Abstract
After decades of intense investigation, the molecular components of mammalian hair-cell mechanotransduction (MET) machinery have started to emerge. Convincing evidences suggested that tip links are composed of two atypical cadherin proteins, protocadherin 15 (PCDH15) and cadherin 23 (CDH23). Meanwhile, the identity of the MET channel is still not confirmative, although several promising candidates have been put forward. In this chapter, we will first introduce the recent progress of our understanding of tip links, as well as the so-called upper and lower tip-link complexes associated with them. Then we will focus on the MET channel that lies at the heart of the MET machinery. TMC1, TMC2, LHFPL5, TMIE, and CIB2 have been suggested to be integral components of the machinery, but confirmative evidences for them as the pore-forming subunits of the channel are still missing. Lastly, we will briefly discuss the recent identification of PIEZO2 as the channel responsible for the reverse-polarity MET currents.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Tip-Link Proteins
Tip links play a critical role in hair-cell MET. The molecular composition of tip links had been the subject of intense debate for decades, until two atypical cadherins, cadherin 23 (CDH23) and protocadherin 15 (PCDH15), were identified as tip-link components [1,2,3]. CDH23 and PCDH15 bind to each other via their extracellular N-terminal ends and make up the upper part and the lower part of tip links, respectively. Their cytoplasmic ends interact with other stereociliary proteins and form the so-called upper tip-link complex and lower tip-link complex.
Mutations of CDH23 and PCDH15 genes cause nonsyndromic autosomal recessive deafness as well as syndromic deafness (Usher syndrome type I) [4,5,6,7,8]. Usher syndrome (USH) is an autosomal recessive genetic disease, characterized by the association of hearing loss with retinitis pigmentosa and occasional balance problems. USH is classified into three types based on the severity of clinical symptoms. In terms of hearing phenotype, USH type I (USH1) patients suffer from congenital severe-to-profound deafness; USH2 patients suffer from congenital moderate-to-severe hearing loss; USH3 patients, the least severe ones, show progressive hearing loss. Ten USH proteins have been identified so far, including five USH1 proteins, three USH2 proteins, one USH3 protein, and one USH modifier protein [9] (Table 4.1). All known USH proteins could be detected in the hair-cell stereocilia and are able to interact with each other and form protein complexes [21, 22]. USH proteins are required for the development and function of hair bundles, and some of them, especially CDH23 and PCDH15, play an important role in MET.
4.1.1 CDH23 and Upper Tip-Link Complex
4.1.1.1 CDH23 Gene and Protein
CDH23 is a large single transmembrane protein, which belongs to atypical cadherin. It consists of multiple extracellular cadherin (EC) repeats, a single transmembrane fragment, and a short cytoplasmic domain (Fig. 4.1). Cdh23 gene contains 69 exons, and transcription starting from different initiation sites gives rise to various transcripts encoding proteins with different numbers of EC repeats [23]. Typical EC repeat is composed of ~110 amino acids that form 7 tightly packed β-strands and adopts a “Greek key” topology that spans approximately 4 nm [24]. The longest CDH23 isoform has 27 EC repeats and hence could span more than 100 nm. In classical cadherins, the linker region between EC repeats binds Ca2+ ions, which makes it very rigid [24]. Molecular dynamics simulations suggested that the EC repeats of CDH23 are also quite stiff [25].
The cytoplasmic part of CDH23 bears no homology with that of classical cadherins. As a result, it does not interact with β-catenin as classical cadherins do. CDH23 contains a type-I PDZ domain-binding interface (PBI) at its C-terminal end, which mediates the interaction with PDZ domains [26,27,28,29]. Exon 68 of Cdh23 gene encodes part of the cytoplasmic domain and is subjected to alternative splicing [30]. The resulting Cdh23 variants have different expression patterns: Cdh23(−68) is expressed broadly in various tissues, whereas Cdh23(+68) is preferentially expressed in the inner ear [1, 28]. The biological significance of the inclusion or exclusion of exon 68 has not been fully understood yet, although it might affect the conformation of CDH23 and protein-protein interaction [27, 29, 31].
4.1.1.2 CDH23 Is an Upper Tip-Link Component
The first evidence of CDH23 as a tip-link component came from its localization near the tips of stereocilia [1]. Immunostaining with specific antibodies localized CDH23 on the stereocilia, at a position corresponding to the upper end of tip links [3]. CDH23 was also associated with transient lateral links as well as kinociliary links, both of which are important extracellular links during hair bundle development [32]. Moreover, the recombinant extracellular fragment of CDH23 formed coiled homodimers, in which the membrane-proximal C-terminus separated into two strands [3], just as the upper insertion site of tip links did [33]. CDH23 interacts with the lower tip-link component PCDH15 via their most N-terminal two EC repeats, forming a CDH23-PCDH15 heterotetramer that is about 180 nm long, consistent with the length of tip links [3, 34]. As mentioned in Chap. 2, tip links were disrupted by application of Ca2+ chelators such as EGTA, while exogenous application of CDH23 fragments encompassing the extracellular EC1 inhibited tip-link recovery from EGTA-induced disruption [35].
If CDH23 is a component of tip links, inactivation of Cdh23 gene in mice would cause tip-link disruption as well as MET deficit. Indeed, in waltzer mice, a Cdh23-null mutant mouse line, tip links were missing, and MET was affected, consistent with the role of CDH23 as a tip-link component [36]. However, in the Cdh23-null mice, the development of hair bundle was also severely affected, and the stereocilia were never developed into a mature morphology [37]; hence it’s difficult to separate the role of CDH23 in hair bundle development from tip-link formation and MET. The final conclusive proof came from a mutant mouse line named salsa that harbours a point mutation in Cdh23 gene [38]. In salsa mice, hair bundles were developed normally, and tip links were formed during early developmental stages and then started to disappear after postnatal day 10 (P10), accompanied by increased hearing threshold [38]. Similar phenotypes were also observed later in another mutant mouse line jera that contains different point mutation in Cdh23 gene [39].
4.1.1.3 CDH23 and Upper Tip-Link Complex
As mentioned in Chap. 2, the upper tip-link insertion sites on the lateral shaft of taller stereocilia display electron-dense plaques, referred as upper tip-link density (UTLD). Several proteins have been shown to interact with the cytoplasmic domain of CDH23 and participate in the formation of UTLD. USH1C protein harmonin is a scaffold protein containing three PDZ domains, of which PDZ2 binds the C-terminal PBI of CDH23 [26, 27]. Additionally, a highly conserved N-terminal fragment preceding PDZ1 of harmonin binds an internal fragment of CDH23 [40]. Immunostaining showed that harmonin localized at UTLD [41], and this localization was dependent on the presence of CDH23 [42]. Deletion of the coiled-coil and PST domains between harmonin PDZ2 and PDZ3 in deaf circler (dfcr) mice prevented the formation of UTLD, suggesting that harmonin plays a central role in UTLD [41]. Harmonin could also bind F-actin; hence it might connect tip links to the cytoskeleton [26].
Two other USH1 proteins, USH1B protein myosin 7a (MYO7A) and USH1G protein SANS, also bind the cytoplasmic domain of CDH23 [42, 43]. Both MYO7A and SANS localized at the UTLD, whereas in Cdh23-null mutant mice, their stereociliary localization was altered or completely absent [42, 44]. Myo7a mutation in mice resulted in MET deficits and altered adaptation [45], while sans inactivation in mice led to loss of tip links and MET deficits [43]. Taken together, CDH23, harmonin, MYO7A, and SANS interact with each other and form the core of upper tip-link complex.
4.1.1.4 Upper Tip-Link Complex and Slow Adaptation
Slow adaptation occurs on a timescale of several milliseconds. According to the commonly accepted adaptation motor model, slow adaptation is mediated by a motor that sets the MET channel operating point [46, 47]. The motor is associated with tip links as well as the F-actin core near the upper tip-link insertion site and hence regulates the tension of tip links and sets the operating point of the MET channels. In the resting state, the motor climbs up along the stereocilia and hence sets the resting tension of the tip links. When the stereocilia are deflected in the positive direction, tension of tip links increases, and Ca2+ enters stereocilia through the MET channels. Ca2+ then diffuses to the upper tip-link insertion site and causes the motor to slide down the stereocilia through an unknown mechanism. This movement of motor decreases the tension of tip links and resets the MET channel operating point to the resting level.
