Abstract
Peroxisomes are dynamic organelles of eukaryotic cells performing a wide range of functions including fatty acid oxidation, peroxide detoxification and ether-lipid synthesis in mammals. Peroxisomes lack their own DNA and therefore have to import proteins post-translationally. Peroxisomes can import folded, co-factor bound and even oligomeric proteins. The involvement of cycling receptors is a special feature of peroxisomal protein import. Complex machineries of peroxin (PEX) proteins mediate peroxisomal matrix and membrane protein import. Identification of PEX genes was dominated by forward genetic techniques in the early 90s. However, recent developments in proteomic techniques has revolutionized the detailed characterization of peroxisomal protein import. Here, we summarize the current knowledge on peroxisomal protein import with emphasis on the contribution of proteomic approaches to our understanding of the composition and function of the peroxisomal protein import machineries.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Peroxisome
- Peroxin
- Peroxisome targeting signal
- Piggyback protein import
- Cycling import receptors
- Peroxisomal import pores
1 Peroxisomal Protein Import
Peroxisomes are the ubiquitous organelles of eukaryotic cells, which compartmentalize diverse metabolic functions, including fatty acid oxidation and detoxification of reactive oxygen species. Peroxisomes are dynamic organelles, which can adapt their protein composition to the cellular needs. This feature of peroxisomes contributes to the capacity of cells to adapt to environmental changes. Peroxisome-like organelles are present in certain organisms and include Woronin bodies in filamentous fungi, glyoxysomes in plants and glycosomes in trypanosomatid parasites. Peroxisomes are single membrane bound organelles devoid of their own DNA, requiring that all peroxisomal proteins are encoded in the nucleus and that they have to be targeted to peroxisomes. Accordingly, peroxisomal matrix proteins are synthesized on free ribosomes in the cytosol and imported post-translationally. Hallmark features of peroxisomal protein import are the involvement of cycling receptors and the import of folded, co-factor bound even oligomeric proteins/complexes (Fig. 1).
Peroxisomes can post-translationally import folded proteins, co-factor bound proteins, homo- or heteromeric protein complexes, even gold nanoparticles coated with peroxisome targeting signals. Some proteins lack a peroxisome targeting signal but can enter peroxisomes by piggybacking. These proteins hitchhike to peroxisomes by binding to PTS-containing proteins in the cytosol
Proteins involved in the biogenesis of peroxisomes are termed peroxins (PEX). In the early 90s, forward genetic screening of yeast mutants (Erdmann et al. 1989) or mutant mammalian cells (Tsukamoto et al. 1990) defective in peroxisome assembly was instrumental for the identification of majority of PEX genes. Later on, further PEX genes were originally identified by a complementary reverse genetic approach based on the isolation of peroxisomal membrane proteins and partial protein sequencing and the use of the sequence information for cloning of the corresponding genes by PCR using degenerated primers (Erdmann and Blobel 1995). This reverse genetic approach to study peroxisome biogenesis requires the isolation of purified peroxisomal membranes or PEX protein-complexes. The strategies for these are reviewed in this issue by Islinger et al. (Chap. 4), Klümper et al. (Chap. 11), and Okumoto et al. (Chap. 12). Genetic defects in PEX genes are responsible for several severe inborn disorders in humans, like the Zellweger syndrome. These patients usually die in early infancy. Glycosomes are peroxisome-like organelles of trypanosomatid parasites, which compartmentalize glycolysis inside the organelle. Glycosomes are essential for parasite survival and differences in peroxisome and glycosome biogenesis recently have been exploited as drug targets against trypanosomatid infections (Dawidowski et al. 2017).
So far, 36 peroxins have been identified in yeast, 14 in humans and 16 in plants. Of these, three peroxins are involved in membrane protein targeting, while majority play a role in targeting of peroxisomal matrix proteins (detailed in later sections) or other aspects of peroxisome maintenance (Table 1). Targeting of peroxisomal matrix proteins to the organelle depends on the presence of peroxisomal targeting signals (PTS), or piggybacking on proteins, which harbor PTS. Likewise, membrane proteins destined for peroxisomal membrane contain membrane peroxisome targeting signal (mPTS).
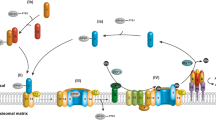
Peroxisomal protein targeting involves distinct steps; cargo recognition by the targeting receptor, docking of the cargo-receptor complex at the peroxisomal membrane, translocation of matrix proteins or insertion of membrane proteins and recycling of the receptor (Fig. 2). These steps involve the interaction of distinct sets of peroxins. Here, we review the current knowledge of peroxisomal protein import machineries, with emphasis on the proteomic approaches, which contributed to our understanding of their composition and function.
Peroxisomal matrix proteins are synthesized in the cytosol on free ribosomes and imported post-translationally into peroxisomes. Peroxisomal protein import requires peroxin (PEX) proteins (individual PEX protein shown in bold numbers) and involves different steps (Roman numerals in brackets): (I) Cytosolic import receptors Pex5p and Pex7p along with co-receptor Pex18p/Pex21p recognize the cargo proteins via peroxisome targeting signals, PTS1 and PTS2 respectively. (II) The cargo-receptor complexes are transported to the peroxisomal membrane by binding to the docking complex comprised of Pex14p, Pex13p and Pex17p. In this scheme, emphasis is on the import of PTS1 proteins. (III) Association of Pex14p with Pex5p or Pex18p results in the formation of respective distinct transient PTS1 and PTS2 protein import pores. After the translocation of cargo proteins into the peroxisomal lumen, the import receptors are released from the peroxisomal membrane and recycled for the next round of protein import. (IV) Receptor recycling involves ubiquitination by the ubiquitin-conjugating (E2) enzyme Pex4p and its membrane anchor Pex22p, as well as the complex of RING (Really Interesting New Gene) finger domain containing E3 ubiquitin ligases Pex2p, Pex10p, Pex12p. Intra-peroxisomal Pex8p connects the docking complex to the RING-finger complex. (V) Mono-ubiquitination of Pex5p signals its ATP-dependent extraction from the membrane by the hetero-hexameric AAA-complex (Pex1p-Pex6p), which is anchored to the peroxisomal membrane by Pex15p. Poly-ubiquitination of Pex5p serves as a quality control mechanism that directs a supposedly non-functional receptor to proteasomal degradation
2 Peroxisomal Matrix Protein Import
2.1 Cargo Recognition
Proteins destined for import into peroxisomal matrix usually contain a type 1 (PTS1) or type 2 (PTS2) peroxisome targeting signal. The originally identified PTS1 is a conserved C-terminal tripeptide with the sequence SKL, first identified in Firefly luciferase (Gould et al. 1987). Sequence analysis of PTS1-containing proteins from different organisms and species led to a refined PTS1 consensus sequence [S/A/C][K/R/H][L/M] (Lametschwandtner et al. 1998). Additional residues upstream of the PTS1 tripeptide are less conserved but turned out to be relevant for recognition of the PTS1 signal (Brocard and Hartig 2006; Hagen et al. 2015). Pex5p is the cytosolic receptor for PTS1-containing proteins and it recognizes the PTS1-signal via its carboxy-terminal tetratricopeptide repeat (TPR) domains (Terlecky et al. 1995).
The type 2 peroxisome targeting signal (PTS2) is a nonapeptide located near the N-terminal part of the proteins (Swinkels et al. 1991) with consensus sequence R[L/V/I/Q]xx[L/V/I/H][L/S/G/A]x[H/Q][L/A] (Petriv et al. 2004). Pex7p is the cytosolic receptor for PTS2 containing proteins and the targeting signal is recognized by tryptophan-aspartate (WD-40) repeats of the receptor (Marzioch et al. 1994; Pan et al. 2013). Compared to PTS1, only few mammalian peroxisomal proteins harbor PTS2 and only two PTS2 proteins are known in yeast, thiolase (Fox3p) and glycerol phosphate dehydrogenase (Gpd1p) (Glover et al. 1994; Jung et al. 2010). However, one-third of peroxisomal proteins in plants contain a PTS2 signal (Reumann 2004).
Reumann et al. (2007) isolated highly purified peroxisomes from Arabidopsis thaliana leaves and performed proteomic analysis using gel-based and gel-independent approaches. The study identified 78 non-redundant proteins of which 42 novel proteins were previously not known to be associated with plant peroxisomes. Sequence analysis and fluorescence microscopy of full-length fusion proteins led to the identification of functional PTS1-related peptides in plants. The PTS1 signal typically harbors a positively charged residue in the middle of the PTS1-tripeptide (Lametschwandtner et al. 1998). Plant peroxisomes could import proteins terminating with SSL, SSI, and ASL residues, thus demonstrating that PTS1-signal in plants can be highly degenerated (Reumann et al. 2007). The PTS2 signal is normally localized close to the N-terminus of proteins (Swinkels et al. 1991). The proteomic study by Reumann et al. (2007) also identified an internal PTS2-like signal in transthyretin-like protein (TLP, At5g58220). TLP is a bifunctional protein and the PTS2-like signal is located at position 182-190 between the N-terminal decarboxylase and the C-terminal hydrolase domain. Deletion of the internal PTS2-like signal in TLP abolished its peroxisomal targeting, suggesting that an internal sequence can function as PTS2 signal (Reumann et al. 2007).
