Abstract
Bioremediation technology involves the use of living organisms like microbes and plants to reduce/degrade, eliminate and transform contaminants present in soils, sediments and water. The technology has gained wider acceptance in the recent years because of its potential to remove various organic and inorganic contaminants from various components of the environment. The technology provides an effective treatment of inorganic and organic contaminants under in situ and ex situ conditions by natural means. Potential of microbes and plants both have been exploited to achieve maximum removal/remediation of inorganic and organic contaminants. The biotechnological approaches and genetic engineering strategies have been employed by researchers to improve the efficacy of this technique for achieving complete degradation of contaminants. Enhancement in potential of both plants and microbes for achieving complete remediation of one or more than one pollutant can prove an asset for remediating contaminated sites. The present chapter highlights the role of microbial and phytoremediation in removal of pollutants from the environment.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
5.1 Introduction
Environmental contamination with inorganic and organic toxicants has increased over the years due to rapid industrialization, urbanization and anthropogenic activities. The organic contaminants such as petroleum hydrocarbons, pesticides, agrochemicals, pharmaceutical product and inorganic pollutants such as heavy metals are constantly added in the environment (Agarwal 1998; Zeyaullah et al. 2009). Most of the xenobiotic compounds resist degradation. The remediation or treatment of contaminants done by conventional methods (both physical and chemical) is a costly, time-consuming, invasive approach and causes environmental deterioration (EPA 1999, 2003). According to an estimate, the cleaning/restoring of the contaminated sites in the USA requires a capital investment of approximately US $1.7 trillion. Bioremediation has emerged as a safe, reliable, effective, low-cost and environmentally friendly alternative technology to achieve sustainable remediation of hazardous and recalcitrant pollutants. In this technique, treatment of contaminants can be done at site in a cost-effective, less disruptive, eco-friendly (no by-products, no requirement of complex setups and operations) manner.
The bioremediation technology uses biological processes and naturally occurring catabolic activity of microbes and plants to eliminate, attenuate, or transform inorganic and organic contaminants to less hazardous products such as carbon dioxide and water (Abruscia et al. 2007; Pandey and Fulekar 2012). Biological agents such as yeast, fungi, bacteria and plants remove contaminants by biotransformation and biodegradation mechanisms. The physiological and metabolic capabilities of organisms assist in degrading the pollutants converting them to nontoxic and environmentally safe products. In this technology, target compound is used as a carbon source. The complete mineralization of contaminants results in the formation of H2O and CO2 (Strong and Burgess 2008; Sharma and Fulekar 2009).
5.2 Bioremediation
Bioremediation processes have been broadly categorized into two groups.
5.2.1 Ex Situ Bioremediation
In this type of remediation, removal of the contaminant from soil and groundwater is done away from the site (Maheshwari et al. 2014). The treatment of contaminants has been done away from site. This includes bioreactors, biofilters, land farming, bioventing, biosparging, biostimulation and composting methods (Olaniran et al. 2006).
Ex situ bioremediation is of two types.
5.2.1.1 Solid Phase Treatment
It is a treatment process for land and soil contaminated with organic, industrial wastes, municipal wastes and sewage sludge. It includes:
-
Land Farming: In this technique, contaminated soil is excavated and spread over a prepared bed and periodically tilled to achieve degradation of pollutants. Microorganisms facilitate aerobic degradation of contaminants.
-
Composting: In this technique, contaminated soil is mixed with nonhazardous organic amendments such as manure or agricultural wastes. The presence of organic materials supports the growth of microbial population.
-
Biopiles: Biopiles are a hybrid of land farming and composting. Engineered cells are constructed as aerated composted piles. Contaminated material is mixed with a bulking agent and aerobic, thermophilic bacteria are used in the treatment process.
-
Bioreactors: In this technique, biodegradation is carried out by microbes in a container. It is used to treat organic contaminants from liquids or slurries.
-
Bioventing: It involves supplying air and nutrients through wells to contaminated soil to stimulate the indigenous bacteria. The low airflow rates provide the amount of oxygen necessary for the biodegradation while minimizing volatilization and release of contaminants to the atmosphere. It is used to treat hydrocarbons.
-
Bioaugmentation: It involves introduction of exogenic microorganisms (sourced from outside the soil environment) capable of detoxifying a particular contaminant. The addition of contaminant-degrading organisms accelerates the transformation rates (El Fantroussi and Agathos 2005; Thierry et al. 2008). Enhanced chlorpyrifos biodegradation has been reported via this process.
-
Biosparging: This involves the injection of air under pressure to increase groundwater oxygen concentrations and enhance the rate of biological degradation of contaminants by naturally occurring bacteria.
-
Biostimulation: This involves the addition of soil nutrients, trace minerals, electron acceptors, or electron donors to enhance the biotransformation of a wide range of soil contaminants (Li et al. 2010). Trichloroethene and perchloroethylene are reported to be completely converted to ethane by microorganisms in a short span of time with the addition of lactate as biostimulation (Shan et al. 2010). Electron shuttles such as humic substances (HS) stimulate anaerobic biotransformation of organic pollutants through enhancing the electron transfer speed.
5.2.1.2 Slurry Phase
In this type of bioremediation, contaminated soil is combined with water, other additives and microbes in a bioreactor. Nutrients and oxygen are added, and conditions are controlled to create the optimum environment for the microorganisms to degrade the contaminants. Slurry reactors are used for treatment of contaminated soil and water.
5.2.2 In Situ Bioremediation
In situ technique is applied to treat contaminated soil and groundwater. This involves addition of indigenous or naturally occurring microbial populations by feeding nutrients and oxygen to increase their metabolic activity. Oxygen, electron acceptors, and nutrients (nitrogen and phosphorus) promote microbial growth. The treatment is done on the site without any need to excavate or remove soils or water in order to accomplish remediation (Vidali 2001).
5.3 Microbial Remediation
5.3.1 Contaminants Removed by Microbes
Naturally occurring bacteria and fungi degrade/detoxify hazardous substances. Aerobic and anaerobic bacteria degrade various inorganic and organic contaminants (Kumar et al. 2011). Aerobic bacteria such as Pseudomonas, Alcaligenes, Sphingomonas, Rhodococcus and Mycobacterium degrade pesticides, hydrocarbons, alkanes and polyaromatic compounds and use the contaminant as the sole source of carbon and energy. Anaerobic bacteria degrade polychlorinated biphenyls (PCBs) and organic solvents such as trichloroethylene (TCE) and chloroform. Dioxygenases and monooxygenases are two of the primary enzymes employed by aerobic organisms during transformation and mineralization of xenobiotics, while anaerobic microbes use range of electron acceptors such as NO3-, Fe, Mn, SO4 2− and CO2 depending on their availability and the prevailing redox conditions. Methane monooxygenase degrade various substrates such as chlorinated aliphatic trichloroethylene and 1,2-dichloroethane.
Microbes form an important part of consortium that assist in degrading contaminants. These include Acinetobacter, Alcaligenes, Arthrobacter, Bacillus, Beijerinckia, Flavobacterium, Methylosinus, Mycobacterium, Myxococcus, Nitrosomonas, Nocardia, Penicillium, Phanerochaete, Pseudomonas, Rhizoctonia, Serratia, Trametes and Xanthobacter (Table 5.1). The complete mineralization involves synergism and cometabolism. Cometabolism of xenobiotics is required when the compound cannot serve as a source of carbon and energy. Hydrocarbons and persistent organic pollutants (POPs) such as dichlorodiphenyltrichloroethane (DDT), polychlorinated biphenyls (PCBs), dioxins etc. are degraded in the soil by bacteria present in the rhizosphere (Olson et al. 2003). Acidophilic bacteria like Acidithiobacillus ferrooxidans (Takeuchi et al. 2005) and sulfur-oxidizing bacteria remove high concentrations of As, Cd, Cu, Co, and Zn from contaminated soils. Pesticides have also been successfully removed by bacteria. Providencia stuartii strain depicts potential for degradation of chlorpyrifos (Surekha Rani et al. 2008). Isolates of Bacillus, Staphylococcus, and Stenotrophomonas from cultivated and uncultivated soil are able to degrade dichlorodiphenyltrichloroethane (DDT) (Kanade et al. 2012). Bacterial strains are able to degrade azo dyes under aerobic and anaerobic conditions (Dos Santos et al. 2007).
