini et al. 2017) or sublinear (Jepson et al. 2016) trend. If a threshold concentration must be reached in the membrane to form pores (Melo et al. 2009), it is obvious that more membrane-bound peptide molecules are needed to kill a higher number of cells (Savini et al. 2017, 2018). Therefore, under conditions of relatively high cell densities, where the peptide is completely bound to the cells, a strong cell density dependence of the activity is expected. Data by Jepson et al. (2016) and Snoussi et al. (2018) indicate that this effect might be due also to peptide sequestration by strong binding to killed bacterial cells. We tentatively explained the plateau in the low cell density regime as due to the cell-binding equilibrium. At low cell densities, some of the peptide remains free in solution, and this fraction increases with decreasing concentrations of cells. As a consequence, the two effects (when there are less cells to kill, also a lower fraction of peptide is cell-bound) cancel each other, leading to a plateau in the total peptide concentration needed in the sample to kill the bacteria (Savini et al. 2017, 2018).
Interestingly, we observed a cell density dependence also for the hemolytic activity, with a plateau at densities below 107 cell/mL (Savini et al. 2017).
9.3 Competition for Cell Binding in Co-culture Experiments Might Be Regulated by Kinetic Phenomena
Since both antimicrobial activity and toxicity seem to depend on the density of cells, the effective selectivity also depends on the value of this parameter used in the two assays (MIC and MHC). The quantity of peptide that binds to a type of cell or to the other is determined by the respective affinities but also by the concentration of cells of each type. In the end, the peptide is active and/or toxic if the respective threshold of bound peptide needed for membrane perturbation is reached. In principle, under conditions where the host cells are in large excess (as in the case of systemic treatment of an infection), lack of toxicity is expected even if the affinities for the two cell types are similar (Matsuzaki 2009; Savini et al. 2017).
A limited number of studies have tested peptide activity and toxicity in assays where both bacteria and mammalian cells were present. Mor and coworkers showed that AMPs bound to RBCs are able to transfer to microbial cells, exerting their activity (a phenomenon they termed “affinity-driven molecular transfer”) (Feder et al. 2001). Derivatives of lentivirus lytic peptides killed P. aeruginosa bacteria interacting with cultured human airway epithelial cells, at peptide concentrations that only moderately affected the cell monolayer (Phadke et al. 2003). Fluorescence microscopy images showed that in a co-culture of S. aureus and human cells (endothelial cells or neutrophils), AMPs concentrate on the bacterial cells (Matsuzaki 2009; Akram et al. 2015) (Fig. 11.2a). Chen and Liang (2013) showed that the artificial AMP CL-1 is able to kill selectively S. aureus in co-culture with human cells, and Malmsten and coworkers reported activity without hemoliticity in bacteria-supplemented blood for an engineered AMP (Schmidtchen et al. 2011). A striking evidence of selectivity is provided by the fact that some AMPs (e.g., LL-37) are able to kill S. aureus bacteria internalized into mammalian cells (Noore et al. 2012). Selectivity in co-culture has been reported also for AMP-inspired systems, such as peptidomimetics, cationic peptidopolysaccharides, and peptide hydrogels (Salick et al. 2007; Li et al. 2012; Konai et al. 2018). Some co-culture data have been reported also for antifungal, antiprotozoan, and anticancer activities. For instance, an analogue of the antifungal peptide PAF26 concentrated on fungal cells in co-culture with human lung epithelial cells (Mendive-Tapia et al. 2016); an artificial anticancer peptide concentrated in cancer cells, co-cultured with primary cells (Chen et al. 2014). Some AMPs (e.g., dermaseptins or NK-lysin) are able to kill protozoan parasites such as P. falciparum or T. cruzi inside human cells, disrupting the plasma membrane of the intracellular parasites without harming that of the host (Ghosh et al. 1997; Krugliak et al. 2000; Jacobs et al. 2003; Gelhaus et al. 2008).
Based on association equilibria, when the two cell types are present at the same time, competition for peptide binding should take place: the antimicrobial activity and/or the toxicity of the peptides should be inhibited by sequestration of a fraction of the peptide molecules due to binding to the other cell population. Very surprisingly, we observed that this is not the case (Savini et al. 2017). We measured both bacterial killing and hemolysis for analogues of PMAP-23 and esculentin, in a mixed population of E. coli and erythrocytes, and compared these results with the traditional assays performed on the two cell types separately (Fig. 11.8). The activities on bacteria and on RBCs were essentially unaffected by the presence of the other cell population, in contradiction with the predictions based on binding equilibria. Data from Wimley’s lab provide further support to the conclusion that out of equilibrium, kinetic phenomena are at play when two cell populations are present at the same time. They showed that the results of such experiments depend on the order of addition of the different components: like in our case, no change in the MIC was observed when AMPs were added to a mixture of bacteria and RBCs or to bacteria alone. However, the antimicrobial activity was significantly inhibited when the peptide was incubated with RBCs first, and then both were added to the bacterial culture (Starr et al. 2016) (Fig. 11.8). Actually, while equilibrium processes can be invoked before the cell membranes are perturbed, it is easy to realize that this approach is too simplistic in the case of bilayer disruption, which allows access to multiple additional binding targets (e.g., inside the cell) (Snoussi et al. 2018).
Antimicrobial and hemolytic activity in assays with both bacterial and erythrocytes. Left panel, bactericidal (blue) and hemolytic (red) activities of the AMP DNS-PMAP23 in the presence of both bacteria and erythrocytes (squares) or of one cell type only (circles). Both activities are only slightly affected by the presence of the other cell population. 4.5 × 107 E. coli cells/mL, 4.5 × 108 RBCs/mL. (Reproduced with permission from Savini et al. 2017 https://pubs.acs.org/doi/abs/10.1021/acschembio.6b00910 (Copyright 2017, American Chemical Society)). Further permissions related to the material excerpted should be directed to the ACS. Right panel: effect of incubation time on MIC values in assays with both bacteria and erythrocytes. The artificial AMP ARVA-D was tested against E. coli (red) and S. aureus (blue) under various conditions. “MIC” represents measurements done in the absence of RBCs. All other experiments include 109 human RBC/mL. Time zero represents the experiments in which RBC and bacteria were first mixed, followed by peptide addition, i.e., no preincubation with either cell type. Negative times represent peptide preincubation with bacteria before the addition of RBCs. Positive times represent peptide preincubation with RBCs, followed by addition of bacteria. Points plotted at 20 μM had MIC values ≥20 μM. Significant inhibition of peptide antimicrobial activity due to the presence of RBCs was observed only in the case of preincubation with erythrocytes. Proteolytic degradation effects can be ruled out, since ARVA-D is a peptide comprising all d-amino acids. (Adapted with permission from Starr 2016 (Copyright 2016, American Chemical Society))
Overall, quantitative measurements of peptide interactions with cells confirmed that AMPs have a higher affinity for bacterial than for host cells. Experiments performed with varying cell densities indicated that both activity and toxicity depend on this parameter (even though a plateau is observed at low cell densities) and that therefore the measured selectivity depends on the specific conditions of the experiments. Finally, experiments with mixed bacterial and eukaryotic cells showed that, contrary to expectations based on equilibrium considerations, competition for peptide binding does not lead to a loss in activity. This finding definitely warrants further co-culture studies.
10 Concluding Remarks
A large body of studies has been devoted to characterize, understand, and improve the selectivity of AMPs. These data support the view that selectivity arises due to the different lipid composition of bacterial and host cell membranes. AMPs are able to discriminate between the two types of bilayers, thanks to their physicochemical properties. We can summarize here the main guidelines for optimization of peptide selectivity:
-
Increasing the cationic charge, by C-terminal amidation, substitution of anionic residues, or insertion of cationic amino acids in the polar side of the peptide structure, leads to a better selectivity. However, an excessive increase in positive charge, above a threshold that depends on each specific case, might be ineffective or even detrimental.
-
Reducing hydrophobicity, amphipathicity, and helicity is an effective strategy. These properties are necessary for activity, since they are responsible for the hydrophobic driving force for membrane binding and for insertion in the bilayer. However, several studies have demonstrated that they affect toxicity more than activity. This finding is probably a consequence of the nonadditivity of electrostatic and hydrophobic effects. Therefore, an intermediate range of hydrophobicity and amphipathicity values optimizes selectivity.
-
The real determinants of selectivity are the effective, conformation-dependent, hydrophobicity and amphipathicity values (rather than the parameters determined based on an idealized conformation). They can be optimized by introducing:
-
Polar residues in the hydrophobic side of the peptide helix (imperfect amphipathicity)
-
Helix-breaking residues, such as Pro, Gly, d-amino acids, or peptoids
-
-
Aggregation in the aqueous phase modulates selectivity, too.
In this chapter, we tried to generalize and simplify as much as possible the results collected over many years of studies. However, the readers that had the patience to follow us until here must have realized that the literature on AMP selectivity is full of exceptions and contradictions. Peptide association to bacterial and host cells is modulated (in a nonadditive way) by electrostatic and hydrophobic interactions. The effective peptide hydrophobicity is determined by peptide conformation and aggregation state. Therefore, AMP selectivity is finely regulated by interconnected binding, aggregation and conformational equilibria. Any variation in peptide property will affect all of these phenomena. Therefore, in order to predict, or at least to understand, the effect of peptide modifications on the final selectivity, all the possible processes involved in peptide behavior in the aqueous and membrane phases must be considered.
We still do not fully understand what happens when AMPs act on bacteria and human cells together. Recent data indicate that in this case even the complex scenario outlined above is overly simplified, since kinetic phenomena probably have to be taken into account. Further studies in this area are definitely warranted and hold the promise to provide a better understanding of AMP selectivity.