Slow adaptation rates vary considerably between vestibular and cochlear hair cells, as well as among different species [46, 48,49,50,51]. Hence it is possible that the molecular mechanisms for slow adaptation in vestibular and cochlear hair cells in different species are different. At present, the most prominent candidate of adaptation motor is an unconventional myosin MYO1C. In frogs, MYO1C was concentrated at the UTLD [52, 53]. In mammals, MYO1C was shown to localize along the length of stereocilia [54], and it could interact with the upper tip-link component CDH23 [1]. The most convincing evidence came from a transgenic mouse model that expresses a mutant MYO1C Y61G. The substitution of tyrosine (Y) to glycine (G) in the nucleotide-binding pocket of MYO1C conferred susceptibility to inhibition by N6-modified ADP analogs such as NMB-ADP [55]. The MYO1C Y61G transgenic mice had normal MET in the absence of NMB-ADP; however, when NMB-ADP was present, slow adaptation in the vestibular hair cells was blocked [56]. Similar results were observed in MYO1C Y61G knockin mice, supporting an important role of MYO1C in slow adaptation [57].
The second candidate for the adaptation motor is MYO7A. As mentioned above, MYO7A is a component of the UTLD and could bind CDH23. Moreover, slow adaptation was affected in Myo7a mutant mice [45]. Similarly, another UTLD component harmonin also binds CDH23 and F-actin, and adaptation was significantly slowed in harmonin mutant mice dfcr [41]. Given their actin-binding ability, MYO1C, MYO7A, or harmonin could act as the adaptation motor. Alternatively, they might affect adaptation through regulating the localization or assembly of the adaptation motor.
4.1.2 PCDH15 and Lower Tip-Link Complex
4.1.2.1 PCDH15 Is a Lower Tip-Link Component
Similar to CDH23, PCDH15 is also an atypical cadherin. PCDH15 consists of multiple EC repeats, a single transmembrane fragment, and a short cytoplasmic domain (Fig. 4.1). Several lines of evidence supported PCDH15 as a component of tip links. Immunostaining with specific antibodies localized PCDH15 near the tips of shorter stereocilia, a position corresponding to the lower end of tip links [3]. PCDH15 interacts with the upper tip-link component CDH23 via their N-terminal two EC repeats, and the resulting PCDH15-CDH23 heterotetramer is about 180 nm long, consistent with the length of tip links [3, 34]. Exogenous application of the extracellular fragments of PCDH15 inhibited tip-link recovery from EGTA-induced disruption [35].
Further evidences came from Pcdh15-null or mutant mice. Tip links were substantially reduced in early postnatal cochlear hair cells of Pcdh15-null mutant mice Ames waltzer 3J (av3J) [36]. Comparatively, tip links were reduced to a less extent in early postnatal cochlear hair cells of Pcdh15-mutant mice av6J, in which part of PCDH15 EC9 is deleted [36]. MET currents were reduced in both av3J and av6J mice [36]. Pcdh15-mutant mice noddy harbours an isoleucine-to-asparagine (I108N) mutation in EC1 repeat, which impairs the interaction between PCDH15 and CDH23. In noddy mice, both bundle morphology and tip-link formation were disrupted, and MET in cochlear hair cells was also affected [58].
4.1.2.2 PCDH15-CD2 Is Indispensable for Tip-Link Formation
Pcdh15 gene contains 39 exons, and alternative splicing produces various PCDH15 isoforms with different numbers of EC repeats. The longest PCDH15 isoform contains 11 EC repeats. Different from classical cadherins, some linker regions between adjacent PCDH15 EC repeats are calcium-free or partially calcium-free, which might confer some elasticity to the otherwise rigid tip links [59, 60].
Furthermore, according to the alternative inclusion of exon 35, 38, or 39, Pcdh15 transcript variants are classified into three groups: Pcdh15-CD1, Pcdh15-CD2, and Pcdh15-CD3, respectively [2]. PCDH15-CD1, PCDH15-CD2, and PCDH15-CD3 differ in their C-terminal cytoplasmic domains except for a common region (CR) that is proximal to the transmembrane fragment. Nevertheless, they all contain a type-I PBI at their C-terminal end, which mediates interaction with PDZ domains [21, 61, 62]. The alternative inclusion of exon 26a gives rise to the fourth PCDH15 group without a transmembrane domain and hence might represent a secretory protein and is not likely involved in the formation of tip links.
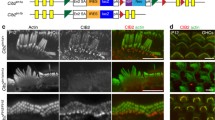
Expression of different PCDH15 isoforms was examined by performing immunostaining, which showed that they have different spatiotemporal expression patterns in the developing and mature inner ear [2]. In the developing organ of Corti, PCDH15-CD2 immunoreactivity was stronger at the apical turn, whereas PCDH15-CD1 and PCDH15-CD3 immunoreactivity was more intense at the basal turn. PCDH15-CD1 was distributed evenly along the length of the stereocilia except for the tips in mature cochlear hair cells. PCDH15-CD3 immunoreactivity detected with one antibody is restricted to the tips of the shorter stereocilia in immature IHCs and OHCs, but not present in the mature hair cells. Another antibody, however, detected PCDH15-CD3 at the tips of all the stereocilia in adult OHCs, but not in IHCs. In the same study, PCDH15-CD2 immunoreactivity was shown to decrease to an undetectable level in mature cochlear stereocilia [2]. However, another study showed that PCDH15-CD2 immunoreactivity localized at the tips of the three rows of stereocilia in both immature and mature IHCs and OHCs [63]. These discrepancies might result from variations in epitopes recognized by different antibodies or posttranslational modifications of the epitopes.
Knockout mice that lack each of the three PCDH15 isoforms (Pcdh15-∆CD1, Pcdh15-∆CD2, and Pcdh15-∆CD3) helped us to learn more about the role of each isoforms [64]. Hair bundle development and auditory function were normal in Pcdh15-∆CD1 and Pcdh15-∆CD3 mice. However, in Pcdh15-∆CD2 mice, kinociliary links were lost, and hair bundles were misorientated, which contributed to profound hearing loss. Nevertheless, tip links were still present in P1 hair cells in all three knockout mice, suggesting that none of the three PCDH15 isoforms is indispensable for tip-link formation in immature hair bundles. To circumvent the functional redundancy among these PCDH15 isoforms in tip-link formation during early development, conditional Pcdh15-CD2 knockout mice were developed so that the CD2-encoded exon 38 was deleted only in adult hair cells [63]. In the conditional Pcdh15-CD2 knockout mice, hair bundles were correctly oriented, and the auditory function was normal by P15. But by P30, the conditional Pcdh15-CD2 knockout mice were profoundly deaf, and tip links were almost completely lost, suggesting that PCDH15-CD2 is essential for tip-link formation in mature auditory hair cells. In line with this, MET was reduced when a dominant negative PCDH15-CD2 fragment was expressed in wild-type hair cells [65].
4.1.2.3 PCDH15 and Lower Tip-Link Complex
The lower tip-link insertion site at the tips of shorter stereocilia displays electron-dense plaques, referred as lower tip-link density (LTLD). Four transmembrane proteins have been reported to localize at the lower tip-link insertion site, which are TMC1, TMC2, LHFPL5, and TMIE. These transmembrane proteins interact with PCDH15 and are considered as integral components of MET machinery. These proteins will be discussed in more details in the following sessions.
LTLD might also involve cytosolic proteins that interact with PCDH15 cytoplasmic domain. SANS interacts with PCDH15-CD2 and PCDH15-CD3, but not PCDH15-CD1, and has been located at the tips of short and middle row stereocilia, close to LTLD [43]. Harmonin and MYO7A bind the cytoplasmic domain of PCDH15-CD1 [21, 61, 66]; however their localization at LTLD has not been reported.
4.2 Mechanotransduction Machinery
Identification of the MET machinery components, especially the pore-forming channels, has been a great challenge because of the scarcity of hair cells and the low expression levels of the channels and associated proteins. For example, in the mouse cochlea, there are about 15,000 hair cells with around 100 functional tip links within each hair cell, and only one or two functional channels are estimated to be associated with each tip link [67, 68]. Genetic studies have suggested several candidates for MET channels, most of which were disapproved later on. Recently, many lines of evidence suggested that four integral transmembrane proteins, TMC1, TMC2, LHFPL5, and TMIE, as well as a soluble protein CIB2, might form the core component of MET machinery. Especially, TMC1 and TMC2 were proposed as the pore-forming subunits of the MET channel. However, further investigation is needed before a final conclusion could be drawn.
4.2.1 TMC1/TMC2
4.2.1.1 TMC1/TMC2 Gene and Protein
Transmembrane channel-like 1 and 2 (TMC1 and TMC2) are homologous proteins with six predicted transmembrane domains and two additional hydrophobic domains that are not predicted to span the membrane [69]. TMC1 and TMC2 are members of TMC protein family that includes eight proteins in humans or mice [70, 71]. Mouse Tmc1 gene contains 24 exons, and exon 2 is subjected to alternative splicing [72]. The two Tmc1 variants use different translation initiation codons in exons 1 and 2, respectively, giving rise to TMC1 protein with different N-termini. Both splicing variants were detected in the mouse inner ear with similar expression levels [72].