Mycoremediation is a form of bioremediation in which fungi especially white rot fungus such as Phanerochaete chrysosporium degrade diverse range of persistent or toxic environmental contaminants (Singh 2006) (Table 5.2). The fungal mycelium secretes extracellular enzymes and acids that break down lignin and cellulose (Eaton 1985). Microfungi transform aromatic organopollutants cometabolically including polycyclic aromatic hydrocarbons (PAHs) and biphenyls, dibenzofurans, nitroaromatics, and various pesticides (Fritsche and Hofrichter 2008). Plant growth-promoting rhizobacteria (PGPR), endophytic bacteria and other rhizospheric bacteria have been shown to potentially degrade toxic organic compounds in contaminated soil (Sylvestre et al. 2009). Pseudomonas sp. specifically has shown potential for hydrocarbon-degrading capacity. Yeast species such as Trichosporon cutaneum also utilize aromatic compounds as growth substrates.
5.4 Mechanisms of Removal of Contaminants by Microbes
The inorganic contaminants removed by bacteria mainly include heavy metals and radionuclides. Heavy metals are removed via biosorption (metal sorption to cell surface by physicochemical mechanisms), bioleaching (heavy metal mobilization through the excretion of organic acids or methylation reactions), biomineralization (heavy metal immobilization through the formation of insoluble sulfides or polymeric complexes), intracellular accumulation and enzyme-catalyzed transformation (redox reactions) mechanisms (Lloyd and Lovley 2001). The resistance to heavy metal toxicity occurs by adsorption, uptake, methylation, oxidation, and reduction mechanism. Metals are also precipitated as insoluble sulfides via metabolic activity of sulfate-reducing bacteria. Heavy metal ions can be entrapped in the cellular structure and subsequently biosorbed onto the binding sites present in the cellular structure. They pass into the cell across the cell membrane through the cell metabolic cycle. Toxic radionuclides such as U and Th from nuclear waste streams are removed by similar mechanisms (PinakiSar et al. 2004).
Both anaerobic and aerobic bacteria are capable of metabolizing organic pollutants. The initial intracellular attack of organic pollutants is an oxidative process, and the enzymatic key reaction is catalyzed by oxygenases and peroxidases. Peripheral degradation pathways convert organic pollutants step by step into intermediates of the central intermediary metabolism. Cytochrome P450 alkane hydroxylases play an important role in the microbial degradation of oil, chlorinated hydrocarbons, fuel additives, and many other compounds. The degradation of hydrocarbons is carried out under aerobic condition and is mediated by specific enzyme system. Enzymes involved in degradation of xenobiotics mainly include oxygenases. Higher chlorinated PCBs are reduced by anaerobic microorganisms, while lower chlorinated biphenyls are oxidized by aerobic bacteria (Seeger et al. 2001). Aerobic catabolic pathway for PCB degradation involves steps catalyzed by enzymes, biphenyl dioxygenase (BphA), dihydrodiol dehydrogenase (BphB), 2,3-dihydroxybiphenyl dioxygenase (DHBD) (BphC) and hydrolase (BphD) (Taguchi et al. 2001).
Fungi are an important part of degrading microbiota because, like bacteria, they metabolize dissolved organic matter. Extracellular multienzyme complexes of fungi are efficient in breaking down the natural polymeric compounds. By means of their hyphal systems, they are also able to colonize and penetrate substrates rapidly and transport and redistribute nutrients within their mycelium (Fritsche and Hofrichter 2005). Hyphal penetration provides a mechanical adjunct to the chemical breakdown affected by the secreted enzymes. The high surface-to-cell ratio characteristic of filaments maximizes both mechanical and enzymatic contact with the environment. Second, the extracellular nature of the degradative enzymes enables fungi to tolerate higher concentrations of toxic chemicals. Among the filamentous fungi, the ligninolytic ones have been specifically investigated because of their extracellular, specific oxidoreductive enzymes that have been already successfully exploited in the degradation of many aromatic pollutants. Studies with Aspergillus niger AB10 Cd and Rhizopus arrhizus M1 have indicated Pb binding occurs via the functional groups on the cell surface. The functional groups act as ligands for metal sequestration (Pal et al. 2010). The proteins in the cell walls of AMF appear to have similar ability to sorb potentially toxic elements by sequestering them. Filamentous fungi may degrade pesticides using two types of enzymatic systems: intracellular (cytochromes P450) and exocellular (lignin-degrading system mainly consisting in peroxidases and lactases) (Chaplain et al. 2011). Yeast species use n-alkanes and other aliphatic hydrocarbons as a sole source of carbon, and energy is mediated by the existence of multiple microsomal cytochrome P450 forms.
5.5 Phytoremediation
The capacity of plants for removing and degrading various inorganic and organic contaminants from different components of the environment is referred as phytoremediation (Salt et al. 1998; Meagher 2000; Pilon-Smits 2005). It is a cost-effective, nonintrusive, aesthetically pleasing technology that removes contaminants via processes such as degradation, sequestration, or transformation mechanisms (Raskin and Ensley 2000; Garbisu et al. 2002; McCutcheon and Schnoor 2003). The major advantage of using this technology is that treatment can be done under in situ. The plants have been successfully used in removing contaminants such as explosives (trinitrotoluene), herbicides, pesticides and metals from different areas such as military areas, agricultural fields, industrial areas, mine tailings, sewage, municipal wastewater, drainage water and landfill leachate. The plants species with an effective remediation potential include mustard, alpine pennycress, hemp, and pigweed. The major concern about phytoremediation technology is that it is a time-consuming process and depends on the plant’s ability to grow and thrive in contaminated environment.
Potential of both terrestrial and aquatic plant species has been exploited for removing contaminants from the environment. Efficacy of phytoremediation varies according to varieties, cultivars, genotypes and type of pollutant (Dipu et al. 2011). The selection of the plant species is very crucial for the success of this technology. Plants with less maintenance and acclimatization in native climate conditions are favored. Each plant species depicts a variation in its ability to remove contaminants from the environment. The selection of plant species depends upon factors such as:
-
Tolerance to the environment
-
Uptake, translocation and accumulation ability of the plant
-
High growth rates and biomass production
-
Tolerance to environmental conditions such as drought, salinity, etc.
-
Availability of the species (annual/perennial)
Among terrestrial plants, trees and grass species with the characteristics such as deep roots, high biomass production, and fast growth are commonly preferred for remediation (EPA 1998; Schnoor 2000). Trees stabilize a pollutant and minimize spread of contaminant. Strong and dense root system (around 3 meters deep) in grasses assists in higher uptake of contaminants. The tolerance to extreme climatic variations such as drought, flood, submergence, fire, and heat and wide range of soil acidity, alkalinity, salinity and sodicity establish plants as ideal candidates for phytoremediation. Populus deltoides (hybrid poplar), Brassica juncea (Indian mustard), Helianthus annuus (sunflower), Thlaspi sp. including T. caerulescens and T. rotundifolium, Vetiveria zizanioides, and Paspalum conjugatum are some of the plant species with high capacity for removal of contaminants.
Among aquatic plant species, free-floating, submerged, and emergent forms exhibit exorbitant capacity for removal of various contaminants including heavy metals, radioactive wastes, nutrients, explosives, organic xenobiotics, and herbicides/pesticides from municipal and industrial wastewater. Features such as easy cultivation, high biomass production, faster growth rate, surplus availability and high tolerance to survive adverse environmental conditions assist in removal of contaminants and make them an ideal and most suitable candidate for use in phytoremediation technology. Aquatic plant species with high contaminant removal ability include Eichhornia crassipes (water hyacinth), Salvinia herzogii, Salvinia minima (water ferns), Pistia stratiotes (water lettuce), Nasturtium officinale (watercress), Spirodela intermedia, Lemna minor (duckweeds), Azolla pinnata (water velvet), Potamogeton pectinatus (American pondweed), Ceratophyllum demersum (coontail or hornwort), Myriophyllum spicatum (parrot feather), Typha latifolia (cattail), Phragmites (common reed) and Scirpus spp. (bulrush) (Dhir et al. 2009; Dhir 2013). Aquatic plants form an important component of constructed wetlands that remove many inorganic contaminants including metals, nitrates, phosphates, cyanides, as well as organic contaminants such as explosives and herbicides (Horne 2000; Jacobson et al. 2003; Dhir 2013).
5.5.1 Types of Phytoremediation
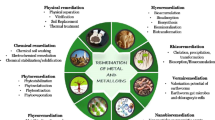
Plants remove contaminants by different processes such as phytoextraction/phytoaccumulation, phytodegradation/phytotransformation, phytovolatilization, rhizofiltration/phytofiltration and phytostabilization (Cunningham et al. 1995; Raskin et al. 1997; Salt et al. 1995a, 1998). Inorganic contaminants are removed by phytoextraction and/or phytostabilization processes, while organic contaminants are most commonly treated by phytodegradation and phytostimulation mechanisms.