References
Abraham T, Lewis RN, Hodges RS, McElhaney RN (2005) Isothermal titration calorimetry studies of the binding of the antimicrobial peptide gramicidin S to phospholipid bilayer membranes. Biochemistry 44(33):11279–11285
Agrawal A, Weisshaar JC (2018) Effects of alterations of the E. coli lipopolysaccharide layer on membrane permeabilization events induced by Cecropin A. Biochim Biophys Acta 1860(7):1470–1479
Ahmad A, Yadav SP, Asthana N, Mitra K, Srivastava SP, Ghosh JK (2006) Utilization of an amphipathic leucine zipper sequence to design antibacterial peptides with simultaneous modulation of toxic activity against human red blood cells. J Biol Chem 281(31):22029–22038
Ahmad A, Asthana N, Azmi S, Srivastava RM, Pandey BK, Yadav V, Ghosh JK (2009a) Structure–function study of cathelicidin-derived bovine antimicrobial peptide BMAP-28: design of its cell-selective analogs by amino acid substitutions in the heptad repeat sequences. Biochim Biophys Acta 1788(11):2411–2420
Ahmad A, Azmi S, Srivastava RM, Srivastava S, Pandey BK, Saxena R, Bajpai VK, Ghosh JK (2009b) Design of nontoxic analogues of cathelicidin-derived bovine antimicrobial peptide BMAP-27: the role of leucine as well as phenylalanine zipper sequences in determining its toxicity. Biochemistry 48(46):10905–10917
Akhtar MS, Qaisar A, Irfanullah J, Iqbal J (2005) Antimicrobial peptide 99mTc-ubiquicidin 29-41 as human infection-imaging agent: clinical trial. J Nucl Med 46(4):567–573
Akhtar MS, Imran MB, Nadeem MA, Shahid A (2012) Antimicrobial peptides as infection imaging agents: better than radiolabeled antibiotics. Int J Pept. https://doi.org/10.1155/2012/965238
Akram AR, Avlonitis N, Lilienkampf A, Perez-Lopez AM, McDonald N, Chankeshwara SV, Scholefield E, Haslett C, Bradley M, Dhaliwal K (2015) A labelled-ubiquicidin antimicrobial peptide for immediate in situ optical detection of live bacteria in human alveolar lung tissue. Chem Sci 6(12):6971–6979
Alba A, López-Abarrategui C, Otero-González AJ (2012) Host defense peptides: an alternative as antiinfective and immunomodulatory therapeutics. Biopolymers 98(4):251–267
Allende D, Simon SA, McIntosh TJ (2005) Melittin-induced bilayer leakage depends on lipid material properties: evidence for toroidal pores. Biophys J 88(3):1828–1837
Aloia RC, Tian H, Jensen FC (1993) Lipid composition and fluidity of the human immunodeficiency virus envelope and host cell plasma membranes. Proc Natl Acad Sci U S A 90(11):5181–5185
Ames GF (1968) Lipids of Salmonella typhimurium and Escherichia coli: structure and metabolism. J Bacteriol 95(3):833–843
Andra J, Goldmann T, Ernst CM, Peschel A, Gutsmann T (2011) Multiple peptide resistance factor (MprF)-mediated resistance of Staphylococcus aureus against antimicrobial peptides coincides with a modulated peptide interaction with artificial membranes comprising lysyl-phosphatidylglycerol. J Biol Chem 286(21):18692–18700
Arcidiacono S, Soares JW, Meehan AM, Marek P, Kirby R (2009) Membrane permeability and antimicrobial kinetics of cecropin P1 against Escherichia coli. J Pept Sci 15(6):398–403
Assadi M, Vahdat K, Nabipour I, Sehhat MR, Hadavand F, Javadi H, Tavakoli A, Saberifard J, Kalantarhormozi MR, Zakani A, Eftekhari M (2011) Diagnostic value of 99mTc-ubiquicidin scintigraphy for osteomyelitis and comparisons with 99mTc-methylene diphosphonate scintigraphy and magnetic resonance imaging. Nucl Med Commun 32(8):716–723
Asthana N, Yadav SP, Ghosh JK (2004) Dissection of antibacterial and toxic activity of Melittin: a leucine zipper motif plays a crucial role in determining its hemolytic activity but not antibacterial activity. J Biol Chem 279(53):55042–55050
Avrahami D, Shai Y (2002) Conjugation of a magainin analogue with lipophilic acids controls hydrophobicity, solution assembly, and cell selectivity. Biochemistry 41(7):2254–2263
Avrahami D, Shai Y (2004) A new group of antifungal and antibacterial lipopeptides derived from non-membrane active peptides conjugated to palmitic acid. J Biol Chem 279(13):12277–12285
Bacalum M, Radu M (2015) Cationic antimicrobial peptides cytotoxicity on mammalian cells: an analysis using therapeutic index integrative concept. Int J Pept Res Ther 21:47–55
Bagheri M, Amininasab M, Dathe M (2018) Arg/Trp-rich cyclic α/β-antimicrobial peptides: the roles of H-bonding and hydrophobic/hydrophilic solvent accessible surface areas upon the activity and membrane selectivity. Chem Eur J. 24(53):14242–14253
Ballas SK, Krasnow SH (1980) Structure of erythrocyte membrane and its transport functions. Ann Clin Lab Sci 10(3):209–219
Beschiaschvili G, Seelig J (1990) Melittin binding to mixed phosphatidylglycerol/phosphatidylcholine membranes. Biochemistry 29(1):52–58
Bessalle R, Kapitkovsky A, Gorea A, Shalit I, Fridkin M (1990) All-D-magainin: chirality, antimicrobial activity and proteolytic resistance. FEBS Lett 274(1–2):151–155
Bessalle R, Haas H, Goria A, Shalit I, Fridkin M (1992) Augmentation of the antibacterial activity of magainin by positive-charge chain extension. Antimicrob Agents Chemother 36(2):313–317
Bhatt J, Mukherjee A, Shinto A, Karuppusamy KK, Korde A, Kumar M, Sarma HD, Repaka K, Dash A (2018) Gallium-68 labeled Ubiquicidin derived octapeptide as a potential infection imaging agent. Nucl Med Biol 62–63:47–53
Bishop DG, Rutberg L, Samuelsson B (1967) The chemical composition of the cytoplasmic membrane of Bacillus subtilis. Eur J Biochem 2(4):448–453
Bland JM, De Lucca AJ, Jacks TJ, Vigo CB (2001) All-D-cecropin B: synthesis, conformation, lipopolysaccharide binding, and antibacterial activity. Mol Cell Biochem 218(1–2):105–111
Blondelle SE, Houghten RA (1991) Hemolytic and antimicrobial activities of the twenty-four individual omission analogs of melittin. Biochemistry 30(19):4671–4678
Bobone S, Piazzon A, Orioni B, Pedersen JZ, Nan YH, Hahm KS, Shin SH, Stella L (2011) The thin line between cell-penetrating and antimicrobial peptides: the case of Pep-1 and Pep-1-K. J Pept Sci 17(5):335–341
Bobone S, Roversi D, Giordano L, De Zotti M, Formaggio F, Toniolo C, Park Y, Stella L (2012) The lipid dependence of antimicrobial peptide activity is an unreliable experimental test for different pore models. Biochemistry 51(51):10124–10126
Bobone S, Bocchinfuso G, Park Y, Palleschi A, Hahm KS, Stella L (2013) The importance of being kinked: role of Pro residues in the selectivity of the helical antimicrobial peptide P5. J Pept Sci 19(12):758–769
Bocchinfuso G, Palleschi A, Orioni B, Grande G, Formaggio F, Toniolo C et al (2009) Different mechanisms of action of antimicrobial peptides: insights from fluorescence spectroscopy experiments and molecular dynamics simulations. J Pept Sci Off Publ Eur Pept Soc 15(9):550–558
Bocchinfuso G, Bobone S, Mazzuca C, Palleschi A, Stella L (2011) Fluorescence spectroscopy and molecular dynamics simulations in studies on the mechanism of membrane destabilization by antimicrobial peptides. Cell Mol Life Sci 68(13):2281–2301
Boyd KJ, Alder NN, May ER (2017) Buckling under pressure: curvature-based lipid segregation and stability modulation in cardiolipin-containing bilayers. Langmuir 33(27):6937–6946
Braun S, Pokorná S, Šachl R, Hof M, Heerklotz H, Hoernke M (2017) Biomembrane permeabilization: statistics of individual leakage events harmonize the interpretation of vesicle leakage. ACS Nano 12(1):813–819
Brender JR, McHenry AJ, Ramamoorthy A (2012) Does cholesterol play a role in the bacterial selectivity of antimicrobial peptides? Front Immunol 3:195
Broekhuyse RM (1969) Quantitative two-dimensional thin-layer chromatography of blood phospholipids. Clin Chim Acta 23(3):457–461
Brouwer CP, Bogaards SJ, Wulferink M, Velders MP, Welling MM (2006) Synthetic peptides derived from human antimicrobial peptide ubiquicidin accumulate at sites of infections and eradicate (multi-drug resistant) Staphylococcus aureus in mice. Peptides 27(11):2585–2591
Brouwer CPJM, Sarda-Mantel L, Meulemans A, Guludec DL, Welling MM (2008) The use of technetium-99m radiolabeled human antimicrobial peptides for infection specific imaging. Mini-Rev Med Chem 8(10):1039–1052
Bulet P, Stocklin R (2005) Insect antimicrobial peptides: structures, properties and gene regulation. Protein Pept Lett 12(1):3–11
Bütikofer P, Lin ZW, Chiu DT, Lubin B, Kuypers FA (1990) Transbilayer distribution and mobility of phosphatidylinositol in human red blood cells. J Biol Chem 265(27):16035–16038
Büttner K, Blondelle SE, Ostresh JM, Houghten RA (1992) Perturbation of peptide conformations induced in anisotropic environments. Biopolymers 32(6):575–583
Carneiro VA, Duarte HS, Prado MGV, Silva ML, Teixeira M, dos Santos YM, Vasconcelos IB, Cunha RMS (2015) Antimicrobial peptides: from synthesis to clinical perspectives. In: The battle against microbial pathogens: basic science, technological advances and educational programs, 1st edn. Formatex Research Center, Spain, pp 81–90
Carotenuto A, Malfi S, Saviello MR, Campiglia P, Gomez-Monterrey I, Mangoni ML, Marcellini Hercolani Gaddi L, Novellino E, Grieco P (2008) A different molecular mechanism underlying antimicrobial and hemolytic actions of temporins A and L. J Med Chem 51(8):2354–2362
Chairatana P, Nolan EM (2014) Molecular basis for self-assembly of a human host-defense peptide that entraps bacterial pathogens. J Am Chem Soc 136(38):13267–13276