Some proteins that are imported into peroxisomes in a Pex5p-dependent manner lack a typical PTS1 signal. Of these, acyl-CoA oxidase (S. cerevisiae, Y. lipolytica) and alcohol oxidase (H. polymorpha) interact with the N-terminal domain of Pex5p (Schäfer et al. 2004; van der Klei and Veenhuis 2006), however, our knowledge on the molecular details of this cargo recognition is still scarce. Another mode of peroxisomal import of non-PTS proteins is the piggy-back transport. Here, proteins that lack a PTS hitch-hike to peroxisomes via binding to proteins that harbor PTS signals. This is possible as peroxisomes can import folded and oligomeric proteins, as pointed out earlier. In yeast, enoyl-CoA isomerase (Eci1p) lacking its own PTS1 signal is imported into peroxisomes by oligomerizing with its PTS1-containing homolog Dci1p (Yang et al. 2001). Mammalian Cu/Zn superoxide dismutase 1 (SOD1) lacks an endogenous PTS but is imported by piggy-backing on its natural PTS1-containing binding partner the copper chaperone of SOD1 (CCS) (Islinger et al. 2009). Also the PTS2-import machinery allows the piggy-back import of non-PTS containing proteins. Examples include hitch-hiking of PTS2-truncated thiolase by dimerizing with endogenous thiolase in S. cerevisiae (Glover et al. 1994) and artificial cargo proteins in plants (Flynn et al. 1998).
A recent proteomic study identified a natural piggy-back import via the PTS2-import machinery (Effelsberg et al. 2015). Pnc1p is a nicotinamidase, which localizes to peroxisomes in Pex7p-dependent manner although it lacks a typical PTS2 signal (Anderson et al. 2003). In response to osmotic stress, yeast PTS2 protein Gpd1p shows a tripartite localization in peroxisomes, cytosol and nucleus. Pnc1p also shows similar dynamic localization pattern (Jung et al. 2010). Effelsberg et al. (2015) isolated Gpd1p from the soluble fractions of high-salt grown yeast cells expressing Gpd1-TPA by affinity purification. The isolated Gpd1-complex contained an additional protein, which was identified as Pnc1p by mass-spectrometry. Size-exclusion chromatography of Gpd1p TEV-eluate revealed the presence of a heterodimeric complex of Gpd1p-Pnc1p. This study showed that the piggyback peroxisomal import of Pnc1p depends on the presence of PTS2 signal in Gpd1p (Effelsberg et al. 2015).
Pex5p recognizes the carboxy-terminal PTS1 tripeptide in cargo proteins and the amino acid residues upstream of PTS1 signal also influence the cargo recognition by Pex5p (Lametschwandtner et al. 1998; Brocard and Hartig 2006). To gain further structural insights into the binding interface of Pex5p and cargo protein Pcs60p, Hagen et al. (2015) performed in vitro photo-crosslinking studies of recombinant proteins and mass-spectrometric analysis of the cross-linked products. The study revealed that Pcs60p lacking the C-terminal tripeptide of the PTS1 is still cross-linked to Pex5p, indicating that also other regions of the cargo are in close proximity to the receptor. High-resolution mass spectrometry of the cross-linked products revealed that the amino acids adjacent to the C-terminal tripeptide contribute to the interaction and the study also identified the corresponding binding interface of Pex5p. Further biophysical and biochemical analysis revealed two modes of Pex5p-Pcs60p interaction. The high affinity binding occurs between the PTS1 tripeptide and Pex5p, while the amino acids upstream of the PTS1 provide a low affinity second binding site and interact with a distinct region of the receptor (Hagen et al. 2015).
2.1.1 Pre-import Complexes
Prior to peroxisomal targeting, the receptor cargo complex is assembled in the cytosol (pre-import complex). In contrast to the PTS1-receptor Pex5p, Pex7p requires additional cytosolic peroxins as co-receptors for PTS2-protein import. The co-receptor family includes Pex18p and Pex21p (S. cerevisiae) and Pex20p (other yeasts and filamentous fungi). In mammals and rice, alternative splicing leads to two isoforms of PEX5, the longer isoform PEX5L and the shorter PEX5S. Arabidopsis and Trypanosomatids encode a single PEX5 transcript, which resembles the mammalian long isoform. Mammalian PEX5L and plant as well as T. brucei PEX5 also act as co-receptor for PEX7. The common theme across Pex7p co-receptors is the presence of Pex7p binding domain (reviewed in Schliebs and Kunau 2006).
To investigate the early steps of the import of PTS2-proteins, Grunau et al. (2009) performed isolation of the pre-import complexes and the mass-spectrometric analysis of their composition. Authors isolated cytosolic complexes of Protein-A tagged co-receptors Pex18p and Pex21p, as well as the PTS2-cargo protein Fox3p. Mass spectrometric analysis identified Fox3p as the only abundant PTS2 protein in oleate-grown yeast cells. The study revealed a sequential assembly of the ternary PTS2 pre-import complex in the cytosol. Pex7p first recognizes the PTS2-containing cargo proteins in the cytosol, followed by binding of the co-receptor to Pex7p-cargo complex and this ternary pre-import complex is targeted to the peroxisomal membrane.
2.2 Docking at the Peroxisomal Membrane
After cargo recognition, the cargo-receptor complex is targeted to the peroxisomes by binding to the docking complex at the peroxisomal membrane. The docking complex comprises Pex14p, Pex13p and Pex17p in yeast. The N-terminal domain of Pex5p contains several di-aromatic pentapeptide motifs with the consensus sequence Wxxx(F/Y), which are also present in PEX7 co-receptors (reviewed in Schliebs and Kunau 2006). These WxxxF motifs bind to the N-terminal domain in Pex14p with high affinity, providing the basis for membrane docking of the cargo-receptor complex. Pex14p interacts with both the PTS1- and the PTS2- receptor and is the point of convergence of both import pathways (Albertini et al. 1997). Pex14 is an integral membrane protein but also partially released by carbonate treatment, which is typical for peripheral membrane proteins. Pex13p is an integral membrane protein of peroxisomes and contains a C-terminal Src Homology 3 (SH3) domain. This SH3-domain faces the cytosol and interacts with Pex14p, which provides a classical proline-rich SH3-binding motif (PxxP) for the interaction (Pires et al. 2003). Absence of docking complex components Pex14p or Pex13p abolishes import of both PTS1 and PTS2 proteins (Albertini et al. 1997; Girzalsky et al. 1999; Gould et al. 1996). The Pex13p-Pex14p interaction is important for the correct peroxisomal localization of Pex14p in different organisms (Girzalsky et al. 1999; Fransen et al. 2004; Brennand et al. 2012). Evidence has been provided that docking of the cargo-loaded receptors is first performed by binding to Pex14p (Urquhart et al. 2000). However, also Pex13p binds the import receptors. The SH3-domain of Pex13p interacts with N-terminal domain of Pex5p in a non-PxxP manner with one of the WxxxF motifs representing the binding site (Bottger et al. 2000; Douangamath et al. 2002; Pires et al. 2003). On the other hand, Pex7p can directly bind to the YG-rich N-terminal domain of Pex13p (Girzalsky et al. 1999). Trypanosomatids are the only organisms, which encode two isoforms of PEX13, PEX13.1 and PEX13.2. Although PEX13.2 lacks the SH3-domain, both isoforms are essential for glycosomal protein import (Brennand et al. 2012).
In addition to the analysis of the cytosolic PTS2 pre-import complexes (see Sect. 2.1.1), Grunau et al. (2009) also performed proteomic analysis of the membrane bound PTS2-import complexes. The study revealed a high-molecular-weight complex of Pex14p and PTS2 co-receptor Pex18p at the peroxisomal membrane, which lacks Pex7p and Fox3p. This indicates that the PTS2 pre-import complex seems to dissociate after docking.