In mammals, eight TMC proteins (TMC1 through 8) form a distinct protein family, sharing six-transmembrane domains as well as a highly conserved TMC motif with unknown function [70] (Fig. 4.1). The topology of TMC1 has been investigated by inserting HA epitope tag at different positions of heterogeneously expressed TMC1. The result showed that TMC1 is an integral membrane protein localized at ER, containing six-transmembrane domains and cytosolic N- and C-termini [73]. Heterogeneously expressed TMC1 or TMC2 is unsuccessful to be targeted to the plasma membrane at present, which hinders further functional examination of these proteins.
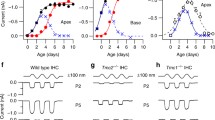
In situ hybridization revealed that Tmc1 and Tmc2 were expressed specifically in auditory and vestibular hair cells in the mouse inner ear [72]. This hair cell-specific expression of Tmc1/Tmc2 was further supported by RNA-seq results (SHIELD; https://shield.hms.harvard.edu) [74, 75]. The temporal expression profile of Tmc1/Tmc2 in the mouse inner ear was examined by quantitative RT-PCR (qPCR) analysis [72]. In the cochlea, Tmc2 expression started at P0–P2, around the same time that hair cells became mechanosensitive (P1–P3) [76]. Then Tmc2 expression declined at P3–P5, while Tmc1 expression started. In the adult mouse cochlea, Tmc2 expression dropped to an undetectable level, whereas Tmc1 expression persisted [72]. Both Tmc1 and Tmc2 were expressed in the vestibular system through development to adulthood [72].
4.2.1.2 TMC1/TMC2 Localizes Near the Lower End of Tip Links and Binds PCDH15
The subcellular localization of TMC1 and TMC2 in hair cells was examined using BAC-transgenic mice that express fluorescence protein-tagged TMC1 and TMC2. The result showed that both TMC1 and TMC2 were concentrated near the lower end of tip links [77], where the transduction channels were suggested to localize [68]. Similar results were obtained with antibodies against the endogenous TMC1 and TMC2 [77]. This localization puts TMC1/TMC2 at the centre of the MET machinery.
Moreover, immunoprecipitation experiments showed that heterogeneously expressed TMC1 and TMC2 interact with the lower tip-link component PCDH15, which further supported the role of TMC1/TMC2 as an integral component of MET machinery [78, 79]. However, caution is needed to be exercised in interpreting these results, since when expressed heterogeneously, TMC1/TMC2 and PCDH15 were distributed in different intracellular compartments.
4.2.1.3 TMC1 Disruption Causes Hearing Loss
TMC1 gene mutations are responsible for recessive and dominant nonsyndromic autosomal deafness, DFNB7/B11 and DFNA36, respectively [69]. Meanwhile, mutations of Tmc1 gene cause recessive hearing loss in deafness (dn) mice as well as semidominant hearing loss in Beethoven (Bth) mice [69, 80]. Dn mice were originally identified as spontaneous mutant mice that are congenitally deaf [81]. Dn mice contains an in-frame deletion of exon 14 (171 bp) of Tmc1 gene [69] and showed significant hair-cell loss by P30 [82]. Bth mice harbour a methionine-to-lysine (M412 K) point mutation in Tmc1 gene. In Bth/+ mice, progressive hair-cell degeneration started from P20, followed by progressive hearing loss from around P30. In Bth/Bth mice, hair-cell degeneration was much more severe, and Bth/Bth mice were congenitally deaf [80]. Mutations in human or mouse TMC2/Tmc2 gene have not been reported so far. Consistently, hearing threshold was normal in Tmc2 knockout mice [72].
4.2.1.4 TMC1/TMC2 Disruption Affects MET
Consistent with the fact that Tmc2 knockout mice had normal hearing threshold, the MET currents were normal in early postnatal auditory hair cells of Tmc2 knockout mice [72]. Surprisingly, despite the fact that homozygous dn and Bth mice were congenitally deaf, the MET currents were normal in early postnatal auditory hair cells of these Tmc1 mutant mice [82]. Similar result was obtained in Tmc1 knockout mice [72]. However, the MET currents were completely absent in Tmc1/Tmc2 double knockout mice; meanwhile, hair bundle morphology and tip links were unaffected [72, 83, 84]. The specific deficit of MET in Tmc1/Tmc2 double knockout mice suggested that TMC1 and TMC2 are important components of the MET machinery.
4.2.1.5 Transportation of TMC1/TMC2 to the Stereocilia Requires TOMT/LRTOMT
Transportation of TMC1/TMC2 to the tips of stereocilia is tightly regulated by other proteins, one of which has recently been identified as transmembrane O-methyltransferase (TOMT) [85, 86]. The human ortholog of Tomt gene is called leucine rich transmembrane and O-methyltransferase domain containing (LRTOMT), which has evolved from the fusion of TOMT gene with the neighbouring LRRC51 gene, and mutations in LRTMOT gene cause nonsyndromic recessive deafness DFNB63 [87, 88]. Similarly, mutations in mouse or zebrafish TOMT gene cause progressive degeneration of hair cells and profound hearing loss [85, 86, 88]. Further investigation revealed that MET currents were completely absent in TOMT-deficient mice and zebrafish [85, 86].
The subcellular localization of TOMT in mouse hair cells was examined by immunostaining, which showed that endogenous TOMT was distributed throughout the cytoplasm of hair cells, but not in the stereocilia [87]. Similar results were obtained when epitope-tagged TOMT/LRTOMT was expressed in transgenic zebrafish or cultured mouse cochlear sensory epithelium [85, 86]. Epitope-tagged TOMT/LRTOMT partially co-localized with the Golgi marker in hair cells of transgenic zebrafish, or ER marker in heterologous cells, suggesting a possible role in the regulation of membrane protein transportation [85, 86]. In line with this, epitope-tagged TOMT was co-immunoprecipitated with TMC1/2, LHFPL5, TMIE, and PCDH15-CD2 in vitro, and TMC1/2 was absent in the stereocilia in TOMT-deficient mice or zebrafish [85, 86]. Taken together, the present data suggested that TOMT plays an essential regulatory role in the transportation of TMC1/2 to the stereocilia and is indispensable for hair-cell MET. Meanwhile, despite of the important role of TOMT in TMC1/2 transportation, co-expression of TOMT did not alter the cytoplasmic localization of TMC1/2 in heterologous cells, suggesting that protein(s) other than TOMT is also necessary to target TMC1/2 to the plasma membrane [85].
4.2.1.6 Are TMC1/TMC2 the MET Channels?
As mentioned above, many lines of evidence are consistent with the hypothesis that TMC1/TMC2 are MET channels of mammalian auditory hair cells. First, TMC1 and TMC2 are expressed during the right time and at the right place. In the mouse cochlea, hair cells become mechanosensitive at P1–P3, whereas Tmc2 expression starts at P0–P2, followed by expression of Tmc1. In the mouse cochlea, Tmc1 and Tmc2 are exclusively expressed in hair cells. Within hair cells, TMC1 and TMC2 were shown to localize near the tips of shorter stereocilia, coincident with the localization of MET channels. Second, heterologously expressed TMC1/TMC2 could be co-immunoprecipitated together with the lower tip-link component PCDH15, which also localizes at the tips of shorter stereocilia. Third, Tmc1/Tmc2 double knockout completely eliminates the MET currents in mouse hair cells while leaving hair bundle and tip links unaffected.
At present the role of TMC1/TMC2 as MET channels is under hot debate [89,90,91]. Other proteins such as LHFPL5 and TMIE behave similarly to TMC1/TMC2: they localize at the tips of shorter stereocilia and interact with PCDH15, and their mutations lead to deafness as well as loss of MET current (discussed below). Probably the biggest challenge comes from the fact that TMC1 or TMC2 so far has not been successfully shown to serve as a channel in a heterologous expression system. Heterogeneously expressed TMC1 localizes at the ER, not on the plasma membrane [73], which hinders further examination of its potential channel activity. Heterologously expressed TMC-1, one of the two TMCs that exist in C. elegans, was reported to have sodium channel activity [92], whereas similar observation has not been reported on mammalian TMCs yet.
4.2.2 LHFPL5
4.2.2.1 LHFPL5 Gene and Protein
LHFPL5 (lipoma HMGIC fusion partner-like 5), also known as TMHS (tetraspan membrane protein of hair-cell stereocilia), is a predicted tetraspan transmembrane stereociliary protein, as its name implicates. Human LHFPL5 gene contains 4 exons, encoding 219 amino acids. LacZ reporter assay and in situ hybridization showed that Lhfpl5 mRNA was expressed in mouse cochlear and vestibular hair cells [93, 94]. The hair cell-specific expression of Lhfpl5 was further supported by RNA-seq results (SHIELD; https://shield.hms.harvard.edu) [74, 75].