Chapuis H, Slaninová J, Bednárová L, Monincová L, Buděšínský M, Čeřovský V (2012) Effect of hydrocarbon stapling on the properties of α-helical antimicrobial peptides isolated from the venom of hymenoptera. Amino Acids 43(5):2047–2058
Chen L, Liang JF (2013) Peptide fibrils with altered stability, activity, and cell selectivity. Biomacromolecules 14(7):2326–2331
Chen Y, Mant CT, Farmer SW, Hancock REW, Michael L, Vasil ML, Hodges RS (2005) Rational design of alpha-helical antimicrobial peptides with enhanced activities and specificity/therapeutic index. J Biol Chem 280:12316–12329
Chen Y, Vasil AI, Rehaume L, Mant CT, Burns JL, Vasil ML, Hancock RE, Hodges RS (2006) Comparison of biophysical and biologic properties of α-helical enantiomeric antimicrobial peptides. Chem Biol Drug Des 67(2):162–173
Chen Y, Guarnieri MT, Vasil AI, Vasil ML, Mant CT, Hodges RS (2007) Role of peptide hydrophobicity in the mechanism of action of α-helical antimicrobial peptides. Antimicrob Agents Chemother 51(4):1398–1406
Chen C, Pan F, Zhang S, Hu J, Cao M, Wang J, Xu H, Zhao X, Lu JR (2010) Antibacterial activities of short designer peptides: a link between propensity for nanostructuring and capacity for membrane destabilization. Biomacromolecules 11(2):402–411
Chen C, Hu J, Zeng P, Pan F, Yaseen M, Xu H, Lu JR (2014) Molecular mechanisms of anticancer action and cell selectivity of short α-helical peptides. Biomaterials 35(5):1552–1561
Chen H, Liu C, Chen D, Madrid K, Peng S, Dong X, Zhang M, Gu Y (2015) Bacteria-targeting conjugates based on antimicrobial peptide for bacteria diagnosis and therapy. Mol Pharm 12(7):2505–2516
Cherry MA, Higgins SK, Melroy H, Lee HS, Pokorny A (2014) Peptides with the same composition, hydrophobicity, and hydrophobic moment bind to phospholipid bilayers with different affinities. J Phys Chem B 118(43):12462–12470
Chu-Kung AF, Bozzelli KN, Lockwood NA, Haseman JR, Mayo KH, Tirrell MV (2004) Promotion of peptide antimicrobial activity by fatty acid conjugation. Bioconjug Chem 15(3):530–535
Chu-Kung AF, Nguyen R, Bozzelli KN, Tirrell M (2010) Chain length dependence of antimicrobial peptide-fatty acid conjugate activity. J Colloid Interface Sci 345(2):160–167
Cornut I, Büttner K, Dasseux JL, Dufourcq J (1994) The amphipathic α-helix concept: application to the de novo design of ideally amphipathic Leu, Lys peptides with hemolytic activity higher than that of melittin. FEBS Lett 349(1):29–33
Dalzini A, Bergamini C, Biondi B, De Zotti M, Panighel G, Fato R, Peggion C, Bortolus M, Maniero AL (2016) The rational search for selective anticancer derivatives of the peptide trichogin GA IV: a multi-technique biophysical approach. Sci Rep 6:24000
Daschbach MM, Negin S, You L, Walsh M, Gokel GW (2012) Aggregation and supramolecular membrane interactions that influence anion transport in tryptophan-containing synthetic peptides. Chem Eur J 18(24):7608–7623
Dathe M, Wieprecht T (1999) Structural features of helical antimicrobial peptides: their potential to modulate activity on model membranes and biological cells. Biochim Biophys Acta 1462(1):71–87
Dathe M, Schümann M, Wieprecht T, Winkler A, Beyermann M, Krause E, Matsuzatki K, Murase O, Bienert M (1996) Peptide helicity and membrane surface charge modulate the balance of electrostatic and hydrophobic interactions with lipid bilayers and biological membranes. Biochemistry 35(38):12612–12622
Dathe M, Wieprecht T, Nikolenko H, Handel L, Maloy WL, MacDonald DL, Beyermann M, Bienert M (1997) Hydrophobicity, hydrophobic moment and angle subtended by charged residues modulate antibacterial and haemolytic activity of amphipathic helical peptides. FEBS Lett 403(2):208–212
Dathe M, Nikolenko H, Meyer J, Beyermann M, Bienert M (2001) Optimization of the antimicrobial activity of magainin peptides by modification of charge. FEBS Lett 501(2–3):146–150
Dathe M, Meyer J, Beyermann M, Maul B, Hoischen C, Bienert M (2002) General aspects of peptide selectivity towards lipid bilayers and cell membranes studied by variation of the structural parameters of amphipathic helical model peptides. Biochim Biophys Acta 1558(2):171–186
Dawson RM, Liu CQ (2011) Analogues of peptide SMAP-29 with comparable antimicrobial potency and reduced toxicity. Int J Antimicrob Agents 37(5):432–437
Dawson RM, Fox MA, Atkins HS, Liu CQ (2011) Potent antimicrobial peptides with selectivity for Bacillus anthracis over human erythrocytes. Int J Antimicrob Agents 38(3):237–242
de Kruijff B (1990) Cholesterol as a target for toxins. Biosci Rep 10(2):127–130
de la Fuente-Núñez C, Reffuveille F, Mansour SC, Reckseidler-Zenteno SL, Hernández D, Brackman G, Coenye T, Hancock REW (2015) D-enantiomeric peptides that eradicate wild-type and multidrug-resistant biofilms and protect against lethal Pseudomonas aeruginosa infections. Chem Biol 22(2):196–205
de Murphy CA, Gemmel F, Balter J (2010) Clinical trial of specific imaging of infections. Nucl Med Commun 31(8):726–733
Dean SN, Bishop BM, Van Hoek ML (2011) Susceptibility of Pseudomonas aeruginosa biofilm to alpha-helical peptides: D-enantiomer of LL-37. Front Microbiol 2:128
DeGrado WF, Kezdy FJ, Kaiser ET (1981) Design, synthesis, and characterization of a cytotoxic peptide with melittin-like activity. J Am Chem Soc 103(3):679–681
Dempsey CE, Ueno S, Avison MB (2003) Enhanced membrane permeabilization and antibacterial activity of a disulfide-dimerized magainin analogue. Biochemistry 42(2):402–409
Dennison SR, Harris F, Bhatt T, Singh J, Phoenix DA (2009) The effect of C-terminal amidation on the efficacy and selectivity of antimicrobial and anticancer peptides. Mol Cell Biochem 332(1–2):43–50
Deslouches B, Hasek ML, Craigo JK, Steckbeck JD, Montelar RC (2016) Comparative functional properties of engineered cationic antimicrobial peptides consisting exclusively of tryptophan and either lysine or arginine. J Med Microbiol 65(6):554–565
Dodge JT, Phillips GB (1967) Composition of phospholipids and of phospholipid fatty acids and aldehydes in human red cells. J Lipid Res 8(6):667–675
Dutta J, Baijnath S, Somboro AM, Nagiah S, Albericio F, de la Torre BG, Marjanovic-Painter B, Zeevaart JR, Sathekge M, Kruger HG, Chuturgoon A, Naicker T, Ebenhan T, Govender T (2017) Synthesis, in vitro evaluation, and 68Ga-radiolabeling of CDP1 toward PET/CT imaging of bacterial infection. Chem Biol Drug Des 90(4):572–579
Ebenhan T, Gheysens O, Kruger HG, Zeevaart JR, Sathekge MM (2014a) Antimicrobial peptides: their role as infection-selective tracers for molecular imaging. Biomed Res Int. https://doi.org/10.1155/2014/867381
Ebenhan T, Chadwick N, Sathekge MM, Govender P, Govender T, Kruger HG, Marjanovic-Painter B, Zeevaart JR (2014b) Peptide synthesis, characterization and 68Ga-radiolabeling of NOTA-conjugated ubiquicidin fragments for prospective infection imaging with PET/CT. Nucl Med Biol 41(5):390–400
Ebenhan T, Sathekge M, Lenagana T, Koole M, Gheysens O, Govender T, Zeevaart JR (2018) 68Ga-NOTA-functionalized ubiquicidin: cytotoxicity, biodistribution, radiation dosimetry and first-in-human positron emission tomography/computed tomography imaging of infections. J Nucl Med 59(2):334–339
Eckert R (2011) Road to clinical efficacy: challenges and novel strategies for antimicrobial peptide development. Future Microbiol 6(6):635–651
Eisenberg D, Weiss RM, Terwilliger TC (1982) The helical hydrophobic moment: a measure of the amphiphilicity of a helix. Nature 299(5881):371–374
Epand RF, Schmitt MA, Gellman SH, Epand RM (2006) Role of membrane lipids in the mechanism of bacterial species selective toxicity by two α/β-antimicrobial peptides. Biochim Biophys Acta 1758(9):1343–1350
Farrotti A, Bocchinfuso G, Palleschi A, Rosato N, Salnikov ES, Voievoda N, Bechinger B, Stella L (2015) Molecular dynamics methods to predict peptide locations in membranes: LAH4 as a stringent test case. Biochim Biophys Acta 1848(2):581–592
Farrotti A, Conflitti P, Srivastava S, Ghosh JK, Palleschi A, Stella L, Bocchinfuso G (2017) Molecular dynamics simulations of the host defense peptide temporin L and its Q3K derivative: an atomic level view from aggregation in water to bilayer perturbation. Molecules 22(7):1235
Feder R, Dagan A, Mor A (2000) Structure-activity relationship study of antimicrobial dermaseptin S4 showing the consequences of peptide oligomerization on selective cytotoxicity. J Biol Chem 275(6):4230–4238
Feder R, Nehushtai R, Mor A (2001) Affinity driven molecular transfer from erythrocyte membrane to target cells. Peptides 22(10):1683–1690
Fernández-Vidal M, Jayasinghe S, Ladokhin AS, White SH (2007) Folding amphipathic helices into membranes: amphiphilicity trumps hydrophobicity. J Mol Biol 370(3):459–470
Ferro-Flores G, de Murphy CA, Pedraza-López M, Meléndez-Alafort L, Zhang YM, Rusckowski M, Hnatowich DJ (2003) In vitro and in vivo assessment of 99mTc-UBI specificity for bacteria. Nucl Med Biol 30(6):597–603
Findlay EG, Currie SM, Davidson DJ (2013) Cationic host defence peptides: potential as antiviral therapeutics. BioDrugs 27(5):479–493
Friedrich CL, Moyles D, Beveridge TJ, Hancock RE (2000) Antibacterial action of structurally diverse cationic peptides on gram-positive bacteria. Antimicrob Agents Chemother 44(8):2086–2092
Gaillard P, Carrupt PA, Testa B, Boudon A (1994) Molecular lipophilicity potential, a tool in 3D QSAR: method and applications. J Comput Aided Mol Des 8(2):83–96