Pex17p is an additional component of docking complex but so far has been identified only in yeast species. To characterize the function of Pex17p, Chan et al. (2016) isolated Protein-A tagged peroxisomal receptor-docking complexes from wild-type and pex17 deletion strains followed by proteomic analysis of the complex composition. The analysis of this Pex14p complex revealed its composition, which included Pex14p, Pex17p as well as the dynein light chain protein Dyn2p as a core component of the complex. Pex17p and Dyn2p eluted with Pex14p in stoichiometric amounts. Previous studies had already implicated a role of Dyn2p in peroxisome biogenesis. First, yeast dyn2 deletion cells displayed a reduced capacity to utilize oleic acid as sole carbon source indicative of defect in peroxisome function (Smith et al. 2006). Moreover, in Yarrowia lipolytica, Dyn2p was found to interact with the peroxisomal docking complex, particularly with Pex17p, and it turned out to be required for efficient peroxisomal protein import (Chang et al. 2013). The study by Chan et al. (2016) demonstrated that Dyn2p is associated with Pex14p and in a Pex17p-dependent manner. In particular, liquid chromatography-mass spectrometry (LC–MS) based protein profiling experiments indicated that Dyn2p fails to associate with Pex14p in the absence of Pex17p. Further, chemical cross-linking experiments combined with MS (XL-MS) revealed the binding interface of Dyn2p and Pex14p but also showed that Dyn2p also interacts with Pex17p. A high-molecular weight Pex14p-subcomplex that is formed in wild-type cells was absent in either pex17 or dyn2 deletion yeast strains.
Pex17p is present in yeasts but it is absent in filamentous fungi and higher eukaryotes. Bioinformatic analysis of PEX genes in fungal genomes indicated the presence of a gene, which encodes a protein with significant similarity to both Pex14p and 17p (Kiel et al. 2006). The N-terminal region of this protein exhibits sequence similarity with the highly conserved N-terminal domain of Pex14p, while the C-terminus shows a weak similarity to yeast Pex17p. Managadze et al. (2010) performed proteomic analysis of the docking complex components of filamentous fungus N. crassa. In this study, hexa-histidine tagged PEX14 (His6-PEX14) was affinity-purified from digitonin-solubilized membranes, followed by further purification by using size-exclusion chromatography. The two-step purified PEX14 complex was then analyzed by liquid chromatography coupled to tandem mass spectrometry (LC/MS/MS), which revealed a novel component of the docking complex designated as PEX33. This protein shows sequence similarity with conserved PEX14 but only in short region of its N-terminal domain. PEX33 was shown to localize to glyoxysomes and to interact with PEX14, with the PTS1-receptor PEX5 as well as with itself. Biogenesis of glyoxysomes and Woronin bodies was defective in the single pex14 or pex33 deletion strains. The study showed that PEX33 plays an essential role in the import of PTS1 proteins and certain non-PTS proteins and the function of PEX33 is non-redundant with PEX14 (Managadze et al. 2010).
2.3 Peroxisomal Import Pores
Unlike mitochondria and chloroplast, peroxisomes can import folded, cofactor-bound or oligomerized proteins. How the transport of proteins across the peroxisomal membrane is achieved without affecting the permeability barrier was unknown for long time. The “transient import pore model” proposed that the cargo-loaded cycling import receptor associates with the peroxisomal membrane to form a translocation pore, and disassembles after the cargo is translocated across the membrane (Erdmann and Schliebs 2005). Transient assembly of the translocon on demand might explain how folded protein import occurs without damaging the integrity of peroxisomes. In this context, a minimal functional importomer was described for the peroxisomal import of Pex8p that only required the presence of the import receptors and Pex14p (Ma et al. 2009). To disclose the identity of the import pore, membrane-bound complexes of the import receptor Pex5p were isolated and tested for channel properties (Meinecke et al. 2010). To this end, higher molecular weight complexes containing Pex5p, PTS1 cargo protein, docking complex components and associated proteins were reconstituted into liposomes and electrophysiological characterization was performed using the planar lipid bilayer technique. The study revealed the ion-channel activity of complexes, which contain Pex5p and Pex14p as the principal components. The water-filled import pore can open up to 9 nm in a cargo-dependent manner, demonstrating the dynamic nature of the protein-conducting channel, which allows the translocation of the PTS1-cargo proteins across the membrane into the peroxisomal lumen.
The PTS1- and PTS2-dependent peroxisomal protein import pathways converge at Pex14p, which is also part of the PTS1 import pore, raising the question whether PTS2-proteins enter peroxisomes via same or distinct import pores. Pex7p is the receptor for PTS2 proteins but requires co-receptors for their peroxisomal targeting. Di-aromatic pentapeptide (WxxxF/Y) motifs in Pex5p that mediate the binding to Pex14p are also found in the N-terminal region of the PTS2 co-receptors (Schliebs and Kunau 2006). Pex20p, the PTS2 co-receptor in Pichia pastoris interacts with Pex14p via one of its WxxxF/Y motif, shown to be involved in the docking of PTS2 complex at peroxisomes (Léon et al. 2006). For the identification of the PTS2 import pores, membrane bound complexes of the PTS2 co-receptor Pex18p were reconstituted into liposomes, and tested for their channel properties using the single-channel planar lipid bilayer technique (Montilla-Martinez et al. 2015). The reconstituted complex, which contained Pex18p and the Pex14p/Pex17p heteromer as main constituents, showed pore-forming activity. The size of this pore (~4.7 nm) was large enough to accommodate import of folded PTS2-proteins. This demonstrated that PTS1- and PTS2-proteins are imported by distinct pores in yeast and that these pores can function independently. Whether PTS1- and PTS2-proteins are imported via distinct pores also in higher eukaryotes still needs to be investigated, as PEX5 is involved in both cases. PEX5 promotes import of proteins either as short isoform (PEX5S), which imports PTS1-proteins, or as a longer isoform (PEX5L), which in addition to its role in targeting PTS1 proteins also is required for the topogenesis of PTS2 proteins. In particular, PEX5L harbors a binding site for the PTS2-receptor PEX7 and thus functions as a co-receptor for its peroxisomal targeting.
2.4 Cargo Release into the Peroxisomal Matrix
Although PTS1 and PTS2 import pores have been identified, it is still poorly understood how the cargo is translocated into the peroxisomal lumen. Evidence has been provided that Pex8p may play a role in cargo release in yeast (Wang et al. 2003). Pex8p is tightly associated with lumenal side of the peroxisome membrane and known to bind Pex5p (Rehling et al. 2000). Pex8p contains both PTS1 and PTS2 signals for peroxisomal targeting (Rehling et al. 2000; Zhang et al. 2006). Fluorescence Correlation Spectroscopy (FCS) based in vitro studies showed that Pex8p can dissociate PTS1 receptor/cargo complex, suggesting that Pex8p plays a similar role in vivo (Wang et al. 2003). Pex8p is present in all fungi but has not been identified in higher eukaryotes, implicating a different mechanism of cargo release. Interestingly, trypanosomatid parasite PEX13.1 contains a C-terminal PTS1-like signal TKL similar to Pex8p. The role of this putative PTS1 in cargo release into glycosomal matrix awaits investigation (Verplaetse et al. 2009). For mammals, a role in cargo release has been attributed to Pex14p (Freitas et al. 2011). Here, the interaction of the receptor PEX5 with cargo protein catalase is disrupted by PEX14, which might indicate that cargo release may take place early in the receptor cycle after docking of PEX5 to the N-terminal domain of PEX14. In P. pastoris, Pex5p dimerization and oligomerization with Pex8p controls binding and release of the cargo (Ma et al. 2013). Particularly, the redox sensitive amino acid cysteine 10 in PpPex5p plays a role in the regulation of cargo binding by Pex5p, cargo release and receptor recycling.
Some proteins are proteolytically processed during or upon release into the peroxisomal matrix in higher eukaryotes. Tysnd1 (trypsin domain containing 1) is an intraperoxisomal protease of mammalian cells, which cleaves the signal peptide from PTS2 precursor proteins and also processes subset PTS1 proteins (Kurochkin et al. 2007). Tysnd1 regulates the peroxisomal fatty acid β-oxidation pathway and the proteolytic activity of oligomeric Tysnd1 is controlled by self-cleavage. The cleaved products are degraded by peroxisomal Lon protease (PsLon) (Okumoto et al. 2011). In the early 1990s, presence of endopeptidase activity and a leucine-aminopeptidase in plant peroxisomal matrix was reported (Corpas et al. 1993). Seven endoprotease isoenzymes have been detected in plant peroxisomes (Distefano et al. 1997). A role for plant peroxisomal proteases during senescence, oxidative stress conditions, metal pollutant exposure has been suggested (Distefano et al. 1999; reviewed in Palma et al. 2002). Like mammalian Tysnd1, Arabidopsis DEG15 serine protease is involved in cleavage of the PTS2 signal from precursor proteins (Helm et al. 2007).
2.5 Receptor Recycling
2.5.1 Exportomer
Upon or after cargo translocation into the peroxisomal matrix, the import receptors are recycled back to the cytosol to facilitate further rounds of peroxisomal protein import. This receptor export machinery involves ubiquitinating enzymes and ATP-dependent extraction of receptors from the peroxisomal membrane. The exportomer comprises ubiquitin conjugating (E2) enzymes (Pex4p in yeasts, plants and trypanosomatids; Ubc4p in yeasts and UbcH5-family in mammals), docking proteins at the peroxisomal membrane (Pex22p), corresponding ubiquitin ligases (E3) (Pex2p, Pex10p, Pex12p) and the AAA-complex (Pex1p and Pex6p) as well as its membrane anchor (Pex15p in fungi, PEX26 in mammals, APEM9 in plants) (reviewed in Platta et al. 2014).
