Abstract
Peroxisomes are ubiquitous cell organelles of eukaryotic cells. Depending on environmental changes and cellular demands, peroxisomes display a high plasticity in metabolic functions. A prerequisite to carry out their physiological tasks is compartmentalization of peroxisomal enzymes in the lumen of this organelle, the peroxisomal matrix. The matrix proteins are synthesized on free polyribosomes in the cytosol and harbor a peroxisomal targeting sequence (PTS). They are targeted to the peroxisomal membrane by soluble PTS-receptors. Following the release of the cargo enzyme into the peroxisomal matrix, the PTS-receptor is ubiquitinated and exported back to the cytosol to facilitate further rounds of matrix protein import. The retrotranslocation of the receptor is facilitated by a molecular machinery that comprises enzymes required for the ubiquitination as well as for the ATP-dependent extraction of the receptor from the membrane. Furthermore, recent evidence indicates that the export machinery of the receptors might function as molecular motor not only for the retrotranslocation of the receptors themselves but also for the import of peroxisomal matrix proteins. This is thought to be achieved by coupling the ATP-dependent removal of the PTS-receptor with the cargo protein translocation into the organelle. In this review, we will discuss the combined data on the architecture and molecular function of the peroxisomal receptor export machinery, the peroxisomal exportomer.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction to the Cellular Functions of Peroxisomes
Peroxisomes are single membrane-bound organelles that can be found in all eukaryotic cells with the exception of spermatozoa and mature erythrocytes (Novikoff et al. 1973). Peroxisomes display a high variability in their enzyme content and metabolic tasks that can be adjusted according to cellular needs. The enzymes in the lumen of peroxisomes are often highly concentrated and tightly packed to form crystalline inclusions that are visible as electron-dense structures. The beta-oxidation of fatty acids and the detoxification of the hydrogen peroxide are regarded as the central and most conserved functions of peroxisomes (Cooper and Beevers 1969; Lazarow and DeDuve 1976). Furthermore, the beta-oxidation pathway is linked to the synthesis of signaling molecules, like phytohormones in plants (Baker et al. 2006; Kienow et al. 2008) and pheromones in Caenorhabditis elegans and insects (Joo et al. 2010; Spiegel et al. 2011). Mammalian peroxisomes have a key function in the biosynthesis of ether lipids and bile acids (Wanders and Waterham 2006a). Peroxisomes house important steps of penicillin biosynthesis in some filamentous fungi (Meijer et al. 2010; Müller et al. 1991), but also certain enzymes required for the biosynthesis of Vitamin K1 in plants (Widhalm et al. 2012) or the synthesis of siderophores required for iron uptake and virulence of Aspergillus species (Gründlinger et al. 2013). Depending on the metabolic state of the cell, the number of peroxisomes can be dynamically regulated either by the proliferation of peroxisomes or their selective autophagic degradation via pexophagy (Grunau et al. 2011; Opaliński et al. 2011; Till et al. 2012; Tower et al. 2011).
Defects in peroxisome function are the molecular cause for human inborn errors that are caused by mutation of single metabolic enzymes (Wanders and Waterham 2006b) or genes coding for proteins that are required for the biogenesis of the organelles (Steinberg et al. 2006). The peroxisomal biogenesis disorders (PBDs) form a spectrum of autosomal recessive metabolic disorders that are collectively characterized by abnormal peroxisome assembly and result in multisystemic disorders that often lead to death in early infancy (Baes and Van Veldhoven 2012; Nagotu et al. 2012; Waterham and Ebberink 2012). Furthermore, the physiological function of peroxisomes contributes to the cellular protection mechanism against the progressive brain damage and cognitive decline caused by Alzheimer’s disease (Fanelli et al. 2013; Kou et al. 2011; Lizard et al. 2012).
The formation of peroxisomes depends on specific biogenesis factors, the peroxins (Distel et al. 1996). To date, 34 different peroxins have been described. In general, they are involved in the six key stages of peroxisomal biogenesis which comprise the (1) de novo formation and (2) proliferation of peroxisomes, (3) their inheritance and (4) regulated degradation by an authophagic process called pexophagy as well as the import of (5) peroxisomal membrane and (6) matrix proteins (Fagarasanu et al. 2010; Islinger et al. 2012; Liu et al. 2012; Platta and Erdmann 2007b; Theodoulou et al. 2013; Hasan et al. 2013).
In this review, we will discuss the peroxisomal matrix protein import with emphasis on the function of the peroxisomal membrane complexes that are involved in the ubiquitination and energy-consuming dislocation of the dynamic import receptors and describe their concerted function as receptor export machinery, the peroxisomal exportomer (Platta et al. 2013).
2 Peroxisomal Matrix Proteins Are Imported by Cycling Receptors
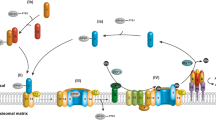
All peroxisomal proteins are encoded in the nucleus, synthesized on free ribosomes in the cytosol and imported posttranslationally. Most remarkably, peroxisomes are capable to accommodate fully folded proteins. Frequently, even oligomeric or cofactor-bound proteins are imported (Leon et al. 2006a; Girzalsky et al. 2009). The import of folded proteins distinguishes peroxisomes from other organelles like chloroplasts or mitochondria, which all import unfolded proteins, but it makes them comparable to the Tat (Twin-arginine translocation) pathways of bacteria and thylakoid membranes (Albiniak et al. 2012; Palmer and Berks 2012). However, in contrast to these translocation systems, peroxisomal matrix protein import is facilitated by dynamic receptors that cycle between a soluble state in the cytosol and a membrane-associated state at the peroxisomal membrane (Hasan et al. 2013; Liu et al. 2012; Platta and Erdmann 2007a). The import cycle can conceptually be divided into five steps, comprising (1) cargo recognition in the cytosol, (2) receptor–cargo docking at the peroxisome, (3) cargo translocation across the membrane, (4) cargo release into the matrix, and (5) receptor ubiquitination and export back to the cytosol (Fig. 15.1).
Peroxisomal PTS1-protein import in Saccharomyces cerevisiae. Peroxisomal matrix proteins are imported into peroxisomes via soluble receptors, which shuttle between the cytosol and the peroxisomal membrane. The matrix proteins are recognized by their peroxisomal targeting signal 1 (PTS1) in the cytosol via the receptor Pex5p, forming a receptor–cargo complex. At the peroxisomal membrane this complex binds to the docking complex (Pex13p, Pex14p and Pex17p), and this binding leads to the formation of a transient pore, whose exact molecular constitution is still under discussion but at least contains Pex5p and Pex14p. In the next step, the cargo is translocated into the peroxisomal lumen in an unknown manner; Pex8p might be involved in the receptor-cargo dissociation. At the end of the receptor cycle, the receptor is removed from the membrane and transported back to the cytosol for another round of import. To this end, Pex5p is monoubiquitinated by the ubiquitin-conjugating Pex4p (E2), which is anchored at the membrane by Pex22p, and by the ubiquitin ligase Pex12p (E3), which forms the RING-finger complex together with the other ubiquitin ligases Pex2p and Pex10p. The ubiquitin signal leads to an ATP-dependent dislocation of Pex5p from the peroxisomal membrane, performed by the Pex15p-anchored AAA peroxins Pex1p and Pex6p. Prior to a new round of import, the ubiquitin moiety is removed from the receptor
The events concerning the cargo transport from the cytosol to the peroxisomal lumen are discussed in detail in Chaps. 13 (Effelsberg et al.) and 14 (Bartel et al.) of this book. In brief, proteins destined for the peroxisomal matrix usually harbor a peroxisomal targeting sequence (PTS). Most peroxisomal matrix proteins carry a C-terminal PTS1-sequence, which is recognized by the PTS1-receptor Pex5p, while a subset of proteins displays an N-terminal PTS2-sequence via which they are ferried to the peroxisomal membrane by the PTS2-receptor Pex7p and its co-receptors, like S. cerevisiae Pex18p or P. pastoris Pex20p. Here, the cargo-bound PTS-receptors interact with constituents of the docking complex (Pex13p, Pex14p), which results in the formation of a transient import pore and finally the transloction and release of the cargo.
Subsequent to the liberation of the cargo, the PTS-receptors return to the cytosol for further rounds of matrix protein import (Fig. 15.2). This retrotranslocation is facilitated by the peroxisomal receptor export machinery, the exportomer (Platta et al. 2013). The monoubiquitination of the PTS-receptors is regarded as a central event in this process and has been shown to depend on the ubiquitin-conjugating enzyme Pex4p, its membrane anchor Pex22p and the presence of the peroxisomal RING–peroxin complex (Pex2p, Pex10p, Pex12p). The ubiquitination is thought to prime the PTS-receptors for the recognition by the AAA-type ATPase complex (Pex1p, Pex6p), which functions as dislocase by extracting the modified PTS-receptors from the membrane. Furthermore, the recent “export-driven-import model” postulates that the ATP-dependent export of the PTS-receptors may be directly linked to the translocation of the cargo proteins into the peroxisomal matrix. In case the monoubiquitination-dependent receptor recycling pathway is blocked, the PTS-receptors become substrates of a polyubiquitination-dependent proteolytic pathway, which promotes their degradation by the 26S proteasome.