Gandomkar M, Najafi R, Shafiei M, Mazidi M, Goudarzi M, Mirfallah SH, Ebrahimi F, Heydarpor HR, Abdie N (2009) Clinical evaluation of antimicrobial peptide [99mTc/Tricine/HYNIC0] ubiquicidin 29–41 as a human-specific infection imaging agent. Nucl Med Biol 36(2):199–205
Gascard P, Tran D, Sauvage M, Sulpice JC, Fukami K, Takenawa T, Claret M, Giraud F (1991) Asymmetric distribution of phosphoinositides and phosphatidic acid in the human erythrocyte membrane. Biochim Biophys Acta 1069(1):27–36
Gaspar D, Veiga AS, Castanho MA (2013) From antimicrobial to anticancer peptides. A review. Front Microbiol 4:294
Gatto E, Mazzuca C, Stella L, Venanzi M, Toniolo C, Pispisa B (2006) Effect of peptide lipidation on membrane perturbing activity: a comparative study on two trichogin analogues. J Phys Chem B 110(45):22813–22818
Gazit E, Boman A, Boman HG, Shai Y (1995) Interaction of the mammalian antibacterial peptide cecropin P1 with phospholipid vesicles. Biochemistry 34(36):11479–11488
Gelhaus C, Jacobs T, Andrä J, Leippe M (2008) The antimicrobial peptide NK-2, the core region of mammalian NK-lysin, kills intraerythrocytic Plasmodium falciparum. Antimicrob Agents Chemother 52(5):1713–1720
Ghosh JK, Shaool D, Guillaud P, Cicéron L, Mazier D, Kustanovich I, Shai Y, Mor A (1997) Selective cytotoxicity of dermaseptin S3 toward intraerythrocytic plasmodium falciparum and the underlying molecular basis. J Biol Chem 272(50):31609–31616
Giacometti A, Cirioni O, Greganti G, Quarta M, Scalise G (1998) In vitro activities of membrane-active peptides against gram-positive and gram-negative aerobic bacteria. Antimicrob Agents Chemother 42(12):3320–3324
Giangaspero A, Sandri L, Tossi A (2001) Amphipathic α helical antimicrobial peptides. A systematic study of the effects of structural and physical properties on biological activity. Eur J Biochem 268(21):5589–5600
Glukhov E, Burrows LL, Deber CM (2008) Membrane interactions of designed cationic antimicrobial peptides: the two thresholds. Biopolymers 89(5):360–371
Golbek TW, Franz J, Elliott Fowler J, Schilke KF, Weidner T, Baio JE (2017) Identifying the selectivity of antimicrobial peptides to cell membranes by sum frequency generation spectroscopy. Biointerphases 12(2):02D406
Gonçalves S, Abade J, Teixeira A, Santos NC (2012) Lipid composition is a determinant for human defensin HNP1 selectivity. Biopolymers 98(4):313–321
Gordesky SE, Marinetti GV (1973) The asymmetric arrangement of phospholipids in the human erythrocyte membrane. Biochem Biophys Res Commun 50(4):1027–1031
Gordesky SE, Marinetti GV, Love R (1975) The reaction of chemical probes with the erythrocyte membrane. J Membr Biol 20(1–2):111–132
Hallock KJ, Lee DK, Omnaas J, Mosberg HI, Ramamoorthy A (2002) Membrane composition determines pardaxin’s mechanism of lipid bilayer disruption. Biophys J 83(2):1004–1013
Hancock RE, Sahl HG (2006) Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies. Nat Biotechnol 24(12):1551
Haney EF, Wu BC, Lee K, Hilchie AL, Hancock RE (2017) Aggregation and its influence on the immunomodulatory activity of synthetic innate defense regulator peptides. Cell Chem Biol 24(8):969–980
Harris F, Dennison SR, Singh J, Phoenix DA (2013) On the selectivity and efficacy of defense peptides with respect to cancer cells. Med Res Rev 33(1):190–234
Hartmann M, Berditsch M, Hawecker J, Ardakani MF, Gerthsen D, Ulrich AS (2010) Damage of the bacterial cell envelope by antimicrobial peptides gramicidin S and PGLa as revealed by transmission and scanning electron microscopy. Antimicrob Agents Chemother 54(8):3132–3142
Hawrani A, Howe RA, Walsh TR, Dempsey CE (2008) Origin of low mammalian cell toxicity in a class of highly active antimicrobial amphipathic helical peptides. J Biol Chem 283(27):18636–18645
Hayami M, Okabe A, Kariyama R, Abe M, Kanemasa Y (1979) Lipid composition of Staphylococcus aureus and its derived L-forms. Microbiol Immunol 23(6):435–442
He J, Krauson AJ, Wimley WC (2014) Toward the de novo design of antimicrobial peptides: lack of correlation between peptide permeabilization of lipid vesicles and antimicrobial, cytolytic, or cytotoxic activity in living cells. Biopolymers 102(1):1–6
Henriksen J, Rowat AC, Brief E, Hsueh YW, Thewalt JL, Zuckermann MJ, Ipsen JH (2006) Universal behavior of membranes with sterols. Biophys J 90(5):1639–1649
Henriksen JR, Etzerodt T, Gjetting T, Andresen TL (2014) Side chain hydrophobicity modulates therapeutic activity and membrane selectivity of antimicrobial peptide mastoparan-X. PLoS One 9:e91007
Hollmann A, Martinez M, Maturana P, Semorile LC, Maffia PC (2018) Antimicrobial peptides: interaction with model and biological membranes and synergism with chemical antibiotics. Front Chem 6:204
Holt A, de Almeida RF, Nyholm TK, Loura LM, Daily AE, Staffhorst RW, Rijkers DTS, Koeppe RE II, Prieto M, Killian JA (2008) Is there a preferential interaction between cholesterol and tryptophan residues in membrane proteins? Biochemistry 47(8):2638–2649
Hong SY, Oh JE, Lee KH (1999) Effect of D-amino acid substitution on the stability, the secondary structure, and the activity of membrane-active peptide. Biochem Pharmacol 58(11):1775–1780
Hoskin DW, Ramamoorthy A (2008) Studies on anticancer activities of antimicrobial peptides. Biochim Biophys Acta 1778(2):357–375
Hoyos-Nogués M, Gil FJ, Mas-Moruno C (2018) Antimicrobial peptides: powerful biorecognition elements to detect bacteria in biosensing technologies. Molecules 23(7):1683
Huang Y, Huang J, Chen Y (2010) Alpha-helical cationic antimicrobial peptides: relationships of structure and function. Protein Cell 1(2):143–152
Huang Y, He L, Li G, Zhai N, Jiang H, Chen Y (2014) Role of helicity of α-helical antimicrobial peptides to improve specificity. Protein Cell 5(8):631–642
Jacobs T, Bruhn H, Gaworski I, Fleischer B, Leippe M (2003) NK-lysin and its shortened analog NK-2 exhibit potent activities against Trypanosoma cruzi. Antimicrob Agents Chemother 47(2):607–613
Javadpour MM, Juban MM, Lo WCJ, Bishop SM, Alberty BJ, Cowell SM, Becker CL, McLaughlin ML (1996) De novo antimicrobial peptides with low mammalian cell toxicity. J Med Chem 39:3107–3113
Jepson AK, Schwarz-Linek J, Ryan L, Ryadnov MG, Poon WC (2016) What is the ‘Minimum Inhibitory Concentration’ (MIC) of Pexiganan acting on Escherichia coli?-A cautionary case study. In: Leake MC (ed) Biophysics of infection. Springer, Cham, pp 33–48
Jiang Z, Vasil AI, Hale JD, Hancock RE, Vasil ML, Hodges RS (2008) Effects of net charge and the number of positively charged residues on the biological activity of amphipathic α-helical cationic antimicrobial peptides. Biopolymers 90(3):369–383
Jiang Z, Vasil AI, Gera L, Vasil M, Hodges RS (2011) Rational design of a-helical antimicrobial peptides to target Gram-negative pathogens, Acinetobacter baumannii and Pseudomonas aeruginosa: utilization of charge, ‘specificity determinants,’ total hydrophobicity, hydrophobe type and location as design parameters to improve the therapeutic ratio. Chem Biol Drug Des 77(4):225–240
Jiang Z, Vasil AI, Vasil ML, Hodges RS (2014) “Specificity Determinants” improve therapeutic indices of two antimicrobial peptides piscidin 1 and dermaseptin s4 against the Gram-negative pathogens Acinetobacter baumannii and Pseudomonas aeruginosa. Pharmaceuticals 7(4):366–391
Jiang Z, Mant CT, Vasil M, Hodges RS (2018) Role of positively charged residues on the polar and non- polar faces of amphipathic α- helical antimicrobial peptides on specificity and selectivity for Gram-negative pathogens. Chem Biol Drug Des 91(1):75–92
Jones EM, Smart A, Bloomberg G, Burgess L, Millar MR (1994) Lactoferricin, a new antimicrobial peptide. J Appl Bacteriol 77(2):208–214
Joshi S, Dewangan RP, Yar MS, Rawat DS, Pasha S (2015) N-terminal aromatic tag induced self assembly of tryptophan–arginine rich ultra short sequences and their potent antibacterial activity. RSC Adv 5(84):68610–68620
Juretic D, Vukicevic D, Ilic N, Antcheva N, Tossi A (2009) Computational design of highly selective antimicrobial peptides. J Chem Inf Model 49(12):2873–2882
Juvvadi P, Vunnam S, Merrifield RB (1996) Synthetic melittin, its enantio, retro, and retroenantio isomers, and selected chimeric analogs: their antibacterial, hemolytic, and lipid bilayer action. J Am Chem Soc 118(38):8989–8997
Kahrom M, Bahar MM, Jangjoo A, Erfani M, Sadeghi R, Zakavi SR (2014) Poor sensitivity of 99mTc-labeled ubiquicidin scintigraphy in diagnosis of acute appendicitis. Eur Surg 46(4):173–176
Kamech N, Vukičević D, Ladram A, Piesse C, Vasseur J, Bojović V, Simunić J, Juretić D (2012) Improving the selectivity of antimicrobial peptides from anuran skin. J Chem Inf Model 52(12):3341–3351
Kaminski HM, Feix JB (2011) Effects of D-lysine substitutions on the activity and selectivity of antimicrobial peptide CM15. Polymers 3(4):2088–2106
Kang JH, Shin SY, Jang SY, Kim KL, Hahm KS (1999) Effects of tryptophan residues of porcine myeloid antibacterial peptide PMAP-23 on antibiotic activity. Biochem Biophys Res Commun 264(1):281–286
Khandelia H, Kaznessis YN (2006) Molecular dynamics investigation of the influence of anionic and zwitterionic interfaces on antimicrobial peptides’ structure: implications for peptide toxicity and activity. Peptides 27(6):1192–1200
Kim S, Kim SS, Lee BJ (2005) Correlation between the activities of α-helical antimicrobial peptides and hydrophobicities represented as RP HPLC retention times. Peptides 26(11):2050–2056
