Abstract
Molecular phylogeography has for decades been a frequently used approach to delineate novel evolutionarily significant units (ESUs) and to study the dynamics of invasive species. Next-generation sequencing technology (NGS) and the use of environmental DNA (eDNA) have the potential to revolutionize our way of understanding biodiversity and to establish rapid protocols for early-stage detection of invasive species. In seaweeds, however, several years of research on iconic invasive taxa of ambiguous taxonomic status (e.g. Caulerpa, Codium, Asparagopsis) have suggested that an integrative approach, namely the combination of multiple lines of evidence (e.g. phylogeographic, ecological, physiological and predictive modelling), is necessary to accurately resolve the taxonomy and their invasive potential. At present, integrative approaches in these fields are often weak because of incongruences among species delineation, newly discovered ESUs which remain undescribed taxonomically, and because databases containing vouchers of barcoded specimens are incomplete. As relocations of marine biota accelerate and climatic changes offer new potential niches for invasive seaweeds, new, transferable and internationally adopted protocols are necessary for exploring, monitoring and managing marine biodiversity. This is particularly urgent in areas of intense maritime traffic, such as the Mediterranean Sea and the Hawaiian archipelago, in order to achieve sustainable socio-economic development without compromising the local marine resources.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Humans have relied on macroalgae for food since the very early stages of civilization. Today, seaweeds also represent a promising resource for biocompounds, alternative energy, bioindicators for environmental health monitoring and bioextractors for recovering eutrophic areas (Gupta and Abu-Ghannam 2011). From an ecological perspective, seaweeds hold a fundamental role in regulating the nutrient composition of the water column, the hydrodynamic forces and sedimentation. Further, as ecosystem engineers , seaweeds provide shelter for the development and maintenance of benthic communities, occupy empty space and are placed at the base of trophic nets (Lüning et al. 1990). However, in the era of global climate change and accelerated trade, seaweeds are of major concern as invasive species (in the sense of Boudouresque and Verlaque 2010), posing severe social and economic threats in the coastal economy of many countries (Andreakis and Schaffelke 2012; Schaffelke and Hewitt 2007; Schaffelke et al. 2006). Estimates indicate that seaweeds represent up to 40 % of the non-native introduced species in the world’s oceans (Schaffelke et al. 2006). The outstanding success of some invasive aliens is mostly attributed to remarkable levels of propagule persistence during transport across the globe, together with a suite of demographic traits that support adaptation and elevated growth rates in the recipient environments (Anderson 2007; Engelen and Santos 2009; Flagella et al. 2006, 2007, 2010; Zanolla et al. 2015).
A combination of environmental variables such as temperature, light, salinity and nutrients affect seaweed survival and distributions locally, regionally and globally (Breeman et al. 1988, 2002; Eggert 2012). These variables are subject to change either slowly through geological time at large geographical scales, or rapidly within decades locally or globally, in response to climate change and ocean acidification (O’Hara et al. 2011; Harley et al. 2012). Historical climatic fluctuations and large scale geological events have often altered the ecophysiological optima for survival in many marine groups including seaweeds, causing extensive shifts in species distributions, and may explain some present-day biogeographic patterns (Maggs et al. 2008; Payo et al. 2013). Signatures of diversification and distributional shifts driven by the reduction of global sea levels by more than a hundred metre, during Pleistocene glacial maxima, can be detected in many marine communities as a periodic, slow, yet naturally occurring process (Ludt and Rocha 2015). The same, however, cannot be assumed for the present-day changes observed on the distribution patterns of many marine groups. In the last decades, we have witnessed unprecedented rates of species range shifts (Chen et al. 2011) and relocation among bioregions both intentionally or accidentally via human-mediated transport (Sorte et al. 2010). The rates and the distances within which species are moving across oceans cannot be compared to the macroecological changes observed in historical times and they have been interpreted as a consequence of climate change (Boudouresque and Verlaque 2002, 2010; Andreakis and Schaffelke 2012).
Phylogeography is concerned with identifying the processes responsible for the geographic distribution of genealogical lineages in space and time. A gene genealogy can be inferred from genetic data extracted at individual, population or species level to test how historical, geological, climatic or ecological events have influenced their distribution patterns (Avise 2000). Methodological approaches based mostly on phylogeographic inference and species spatial distribution modelling have recently become the main tools for identifying introduced species , deciphering sources of introduction and assessing the success and invasive potential of new colonists at multiple stages across the invasion process (Peterson 2003; Booth et al. 2007; Bolton et al. 2011). Given that human-mediated transport and global change facilitate diffusion of biota that would otherwise have limited dispersal potential, it becomes obvious that surveys aiming to identify endemisms and detect introduced species will have profound consequences in conservation biogeography and ecosystem management (Bickford et al. 2007; Andreakis and Schaffelke 2012).
In this chapter, we discuss the importance of properly delineating taxonomic units in phylogeographic research of invasive seaweeds . The case studies discussed below are not necessarily considered pests with demonstrated economic impact, since this has been established only in a few cases (Andreakis and Schaffelke 2012). We further argue that current knowledge and methodologies are insufficient for accurate predictions of whether introduced seaweeds will become invasive, pests or neither of the above. However, in cases of already-established invaders or species of recognized invasive potential, it is possible to accrue evidence for forecasting source populations, direction of range expansion and to predict hypothetical distribution ranges based on environmental suitability. In invasion biology, the adoption of integrative approaches based on species distribution modelling, phylogeographic inference and ecophysiological data is necessary for successful predictions and conservation planning (Fig. 7.1).
The integrative phylogeographic approach for studying biological invasions. Progression of the invasive process (1–4) and the disciplines necessary to study the main variables involved in each stage are reported in brackets. The representation is based on the introduction of multiple Asparagopsis cryptic lineages in Australia
2 The Advantages of Molecular Tools in Delineating Species
Accurate taxonomic delineation is essential for identifying the organisms being transported, understanding the dynamics of the invasion processes, and tracking the species’ historical root or cryptogenic status in situ from the analysis of historical collections (Sherwood 2008). Despite several species definitions proposed previously (Mayden 1997) and approaches for testing them (Leliaert et al. 2009), an unifying concept of species still represents a hot debate within the scientific and environmental community (Wattier and Maggs 2001; Carstens et al. 2013). Modern biology in the post-genomic era calls for a convincing and universal species definition to be used as the basic unit in biodiversity research. This is crucial for corroborating ecological, biological and evolutionary interpretations, executing practical applications such as estimating species richness and bio-invasion control programmes and comparing invasive processes of the same organism from distant areas (Pante et al. 2015). Morphology has traditionally been the dominant criterion to identify taxonomical units in seaweeds (Wattier and Maggs 2001). Molecular markers provide a solid alternative when morphological data are insufficient or inexistent. Indeed, a few cases of new species descriptions are today accepted without molecular corroboration. Genetic species delineation offers several undeniable advantages. First, neutrally evolving DNA regions do not suffer homoplasy compared to morphological and ecophysiological traits which are responsible for the species’ functional reaction to its surrounding environment (Saunders 2005). Second, modern molecular biology platforms allow for barcoding analysis of large numbers of samples for the validation of a specific ESUs or the analysis of eDNA (environmental DNA) for the identification of cryptic species. Third, since DNA composition at the sequence level remains the same across the ontogenetic development of any organism, molecular tools have the power to accurately identify heteromorphic life stages or microscopic forms from within the species life cycle or even fragments of propagules . Disadvantages of molecular taxonomy are mostly related to the adoption of inconsistent laboratory methodologies amongst laboratories and the production of heterogeneous data from multiple DNA regions instead of consistently targeting the same markers across the group in question (Verbruggen et al. 2010).