In phytoextraction/phytoaccumulation process, contaminants are taken up by plants via roots followed by translocation to aboveground plant tissues, which are subsequently harvested (Salt et al. 1995a, b). It is used for removal of contaminants such as metals which cannot be degraded (Cd, Pb, Zn, Ni, Cr, Co, metalloids such as As, Se) and radionuclides (such as 90Sr, 137Cs, 238U). It is also referred as phytoaccumulation, phytoabsorption, phytosequestration, phytomining, or biomining.
In phytodegradation/phytotransformation process, the metabolization and degradation of contaminants takes place within the plant with the help of enzymes produced and released by them. Phytodegradation is most suited for moderately hydrophobic organic chemicals (octanol-water partition coefficients, log Kow = 0.5 ~ 3.0). Plant enzymes such as dehalogenase, peroxidase, nitroreductase, laccase and nitrilase assist in degradation of organic pollutants, such as 2,4,6-trinitrotoluene (TNT) and polychlorinated biphenyls (PCBs), polycyclic aromatic hydrocarbons (PAHs), herbicides, pesticides, BTEX (benzene, toluene, ethylbenzene, and xylene), chlorinated solvents such as trichloroethylene (TCE) and short-chain aliphatic chemicals, explosives such as 2,4,6-trinitrotoluene (TNT), and inorganic nutrients.
In phytovolatilization process, plants take up contaminants through roots followed by their release as volatile chemicals by shoot or leaf surfaces. Biomethylated forms of metals such as Se, As and Hg form volatile molecules (less toxic), which are lost to atmosphere. Selenium is converted to methyl selenate, and the volatile form is released in the atmosphere (Meagher 2000).
In rhizofiltration/phytofiltration process, plant roots absorb, precipitate, and remove contaminants from water in either a hydroponic or a constructed wetland. This process is applicable for removal of inorganic contaminants such as metals (Pb, Cd, Cu, Fe, Ni, Mn, Zn, Cr), nutrients and radionuclide (90Sr, 137Cs, 238U, 236U) present in groundwater, surface water and wastewater (Dushenkov et al. 1995, 1997a, b).
In phytostabilization process, plants immobilize or stabilize contaminants in the soil through accumulation by plant roots or precipitation in the soil by root exudates, thereby reducing the bioavailability of contaminants in the environment. The contaminants are sequestered from the soil and the process is efficient in removing inorganic and organic contaminants from the soils, sediments, and sludges. Contaminants also bind to humic (organic) matter through the process of humification. Phytostabilization of organic contaminants or metabolic by-products is also achieved by attaching to plant components such as lignin which is referred to as “phytolignification” (Cunningham et al. 1995).
5.5.2 Mechanism of Removal of Contaminants
5.5.2.1 Inorganic
Metals and radionuclides are captured by root cells and subsequently translocated to plant parts (symplastic). Metal uptake in plants also involves cation exchange by cell walls (apoplastic) (Williams et al. 2000; Pollard et al. 2000), or transport via symplastic pathway involves the role of membrane transport proteins (Blaylock and Huang 2000; Pollard et al. 2000). Intracellular high-affinity binding sites facilitate metal uptake across the plasma membrane (Dhankher et al. 2002; Hall 2002; Yang et al. 2005a, b). The natural resistance-associated macrophage (Nramp) family of proteins, cation diffusion facilitator (CDF) family proteins and zinc-iron permease (ZIP) family proteins (Williams et al. 2000) assist in metal transportation across the membranes. Metal chelate complexes are transported across the plasma membrane via specialized carriers. Cadmium is actively transported across the tonoplast of roots via a Cd/[H+] antiport. CPx-type heavy metal ATPase transport proteins use ATP to pump variety of charged substrates along Cu and/or Cd across cell membranes (Williams et al. 2000). ZIP proteins mainly transport potentially toxic metals (Zn) as well as nutrients (Fe). These include the iron transporter 1 (ITR1) gene of Arabidopsis which is an iron (Fe [II]) transporter. Subsequent to uptake and translocation, heavy metals are stored in vacuole. Final sequestration of metal ions in chelated form or phytochelatins takes place in vacuole (Kramer et al. 1996). Metals are sequestered by bonding with organic sulfur (R-SH) on the cysteine residues by formation of metallothioneins (MTs) and phytochelatins (PCs). Organic acids, viz., citrate and phytosiderophores such as mugenic and avenic acid chelate metal ions, increase the efficiency for uptake and translocation of metals.
Radionuclides are translocated to the aboveground plant parts through the vascular system via high-affinity K+ transporters. Translocation is followed by compartmentalization and complexation with ligands present in the cell including proteins, cysteine and glutathione. Radionuclides passively bind to negatively charged groups on the cell surface followed by transport to the cell wall. In active process, metabolically dependent penetration of ions through the cell membrane, movement inside cytoplasm and the bioaccumulation of the metal ions onto the protoplasts take place.
5.5.2.2 Organic
Organic contaminants (xenobiotic compounds) are subjected to partial or complete degradation within plants (Sandermann 1994). Plants absorb xenobiotics by simple diffusion primarily through roots and leaves (Wang and Liu 2007). Uptake and metabolism of hydrophobic organic contaminants is rapid. They are bound strongly to the surface of the roots especially by hemicellulose in the cell wall and the lipid bilayer of plant membranes; hence, their translocation within the plant is slow. They are actively transported through plant membranes (Meagher 2002; Pilon-Smits 2005). Several enzymes including monooxygenases, dioxygenases, dehydrogenases, hydrolases, peroxidases, nitroreductases, nitrilases, dehalogenases, phosphatases and carboxylesterases play an important role in degradation of xenobiotics (Dietz and Schnoor 2001; Pilon-Smits 2005). The detoxification of xenobiotic is carried out in three stages, namely, transformation, conjugation and sequestration.
Xenobiotics generally undergo transformation via chemical modification (oxidation, reduction, hydrolysis), conjugation (with glutathione, sugars, amino acids resulting in soluble, polar compounds), and sequestration or compartmentalization (conjugants are converted to other conjugates and deposited in plant vacuoles or bound to the cell wall and lignin) (Cherian and Oliveira 2005). Oxygenation increases water solubility and provides site for conjugation via glycosidic bond formation. The reaction is catalyzed by enzymes such as P450 monooxygenases, carboxylesterases, cytochrome P450 and peroxidases. Oxidation reactions are followed by reduction and/or hydrolysis reactions after which conjugation with glutathione (GSH), sugars, or organic acids takes place. Enzymes such as glutathione S-transferases, carboxylesterases, O-glucosyltransferases, O-malonyltransferases, N-glucosyltransferases and N-malonyltransferases are associated with xenobiotic metabolism.
The conjugation-sequestration involve coupling of glucose or malonyl group to the organic compound followed by the transport of the conjugate to the vacuole or the apoplast. Conjugated xenobiotics are then sequestered as part of insoluble cell wall polymers or in cellular compartments such as vacuoles and further metabolized to form CO2 (Pilon-Smits 2005). Cell compartmentation is mediated by a wide array of glutathione S-transferases (GSTs). ATP-binding cassette (ABC) transporters play a key role in the transfer of conjugates from the cytosol to either the vacuole or the apoplast (Klein et al. 2006).
The metabolism of certain nonagricultural contaminants such as PAHs, TCE, 2,4,6-trinitrotoluene (TNT), glyceryl trinitrate (GTN), and other chlorinated compounds has been well documented in literature (Macek et al. 2000; Alkorta and Garbisu 2001). Poplar trees have shown the potential of oxidizing alkanes, alkenes and methane and their halogenated analogues via dehalogenase enzyme. Dehalogenase(s) ultimately mineralize TCE to CO2 via an oxidative pathway.
5.6 Factors Affecting Bioremediation Process
The bioremediation processes is regulated by many factors. These mainly include metabolic capacity of the organism, availability of contaminants, and the environmental factors such as type of soil, temperature, pH and the presence of oxygen and nutrients. The compounds either serve as primary or secondary substrate to the organism (Boopathy 2000). Type of contaminants, their concentration and the physicochemical bioavailability of pollutants critically regulate the biodegradation potential. The growth and activity of microbes is affected by pH, temperature and moisture. The rate of enzymatic reactions within microorganisms is also regulated by temperature. After every 10 °C rise in temperature, the rate of biochemical reactions gets doubled due to increase in enzymatic activity. Bacteria found in soil are mesophiles and degrade petroleum hydrocarbons at an optimum temperature ranging from 25 °C to 45 °C. Soil pH is one of the important factors because it affects survival of microbial species and also affects availability of nutrients. Biodegradation of organic contaminants is optimal at a pH range of pH 6–8. Moisture influences the rate of contaminant metabolism because it influences the kind and amount of soluble materials that are available as well as the osmotic pressure and pH of terrestrial and aquatic systems.