Kim JK, Lee SA, Shin S, Lee JY, Jeong KW, Nan YH, Park YS, Shin SY, Kim Y (2010) Structural flexibility and the positive charges are the key factors in bacterial cell selectivity and membrane penetration of peptoid-substituted analog of Piscidin 1. Biochim Biophys Acta 1798(10):1913–1925
Kindrachuk J, Napper S (2010) Structure-activity relationships of multifunctional host defence peptides. Mini-Rev Med Chem 10(7):596–614
Koller D, Lohner K (2014) The role of spontaneous lipid curvature in the interaction of interfacially active peptides with membranes. Biochim Biophys Acta 1838(9):2250–2259
Konai MM, Samaddar S, Bocchinfuso G, Santucci V, Stella L, Haldar J (2018) Selectively targeting bacteria by tuning the molecular design of membrane-active peptidomimetic amphiphiles. Chem Commun 54(39):4943–4946
Kondejewski LH, Jelokhani-Niaraki M, Farmer SW, Lix B, Kay CM, Sykes BD, Hancock RE, Hodges RS (1999) Dissociation of antimicrobial and hemolytic activities in cyclic peptide diastereomers by systematic alterations in amphipathicity. J Biol Chem 274(19):13181–13192
Kondejewski LH, Lee DL, Jelokhani-Niaraki M, Farmer SW, Hancock REW, Hodges RS (2002) Optimization of microbial specificity in cyclic peptides by modulation of hydrophobicity within a defined structural framework. J Biol Chem 277(1):67–74
Krause E, Beyermann M, Dathe M, Rothemund S, Bienert M (1995) Location of an amphipathic. alpha-Helix in peptides using reversed-phase HPLC retention behavior of D-amino acid analogs. Anal Chem 67(2):252–258
Krugliak M, Feder R, Zolotarev VY, Gaidukov L, Dagan A, Ginsburg H, Mor A (2000) Antimalarial activities of dermaseptin S4 derivatives. Antimicrob Agents Chemother 44(9):2442–2451
Kustanovich I, Shalev DE, Mikhlin M, Gaidukov L, Mor A (2002) Structural requirements for potent versus selective cytotoxicity for antimicrobial dermaseptin S4 derivatives. J Biol Chem 277(19):16941–16951
Ladokhin AS, White SH (2001) Protein chemistry at membrane interfaces: non-additivity of electrostatic and hydrophobic interactions. J Mol Biol 309(3):543–552
Laverty G, McLaughlin M, Shaw C, Gorman SP, Gilmore BF (2010) Antimicrobial activity of short, synthetic cationic lipopeptides. Chem Biol Drug Des 75(6):563–569
Le Joncour V, Laakkonen P (2018) Seek & destroy, use of targeting peptides for cancer detection and drug delivery. Bioorg Med Chem 26(10):2797–2806
Lee K, Shin SY, Kim K, Lim SS, Hahm KS, Kim Y (2004) Antibiotic activity and structural analysis of the scorpion-derived antimicrobial peptide IsCT and its analogs. Biochem Biophys Res Commun 323(2):712–719
Lee MT, Hung WC, Chen FY, Huang HW (2005) Many-body effect of antimicrobial peptides: on the correlation between lipid’s spontaneous curvature and pore formation. Biophys J 89(6):4006–4016
Lee SA, Kim YK, Lim SS, Zhu WL, Ko H, Shin SY, Hahm KS, Kim Y (2007) Solution structure and cell selectivity of piscidin 1 and its analogues. Biochemistry 46(12):3653–3663
Lei R, Hou J, Chen Q, Yuan W, Cheng B, Sun Y, Jin Y, Ge L, Ben-Sasson SA, Chen J, Wang H, Lu W, Fang X (2018) Self-assembling myristoylated human α-defensin 5 as a next-generation nanobiotics potentiates therapeutic efficacy in bacterial infection. ACS Nano 12(6):5284–5296
Leite NB, Aufderhorst-Roberts A, Palma MS, Connell SD, Neto JR, Beales PA (2015) PE and PS lipids synergistically enhance membrane poration by a peptide with anticancer properties. Biophys J 109(5):936–947
Levison ME, Pitsakis PG, May PL, Johnson CC (1993) The bactericidal activity of magainins against Pseudomonas aeruginosa and Enterococcus faecium. J Antimicrob Chemother 32(4):577–585
Lewis RNAH, McElhaney RN (2005) The mesomorphic phase behavior of lipids. In: Yeagle PL (ed) The structure of biological membranes, 2nd edn. CRC Press, Boca Raton, pp 53–120
Li P, Zhou C, Rayatpisheh S, Ye K, Poon YF, Hammond PT, Duan H, Chan-Park MB (2012) Cationic peptidopolysaccharides show excellent broad-spectrum antimicrobial activities and high selectivity. Adv Mater 24(30):4130–4137
Lin D, Grossfield A (2015) Thermodynamics of micelle formation and membrane fusion modulate antimicrobial lipopeptide activity. Biophys J 109(4):750–759
Liu L, Xu K, Wang H, Tan PJ, Fan W, Venkatraman SS, Li L, Yang YY (2009) Self-assembled cationic peptide nanoparticles as an efficient antimicrobial agent. Nat Nanotechnol 4(7):457–463
Lorian V (2005) Antibiotics in laboratory medicine, 5th edn. Lippincott Williams & Wilkins, Philadelphia
Luo Y, McLean DT, Linden GJ, McAuley DF, McMullan R, Lundy FT (2017) The naturally occurring host defense peptide, LL-37, and its truncated mimetics KE-18 and KR-12 have selected biocidal and antibiofilm activities against Candida albicans, Staphylococcus aureus, and Escherichia coli in vitro. Front Microbiol 8:544
Lupetti A, Welling MM, Pauwels EK, Nibbering PH (2003) Radiolabelled antimicrobial peptides for infection detection. Lancet Infect Dis 3(4):223–229
Lupetti A, Van Dissel JT, Brouwer CP, Nibbering PH (2008) Human antimicrobial peptides’ antifungal activity against Aspergillus fumigatus. Eur J Clin Microbiol Infect Dis 27(11):1125–1129
Lyu Y, Yang Y, Lyu X, Dong N, Shan A (2016) Antimicrobial activity, improved cell selectivity and mode of action of short PMAP-36-derived peptides against bacteria and Candida. Sci Rep 6:27258
Mahlapuu M, Håkansson J, Ringstad L, Björn C (2016) Antimicrobial peptides: an emerging category of therapeutic agents. Front Cell Infect Microbiol 6:194
Makovitzki A, Avrahami D, Shai Y (2006) Ultrashort antibacterial and antifungal lipopeptides. Proc Natl Acad Sci U S A 103(43):15997–16002
Makovitzki A, Baram J, Shai Y (2008) Antimicrobial lipopolypeptides composed of palmitoyl di-and tricationic peptides: in vitro and in vivo activities, self-assembly to nanostructures, and a plausible mode of action. Biochemistry 47(40):10630–10636
Malanovic N, Lohner K (2016) Antimicrobial peptides targeting gram-positive bacteria. Pharmaceuticals 9(3):59
Malina A, Shai Y (2005) Conjugation of fatty acids with different lengths modulates the antibacterial and antifungal activity of a cationic biologically inactive peptide. Biochem J 390(3):695–702
Mangoni ML, Shai Y (2009) Temporins and their synergism against Gram-negative bacteria and in lipopolysaccharide detoxification. Biochim Biophys Acta 1788(8):1610–1619
Mangoni ML, Carotenuto A, Auriemma L, Saviello MR, Campiglia P, Gomez-Monterrey I, Malfi S, Marcellini L, Barra D, Novellino E, Grieco P (2011) Structure-activity relationship, conformational and biological studies of Temporin L analogues. J Med Chem 54(5):1298–1307
Mannoor MS, Zhang S, Link AJ, McAlpine MC (2010) Electrical detection of pathogenic bacteria via immobilized antimicrobial peptides. Proc Natl Acad Sci U S A 107(45):19207–19212
Marsh D (1990) CRC handbook of lipid bilayers. CRC Press, Boca Raton
Matsuzaki K (2009) Control of cell selectivity of antimicrobial peptides. Biochim Biophys Acta 1788(8):1687–1692
Matsuzaki K, Harada M, Handa T, Funakoshi S, Fujii N, Yajima H, Miyajima K (1989) Magainin 1-induced leakage of entrapped calcein out of negatively-charged lipid vesicles. Biochim Biophys Acta 981(1):130–134
Matsuzaki K, Sugishita K, Fujii N, Miyajima K (1995) Molecular basis for membrane selectivity of an antimicrobial peptide, magainin 2. Biochemistry 34(10):3423–3429
Matsuzaki K, Sugishita KI, Harada M, Fujii N, Miyajima K (1997) Interactions of an antimicrobial peptide, magainin 2, with outer and inner membranes of Gram-negative bacteria. Biochim Biophys Acta 1327(1):119–130
Matsuzaki K, Sugishita KI, Ishibe N, Ueha M, Nakata S, Miyajima K, Epand RM (1998) Relationship of membrane curvature to the formation of pores by magainin 2. Biochemistry 37(34):11856–11863
Maturana P, Martinez M, Noguera ME, Santos NC, Disalvo EA, Semorile L, Maffia PC, Hollmann A (2017) Lipid selectivity in novel antimicrobial peptides: implication on antimicrobial and hemolytic activity. Colloids Surf B Biointerfaces 153:152–159
Mazzuca C, Stella L, Venanzi M, Formaggio F, Toniolo C, Pispisa B (2005) Mechanism of membrane activity of the antibiotic trichogin GA IV: a two-state transition controlled by peptide concentration. Biophys J 88(5):3411–3421
McCloskey AP, Gilmore BF, Laverty G (2014) Evolution of antimicrobial peptides to self-assembled peptides for biomaterial applications. Pathogens 3(4):791–821
McHenry AJ, Sciacca MF, Brender JR, Ramamoorthy A (2012) Does cholesterol suppress the antimicrobial peptide induced disruption of lipid raft containing membranes? Biochim Biophys Acta 1818(12):3019–3024
McIntosh TJ, Vidal A, Simon SA (2002) The energetics of peptide-lipid interactions: modulation by interfacial dipoles and cholesterol. In: Peptide-lipid interactions, current topics in membranes, vol 52. Elsevier, Amsterdam, pp 309–338
McMahon HT, Boucrot E (2015) Membrane curvature at a glance. J Cell Sci 128(6):1065–1070
Meléndez-Alafort L, Rodríguez-Cortés J, Ferro-Flores G, De Murphy CA, Herrera-Rodríguez R, Mitsoura E, Martínez-Duncker C (2004) Biokinetics of 99mTc-UBI 29-41 in humans. Nucl Med Biol 31(3):373–379
Melo MN, Ferre R, Castanho MA (2009) Antimicrobial peptides: linking partition, activity and high membrane-bound concentrations. Nat Rev Microbiol 7(3):245–250