Molecular taxonomy involves (a) the collection of molecular data (e.g. DNA sequences from one or more genomic regions) from multiple specimens of the group under examination and (b) their phylogenetic analysis, in order to identify well-supported clusters of closely related specimens corresponding to ESUs (Moritz 1994). ESUs are here intended as taxa which are reciprocally monophyletic for mtDNA alleles and show significant divergence of allele frequencies at nuclear loci (Moritz 1994). Different molecular markers are characterized by different evolutionary rates. The choice is based on their ability to achieve unambiguous identifications given the taxonomical level questioned (Hebert et al. 2003). Examples of different markers developed for systematic and evolutionary studies in seaweeds are shown in Table 7.1. In phylogeography of invasive seaweeds, the choice of the marker is a fundamental decision for identifying introduced species . The selected molecular marker should have a suitable resolution for detecting the target taxa either in their vectors of transport or within their suspected introduced range (Geller et al. 2010).
Molecular taxonomy and phylogenetics have revolutionized our perception of seaweed biodiversity by revealing hitherto unknown levels of diversity, and have provided a statistically robust framework for testing evolutionary hypotheses (Bickford et al. 2007). Cryptic speciation is common in many marine organisms (Knowlton 1993). Leaving morphological delineation aside, a preliminary genetic screening is always necessary to draw the bottom line when it comes to investigations within widely distributed taxa (Geller et al. 2010). In light of combined molecular and morphological evidence, a choice must be made on which taxonomical level to focus on and bypass potential incongruences that may exist between molecular data, morphological descriptions and previous reports. A classic example of an extremely morphologically plastic group, in which genetic surveys have been essential tools for species delineation , is the green algal genus Caulerpa J.V. Lamouroux. Caulerpa represents an iconic case in which the phylogenetic approach has led to controversy because of differences in the chosen molecular markers (and their resolution) among studies. The original results of the first studies based mostly on morphology led to misidentifications and constant taxonomic restructuring. Previous morphological identifications of Caulerpa cylindracea Sonder led to erroneous records of the invasive strain (Verlaque et al. 2003; Yeh and Chen 2004; Nuber et al. 2007), later emended by molecular studies based on the ITS marker (Klein and Verlaque 2008). Further, C. cylindracea started as a “variety” (C. racemosa var. cylindracea (Sonder) Verlaque, Huisman & Boudouresque), went through a “form” (C. racemosa f. complanata (J. Agardh) Weber-van Bosse), and ended as a “species” into which a new cryptic variety has been included (Belton et al. 2014).
3 Multiple Cryptic Endemisms or Introduced Lineages Within Cosmopolitan Species?
Sorting out endemic lineages from cryptic introductions may be difficult without information gathered from historical collections, since the discovery of a new species may be the result of unrecognized parapatric or sympatric speciation (Voisin et al. 2005; Schaffelke et al. 2006; Andreakis and Schaffelke 2012). Unnoticed propagules of cryptic lineages can successfully establish founder populations following human-mediated long-range dispersal in areas of environmental suitability for that lineage (Breeman 1988). Megadiverse areas such as the coral triangle and the Great Barrier Reef or bioregions affected by intense maritime traffic such as the Mediterranean Sea and the Hawaiian archipelago are considered highly vulnerable to biological invasions . In these areas, the identification of cryptic endemisms , versus cryptic introductions, represents both a major endeavour for conservation biogeography and a knowledge gap capable of compromising future biodiversity management strategies (Bickford et al. 2007; Trontelj and Fišer 2009). Misidentifications of one or the other will result in a complete overlook of either the invasive organism, leading to a cryptic invasion, or unrecognized endemic lineage (McIvor et al. 2001; Geller et al. 2010). Given the importance of biodiversity fluctuations for ecological marine conservation, control plans and assessments of the long-term ecological and evolutionary consequences of cryptic species must be a priority for management agencies.
4 The Impact of Multiple Invasive Life Stages
Life-history strategies characterized by multiple phases of distinct morphology and ploidy level are common in seaweeds (Drew 1955). Each of the life stages can therefore contribute differently in the expansion potential of the species across the course of an invasion , assuring its success acting either as dispersal units (Hewitt et al. 2007; Zanolla et al. 2015) or seed banks (Hewitt et al. 2005). For instance, the red seaweed genus Asparagopsis has a triphasic diplohaplontic heteromorphic life cycle (Fig. 7.2), in which gametophytes, microscopic carposporophytes and filamentous tetrasporophytes (Falkenbergia) of unknown ploidy level alternate (Feldmann and Feldmann 1942; Rojas et al. 1982).
Depending on how pronounced the heteromorphy is, each of the life-history stages may eventually belong to a different functional group and is thus expected to present different thermal ranges of reproduction, growth and survival (Breeman 1988; Eggert 2012), as well as to be subject to distinct biotic and abiotic pressures (Littler and Littler 1980). Microscopic life stages are believed to be more resistant (Breeman 1988) and are thus considered good candidates for long distance dispersal. Macroscopic phases on the other hand would be responsible for increased biomass production and will guarantee population persistence and short distance dispersal . The contribution to the population persistence of each of the life-history stages, ploidy levels, clones or sexually reproduced individuals may vary. Their impact will depend on the functional group affected and may reflect shifts in the dominance of one versus the other according to their adaptation potentials (Van der Strate et al. 2002) and the environmental characteristics of the invaded location.