2.5.2 Receptor Ubiquitination
The import receptor Pex5p and the PTS2 co-receptor Pex18p is mono- or poly-ubiquitinated at its N-terminal region (Kragt et al. 2005; Hensel et al. 2011). Mono-ubiquitination of Pex5p involves the attachment of a single ubiquitin via a thioester linkage to a conserved cysteine reside, while poly-ubiquitination occurs at conserved lysine residues in the N-terminal region. When the receptor is non-functional, poly-ubiquitination is supposed to trigger its export and proteasomal degradation, serving as a quality control mechanism. On the other hand, mono-ubiquitination signals export of the receptor for further rounds of peroxisomal protein import. The activation of the ubiquitin and the export of ubiquitinated receptors are the only ATP-dependent steps in peroxisomal matrix protein import.
-
Mono-ubiquitination
The receptor ubiquitination machinery is well characterized. Pex4p was the first E2 enzyme identified to be essential for the peroxisome biogenesis and involved in both PTS1 and PTS2 import (Wiebel and Kunau 1992; van der Klei et al. 1998). Pex4p is anchored to the peroxisomal membrane by the integral membrane protein Pex22p (Koller et al. 1999). Association of Pex4p to the cytosolic soluble domain of Pex22p enables assembly of functional E2-complex (Williams et al. 2012). Pex4p and Pex22p are well conserved in yeast, plants and trypanosomatids where they mediate mono-ubiquitination of Pex5p. However, Pex4p and Pex22p orthologues apparently do not exist in mammals and their function is fulfilled by other proteins.
To identify mammalian PEX5 ubiquitin conjugating enzyme (E2) i.e. counterpart of yeast Pex4p, a proteomic approach was used (Grou et al. 2008). Authors utilized an in vitro system comprising highly purified rat liver peroxisomes, recombinant E1, E2s and GST-Ubiquitin. The substrate for ubiquitination in this assay is a truncated PEX5 (1-324), containing single conserved cysteine, which is synthesized in vitro and radio-labelled to monitor ubiquitinated PEX5 by autoradiography. Supplementation of cytosol to the import reaction with purified peroxisomes resulted in appearance of thiol-sensitive ubiquitinated PEX5, indicating the presence of an E2 in the cytosol that ubiquitinates the cysteine of PEX5. The authors then fractionated the cytosol guided by peroxisome-dependent PEX5-ubiquitination activity to yield a fraction, which migrated as two protein bands in SDS-PAGE analysis. Mass spectrometry analysis of these bands identified E2-enzymes UbcH7, UbcH13 and three closely related UbcH5a-c. Using recombinant E2 enzymes, only the UbcH5 family members were found to mono-ubiquitinate PEX5 (Grou et al. 2008). Therefore, UbcH5 proteins are functional counterparts of yeast/plant Pex4p. But unlike peroxisome-specific Pex4p, the functions of the mammalian UbcH5 proteins are not restricted to peroxisomes as they are also involved in the ubiquitination of cellular targets that are evidently not related to peroxisomes (Brzovic and Klevit 2006).
The ubiquitination of Pex5p requires RING-domain containing ubiquitin-protein ligases (E3). Three conserved peroxins, Pex2p, Pex10p and Pex12p, are ubiquitin ligases that associate at the peroxisomal membrane to form the RING-finger complex (Platta et al. 2009). Pex12p facilitates Pex4p-dependent mono-ubiquitination of Pex5p, which may also involve Pex10p (Platta et al. 2009; El Magraoui et al. 2012). Like Pex5p, also the PTS2 co-receptors Pex18p (S. cerevisiae) and Pex20p (P. pastoris) are mono-ubiquitinated at a conserved cysteine residue (Hensel et al. 2011; Léon et al. 2006). Pex18p mono-ubiquitination is mediated by Pex4p as responsible E2 enzyme and the E3 ligases Pex10p and Pex12p (El Magraoui et al. 2013). Ubiquitin receptor proteins DSK2a and DSK2b interact with PEX12 in plants, although the role in receptor recycling is not clear (Kaur et al. 2013).
-
Poly-ubiquitination
Poly-ubiquitination of the PTS1-receptor serves as a quality control mechanism when the receptor is non-functional or when the normal mono-ubiquitination dependent recycling is defective (Platta et al. 2004, Kiel et al. 2005). This pathway has also been named “RADAR” pathway, standing for Receptor Accumulation and Degradation in Absence of Recycling (Léon and Subramani 2007). Here, Pex5p is poly-ubiquitinated by the attachment of K48-linked poly-ubiquitin chains to two conserved lysine residues in Pex5p, which primes the receptor for proteasomal degradation. E2 ubiquitin conjugating Ubc4p and E3 Ubiquitin ligases Pex10p and Pex2p are involved in the poly-ubiquitination of yeast Pex5 (Platta et al. 2004; Williams et al. 2008; Platta et al. 2009). PTS2 co-receptor Pex18p is also poly-ubiquitinated by Ubc4 and E3 enzymes Pex2p and Pex10p, leading to rapid turnover by proteasomal degradation (Hensel et al. 2011; El Magraoui et al. 2013). However, poly-ubiquitination of P. pastoris PTS2 co-receptor Pex20p is dependent on Pex4p and all three members of the RING-complex (Liu and Subramani 2013). In plants, Pex2p also interacts with ubiquitin-adapter DSK2 proteins (Kaur et al. 2013).
2.5.3 Membrane Dislocation of the Receptors
Pex1p and Pex6p are AAA-ATPases which function as dislocase for the export of Pex5p back to the cytosol (Platta et al. 2005). Pex1p and Pex6p form a heterohexameric AAA-complex (Ciniawsky et al. 2015), which is anchored to the peroxisomal membrane by Pex15p (yeast), PEX26 (human) or APEM9 (plants). Genetic mutations in PEX1, PEX6 or PEX26 abrogates peroxisomal protein import, which is the most common cause of Peroxisome Biogenesis Disorders (PBDs) in humans. Pex15p/PEX26 proteins show very low sequence conservation. To investigate the role of a putative PEX26 in Neurospora crassa, PEX26-complexes were isolated, and their composition was analyzed by mass-spectroscopy, which identified PEX6 and PEX1 as the prominent binding partners (Liu et al. 2011). This revealed that despite the low degree of sequence conservation, the Nc PEX26 is an orthologue of PEX26.
2.5.4 Receptor Deubiquitination
After or during cargo translocation, the PTS1 receptor is deubiquitinated. For mammalian cells, it was shown that the mono-ubiquitin, which is linked to the conserved cysteine via thioester-linkage, can be removed non-enzymatically by glutathione (GSH) in vitro (Grou et al. 2009). GSH also is a major nucleophile in the cytosol of mammalian cells, indicating that the non-enzymatic cleavage might also occur in vivo. However, numerous de-ubiquitinating enzymes (DUBs) are predicted in yeast and human genomes (Hutchins et al. 2013). DUBs acting on peroxisomal import receptors have been investigated in yeast (Debelyy et al. 2011) and mammalian cells (Grou et al. 2012).
In S. cerevisiae, the purified cytosolic complex of Pex1p-Pex6p (AAA-complex) was found to de-ubiquitinate membrane-bound monoUb-Pex5p (Debelyy et al. 2011). To identify the unknown associated factor or hydrolase, the authors isolated cytosolic AAA-complexes from yeast cells overexpressing His6-Pex6p (Debelyy et al. 2011). Analysis of the complex revealed the presence of two dominant proteins, which were identified by mass spectrometry as Ubiquitin hydrolase Ubp15 and ubiquitin-adapter protein Ecm21p. In a complementary approach, mass-spectrometric analysis of isolated genomically tagged Pex1-complexes also revealed their association with Ubp15p. Finally, affinity purified cytosolic complexes of His6-GST-Ubp15p also contained Pex1p and Pex6p. These studies identified Ubp15p as a novel component of the AAA-complex, which localizes to peroxisomes and is capable to cleave ubiquitin moieties from Pex5p. Accordingly, Ubp15p deficient yeast cells display a hydrogen peroxide stress-dependent PTS1 import defect (Debelyy et al. 2011).
In mammalian cells, biochemical studies indicate that mono-ubiquitinated PEX5 is the substrate of ubiquitin-specific protease 9X (USP9x) (Grou et al. 2012). However, USP9x function is not restricted to peroxisomal protein import as it also has other cellular targets. Apart from yeast Ubp15p or mammalian USP9x, additional or redundant ubiquitin hydrolases may also be involved in Pex5p de-ubiquitination.