Ubiquitination and export of the PTS1-receptor at the peroxisomal membrane in Saccharomyces cerevisiae. At the peroxisomal membrane, the PTS1-receptor Pex5p is either mono- or polyubiquitinated. Both ubiquitination cascades are initiated by the ATP-dependent ubiquitin-activating enzyme Uba1p (E1). For monoubiquitination of Pex5p as part of the typical receptor cycle (left), the activated ubiquitin is transferred to the ubiquitin-conjugating enzyme Pex4p (E2) and then attached to a conserved cysteine of the receptor by assistance of the RING-ligase Pex12p (E3). The ubiquitinated receptor is ATP-dependent export of Pex5p catalyzed by the Pex15p-anchored AAA-peroxins Pex1p and Pex6p. For a new round of import the ubiquitin is cleaved off by the deubiquitinating enzyme Ubp15p. For polyubiquitination of Pex5p as part of a quality control pathway (right), the activated ubiquitin is transferred to the ubiquitin-conjugating enzyme Ubc4p (E2) and then delivered to conserved lysines of the receptor by the RING-ligase Pex2p (E3). The following export of Pex5p is also performed by the AAA peroxins Pex1p and Pex6p and the polyubiquitination signal leads to a degradation of Pex5p by the 26S proteasome
3 The Peroxisomal Receptor Export Machinery: The Exportomer
The import of matrix proteins depends on the cycle of the PTS-receptors, which itself relies on the function of each constituent of the membrane-bound peroxins. Previous studies elucidated the composition of the peroxisomal membrane-bound subcomplexes, the docking- and the RING complex (Agne et al. 2003; Hazra et al. 2002). These two complexes were referred to as the “importomer” because both are physically connected by Pex8p in S. cerevisiae and both are required for matrix protein import (Agne et al. 2003). However, later work revealed that also constituents of the AAA complex and Pex4p complex could be co-purified with the importomer components, which strongly suggests that all membrane-associated peroxin complexes required for matrix protein import are dynamically interconnected (Oeljeklaus et al. 2012; Rosenkranz et al. 2006; Platta et al. 2009). Therefore, an alternative approach to define functionally related subcomplexes could be based on the steps of the PTS-receptor cycle at the membrane in general and on the energy dependence in particular. The current view is that the association of the PTS-receptors with the peroxisomal membrane at the site of the importomer is ATP independent (Miyata and Fujiki 2005; Miyata et al. 2009; Oliveira et al. 2003; Platta et al. 2005). However, the RING complex (Pex2p, Pex10p, Pex12p) as well as the Ubc components (Pex22p, Pex4p, Ubc4p family, UbcH5 family) belong to the ATP-dependent ubiquitination cascade (El Magraoui et al. 2012; Grou et al. 2008; Platta et al. 2007a, b, 2009; Williams et al. 2007, 2008, 2012; Liu and Subramani 2013; Kaur et al. 2013) and form together with the AAA-type ATPase complex (Pex1p, Pex6p, Pex15p, Ubp15p, AWP1) (Debelyy et al. 2011; Hensel et al. 2011; Leon et al. 2006b; Miyata and Fujiki 2005; Miyata et al. 2012) the receptor export machinery, or alternatively, peroxisomal exportomer (Table 15.1).
3.1 The Ubiquitin-Conjugating Enzymes Required for Monoubiquitination of the Receptors
Ubiquitination is a posttranslational protein modification that is mediated by a three-step enzyme cascade. The ubiquitin-activating enzyme (E1) activates ubiquitin via an AMP-bound intermediate and transfers it to an ubiquitin-conjugating enzyme (E2). Finally, an ubiquitin-protein ligase (E3) binds the ubiquitin-charged E2 as well as the substrate protein, thereby enabling the transfer of the ubiquitin moiety to the target amino acid residue of the substrate (Kerscher et al. 2006; Ravid and Hochstrasser 2008). Regularly, the epsilon-amino group of a lysine within the target protein is covalently linked to ubiquitin via an isopeptide bond. Interestingly, ubiquitin can also be attached via a peptide bond to the alpha-amino group to the N-terminal amino acid, or via an oxyester bond to a threonine or serine, or even via a thioester bond to a cysteine (Wang et al. 2012).
The peroxisomal matrix protein import depends on the unusual ubiquitination of a conserved cysteine of the PTS1-receptor Pex5p (Carvalho et al. 2007; Okumoto et al. 2011; Williams et al. 2007) and of the PTS2-co-receptors Pex18p (Hensel et al. 2011) or Pex20p (Liu and Subramani 2013). The E2-enzyme that has been demonstrated to catalyze the cysteine-dependent monoubiquitination of S. cerevisiae Pex5p both in vivo and in vitro is Pex4p (Ubc10p; Platta et al. 2007a; Williams et al. 2007). Recently, the monoubiquitination of S. cerevisiae Pex18p (El Magraoui et al. 2013) and P. pastoris Pex20p (Liu and Subramani 2013) has been demonstrated to depend on the presence of Pex4p as well.
The soluble E2-enzyme Pex4p is essential for the import of both PTS1 and PTS2 proteins and therefore was the first E2-enzyme shown to be essential for the biogenesis of an organelle (Crane et al. 1994; van der Klei et al. 1998; Wiebel and Kunau 1992; Zolman et al. 2005). Pex4p is anchored to peroxisomes via the membrane protein Pex22p (Koller et al. 1999; Zolman et al. 2005). The crystal structure of S. cerevisiae Pex4p complexed to Pex22p (without its membrane domain) revealed that the Pex22p-binding site in Pex4p does not resemble a common substrate-binding motif and therefore it has been suggested that Pex22p may act as a co-activator of this E2-enzyme (Williams et al. 2012).
The molecular function of the Pex4p-catalyzed monoubiquitination of the membrane-bound Pex5p is to prime the PTS1-receptor for export (Platta et al. 2007a). While Pex4p and Pex22p are well conserved in yeasts and plants, they are absent in the genomes of mammals (Kiel et al. 2006). Instead, members of the E2D family of E2-enzymes (UbcH5a, UbcH5b and UbcH5c) fulfill the function of Pex4p in mammals (Grou et al. 2008). They catalyze the monoubiquitination of mammalian Pex5p on the conserved cysteine and therefore are required for the receptor export in vitro (Grou et al. 2008). Even though the three UbcH5 proteins carry out a central task in peroxisome biogenesis, their cellular targets are not restricted to this organelle (Brzovic and Klevit 2006; Gonen et al. 1999; Saville et al. 2004). Future work may reveal why the monoubiquitination of Pex5p has been transferred to the promiscuous UbcH5 proteins. One possible explanation could be that they control cellular events that are interconnected with peroxisome function in a concerted manner.
3.2 The RING–Peroxin Complex
The import of peroxisomal matrix proteins requires the presence of the three RING-finger proteins Pex2p, Pex10p, and Pex12p (Albertini et al. 2001; Chang et al. 1999; Eckert and Johnsson 2003; Okumoto et al. 2000; Berteaux-Lecellier et al. 1995; Krazy and Michels 2006; Peraza-Reyes et al. 2008; Sparkes et al. 2003). They have been found to form a distinct subcomplex at the peroxisomal membrane (Agne et al. 2003; Hazra et al. 2002). Defects in the assembly of the human RING complex are the second most common cause of peroxisomal biogenesis disorders (Ebberink et al. 2011; Steinberg et al. 2006). The mammalian Pex2p (formerly PAF-1) was the first gene that could be linked to PBDs (Shimozawa et al. 1992; Tsukamoto et al. 1991). Work from S. cerevisiae and A. thaliana in recent years has uncovered that all three peroxins display ubiquitin-protein ligases activity (Kaur et al. 2013; Platta et al. 2009; Williams et al. 2008). The RING–peroxins are directly involved in the ubiquitination of the PTS1-receptor Pex5p in S. cerevisiae (Platta et al. 2009; Williams et al. 2008). Their activity is also required for the formation of ubiquitinated Pex20p in P. pastoris (Liu and Subramani 2013).
In general, E3-enzymes determine the substrate specificity of ubiquitination reactions because they bind the ubiquitin-charged E2-enzyme and the substrate, thereby insuring a specific transfer of ubiquitin to the target amino acid. RING-type E3-enzymes catalyze the direct transfer of ubiquitin from the E2-enzyme to the substrate (Deshaies and Joazeiro 2009). They belong to the superfamily of Treble-Clef fold-containing proteins. This scaffold structure, which is stabilized by a Zn2+-ion, functions as an interaction motif in diverse proteins even outside the ubiquitin system (Burroughs et al. 2011). The canonical RING-finger domain (Freemont et al. 1991) binds two Zn2+-ions through its conserved Cys and His residues in a “cross-brace” arranged manner (Deshaies and Joazeiro 2009). The RING domains of Pex2p and Pex10p coordinate two Zn2+-ions, whereas the RING-finger of Pex12p binds only one Zn2+-ion (Koellensperger et al. 2007). It is interesting to note that also several members of the RBR-(RING-between-RING) family of E3-enzymes, e.g., the Parkin-like Ariadne, contain an active RING domain at their carboxy-terminus containing a single Zn2+-ion (Eisenhaber et al. 2007).
The RING–peroxins Pex2p, Pex10p, and Pex12p assemble to a distinct complex and stabilize each other in vivo (Agne et al. 2003; Hazra et al. 2002). Based on earlier binary interaction studies (Albertini et al. 2001; Chang et al. 1999; Eckert and Johnsson 2003; Okumoto et al. 2000) and recent in vitro interaction data on all three RING domains (El Magraoui et al. 2012), the RING–peroxins are thought to form a heterotrimeric complex. Pex10p (RING) functions as central component of the ternary complex as it directly binds to Pex2p(RING) and Pex12p(RING) thereby bridging the indirect interaction between these two RING domains (El Magraoui et al. 2012). The heteromeric architecture of the RING complex has a direct influence on the E3-ligase activity of the RING–peroxins because the ubiquitination activity of the combined Pex10p/Pex12p RING-domains is enhanced in presence of Pex4p in vitro (El Magraoui et al. 2012).
Pex10p also fulfills additional tasks that are distinct from Pex2p and Pex12p. A systematic functional screen of all peroxins in A. thaliana uncovered that only Pex10p has a pleiotropic growth phenotype (Nito et al. 2007). Furthermore, overexpression experiments of proteins with mutated RING domain in wild-type background suggested that A. thaliana Pex10p but not Pex2p or Pex12p are required for the contact of peroxisomes to chloroplasts during photorespiration (Prestele et al. 2010; Schumann et al. 2007). However, whether this association is due to a physical interaction or due to a functional interaction via ubiquitination events remains to be investigated.