Mendive-Tapia L, Zhao C, Akram AR, Preciado S, Albericio F, Lee M, Serrels A, Kielland N, Read ND, Lavilla R, Vendrell M (2016) Spacer-free BODIPY fluorogens in antimicrobial peptides for direct imaging of fungal infection in human tissue. Nat Commun 7:10940
Miyazaki Y, Aoki M, Yano Y, Matsuzaki K (2012) Interaction of antimicrobial peptide magainin 2 with gangliosides as a target for human cell binding. Biochemistry 51(51):10229–10235
Mojsoska B, Zuckermann RN, Jenssen H (2015) Structure-activity relationship study of novel peptoids that mimic the structure of antimicrobial peptides. Antimicrob Agents Chemother 59(7):4112–4120
Morein S, Andersson AS, Rilfors L, Lindblom G (1996) Wild-type Escherichia coli cells regulate the membrane lipid composition in a window between gel and non-lamellar structures. J Biol Chem 271(12):6801–6809
Mouritsen OG, Zuckermann MJ (2004) What’s so special about cholesterol? Lipids 39(11):1101–1113
Mura M, Wang J, Zhou Y, Pinna M, Zvelindovsky AV, Dennison SR, Phoenix DA (2016) The effect of amidation on the behaviour of antimicrobial peptides. Eur Biophys J 45(3):195–207
Nan YH, Bang JK, Shin SY (2009) Design of novel indolicidin-derived antimicrobial peptides with enhanced cell specificity and potent anti-inflammatory activity. Peptides 30(5):832–838
Nan YH, Lee BJ, Shin SY (2012) Prokaryotic selectivity, anti-endotoxic activity and protease stability of diastereomeric and enantiomeric analogs of human antimicrobial peptide LL-37. Bull Kor Chem Soc 33(9):2883–2889
Nicolas P (2009) Multifunctional host defense peptides: intracellular-targeting antimicrobial peptides. FEBS J 276(22):6483–6496
Nishi H, Komatsuzawa H, Fujiwara T, McCallum N, Sugai M (2004) Reduced content of lysyl-phosphatidylglycerol in the cytoplasmic membrane affects susceptibility to moenomycin, as well as vancomycin, gentamicin, and antimicrobial peptides, in Staphylococcus aureus. Antimicrob Agents Chemother 48(12):4800–4807
Noore J, Noore A, Li B (2012) Cationic antimicrobial peptide LL-37 is effective against both extra-and intra-cellular staphylococcus aureus. Antimicrob Agents Chemother 57(3):1283–1290
Oddo A, Hansen PR (2017) Hemolytic activity of antimicrobial peptides. In: Hansen PR (ed) Antimicrobial peptides: methods and protocols. Methods in molecular biology, vol 1548. Humana Press, New York, pp 427–435
Oh H, Hedberg M, Wade D, Edlund C (2000) Activities of synthetic hybrid peptides against anaerobic bacteria: aspects of methodology and stability. Antimicrob Agents Chemother 44(1):68–72
Oliver PM, Crooks JA, Leidl M, Yoon EJ, Saghatelian A, Weibel DB (2014) Localization of anionic phospholipids in Escherichia coli cells. J Bacteriol 196(19):3386–3398
Op den Kamp JAF, Redai I, van Deenen LLM (1969) Phospholipid composition of Bacillus subtilis. J Bacteriol 99(1):298–303
Oren Z, Shai Y (1997) Selective lysis of bacteria but not mammalian cells by diastereomers of melittin: structure-function study. Biochemistry 36(7):1826–1835
Oren Z, Shai Y (2000) Cyclization of a cytolytic amphipathic α-helical peptide and its diastereomer: effect on structure, interaction with model membranes, and biological function. Biochemistry 39(20):6103–6114
Oren Z, Lerman JC, Gudmundsson GH, Agerberth B, Shai Y (1999) Structure and organization of the human antimicrobial peptide LL-37 in phospholipid membranes: relevance to the molecular basis for its non-cell-selective activity. Biochem J 341(3):501–513
Orioni B, Bocchinfuso G, Kim JY, Palleschi A, Grande G, Bobone S, Park Y, Kim JI, Hahm KS, Stella L (2009) Membrane perturbation by the antimicrobial peptide PMAP-23: a fluorescence and molecular dynamics study. Biochim Biophys Acta 1788(7):1523–1533
Osborn MJ, Gander JE, Parisi E, Carson J (1972) Mechanism of assembly of the outer membrane of Salmonella typhimurium isolation and characterization of cytoplasmic and outer membrane. J Biol Chem 247(12):3962–3972
Ostovar A, Assadi M, Vahdat K, Nabipour I, Javadi H, Eftekhari M, Assadi M (2013) A pooled analysis of diagnostic value of 99mTc-ubiquicidin (UBI) scintigraphy in detection of an infectious process. Clin Nucl Med 38(6):413–416
Otvos L (2017) Racing on the wrong track. Front Chem 5:42
Otvos L Jr, Bokonyi K, Varga I, Otvos BI, Hoffmann R, Ertl HC, Wade JD, McManus AM, Craik DJ, Bulet P (2000) Insect peptides with improved protease-resistance protect mice against bacterial infection. Protein Sci 9(4):742–749
Pachón-Ibáñez ME, Smani Y, Pachón J, Sánchez-Céspedes J (2017) Perspectives for clinical use of engineered human host defense antimicrobial peptides. FEMS Microbiol Rev 41(3):323–342
Pandey BK, Ahmad A, Asthana N, Azmi S, Srivastava RM, Srivastava S, Verma R, Vishwakarma AL, Ghosh JK (2010) Cell-selective lysis by novel analogues of melittin against human red blood cells and Escherichia coli. Biochemistry 49(36):7920–7929
Pandey BK, Srivastava S, Singh M, Ghosh JK (2011) Inducing toxicity by introducing a leucine-zipper-like motif in frog antimicrobial peptide, magainin 2. Biochem J 436(3):609–620
Panteleev P, Bolosov IA, Balandin SV, Ovchinnikova TV (2015) Design of antimicrobial peptide arecinin analogs with improved therapeutic indices. J Pept Sci 21(2):105–113
Papo N, Oren Z, Pag U, Sahl HG, Shai Y (2002) The consequence of sequence alteration of an amphipathic α-helical antimicrobial peptide and its diastereomers. J Biol Chem 277(37):33913–33921
Park Y, Lee DG, Jang SH, Woo EH, Jeong HG, Choi CH, Hahm KS (2003) A Leu-Lys-rich antimicrobial peptide: activity and mechanism. Biochim Biophys Acta 1645(2):172–182
Pasupuleti M, Schmidtchen A, Chalupka A, Ringstad L, Malmsten M (2009) End-tagging of ultra-short antimicrobial peptides by W/F stretches to facilitate bacterial killing. PLoS One 4(4):e5285
Patel JB, Cockerill FR, Bradford PA, Eliopulos GM, Hindler JA, Jenkins SG, Lewis II JS, Limbago B, Miller LA, Nicolau DP, Powell DP, Swenson JM, Traczewski MM, Turnidge JD, Weistein MP, Zimmer BL (2012) Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically, approved standard, 10th edn. Clinical and Laboratory Standards Institute CLSI document M07-A10
Perron GG, Zasloff M, Bell G (2006) Experimental evolution of resistance to an antimicrobial peptide. Proc Biol Sci 273(1583):251–256
Phadke SM, Islam K, Deslouches B, Kapoor SA, Stolz DB, Watkins SC, Montelaro RC, Pilewski JM, Mietzner TA (2003) Selective toxicity of engineered lentivirus lytic peptides in a CF airway cell model. Peptides 24(8):1099–1107
Phoenix DA, Harris F (2002) The hydrophobic moment and its use in the classification of amphiphilic structures. Mol Membr Biol 19(1):1–10
Phoenix DA, Dennison SR, Harris F (eds) (2012) Antimicrobial peptides. Wiley, New York
Phoenix DA, Harris F, Mura M, Dennison SR (2015) The increasing role of phosphatidylethanolamine as a lipid receptor in the action of host defence peptides. Prog Lipid Res 59:26–37
Qiao Z, Lei C, Fu Y, Li Y (2017) Rapid and sensitive detection of E. coli O157: H7 based on antimicrobial peptide functionalized magnetic nanoparticles and urease-catalyzed signal amplification. Anal Methods 9(35):5204–5210
Raetz CR (1986) Molecular genetics of membrane phospholipid synthesis. Annu Rev Genet 20(1):253–291
Raimondo D, Andreotti G, Saint N, Amodeo P, Renzone G, Sanseverino M, Zocchi I, Molle G, Motta A, Scaloni A (2005) A folding-dependent mechanism of antimicrobial peptide resistance to degradation unveiled by solution structure of distinction. Proc Natl Acad Sci U S A 102(18):6309–6314
Rautenbach M, Troskie AM, Vosloo JA (2016) Antifungal peptides: to be or not to be membrane active. Biochimie 130:132–145
Ravi J, Bella A, Correia AJ, Lamarre B, Ryadnov MG (2015) Supramolecular amphipathicity for probing antimicrobial propensity of host defence peptides. Phys Chem Chem Phys 17(24):15608–15614
Renner LD, Weibel DB (2011) Cardiolipin microdomains localize to negatively curved regions of Escherichia coli membranes. Proc Natl Acad Sci U S A 108(15):6264–6269
Rivas L, Luque-Ortega JR, Andreu D (2009) Amphibian antimicrobial peptides and Protozoa: lessons from parasites. Biochim Biophys Acta 1788(8):1570–1581
Rowlett VW, Mallampalli VK, Karlstaedt A, Dowhan W, Taegtmeyer H, Margolin W, Vitrac H (2017) The impact of membrane phospholipid alterations in Escherichia coli on cellular function and bacterial stress adaptation. J Bacteriol 199(13). https://doi.org/10.1128/JB.00849-16
Ruiz J, Calderon J, Rondón-Villarreal P, Torres R (2014) Analysis of structure and hemolytic activity relationships of antimicrobial peptides (AMPs). In: Advances in computational biology. Springer, Cham, pp 253–258
Russell AL, Kennedy AM, Spuches AM, Venugopal D, Bhonsle JB, Hicks RP (2010) Spectroscopic and thermodynamic evidence for antimicrobial peptide membrane selectivity. Chem Phys Lipids 163(6):488–497
Saeed S, Zafar J, Khan B, Akhtar A, Qurieshi S, Fatima S, Ahmad N, Irfanullah J (2013) Utility of 99mTc-labelled antimicrobial peptide ubiquicidin (29-41) in the diagnosis of diabetic foot infection. Eur J Nucl Med Mol Imaging 40(5):737–743
Salick DA, Kretsinger JK, Pochan DJ, Schneider JP (2007) Inherent antibacterial activity of a peptide-based β-hairpin hydrogel. J Am Chem Soc 129(47):14793–14799
Sal-Man N, Oren Z, Shai Y (2002) Preassembly of membrane-active peptides is an important factor in their selectivity toward target cells. Biochemistry 41(39):11921–11930