2.5.5 Connecting Docking and Export Complexes
Beside its role in cargo release (see Sect. 2.4), S. cerevisiae Pex8p plays a major role in connecting the docking complex or import pore with the export complex consisting of the receptor ubiquitination and export machinery (Agne et al. 2003). Proteomic approaches were applied to define the components of two purified complexes of the peroxisomal import machinery, the docking complex, isolated by affinity chromatography via genomically tagged Pex14p, and the RING finger complex isolated in a similar way via genomically tagged Pex2p (Agne et al. 2003). Mass-spectrometry analysis of isolated complexes revealed that Pex14p and Pex2p bind same set of peroxins. Proteomic analysis of the Pex8p complex revealed the presence of the docking as well as the export complex. However, In the absence of Pex8p the docking complex and the export complex could not be co-isolated. In the absence of individual RING-finger peroxins, Pex8p still associates with the docking complex. Taken together, the data indicate that the association of the docking complex and the export complex to form a larger complex requires the presence of the intra-peroxisomal Pex8p (Agne et al. 2003). However, in P. pastoris, this networking function is apparently performed by Pex3p (Hazra et al. 2002).
2.6 Peroxisomal Importomer—Stable Versus Transient Interactors
Peroxisomal importomer consists of the docking complex (Pex14p, Pex17p, Pex13p) linked to the RING finger complex (Pex2p, Pex10p, Pex12p) via intraperoxisomal Pex8p. A comprehensive analysis of the stable and transient interaction partners of Pex14p (the central component of importomer) was performed by Oeljeklaus et al. (2012). The quantitative proteomic study involved epitope tagged Pex14p combined with dual-track stable isotope labeling with amino acid in culture-mass spectrometry (SILAC-MS) analysis of the affinity-isolated Pex14p complexes. The study identified 9 core components and additional 12 transient components of the Pex14p interactome. Pex8p, Pex11p and Dyn2p were part of the core complex in addition to the known docking and RING complex proteins. The study also identified Pex25p, Hrr25p, Esl2p and prohibitin as novel transient interaction partners (Oeljeklaus et al. 2012).
3 Peroxisomal Membrane Protein Import
In contrast to the large set of peroxins required for matrix protein import, peroxisomal membrane protein (PMP) import involves only three peroxins, Pex19p, Pex3p and Pex16p. Defects in any one of these result in complete loss of peroxisomes, mislocalisation of matrix proteins to the cytosol and membrane protein mislocalisation to other cellular membranes. Genetic defects in these peroxins also causes severe peroxisome biogenesis disorders in humans.
Pex19p is the cytosolic receptor for peroxisomal targeting of newly synthesized PMPs and possesses chaperone activity to stabilize the PMPs in the cytosol (Jones et al. 2004). Most PMPs contain membrane peroxisome targeting signal (mPTS) comprised of a short α-helical segment with positively charged and hydrophobic residues, and additionally at least one transmembrane segment (Jones et al. 2001; Honsho and Fujiki 2001; Rottensteiner et al. 2004). The N-terminal domain of Pex19p contains binding sites for Pex3p and/or Pex14p, whereas mPTS-PMP binding site is located in the C-terminal domain of Pex19p (Sato et al. 2010; Neufeld et al. 2009). Pex19p proteins contain a C-terminal CaaX motif for farnesylation. This modification of Pex19p is important for its structural integrity and PMP recognition (Rucktäschel et al. 2009; Emmanouilidis et al. 2017). Interestingly, Trypanosomatid PEX19 proteins lack such a C-terminal CaaX motif.
Docking of the Pex19p-PMP receptor-cargo complex at the peroxisomal membrane is mediated by the integral membrane protein Pex3p (Fang et al. 2004). The transmembrane domain at the N-terminus of Pex3p anchors it to peroxisomal membrane, while its soluble domain facing the cytosol binds to Pex19p. To gain further insight into the Pex3p-Pex19p interaction, Hattula et al. (2014) used hydrogen exchange mass spectrometry (HXMS) to monitor conformational changes during Pex3p-Pex19p complex formation in vitro. The study disclosed that although Pex19p remains very flexible during interaction with Pex3p, the N- and C-terminus along with a short stretch in the middle (F64-L74) of Pex19p are shielded from hydrogen exchange upon interaction with Pex3p. The authors suggest that Pex19p stabilizes Pex3p within the cell by preventing its aggregation.
In mammalian cells, PEX3 can directly sort to peroxisomes, which is PEX19- as well as PEX16- dependent and PEX16 functions as membrane receptor for the PEX3-PEX19 complexes (Matsuzaki and Fujiki 2008). Colasante et al. (2013) performed proteomic analysis of purified glycosomal membranes from Leishmania parasites, which identified 464 proteins in the glycosomal membrane fraction including known PMPs and a protein with weak homology to PEX16. Further studies confirmed the putative protein as trypanosomal PEX16 involved in glycosome biogenesis in Trypanosoma parasites (Kalel et al. 2015). Although PEX16 is present in higher eukaryotes, plants, fungi and Trypanosoma, it has not yet been identified in S. cerevisiae. However, recently a novel peroxin (Pex36p) was identified in Komagataella phaffii (P. pastoris) (Farre et al. 2017). Remarkably, the growth defect observed in cells lacking Pex36p could be restored by human PEX16 or yeast Pex34p, opening the possibility of an analogous or orthologous relationship of the three proteins. Interestingly, so far PEX3 has not been identified in trypanosomatid parasites.
Membrane biogenesis peroxins Pex19p, Pex3p and Pex16p are also involved in de novo biogenesis of peroxisomes from the endoplasmic reticulum (ER) (reviewed in Agrawal and Subramani 2016). Contribution of the ER to de novo peroxisome formation was proposed based on the observation that in S. cerevisiae pex3 deletion cells, re-introduced YFP-tagged Pex3p first sorts to ER, concentrates as foci, and results in a subsequent Pex19p-dependent budding and maturation of fully functional peroxisomes (Hoepfner et al. 2005). However, recent studies show that also in absence of Pex3p, PMPs accumulate in pre-peroxisomal vesicles, which are morphologically distinct from the ER (Knoops et al. 2014). These vesicles contained Pex14p and to investigate the composition of these pre-peroxisomal structures, Wroblewska et al. (2017) performed the proteomic analysis of affinity-purified Pex14p complexes from pex3 deletion yeast cells. These complexes contained Pex14p and docking components Pex13p, Pex17p as well as import receptors Pex5p, Pex7p and Pex4p/22p. However, the absence of Pex8p and the RING finger peroxins in this complex indicates that a functional importomer has not yet been formed in these prestructures. The study suggests that some PMPs are targeted to the pre-peroxisomal vesicles in a Pex3p-independent manner (Wroblewska et al. 2017).
4 Regulation of Peroxisomal Import Machineries
Reversible protein phosphorylation mediated by protein kinases and phosphatases is a common post-translational modification, regulating diverse cellular processes. Phosphorylation of the mitochondrial preprotein translocase proteins by cytosolic kinases regulate the mitochondrial protein import (Schmidt et al. 2011). There is increasing evidence that protein phosphorylation also plays an important role in peroxisome dynamics. Phosphorylation close to the PTS2-signal in Gpd1p is required for its dynamic localization to the peroxisomes (Jung et al. 2010). Analysis of the public protein databases demonstrates that numerous yeast and mammalian peroxisome biogenesis proteins are phosphorylated in vivo (Oeljeklaus et al. 2016). Currently, the exact role of these modifications in regulating the peroxisomal protein import is still lacking.
Recently, a strategy for a targeted analysis of peroxisomal phospho-proteome by affinity purification of epitope-tagged peroxisomal proteins using Phos-tag SDS-PAGE and high-resolution mass spectrometry has been described. Additionally, a protocol for MS-based in vitro kinase assay with recombinant peroxisomal proteins and specific kinase has been described, which allows the identification and localization of phospho-sites in peroxisomal proteins in vivo and identification of specific substrate-kinase relationships (Schummer et al. 2017). These emerging proteomics-based technologies will contribute to a better understanding of the regulation of peroxisomal protein import and of the resulting molecular dynamics of these fascinating multi-purpose organelles.