Early studies already linked the function of the RING–peroxins to the recycling of Pex5p (Chang et al. 1999; Dodt and Gould 1996) and Pex20p (Leon et al. 2006b) as these receptors accumulate at the peroxisomal membrane in cells with disrupted RING complex. Because the monoubiquitination of Pex5p is reported to be essential for the export of Pex5p (Grou et al. 2008; Platta et al. 2007a; Okumoto et al. 2011) and as Pex12p (RING) cooperates with Pex10p(RING) in vitro (El Magraoui et al. 2012) and catalyzes this Pex4p-dependent modification in vivo (Platta et al. 2009), the Pex10p/Pex12p unit may function as the physiologic active ligase complex dedicated to the monoubiquitination-mediated export of the PTS1-receptor.
3.3 The Peroxisomal AAA-Type ATPase Complex
The ubiquitinated PTS1-receptor Pex5p is substrate for the peroxisomal AAA-type ATPase complex, which functions as dislocase that extracts Pex5p from the membrane and thereby exports it back to the cytosol (Fujiki et al. 2012; Grimm et al. 2012; Miyata and Fujiki 2005; Platta et al. 2005, 2008). The two peroxisomal AAA proteins Pex1p and Pex6p display a non-redundant and essential function in this process (Birschmann et al. 2005; Kiel et al. 1999, 2000; Tamura et al. 1998; Tamura et al. 2006).
The AAA peroxins associate with peroxisomes via an interaction of Pex6p to the tail-anchored membrane protein Pex15p in yeast and the orthologous Pex26p in mammals as well as APEM9 in plants (Birschmann et al. 2003; Furuki et al. 2006; Goto et al. 2011; Matsumoto et al. 2003a, b). An impaired assembly of the human AAA complex is the most common cause of Zellweger syndrome spectrum disorders (Geisbrecht et al. 1998; Steinberg et al. 2006). It is interesting to point out that Pex1p (formerly PAS1) was the first peroxin to be identified and also one of the founding members of the AAA family (Beyer 1997; Erdmann et al. 1991; Kunau et al. 1993).
In general, AAA proteins are characterized by a conserved modular architecture. They can be classified as P-loop NTPases, which are characterized by conserved motifs for NTP binding (Walker A motif) and hydrolysis (Walker B motif; Walker et al. 1982). AAA proteins in particular are defined by the evolutionary conserved AAA domain that contains the Walker A and B motifs as well as other conserved regions like the Second Region of Homology (SRH; Beyer 1997; Neuwald et al. 1999; Wendler et al. 2012). Pex1p and Pex6p harbor two AAA domains (AAA-D1 and AAA-D2) as well as an N-terminal domain (NTD). The binding and hydrolysis of ATP by the AAA peroxins are thought to result in conformational changes, as shown for p97 (Beuron et al. 2003), ClpX (Stinson et al. 2013) or NSF (Cipriano et al. 2013).
Most AAA proteins form active oligomers with predominantly hexameric constitution (Iyer et al. 2004). However, the current knowledge on the structural assembly of the AAA peroxins Pex1p and Pex6p is still scarce and even though they are thought to form a hetero-oligomeric complex, the stoichiometry has not yet been solved. Distinct ATP-binding and hydrolysis sites contribute to the assembly of the AAA complex (Birschmann et al. 2003, 2005; Nashiro et al. 2011; Tamura et al. 2006; Saffian et al. 2012). In yeast, ATP binding and hydrolysis in Pex6p regulate the assembly and disassembly with Pex15p (Birschmann et al. 2003), while the Pex1p–Pex6p interaction is influenced by ATP binding in D2 of Pex1p (Birschmann et al. 2005). Furthermore, the release of the AAA peroxins from the peroxisomal membrane might be regulated by the E2-enzyme Pex4p, because Pex1p and Pex6p accumulate at the peroxisome in Pex4p-deficient yeast cells. This might indicate that the ubiquitin-dependent PTS1-receptor cycle and the dynamic ATPase cycle of the AAA peroxins are interconnected (Rosenkranz et al. 2006).
In addition to their involvement in matrix protein import, the AAA peroxins have been suggested to function in the fusion of pre-peroxisomal vesicles in yeasts (Titorenko and Rachubinski 2000; van der Zand et al. 2012), while Pex6p seems to be involved in the suppression of different cell death mechanisms (Jungwirth et al. 2008; Seo et al. 2007; Warner et al. 2003). However, the best analyzed function of Pex1p and Pex6p to date is their role in peroxisomal matrix protein import (Miyata and Fujiki 2005; Platta et al. 2005; Grimm et al. 2012; Fujiki et al. 2012).
While accumulating evidence strongly indicates that the purpose of monoubiquitination is to prime Pex5p for AAA complex-mediated dislocation, the direct mechanistic purpose of this modification remains elusive. In this context, it is interesting to note that the X-ray structure of the N-domain of murine Pex1p contains a double-psi-beta-barrel fold (Shiozawa et al. 2004). This fold is also present in the N-domain of p97, where it functions as binding module for ubiquitin (Park et al. 2005). However, if the domain found in Pex1p carries out a similar function still has to be investigated. AWP1 (Associated with PRK1) has been identified as a novel binding protein of human Pex6p (Miyata et al. 2012) and is supposed to contribute to linking of the AAA peroxins to the ubiquitinated Pex5p. Accordingly, AWP1 is required for peroxisomal biogenesis in vivo and the protein interacts with both Pex6p as well as with monoubiquitinated Pex5p (Miyata et al. 2012). Thus, AWP1 might function as specific adaptor, which links the modified Pex5p to the AAA peroxins and enables them to transfer their suggested pulling force to the monoubiquitinated PTS1-receptor. Interestingly, AWP1 has also been described as an ubiquitin-binding modulator of NF-kappaB (Fenner et al. 2009).
3.4 Deubiquitination of the Receptor
The ubiquitin moiety is removed from the PTS1-receptor during or shortly after the export step but certainly prior to a new round of matrix protein import. In general, the cleavage of ubiquitin from a substrate protein is catalyzed by ubiquitin hydrolases that are called deubiquitinating enzymes (Amerik and Hochstrasser 2004). The ubiquitin hydrolase Ubp15p has been identified as a binding partner of Pex6p in S. cerevisiae (Debelyy et al. 2011). Ubp15p functions as deubiquitinating enzyme acting on Pex5p, which represents the first characterized target of this enzyme (Debelyy et al. 2011). Work based on an in vitro system with mammalian proteins suggests that the thioester bond between ubiquitin and Pex5p can be cleaved either non-enzymatically via a nucleophilic attack of glutathione or, as the major pathway, enzyme-catalyzed by ubiquitin hydrolases (Grou et al. 2009b). USP9X has been described as the main deubiquitinating enzyme acting on mammalian Pex5p (Grou et al. 2012). USP9X is a cytosolic protein whose function is not restricted to peroxisomal protein import because it has been described to take part in the regulation of the transforming growth factor beta (TGFbeta) pathway (Dupont et al. 2009).
4 Functional Link Between Receptor Export and Cargo Release
Early work has defined that the import of peroxisomal matrix proteins requires the hydrolysis of ATP (Imanaka et al. 1987). Later studies identified the export of the receptor back to the cytosol as the energy-dependent step (Oliveira et al. 2003; Gouveia et al. 2003). In recent years, it has become evident that the ubiquitination machinery (Carvalho et al. 2007; Grou et al. 2008, 2009b; Okumoto et al. 2011; Platta et al. 2007a) as well as the AAA complex (Leon et al. 2006b; Miyata and Fujiki 2005; Miyata et al. 2012; Platta et al. 2005, 2007a; Kerssen et al. 2006) can be regarded as the only ATP-consuming factors of the peroxisomal protein import machinery. This indicates that energy consumption, matrix-protein import, and PTS-receptor export merge at the exportomer.
In this respect, it is interesting to note that the protein composition of the exportomer is functionally and evolutionary related to the proteins of the endoplasmic reticulum associated degradation (ERAD) machinery (Gabaldon et al. 2006; Schluter et al. 2006). ERAD can be defined as a mechanism by which misfolded proteins are polyubiquitinated and extracted from the ER in order to be disposed by the 26S proteasome in the cytosol (Hampton and Sommer 2012). Translocation systems that are in many aspects comparable to the exportomer and ERAD are the mitochondria associated degradation (MAD) for proteins of the outer mitochondrial membrane (Taylor and Rutter 2011) as well as the pre-protein translocator of complex plastids called symbiont-derived ERAD-like machinery (SELMA; Bolte et al. 2011). Therefore, a mechanistic parallel can be drawn between the exportomer, ERAD-, MAD-, and SELMA substrates because all are extracted by mechanoenzymes of the AAA-type ATPase family in an ubiquitination-dependent manner (Bolte et al. 2011; Platta et al. 2007b; Schliebs et al. 2010).
Based on this similarity, a model has been proposed that draws a direct interconnection of receptor export and the translocation of matrix proteins across the peroxisomal membrane (Schliebs et al. 2010). This “export-driven import model” is supported by the fact that the presence of a functional exportomer is a prerequisite for the import of matrix proteins. This, ATP is required for the ubiquitin- and AAA-driven extraction of the receptor and might be mechanically coupled to the translocation of the cargo proteins over the membrane.