5 Vectors of Introduction Promote Relocations of Seaweeds: Range Shifts Versus Niche Shifts
A major cause of increased invasion rates is the reinforcement of multiple introduction vectors rather than global climate change itself (Boudouresque and Verlaque 2010). Aquaculture, maritime transport and the opening of interoceanic canals, to mention a few, have been widely accepted as major causes for seaweed introductions (Williams and Smith 2007). In Europe, globalization has promoted an increased number of maritime commercial shipping routes, considered responsible for more than half of the introduced species in the marine environment, followed by the opening of artificial water corridors between basins. Intentional or unintentional introductions of invasive species through aquaculture and aquarium trade are considered less relevant pathways for the spread of invasive species in Europe (<1.2 %) (Katsanevakis et al. 2013). Invasive species provide unique model systems for ecology and evolutionary biology since the species range shift following introduction can potentially be accompanied by niche shifts because of the species’ environmental tolerance in the introduced region (Rödder and Lötters 2009; Tingley et al. 2014). A ‘niche shift’ refers to “any change in the position of either the fundamental or realized (Hutchinsonian) niche of a species” (Pearman et al. 2008). As the exploitation of novel habitats and niches may lead to adaptive radiations, the study of such organisms can provide important insights in understanding species diversification . Further, ecological niche models for predicting potential distribution of invasive species may allow us to anticipate invasions (Guisan et al. 2014). Niche conservatism, i.e. the extent to which niches are conserved over space and time, is a useful approach for extrapolating invasive species’ distribution ranges and predicting their invasive risk. Niche shifts among seaweeds have only been rarely documented. For instance, in Halimeda J.V. Lamouroux (Verbruggen et al. 2009), despite the dispersal limitations and the conservatism of the genus to tropical habitats, successful colonisation of colder environments has occurred than once in the past. However, range shifts of tropical seaweeds are expected to occur more frequently as a result of ongoing global warming (Thuiller et al. 2005; Boudouresque and Verlaque 2010).
6 Multiple Introductions from a Single Source
Biological invasions may involve the introduction of multiple contingents of individuals from the same source or multiple introduced individuals from more than one source. Asparagopsis armata Harvey is common in temperate seas and is believed to be native in Southern Australia and New Zealand (Dixon 1964). Two cryptic lineages have been recently reported within this species (lineages 1 and 2) (Dijoux et al. 2014). Lineage 1 has been introduced to northern Mediterranean Sea, Western Europe, Japan and the US west coast. Analysis of nuclear, mitochondrial, and plastid molecular markers revealed a unique southern Australian origin of invasive Mediterranean populations of lineage 1 of A. armata (Andreakis et al. 2007b). This conclusion is supported by the lack of genetic structuring among invasive populations, the shared haplotypes between recipient and donor regions and the invasive history of this species (Feldmann and Feldmann 1942; Andreakis et al. 2007b).
Codium fragile subsp. tomentosoides (van Goor) P.C. Silva is a well-recognized invasive green seaweed. This taxon is characterized by low genetic variation both within its introduced (Mediterranean Sea, Northwest Atlantic, Northern European and South Pacific) and native (Japan) range although parthenogenesis, prevalent in this genus, may also contribute to this (Prince and Trowbridge 2004). Introduced Codium populations shared unique haplotypes with the subspecies donor region. Eight different haplotypes were recovered in Japan and only one of them could be found exclusively in the Mediterranean Sea. The latter was clearly different from the haplotypes present in other introduced locations (Provan et al. 2005, 2008).
The “aquarium-Mediterranean strain” of Caulerpa taxifolia (M. Vahl) C. Agardh has invaded the Mediterranean Sea through aquarium trade (Meusnier et al. 2002). Phylogenetic studies of the aforementioned strain which was released from aquarium facilities revealed the origin of these invasive populations to be the tropical and subtropical region of Australia (Meusnier et al. 2002). However, invasive Australian populations of this same taxon were suggested to be derived from different source regions (Schaffelke et al. 2002).
7 Introductions from Multiple Sources
In many cases, invasive populations are the result of introductions from multiple sources. This type of introduction is often overlooked when the so-called cosmopolitan species are believed to be native in many regions or when successfully established invasive populations act as propagule donors. Asparagopsis taxiformis (Delile) Trevisan de Saint León represents a clear example (Fig. 7.3). Multiple lineages are known from the Mediterranean Sea, the Atlantic and the Indo-Pacific Oceans. Interestingly, invasive behaviour has been reported for solely lineages 2 and 3 in the Mediterranean region (Andreakis et al. 2007b; Boudouresque and Verlaque 2010) and South Africa (Bolton et al. 2011), respectively. Further, lineage 2 recently expanded its distributional range in southern Portugal and the Mediterranean Sea, and it is considered an invasive taxon characterized by high genotypic diversity (Andreakis et al. 2009). Two more lineages have been recently described, both confined to South Pacific and Western Australia (Dijoux et al. 2014; Andreakis et al. 2016). To what extent these Asparagopsis lineages represent biologically different species, ESUs or groups of distinct genotypes, still requires further assessments (Dijoux et al. 2014; Zanolla et al. 2014, 2015). However, Mediterranean strains of lineage 2 might represent a distinct ecotype for that lineage found in Australia (Andreakis et al. 2007b; Dijoux et al. 2014) and the Hawaiian Islands (Sherwood 2008) and not a distinct genetic variant. This is confirmed by its distinct morphology, photosynthetic performance and the identical mitochondrial haplotypes shared among Mediterranean Australian, Hawaiian and African isolates (Zanolla et al. 2014, 2015).
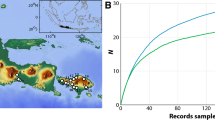
Updated distribution occurrence map in the Mediterranean Sea (a), the Alboran Sea (b) and the Hawaiian Islands (c) for each of the Asparagopsis lineages based on genetically delineated specimens (Andreakis et al. 2004, 2007b; Sherwood 2008; Dijoux et al. 2014). A. armata (filled triangle); A. taxiformis L1 (X); A. taxiformis L2 (filled circle); A. taxiformis L3 (filled square); A. taxiformis L4 (filled diamond), modified from Zanolla et al. (Submitted)
8 Differences Between Donor and Introduced Populations
Genetic variation of introduced seaweed populations may differ significantly from the donor populations because (a) individuals transported under suboptimal conditions are subjected to negative selection, (b) introduction events may be multiple (propagule pressure) and (c) introduced species undergo strong bottleneck events following introduction . Consequently, as a general trend, the genetic profile of a recently introduced population may appear less variable due to the perpetuation of few successful genotypes (Voisin et al. 2005). In addition, genetic variability among invasive populations may differ because of dissimilar population propagation mechanisms (i.e. sexual vs. vegetative propagation ) and/or introduction events from multiple sources (Andreakis et al. 2009).