Abbreviations
- AAA:
-
ATPase Associated with diverse cellular Activities
- DUBs:
-
De-ubiquitinating enzymes
- mPTS:
-
membrane Peroxisome Targeting Signal
- PBDs:
-
Peroxisome Biogenesis Disorders
- PEX:
-
Peroxin
- PMPs:
-
Peroxisomal Membrane Proteins
- PTS:
-
Peroxisome Targeting Signal
- RING:
-
Really Interesting New Gene
- SH3:
-
Src Homology 3
- TEV:
-
Tobacco Etch Virus protease
- TPA:
-
TEV cleavage site—protein A
- TPR:
-
Tetratrico-Peptide Repeat
References
Agne B, Meindl NM, Niederhoff K, Einwachter H, Rehling P, Sickmann A, Meyer HE, Girzalsky W, Kunau WH (2003) Pex8p: an intraperoxisomal organizer of the peroxisomal import machinery. Mol Cell 11:635–646
Agrawal G, Subramani S (2016) De novo peroxisome biogenesis: evolving concepts and conundrums. Biochim Biophys Acta 1863:892–901
Albertini M, Rehling P, Erdmann R, Girzalsky W, Kiel JA, Veenhuis M, Kunau WH (1997) Pex14p, a peroxisomal membrane protein binding both receptors of the two PTS-dependent import pathways. Cell 89:83–92
Anderson RM, Bitterman KJ, Wood JG, Medvedik O, Sinclair DA (2003) Nicotinamide and PNC1 govern lifespan extension by calorie restriction in Saccharomyces cerevisiae. Nature 423:181–185
Bottger G, Barnett P, Klein AT, Kragt A, Tabak HF, Distel B (2000) Saccharomyces cerevisiae PTS1 receptor Pex5p interacts with the SH3 domain of the peroxisomal membrane protein Pex13p in an unconventional, non-PXXP-related manner. Mol Biol Cell 11:3963–3976
Brennand A, Rigden DJ, Michels PA (2012) Trypanosomes contain two highly different isoforms of peroxin PEX13 involved in glycosome biogenesis. FEBS Lett 586:1765–1771
Brocard C, Hartig A (2006) Peroxisome targeting signal 1: is it really a simple tripeptide? Biochim Biophys Acta 1763:1565–1573
Brzovic PS, Klevit RE (2006) Ubiquitin transfer from the E2 perspective: why is UbcH5 so promiscuous? Cell Cycle 5:2867–2873
Chan A, Schummer A, Fischer S, Schröter T, Cruz-Zaragoza LD, Bender J, Drepper F, Oeljeklaus S, Kunau WH, Girzalsky W, Warscheid B, Erdmann R (2016) Pex17p-dependent assembly of Pex14p/Dyn2p-subcomplexes of the peroxisomal protein import machinery. Eur J Cell Biol 95:585–597
Chang J, Tower RJ, Lancaster DL, Rachubinski RA (2013) Dynein light chain interaction with the peroxisomal import docking complex modulates peroxisome biogenesis in yeast. J Cell Sci 126:4698–4706
Ciniawsky S, Grimm I, Saffian D, Girzalsky W, Erdmann R, Wendler P (2015) Molecular snapshots of the Pex1/6 AAA+ complex in action. Nat Commun 6:7331
Colasante C, Voncken F, Manful T, Ruppert T, Tielens AGM, van Hellemond JJ, Clayton C (2013) Proteins and lipids of glycosomal membranes from Leishmania tarentolae and Trypanosoma brucei. F1000Res 2:27
Corpas FJ, Palma JM, del Río LA (1993) Evidence for the presence of proteolytic activity in peroxisomes. Eur J Cell Biol 61:81–85
Dawidowski M, Emmanouilidis L, Kalel VC, Tripsianes K, Schorpp K, Hadian K, Kaiser M, Mäser P, Kolonko M, Tanghe S, Rodriguez A, Schliebs W, Erdmann R, Sattler M, Popowicz GM (2017) Inhibitors of PEX14 disrupt protein import into glycosomes and kill Trypanosoma parasites. Science 355:1416–1420
Debelyy MO, Platta HW, Saffian D, Hensel A, Thoms S, Meyer HE, Warscheid B, Girzalsky W, Erdmann R (2011) Ubp15p, a ubiquitin hydrolase associated with the peroxisomal export machinery. J Biol Chem 286:28223–28234
Distefano S, Palma JM, Gómez M, del Río LA (1997) Characterization of endoproteases from plant peroxisomes. Biochem J 327:399–405
Distefano S, Palma JM, McCarthy I, del Río LA (1999) Proteolytic cleavage of plant proteins by peroxisomal endoproteases from senescent pea leaves. Planta 209:308–313
Douangamath A, Filipp FV, Klein AT, Barnett P, Zou P, Voorn-Brouwer T, Vega MC, Mayans OM, Sattler M, Distel B, Wilmanns M (2002) Topography for independent binding of alpha-helical and PPII-helical ligands to a peroxisomal SH3 domain. Mol Cell 10:1007–1017
Effelsberg D, Cruz-Zaragoza LD, Tonillo J, Schliebs W, Erdmann R (2015) Role of Pex21p for piggyback import of Gpd1p and Pnc1p into peroxisomes of Saccharomyces cerevisiae. J Biol Chem 290:25333–25342
El Magraoui F, Baumer BE, Platta HW, Baumann JS, Girzalsky W, Erdmann R (2012) The RING-type ubiquitin ligases Pex2p, Pex10p and Pex12p form a heteromeric complex that displays enhanced activity in an ubiquitin conjugating enzyme-selective manner. FEBS J 279:2060–2070
El Magraoui F, Brinkmeier R, Schrotter A, Girzalsky W, Muller T, Marcus K, Meyer HE, Erdmann R, Platta HW (2013) Distinct ubiquitination cascades act on the peroxisomal targeting signal type 2 co-receptor Pex18p. Traffic 14:1290–1301
Emmanouilidis L, Schutz U, Tripsianes K, Madl T, Radke J, Rucktaschel R, Wilmanns M, Schliebs W, Erdmann R, Sattler M (2017) Allosteric modulation of peroxisomal membrane protein recognition by farnesylation of the peroxisomal import receptor PEX19. Nat Commun 8:14635
Erdmann R, Blobel G (1995) Giant peroxisomes in oleic acid-induced Saccharomyces cerevisiae lacking the peroxisomal membrane protein Pmp27p. J Cell Biol 128:509–523
Erdmann R, Schliebs W (2005) Peroxisomal matrix protein import: the transient pore model. Nat Rev Mol Cell Biol 6:738–742
Erdmann R, Veenhuis M, Mertens D, Kunau WH (1989) Isolation of peroxisome-deficient mutants of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 86:5419–5423
Fang Y, Morrell JC, Jones JM, Gould SJ (2004) PEX3 functions as a PEX19 docking factor in the import of class I peroxisomal membrane proteins. J Cell Biol 164:863–875
Farre JC, Carolino K, Stasyk OV, Stasyk OG, Hodzic Z, Agrawal G, Till A, Proietto M, Cregg J, Sibirny AA, Subramani S (2017) A new yeast peroxin, Pex36, a functional homolog of mammalian PEX16, functions in the ER-to-peroxisome traffic of peroxisomal membrane proteins. J Mol Biol 429:3743–3762
Flynn CR, Mullen RT, Trelease RN (1998) Mutational analyses of a type 2 peroxisomal targeting signal that is capable of directing oligomeric protein import into tobacco BY-2 glyoxysomes. Plant J 16:709–720
Fransen M, Vastiau I, Brees C, Brys V, Mannaerts GP, Van Veldhoven PP (2004) Potential role for Pex19p in assembly of PTS-receptor docking complexes. J Biol Chem 279:12615–12624
Freitas MO, Francisco T, Rodrigues TA, Alencastre IS, Pinto MP, Grou CP, Carvalho AF, Fransen M, Sa-Miranda C, Azevedo JE (2011) PEX5 protein binds monomeric catalase blocking its tetramerization and releases it upon binding the N-terminal domain of PEX14. J Biol Chem 286:40509–40519
Girzalsky W, Rehling P, Stein K, Kipper J, Blank L, Kunau WH, Erdmann R (1999) Involvement of Pex13p in Pex14p localization and peroxisomal targeting signal 2-dependent protein import into peroxisomes. J Cell Biol 144:1151–1162
Glover JR, Andrews DW, Rachubinski RA (1994) Saccharomyces cerevisiae peroxisomal thiolase is imported as a dimer. Proc Natl Acad Sci U S A 91:10541–10545
Gould SG, Keller GA, Subramani S (1987) Identification of a peroxisomal targeting signal at the carboxy terminus of firefly luciferase. J Cell Biol 105:2923–2931
Gould SJ, Kalish JE, Morrell JC, Bjorkman J, Urquhart AJ, Crane DI (1996) Pex13p is an SH3 protein of the peroxisome membrane and a docking factor for the predominantly cytoplasmic PTS1 receptor. J Cell Biol 135:85–95
Grou CP, Carvalho AF, Pinto MP, Wiese S, Piechura H, Meyer HE, Warscheid B, Sa-Miranda C, Azevedo JE (2008) Members of the E2D (UbcH5) family mediate the ubiquitination of the conserved cysteine of Pex5p, the peroxisomal import receptor. J Biol Chem 283:14190–14197
Grou CP, Carvalho AF, Pinto MP, Huybrechts SJ, Sa-Miranda C, Fransen M, Azevedo JE (2009) Properties of the ubiquitin-pex5p thiol ester conjugate. J Biol Chem 284:10504–10513