Accordingly, the import defects observed in mutants of the exportomer can be explained in two ways. First, the binding capacity for functional PTS-receptors at the peroxisomal membrane seems to be limited. In fact, a decreased rate of receptor export caused by the functional impairment of the export machinery leads to an accumulation of PTS-receptors at the membrane (Leon et al. 2006b; Platta et al. 2004) and therefore would block the docking of new receptor–cargo complexes from the cytosol. In A. thaliana, the physiological defects of mutated and only insufficiently active Pex6p could be partially overcome when it was co-expressed with a weak allele of the docking protein Pex13p (Ratzel et al. 2011). This finding strongly indicates that the import and export rates of the PTS-receptors need to be balanced. Second, this model suggests that export of the receptor and the release of the cargo might be directly linked by a concerted mechanism. Work on the ubiquitination of the S. cerevisiae PTS2-co-receptor Pex18p delivered first direct evidence for such a connection (Hensel et al. 2011). Based on protease-protection assays, it was revealed that Pex7p is partially protease protected in wild-type cells, while Pex18p remains accessible. This topology is reversed when the cysteine of Pex18p is mutated or the AAA peroxins Pex1p/Pex6p are deleted (Hensel et al. 2011). This finding strongly indicates that monoubiquitination of Pex18p as well as AAA complex governs the import of cargo-loaded Pex7p. However, in the mammalian system, it is not yet clear whether the cargo release step itself requires ATP hydrolysis (Miyata et al. 2009) or does not (Alencastre et al. 2009).
In conclusion, the receptor export machinery is thought to function as the energy-consuming import motor for matrix proteins, either indirectly via balanced receptor import/export rates and/or directly via an interconnection of receptor export and cargo translocation.
5 Polyubiquitination of the PTS-Receptors
Under certain conditions, the PTS-receptors Pex5p, Pex18p, and Pex20p are polyubiquitinated on lysine residues in order to mark them for the degradation by the 26S proteasome (Hensel et al. 2011; Kiel et al. 2005b; Leon et al. 2006b; Platta et al. 2007a; Williams et al. 2007). This proteolytic pathway is induced when the normal monoubiquitination-dependent recycling pathway is blocked, as it is the case when constituents of the Pex4p or AAA complexes are deleted or the conserved cysteine of the PTS-receptor is mutated (Kiel et al. 2005a; Kragt et al. 2005; Platta et al. 2004; Leon and Subramani 2007; Hensel et al. 2011). The polyubiquitination of the S. cerevisiae PTS1-receptor Pex5p is predominantly catalyzed by Ubc4p and to a minor portion by the partial redundant enzymes Ubc5p and Ubc1p (Kiel et al. 2005a; Kragt et al. 2005; Platta et al. 2004). These three ubiquitin-conjugating enzymes display a high sequence similarity and are involved in diverse other cellular processes as well (Seufert and Jentsch 1990; Seufert et al. 1990). Both Pex10p (Williams et al. 2008) as well as Pex2p (Platta et al. 2009) have been suggested to function as E3 enzymes for the polyubiquitination of Pex5p. In this respect, it is interesting to note that a recent in vitro study demonstrates that Pex10p (RING) can synergistically enhance the ubiquitination activity of the Ubc4p–Pex2p (RING) enzyme pair (El Magraoui et al. 2012). This result suggests that both RING–peroxins may act together in the Ubc4p-dependent generation of K48-linked polyubiquitin chains on Pex5p.
Receptor polyubiquitination is enhanced when the export machinery is affected in its function and therefore the purpose of this modification is likely to remove the aberrant receptor molecules from the membrane when the normal extraction and recycling reaction is hampered. However, mutagenesis of the lysine residues required for polyubiquitination of S. cerevisiae Pex5p does not lead to a growth defect on oleate medium (Platta et al. 2007a; Williams et al. 2007). Interestingly, polyubiquitination of Pex5p can also be regarded as an alternative export signal. In vitro export assays demonstrated that a fraction of Pex5p is still exported even in a Pex4p-deficient system when the two conserved lysine residues required for polyubiquitination were still present (Platta et al. 2007a). Moreover, mutation of the conserved cysteine in P. pastoris Pex20p (Leon and Subramani 2007) induces polyubiquitination of Pex20p but still retains a partial functional receptor molecule that displays partial complementation in growth tests. Interestingly, only both the non-essential lysine targets for polyubiquitination of Pex20p as well as to the typically monoubiquitinated cysteine are mutated, the receptor completely loses its functionality (Leon and Subramani 2007). These data demonstrate that the enhanced degradation of Pex20p can restore the matrix protein import to a certain extent, supposedly because the receptors are removed efficiently enough to allow the docking of further cargo-bound receptors. This mechanism has been described as RADAR (receptor accumulation and degradation in the absence of recycling; Leon et al. 2006a, b) in order to distinguish it from the non-essential quality control. However, it should be noted that the mutation of the conserved cysteine of S. cerevisiae Pex5p and Pex18p alone is already sufficient to fully abolish the function of these receptors (Hensel et al. 2011; Williams et al. 2007). In this respect, it is interesting to note that degradation of Pex5p occurs much slower in S. cerevisiae than in most other species (Collins et al. 2000; Dodt and Gould 1996; van der Klei et al. 1998; Zolman and Bartel 2004; Zolman et al. 2005). Therefore, the observed instability of Pex5p in exportomer mutants in these species is most likely due to rapid degradation via K48-linked polyubiquitination as described for the PTS1-receptor of H. polymorpha (Kiel et al. 2005b).
The S. cerevisiae PTS2-co-receptor Pex18p behaves somewhat different from the PTS2-co-receptor Pex20p in P. pastoris and H. polymorpha because Pex18p shows a constitutive turnover already under wild-type conditions (Hensel et al. 2011; Leon et al. 2006b; Otzen et al. 2005; Purdue and Lazarow 2001). Currently, the functional impact of this instability is not known. In contrast to Pex18p, the PTS2-receptor Pex7p of S. cerevisiae is a stable protein (Hensel et al. 2011). So far, no indications for an ubiquitination of yeast Pex7p have been found. Interestingly, a recent report describes the polyubiquitination and degradation of Arabidopsis Pex7p when the dominant-negative GFP-Pex7p species is expressed in the cell (Cui et al. 2013). However, it is not clear if this mechanism is conserved in other organisms.
In general, the removal of the PTS-receptors via polyubiquitination is initiated when the monoubiquitination-dependent recycling pathway is blocked and therefore may function as alternative export signal.
6 Concluding Remarks
The combined work of several laboratories on the ubiquitination and recycling of the PTS-receptors has helped to uncover the functional contribution of distinct peroxisomal subcomplexes to the dislocation step and therefore enabled the definition of the peroxisomal receptor export machinery, the exportomer (Platta et al. 2013).
Certainly, many open questions remain to be answered and one of the most intriguing ones concerns the finding that Pex5p, Pex18p, and Pex20p are monoubiquitinated on a cysteine via a thioester bond and not by a more common isopeptide bond to a lysine. The first evidence that ubiquitin can be attached to cysteine, serine or threonine residues came from studies of viral MARCH (Membrane-associated RING-CH) E3 ligases that ubiquitinate MHC I (Major Histocompatibility Complex I) molecules (Cadwell and Coscoy 2005; Wang et al. 2007), and recent work demonstrates that this uncommon ubiquitination can also take place during ERAD (Ishikura et al. 2010; Shimizu et al. 2010). However, it is unclear how the specificity for these non-lysine ubiquitination reactions is ensured because the E2 and E3 enzymes involved are not restricted to this kind of modification and can also modify lysine residues (Wang et al. 2012). Interestingly, the cysteine of mammalian Pex5p (Grou et al. 2009b, 2012) and P. pastoris Pex20p (Leon and Subramani 2007) can be replaced by a lysine, which results in a still largely functional protein. Thus, even though the cysteine and the thioester-bond mediated ubiquitination of the PTS-receptors are evolutionary conserved, they are not essential for the principle export mechanisms and therefore may mainly represent an important regulatory device.
There are several different possibilities to explain the function of the conserved cysteine of the peroxisomal receptors. (1) The first concept is based on the fact that thioester bonds are less stable in comparison to isopeptide bonds. Therefore, the duration of the ubiquitin moiety at the PTS-receptor might be restricted in order to disable the formation of a polyubiquitin chain or to prevent the recognition by proteasomal adaptors. The rapid non-enzymatic disruption of the thioester bond of Ub-Pex5p in a mammalian in vitro system supports the idea that the cysteine-ubiquitination protects the PTS-receptors against degradation (Grou et al. 2009a, b). (2) Another concept takes into account that that certain E3 enzymes, like HECT-type ligases (Kee and Huibregtse 2007) or the RBR-type ligases (Wenzel et al. 2011) form an ubiquitin-thioester intermediate on a cysteine before this ubiquitin molecule is finally transferred to the substrate protein. One hypothetical model could be that once Pex12p/Pex10p have modified one of the receptor molecules of the oligomeric pore, Pex5p itself could catalyze an intra-oligomeric ubiquitin transfer in a relay-like system in order to accelerate the decomposition of the pore (Erdmann and Schliebs 2005; Platta et al. 2013). Interestingly, the E2-enzyme E2-230 K represents an example of an intramolecular ubiquitin transfer (Berleth and Pickart 1996), where ubiquitin is transferred from the first cysteine to a second cysteine of E2-230 K prior to attachment of the ubiquitin to the target protein. (3) A third concept is related to recent work on the regulation of the peroxisomal redox balance (Ivashchenko et al. 2011), which contributes to the general functional role of peroxisomes in the control of the cellular levels of reactive oxygen species (Bonekamp et al. 2009). One possibility is that the cysteine required for monoubiquitination might be accessible for redox changes. This might have a direct impact on the availability of this residue for the monoubiquitination and therefore could control the import/export rates of the receptor (Fransen et al. 2012).
In conclusion, the understanding of the late acting peroxins as concerted acting components of the exportomer will be instrumental to uncover the molecular mechanism underlying peroxisomal matrix protein import.