Aerobic or anaerobic conditions also decide the rate and extent of biodegradation process. Hydrocarbons are readily degraded under aerobic conditions, whereas chlorinate compounds are degraded only in anaerobic ones. Stimulation of microorganisms is achieved by the addition of growth substances, nutrients, terminal electron acceptor/donors, or some combination, thereby resulting in an increase in organic pollutant degradation and biotransformation. The process of bioremediation can be enhanced by supplementing microorganisms with nutrients, carbon sources, or electron donors. Establishment of such microbial consortia can be done in several ways, e.g., by promoting growth through addition of nutrients, by adding terminal electron acceptor, or by controlling moisture and temperature conditions (Agarwal 1998). Addition of supplements such as fertilizers, oxygen, etc. assists in bioremediation as they act as biostimulants. Sufficient amount of nutrient and oxygen must be available in a usable form and in proper proportions for unrestricted microbial growth to occur.
Among the biological factors, metabolic ability of microorganisms affects the microbial degradation of organic compounds. The capacity of the plants to remove contaminants varies according to varieties, cultivars or genotypes and type of pollutant (Dipu et al. 2011). The selection of the plant species is very crucial. Plants with less maintenance, acclimatization to varied climate conditions, and increased biomass production are favored. The tolerance to contaminant also regulates the extent of contaminant removal capacity of plants.
5.7 Success Stories in Bioremediation
In situ bioremediation of U-contaminated sites has been conducted successfully with Desulfosporosinus spp. and Closteridium spp. (Bruschi and Florence 2006). Consortium of SRB (sulfate-reducing bacteria) has been used successfully to remove Zn and sulfate. The metals were precipitated as sulfides. Eight months after project implementation, 80% reduction in Site COC comprised a complex mixture of halogenated organic compound (mixture of brominated and chlorinated organic compounds). A company named TMPD technologies, Lafayette, LA, treated acres of land with multiple contaminants ranging from PCBs to hydrocarbons using microbes. It also removed oil spill from Lake Charles Refinery in Lake Charles, LA, via bioremediation techniques involving biostimulation and bioaugmentation. The Microbiological Resource Centers (MIRCENs) at Cairo, Egypt, is examining the use of microbes in degrading persistent pesticides pollutants.
The companies such as Edenspace Systems Corporation of the USA have successfully used Indian mustard plant to treat the soil contaminated with radionuclide strontium (Sr89/90) at Fort Greely in Alaska, USA and Cs137 from the contaminated pond waters (Singh et al. 2006). Indian mustard plant was used with sunflower (Helianthus annuus) to phytoremediate the Pb-contaminated soil at industrial facility in Connecticut, USA (Singh et al. 2006). Plants remove contamination by bioaccumulation in aerial parts. The Phytotech, Florida, USA, used the Indian mustard plant to remediate Pb and Cd from contaminated soil at the Czechowice Oil Refinery, Katowice, in Poland (Singh et al. 2006). In Milwaukee, Wisconsin, USA, the Ecolotree Inc. used the hybrid poplar trees to phytoremediate soil and groundwater contamination with petroleum-related organics, PAHs, and chlorinated organic compounds. In Illinois, USA, the Ecolotree Inc. used the hybrid poplar to treat soil contaminated with chemical fertilizer and pesticides. Hybrid polar was successfully used by Phytokinetics Inc., USA, to treat groundwater contaminated with chlorinated volatile organics including dichlorobenzidines and soils contaminated by gasoline and diesel compounds at old gas filling station at Axelved, Denmark, and cyanide, PAHs, oil, and BTEX (benzene, toluene, ethylbenzene, and xylene) in Denmark.
5.8 Genetic Engineering Approach for Improving Bioremediation
The genetic engineering technology has proved useful in improving the bioremediation process (Rugh et al. 1998; Bizily et al. 1999; Joutey et al. 2014). Recombinant DNA techniques enhance the capacity of organisms for degradation and breakdown of toxicants such as hydrocarbons and pesticides. Recombinant DNA techniques help to create organism with an ability to metabolize xenobiotics by detection of genes responsible for enzymes involved in degradation. Transgenic plants show improved metal tolerance, accumulation and enhanced capacity for degradation of organic compounds (Meagher 2000; Kramer and Chardonnens 2001; Pilon-Smits 2005). The genes encoding for biodegradative enzymes are present in chromosomal and extrachromosomal DNA of microbes. Plasmid exchange results in the production of novel microbial strains with a large number of degradative capabilities.
Inorganic contaminant removal is achieved via plants engineered to improve pollutant uptake by overexpression or knockdown of specific membrane transporter proteins or enzymes, root-shoot translocation abilities, sequestration and volatilization. The expression of the introduced gene is regulated by promoters. The protein may be directed to different cellular compartments, such as the chloroplast, the vacuole, or the cell wall. Various transgenic plants were created with metal tolerance and accumulation properties, either by overexpression of membrane transporter proteins (Hirschi et al. 2000; Song et al. 2003) or by overproduction of chelator molecules (Zhu et al. 1999a, b; Dhankher et al. 2002). Transgenic plants have been raised by transfer of metal hyperaccumulator genes to high-biomass, fast-growing species (Chaney et al. 2000; LeDuc et al. 2004). Synthesis of metal chelators leading to enhanced metal uptake, translocation, and sequestration has been overexpressed in plants (Cherian and Oliveira 2005; Pilon-Smits 2005). The biosynthesis of MTs is regulated at the transcriptional level and is induced by several factors, such as hormones, cytotoxic agents, and metals, including Cd, Zn, Hg, Cu, Au, Ag, Co, Ni, and Bi (Yang et al. 2005a, b). Phytochelatins are a class of posttranslationally synthesized (cysteine-rich metal-chelating) peptides that play a pivotal role in heavy metal tolerance in plants by chelating these substances and decreasing their free concentrations (Vatamaniuk et al. 1999). Metal-tolerant tobacco (Nicotiana tabacum) has been developed by expressing a yeast metallothionein gene for higher tolerance to Cd. Brassica juncea was genetically engineered with E. coli gshl gene for increased glutathione and phytochelatin production for high Cd tolerance and high concentrations of phytochelatins (Fulekar et al. 2009). Overexpression of a bacterial glutathione synthetase (GS) for higher GSH and PC concentrations and increased Cd tolerance/accumulation by Brassica juncea has also been noted. Overexpression of plant phytochelatin synthase (PS) in transgenic yeast increases tolerance and accumulation of Cd. Manipulation of GSH and PC concentrations increases potential for increasing the accumulation of toxic metals by plants. Abhilash et al. (2009) reported the introduction of genes for enzyme glutathione S-transferase (GST) (responsible for GSH synthesis), by introduction of a [gamma]-glutathione synthetase into Populus trichocarpa (Gullner et al. 2001). For heavy metals, Sriprang et al. (2003) introduced Arabidopsis thaliana gene for phytochelatin synthase (PCS; PCSAt) into Mesorhizobium huakuii subsp. rengei strain B3 and then established the symbiosis between M. huakuii subsp. rengei strain B3 and Astragalus sinicus. The gene was expressed to produce phytochelatins and accumulate Cd2+, under the control of bacteroid-specific promoter, the nifH gene.
Some genes for increased heavy metal (Cd) resistance and uptake, like AtNramps (Thomine et al. 2000), AtPcrs (Song et al. 2004) and CAD1 (Ha et al. 1999) from Arabidopsis thaliana; gshI, gshII (Zhu et al. 1999a), and PCS cDNA clone (Heiss et al. 2003) from Brassica juncea, tobacco (Goto et al. 1998) and rice (Goto et al. 1998); ferritin from soybean for increased Fe accumulation; and merA from bacteria to A. thaliana and tobacco for resistance to Hg with gene (Bizily et al. 1999; Eapen and D’Souza 2005), have been introduced into plants. Transgenics have also been raised for Se tolerance with a bacterial glutathione reductase in the cytoplasm and chloroplast for Indian mustard. Transgenic A. thaliana plants expressing SRSIp/ArsC and ACT 2p/γ-ECS with high tolerance to As than wild plants and transgenic plants expressing γ-ECS or ArsC alone have also been reported (Dhankher et al. 2002; Mello-Farias and Chaves 2008). Studies also report overexpression of 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase for an enhanced metal accumulation (Eapen and D’Souza 2005).