Invasive species characterized by identical genetic profiles between their native and introduced strains may develop adaptive phenotypic plasticity and ecophysiological tolerance in response to the novel environmental conditions (Eggert 2012). Rapid adaptation will promote a superior fitness to the introduced individuals which will be characterized by distinct ecophysiological and/or morphological traits, compared to populations found in the species native range (Andreakis and Schaffelke 2012). Adaptive plasticity can therefore confer evolutionary advantages to invasive species by optimizing their acclimation mechanisms (Davidson et al. 2011).
9 Integrative Taxonomy and Phylogeography : Combining Multiple Lines of Evidence
In several cases, a molecular phylogeographic approach has been decisive for the identification of cryptic invaders, the detection of organisms imported via vectors of transport such as ballast tanks and to infer the colonization route and the donor population of introduced taxa (Deagle et al. 2003; Bolton et al. 2011). However, for robust species delineation and prediction making, the combination of data from multiple lines of evidence will give more realistic results (Figs. 7.1 and 7.4). Molecular, morphological, physiological and geographic distribution data tested against multiple species concepts (e.g. phenetic, biological and phylogenetic ) render species delineation unambiguous. A species integrative profile can thereafter be tested against observed phenetic, physiological or behavioural variants for that species and assess whether these changes are (a) part of the species’ adaptation potential within its plasticity range, (b) associated with genome level variations or specific gene expression profiles or (c) are the result of transgenerational adaptation mechanisms based on epigenetic modifications. The latter is often induced by diversifying selection among populations invading different habitats (Weinig 2000; Lee 2002).
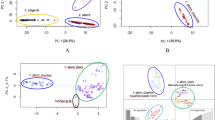
The invasive red seaweed genus Asparagopsis (Montagne) provides a good example of combining molecular, morphological and ecophysiological data in resolving taxonomic issues. Previous reports on the presence of Asparagopsis species worldwide were limited to reporting the species sensu lato rather than the corresponding cryptic lineage itself. This misidentification resulted in erroneous distribution maps and a general failure to identify species where molecular studies had not been employed. Immediate Asparagopsis lineage delineation is now possible, even without molecular screening, by means of a set of vegetative and reproductive diagnostic morphological characters. These characters have been identified from morphological studies performed on genetically delineated specimens of Asparagopsis lineages collected from sites where these lineages are considered either introduced or native (Zanolla et al. 2014).
10 The Utility of Combining Multiple Lines of Evidence in the Study of Invasive Seaweeds
Species distribution modelling (SDM) calculates the similarities of environmental affinities between locations where a species occurs and locations where the same species has never been reported. As a consequence, SDM can be used to forecast distribution ranges using environmental variables as predictors (Austin 2002). In invasion biology, SDMs are routinely used to identify potentially suitable settlement areas of invasive species. However, since SDMs require the compilation of georeferenced data (species presence or absence), the precise taxonomic identification and deep knowledge of the target species’ ecophysiological optima have become of paramount importance for the reliable performance of the models. Accurate information on the taxonomy, distribution and ecophysiological limits of species or lineages can be combined in SDM to provide crucial knowledge in biodiversity research and invasion biology. For instance, in the case of a cosmopolitan species, the occurrence of that species in locations not suggested by the models may indicate a novel cryptic ESU , which can represent either an endemism or a cryptic introduction that requires further attention for management and conservation.
A combination of field and laboratory work, aiming to identify the taxonomic position and understand the physiology, demography, phenology, population dynamics and impact on the local community of an invasive organism, is necessary to detect biological invasions but also designates them as pests (Meinesz 2007). The astonishing success of invasive species relies on their extraordinary adaptation potential. As a process, adaptation relies on the improvement of multiple functional traits in the life cycle of an organism capable of enhancing its fitness, survival and resilience against novel environmental conditions. Consequently, adaptation potential cannot be understood by means of one approach. The extreme adaptive responses in invasive seaweeds can be clearly visible (e.g. noticeable ecophysiological and morphological differences between native and introduced populations) or invisible. The latter category includes adaptive responses associated with positively selected genetic variants in response to novel conditions or responses associated with transgenerationally inherited epigenetic polymorphisms. The latter in their turn are induced by environmental cues, and might not be detectable by traditional sequence analysis approaches (i.e. no differences at the DNA sequence level between native and introduced populations). Thus, the invasive behaviour observed in an organism cannot be explained on the basis of one line of evidence. The importance of the integrative approach becomes further obvious when the communication of scientific knowledge goes through stakeholders responsible for the implementation of management plans and decision making.
An integrative approach has been implemented in the genus Asparagopsis , one of the few cases where haplotype analysis, historical demography and SDM were combined to assess lineage-specific invasive risk (Zanolla et al., submitted), lineage-specific photosynthetic plasticity in response to a range of temperatures (Zanolla et al. 2015) and morphological differentiation among cryptic lineages of genetically delineated tetrasporophytes and gametophytes locally and globally (Zanolla et al. 2014). Further, vegetative and reproductive traits were examined taking as a model an established population of this invasive taxon (Zanolla et al., submitted). This comprehensive multifaceted approach allowed characterization of the genetic composition, colonization strategy and lineage-specific potential for short and long distance dispersals as well as invasion risk.
11 Modern Technology and Metabarcoding in the Study of Invasive Seaweeds
Global climate change , human-mediated transport and extensive urban development of coastal zones are responsible for increasing rates of marine species introductions but also conspicuous behavioural and/or phenetic changes in many endemic, bloom-forming taxa. Because of eutrophication or accidental release, both native and introduced organisms can possibly and unexpectedly turn into pests. Therefore, the complete prevention of seaweeds invasions or native algal blooms appears nearly impossible in the long term. However, focusing on taxonomic groups associated with high rates of invasibility risk and the establishment of early warning protocols, especially in vulnerable areas, can help reduce the speed and ecological impact of invasions. Modern high throughput molecular approaches possess the resolution power and the affordability to unravel many aspects of the biology of invasive organisms at functional and molecular levels. Novel technology has the potential to address questions and examine behavioural changes associated with the metabolomic, proteomic, genomic and transcriptomic profiles of an invasive organism but also the influences of bacterial symbionts in the species response against environmental cues. A major outcome is expected in genomic and/or proteomic identification of biomarkers to be used for assessing invasive potential in high-risk groups. The same marker profiles can be used to monitor the possibilities of invasiveness in endemic taxa. In addition, genomic approaches are expected to resolve the influence of polyploidy in inducing invasiveness in seaweeds. Polyploidy has already been proven to be relevant in the invasiveness of higher plants (Pandit et al. 2011) and has been proposed as a mechanism capable of supporting invasiveness in green (Caulerpa) and red (Asparagopsis) algal genera (Andreakis et al. 2007a; Varela-Álvarez et al. 2012). The correlation between invasive potential and increased ploidy levels in these species is likely explained by increased levels of heterozygosity associated with polyploidy (Brochmann et al. 2004). Increased heterozygosity may support the ecological success of an introduced alien by balancing the loss of diversity due to population bottleneck and low sexual recombination that take place at early stages of introduction (reviewed in Varela-Alvarez et al. 2012).