Grou CP, Francisco T, Rodrigues TA, Freitas MO, Pinto MP, Carvalho AF, Domingues P, Wood SA, Rodriguez-Borges JE, Sa-Miranda C, Fransen M, Azevedo JE (2012) Identification of ubiquitin-specific protease 9X (USP9X) as a deubiquitinase acting on ubiquitin-peroxin 5 (PEX5) thioester conjugate. J Biol Chem 287:12815–12827
Grunau S, Schliebs W, Linnepe R, Neufeld C, Cizmowski C, Reinartz B, Meyer HE, Warscheid B, Girzalsky W, Erdmann R (2009) Peroxisomal targeting of PTS2 pre-import complexes in the yeast Saccharomyces cerevisiae. Traffic 10:451–460
Hagen S, Drepper F, Fischer S, Fodor K, Passon D, Platta HW, Zenn M, Schliebs W, Girzalsky W, Wilmanns M, Warscheid B, Erdmann R (2015) Structural insights into cargo recognition by the yeast PTS1 receptor. J Biol Chem 290:26610–26626
Hattula K, Hirschberg D, Kalkkinen N, Butcher SJ, Ora A (2014) Association between the intrinsically disordered protein PEX19 and PEX3. PLoS ONE 9:e103101
Hazra PP, Suriapranata I, Snyder WB, Subramani S (2002) Peroxisome remnants in pex3delta cells and the requirement of Pex3p for interactions between the peroxisomal docking and translocation subcomplexes. Traffic 3:560–574
Helm M, Luck C, Prestele J, Hierl G, Huesgen PF, Frohlich T, Arnold GJ, Adamska I, Gorg A, Lottspeich F, Gietl C (2007) Dual specificities of the glyoxysomal/peroxisomal processing protease Deg15 in higher plants. Proc Natl Acad Sci U S A 104:11501–11506
Hensel A, Beck S, El Magraoui F, Platta HW, Girzalsky W, Erdmann R (2011) Cysteine-dependent ubiquitination of Pex18p is linked to cargo translocation across the peroxisomal membrane. J Biol Chem 286:43495–43505
Hoepfner D, Schildknegt D, Braakman I, Philippsen P, Tabak HF (2005) Contribution of the endoplasmic reticulum to peroxisome formation. Cell 122:85–95
Honsho M, Fujiki Y (2001) Topogenesis of peroxisomal membrane protein requires a short, positively charged intervening-loop sequence and flanking hydrophobic segments. Study using human membrane protein PMP34. J Biol Chem 276:9375–9382
Hutchins AP, Liu S, Diez D, Miranda-Saavedra D (2013) The repertoires of ubiquitinating and deubiquitinating enzymes in eukaryotic genomes. Mol Biol Evol 30:1172–1187
Islinger M, Li KW, Seitz J, Volkl A, Luers GH (2009) Hitchhiking of Cu/Zn superoxide dismutase to peroxisomes–evidence for a natural piggyback import mechanism in mammals. Traffic 10:1711–1721
Jones JM, Morrell JC, Gould SJ (2001) Multiple distinct targeting signals in integral peroxisomal membrane proteins. J Cell Biol 153:1141–1150
Jones JM, Morrell JC, Gould SJ (2004) PEX19 is a predominantly cytosolic chaperone and import receptor for class 1 peroxisomal membrane proteins. J Cell Biol 164:57–67
Jung S, Marelli M, Rachubinski RA, Goodlett DR, Aitchison JD (2010) Dynamic changes in the subcellular distribution of Gpd1p in response to cell stress. J Biol Chem 285:6739–6749
Kalel VC, Schliebs W, Erdmann R (2015) Identification and functional characterization of Trypanosoma brucei peroxin 16. Biochim Biophys Acta 1853:2326–2337
Kaur N, Zhao Q, Xie Q, Hu J (2013) Arabidopsis RING peroxins are E3 ubiquitin ligases that interact with two homologous ubiquitin receptor proteins(F). J Integr Plant Biol 55:108–120
Kiel JA, Emmrich K, Meyer HE, Kunau WH (2005) Ubiquitination of the peroxisomal targeting signal type 1 receptor, Pex5p, suggests the presence of a quality control mechanism during peroxisomal matrix protein import. J Biol Chem 280:1921–1930
Kiel JA, Veenhuis M, van der Klei IJ (2006) PEX genes in fungal genomes: common, rare or redundant. Traffic 7:1291–1303
Knoops K, Manivannan S, Cepinska MN, Krikken AM, Kram AM, Veenhuis M, van der Klei IJ (2014) Preperoxisomal vesicles can form in the absence of Pex3. J Cell Biol 204:659–668
Koller A, Snyder WB, Faber KN, Wenzel TJ, Rangell L, Keller GA, Subramani S (1999) Pex22p of Pichia pastoris, essential for peroxisomal matrix protein import, anchors the ubiquitin-conjugating enzyme, Pex4p, on the peroxisomal membrane. J Cell Biol 146:99–112
Kragt A, Voorn-Brouwer T, van den Berg M, Distel B (2005) The Saccharomyces cerevisiae peroxisomal import receptor Pex5p is monoubiquitinated in wild type cells. J Biol Chem 280:7867–7874
Kurochkin IV, Mizuno Y, Konagaya A, Sakaki Y, Schonbach C, Okazaki Y (2007) Novel peroxisomal protease Tysnd1 processes PTS1- and PTS2-containing enzymes involved in beta-oxidation of fatty acids. EMBO J 26:835–845
Lametschwandtner G, Brocard C, Fransen M, Van Veldhoven P, Berger J, Hartig A (1998) The difference in recognition of terminal tripeptides as peroxisomal targeting signal 1 between yeast and human is due to different affinities of their receptor Pex5p to the cognate signal and to residues adjacent to it. J Biol Chem 273:33635–33643
Léon S, Subramani S (2007) A conserved cysteine residue of Pichia pastoris Pex20p is essential for its recycling from the peroxisome to the cytosol. J Biol Chem 282:7424–7430
Léon S, Zhang L, McDonald WH, Yates J 3rd, Cregg JM, Subramani S (2006) Dynamics of the peroxisomal import cycle of PpPex20p: ubiquitin-dependent localization and regulation. J Cell Biol 172:67–78
Liu X, Subramani S (2013) Unique requirements for mono- and polyubiquitination of the peroxisomal targeting signal co-receptor, Pex20. J Biol Chem 288:7230–7240
Liu F, Lu Y, Pieuchot L, Dhavale T, Jedd G (2011) Import oligomers induce positive feedback to promote peroxisome differentiation and control organelle abundance. Dev Cell 21:457–468
Ma C, Schumann U, Rayapuram N, Subramani S (2009) The peroxisomal matrix import of Pex8p requires only PTS receptors and Pex14p. Mol Biol Cell 20:3680–3689
Ma C, Hagstrom D, Polley SG, Subramani S (2013) Redox-regulated cargo binding and release by the peroxisomal targeting signal receptor, Pex5. J Biol Chem 288:27220–27231
Managadze D, Wurtz C, Wiese S, Schneider M, Girzalsky W, Meyer HE, Erdmann R, Warscheid B, Rottensteiner H (2010) Identification of PEX33, a novel component of the peroxisomal docking complex in the filamentous fungus Neurospora crassa. Eur J Cell Biol 89:955–964
Marzioch M, Erdmann R, Veenhuis M, Kunau WH (1994) PAS7 encodes a novel yeast member of the WD-40 protein family essential for import of 3-oxoacyl-CoA thiolase, a PTS2-containing protein, into peroxisomes. EMBO J 13:4908–4918
Matsuzaki T, Fujiki Y (2008) The peroxisomal membrane protein import receptor Pex3p is directly transported to peroxisomes by a novel Pex19p- and Pex16p-dependent pathway. J Cell Biol 183:1275–1286
Meinecke M, Cizmowski C, Schliebs W, Kruger V, Beck S, Wagner R, Erdmann R (2010) The peroxisomal importomer constitutes a large and highly dynamic pore. Nat Cell Biol 12:273–277
Montilla-Martinez M, Beck S, Klumper J, Meinecke M, Schliebs W, Wagner R, Erdmann R (2015) Distinct pores for peroxisomal import of PTS1 and PTS2 proteins. Cell Rep 13:2126–2134
Neufeld C, Filipp FV, Simon B, Neuhaus A, Schuller N, David C, Kooshapur H, Madl T, Erdmann R, Schliebs W, Wilmanns M, Sattler M (2009) Structural basis for competitive interactions of Pex14 with the import receptors Pex5 and Pex19. EMBO J 28:745–754
Oeljeklaus S, Reinartz BS, Wolf J, Wiese S, Tonillo J, Podwojski K, Kuhlmann K, Stephan C, Meyer HE, Schliebs W, Brocard C, Erdmann R, Warscheid B (2012) Identification of core components and transient interactors of the peroxisomal importomer by dual-track stable isotope labeling with amino acids in cell culture analysis. J Proteome Res 11:2567–2580
Oeljeklaus S, Schummer A, Mastalski T, Platta HW, Warscheid B (2016) Regulation of peroxisome dynamics by phosphorylation. Biochim Biophys Acta 1863:1027–1037