The genes for phytovolatilization have also been introduced into plants. Introduction of bacterial mercury reductase (MerA) and organomercurial lyase (MerB) genes into plants such as Arabidopsis thaliana increases plants’ tolerance to Hg. Toxic organic mercuric compounds are converted into volatile elemental Hg (Rugh et al. 1996; Bizily et al. 2000; Dhankher et al. 2002; Eapen and D’Souza 2005). Overexpression of two key enzymes, cystathionine gamma-synthase and selenocysteine methyltransferase, which promote the conversion of selenocysteine to volatile Se has also been reported (van Huysen et al. 2003; LeDuc et al. 2004). Transgenic plants engineered to have enhanced sulfate/selenate reduction showed fivefold higher Se accumulation in the field (Bañuelos et al. 2005). Transgenic Arabidopsis plants which could transport oxyanion arsenate to aboveground, reduce to arsenite, and sequester it to thiol peptide complexes by transfer of Escherichia coli C and γ-ECS genes have been developed (Eapen and D’Souza 2005).
The degradation of organic pollutant can be improved by overexpressing enzymes that facilitate degradation in plant tissue or rhizosphere. The genes procured from other organisms such as bacteria or mammals are introduced in plants. The transformed organisms possess the enzymatic machinery required to achieve a complete mineralization of organic molecules. Specific proteins or peptides for binding and transporting xenobiotics and enzymes involved in biodegradation have been introduced or overexpressed in plants to achieve compete degradation. The genetically transformed plants for degrading herbicides, organomercurials, phenolic compounds, PCBs and nitroaromatics (Bizily et al. 1999; Karavangeli et al. 2005; Rylott et al. 2006; Mohammadi et al. 2007) include Arabidopsis, tobacco (Nicotiana tabacum), Indian mustard (Brassica juncea), hybrid poplar (Populus sp.), and yellow poplar (Liriodendron sp.). Transgenic wetland species include Spartina spp., reeds and Typha spp. (Czako et al. 2005). Abhilash et al. (2009) reported the introduction of genes and enzymes such as mammalian cytochrome p450s gene into rice plant.
The genes coding for cytochrome P450 and GST for the enhanced degradation and remediation of herbicides, explosives, PCBs etc. have been overexpressed in plants. Increased expression of extracellular enzymes laccases, peroxidases, and cytochrome P450 has been proposed as an approach for remediation of small organic compounds (Doty 2008). Pseudomonas putida MHF 7109 isolated from cow dung has shown ability for biodegradation of petroleum hydrocarbon compounds – benzene, toluene and o-xylene (BTX). The bacterium Deinococcus radiodurans (the most radioresistant organism known) has been modified to consume and digest toluene and Hg from highly radioactive nuclear waste. Transgenic poplar trees and tobacco plants overexpressing a mammalian cytochrome P450 2E1 (CYP2E1) and human cytochrome P450 2E1 were developed with the capacity for metabolizing trichloroethylene (TCE). Rabbit cytochrome P450 has been introduced in Atropa belladonna to facilitate faster metabolism of TCE. Transgenic plants removed organic compounds as high as 79% of TCE, 49% of vinyl chloride, and 40% of benzene in comparison to 10–30% controls. Bacterial genes dhlAB from Xanthobacter improved removal and degradation of 1,2-dichlorethane in plants. Higher expression of genes responsible for root development has been targeted for effective remediation of atrazine and alachlor. The expression of bacterial genes atrazine chlorohydrolase (AtzA) and 1-aminocyclopropane-l-carboxylate deaminase has shown promising role in remediation of atrazine and alachlor (Wang et al. 2008). Hydrophilic organics cannot pass the hydrophobic interior of membranes passively as there is no suitable transporter available in the plant. Hydrophobic organic contaminants stick to soil particles, thereby reducing their bioavailability, or become stuck inside root membranes preventing their movement into the cell’s interior. Rhizoremediation utilizes the capacity of plant-associated microbes that have been proposed for remediation of PCBs (Doty 2008; Rylott and Bruce 2009). The degradation of PCBs takes place in two steps. In the first step, PCB degradation takes place by expressing the genes of first multicomponent enzyme biphenyl 2,3-dioxygenase in degradation pathway. The released intermediate compounds undergo further transformation by rhizospheric bacteria. In the second step, expression of 2,3-dihydroxybiphenyl dioxygenase enzyme harbors bphC and avoids plants’ inability to cleave toxic dihydroxybiphenyls. These transgenic plants are more resistant to PCBs than wild type indicating the potential utility of plants for effective rhizoremediation of PCBs.
Shiota et al. (1994) made transgenic tobacco plants by fusing rat P450 1A1 to yeast NADPH P450 oxidoreductase for metabolizing the herbicide chlortoluron. Helianthus tuberosus CYP76B1 and Glycine max CYP71A10 were the first transgenic plant with enzymes to actively metabolize organic contaminants (Siminszky et al. 1999). Human P450s have been shown to significantly enhance herbicide tolerance in transgenic potato (Solanum tuberosum L.) (Inui et al. 2001), rice (Oryza sativa L.) (Kawahigashi et al. 2007), Arabidopsis and tobacco (Nicotiana tabacum L.) (Didierjean et al. 2002).
Transgenic plants have been developed by introducing genes that are able to degrade explosive nitrate esters and NACs by introducing the bacterial enzyme pentaerythritol tetranitrate reductase (French et al. 1999). Van Aken (2008) reported the development of transgenic plants for remediation of 2,4,6-trinitrotoluene, hexahydro-l,3,5-trinitro-l,3,5-triazine and glyceroltrinitrate by introducing and expressing bacterial nitroreductases and cytochrome P450s. Plants expressing these genes show significantly increased tolerance, uptake and detoxification of the targeted explosives. The introduction of the pnrA gene encoding for nitroreductase from Pseudomonas putida into a fast-growing tree aspen has shown promising results for remediation of explosives in contaminated field conditions (Rylott and Bruce 2009; James and Strand 2009). Transgenic approaches increased the ability of tobacco to degrade explosives such as GTN and TNT by overexpressing a bacterial NADPH-dependent nitroreductase (French et al. 1999). The genes encoding a nitroreductase from a bacterium have been inserted in tobacco, and the transformed species showed faster removal of TNT and enhanced resistance to the toxic effects of TNT.
Genetically engineered microorganisms (GEMs) have enhanced degrading capabilities of a wide range of chemical contaminants. The principles involved in the development of GEM plants include (1) modification of enzyme specificity and affinity; (2) pathway construction and regulation; (3) bioprocess development, monitoring, and control; and (4) bioaffinity bioreporter sensor applications for chemical sensing, toxicity reduction and end point analysis. Genes responsible for degradation of environmental pollutants, for example, toluene, chlorobenzene acids and other halogenated pesticides and toxic wastes, have been identified. For every compound, one separate plasmid is required. It is not like that one plasmid can degrade all the toxic compounds of different groups. The plasmids are grouped into four categories: (1) OCT plasmid which degrades octane, hexane and decane, (2) XYL plasmid which degrades xylene and toluenes, (3) CAM plasmid that decomposes camphor, and (4) NAH plasmid which degrades naphthalene. The potential for creating, through genetic manipulation, microbial strains able to degrade a variety of different types of hydrocarbons has been demonstrated. They successfully produced a multiplasmid-containing Pseudomonas strain capable of oxidizing aliphatic, aromatic, terpenic, and polyaromatic hydrocarbons. Pseudomonas putida that contained the XYL and NAH plasmid as well as a hybrid plasmid derived by recombinating parts of CAM and OCT developed by conjugation could degrade camphor, octane, salicylate, and naphthalene and could grow rapidly on crude oil because it was capable of metabolizing hydrocarbons more efficiently than any other single plasmid. This product of genetic engineering was called as superbug (oil eating bug). The plasmids of P. putida degrading various chemical compounds are TOL (for toluene and xylene), RA500 (for 3,5-xylene), pAC 25 (for 3-cne chlorobenxoate), and pKF439 (for salicylate toluene). Plasmid WWO of P. putida is one member of a set of plasmids now termed as TOL plasmid. Alcaligenes eutrophus AE104 (pEBZ141) was used for chromium removal from industrial wastewater, and the recombinant photosynthetic bacterium, Rhodopseudomonas palustris, was constructed to simultaneously express mercury transport system and metallothionein for Hg2+ removal from heavy metal wastewater. For polychlorinated biphenyl degradation, chromosomally located PCB catabolic genes of R. eutropha A5, Achromobacter sp. LBS1C1, and A. denitrificans JB1 were transferred into a heavy metal-resistant strain R. eutropha CH34 through natural conjugation.