LHFPL5 is predicted to contain four transmembrane helices as well as two extracellular loops (Fig. 4.1). LHFPL5 belongs to a small tetraspan transmembrane protein family that includes LHFP as well as LHFP-like 1–5 (LHFPL1–5) [95, 96], whose biological functions largely remain unknown. In general, tetraspan proteins constitute a large protein superfamily, which includes claudin tight junction proteins, connexin gap junction proteins, clarins, etc. Some tetraspan proteins are involved in channel activity. For example, some claudins act as channels that specifically allow certain ions to cross tight junctions [97]. Connexins are subunits of gap junctions that form transmembrane channels connecting the cytoplasm of adjacent cells [98]. LRRC8 family tetraspan proteins form the pore of volume-regulated anion channels (VRAC) that sense ionic strength [99]. Additionally, tetraspan proteins TARPs regulate AMPA channel activity as auxiliary subunits [100]. It awaits further investigation whether LHFPL5 could act as pore-forming subunit or auxiliary subunit of channels.
4.2.2.2 LHFPL5 Localizes Near the Lower End of Tip Links and Binds PCDH15
Immunostaining showed that LHFPL5 localized near the lower end of tip links in hair cells [94], putting LHFPL5 at the position where MET machinery localizes [68]. Moreover, LHFPL5 directly binds to the transmembrane and membrane-proximal domains of PCDH15 [94]. Further investigation demonstrated that LHFPL5 was necessary for the localization of PCDH15 and TMC1 on the stereocilia, thereby controlling the formation of tip link and MET machinery [79, 94].
4.2.2.3 LHFPL5 Disruption Causes Hearing Loss
Mutations in LHFPL5 gene cause autosomal recessive nonsyndromic deafness DFNB67 [101, 102]. Lhfpl5 mutation is also responsible for profound hearing loss and balance deficits in hurry-scurry (hscy) mice [96]. Hscy mice harbour a cysteine-to-phenylalanine (C161F) missense mutation in LHFPL5, which occurs in the second extracellular loop. The mutant LHFPL5 is unstable and is undetected in the inner ear of hscy mice. In hscy mice, stereocilia were disorganized when examined at P8, followed by hair-cell degeneration [96]. Lhfpl5 knockout mice were also developed and showed phenotype identical to that of the hscy mice [93].
4.2.2.4 LHFPL5 Disruption Affects MET
Recently, Lhfpl5 mutation was shown to lead to a nearly 90% reduction in MET in cochlear hair cells of hscy mice and Lhfpl5 knockout mice [94]. Single-channel recordings revealed that the conductance of MET was reduced and adaptation was severely impaired in the absence of LHFPL5, suggesting that LHFPL5 is an integral component of the MET machinery [94]. Moreover, the tonotopic gradient in the conductance of the transducer channels was also blunted in Lhfpl5 knockout mice [79].
4.2.3 TMIE
4.2.3.1 TMIE Gene and Protein
Human TMIE gene consists of 6 exons that encode a small protein of 156 amino acids. Northern blot showed that three Tmie transcripts were expressed in various mouse tissues with a molecular weight of 2.2 kb, 2.8 kb, and 3.2 kb, respectively [103]. RNA sequencing revealed that Tmie was enriched in hair cells in mouse organ of Corti (SHIELD; https://shield.hms.harvard.edu) [74, 75]. LacZ reporter assay also showed that Tmie mRNA was specifically expressed in cochlear and vestibular hair cells [65].
TMIE contains two predicted transmembrane domains (Fig. 4.1). Consistently, heterologously expressed TMIE localized on the plasma membrane of HEK293 cells [65, 104]. Western blot with a polyclonal antibody detected a single band of approximately 17 kD in rat tissues [105]. TMIE shows no homology with any known proteins, and at present its molecular function remains elusive. It is worth noticing that proteins with two transmembrane domains such as ENaC/DEG have been shown to form the pore of MET channels in C. elegans [106].
4.2.3.2 TMIE Localizes Near the Lower End of Tip Links and Binds PCDH15
Immunostaining revealed that TMIE localized at the tips of shorter stereocilia, where the lower end of tip links inserts [65]. In cultured cochlear explants that injectoporated with TMIE-HA expression plasmid, HA immunoreactivity accumulated at the tips of the shorter rows of stereocilia, confirming the localization of TMIE near the lower end of tip links [65]. Furthermore, yeast two-hybrid screen and co-immunoprecipitation experiments showed that TMIE directly binds LHFPL5 as well as PCDH15-CD2 [65].
The localization of TMIE and its interaction with known MET components suggested that it is an integral component of hair-cell MET machinery. TMIE, LHFPL5, and PCDH15 could form a ternary complex, but the geometry of this protein complex differs when different PCDH15 isoforms are present. When PCDH15-CD2 is present, TMIE, LHFPL5, and PCDH15-CD2 bind to one another directly. On the other hand, when PCDH15-CD1 or PCDH15-CD3 is present, TMIE and PCDH15-CD1/CD3 bind to each other indirectly via LHFPL5. The TMIE/LHFPL5/PCDH15 complex might further connect to TMC1/TMC2 via PCDH15. Hence LHFPL5 seems to play a central role in the organization of this protein complex. Consistently, disruption of LHFPL5, but not TMIE, affected tip-link assembly as well as the expression of TMC1/TMC2 on stereocilia [65, 79, 94].
4.2.3.3 TMIE Disruption Causes Hearing Loss
Mutations of TMIE cause autosomal recessive nonsyndromic deafness DFNB6 [107]. Tmie mutations are also responsible for hearing loss and vestibular dysfunction in spinner (sr) as well as circling (cir) mutant mice [103, 108]. Sr and cir mice have deletions in the genome that include the entire Tmie gene, while sr J mice contain a single-base-pair substitution that truncates TMIE protein. Both sr and cir mice showed disorganized stereocilia of auditory hair cells after P10 [103, 109]. Similarly, when selective exons of Tmie gene were deleted through homologous recombination, the mice showed profound hearing deficits [65].
4.2.3.4 TMIE Disruption Affects MET
MET was completely abolished in hair cells of Tmie knockout mice [65]. Moreover, MET was reduced when a dominant negative TMIE fragment was expressed in wild-type hair cells [65]. These results suggested that TIME is indispensable for MET. Surprisingly, although TMIE disruption completely abolished MET in hair cells, it didn’t affect the localization of other known components of the MET machinery in the stereocilia [65]. The mechanism by which TMIE affects MET awaits further investigation.
4.2.4 CIB2
4.2.4.1 CIB2 Gene and Protein
Human CIB2 (calcium- and integrin-binding protein 2) gene contains 6 exons encoding 3 isoforms, the longest of which consists of 210 amino acids [110, 111]. Cib2 was expressed ubiquitously in multiple tissues such as skeletal muscle, the brain, the eye, and the inner ear [110,111,112]. RNA sequencing revealed that Cib2 was enriched in mouse auditory and vestibular hair cells (SHIELD; https://shield.hms.harvard.edu) [74, 75]. Consistently, LacZ reporter assay showed that Cib2 mRNA was specifically expressed in hair cells in mouse inner ear [113].
Different from the MET components discussed above, CIB2 is a soluble protein. CIB2 belongs to a protein family that includes CIB1 through CIB4, which is characterized by multiple calcium-binding EF-hand domains [114]. CIB2 contains three EF-hand domains and binds calcium through the last two EF-hand domains [115] (Fig. 4.1). Moreover, CIB2 underwent N-myristoylation and was associated with intracellular membranes in neurons, co-localizing with Golgi apparatus and dendrite markers [115]. Known CIB2-binding partners include integrins αIIb and α7b, myosin VIIA, whirlin, and sphingosine kinase 1 (SK1) [111, 112, 116, 117]. CIB2 has been suggested to regulate HIV-1 entry through affecting the surface receptor such as CXCR4, CCR5, and integrin α4β7, but the detailed mechanism remains elusive [118, 119].
4.2.4.2 CIB2 Is Concentrated Near the Tips of Stereocilia and Interacts with TMC1/TMC2
In the inner ear, CIB2 immunoreactivity localized along the length of hair-cell stereocilia and was concentrated at the tips of stereocilia [111, 113]. Concentrated localization at the stereocilia tips was also observed when CIB2-GFP was transfected into auditory or vestibular hair cells using gene gun [111]. Notably, both endogenous and exogenous CIB2 were more concentrated at the tips of shorter row stereocilia [111, 113]. This localization suggested that CIB2 might associate with the MET machinery. Consistent with this hypothesis, fluorescence resonance energy transfer (FRET) and co-immunoprecipitation experiments showed that CIB2 interacts with TMC1/TMC2 in vitro, and the interaction was affected by deafness-associated CIB2 mutations [113]. However, the localization of MET components TMC1/2 and PCDH15 on stereocilia was not affected by CIB2 deficiency, suggesting that CIB2 is not essential for transporting or stabilizing these proteins to stereocilia [113].