Next-generation sequencing-based eDNA/RNA metabarcoding from environmental samples is expected to play a crucial role in the monitoring, detection and identification of introduced, transported or invasive species (Chown et al. 2015). Although not extensively applied in assessing biological invasions , eDNA/RNA metabarcoding has so far showed promising results (Armstrong and Ball 2005; Ficetola et al. 2008; Saunders 2009). The technology relies on the same general principle of the regular DNA barcoding for species identification. Short DNA sequences (barcodes ) originated upon a previously agreed, high-resolution DNA marker are compared to barcodes produced from well-identified voucher specimens deposited in reference databases. This approach can be designed to capture more than one species and can be applied in all steps of the invasion history (Fig. 7.1), or the species life stages (Fig. 7.2), to provide constant monitoring support and information for early detection and identification of dispersal vectors, therefore allowing estimates on the demography, population dynamics and dispersal of the invasive organism (Metzker 2010). Although initially the short DNA reads (ca. 100 bp) limited the use of NGS for DNA metabarcoding purposes, this downside has been now attenuated by the sequencing of longer reads produced by many platforms (e.g. >500 bp; Illumina, MiSeq, http://www.illumina.com/systems/miseq/performance_specifications.ilmn). At present, however, a real drawback of metabarcoding remains the current limited availability of reference databases (Cristescu 2014).
12 Conclusion
The combination of novel DNA-based analytical methodologies with traditional approaches has the potential to alleviate the methodological and conceptual critiques charged at invasion science (Richardson and Ricciardi 2013).
Several European countries have officially recognized the need for surveillance and monitoring of invasive species (No 1143/2014 of 22 October 2014; http://ec.europa.eu/environment/nature/invasivealien/index_en.htm). With the rapid identification and detection of non-native species being the top priority, DNA metabarcoding, in particular, represents an efficient and affordable method to identify transported species or detect the presence of invasive pests from minute quantities of DNA from environmental samples (Darling and Mahon 2011). To be successful, this approach requires the rapid description of the cryptic species and the parallel development of accurate reference databases, consisting of type voucher specimens associated with specific environmental profiles and lineage-specific barcodes (e.g. BoLD) (Ratnasingham and Hebert 2007).
An integrative approach to biological invasions can provide realistic solutions by documenting global patterns of the invasion process and by identifying areas into which direct management actions are immediately required (i.e. introduction vectors, marine protected zones, areas of intense maritime traffic ). As the vertiginous socio-economic development of many countries of the world struggles to stay ecologically sustainable, the consequences of biological invasions still remain global and irreversible. This interface requires the use of common language and sense from both scientists and politicians towards rapid and effective balancing of management actions at national and international levels.
References
Anderson LW. Control of invasive seaweeds. Bot Mar. 2007;50:418–37.
Andreakis N, Procaccini G, Kooistra WHCF. Asparagopsis taxiformis and Asparagopsis armata (Bonnemaisoniales, Rhodophyta): genetic and morphological identification of Mediterranean populations. Eur J Phycol. 2004;39:273–83.
Andreakis N, Koolstra WHCF, Procaccini G. Microsatellite markers in an invasive strain of Asparagopsis taxiformis (Bonnemaisoniales, Rhodophyta): insights in ploidy level and sexual reproduction. Gene. 2007a;406:144–51.
Andreakis N, Procaccini G, Maggs C, Kooistra WHCF. Phylogeography of the invasive seaweed Asparagopsis (Bonnemaisoniales, Rhodophyta) reveals cryptic diversity. Mol Ecol. 2007b;16:2285–99.
Andreakis N, Kooistra WHCF, Procaccini G. High genetic diversity and connectivity in the polyploid invasive seaweed Asparagopsis taxiformis (Bonnemaisoniales) in the Mediterranean, explored with microsatellite alleles and multilocus genotypes. Mol Ecol. 2009;18:212–26.
Andreakis N, Schaffelke B. Invasive marine seaweeds: pest or prize? In: Wiencke C, Bischof K, editors. Seaweed biology. Berlin: Springer; 2012. p. 235–62.
Andreakis N, Costello P, Zanolla M, Saunders GW, Mata L. Endemic or introduced? Phylogeography of Asparagopsis (Florideophyceae) in Australia reveals multiple introductions and a new mitochondrial lineage. J Phycol. 52:2016. doi:10.1111/jpy.12373-15-166.
Armstrong K, Ball S. DNA barcodes for biosecurity: invasive species identification. Phil Trans R Soc Lond B Biol Sci. 2005;360:1813–23.
Austin M. Spatial prediction of species distribution: an interface between ecological theory and statistical modelling. Ecol Model. 2002;157:101–18.
Avise JC. Phylogeography: the history and formation of species. Massachusetts: Harvard University Press; 2000.
Belton GS, Reine WF, Huisman JM, Draisma SG, Gurgel D, Frederico C. Resolving phenotypic plasticity and species designation in the morphologically challenging Caulerpa racemosa–peltata complex (Chlorophyta, Caulerpaceae). J Phycol. 2014;50:32–54.
Bickford D, Lohman DJ, Sodhi NS, Ng PKL, Meier R, Winker K, Ingram KK, Das I. Cryptic species as a window on diversity and conservation. Rend Ecol Evol. 2007;22:148–55.
Bolton JJ, Andreakis N, Anderson RJ. Molecular evidence for three separate cryptic introductions of the red seaweed Asparagopsis (Bonnemaisoniales, Rhodophyta) in South Africa. Afr J Mar Sci. 2011;33:263–71.
Booth D, Provan J, Maggs CA. Molecular approaches to the study of invasive seaweeds. Bot Mar. 2007;50:385–96.
Boudouresque C, Verlaque M. Assessing scale and impact of ship-transported alien macrophytes in the Mediterranean Sea CIESM Workshop Monographs. 2002;53–61.
Boudouresque CF, Verlaque M. Is global warming involved in the success of seaweed introductions in the Mediterranean Sea? In seaweeds and their role in globally changing environments. Berlin: Springer; 2010. p. 31–50.
Breeman AM. Relative importance of temperature and other factors in determining geographic boundaries of seaweeds—experimental and phenological evidence. Helgol Meeresunters. 1988;42:199–241.
Breeman A, Oh YS, Hwang MS, Van Den Hoek C. Evolution of temperature responses in the Cladophora vagabunda complex and the C. albida/sericea complex (Chlorophyta). Eur J Phycol. 2002;37:45–58.