5.9 Conclusions
Bioremediation is a natural process utilizing bacteria and fungi or plants to degrade or detoxify substances hazardous to human health and/or the environment. The microorganisms indigenous to a contaminated area or site aid in removal of contaminants. Biotechnology utilizes the application of genetic engineering to improve the efficiency of microorganisms to reduce the toxic substances. Bioremediation must be tailored to the site-specific conditions. More research is needed to develop and engineer bioremediation technologies that are appropriate for sites with complex mixtures of contaminants and are not evenly dispersed in the environment. This technology can be applied both in situ and ex situ for removing broad range of environmental contaminants, viz., organic and inorganic. Environmental conditions regulate the growth and degradation ability of organism. Resistance to degradation is some of the major concerns for bioremediation technology. A comprehensive understanding of the transport and sequestration mechanisms in plant cells is essential for formulating effective strategies to develop genetically engineered plants with higher phytoremediation efficiency. Genetic engineering of endophytic and rhizospheric bacteria can be used in plant-associated degradation of toxic compounds in soil and is considered one of the most promising new technologies for remediation of contaminated environmental sites.
References
Abhilash PC, Jamil S, Singh N (2009) Transgenic plants for enhanced biodegradation and phytoremediation of organic xenobiotics. Biotechnol Adv 27:474–488
Abruscia C, Marquinaa D, Del Amob A, Catalina F (2007) Biodegradation of cinematographic gelatin emulsion by bacteria and filamentous fungi using indirect impedance technique. Int Biodet Biodegr 60:137–114
Agarwal SK (1998) Environmental Biotechnology, 1st edn. APH Publishing Corporation, New Delhi, pp 267–289
Alkorta I, Garbisu C (2001) Phytoremediation of organic contaminants in soils. Bioresource Technol 79:273–276
Bañuelos G, Terr N, LeDuc DL et al (2005) Field trial of transgenic Indian mustard plants shows enhanced phytoremediation of selenium-contaminated sediment. Environ Sci Technol 39:1771–1777
Bizily SP, Rugh CL, Meagher RB (2000) Phytodetoxification of hazardous organomercurials by genetically engineered plants. Biotechnol 18:213–217
Bizily S, Rugh CL, Summers AO, Meagher RB (1999) Phytoremediation of methylmercury pollution: Mer B expression in Arabidopsis thaliana confers resistance to organomercurials. Proc Natl Acad Sci U S A 96:6808–6813
Blaylock MJ, Huang JW (2000) Phytoextraction of metals. In: Raskin I, Ensley BD (eds) Phytoremediation of toxic metals. Using plants to clean up the environment. Wiley, New York, pp 53–70
Boopathy R (2000) Factors limiting bioremediation technologies. Biores Technol 74:63–67
Bruschi M, Florence G (2006) New Bioremediation technologies to remove heavy metals and radionuclides using Fe (III)-sulfate- and sulfur reducing bacteria. In: Singh SN, Tripathi RD (eds) Environmental bioremediation technologies. Springer, New York, pp 35–55
de Cássia Miranda R, de Souza CS, de Barros Gomes E, Barros Lovaglio R, Edison Lopes C, de Fátima Vieira de Queiroz Sousa M (2007) Biodegradation of diesel oil by yeasts isolated from the vicinity of Suape Port in the state of Pernambuco – Brazil. Braz Arch Biol Technol 50:147–152
Chaney RL, Brown SL, Li YM, Angle JS, Stuczynski TI, Daniels WL, Henry CL, Siebelec G, Malik M, Ryan JA, Compton H (2000) Progress in risk assessment for soil metals, and in-situ remediation and phytoextraction of metals from hazardous contaminated soils. USEPA Phytoremediation: State of Science, Boston
Chaplain V, Défossez P, Richard G, Tessier D, Roger-Estrade J (2011) Contrasted effects of no-till on bulk density of soil and mechanical resistance. Soil Tillage Res 111:105–114
Cherian S, Oliveira MM (2005) Transgenic plants in phytoremediation: recent advances and new possibilities. Environ Sci Technol 39(24):9377–9390
Cunningham SD, Berti WR, Huang JW (1995) Phytoremediation of contaminated soils. Trends Biotechnol 13:393–397
Czako M, Feng X, He Y (2005) Genetic modification of wetland grasses for phytoremediation. Z Naturforsch 60:285–291
Dhankher OP, Li Y, Rosen BP, Shi J, Salt D, Senecoff JF, Sashti NA, Meagher RB (2002) Engineering tolerance and hyperaccumulation of arsenic in plants by combining arsenate reductase and gamma-glutamylcysteine synthetase expression. Nat Biotechnol 20:1140–1145
Dhir B (2013) Phytoremediation: role of aquatic plants in environmental clean-up. Springer. doi:10.1007/978–81–322-1307-9
Dhir B, Sharmila P, Pardha Saradhi P (2009) Potential of aquatic macrophytes for removing contaminants from the environment. Crit Rev Environ Sci Technol 39:754–781
Didierjean I, Gondet L, Perkins R, Lau SM, Schaller H, O’keefe DP, Werck-reichhart D (2002) Engineering herbicide metabolism in tobacco and Arabidopsis with CYP76B1, a cytochrome P450 enzyme from Jerusalem artichoke. Plant Physiol 130:179–189
Dietz AC, Schnoor JL (2001) Advances in phytoremediation. Environ Health Perspect 109:163–168
Dipu S, Kumar AA, Thanga VSG (2011) Phytoremediation of dairy effluent by constructed wetland technology. Environmentalist 31:263–278
Dos Santos AB, Cervantes JF, Van Lier BJ (2007) Review paper on current technologies for decolourisation of textile wastewaters: perspectives for anaerobic biotechnology. Biomagn Res Technol 98:2369–2385
Doty SL (2008) Enhancing phytoremediation through the use of transgenics and endophytes. New Phytol 179:318–333
Dushenkov V, Kumar PBAN, Motto H, Raskin I (1995) Rhizofiltration: the use of plants to remove heavy metals from aqueous streams. Environ Sci Technol 29:1239–1245
Dushenkov S, Vasudev D, Kapulnik Y, Gleba D, Fleisher D, Ting KC, Ensley B (1997a) Removal of uranium from water using terrestrial plants. Environ Sci Technol 31:3468–3474
Dushenkov S, Vasudev D, Kapulnik Y, Gleba D, Fleisher D, Ting KC, Ensley B (1997b) Phytoremediation: a novel approach to an old problem. In: Wise DL (ed) Global environmental biotechnology. Elsevier Science BV, Amsterdam, pp 563–572
Eapen S, D’Souza SF (2005) Prospects of genetic engineering of plants for phytoremediation of toxic metals. Biotechnol Adv 23:97–114
Eaton DC (1985) Mineralization of polychlorinated biphenyls by Phanerochaete chrysosporium: a ligninolytic fungus. Enzym Microb Technol 7:194–196
El Fantroussi S, Agathos SN (2005) Is bioaugmentation a feasible strategy for pollutant removal and site remediation? Curr Opin Microbiol 8:268–275
El Fantroussi S, Belkacemi M, Top EM, Mahillon J, Naveau H, Agathos SN (1999) Bioaugmentation of a soil bioreactor designed for pilot-scale anaerobic bioremediation studies. Environ Sci Technol 33:2992–3001
EPA (Environmental Protection Agency) (1998) A citizen’s guide to phytoremediation. EPA Publication, Washington, DC. 542-F-98-011
EPA (Environmental Protection Agency) (1999) Phytoremediation resource guide. EPA Publication, Washington, DC. 542-B-99-003
EPA (2003) Annual report: revised draft. Environmental Protection Agency, Accra
French CE, Rosser SJ, Davies GJ, Nicklin S, Bruce NC (1999) Biodegradation of explosives by transgenic plants expressing pentaerythritol tetranitrate reductase. Nat Biotechnol 17:491–494
Fritsche W, Hofrichter M (2005) Aerobic degradation of recalcitrant organic compounds by microorganisms. In: Jördening HJ, Winter J (eds) Environmental biotechnology, concepts and applications. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim. doi: 10.1002/3527604286.ch7