4.2.4.3 CIB2 Disruption Causes Hearing Loss
Mutations in the CIB2 gene are associated with nonsyndromic hearing loss DFNB48 [111, 120, 121]. CIB2 mutation has also been reported to lead to syndromic hearing loss USH1J [111], but recently it was suggested that CIB2 mutation might not cause Usher syndrome [122, 123]. Morpholino knockdown of Cib2 expression in the zebrafish embryo resulted in reduced or even absent response to acoustic stimuli as well as balancing problems [111]. To further explore the role of CIB2 in hearing, several lines of Cib2-deficient mice have been established, including Cib2 tm1a/Cib2 tm1b, Cib2 F91S, Cib2 Δex4, and Cib2 Δ17bp. Cib2 tm1a mice were generated by inserting a gene trap cassette containing lacZ and neomycin resistance genes between Cib2 exons 3 and 4 [113, 124]. Cib2 tm1b mice were obtained by crossing Cib2 tm1a mice with Cre-expressing mice to delete the neomycin cassette and exon 4 of Cib2 [113]. Cib2 F91S knockin mice carry a p.F91S missense mutation, which is the most prevalent CIB2 mutation that causes nonsyndromic deafness [113]. Cib2 Δex4 mice were generated by putting loxP sites flanking exon 4 of Cib2 gene and crossing with Cre-expressing mice to delete exon 4 [122]. Lastly, Cib2 Δ17bp mice were generated using CRISPR/Cas9 technique that frameshift deletions (8 and 9 bp, respectively) were introduced into exon 4 of Cib2 gene [125].
All the Cib2-deficient mice showed profound hearing loss, confirming that CIB2 plays an important role in hearing transduction [113, 122, 124, 125]. Interestingly, the shorter row stereocilia in Cib2-deficient OHCs and IHCs were over-elongated, whereas the tallest stereocilia remained unaffected [113, 125]. Furthermore, the kinocilia in Cib2-deficient IHCs do not regress properly during development [113, 122, 125].
4.2.4.4 CIB2 Disruption Affects MET
FM1-43 dye uptake and microphonic potential were reduced in the lateral-line hair cells of Cib2 zebrafish morphants, suggesting that MET is affected by Cib2 knockdown [111]. In Cib2 tm1a hair cells, FM1-43 dye uptake was even completely abolished [113]. Consistently, whole-cell patch-clamp recordings showed that the conventional MET currents in Cib2-deficient IHCs and OHCs were completely absent, whereas the reverse-polarity MET was unaffected [113, 122, 125]. Taken together, these results suggest that CIB2 plays an indispensable role in hair-cell MET.
4.2.5 PIEZO2
In mature hair cells, deflection of the hair bundles towards the tallest stereocilia increases the open probability of the sensory MET channels, while deflection in the opposite direction decreases the open probability [126]. However, in the developing immature hair cells, the hair bundles are less directionally sensitive, and transducer currents can be evoked by deflection of the hair bundles in both directions [127, 128]. Additionally, reverse-polarity currents can also be evoked in hair cells lacking tip links, as well as in hair cells deficient for MYO7A, MYO15A, ADGRV1, TMC1/TMC2, LHFPL5, or TMIE [36, 65, 79, 83, 128,129,130]. The ion selectivity and responsiveness to pharmacological blockers of the reverse-polarity current are similar but not identical to that of the regular MET current [83, 131]. High-speed Ca2+ imaging suggested that the reverse-polarity channels are not localized to the hair bundle but distributed at the apical surface of hair cells [132]. These data suggest that the reverse-polarity MET currents are mediated by channels different from the regular MET channels associated with the lower end of tip links.
Recently, PIEZO2 was suggested to constitute the MET channel responsible for the reverse-polarity currents in mouse hair cells [133]. PIEZO2 and its close homolog PIEZO1 are the first mammalian mechanosensitive ion channels identified so far [134]. Both proteins consist of more than 2500 amino acids that are predicted to encompass 26–40 transmembrane domains. PIEZO2 mediated MET in mouse Merkel cells, dorsal root ganglion cells, proprioceptors, and airway-innervating sensory neurons, and deletion of Piezo2 gene caused loss of touch sensation and proprioception as well as lung inflation-induced apnoea [135,136,137,138]. Mutations of human PIEZO2 gene are associated with distal arthrogryposis, muscular atrophy, or loss of discriminative touch perception [139,140,141,142].
In the mouse inner ear, Piezo2 expression was detected in OHCs and vestibular hair cells, but not in IHCs. Immunostaining revealed that PIEZO2 localized at the apical surface of OHCs near the tallest stereocilia [133]. In conditional Piezo2 knockout mice that Piezo2 was inactivated in the inner ear, the patterning of organ of Corti as well as hair bundle morphology was not affected, whereas OHC function was mildly affected, but no obvious vestibular defect was observed [133]. Interestingly, Piezo2 disruption did not affect regular hair-cell MET but abolished the reverse-polarity currents in hair cells lacking tip links or in immature hair cells [133]. These data suggest that PIEZO2 is responsible for the reverse-polarity currents in mouse hair cells.
4.3 Discussion
The mechanism of mammalian hair-cell MET has been under intense investigation in the recent several decades, and significant advances have been achieved. At present, researchers have identified the proteins that constitute the tip links as well as other MET components and are on the way to discover the MET channel itself. However, as we pointed out in the above sections, there is still a long way to go before we can fully understand how MET happens in hair cells.
Tip links play a pivotal role in hair-cell MET. Convincing evidences have been provided suggesting that PCDH15 and CDH23 bind each other via their N-termini and constitute the lower and upper component of tip links, respectively. It’s worth noting that tip links might not always be composed of PCDH15 and CDH23. For example, during regeneration of disrupted tip links, shorter tip links containing only PCDH15 appeared first and then were replaced by mature PCDH15/CDH23 tip links [143]. It also remains elusive whether tip links could act as the gating spring during MET. High-resolution electron microscopy suggested that tip links are relatively stiff and inextensible [33]. Molecular dynamics simulations based on the structure of the CDH23 EC1–EC2 fragment suggested that the EC repeats of CDH23 are quite stiff [25]. Recently, however, some of the PCDH15 EC repeat linker regions were shown to be calcium-free or partially calcium-free, which might confer some elasticity to the otherwise rigid tip links [59, 60].
PCDH15 and CDH23 interact with other proteins via their transmembrane domains or cytoplasmic domains and form the so-called lower tip-link complex and upper tip-link complex, respectively. The molecular composition of these complexes is just emerging. MYO7A, harmonin, SANS, and possibly MYO1C are considered as components of upper tip-link complex. The adaption motor is positioned near the upper tip-link complex. Through moving along the stereocilia, the motor regulates the tension of tip links and adjusts the operating point of the MET channels. MYO1C is the most prominent candidate for the adaptation motor, whereas MYO7A and harmonin are also suggested to regulate slow adaptation directly or indirectly.
So far four transmembrane proteins have been localized near the lower tip-link insertion site, namely, TMC1, TMC2, LHFPL5, and TMIE. They all interact with PCDH15 in vitro and are likely part of the lower tip-link complex. TMC1 and TMC2 are potential six-transmembrane proteins and have been proposed as pore-forming subunits of hair-cell MET channel, although compelling evidence is needed to fully support this conclusion. LHFPL5 and TMIE contain four and two predicted transmembrane domains, respectively, and are suggested to be integral components of hair-cell MET machinery. Another candidate MET component is CIB2, a soluble protein that is concentrated at the tips of shorter row stereocilia and binds TMC1/TMC2. Characterization of the MET channel surely is the most exiting task in this field in the coming several years.
References
Siemens, J., et al. 2004. Cadherin 23 is a component of the tip link in hair-cell stereocilia. Nature 428 (6986): 950–955.
Ahmed, Z.M., et al. 2006. The tip-link antigen, a protein associated with the transduction complex of sensory hair cells, is protocadherin-15. The Journal of Neuroscience 26 (26): 7022–7034.
Kazmierczak, P., et al. 2007. Cadherin 23 and protocadherin 15 interact to form tip-link filaments in sensory hair cells. Nature 449 (7158): 87–91.
Bolz, H., et al. 2001. Mutation of CDH23, encoding a new member of the cadherin gene family, causes Usher syndrome type 1D. Nature Genetics 27 (1): 108–112.
Bork, J.M., et al. 2001. Usher syndrome 1D and nonsyndromic autosomal recessive deafness DFNB12 are caused by allelic mutations of the novel cadherin-like gene CDH23. American Journal of Human Genetics 68 (1): 26–37.
Ahmed, Z.M., et al. 2001. Mutations of the protocadherin gene PCDH15 cause Usher syndrome type 1F. American Journal of Human Genetics 69 (1): 25–34.
Alagramam, K.N., et al. 2001. Mutations in the novel protocadherin PCDH15 cause Usher syndrome type 1F. Human Molecular Genetics 10 (16): 1709–1718.
Ahmed, Z.M., et al. 2003. PCDH15 is expressed in the neurosensory epithelium of the eye and ear and mutant alleles are responsible for both USH1F and DFNB23. Human Molecular Genetics 12 (24): 3215–3223.