References
Agne B, Meindl NM, Niederhoff K, Einwächter H, Rehling P, Sickmann A, Meyer HE, Girzalsky W, Kunau WH (2003) Pex8p. An intraperoxisomal organizer of the peroxisomal import machinery. Mol Cell 11(3):635–646
Albertini M, Girzalsky W, Veenhuis M, Kunau W-H (2001) Pex12p of Saccharomyces cerevisiae is a component of a multi-protein complex essential for peroxisomal matrix protein import. Eur J Cell Biol 80(4):257–270
Albiniak AM, Baglieri J, Robinson C (2012) Targeting of lumenal proteins across the thylakoid membrane. J Exp Bot 63(4):1689–1698
Alencastre IS, Rodrigues TA, Grou CP, Fransen M, Sá-Miranda C, Azevedo JE (2009) Mapping the Cargo Protein Membrane Translocation Step into the PEX5 Cycling Pathway. J Biol Chem 284(40):27243–27251
Amerik AY, Hochstrasser M (2004) Mechanism and function of deubiquitinating enzymes. Biochim Biophys Acta 1695:189–207
Baes M, Van Veldhoven PP (2012) Mouse models for peroxisome biogenesis defects and β-oxidation enzyme deficiencies. Biochim Biophys Acta 1822(9):1489–1500
Baker A, Graham IA, Holdsworth M, Smith SM, Theodoulou FL (2006) Chewing the fat: beta-oxidation in signalling and development. Trends Plant Sci 11(3):124–132
Berleth ES, Pickart CM (1996) Mechanism of ubiquitin conjugating enzyme E2-230 K: catalysis involving a thiol relay? Biochemistry 35(5):1664–1671
Berteaux-Lecellier V, Picard M, Thompson-Coffe C, Zickler D, Panvier-Adoutte A, Simonet JM (1995) A nonmammalian homolog of the PAF1 gene (Zellweger syndrome) discovered as a gene involved in caryogamy in the fungus Podospora anserina. Cell 81(7):1043–1051
Beuron F, Flynn TC, Ma J, Kondo H, Zhang X, Freemont PS (2003) Motions and negative cooperativity between p97 domains revealed by cryo-electron microscopy and quantised elastic deformational model. J Mol Biol 327(3):619–629
Beyer A (1997) Sequence analysis of the AAA protein family. Protein Sci 6(10):2043–2058
Birschmann I, Stroobants AK, Van Den Berg M, Schäfer A, Rosenkranz K, Kunau WH, Tabak HF (2003) Pex15p of Saccharomyces cerevisiae Provides a Molecular Basis for Recruitment of the AAA Peroxin Pex6p to Peroxisomal Membranes. Mol Biol Cell 14(6):2226–2236
Birschmann I, Rosenkranz K, Erdmann R, Kunau WH (2005) Structural and functional analysis of the interaction of the AAA-peroxins Pex1p and Pex6p. FEBS J 272(1):47–58
Bolte K, Gruenheit N, Felsner G, Sommer MS, Maier UG, Hempel F (2011) Making new of old: recycling and modification of an ancient protein translocation system during eukaryotic evolution. Mechanistic comparison and phylogenetic analysis of ERAD, SELMA and the peroxisomal importomer. Bioessays 33(5):368–376
Bonekamp NA, Völkl A, Fahimi HD, Schrader M (2009) Reactive oxygen species and peroxisomes: struggling for balance. Biofactors 35(4):346–355
Brzovic PS, Klevit RE (2006) Ubiquitin transfer from the E2 perspective: why is UbcH5 so promiscuous? Cell Cycle 5:2867–2873
Burroughs AM, Iyer LM, Aravind L (2011) Functional diversification of the RING finger and other binuclear treble clef domains in prokaryotes and the early evolution of the ubiquitin system. Mol Biosyst 7(7):2261–2277
Cadwell K, Coscoy L (2005) Ubiquitination on nonlysine residues by a viral E3 ubiquitin ligase. Science 309:127–130
Carvalho AF, Pinto MP, Grou CP, Alencastre IS, Fransen M, Sa-Miranda C, Azevedo JE (2007) Ubiquitination of mammalian Pex5p, the peroxisomal import receptor. J Biol Chem 282(43):31267–31272
Chang CC, Warren DS, Sacksteder KA, Gould SJ (1999) PEX12 Interacts with PEX5 and PEX10 and Acts Downstream of Receptor Docking in Peroxisomal Matrix Protein Import. J Cell Biol 147(4):761–774
Cipriano DJ, Jung J, Vivona S, Fenn TD, Brunger AT, Bryant Z (2013) Processive ATP-driven disassembly of SNARE complexes by the N-ethylmaleimide sensitive factor molecular machine. J Biol Chem. doi:10.1074/jbc.M1113.476705
Collins CS, Kalish JE, Morrell JC, McCaffery JM, Gould SJ (2000) The peroxisome biogenesis factors Pex4p, Pex22p, Pex1p, and Pex6p Act in the terminal steps of peroxisomal matrix protein import. Mol Cell Biol 20(20):7516–7526
Cooper TG, Beevers H (1969) Beta oxidation in glyoxysomes from castor bean endosperm. J Biol Chem 244(13):3514–3520
Crane DI, Kalish JE, Gould SJ (1994) The Pichia pastoris PAS4 gene encodes a ubiquitin-conjugation enzyme required for peroxisome assembly. J Biol Chem 269(34):21835–21844
Cui S, Fukao Y, Mano S, Yamada K, Hayashi M, Nishimura M (2013) Proteomic analysis reveals that the Rab GTPase RabE1c is involved in the degradation of the peroxisomal protein receptor PEX7 (peroxin 7). J Biol Chem 288(8):doi: 1.1074/jbc.M1112.438143
Debelyy MO, Platta HW, Saffian D, Hensel A, Thoms S, Meyer HE, Warscheid B, Girzalsky W, Erdmann R (2011) Ubp15p, a ubiquitin hydrolase associated with the peroxisomal export machinery. J Biol Chem 286(32):28223–28234
Deshaies RJ, Joazeiro CA (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78:399–434
Distel B, Erdmann R, Gould SJ, Blobel G, Crane DI, Cregg JM, Dodt G, Fujiki Y, Goodman JM, Just WW, Kiel JA, Kunau WH, Lazarow PB, Mannaerts GP, Moser HW, Osumi T, Rachubinski RA, Roscher A, Subramani S, Tabak HF, Tsukamoto T, Valle D, van der Klei I, van Veldhoven PP, Veenhuis M (1996) A unified nomenclature for peroxisome biogenesis factors. J Cell Biol 135(1):1–3
Dodt G, Gould SJ (1996) Multiple PEX genes are required for proper subcellular distribution and stability of Pex5p, the PTS1 receptor: Evidence that PTS1 protein import is mediated by a cycling receptor. J Cell Biol 135:1763–1774
Dupont S, Mamidi A, Cordenonsi M, Montagner M, Zacchigna L, Adorno M, Martello G, Stinchfield MJ, Soligo S, Morsut L, Inui M, Moro S, Modena N, Argenton F, Newfeld SJ, Piccolo S (2009) FAM/USP9x, a deubiquitinating enzyme essential for TGFbeta signaling, controls Smad4 monoubiquitination. Cell 136(1):123–135
Ebberink MS, Mooijer PA, Gootjes J, Koster J, Wanders RJ, Waterham HR (2011) Genetic classification and mutational spectrum of more than 600 patients with a Zellweger syndrome spectrum disorder. Hum Mutat 32(1):59–69
Eckert JH, Johnsson N (2003) Pex10p links the ubiquitin conjugating enzyme Pex4p to the protein import machinery of the peroxisome. J Cell Sci 116(17):3623–3634
Eisenhaber B, Chumak N, Eisenhaber F, Hauser MT (2007) The ring between ring fingers (RBR) protein family. Genome Biol 8(3):209
El Magraoui F, Bäumer BE, Platta HW, Baumann JS, Girzalsky W, Erdmann R (2012) The RING-type ubiquitin ligases Pex2p, Pex10p and Pex12p form a heteromeric complex that displays enhanced activity in an ubiquitin conjugating enzyme-selective manner. FEBS J 279(11):2060–2070
El Magraoui F, Brinkmeier R, Schrötter A, Girzalsky W, Müller T, Marcus K, Meyer HE, Erdmann R, Platta HW (2013) Distinct ubiquitination cascades act on the peroxisomal targeting signal type 2 co-receptor Pex18p. Traffic 14(12):1290–1301
Erdmann R, Schliebs W (2005) Peroxisomal matrix protein import: the transient pore model. Nat Rev Mol Cell Biol 6(9):738–742
Erdmann R, Wiebel FF, Flessau A, Rytka J, Beyer A, Fröhlich KU, Kunau W-H (1991) PAS1, a yeast gene required for peroxisome biogenesis, encodes a member of a novel family of putative ATPases. Cell 64(3):499–510
Fagarasanu A, Mast FD, Knoblach B, Rachubinski RA (2010) Molecular mechanism of organelle inheritance: lessons from peroxisomes in yeast. Nat Rev Mol Cell Biol 11(9):644–654
Fanelli F, Sepe S, D’Amelio M, Bernardi C, Cristiano L, Cimini A, Cecconi F, Ceru’ MP, Moreno S (2013) Age-dependent roles of peroxisomes in the hippocampus of a transgenic mouse model of Alzheimer’s disease. Mol Neurodegener 8 (8):doi:10.1186/1750-1326-1188-1188
Fenner BJ, Scannell M, Prehn JH (2009) Identification of polyubiquitin binding proteins involved in NF-kappaB signaling using protein arrays. Biochim Biophys Acta 1794(7):1010–1016
Fransen M, Nordgren M, Wang B, Apanasets O (2012) Role of peroxisomes in ROS/RNS-metabolism: Implications for human disease. Biochim Biophys Acta 1822(9):1363–1373
Freemont PS, Hanson IM, Trowsdale J (1991) A novel cysteine-rich sequence motif. Cell 64(3):483–484