Brochmann C, Brysting A, Alsos I, Borgen L, Grundt H, Scheen AC, Elven R. Polyploidy in arctic plants. Biol J Linnean Soc. 2004;82:521–36.
Carstens BC, Pelletier TA, Reid NM, Satler JD. How to fail at species delimitation. Mol Ecol. 2013;22:4369.
Cheang CC, Chu KH, Ang PO. Phylogeography of the marine macroalga Sargassum hemiphyllum (Phaeophyceae, Heterokontophyta) in northwestern Pacific. Mol Ecol. 2010;19(14):2933–48.
Chen I-C, Hill JK, Ohlemüller R, Roy DB, Thomas CD. Rapid range shifts of species associated with high levels of climate warming. Science. 2011;333(6045):1024–6.
Chown SL, Hodgins KA, Griffin PC, Oakeshott JG, Byrne M, Hoffmann AA. Biological invasions, climate change and genomics. Evol Appl. 2015;8:23–46.
Conklin KY, Kurihara A, Sherwood AR. A molecular method for identification of the morphologically plastic invasive algal genera Eucheuma and Kappaphycus (Rhodophyta, Gigartinales) in Hawaii. J Appl Phycol. 2009;21(6):691–9.
Coyer JA, Hoarau G, Stam WT, Olsen JL. Geographically specific heteroplasmy of mitochondrial DNA in the seaweed, Fucus serratus (Heterokontophyta: Phaeophyceae, Fucales). Mol Ecol. 2004;13(5):1323–6.
Cristescu ME. From barcoding single individuals to metabarcoding biological communities: towards an integrative approach to the study of global biodiversity. Trends Ecol Evol. 2014;29(10):566–71.
Daguin C, Voisin M, Engel C, Viard F. Microsatellites isolation and polymorphism in introduced populations of the cultivated seaweed Undaria pinnatifida (Phaeophyceae, Laminariales). Conserv Genet. 2005;6(4):647–50.
D’Archino R, Nelson WA, Zuccarello GC. Invasive marine red alga introduced to New Zealand waters: first record of Grateloupia turuturu (Halymeniaceae, Rhodophyta). New Zeal J Mar Fresh Res. 2007;41(1):35–42.
Darling JA, Mahon AR. From molecules to management: adopting DNA-based methods for monitoring biological invasions in aquatic environments. Env Res. 2011;111:978–88.
Davidson AM, Jennions M, Nicotra AB. Do invasive species show higher phenotypic plasticity than native species and if so, is it adaptive? A meta-analysis. Ecol Lett. 2011;14:419–31.
Deagle B, Bax N, Hewitt C, Patil J. Development and evaluation of a PCR-based test for detection of Asterias (Echinodermata: Asteroidea) larvae in Australian plankton samples from ballast water. Mar Fresh Res. 2003;54:709–19.
Dijoux L, Viard F, Payri C. The more we search, the more we find: discovery of a new lineage and a new species complex in the genus Asparagopsis. PLoS ONE. 2014;9:e103826.
Dixon PS. Asparagopsis in Europe. Nature. 1964;204:902.
Drew KM. Life histories in the algae with special reference to the Chlorophyta, Phaeophyta and Rhodophyta. Biol Rev. 1955;30:343–90.
Eggert A. Seaweed responses to temperature seaweed biology. Berlin: Springer; 2012. p. 47–66.
Engelen A, Santos R. Which demographic traits determine population growth in the invasive brown seaweed Sargassum muticum? J Ecol. 2009;97(4):675–84.
Feldmann J, Feldmann G. Recherches sur les Bonnemaisoniacées et leur alternance de générations Annales des Sciences Naturelles. Bot Sér. 1942;11:75–175.
Ficetola GF, Miaud C, Pompanon F, Taberlet P. Species detection using environmental DNA from water samples. Biol Lett. 2008;4:423–5.
Flagella MM, Soria A, Buia MC. Shipping traffic and introduction of non-indigenous organisms: study case in two Italian harbours. Ocean Coast Manag. 2006;49:947–60.
Flagella MM, Verlaque M, Soria A, Buia MC. Macroalgal survival in ballast water tanks. Mar Poll Bull. 2007;54(9):1395–401.
Flagella MM, Andreakis N, Hiraoka M, Verlaque M, Buia MC. Identification of cryptic Ulva species (Chlorophyta, Ulvales) transported by ballast water. J Biol Res. 2010;13:47–57.
Geller JB, Darling JA, Carlton JT. Genetic perspectives on marine biological invasions. Annu Rev Mar Sci. 2010;2:367–93.
Geoffroy A, Le Gall L, Destombe C. Cryptic introduction of the red alga Polysiphonia morrowii Harvey (Rhodomelaceae, Rhodophyta) in the North Atlantic Ocean highlighted by a DNA barcoding approach. Aq Bot. 2012;100:67–71.
Guisan A, Petitpierre B, Broennimann O, Daehler C, Kueffer C. Unifying niche shift studies: insights from biological invasions. Trends Ecol Evol. 2014;29(5):260–69.
Gupta S, Abu-Ghannam N. Recent developments in the application of seaweeds or seaweed extracts as a means for enhancing the safety and quality attributes of foods. Innov Food Sci Emerg. 2011;12:600–9.
Halling C, Wikström SA, Lilliesköld-Sjöö G, Mörk E, Lundsør E, Zuccarello GC. Introduction of Asian strains and low genetic variation in farmed seaweeds: indications for new management practices. J Appl Phycol. 2013;25(1):89–95.
Harley CD, Anderson KM, Demes KW, Jorve JP, Kordas RL, Coyle TA, Graham MH. Effects of climate change on global seaweed communities. J Phycol. 2012;48(5):1064–78.
Hebert PDN, Cywinska A, Ball SL, DeWaard JR. Biological identifications through DNA barcodes. P R Soc London B Bio. 2003;270:313–21.
Hewitt CL, Campbell ML, McEnnulty F, Moore KM, Murfet NB, Robertson B, Schaffelke B. Efficacy of physical removal of a marine pest: the introduced kelp Undaria pinnatifida in a Tasmanian Marine Reserve. Biol Inv. 2005;7:251–63.
Hewitt CL, Campbell ML, Schaffelke B. Introductions of seaweeds: accidental transfer pathways and mechanisms. Bot Mar. 2007;50:326–37.
Johnson MT, Carpenter EJ, Zhijian Tian, et al. Evaluating methods for isolating total RNA and predicting the success of sequencing phylogenetically diverse plant transcriptomes. PLoS ONE. 2012;7:e50226.
Jousson O, Pawlowski J, Zaninetti L, Meinesz A, Boudouresque CF. Molecular evidence for the aquarium origin of the green alga Caulerpa taxifolia introduced to the Mediterranean Sea. Mar Ecol Progr Ser. 1998;172:275–80.