Mathur, P., and J. Yang. 2015. Usher syndrome: hearing loss, retinal degeneration and associated abnormalities. Biochimica et Biophysica Acta 1852 (3): 406–420.
Weil, D., et al. 1995. Defective myosin VIIA gene responsible for Usher syndrome type 1B. Nature 374 (6517): 60–61.
Verpy, E., et al. 2000. A defect in harmonin, a PDZ domain-containing protein expressed in the inner ear sensory hair cells, underlies Usher syndrome type 1C. Nature Genetics 26 (1): 51–55.
Bitner-Glindzicz, M., et al. 2000. A recessive contiguous gene deletion causing infantile hyperinsulinism, enteropathy and deafness identifies the Usher type 1C gene. Nature Genetics 26 (1): 56–60.
Weil, D., et al. 2003. Usher syndrome type I G (USH1G) is caused by mutations in the gene encoding SANS, a protein that associates with the USH1C protein, harmonin. Human Molecular Genetics 12 (5): 463–471.
Eudy, J.D., et al. 1998. Mutation of a gene encoding a protein with extracellular matrix motifs in Usher syndrome type IIa. Science 280 (5370): 1753–1757.
Weston, M.D., et al. 2004. Mutations in the VLGR1 gene implicate G-protein signaling in the pathogenesis of Usher syndrome type II. American Journal of Human Genetics 74 (2): 357–366.
Ebermann, I., et al. 2007. A novel gene for Usher syndrome type 2: mutations in the long isoform of whirlin are associated with retinitis pigmentosa and sensorineural hearing loss. Human Genetics 121 (2): 203–211.
Joensuu, T., et al. 2001. Mutations in a novel gene with transmembrane domains underlie Usher syndrome type 3. American Journal of Human Genetics 69 (4): 673–684.
Fields, R.R., et al. 2002. Usher syndrome type III: revised genomic structure of the USH3 gene and identification of novel mutations. American Journal of Human Genetics 71 (3): 607–617.
Adato, A., et al. 2002. USH3A transcripts encode clarin-1, a four-transmembrane-domain protein with a possible role in sensory synapses. European Journal of Human Genetics 10 (6): 339–350.
Ebermann, I., et al. 2010. PDZD7 is a modifier of retinal disease and a contributor to digenic Usher syndrome. The Journal of Clinical Investigation 120 (6): 1812–1823.
Adato, A., et al. 2005. Interactions in the network of Usher syndrome type 1 proteins. Human Molecular Genetics 14 (3): 347–356.
Chen, Q., et al. 2014. Whirlin and PDZ domain-containing 7 (PDZD7) proteins are both required to form the quaternary protein complex associated with Usher syndrome type 2. The Journal of Biological Chemistry 289 (52): 36070–36088.
Lagziel, A., et al. 2005. Spatiotemporal pattern and isoforms of cadherin 23 in wild type and waltzer mice during inner ear hair cell development. Developmental Biology 280 (2): 295–306.
Boggon, T.J., et al. 2002. C-cadherin ectodomain structure and implications for cell adhesion mechanisms. Science 296 (5571): 1308–1313.
Sotomayor, M., et al. 2010. Structural determinants of cadherin-23 function in hearing and deafness. Neuron 66 (1): 85–100.
Boeda, B., et al. 2002. Myosin VIIa, harmonin and cadherin 23, three Usher I gene products that cooperate to shape the sensory hair cell bundle. The EMBO Journal 21 (24): 6689–6699.
Siemens, J., et al. 2002. The Usher syndrome proteins cadherin 23 and harmonin form a complex by means of PDZ-domain interactions. Proceedings of the National Academy of Sciences of the United States of America 99 (23): 14946–14951.
Xu, Z., et al. 2008. MAGI-1, a candidate stereociliary scaffolding protein, associates with the tip-link component cadherin 23. The Journal of Neuroscience 28 (44): 11269–11276.
Xu, Z., K. Oshima, and S. Heller. 2010. PIST regulates the intracellular trafficking and plasma membrane expression of cadherin 23. BMC Cell Biology 11: 80.
Di Palma, F., R. Pellegrino, and K. Noben-Trauth. 2001. Genomic structure, alternative splice forms and normal and mutant alleles of cadherin 23 (Cdh23). Gene 281 (1-2): 31–41.
Yonezawa, S., et al. 2008. Redox-dependent structural ambivalence of the cytoplasmic domain in the inner ear-specific cadherin 23 isoform. Biochemical and Biophysical Research Communications 366 (1): 92–97.
Michel, V., et al. 2005. Cadherin 23 is a component of the transient lateral links in the developing hair bundles of cochlear sensory cells. Developmental Biology 280 (2): 281–294.
Kachar, B., et al. 2000. High-resolution structure of hair-cell tip links. Proceedings of the National Academy of Sciences of the United States of America 97 (24): 13336–13341.
Sotomayor, M., et al. 2012. Structure of a force-conveying cadherin bond essential for inner-ear mechanotransduction. Nature 492 (7427): 128–132.
Lelli, A., et al. 2010. Development and regeneration of sensory transduction in auditory hair cells requires functional interaction between cadherin-23 and protocadherin-15. The Journal of Neuroscience 30 (34): 11259–11269.
Alagramam, K.N., et al. 2011. Mutations in protocadherin 15 and cadherin 23 affect tip links and mechanotransduction in mammalian sensory hair cells. PLoS One 6 (4): e19183.
Di Palma, F., et al. 2001. Mutations in Cdh23, encoding a new type of cadherin, cause stereocilia disorganization in waltzer, the mouse model for Usher syndrome type 1D. Nature Genetics 27 (1): 103–107.
Schwander, M., et al. 2009. A mouse model for nonsyndromic deafness (DFNB12) links hearing loss to defects in tip links of mechanosensory hair cells. Proceedings of the National Academy of Sciences of the United States of America 106 (13): 5252–5257.
Manji, S.S., et al. 2011. An ENU-induced mutation of Cdh23 causes congenital hearing loss, but no vestibular dysfunction, in mice. The American Journal of Pathology 179 (2): 903–914.
Pan, L., et al. 2009. Assembling stable hair cell tip link complex via multidentate interactions between harmonin and cadherin 23. Proceedings of the National Academy of Sciences of the United States of America 106 (14): 5575–5580.
Grillet, N., et al. 2009. Harmonin mutations cause mechanotransduction defects in cochlear hair cells. Neuron 62 (3): 375–387.
Bahloul, A., et al. 2010. Cadherin-23, myosin VIIa and harmonin, encoded by Usher syndrome type I genes, form a ternary complex and interact with membrane phospholipids. Human Molecular Genetics 19 (18): 3557–3565.
Caberlotto, E., et al. 2011. Usher type 1G protein sans is a critical component of the tip-link complex, a structure controlling actin polymerization in stereocilia. Proceedings of the National Academy of Sciences of the United States of America 108 (14): 5825–5830.
Grati, M., and B. Kachar. 2011. Myosin VIIa and sans localization at stereocilia upper tip-link density implicates these Usher syndrome proteins in mechanotransduction. Proceedings of the National Academy of Sciences of the United States of America 108 (28): 11476–11481.
Kros, C.J., et al. 2002. Reduced climbing and increased slipping adaptation in cochlear hair cells of mice with Myo7a mutations. Nature Neuroscience 5 (1): 41–47.
Assad, J.A., and D.P. Corey. 1992. An active motor model for adaptation by vertebrate hair cells. The Journal of Neuroscience 12 (9): 3291–3309.
Yamoah, E.N., and P.G. Gillespie. 1996. Phosphate analogs block adaptation in hair cells by inhibiting adaptation-motor force production. Neuron 17 (3): 523–533.
Eatock, R.A., D.P. Corey, and A.J. Hudspeth. 1987. Adaptation of mechanoelectrical transduction in hair cells of the bullfrog’s sacculus. The Journal of Neuroscience 7 (9): 2821–2836.
Holt, J.R., D.P. Corey, and R.A. Eatock. 1997. Mechanoelectrical transduction and adaptation in hair cells of the mouse utricle, a low-frequency vestibular organ. The Journal of Neuroscience 17 (22): 8739–8748.
Wu, Y.C., A.J. Ricci, and R. Fettiplace. 1999. Two components of transducer adaptation in auditory hair cells. Journal of Neurophysiology 82 (5): 2171–2181.
Stauffer, E.A., and J.R. Holt. 2007. Sensory transduction and adaptation in inner and outer hair cells of the mouse auditory system. Journal of Neurophysiology 98 (6): 3360–3369.
Steyger, P.S., P.G. Gillespie, and R.A. Baird. 1998. Myosin Ibeta is located at tip link anchors in vestibular hair bundles. The Journal of Neuroscience 18 (12): 4603–4615.