Santos NC, Prieto M, Castanho MA (2003) Quantifying molecular partition into model systems of biomembranes: an emphasis on optical spectroscopic methods. Biochim Biophys Acta 1612(2):123–135
Savini F, Luca V, Bocedi A, Massoud R, Park Y, Mangoni ML, Stella L (2017) Cell-density dependence of host-defense peptide activity and selectivity in the presence of host cells. ACS Chem Biol 12(1):52–56
Savini F, Bobone S, Roversi D, Mangoni ML, Stella L (2018) From liposomes to cells: filling the gap between physicochemical and microbiological studies of the activity and selectivity of host-defense peptides. Pept Sci 110(5):e24041
Schmidtchen A, Pasupuleti M, Mörgelin M, Davoudi M, Alenfall J, Chalupka A, Malmsten M (2009) Boosting antimicrobial peptides by hydrophobic oligopeptide end-tags. J Biol Chem 284(26):17584–17594
Schmidtchen A, Ringstad L, Kasetty G, Mizuno H, Rutland MW, Malmsten M (2011) Membrane selectivity by W-tagging of antimicrobial peptides. Biochim Biophys Acta 1808(4):1081–1091
Schmidtchen A, Pasupuleti M, Malmsten M (2014) Effect of hydrophobic modifications in antimicrobial peptides. Adv Colloid Interf Sci 205:265–274
Schröder-Borm H, Willumeit R, Brandenburg K, Andrä J (2003) Molecular basis for membrane selectivity of NK-2, a potent peptide antibiotic derived from NK-lysin. Biochim Biophys Acta 1612(2):164–171
Schweizer F (2009) Cationic amphiphilic peptides with cancer-selective toxicity. Eur J Pharmacol 625(1–3):190–194
Seelig J (2004) Thermodynamics of lipid–peptide interactions. Biochim Biophys Acta 1666(1):40–50
Seo MD, Won HS, Kim JH, Mishig-Ochir T, Lee BJ (2012) Antimicrobial peptides for therapeutic applications: a review. Molecules 17(10):12276–12286
Shai Y, Oren Z (1996) Diastereomers of cytolysins, a novel class of potent antibacterial peptides. J Biol Chem 271(13):7305–7308
Shai Y, Oren Z (2001) From “carpet” mechanism to de-novo designed diastereomeric cell-selective antimicrobial peptides. Peptides 22(10):1629–1641
Shankar SS, Benke SN, Nagendra N, Srivastava PL, Thulasiram HV, Gopi HN (2013) Self-assembly to function: design, synthesis, and broad spectrum antimicrobial properties of short hybrid E-vinylogous lipopeptides. J Med Chem 56(21):8468–8474
Shi X, Zhang X, Yao Q, He F (2017) A novel method for the rapid detection of microbes in blood using pleurocidin antimicrobial peptide functionalized piezoelectric sensor. J Microbiol Methods 133:69–75
Shin SY, Yang ST, Park EJ, Eom SH, Song WK, Kim JI, Lee SH, Lee MK, Lee DG, Hahm KS, Kim Y (2001) Antibacterial, antitumor and hemolytic activities of α-helical antibiotic peptide, P18 and its analogs. J Pept Res 58(6):504–514
Shriver-Lake LC, North SH, Dean SN, Taitt CR (2012) Antimicrobial peptides for detection and diagnostic assays. In: Designing receptors for the next generation of biosensors. Springer, Berlin, pp 85–104
Silva RR, Avelino KY, Ribeiro KL, Franco OL, Oliveira MD, Andrade CA (2014) Optical and dielectric sensors based on antimicrobial peptides for microorganism diagnosis. Front Microbiol 5:443. https://doi.org/10.3389/fmicb.2014.00443
Simon SA, McIntosh TJ (2002) Peptide-lipid interactions, current topics in membranes, vol 52. Elsevier, Amsterdam
Skerlavaj B, Renato Gennaro R, Luigi Bagella L, Laura Merluzzi L, Angela Risso A, Zanetti M (1996) Biological characterization of two novel cathelicidin-derived peptides and identification of structural requirements for their antimicrobial and cell lytic activities. J Biol Chem 271(45):28375–28381
Slaninová J, Mlsová V, Kroupová H, Alán L, Tůmová T, Monincová L, Borovičková L, Fučík V, Ceřovský V (2012) Toxicity study of antimicrobial peptides from wild bee venom and their analogs toward mammalian normal and cancer cells. Peptides 33(1):18–26
Snoussi M, Talledo JP, Del Rosario NA, Ha BY, Kosmrlj A, Taheri-Araghi S (2018) Heterogeneous absorption of antimicrobial peptide LL37 in Escherichia coli cells enhances population survivability. eLife 7:e38174
Son M, Lee Y, Hwang H, Hyun S, Yu J (2013) Disruption of interactions between hydrophobic residues on nonpolar faces is a key determinant in decreasing hemolysis and increasing antimicrobial activities of α-helical amphipathic peptides. ChemMedChem 8(10):1638–1642
Song YM, Yang ST, Lim SS, Kim Y, Hahm KS, Kim JI, Shin SY (2004) Effects of L-or D-Pro incorporation into hydrophobic or hydrophilic helix face of amphipathic α-helical model peptide on structure and cell selectivity. Biochem Biophys Res Commun 314(2):615–621
Song YM, Park Y, Lim SS, Yang ST, Woo ER, Park IS, Lee JS, Kim JI, Hahm KS, Kim Y, Shin SY (2005) Cell selectivity and mechanism of action of antimicrobial model peptides containing peptoid residues. Biochemistry 44(36):12094–12106
Sood R, Kinnunen PK (2008) Cholesterol, lanosterol, and ergosterol attenuate the membrane association of LL-37 (W27F) and temporin L. Biochim Biophys Acta 1778(6):1460–1466
Sood R, Domanov Y, Pietiäinen M, Kontinen VP, Kinnunen PK (2008) Binding of LL-37 to model biomembranes: insight into target vs host cell recognition. Biochim Biophys Acta 1778(4):983–996
Stark M, Liu LP, Deber CM (2002) Cationic hydrophobic peptides with antimicrobial activity. Antimicrob Agents Chemother 46(11):3585–3590
Starr CG, He J, Wimley WC (2016) Host cell interactions are a significant barrier to the clinical utility of peptide antibiotics. ACS Chem Biol 11(12):3391–3399
Steiner H, Andreu D, Merrifield RB (1988) Binding and action of cecropin and cecropin analogues: antibacterial peptides from insects. Biochim Biophys Acta 939(2):260–266
Stella L, Mazzuca C, Venanzi M, Palleschi A, Didone M, Formaggio F, Toniolo C, Pispisa B (2004) Aggregation and water-membrane partition as major determinants of the activity of the antibiotic peptide trichogin GA IV. Biophys J 86(2):936–945
Storch J, Kleinfeld AM (1985) The lipid structure of biological membranes. Trends Biochem Sci 10(11):418–421
Strandberg E, Tiltak D, Ieronimo M, Kanithasen N, Wadhwani P, Ulrich AS (2007) Influence of C-terminal amidation on the antimicrobial and hemolytic activities of cationic -helical peptides. Pure Appl Chem 79(4):717–728
Strömstedt AA, Ringstad L, Schmidtchen A, Malmsten M (2010) Interaction between amphiphilic peptides and phospholipid membranes. Curr Opin Colloid Interface Sci 15(6):467–478
Swierstra J, Kapoerchan V, Knijnenburg A, van Belkum A, Overhand M (2016) Structure, toxicity and antibiotic activity of gramicidin S and derivatives. Eur J Clin Microbiol Infect Dis 35(5):763–769
Tachi T, Epand RF, Epand RM, Matsuzaki K (2002) Position-dependent hydrophobicity of the antimicrobial magainin peptide affects the mode of peptide-lipid interactions and selective toxicity. Biochemistry 41(34):10723–10731
Takahashi D, Shukla SK, Prakash O, Zhang G (2010) Structural determinants of host defense peptides for antimicrobial activity and target cell selectivity. Biochimie 92(9):1236–1241
Teixeira V, Feio MJ, Bastos M (2012) Role of lipids in the interaction of antimicrobial peptides with membranes. Prog Lipid Res 51(2):149–177
Thennarasu S, Nagaraj R (1996) Specific antimicrobial and hemolytic activities of 18-residue peptides derived from the amino terminal region of the toxin pardaxin. Protein Eng Des Sel 9(12):1219–1224
Tian X, Sun F, Zhou XR, Luo SZ, Chen L (2015) Role of peptide self-assembly in antimicrobial peptides. J Pept Sci 21(7):530–539
Tiozzo E, Rocco G, Tossi A, Romeo D (1998) Wide-spectrum antibiotic activity of synthetic, amphipathic peptides. Biochem Biophys Res Commun 249(1):202–206
Toniolo C, Crisma M, Formaggio F, Peggion C, Monaco V, Goulard C, Rebuffat S, Bodo B (1996) Effect of N α-acyl chain length on the membrane-modifying properties of synthetic analogs of the lipopeptaibol trichogin GA IV. J Am Chem Soc 118(21):4952–4958
Tossi A (2011) Design and engineering strategies for synthetic antimicrobial peptides. In: Prokaryotic antimicrobial peptides. Springer, New York, pp 81–98
Tossi A, Sandri L, Giangaspero A (2000) Amphipathic, α-helical antimicrobial peptides. Biopolymers 55(1):4–30
Tu Z, Hao J, Kharidia R, Meng XG, Liang JF (2007) Improved stability and selectivity of lytic peptides through self-assembly. Biochem Biophys Res Commun 361(3):712–717
Tytler EM, Anantharamaiah GM, Walker DE, Mishra VK, Palgunachari MN, Segrest JP (1995) Molecular basis for prokaryotic specificity of magainin-induced lysis. Biochemistry 34(13):4393–4401
Uematsu N, Matsuzaki K (2000) Polar angle as a determinant of amphipathic α-helix-lipid interactions: a model peptide study. Biophys J 79(4):2075–2083
Uggerhøj LE, Poulsen TJ, Munk JK, Fredborg M, Sondergaard TE, Frimodt-Moller N, Hansen PR, Wimmer R (2015) Rational design of alpha-helical antimicrobial peptides: do’s and don’ts. ChemBioChem 16(2):242–253
Vallejo E, Martinez I, Tejero A, Hernandez S, Jimenez L, Bialostozky D, Sanchez G, Ilarraza H, Ferro-Flores G (2008) Clinical utility of 99mTc-labeled ubiquicidin 29–41 antimicrobial peptide for the scintigraphic detection of mediastinitis after cardiac surgery. Arch Med Res 39(8):768–774
van der Weerden NL, Bleackley MR, Anderson MA (2013) Properties and mechanisms of action of naturally occurring antifungal peptides. Cell Mol Life Sci 70(19):3545–3570
Veiga AS, Sinthuvanich C, Gaspar D, Franquelim HG, Castanho MA, Schneider JP (2012) Arginine-rich self-assembling peptides as potent antibacterial gels. Biomaterials 33(35):8907–8916