Fritsche W, Hofrichter M (2008) Aerobic degradation by microorganisms. In: Rehm HJ, Reed G (eds) Biotechnology set, 2nd edn. Wiley-VCH Verlag GmbH, Weinheim. doi:10.1002/9783527620999.ch6m
Fulekar MH, Geetha M, Sharma J (2009) Bioremediation of Trichlorpyr Butoxyethyl Ester (TBEE) in bioreactor using adapted Pseudomonas aeruginosa in scale up process technique. Biol Med 1(3):1–6
Garbisu C, Hernández-Allica J, Barrutia O, Alkorta I, Becerril JM (2002) Phytoremediation: a technology that uses green plants to remove contaminants from polluted areas. Rev. Environ Sci Health 17:173–188
Goto F, Yoshihara T, Saiki H (1998) Iron accumulation in tobacco plants expressing soybean ferritin gene. Transgenic Res 7:173–180
Gullner G, Kömives T, Rennenberg H (2001) Enhanced tolerance of transgenic poplar plants overexpressing gamma-glutamylcysteine synthetase towards chloroacetanilide herbicides. J Exp Bot 52:971–979
Ha SB, Smith AP, Howden R, Dietrich WM, Bugg S, O’Connell MJ, Goldsbrough PB, Cobbett CS (1999) Phytochelatin synthase genes from Arabidopsis and the yeast Schizosaccharomyces pombe. Plant Cell 11:1153–1163
Hall JL (2002) Cellular mechanisms for heavy metal detoxification and tolerance. J Exp Bot 53:1–11
Heiss S, Wachter A, Bogs J, Cobbett C, Rausch T (2003) Phytochelatin synthase (PCS) protein is induced in Brassica juncea leaves after prolonged Cd exposure. J Exp Bot 54:1833–1839
Hirschi KD, Korenkov VD, Wilganowski NL, Wagner GJ (2000) Expression of Arabidopsis CAX2 in tobacco altered metal accumulation and increased manganese tolerance. Plant Physiol 24:125–133
Horne AJ (2000) Phytoremediation by constructed wetlands. In: Terry N, Banuelos G (eds) Phytoremediation of contaminated soil and water. Lewis, Boca Raton, pp 13–40
Inui H, Shiota N, Motoi Y, Ido Y, Inoue T, Kodama T, Ohkawa Y, Ohkawa H (2001) Metabolism of herbicides and other chemicals in human cytochrome P450 species and in transgenic potato plants co-expressing human CYP1A1, CYP2B6 and CYP2C19. J Pest Sci 26:28–40
Jacobson ME, Chiang SY, Gueriguian L, Weshtholm LR, Pierson J (2003) Transformation kinetics of trinitrotoluene conversion in aquatic plants. In: McCutcheon SC, Schnoor JL (eds) Phytoremediation: transformation and control of contaminants. Wiley, New York, pp 409–427
James CA, Strand SE (2009) Phytoremediation of small organic contaminants using transgenic plants. Curr Opin Biotechnol 20(2):237–241
Joutey NT, Bahafid W, Saye H, El Ghachtouli N (2014) Biodegradation: involved microorganisms and genetically engineered microorganisms. In: Chamy R, Rosenkranz F (eds) Biodegradation – life of science. Intech, Rijeka. ISBN 978-953-51-1154-2
Kanade SN, Ade AB, Khilare VC (2012) Malathion degradation by Azospirillum lipoferum Beijerinck. Sci Res Rep 2(1):94–103
Karavangeli M, Labrou NE, Clonis YD, Tsaftarisa A (2005) Development of transgenic tobacco plants overexpressing maize glutathione S-transferase. Biomol Eng 22:121–128
Kawahigashi H, Hirose S, Ohkawa H, Ohkawa Y (2007) Herbicide resistance of transgenic rice plants expressing human CYP1A1. Biotechnol Adv 25:75–84
Klein M, Burla B, MAartinoia E (2006) The multidrug resistance-associated protein (MRP/ABCC) subfamily of ATP-binding cassette transporters in plants. FEBS Lett 580:1112–1122
Kramer U, Chardonnens AN (2001) The use of transgenic plants in the bioremediation of soils contaminated with trace elements. Appl Microbiol Biotechnol 55:661–672
Kramer U, Cotter-Howells JD, Charnock JM, AJM B, JAC S (1996) Free histidine as a metal chelator in plants that accumulate nickel. Nature 379:635–638
Kumar A, Bisht BS, Joshi VD, Dhewa T (2011) Review on bioremediation of polluted environment: a management tool. Int J Environ Sci 1:1079–1093
LeDuc DL, Tarun AS, Montes-Bayon M et al (2004) Overexpression of selenocysteine methyltransferase in Arabidopsis and Indian mustard increases selenium tolerance and accumulation. Plant Physiol 135:377–383
Li CH, Wong YS, Tam NF (2010) Anaerobic biodegradation of polycyclic aromatic hydro- carbons with amendment of iron(III) in mangrove sediment slurry. Bioresour Technol 101:8083–8092
Lloyd JR, Lovley DR (2001) Microbial detoxification of metals and radionuclides. Curr Opin Biotechnol 12:248–253
Macek T, Mackova M, Kas J (2000) Exploitation of plants for the removal of organics in environmental remediation. Biotechnol Adv 18:23–34
Maheshwari R, Singh U, Singh P, Singh N, Jat BL, Rani B (2014) To decontaminate wastewater employing bioremediation technologies. J Adv Sci Res 5(2):7–15
McCutcheon SC, Schnoor JL (2003) Overview of phytotransformation and control of wastes. In: SC MC, Schnoor J (eds) Phytoremediation: transformation and control of contaminants. Wiley, New York, pp 53–58
Meagher RB (2000) Phytoremediation of toxic elemental and organic pollutants. Curr Opin Plant Biol 3:153–162
Meagher M, Taper M, Jerde C (2002) Recent changes in population distribution: the Pelican bison and the domino effect. In: Anderson RJ, Harmon D (eds) Yellowstone Lake: hotbed of chaos or reservoir of resilience? Proceedings of the 6th biennial scientific conference on the Greater Yellowstone Ecosystem October 8–10, 2001, Mammoth Hot Springs Hotel, Yellowstone National Park, Wyo., and Hancock, Mich.: Yellowstone Center for Resources and the George Wright Society, pp 135–147
Mello-Farias PC, Chaves ALS (2008) Biochemical and molecular aspects of toxic metals phytoremediation using transgenic plants. In: Tiznado-Hernandez ME, Troncoso-Rojas R, Rivera-Domínguez MA (eds) Transgenic approach in plant biochemistry and physiology. Research Signpost, Kerala, pp 253–266
Mohammadi M, Chalavi V, Novakova-Sura M, Laliberte JF, Sylvestre M (2007) Expression of bacterial biphenyl-chlorobiphenyl dioxygenase genes in tobacco plants. Biotechnol Bioeng 97:496–505
Mucha AP, Almeida CMR, Bordalo AA, Vasconcelos MTSD (2010) LMWOA (low molecular weight organic acid) exudation by salt marsh plants: natural variation and response to Cu contamination. Estuar Coast Shelf Sci 88:63–70
Olaniran AO, Pillay D, Pillay B (2006) Biostimulation and bioaugmentation enhances aero- bic biodegradation of dichloroethenes. Chemosphere 63:600–608
Olson PE, Reerdon KF, Pillon-Smith EAH (2003) Ecology of rhizosphere bioremediation. In: McCutcheon SC, Schnoor JL (eds) Phytoremediation transformation and control of contaminants. Wiley, Hoboken, pp 317–353
Pal TK, Bhattacharyya S, Basumajumdar A (2010) Cellular distribution of bioaccumulated toxic heavy metals in Aspergillus niger and Rhizopus arrhizus. Int J Pharma Biol Sci 1:1–6
Pandey B, Fulekar MH (2012) Bioremediation technology: a new horizon for environmental clean-up. Biol Med 4(1):51–59
Pilon-Smits E (2005) Phytoremediation. Annu Rev Plant Biol 56:15–39
PinakiSar S, Kazy K, D’Souza SF (2004) Radionuclide remediation using a bacterial biosorbent. Int Biodeter Biodegr 54(2–3):193–202
Pollard AJ, Dandridge KL, Jhee EM (2000) Ecological genetics and the evolution of trace element hyperaccumulation in plants. In: Terry N, Bañuelos G (eds) Phytoremediation of contaminated soil and water. Lewis Publishers, Boca Raton, pp 251–264
Raskin I, Ensley BD (2000) Phytoremediation of toxic metals: using plants to clean up the environment. Wiley, New York
Raskin I, Smith RD, Salt DE (1997) Phytoremediation of metals: using plants to remove pollutants from the environment. Curr Opin Biotechnol 8:221–226
Rugh CL, Wilde HD, Stack NM, Thompson MD, Summers AO, Meagher RB (1996) Mercuric ion reduction and the resistance in transgenic Arabidopsis thaliana plants expressing a modified bacterial mer A gene. Proc Natl Acad Sci U S A 93:3182–3187