Garcia, J.A., et al. 1998. Localization of myosin-Ibeta near both ends of tip links in frog saccular hair cells. The Journal of Neuroscience 18 (21): 8637–8647.
Schneider, M.E., et al. 2006. A new compartment at stereocilia tips defined by spatial and temporal patterns of myosin IIIa expression. The Journal of Neuroscience 26 (40): 10243–10252.
Gillespie, P.G., et al. 1999. Engineering of the myosin-ibeta nucleotide-binding pocket to create selective sensitivity to N(6)-modified ADP analogs. The Journal of Biological Chemistry 274 (44): 31373–31381.
Holt, J.R., et al. 2002. A chemical-genetic strategy implicates myosin-1c in adaptation by hair cells. Cell 108 (3): 371–381.
Stauffer, E.A., et al. 2005. Fast adaptation in vestibular hair cells requires myosin-1c activity. Neuron 47 (4): 541–553.
Geng, R., et al. 2013. Noddy, a mouse harboring a missense mutation in protocadherin-15, reveals the impact of disrupting a critical interaction site between tip-link cadherins in inner ear hair cells. The Journal of Neuroscience 33 (10): 4395–4404.
Araya-Secchi, R., B.L. Neel, and M. Sotomayor. 2016. An elastic element in the protocadherin-15 tip link of the inner ear. Nature Communications 7: 13458.
Powers, R.E., R. Gaudet, and M. Sotomayor. 2017. A partial calcium-free linker confers flexibility to inner-ear protocadherin-15. Structure 25: 482–495.
Reiners, J., et al. 2005. Photoreceptor expression of the Usher syndrome type 1 protein protocadherin 15 (USH1F) and its interaction with the scaffold protein harmonin (USH1C). Molecular Vision 11: 347–355.
Nie, H., et al. 2016. Plasma membrane targeting of protocadherin 15 is regulated by the Golgi-associated chaperone protein PIST. Neural Plasticity 2016: 8580675.
Pepermans, E., et al. 2014. The CD2 isoform of protocadherin-15 is an essential component of the tip-link complex in mature auditory hair cells. EMBO Molecular Medicine 6 (7): 984–992.
Webb, S.W., et al. 2011. Regulation of PCDH15 function in mechanosensory hair cells by alternative splicing of the cytoplasmic domain. Development 138 (8): 1607–1617.
Zhao, B., et al. 2014. TMIE is an essential component of the mechanotransduction machinery of cochlear hair cells. Neuron 84 (5): 954–967.
Senften, M., et al. 2006. Physical and functional interaction between protocadherin 15 and myosin VIIa in mechanosensory hair cells. The Journal of Neuroscience 26 (7): 2060–2071.
Ricci, A.J., A.C. Crawford, and R. Fettiplace. 2003. Tonotopic variation in the conductance of the hair cell mechanotransducer channel. Neuron 40 (5): 983–990.
Beurg, M., et al. 2009. Localization of inner hair cell mechanotransducer channels using high-speed calcium imaging. Nature Neuroscience 12 (5): 553–558.
Kurima, K., et al. 2002. Dominant and recessive deafness caused by mutations of a novel gene, TMC1, required for cochlear hair-cell function. Nature Genetics 30 (3): 277–284.
Keresztes, G., H. Mutai, and S. Heller. 2003. TMC and EVER genes belong to a larger novel family, the TMC gene family encoding transmembrane proteins. BMC Genomics 4 (1): 24.
Kurima, K., et al. 2003. Characterization of the transmembrane channel-like (TMC) gene family: functional clues from hearing loss and epidermodysplasia verruciformis. Genomics 82 (3): 300–308.
Kawashima, Y., et al. 2011. Mechanotransduction in mouse inner ear hair cells requires transmembrane channel-like genes. The Journal of Clinical Investigation 121 (12): 4796–4809.
Labay, V., et al. 2010. Topology of transmembrane channel-like gene 1 protein. Biochemistry 49 (39): 8592–8598.
Scheffer, D.I., et al. 2015. Gene expression by mouse inner ear hair cells during development. The Journal of Neuroscience 35 (16): 6366–6380.
Shen, J., et al. 2015. SHIELD: an integrative gene expression database for inner ear research. Database: The Journal of Biological Databases and Curation 2015: bav071.
Geleoc, G.S., and J.R. Holt. 2003. Developmental acquisition of sensory transduction in hair cells of the mouse inner ear. Nature Neuroscience 6 (10): 1019–1020.
Kurima, K., et al. 2015. TMC1 and TMC2 localize at the site of mechanotransduction in mammalian inner ear hair cell stereocilia. Cell Reports 12 (10): 1606–1617.
Maeda, R., et al. 2014. Tip-link protein protocadherin 15 interacts with transmembrane channel-like proteins TMC1 and TMC2. Proceedings of the National Academy of Sciences of the United States of America 111 (35): 12907–12912.
Beurg, M., et al. 2015. Subunit determination of the conductance of hair-cell mechanotransducer channels. Proceedings of the National Academy of Sciences of the United States of America 112 (5): 1589–1594.
Vreugde, S., et al. 2002. Beethoven, a mouse model for dominant, progressive hearing loss DFNA36. Nature Genetics 30 (3): 257–258.
Steel, K.P., and G.R. Bock. 1980. The nature of inherited deafness in deafness mice. Nature 288 (5787): 159–161.
Marcotti, W., et al. 2006. Tmc1 is necessary for normal functional maturation and survival of inner and outer hair cells in the mouse cochlea. The Journal of Physiology 574 (Pt 3): 677–698.
Kim, K.X., et al. 2013. The role of transmembrane channel-like proteins in the operation of hair cell mechanotransducer channels. The Journal of General Physiology 142 (5): 493–505.
Pan, B., et al. 2013. TMC1 and TMC2 are components of the mechanotransduction channel in hair cells of the mammalian inner ear. Neuron 79 (3): 504–515.
Cunningham, C.L., et al. 2017. The murine catecholamine methyltransferase mTOMT is essential for mechanotransduction by cochlear hair cells. eLife 6: e24318.
Erickson, T., et al. 2017. Integration of Tmc1/2 into the mechanotransduction complex in zebrafish hair cells is regulated by transmembrane O-methyltransferase (Tomt). eLife 6: e28474.
Ahmed, Z.M., et al. 2008. Mutations of LRTOMT, a fusion gene with alternative reading frames, cause nonsyndromic deafness in humans. Nature Genetics 40 (11): 1335–1340.
Du, X., et al. 2008. A catechol-O-methyltransferase that is essential for auditory function in mice and humans. Proceedings of the National Academy of Sciences of the United States of America 105 (38): 14609–14614.
Fettiplace, R. 2016. Is TMC1 the hair cell mechanotransducer channel? Biophysical Journal 111 (1): 3–9.
Corey, D.P., and J.R. Holt. 2016. Are TMCs the mechanotransduction channels of vertebrate hair cells? The Journal of Neuroscience 36 (43): 10921–10926.
Wu, Z., and U. Muller. 2016. Molecular identity of the mechanotransduction channel in hair cells: not quiet there yet. The Journal of Neuroscience 36 (43): 10927–10934.
Chatzigeorgiou, M., et al. 2013. tmc-1 encodes a sodium-sensitive channel required for salt chemosensation in C. elegans. Nature 494 (7435): 95–99.
Longo-Guess, C.M., et al. 2007. Targeted knockout and lacZ reporter expression of the mouse Tmhs deafness gene and characterization of the hscy-2J mutation. Mammalian Genome 18 (9): 646–656.
Xiong, W., et al. 2012. TMHS is an integral component of the mechanotransduction machinery of cochlear hair cells. Cell 151 (6): 1283–1295.
Petit, M.M., et al. 1999. LHFP, a novel translocation partner gene of HMGIC in a lipoma, is a member of a new family of LHFP-like genes. Genomics 57 (3): 438–441.
Longo-Guess, C.M., et al. 2005. A missense mutation in the previously undescribed gene Tmhs underlies deafness in hurry-scurry (hscy) mice. Proceedings of the National Academy of Sciences of the United States of America 102 (22): 7894–7899.
Gunzel, D. 2017. Claudins: vital partners in transcellular and paracellular transport coupling. Pflügers Archiv 469 (1): 35–44.
Kar, R., et al. 2012. Biological role of connexin intercellular channels and hemichannels. Archives of Biochemistry and Biophysics 524 (1): 2–15.
Syeda, R., et al. 2016. LRRC8 proteins form volume-regulated anion channels that sense ionic strength. Cell 164 (3): 499–511.
Jackson, A.C., and R.A. Nicoll. 2011. The expanding social network of ionotropic glutamate receptors: TARPs and other transmembrane auxiliary subunits. Neuron 70 (2): 178–199.
Kalay, E., et al. 2006. Mutations in the lipoma HMGIC fusion partner-like 5 (LHFPL5) gene cause autosomal recessive nonsyndromic hearing loss. Human Mutation 27 (7): 633–639.