Fujiki Y, Nashiro C, Miyata N, Tamura S, Okumoto K (2012) New insights into dynamic and functional assembly of the AAA peroxins, Pex1p and Pex6p, and their membrane receptor Pex26p in shuttling of PTS1-receptor Pex5p during peroxisome biogenesis. Biochim Biophys Acta 1823(1):145–149
Furuki S, Tamura S, Matsumoto N, Miyata N, Moser A, Moser HW, Fujiki Y (2006) Mutations in the peroxin Pex26p responsible for peroxisome biogenesis disorders of complementation group 8 impair its stability, peroxisomal localization, and interaction with the Pex1p x Pex6p complex. J Biol Chem 281(3):1317–1323
Gabaldon T, Snel B, van Zimmeren F, Hemrika W, Tabak H, Huynen MA (2006) Origin and evolution of the peroxisomal proteome. Biol Direct 1:8
Geisbrecht BV, Collins CS, Reuber BE, Gould SJ (1998) Disruption of a PEX1-PEX6 interaction is the most common cause of the neurologic disorders Zellweger syndrome, neonatal adrenoleukodystrophy, and infantile Refsum disease. Proc Natl Acad Sci U S A 95(15):8630–8635
Girzalsky W, Platta HW, Erdmann R (2009) Protein transport across the peroxisomal membrane. Biol Chem 390(8):745–751
Gonen H, Bercovich B, Orian A, Carrano A, Takizawa C, Yamanaka K, Pagano M, Iwai K, Ciechanover A (1999) Identification of the ubiquitin carrier proteins, E2s, involved in signal-induced conjugation and subsequent degradation of IkappaBalpha. J Biol Chem 274(21):14823–14830
Goto S, Mano S, Nakamori C, Nishimura M (2011) Arabidopsis ABERRANT PEROXISOME MORPHOLOGY9 is a peroxin that recruits the PEX1-PEX6 complex to peroxisomes. Plant Cell 23(4):1573–1587
Gouveia AM, Guimaraes CP, Oliveira ME, Reguenga C, Sa-Miranda C, Azevedo JE (2003) Characterization of the peroxisomal cycling receptor Pex5p import pathway. Adv Exp Med Biol 544:213–220
Grimm I, Saffian D, Platta HW, Erdmann R (2012) The AAA-type ATPases Pex1p and Pex6p and their role in peroxisomal matrix protein import in Saccharomyces cerevisiae. Biochim Biophys Acta 1823(1):150–158
Grou CP, Carvalho AF, Pinto MP, Wiese S, Piechura H, Meyer HE, Warscheid B, Sa-Miranda C, Azevedo JE (2008) Members of the E2D (UbcH5) family mediate the ubiquitination of the conserved cysteine of Pex5p, the peroxisomal import receptor. J Biol Chem 283(21):14190–14197
Grou CP, Carvalho AF, Pinto MP, Alencastre IS, Rodrigues TA, Freitas MO, Francisco T, Sa-Miranda C, Azevedo JE (2009a) The peroxisomal protein import machinery–a case report of transient ubiquitination with a new flavor. Cell Mol Life Sci 66(2):254–262
Grou CP, Carvalho AF, Pinto MP, Huybrechts SJ, Sa-Miranda C, Fransen M, Azevedo JE (2009b) Properties of the ubiquitin-Pex5p thiol ester conjugate. J Biol Chem 284(16):10504–10513
Grou CP, Francisco T, Rodrigues TA, Freitas MO, Pinto MP, Carvalho AF, Domingues P, Wood SA, Rodríguez-Borges JE, Sá-Miranda C, Fransen M, Azevedo JE (2012) Identification of ubiquitin-specific protease 9X (USP9X) as a deubiquitinase acting on the ubiquitin-peroxin 5 (PEX5) thioester conjugate. J Biol Chem 287(16):12815–12827
Grunau S, Lay D, Mindthoff S, Platta HW, Girzalsky W, Just WW, Erdmann R (2011) The Phosphoinositide-3-kinase Vps34p is required for pexophagy in Saccharomyces cerevisiae. Biochem J 434:161–170
Gründlinger M, Yasmin S, Lechner BE, Geley S, Schrettl M, Hynes M, Haas H (2013) Fungal siderophore biosynthesis is partially localized in peroxisomes. Mol Microbiol 88(5):862–875
Hampton RY, Sommer T (2012) Finding the will and the way of ERAD substrate retrotranslocation. Curr Opin Cell Biol 24(4):460–466
Hasan S, Platta HW, Erdmann R (2013) Import of proteins into the peroxisomal matrix. Front Physiol 4:261
Hazra PP, Suriapranata I, Snyder WB, Subramani S (2002) Peroxisome remnants in pex3Delta cells and the requirement of Pex3p for interactions between the peroxisomal docking and translocation subcomplexes. Traffic 3(8):560–574
Hensel A, Beck S, El Magraoui F, Platta HW, Girzalsky W, Erdmann R (2011) Cysteine-dependent ubiquitination of Pex18p is linked to cargo translocation across the peroxisomal membrane. J Biol Chem 286:43495–43505
Imanaka T, Small GM, Lazarow PB (1987) Translocation of acyl-CoA oxidase into peroxisomes requires ATP hydrolysis but not a membrane potential. J Cell Biol 105:2915–2922
Ishikura S, Weissman AM, Bonifacino JS (2010) Serine residues in the cytosolic tail of the T-cell antigen receptor alpha-chain mediate ubiquitination and endoplasmic reticulum-associated degradation of the unassembled protein. J Biol Chem 285(31):23916–23924
Islinger M, Grille S, Fahimi HD, Schrader M (2012) The peroxisome: an update on mysteries. Histochem Cell Biol 137(5):547–574
Ivashchenko O, Van Veldhoven PP, Brees C, Ho YS, Terlecky SR, Fransen M (2011) Intraperoxisomal redox balance in mammalian cells: oxidative stress and interorganellar cross-talk. Mol Biol Cell 22(9):1440–1451
Iyer LM, Leipe DD, Koonin EV, Aravind L (2004) Evolutionary history and higher order classification of AAA + ATPases. J Struct Biol 146(1–2):11–31
Joo HJ, Kim KY, Yim YH, Jin YX, Kim H, Kim MY, Paik YK (2010) Contribution of the peroxisomal acox gene to the dynamic balance of daumone production in Caenorhabditis elegans. J Biol Chem 285(38):29319–29325
Jungwirth H, Ring J, Mayer T, Schauer A, Buttner S, Eisenberg T, Carmona-Gutierrez D, Kuchler K, Madeo F (2008) Loss of peroxisome function triggers necrosis. FEBS Lett 582(19):2882–2886
Kaur N, Zhao Q, Xie Q, Hu J (2013) Arabidopsis RING peroxins are E3 ubiquitin ligases that interact with two homologous ubiquitin receptor proteins(F). J Integr Plant Biol 55(1):108–120
Kee Y, Huibregtse JM (2007) Regulation of catalytic activities of HECT ubiquitin ligases. Biochem Biophys Res Commun 354(2):329–333
Kerscher O, Felberbaum R, Hochstrasser M (2006) Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu Rev Cell Dev Biol 22:159–180
Kerssen D, Hambruch E, Klaas W, Platta HW, de Kruijff B, Erdmann R, Kunau WH, Schliebs W (2006) Membrane association of the cycling peroxisome import receptor Pex5p. J Biol Chem 281(37):27003–27015
Kiel JA, Hilbrands RE, van der Klei IJ, Rasmussen SW, Salomons FA, van der Heide M, Faber KN, Cregg JM, Veenhuis M (1999) Hansenula polymorpha Pex1p and Pex6p are peroxisome-associated AAA proteins that functionally and physically interact. Yeast 15(11):1059–1078
Kiel JA, Hilbrands RE, Bovenberg RA, Veenhuis M (2000) Isolation of Penicillium chrysogenum PEX1 and PEX6 encoding AAA proteins involved in peroxisome biogenesis. Appl Microbiol Biotechnol 54(2):238–242
Kiel JA, Emmrich K, Meyer HE, Kunau WH (2005a) Ubiquitination of the peroxisomal targeting signal type 1 receptor, Pex5p, suggests the presence of a quality control mechanism during peroxisomal matrix protein import. J Biol Chem 280(3):1921–1930
Kiel JA, Otzen M, Veenhuis M, van der Klei IJ (2005b) Obstruction of polyubiquitination affects PTS1 peroxisomal matrix protein import. Biochim Biophys Acta 1745(2):176–186
Kiel JA, Veenhuis M, van der Klei IJ (2006) PEX genes in fungal genomes: common, rare or redundant. Traffic 7(10):1291–1303
Kienow L, Schneider K, Bartsch M, Stuible HP, Weng H, Miersch O, Wasternack C, Kombrink E (2008) Jasmonates meet fatty acids: functional analysis of a new acyl-coenzyme A synthetase family from Arabidopsis thaliana. J Exp Bot 59(2):403–419
Koellensperger G, Daubert S, Erdmann R, Hann S, Rottensteiner H (2007) Characterisation of zinc-binding domains of peroxisomal RING finger proteins using size exclusion chromatography/inductively coupled plasma-mass spectrometry. Biol Chem 388(11):1209–1214
Koller A, Snyder WB, Faber KN, Wenzel TJ, Rangell L, Keller GA, Subramani S (1999) Pex22p of Pichia pastoris, essential for peroxisomal matrix protein import, anchors the ubiquitin-conjugating enzyme, Pex4p, on the peroxisomal membrane. J Cell Biol 146(1):99–112
Kou J, Kovacs GG, Höftberger R, Kulik W, Brodde A, Forss-Petter S, Hönigschnabl S, Gleiss A, Brügger B, Wanders R, Just W, Budka H, Jungwirth S, Fischer P, Berger J (2011) Peroxisomal alterations in Alzheimer’s disease. Acta Neuropathol 122(3):271–283