Rugh CL, Senecoff JF, Meagher RB, Merkle SA (1998) Development of transgenic yellow poplar for mercury phytoremediation. Nat Biotechnol 16(10):925–928
Rylott EL, Bruce NC (2009) Plants disarm soil: engineering plants for phytoremediation of explosives. Trends Biotechnol 29:73–81
Rylott EL, Jackson RG, Edwards J, Womack GL, Seth-Smith HMB, Rathbone DA, Strand SE, Bruc NC (2006) An explosive-degrading cytochrome P450 activity and its targeted application for the phytoremediation of RDX. Nat Biotechnol 24:216–219
Salt DE, Blaylock MB, Kumar NP, Dushenkov V, Ensley BD, Chet I, Raskin I (1995a) Phytoremediation: a novel strategy for the removal of toxic metals from the environment using plants. Biotechnology 13:468–474
Salt DE, Prince RC, Pickering IJ, Raskin I (1995b) Mechanisms of cadmium mobility and accumulation in Indian mustard. Plant Physiol 109:1427–1433
Salt DE, Smith RD, Raskin I (1998) Phytoremediation. Annu Rev Plant Physiol Plant Mol Biol 49:643–668
Sandermann H (1994) Higher plant metabolism of xenobiotics: the green liver concept. Pharmacogenetics 4:225–241
Sasek V, Volfova O, Erbanova P, Vyas BRM, Matucha M (1993) Degradation of PCBs by white rot fungi, methylotrophic and hydrocarbon utilizing yeasts and bacteria. Biotechnol Lett 15:521–526
Schnoor JL (2000) Phytostabilization of metals using hybrid poplar trees. In: Raskin I, Ensley BD (eds) Phytoremediation of toxic metals – using plants to clean-up the environment. Wiley, New York, pp 133–150
Seeger M, Cámara B, Hofer B (2001) Dehalogenation, denitration, dehydroxylation, and an- gular attack on substituted biphenyls and related compounds by a biphenyl dioxygenase. J Bacteriol 183:3548–3555
Shan HF, Kurtz HD, Freedman DL (2010) Evaluation of strategies for anaerobic bioremediation of high concentrations of halomethanes. Water Res 44:1317–1328
Sharma J, Fulekar MH (2009) Potential of Citrobacter freundii for bioaccumulation of heavy metal – copper. Biol Med 1(3):7–14
Shiota N, Nagasawa A, Sakaki T, Yabusaki Y, Ohkawa H (1994) Herbicide-resistant plants expressing the fused enzyme between rat cytochrome P4501A1 (CYP1A1) and yeast NADPH-cytochrome P450 reductase. Plant Physiol 106:17–23
Siminszky B, Corbinf T, Warde R, Fleischmann TJ, Dewey RE (1999) Expression of a soybean cytochrome P450 monooxygenase cDNA in yeast and tobacco enhances the metabolism of phenylurea herbicides. Proc Natl Acad Sci U S A 96:1750–1755
Singh H (2006) Mycoremediation: fungal bioremediation. Wiley-Interscience, New York
Singh D, Fulekar MH (2009) Benzene bioremediation using cow dung microflora in two phase partitioning bioreactor. J Hazard Mater 175:336–343
Singh RP, Dhania G, Sharma PA, Jaiwal K (2006) Biotechnological approaches to improve phytoremediation efficiency for environment contaminants. In: Singh SN, Tripathi RD (eds) Environmental bioremediation technologies. Springer, New York, pp 223–258
Song WY, Sohn EJ, Martinoia E, Lee YJ, Yang YY, Jasinski M, Forestier C, Hwang I, Lee Y (2003) Engineering tolerance and accumulation of lead and cadmium in transgenic plants. Nat Biotechnol 21:914–919
Song WY, Martinoia E, Lee J, Kim D, Kim DY, Vogt E, Shim D, Choi KS, Hwang I, Lee Y (2004) A novel family of cys-rich membrane proteins mediates cadmium resistance in Arabidopsis. Plant Physiol 135:1027–1039
Sriprang R, Hayashi M, Hisayo O, Masahiro T, Kazumasa H, Yoshikatsu M (2003) Enhanced accumulation of Cd by a Mesorhizobium sp. transformed with a gene from Arabidopsis thaliana coding for phytochelatin synthase. Appl Environ Microbiol 69:1791–1796
Strong PJ, Burgess JE (2008) Treatment methods for wine-related and distillery wastewaters review. Bioremed J 12:70–87
Surekha Rani M, Vijaya Lakshmi K, Devi SP, Jaya MR, Aruna S, Jyothi K, Narasimha G, Venkateswarlu K (2008) Isolation and characterization of a chlorpyrifos degrading bacterium from agricultural soil and its growth response. Afr J Microbiol Res 2:26–031
Sylvestre M., Macek T, Mackova M (2009) Transgenic plants to improve rhizoremediation of polychlorinated biphenyls (PCBs). Curr Opin Biotechnol 20: 242–247
Taguchi K, Motoyama M, Kudo T (2001) PCB/biphenyl degradation gene cluster in Rhodococcus rhodochrousK37 is different from the well-known bph gene clusters in Rhodococcus sp. P6, RHA1, and TA42. RIKEN Rev 42:23–26
Takeuchi M, Nanba K, Iwamoto H, Nirei H, Kusuda T, Kazaoka O, Owaki M, Furuya K (2005) In situ bioremediation of a cis-dichloroethylene-contaminated aquifer utilizing methane-rich groundwater from an uncontaminated aquifer. Water Res 39:2438–2444
Thierry L, Armelle B, Karine J (2008) Performance of bioaugmentation-assisted phytoextraction applied to metal contaminated soils: a review. Environ Pollut 153:497–522
Thomine S, Wang R, Ward JM, Crawford NM, Schroeder JI (2000) Cadmium and iron transport by members of a plant metal transporter family in Arabidopsis with homology to Nramp genes. Proc Natl Acad Sci 97:4991–4996
Treen-Sears ME, Martin SM, Volesky B (1998) Propagation of Rhiloprzs juvanicus biosorbent. Appld Environ Microbiol 448:137–141
Van Aken B (2008) Transgenic plants for phytoremediation: helping nature to clean up environmental pollution. Trends Biotechnol 26:225–227
Van Huysen T, Abdel-Ghany S, Hale KL, Le Duc D, Terry N, Pilon-Smits EAH (2003) Overexpression of cystathionine-y-synthase enhances selenium volatilization in Brassica juncea. Planta 218:71–78
Vatamaniuk OK, Mari S, Lu YP, Rea PA (1999) At PCS1, a phytochelatin synthase from Arabidopsis: isolation and in vitro reconstitution. Proc Natl Acad Sci 96(12):7110–7115
Vidali M (2001) Bioremediation – an overview. Pure Appl Chem 73(7):1163–1172
Wang C, Liu ZQ (2007) Foliar uptake of pesticides: present status and future challenge. Pest Biochem Physiol 87:1–8
Wang KS, Huang LC, Lee HS, Chen PY, Chang SH (2008) Phytoextraction of cadmium by Ipomoea aquatica (water spinach) in hydroponic solution: effects of cadmium speciation. Chemosphere 72:666–672
White C, Sharman AK, Gadd GM (1998) An integrated microbial process for the bioremediation of soil contaminated with toxic metals. Nat Biotechnol 16:572–575
Williams LE, Pittman JK, Hall JL (2000) Emerging mechanisms for heavy metal transport in plants. Biochim Biophys Acta 1465:104–126
Yang X, Jin XF, Feng Y, Islam E (2005a) Molecular mechanisms and genetic bases of heavy metal tolerance/hyperaccumulation in plants. J Integr Plant Biol 47:1025–1035
Yang X, Feng Y, He Z, Stoffella P (2005b) Molecular mechanisms of heavy metal hyperaccumulation and phytoremediation. J Trace Elem Med Biol 18:339–353
Zeyaullah M, Atif M, Badrul I, Abdelkafe AS, Sultan P, ElSaady MA, Ali A (2009) Bioremediation: a tool for environmental cleaning. Afr J Microbiol Res 3(6):310–314
Zhu Y, Pilon-Smits EAH, Jouanin L, Terry N (1999a) Overexpression of glutathione synthetase in Brassica juncea enhances cadmium tolerance and accumulation. Plant Physiol 119:73–79
Zhu Y, Pilon-Smits EAH, Tarun A (1999b) Cadmium tolerance and accumulation in Indian mustard is enhanced by overexpressing glutamylcysteine synthetase. Plant Physiol 121:1169–1177
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Dhir, B. (2017). Bioremediation Technologies for the Removal of Pollutants. In: Kumar, R., Sharma, A., Ahluwalia, S. (eds) Advances in Environmental Biotechnology. Springer, Singapore. https://doi.org/10.1007/978-981-10-4041-2_5
Download citation
DOI: https://doi.org/10.1007/978-981-10-4041-2_5
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-4040-5
Online ISBN: 978-981-10-4041-2
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)