Shabbir, M.I., et al. 2006. Mutations of human TMHS cause recessively inherited non-syndromic hearing loss. Journal of Medical Genetics 43 (8): 634–640.
Mitchem, K.L., et al. 2002. Mutation of the novel gene Tmie results in sensory cell defects in the inner ear of spinner, a mouse model of human hearing loss DFNB6. Human Molecular Genetics 11 (16): 1887–1898.
Karuppasamy, S., et al. 2011. Subcellular localization of the transmembrane inner ear (Tmie) protein in a stable Tmie-expressing cell line. Laboratory Animal Research 27 (4): 339–342.
Su, M.C., et al. 2008. Expression and localization of Tmie in adult rat cochlea. Histochemistry and Cell Biology 130 (1): 119–126.
O’Hagan, R., M. Chalfie, and M.B. Goodman. 2005. The MEC-4 DEG/ENaC channel of Caenorhabditis elegans touch receptor neurons transduces mechanical signals. Nature Neuroscience 8 (1): 43–50.
Naz, S., et al. 2002. Mutations in a novel gene, TMIE, are associated with hearing loss linked to the DFNB6 locus. American Journal of Human Genetics 71 (3): 632–636.
Cho, K.I., et al. 2006. The circling mouse (C57BL/6J-cir) has a 40-kilobase genomic deletion that includes the transmembrane inner ear (tmie) gene. Comparative Medicine 56 (6): 476–481.
Chung, W.H., et al. 2007. Cochlear pathology of the circling mouse: a new mouse model of DFNB6. Acta Oto-Laryngologica 127 (3): 244–251.
Seki, N., et al. 1999. Structure, expression profile and chromosomal location of an isolog of DNA-PKcs interacting protein (KIP) gene. Biochimica et Biophysica Acta 1444 (1): 143–147.
Riazuddin, S., et al. 2012. Alterations of the CIB2 calcium- and integrin-binding protein cause Usher syndrome type 1J and nonsyndromic deafness DFNB48. Nature Genetics 44 (11): 1265–1271.
Hager, M., et al. 2008. Cib2 binds integrin alpha7Bbeta1D and is reduced in laminin alpha2 chain-deficient muscular dystrophy. The Journal of Biological Chemistry 283 (36): 24760–24769.
Giese, A.P.J., et al. 2017. CIB2 interacts with TMC1 and TMC2 and is essential for mechanotransduction in auditory hair cells. Nature Communications 8 (1): 43.
Gentry, H.R., et al. 2005. Structural and biochemical characterization of CIB1 delineates a new family of EF-hand-containing proteins. The Journal of Biological Chemistry 280 (9): 8407–8415.
Blazejczyk, M., et al. 2009. Biochemical characterization and expression analysis of a novel EF-hand Ca2+ binding protein calmyrin2 (Cib2) in brain indicates its function in NMDA receptor mediated Ca2+ signaling. Archives of Biochemistry and Biophysics 487 (1): 66–78.
Huang, H., J.N. Bogstie, and H.J. Vogel. 2012. Biophysical and structural studies of the human calcium- and integrin-binding protein family: understanding their functional similarities and differences. Biochemistry and Cell Biology 90 (5): 646–656.
Zhu, W., et al. 2017. CIB2 negatively regulates oncogenic signaling in ovarian cancer via sphingosine kinase 1. Cancer Research 77 (18): 4823–4834.
Rato, S., et al. 2010. Novel HIV-1 knockdown targets identified by an enriched kinases/phosphatases shRNA library using a long-term iterative screen in Jurkat T-cells. PLoS One 5 (2): e9276.
Godinho-Santos, A., et al. 2016. CIB1 and CIB2 are HIV-1 helper factors involved in viral entry. Scientific Reports 6: 30927.
Patel, K., et al. 2015. A novel C-terminal CIB2 (Calcium and Integrin Binding Protein 2) mutation associated with non-syndromic hearing loss in a hispanic family. PLoS One 10 (10): e0133082.
Seco, C.Z., et al. 2016. Novel and recurrent CIB2 variants, associated with nonsyndromic deafness, do not affect calcium buffering and localization in hair cells. European Journal of Human Genetics 24 (4): 542–549.
Michel, V., et al. 2017. CIB2, defective in isolated deafness, is key for auditory hair cell mechanotransduction and survival. EMBO Molecular Medicine 9 (12): 1711–1731.
Booth, K.T., et al. 2018. Variants in CIB2 cause DFNB48 and not USH1J. Clinical Genetics. https://doi.org/10.1111/cge.13170.
Zou, J., et al. 2017. The roles of USH1 proteins and PDZ domain-containing USH proteins in USH2 complex integrity in cochlear hair cells. Human Molecular Genetics 26 (3): 624–636.
Wang, Y., et al. 2017. Loss of CIB2 causes profound hearing loss and abolishes mechanoelectrical transduction in mice. Frontiers in Molecular Neuroscience 10: 401.
Shotwell, S.L., R. Jacobs, and A.J. Hudspeth. 1981. Directional sensitivity of individual vertebrate hair cells to controlled deflection of their hair bundles. Annals of the New York Academy of Sciences 374: 1–10.
Kindt, K.S., G. Finch, and T. Nicolson. 2012. Kinocilia mediate mechanosensitivity in developing zebrafish hair cells. Developmental Cell 23 (2): 329–341.
Marcotti, W., et al. 2014. Transduction without tip links in cochlear hair cells is mediated by ion channels with permeation properties distinct from those of the mechano-electrical transducer channel. The Journal of Neuroscience 34 (16): 5505–5514.
Michalski, N., et al. 2007. Molecular characterization of the ankle-link complex in cochlear hair cells and its role in the hair bundle functioning. The Journal of Neuroscience 27 (24): 6478–6488.
Stepanyan, R., and G.I. Frolenkov. 2009. Fast adaptation and Ca2+ sensitivity of the mechanotransducer require myosin-XVa in inner but not outer cochlear hair cells. The Journal of Neuroscience 29 (13): 4023–4034.
Beurg, M., K.X. Kim, and R. Fettiplace. 2014. Conductance and block of hair-cell mechanotransducer channels in transmembrane channel-like protein mutants. The Journal of General Physiology 144 (1): 55–69.
Beurg, M., et al. 2016. Development and localization of reverse-polarity mechanotransducer channels in cochlear hair cells. Proceedings of the National Academy of Sciences of the United States of America 113 (24): 6767–6772.
Wu, Z., et al. 2017. Mechanosensory hair cells express two molecularly distinct mechanotransduction channels. Nature Neuroscience 20 (1): 24–33.
Coste, B., et al. 2010. Piezo1 and Piezo2 are essential components of distinct mechanically activated cation channels. Science 330 (6000): 55–60.
Woo, S.H., et al. 2014. Piezo2 is required for Merkel-cell mechanotransduction. Nature 509 (7502): 622–626.
Ranade, S.S., et al. 2014. Piezo2 is the major transducer of mechanical forces for touch sensation in mice. Nature 516 (7529): 121–125.
Woo, S.H., et al. 2015. Piezo2 is the principal mechanotransduction channel for proprioception. Nature Neuroscience 18 (12): 1756–1762.
Nonomura, K., et al. 2017. Piezo2 senses airway stretch and mediates lung inflation-induced apnoea. Nature 541 (7636): 176–181.
Coste, B., et al. 2013. Gain-of-function mutations in the mechanically activated ion channel PIEZO2 cause a subtype of Distal Arthrogryposis. Proceedings of the National Academy of Sciences of the United States of America 110 (12): 4667–4672.
McMillin, M.J., et al. 2014. Mutations in PIEZO2 cause Gordon syndrome, Marden-Walker syndrome, and distal arthrogryposis type 5. American Journal of Human Genetics 94 (5): 734–744.
Delle Vedove, A., et al. 2016. Biallelic loss of proprioception-related PIEZO2 causes muscular atrophy with perinatal respiratory distress, arthrogryposis, and scoliosis. American Journal of Human Genetics 99 (5): 1206–1216.
Chesler, A.T., et al. 2016. The role of PIEZO2 in human mechanosensation. The New England Journal of Medicine 375 (14): 1355–1364.
Indzhykulian, A.A., et al. 2013. Molecular remodeling of tip links underlies mechanosensory regeneration in auditory hair cells. PLoS Biology 11 (6): e1001583.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Copyright information
© 2018 The Author(s)
About this chapter
Cite this chapter
Xu, Z. (2018). Molecular Components of Mechanotransduction Machinery. In: Mechanotransduction of the Hair Cell. SpringerBriefs in Biochemistry and Molecular Biology. Springer, Singapore. https://doi.org/10.1007/978-981-10-8557-4_4
Download citation
DOI: https://doi.org/10.1007/978-981-10-8557-4_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-8556-7
Online ISBN: 978-981-10-8557-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)