Velduhizen EJA, Scheenstra MR, Tjeerdsma-van Bokhoven JLM, Coorens M, Schneider VAF, Bikker FJ, van Dijk A, Haagsman HP (2017) Antimicrobial and immunomodulatory activity of PMAP-23 derived peptides. Protein Pept Lett 24(7):609–616
Verkleij AJ, Zwaal RFA, Roelofsen B, Comfurius P, Kastelijn D, van Deenen LLM (1973) The asymmetric distribution of phospholipids in the human red cell membrane. A combined study using phospholipases and freeze-etch electron microscopy. Biochim Biophys Acta 323(2):178–193
Verly RM, Rodrigues MA, Daghastanli KRP, Denadai AML, Cuccovia IM, Bloch C Jr, Frezard F, Santoro MM, Pilo-Veloso D, Bemquerer MP (2008) Effect of cholesterol on the interaction of the amphibian antimicrobial peptide DD K with liposomes. Peptides 29(1):15–24
Vermeer LS, Lan Y, Abbate V, Ruh E, Bui TT, Wilkinson L, Jumagulova E, Kozlowska J, Patel J, McIntyre CA, Yam WC, Siu GKH, Atkinson RA, Lam JKW, Bansal SS, Drake AF, Mitchell GH, Mason AJ (2012) Conformational flexibility determines selectivity and Antibacterial, Antiplasmodium, and Anticancer potency of cationic α-helical peptides. J Biol Chem 287(41):34120–34133
Virtanen JA, Cheng KH, Somerharju P (1998) Phospholipid composition of the mammalian red cell membrane can be rationalized by a superlattice model. Proc Natl Acad Sci U S A 95(9):4964–4969
Wade D, Boman A, Wåhlin B, Drain CM, Andreu D, Boman HG, Merrifield RB (1990) All-D amino acid-containing channel-forming antibiotic peptides. Proc Natl Acad Sci U S A 87(12):4761–4765
Wade D, Silberring J, Soliymani R, Heikkinen S, Kilpeläinen I, Lankinen H, Kuusela P (2000) Antibacterial activities of temporin A analogs. FEBS Lett 479(1–2):6–9
Wakabayashi H, Matsumoto H, Hashimoto K, Teraguchi S, Takase M, Hayasawa H (1999) N-Acylated and D enantiomer derivatives of a nonamer core peptide of lactoferricin B showing improved antimicrobial activity. Antimicrob Agents Chemother 43(5):1267–1269
Wang G (ed) (2017) Antimicrobial peptides: discovery, design and novel therapeutic strategies. CABI Publishing, Cambridge, MA
Wang H, Xu K, Liu L, Tan JP, Chen Y, Li Y, Fan W, Wei Z, Sheng J, Yang YY, Li L (2010) The efficacy of self-assembled cationic antimicrobial peptide nanoparticles against Cryptococcus neoformans for the treatment of meningitis. Biomaterials 31(10):2874–2881
Wang J, Chou S, Xu L, Zhu X, Dong N, Shan A, Chen Z (2015) High specific selectivity and membrane-active mechanism of the synthetic centrosymmetric α-helical peptides with Gly-Gly pairs. Sci Rep 5:15963
Wang J, Chou S, Yang Z, Yang Y, Wang Z, Song J, Dou X, Shan A (2018) Combating drug-resistant fungi with novel imperfectly amphipathic palindromic peptides. J Med Chem 61(9):3889–3907
Welling MM, Nibbering PH, Paulusma-Annema A, Hiemstra PS, Pauwels EK, Calame W (1999) Imaging of bacterial infections with 99mTc-labeled human neutrophil peptide-1. J Nucl Med 40(12):2073–2080
Welling MM, Paulusma-Annema A, Balter HS, Pauwels EK, Nibbering PH (2000) Technetium-99m labelled antimicrobial peptides discriminate between bacterial infections and sterile inflammations. Eur J Nucl Med 27(3):292–301
White DA (1973) The phospholipid composition of mammalian tissues. In: Ansell GB, Hawthorne JN, Dawson RMC (eds) Form and function of phospholipids. Elsevier, Amsterdam, pp 441–482
White SH, Wimley WC (1999) Membrane protein folding and stability: physical principles. Annu Rev Biophys Biomol Struct 28(1):319–365
Wiegand I, Hilpert K, Hancock REW (2008) Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc 3(2):163–175
Wieprecht T, Seelig J (2002) Isothermal titration calorimetry for studying interactions between peptides and lipid membranes. Curr Top Membr 52:31–56
Wieprecht T, Dathe M, Beyermann M, Krause E, Maloy WL, MacDonald DL, Bienert M (1997a) Peptide hydrophobicity controls the activity and selectivity of magainin 2 amide in interaction with membranes. Biochemistry 36(20):6124–6132
Wieprecht T, Dathe M, Krause E, Beyermann M, Maloy WL, MacDonald DL, Bienert M (1997b) Modulation of membrane activity of amphipathic, antibacterial peptides by slight modifications of the hydrophobic moment. FEBS Lett 417(1):135–140
Wieprecht T, Dathe M, Epand RM, Beyermann M, Krause E, Maloy WL, MacDonald DL, Bienert M (1997c) Influence of the angle subtended by the positively charged helix face on the membrane activity of amphipathic, antibacterial peptides. Biochemistry 36(42):12869–12880
Wimley WC (2010a) Energetics of peptide and protein binding to lipid membranes. In: Proteins membrane binding and pore formation. Springer, New York, pp 14–23
Wimley WC (2010b) Describing the mechanism of antimicrobial peptide action with the interfacial activity model. ACS Chem Biol 5(10):905–917
Wimley WC, Hristova K (2011) Antimicrobial peptides: successes, challenges and unanswered questions. J Membr Biol 239(1–2):27–34
Wu G, Wu H, Fan X, Zhao R, Li X, Wang S, Ma Y, Shen Z, Xi T (2010) Selective toxicity of antimicrobial peptide S-thanatin on bacteria. Peptides 31(9):1669–1673
Yang ST, Shin SY, Kim YC, Kim Y, Hahm KS, Kim JI (2002) Conformation-dependent antibiotic activity of tritrpticin, a cathelicidin-derived antimicrobial peptide. Biochem Biophys Res Commun 296(5):1044–1050
Yang ST, Lee JY, Kim HJ, Eu YJ, Shin SY, Hahm KS, Kim JI (2006a) Contribution of a central proline in model amphipathic α-helical peptides to self-association, interaction with phospholipids, and antimicrobial mode of action. FEBS J 273(17):4040–4054
Yang ST, Jeon JH, Kim Y, Shin SY, Hahm KS, Kim JI (2006b) Possible role of a PXXP central hinge in the antibacterial activity and membrane interaction of PMAP-23, a member of cathelicidin family. Biochemistry 45(6):1775–1784
Yau WM, Wimley WC, Gawrisch K, White SH (1998) The preference of tryptophan for membrane interfaces. Biochemistry 37(42):14713–14718
Yeaman MR, Yount NY (2003) Mechanisms of antimicrobial peptide action and resistance. Pharmacol Rev 55(1):27–55
Yeung AT, Gellatly SL, Hancock REW (2011) Multifunctional cationic host defence peptides and their clinical applications. Cell Mol Life Sci 68(13):2161
Zelezetsky I, Tossi A (2006) Alpha-helical antimicrobial peptides-using a sequence template to guide structure–activity relationship studies. Biochim Biophys Acta 1758(9):1436–1449
Zelezetsky I, Pacor S, Pag U, Papo N, Shai Y, Sahl HG, Tossi A (2005) Controlled alteration of the shape and conformational stability of α-helical cell-lytic peptides: effect on mode of action and cell specificity. Biochem J 390(1):177–188
Zhang L, Benz R, Hancock REW (1999) Influence of proline residues on the antibacterial and synergistic activities of R-helical peptides. Biochemistry 38(25):8102–8111
Zhang Y, Lu H, Lin Y, Cheng J (2011) Water-soluble polypeptides with elongated, charged side chains adopt ultrastable helical conformations. Macromolecules 44(17):6641–6644
Zhang SK, Song JW, Gong F, Li SB, Chang HY, Xie HM, Gao HW, Tan YX, Ji SP (2016) Design of an α-helical antimicrobial peptide with improved cell-selective and potent anti-biofilm activity. Sci Rep 6:27394
Zhou NE, Mant CT, Hodges RS (1990) Effect of preferred binding domains on peptide retention behavior in reversed-phase chromatography: amphipathic alpha-helices. Pept Res 3(1):8–20
Zhu WL, Hahm K, Shin SY (2007a) Cathelicidin-derived Trp/Pro-rich antimicrobial peptides with lysine peptoid residue (Nlys): therapeutic index and plausible mode of action. J Pept Sci 13(8):529–535
Zhu WL, Song YM, Park Y, Park KH, Yang ST, Kim JI, Park IS, Hahm KS, Shin SY (2007b) Substitution of the leucine zipper sequence in melittin with peptoid residues affects self-association, cell selectivity, and mode of action. Biochim Biophys Acta 1768(6):1506–1517
Zhu WL, Nan YH, Hahm K, Shin SY (2007c) Cell selectivity of an antimicrobial peptide melittin diastereomer with D-amino acid in the leucine zipper sequence. J Biochem Mol Biol 40(6):1090–1094
Zhu WL, Hahm KS, Shin SY (2009) Cell selectivity and mechanism of action of short antimicrobial peptides designed from the cell-penetrating peptide Pep-1. J Pept Sci 15(9):569–575
Zou R, Zhu X, Tu Y, Wu J, Landry MP (2018) Activity of antimicrobial peptide aggregates decreases with increased cell membrane embedding free energy cost. Biochemistry 57(18):2606–2610
Zwaal RFA, Roelofsen B, Colley CM (1973) Localization of red cell membrane constituents. Biochim Biophys Acta 300(2):159–182
Zwaal RFA, Roelsfsen B, Comfurius P, van Deenen LLM (1975) Organization of phospholipids in human red cell membranes as detected by the action of various purified phospholipases. Biochim Biophys Acta 406(1):83–96
Acknowledgments
The authors gratefully thank Dr. F. Savini and Dr. A. Papi for their help with Figs. 11.3 and 11.5. Research in our lab is currently supported by the Italian Ministry for Education, University and Research (grant PRIN 20157WW5EH_007), and by the Italian Association for Cancer Research (AIRC grant IG 2016 19171).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Bobone, S., Stella, L. (2019). Selectivity of Antimicrobial Peptides: A Complex Interplay of Multiple Equilibria. In: Matsuzaki, K. (eds) Antimicrobial Peptides. Advances in Experimental Medicine and Biology, vol 1117. Springer, Singapore. https://doi.org/10.1007/978-981-13-3588-4_11
Download citation
DOI: https://doi.org/10.1007/978-981-13-3588-4_11
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-3587-7
Online ISBN: 978-981-13-3588-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)