Katsanevakis S, Zenetos A, Belchior C, Cardoso AC. Invading European seas: assessing pathways of introduction of marine aliens. Ocean Coast Manag. 2013;76:64–74.
Kim SY, Weinberger F, Boo SM. Genetic data hint at a common donor region for the invasive Atlantic and Pacific populations of Gracilaria vermiculophylla (Gracilariales, Rhodophyta). J Phycol. 2010;46(6):1346–9.
Klein J, Verlaque M. The Caulerpa racemosa invasion: a critical review. Mar Poll Bull. 2008;56:205–25.
Knowlton N. Sibling Species in the Sea. Ann Rev Ecol Systemat. 1993;24:189–216.
Lee CE. Evolutionary genetics of invasive species. Trends Ecol Evol. 2002;17:386–91.
Leliaert F, Verbruggen H, Wysor B, De Clerck O. DNA taxonomy in morphologically plastic taxa: algorithmic species delimitation in the Boodlea complex (Chlorophyta: Cladophorales). Mol Phylogenet Evol. 2009;53:122–33.
Littler MM, Littler DS. The evolution of thallus form and survival strategies in benthic marine macroalgae: field and laboratory tests of a functional form model. Am Nat. 1980;116:25–44.
Ludt WB, Rocha LA. Shifting seas: the impacts of Pleistocene sea-level fluctuations on the evolution of tropical marine taxa. J Biogeogr. 2015;42:25–38.
Lüning K, Yarish C, Kirkman H. Seaweeds: their environment, biogeography, and ecophysiology. New York: Wiley; 1990.
Maggs CA, Castilho R, Foltz D, Henzler C, Jolly MT, Kelly J, Olsen J, Perez KE, Stam W, Vainola R, Viard F, Wares J. Evaluating signatures of glacial refugia for North Atlantic Benthic Marine Taxa. Ecology. 2008;89:S108–22.
Marston M, Villalard-Bohnsack M. Genetic variability and potential sources of Grateloupia doryphora (Halymeniaceae, Rhodophyta), an invasive species in Rhode Island waters (USA). J Phycol. 2002;38(4):649–58.
Mayden RL. A hierarchy of species concepts: the denouement in the saga of the species problem. In: Claridge MF, Dawah HA, Wilson MR, editors. Species: the units of biodiversity. New York: Chapman & Hall; 1997. p. 381–424.
McDevit DC, Saunders GW. On the utility of DNA barcoding for species differentiation among brown macroalgae (Phaeophyceae) including a novel extraction protocol. Phycol Res. 2009;57(2):131–41.
McIvor L, Maggs CA, Provan J, Stanhope MJ. rbcL sequences reveal multiple cryptic introductions of the Japanese red alga Polysiphonia harveyi. Mol Ecol. 2001;10:911–9.
Meinesz A. Methods for identifying and tracking seaweed invasions. Bot Mar. 2007;50:373–84.
Metzker ML. Sequencing technologies—the next generation. Nat Rev Gen. 2010;11:31–46.
Meusnier I, Valero M, Destombe C, Gode C, Desmarais E, Bonhomme F, Stam W, Olsen J. Polymerase chain reaction–single strand conformation polymorphism analyses of nuclear and chloroplast DNA provide evidence for recombination, multiple introductions and nascent speciation in the Caulerpa taxifolia complex. Mol Ecol. 2002;11:2317–25.
Meusnier I, Valero M, Olsen JL, Stam WT. Analysis of rDNA ITS1 indels in Caulerpa taxifolia (Chlorophyta) supports a derived, incipient species status for the invasive strain. Eur J Phycol. 2004;39(1):83–92.
Moritz C. Defining ‘evolutionarily significant units’ for conservation. Trends Ecol Evol. 1994;9:373–5.
Nuber N, Gornik O, Lauc G, Bauer N, Zuljevic A, Papes D, Zoldos V. Genetic evidence for the identity of Caulerpa racemosa (Forsskal) J. Agardh (Caulerpales, Chlorophyta) in the Adriatic sea. Eur J Phycol. 2007;42:113–20.
O’Hara T, Gledhill D, Butler A, Bax N, Wilson R, Poore G, McCallum A, Last P, England P, Andreakis N. Report on application of evolutionary history to inform how species and/or communities might respond to changes in climate. CERF Marine Biodiversity Hub. 2011.
O’Doherty DC, Sherwood AR. Genetic population structure of the Hawaiian Alien invasive seaweed Acanthophora spicifera (Rhodophyta) as revealed by DNA sequencing and ISSR analyses. Pac Sci. 2007;61(2):223–33.
Olsen JL, Valero M, Meusnier I, Boele-Bos S, Stam WT. Mediterranean Caulerpa taxifolia and C. mexicana (Chlorophyta) are not conspecific. J Phycol. 1998;34(5):850–6.
Pandit MK, Pocock MJ, Kunin WE. Ploidy influences rarity and invasiveness in plants. J Ecol. 2011;99:1108–15.
Pante E, Puillandre N, Viricel A, Arnaud-Haond S, Aurelle D, Castelin M, Chenuil A, Destombe C, Forcioli D, Valero M. Species are hypotheses: avoid connectivity assessments based on pillars of sand. Mol Ecol. 2015;24:525–44.
Payo DA, Leliaert F, Verbruggen H, D’hondt S, Calumpong HP, De Clerck O. Extensive cryptic species diversity and fine-scale endemism in the marine red alga Portieria in the Philippines. Proc R Soc B. 2013;280:20122660.
Pearman PB, Guisan A, Broennimann O, Randin CF. Niche dynamics in space and time. Trends Ecol Evol. 2008;23(3):149–58.
Peterson AT. Predicting the geography of species’ invasions via ecological niche modeling. Quart Rev Biol. 2003;78:419–33.
Prince JS, Trowbridge CD. Reproduction in the green macroalga Codium (Chlorophyta): characterization of gametes. Bot Mar. 2004;47:461–70.
Provan J, Murphy S, Maggs CA. Tracking the invasive history of the green alga Codium fragile ssp tomentosoides. Mol Ecol. 2005;14:189–94.
Provan J, Booth D, Todd NP, Beatty GE, Maggs CA. Tracking biological invasions in space and time: elucidating the invasive history of the green alga Codium fragile using old DNA. Divers Distrib. 2008;14:343–54.
Ratnasingham S, Hebert PD. BOLD: the barcode of life data system. Mol Ecol Notes. 2007;7:355–64. http://www.barcodinglife.org.
Richardson DM, Ricciardi A. Misleading criticisms of invasion science: a field guide. Divers Distrib. 2013;19:1461–7.