Okumoto K, Kametani Y, Fujiki Y (2011) Two proteases, trypsin domain-containing 1 (Tysnd1) and peroxisomal lon protease (PsLon), cooperatively regulate fatty acid beta-oxidation in peroxisomal matrix. J Biol Chem 286:44367–44379
Palma JM, Sandalio LM, Corpas FJ, Romero-Puertas MC, McCarthy I, del Río LA (2002) Plant proteases, protein degradation and oxidative stress: role of peroxisomes. Plant Physiol Biochem 40:521–530
Pan D, Nakatsu T, Kato H (2013) Crystal structure of peroxisomal targeting signal-2 bound to its receptor complex Pex7p-Pex21p. Nat Struct Mol Biol 20:987–993
Petriv OI, Tang L, Titorenko VI, Rachubinski RA (2004) A new definition for the consensus sequence of the peroxisome targeting signal type 2. J Mol Biol 341:119–134
Pires JR, Hong X, Brockmann C, Volkmer-Engert R, Schneider-Mergener J, Oschkinat H, Erdmann R (2003) The ScPex13p SH3 domain exposes two distinct binding sites for Pex5p and Pex14p. J Mol Biol 326:1427–1435
Platta HW, Girzalsky W, Erdmann R (2004) Ubiquitination of the peroxisomal import receptor Pex5p. Biochem J 384:37–45
Platta HW, Grunau S, Rosenkranz K, Girzalsky W, Erdmann R (2005) Functional role of the AAA peroxins in dislocation of the cycling PTS1 receptor back to the cytosol. Nat Cell Biol 7:817–822
Platta HW, El Magraoui F, Baumer BE, Schlee D, Girzalsky W, Erdmann R (2009) Pex2 and pex12 function as protein-ubiquitin ligases in peroxisomal protein import. Mol Cell Biol 29:5505–5516
Platta HW, Hagen S, Reidick C, Erdmann R (2014) The peroxisomal receptor dislocation pathway: to the exportomer and beyond. Biochimie 98:16–28
Rehling P, Skaletz-Rorowski A, Girzalsky W, Voorn-Brouwer T, Franse MM, Distel B, Veenhuis M, Kunau WH, Erdmann R (2000) Pex8p, an intraperoxisomal peroxin of Saccharomyces cerevisiae required for protein transport into peroxisomes binds the PTS1 receptor pex5p. J Biol Chem 275:3593–3602
Reumann S (2004) Specification of the peroxisome targeting signals type 1 and type 2 of plant peroxisomes by bioinformatics analyses. Plant Physiol 135:783–800
Reumann S, Babujee L, Ma C, Wienkoop S, Siemsen T, Antonicelli GE, Rasche N, Luder F, Weckwerth W, Jahn O (2007) Proteome analysis of Arabidopsis leaf peroxisomes reveals novel targeting peptides, metabolic pathways, and defense mechanisms. Plant Cell 19:3170–3193
Rottensteiner H, Kramer A, Lorenzen S, Stein K, Landgraf C, Volkmer-Engert R, Erdmann R (2004) Peroxisomal membrane proteins contain common Pex19p-binding sites that are an integral part of their targeting signals. Mol Biol Cell 15:3406–3417
Rucktäschel R, Thoms S, Sidorovitch V, Halbach A, Pechlivanis M, Volkmer R, Alexandrov K, Kuhlmann J, Rottensteiner H, Erdmann R (2009) Farnesylation of pex19p is required for its structural integrity and function in peroxisome biogenesis. J Biol Chem 284:20885–20896
Sato Y, Shibata H, Nakatsu T, Nakano H, Kashiwayama Y, Imanaka T, Kato H (2010) Structural basis for docking of peroxisomal membrane protein carrier Pex19p onto its receptor Pex3p. EMBO J 29:4083–4093
Schäfer A, Kerssen D, Veenhuis M, Kunau WH, Schliebs W (2004) Functional similarity between the peroxisomal PTS2 receptor binding protein Pex18p and the N-terminal half of the PTS1 receptor Pex5p. Mol Cell Biol 24:8895–8906
Schliebs W, Kunau WH (2006) PTS2 co-receptors: diverse proteins with common features. Biochim Biophys Acta 1763:1605–1612
Schmidt O, Harbauer AB, Rao S, Eyrich B, Zahedi RP, Stojanovski D, Schonfisch B, Guiard B, Sickmann A, Pfanner N, Meisinger C (2011) Regulation of mitochondrial protein import by cytosolic kinases. Cell 144:227–239
Schummer A, Fischer S, Oeljeklaus S, Warscheid B (2017) Study of peroxisomal protein phosphorylation by functional proteomics. Methods Mol Biol 1595:267–289
Smith JJ, Sydorskyy Y, Marelli M, Hwang D, Bolouri H, Rachubinski RA, Aitchison JD (2006) Expression and functional profiling reveal distinct gene classes involved in fatty acid metabolism. Mol Syst Biol 2(2006):0009
Swinkels BW, Gould SJ, Bodnar AG, Rachubinski RA, Subramani S (1991) A novel, cleavable peroxisomal targeting signal at the amino-terminus of the rat 3-ketoacyl-CoA thiolase. EMBO J 10:3255–3262
Terlecky SR, Nuttley WM, McCollum D, Sock E, Subramani S (1995) The Pichia pastoris peroxisomal protein PAS8p is the receptor for the C-terminal tripeptide peroxisomal targeting signal. EMBO J 14:3627–3634
Tsukamoto T, Yokota S, Fujiki Y (1990) Isolation and characterization of Chinese hamster ovary cell mutants defective in assembly of peroxisomes. J Cell Biol 110:651–660
Urquhart AJ, Kennedy D, Gould SJ, Crane DI (2000) Interaction of Pex5p, the type 1 peroxisome targeting signal receptor, with the peroxisomal membrane proteins Pex14p and Pex13p. J Biol Chem 275:4127–4136
van der Klei IJ, Veenhuis M (2006) PTS1-independent sorting of peroxisomal matrix proteins by Pex5p. Biochim Biophys Acta 1763:1794–1800
van der Klei IJ, Hilbrands RE, Kiel JA, Rasmussen SW, Cregg JM, Veenhuis M (1998) The ubiquitin-conjugating enzyme Pex4p of Hansenula polymorpha is required for efficient functioning of the PTS1 import machinery. EMBO J 17:3608–3618
Verplaetse E, Rigden DJ, Michels PA (2009) Identification, characterization and essentiality of the unusual peroxin 13 from Trypanosoma brucei. Biochim Biophys Acta 1793:516–527
Wang D, Visser NV, Veenhuis M, van der Klei IJ (2003) Physical interactions of the peroxisomal targeting signal 1 receptor pex5p, studied by fluorescence correlation spectroscopy. J Biol Chem 278:43340–43345
Wiebel FF, Kunau WH (1992) The Pas2 protein essential for peroxisome biogenesis is related to ubiquitin-conjugating enzymes. Nature 359:73–76
Williams C, van den Berg M, Geers E, Distel B (2008) Pex10p functions as an E3 ligase for the Ubc4p-dependent ubiquitination of Pex5p. Biochem Biophys Res Commun 374:620–624
Williams C, van den Berg M, Panjikar S, Stanley WA, Distel B, Wilmanns M (2012) Insights into ubiquitin-conjugating enzyme/co-activator interactions from the structure of the Pex4p:Pex22p complex. EMBO J 31:391–402
Wroblewska JP, Cruz-Zaragoza LD, Yuan W, Schummer A, Chuartzman SG, de Boer R, Oeljeklaus S, Schuldiner M, Zalckvar E, Warscheid B, Erdmann R, van der Klei IJ (2017) Saccharomyces cerevisiae cells lacking Pex3 contain membrane vesicles that harbor a subset of peroxisomal membrane proteins. Biochim Biophys Acta 1864:1656–1667
Yang X, Purdue PE, Lazarow PB (2001) Eci1p uses a PTS1 to enter peroxisomes: either its own or that of a partner, Dci1p. Eur J Cell Biol 80:126–138
Zhang L, Leon S, Subramani S (2006) Two independent pathways traffic the intraperoxisomal peroxin PpPex8p into peroxisomes: mechanism and evolutionary implications. Mol Biol Cell 17:690–699
Acknowledgements
The work is supported by a FoRUM Grant (F883-2016, F913-2017) of the Ruhr-University Bochum and by the Deutsche Forschungsgemeinschaft (FOR1905). The authors apologize to all those in the community whose work could not be discussed due to space limitations.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Kalel, V.C., Erdmann, R. (2018). Unraveling of the Structure and Function of Peroxisomal Protein Import Machineries. In: del Río, L., Schrader, M. (eds) Proteomics of Peroxisomes. Subcellular Biochemistry, vol 89. Springer, Singapore. https://doi.org/10.1007/978-981-13-2233-4_13
Download citation
DOI: https://doi.org/10.1007/978-981-13-2233-4_13
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-2232-7
Online ISBN: 978-981-13-2233-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)