Kragt A, Voorn-Brouwer T, van den Berg M, Distel B (2005) The Saccharomyces cerevisiae peroxisomal import receptor Pex5p is monoubiquitinated in wild type cells. J Biol Chem 280(9):7867–7874
Krazy H, Michels PA (2006) Identification and characterization of three peroxins–PEX6, PEX10 and PEX12–involved in glycosome biogenesis in Trypanosoma brucei. Biochim Biophys Acta 1763(1):6–17
Kunau W-H, Beyer A, Franken T, Götte K, Marzioch M, Saidowsky J, Skaletz-Rorowski A, Wiebel FF (1993) Two complementary approaches to study peroxisome biogenesis in Saccharomyces cerevisiae: Forward and reversed genetics. Biochimie 75:209–224
Lazarow PB, DeDuve C (1976) A fatty acyl-CoA oxidazing system in rat liver peroxisomes; enhancement by clofibrate, a hypolipidemic drug. Proc Natl Acad Sci U S A 73:2043–2046
Leon S, Subramani S (2007) A conserved cysteine residue of Pichia pastoris Pex20p is essential for its recycling from the peroxisome to the cytosol. J Biol Chem 282(10):7424–7430
Leon S, Goodman JM, Subramani S (2006a) Uniqueness of the mechanism of protein import into the peroxisome matrix: transport of folded, co-factor-bound and oligomeric proteins by shuttling receptors. Biochim Biophys Acta 1763(12):1552–1564
Leon S, Zhang L, McDonald WH, Yates J 3rd, Cregg JM, Subramani S (2006b) Dynamics of the peroxisomal import cycle of PpPex20p: ubiquitin-dependent localization and regulation. J Cell Biol 172(1):67–78
Liu X, Subramani S (2013) Unique requirements for mono- and polyubiquitination of the peroxisomal targeting signal co-receptor, Pex20. J Biol Chem 288(10):7230–7240
Liu X, Ma C, Subramani S (2012) Recent advances in peroxisomal matrix protein import. Curr Opin Cell Biol 24(4):484–489
Lizard G, Rouaud O, Demarquoy J, Cherkaoui-Malki M, Iuliano L (2012) Potential roles of peroxisomes in Alzheimer’s disease and in dementia of the Alzheimer’s type. J Alzheimers Dis 29(2):241–254
Matsumoto N, Tamura S, Fujiki Y (2003a) The pathogenic peroxin Pex26p recruits the Pex1p-Pex6p AAA ATPase complexes to peroxisomes. Nat Cell Biol 5(5):454–460
Matsumoto N, Tamura S, Furuki S, Miyata N, Moser A, Shimozawa N, Moser HW, Suzuki Y, Kondo N, Fujiki Y (2003b) Mutations in novel peroxin gene PEX26 that cause peroxisome-biogenesis disorders of complementation group 8 provide a genotype-phenotype correlation. Am J Hum Genet 73(2):233–246
Meijer WH, Gidijala L, Fekken S, Kiel JA, van den Berg MA, Lascaris R, Bovenberg RA, van der Klei IJ (2010) Peroxisomes are required for efficient penicillin biosynthesis in Penicillium chrysogenum. Appl Environ Micobiol 76(17):5702–5709
Miyata N, Fujiki Y (2005) Shuttling mechanism of peroxisome targeting signal type 1 receptor Pex5: ATP-independent import and ATP-dependent export. Mol Cell Biol 25(24):10822–10832
Miyata N, Hosoi K, Mukai S, Fujiki Y (2009) In vitro import of peroxisome-targeting signal type 2 (PTS2) receptor Pex7p into peroxisomes. Biochim Biophys Acta 1793(5):860–870
Miyata N, Okumoto K, Mukai S, Noguchi M, Fujiki Y (2012) AWP1/ZFAND6 Functions in Pex5 Export by Interacting with Cys-monoubiquitinated Pex5 and Pex6 AAA ATPase. Traffic 13(1):168–183
Müller WH, van der Krift TP, Krouwer AJ, Wösten HA, van der Voort LH, Smaal EB, Verkleij AJ (1991) Localization of the pathway of the penicillin biosynthesis in Penicillium chrysogenum. EMBO J 10(2):489–495
Nagotu S, Kalel VC, Erdmann R, Platta HW (2012) Molecular basis of peroxisomal biogenesis disorders caused by defects in peroxisomal matrix protein import. Biochim Biophys Acta 1822(9):1326–1336
Nashiro C, Kashiwagi A, Matsuzaki T, Tamura S, Fujiki Y (2011) Recruiting Mechanism of the AAA peroxins, Pex1p and Pex6p, to Pex26p on Peroxisome Membrane. Traffic 12(6):774–788
Neuwald AF, Aravind L, Spouge JL, Koonin EV (1999) AAA+: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res 9(1):27–43
Nito K, Kamigaki A, Kondo M, Hayashi M, Nishimura M (2007) Functional classification of Arabidopsis peroxisome biogenesis factors proposed from analyses of knockdown mutants. Plant Cell Physiol 48(6):763–774
Novikoff AB, Novikoff PM, Davis C, Quintana N (1973) Studies on microperoxisomes. V are microperoxisomes ubiquitous in mammalian cells? J Histochem Cytochem 21(8):737–755
Oeljeklaus S, Reinartz BS, Wolf J, Wiese S, Tonillo J, Podwojski K, Kuhlmann K, Stephan C, Meyer HE, Schliebs W, Brocard C, Erdmann R, Warscheid B (2012) Identification of core components and transient interactors of the peroxisomal importomer by dual-track stable isotope labeling with amino acids in cell culture analysis. J Proteome Res 11(4):2567–2580
Okumoto K, Abe I, Fujiki Y (2000) Molecular anatomy of the peroxin Pex12p: RING finger domain is essential for the Pex12p function and interacts with the peroxisome targeting signal type 1-receptor Pex5p and a RING peroxin, Pex10p. J Biol Chem 275(33):25700–25710
Okumoto K, Misono S, Miyata N, Matsumoto Y, Mukai S, Fujiki Y (2011) Cysteine ubiquitination of PTS1 receptor Pex5p regulates Pex5p recycling. Traffic 12(8):1067–1083
Oliveira ME, Gouveia AM, Pinto RA, Sa-Miranda C, Azevedo JE (2003) The energetics of Pex5p-mediated peroxisomal protein import. J Biol Chem 278(41):39483–39488
Opaliński L, Veenhuis M, van der Klei IJ (2011) Peroxisomes: membrane events accompanying peroxisome proliferation. Int J Biochem Cell Biol 43(6):847–851
Otzen M, Wang D, Lunenborg MG, van der Klei IJ (2005) Hansenula polymorpha Pex20p is an oligomer that binds the peroxisomal targeting signal 2 (PTS2). J Cell Sci 118(Pt 15):3409–3418
Palmer T, Berks BC (2012) The twin-arginine translocation (Tat) protein export pathway. Nat Rev Microbiol 10(7):483–964
Park S, Isaacson R, Kim HT, Silver PA, Wagner G (2005) Ufd1 exhibits the AAA-ATPase fold with two distinct ubiquitin interaction sites. Structure 13(7):995–1005
Peraza-Reyes L, Zickler D, Berteaux-Lecellier V (2008) The peroxisome RING-finger complex is required for meiocyte formation in the fungus Podospora anserina. Traffic 9(11):1998–2009
Platta HW, Erdmann R (2007a) Peroxisomal dynamics. Trends Cell Biol 17(10):474–484
Platta HW, Erdmann R (2007b) The peroxisomal protein import machinery. FEBS Lett 581(15):2811–2819
Platta HW, Girzalsky W, Erdmann R (2004) Ubiquitination of the peroxisomal import receptor Pex5p. Biochem J 384(Pt 1):37–45
Platta HW, Grunau S, Rosenkranz K, Girzalsky W, Erdmann R (2005) Functional role of the AAA peroxins in dislocation of the cycling PTS1 receptor back to the cytosol. Nat Cell Biol 7(8):817–822
Platta HW, El Magraoui F, Schlee D, Grunau S, Girzalsky W, Erdmann R (2007a) Ubiquitination of the peroxisomal import receptor Pex5p is required for its recycling. J Cell Biol 177(2):197–204
Platta HW, Thoms S, Kunau WH, Erdmann R (2007b) Function of the ubiquitin-conjugating enzyme Pex4p and the AAA peroxins Pex1p and Pex6p in peroxisomal protein transport, vol 25. The Enzymes Elsevier Academic Press, Amsterdam
Platta HW, Debelyy MO, El Magraoui F, Erdmann R (2008) The AAA peroxins Pex1p and Pex6p function as dislocases for the ubiquitinated peroxisomal import receptor Pex5p. Biochem Soc Trans 36:99–104
Platta HW, El Magraoui F, Baumer BE, Schlee D, Girzalsky W, Erdmann R (2009) Pex2 and Pex12 function as protein-ubiquitin ligases in peroxisomal protein import. Mol Cell Biol 29(20):5505–5516
Platta HW, Hagen S, Erdmann R (2013) The exportomer: the peroxisomal receptor export machinery. Cell Mol Life Sci 70(8):1393–1411
Prestele J, Hierl G, Scherling C, Hetkamp S, Schwechheimer C, Isono E, Weckwerth W, Wanner G, Gietl C (2010) Different functions of the C3HC4 zinc RING finger peroxins PEX10, PEX2, and PEX12 in peroxisome formation and matrix protein import. Proc Natl Acad Sci U S A 107(33):14915–14920
Purdue PE, Lazarow PB (2001) Pex18p is constitutively degraded during peroxisome biogenesis. J Biol Chem 276(50):47684–47689
Ratzel SE, Lingard MJ, Woodward AW, Bartel B (2011) Reducing PEX13 expression ameliorates physiological defects of late-acting peroxin mutants. Traffic 12(1):121–134
Ravid T, Hochstrasser M (2008) Diversity of degradation signals in the ubiquitin-proteasome system. Nat Rev Mol Cell Biol 9(9):679–690
Rosenkranz K, Birschmann I, Grunau S, Girzalsky W, Kunau WH, Erdmann R (2006) Functional association of the AAA complex and the peroxisomal importomer. FEBS J 273(16):3804–3815
Saffian D, Grimm I, Girzalsky W, Erdmann R (2012) ATP-dependent assembly of the heteromeric Pex1p-Pex6p-complex of the peroxisomal matrix protein import machinery. J Struct Biol 179(2):126–132
Saville MK, Sparks A, Xirodimas DP, Wardrop J, Stevenson LF, Bourdon JC, Woods YL, Lane DP (2004) Regulation of p53 by the ubiquitin-conjugating enzymes UbcH5B/C in vivo. J Biol Chem 279(40):42169–42181
Schliebs W, Girzalsky W, Erdmann R (2010) Peroxisomal protein import and ERAD: variations on a common theme. Nat Rev Mol Cell Biol 11(12):885–890
Schluter A, Fourcade S, Ripp R, Mandel JL, Poch O, Pujol A (2006) The evolutionary origin of peroxisomes: an ER-peroxisome connection. Mol Biol Evol 23(4):838–845
Schumann U, Prestele J, O’Geen H, Brueggeman R, Wanner G, Gietl C (2007) Requirement of the C3HC4 zinc RING finger of the Arabidopsis PEX10 for photorespiration and leaf peroxisome contact with chloroplasts. Proc Natl Acad Sci U S A 104(3):1069–1074
Seo JG, Lai CY, Miceli MV, Jazwinski SM (2007) A novel role of peroxin PEX6: suppression of aging defects in mitochondria. Aging Cell 6(3):405–413
Seufert W, Jentsch S (1990) Ubiquitin-conjugating enzymes UBC4 and UBC5 mediate selective degradation of short-lived and abnormal proteins. EMBO J 9(2):543–550
Seufert W, McGrath JP, Jentsch S (1990) UBC1 encodes a novel member of an essential subfamily of yeast ubiquitin-conjugating enzymes involved in protein degradation. EMBO J 9(13):4535–3541
Shimizu Y, Okuda-Shimizu Y, Hendershot LM (2010) Ubiquitylation of an ERAD substrate occurs on multiple types of amino acids. Mol Cell 40(6):917–926
Shimozawa N, Tsukamoto T, Suzuki Y, Orii T, Shirayoshi Y, Mori T, Fujiki Y (1992) A human gene responsible for Zellweger syndrome that affects peroxisome assembly. Science 255(5048):1132–1134
Shiozawa K, Maita N, Tomii K, Seto A, Goda N, Akiyama Y, Shimizu T, Shirakawa M, Hiroaki H (2004) Structure of the N-terminal domain of PEX1 AAA-ATPase. Characterization of a putative adaptor-binding domain. J Biol Chem 279(48):50060–50068
Sparkes IA, Brandizzi F, Slocombe SP, El-Shami M, Hawes C, Baker A (2003) An arabidopsis pex10 null mutant is embryo lethal, implicating peroxisomes in an essential role during plant embryogenesis. Plant Physiol 133(4):1809–1819
Spiegel CN, Batista-Pereira LG, Bretas JA, Eiras AE, Hooper AM, Peixoto AA, Soares MJ (2011) Pheromone gland development and pheromone production in lutzomyia longipalpis (Diptera: Psychodidae: Phlebotominae). J Med Entomol 48(3):489–495
Steinberg SJ, Dodt G, Raymond GV, Braverman NE, Moser AB, Moser HW (2006) Peroxisome biogenesis disorders. Biochim Biophys Acta 1763(12):1733–1748
Stinson BM, Nager AR, Glynn SE, Schmitz KR, Baker TA, Sauer RT (2013) Nucleotide binding and conformational switching in the hexameric ring of a AAA + machine. Cell 153(3):628–639
Tamura S, Shimozawa N, Suzuki Y, Tsukamoto T, Osumi T, Fujiki Y (1998) A cytoplasmic AAA family peroxin, Pex1p, interacts with Pex6p. Biochem Biophys Res Commun 245(3):883–886
Tamura S, Yasutake S, Matsumoto N, Fujiki Y (2006) Dynamic and functional assembly of the AAA peroxins, Pex1p and Pex6p, and their membrane receptor Pex26p. J Biol Chem 281(38):27693–27704
Taylor EB, Rutter J (2011) Mitochondrial quality control by the ubiquitin-proteasome system. Biochem Soc Trans 39(5):1509–1513
Theodoulou FL, Bernhardt K, Linka N, Baker A (2013) Peroxisome membrane proteins: multiple trafficking routes and multiple functions? Biochem J 451(3):345–352
Till A, Lakhani R, Burnett SF, Subramani S (2012) Pexophagy: the selective degradation of peroxisomes. Int J Cell Biol 2012:512721
Titorenko VI, Rachubinski RA (2000) Peroxisomal membrane fusion requires two AAA family ATPases, Pex1p and Pex6p. J Cell Biol 150(4):881–886
Tower RJ, Fagarasanu A, Aitchison JD, Rachubinski RA (2011) The Peroxin Pex34p functions with the Pex11 family of peroxisomal divisional proteins to regulate the peroxisome population in yeast. Mol Biol Cell 22(10):1727–1738
Tsukamoto T, Miura S, Fujiki Y (1991) Restoration by a 35 K membrane protein of peroxisome assembly in a peroxisome-deficient mammalian cell mutant. Nature 350(6313):77–81
van der Klei IJ, Hilbrands RE, Kiel JAKW, Rasmussen SW, Cregg JM, Veenhuis M (1998) The ubiquitin-conjugating enzyme Pex4p of Hansenula polymorpha is required for efficient functioning of the PTS1 import machinery. EMBO J 17(13):3608–3618
van der Zand A, Gent J, Braakman I, Tabak HF (2012) Biochemically distinct vesicles from the endoplasmic reticulum fuse to form peroxisomes. Cell 149(2):397–409
Walker JE, Saraste M, Runswick MJ, Gay NJ (1982) Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. Embo J 1(8):945–951
Wanders RJ, Waterham HR (2006a) Biochemistry of mammalian peroxisomes revisited. Annu Rev Biochem 75:295–332
Wanders RJ, Waterham HR (2006b) Peroxisomal disorders: the single peroxisomal enzyme deficiencies. Biochim Biophys Acta 1763(12):1707–1720
Wang X, Herr RA, Chua WJ, Lybarger L, Wiertz EJ, Hansen TH (2007) Ubiquitination of serine, threonine, or lysine residues on the cytoplasmic tail can induce ERAD of MHC-I by viral E3 ligase mK3. J Cell Biol 177(4):613–624
Wang X, Herr RA, Hansen TH (2012) Ubiquitination of substrates by esterification. Traffic 13(1):19–24
Warner DR, Roberts EA, Greene RM, Pisano MM (2003) Identification of novel Smad binding proteins. Biochem Biophys Res Commun 312(4):1185–1190
Waterham HR, Ebberink MS (2012) Genetics and molecular basis of human peroxisome biogenesis disorders. Biochim Biophys Acta 1822(9):1430–1441
Wendler P, Ciniawsky S, Kock M, Kube S (2012) Structure and function of the AAA + nucleotide binding pocket. Biochim Biophys Acta 1823(1):2–14
Wenzel DM, Lissounov A, Brzovic PS, Klevit RE (2011) UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids. Nature 474:105–108
Widhalm JR, Ducluzeau AL, Buller NE, Elowsky CG, Olsen LJ, Basset GJ (2012) Phylloquinone (vitamin K(1) ) biosynthesis in plants: two peroxisomal thioesterases of lactobacillales origin hydrolyze 1,4-dihydroxy-2-naphthoyl-Coa. Plant J 71(2):205–215
Wiebel FF, Kunau W-H (1992) The PAS2 protein essential for peroxisome biogenesis is related to ubiquitin-conjugating enzymes. Nature 359(6390):73–76
Williams C, van den Berg M, Sprenger RR, Distel B (2007) A conserved cysteine is essential for Pex4p-dependent ubiquitination of the peroxisomal import receptor Pex5p. J Biol Chem 282(31):22534–22543
Williams C, van den Berg M, Geers E, Distel B (2008) Pex10p functions as an E3 ligase for the Ubc4p-dependent ubiquitination of Pex5p. Biochem Biophys Res Commun 374(4):620–624
Williams C, van den Berg M, Panjikar S, Stanley WA, Distel B, Wilmanns M (2012) Insights into ubiquitin-conjugating enzyme/ co-activator interactions from the structure of the Pex4p:Pex22p complex. EMBO J 31(2):391–402
Zolman BK, Bartel B (2004) An arabidopsis indole-3-butyric acid-response mutant defective in PEROXIN6, an apparent ATPase implicated in peroxisomal function. Proc Natl Acad Sci U S A 101(6):1786–1791
Zolman BK, Monroe-Augustus M, Silva ID, Bartel B (2005) Identification and functional characterization of arabidopsis PEROXIN4 and the interacting protein PEROXIN22. Plant Cell 17(12):3422–3435
Acknowledgements
We apologize to all the scientists whose work could not be cited due to space limitations. This work was supported by grants of the Deutsche Forschungsgemeinschaft (SFB 642 and FOR 1905) to RE and HWP.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Wien
About this chapter
Cite this chapter
Platta, H.W., Hagen, S., Erdmann, R. (2014). The Peroxisomal Exportomer. In: Brocard, C., Hartig, A. (eds) Molecular Machines Involved in Peroxisome Biogenesis and Maintenance. Springer, Vienna. https://doi.org/10.1007/978-3-7091-1788-0_15
Download citation
DOI: https://doi.org/10.1007/978-3-7091-1788-0_15
Published:
Publisher Name: Springer, Vienna
Print ISBN: 978-3-7091-1787-3
Online ISBN: 978-3-7091-1788-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)