Rödder D, Lötters S. Niche shift versus niche conservatism? Climatic characteristics of the native and invasive ranges of the Mediterranean house gecko (Hemidactylus turcicus). Glob Ecol Biogeogr. 2009;18:674–87.
Rojas J, Lemus A, Ganesan E. El ciclo vital “in vitro” del alga marina roja Asparagopsis taxiformis (Delile) Collins & Hervey (Bonnemaisoniales, Rhodophyta) del Mar Caribe. Bol Inst Oceanogr (Cumaná). 1982;21:101–12.
Saunders GW. Applying DNA barcoding to red macroalgae: a preliminary appraisal holds promise for future applications. Phil Tran R Soc B. 2005;360:1879–88.
Saunders GW. Routine DNA barcoding of Canadian Gracilariales (Rhodophyta) reveals the invasive species Gracilaria vermiculophylla in British Columbia. Mol Ecol Res. 2009;9:140–50.
Schaffelke B, Murphy N, Uthicke S. Using genetic techniques to investigate the sources of the invasive alga Caulerpa taxifolia in three new locations in Australia. Mar Poll Bull. 2002;44:204–10.
Schaffelke B, Smith JE, Hewitt CL. Introduced macroalgae—a growing concern. J Appl Phycol. 2006;18:529–41.
Schaffelke B, Hewitt CL. Impacts of introduced seaweeds. Bot Mar. 2007;50:397–17.
Sherwood AR. Phylogeography of Asparagopsis taxiformis (Bonnemaisoniales, Rhodophyta) in the Hawaiian Islands: two mtDNA markers support three separate introductions. Phycologia. 2008;47:79–88.
Sorte CJ, Williams SL, Carlton JT. Marine range shifts and species introductions: comparative spread rates and community impacts. Global Ecol Biogeogr. 2010;19:303–16.
Thomsen MS, Gurgel CFD, Fredericq S, McGlathery KJ. Gracilaria vermiculophylla (Rhodophyta, Gracilariales) in Hog Island Bay, Virginia: a cryptic alien and invasive macroalga and taxonomic correction. J Phycol. 2005;42:139–41.
Thuiller W, Richardson DM, Pysek P, Midgley GF, Hughes GO, Rouget M. Niche-based modelling as a tool for predicting the risk of alien plant invasions at a global scale. Glob Change Biol. 2005;11:2234–50.
Tingley R, Vallinoto M, Sequeira F, Kearney MR. Realized niche shift during a global biological invasion. Proc Natl Acad Sci USA. 2014;111:10233–8.
Trontelj P, Fišer C. Perspectives: cryptic species diversity should not be trivialised. Syst Biodivers. 2009;7:1–3.
Uwai S, Nelson W, Neill K, Wang WD, Aguilar-Rosas LE, Boo SM, Kawai H. Genetic diversity in Undaria pinnatifida (Laminariales, Phaeophyceae) deduced from mitochondria genes-origins and succession of introduced populations. Phycologia. 2006;45(6):687–95.
Van der Strate HJ, Van de Zande L, Stam WT, Olsen JL. The contribution of haploids, diploids and clones to fine-scale population structure in the seaweed Cladophoropsis membranacea (Chlorophyta). Mol Ecol. 2002;11:329–45.
Varela-Álvarez E, Gómez Garreta A, Rull Lluch J, Salvador Soler N, Serrao EA, Siguán MAR. Mediterranean species of Caulerpa are polyploid with smaller genomes in the invasive ones. PLoS ONE. 2012;7:e47728.
Verbruggen H, Leliaert F, Maggs CA, Shimada S, Schils T, Provan J, Coppejans E. Species boundaries and phylogenetic relationships within the green algal genus Codium (Bryopsidales) based on plastid DNA sequences. Mol Phyl Evol. 2007;44(1):240–54.
Verbruggen H, Tyberghein L, Pauly K, Vlaeminck C, Nieuwenhuyze KV, Kooistra WH, Leliaert F, Clerck OD. Macroecology meets macroevolution: evolutionary niche dynamics in the seaweed Halimeda. Glob Ecol Biogeogr. 2009;18:393–405.
Verbruggen H, Maggs CA, Saunders GW, Le Gall L, Yoon HS, De Clerck O. Data mining approach identifies research priorities and data requirements for resolving the red algal tree of life. BMC Evol Biol. 2010;10(1):16.
Verlaque M, Durand C, Huisman JM, Boudouresque CF, Le Parco Y. On the identity and origin of the Mediterranean invasive Caulerpla racemosa (Caulerpales, Chlorophyta). Eur J Phycol. 2003;38:325–39.
Voisin M, Engel CR, Viard F. Differential shuffling of native genetic diversity across introduced regions in a brown alga: aquaculture vs. maritime traffic effects. Proc Natl Acad Sci USA. 2005;102:5432–7.
Wattier R, Maggs CA. Intraspecific variation in seaweeds: the application of new tools and approaches. Adv Bot Res. 2001;35:171–212.
Weinig C. Plasticity versus canalization: population differences in the timing of shade-avoidance responses. Evolution. 2000;54(2):441–51.
Williams SL, Smith JE. A global review of the distribution, taxonomy, and impacts of introduced seaweeds. Ann Rev Ecol Evol Syst. 2007;38:327–59.
Yeh WJ, Chen GY. Nuclear rDNA and internal transcribed spacer sequences clarify Caulerpa racemosa vars. from other Caulerpa species. Aq Bot. 2004;80:193–207.
Zanolla M, Carmona R, De La Rosa J, Salvador N, Sherwood AR, Andreakis N, Altamirano M. Morphological differentiation of cryptic lineages within the invasive genus Asparagopsis (Bonnemaisoniales, Rhodophyta). Phycologia. 2014;53(3):233.
Zanolla M, Altamirano Jeschke M, Carmona R, de la Rosa Álamos JC, Sherwood AR, Andreakis N. Photosynthetic plasticity of the genus Asparagopsis (Bonnemaisoniales, Rhodophyta) in response to temperature: implications for invasiveness. Biol Inv. 2015;17(5):1–13.
Acknowledgments
Marianela Zanolla is supported by the project P09-RNM-5187 from the Consejería de Innovación, Ciencia y Empresa, Junta de Andalucía, Spain; NA by the Commonwealth Research and Environment Facilities (CERF) Marine Biodiversity Hub. The CERF programme is an Australian Government initiative supporting world class, public good research and is a collaborative partnership between the University of Tasmania, CSIRO Wealth from Oceans Flagship, Geoscience Australia, Australian Institute of Marine Science and Museum Victoria.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Zanolla, M., Andreakis, N. (2016). Towards an Integrative Phylogeography of Invasive Marine Seaweeds, Based on Multiple Lines of Evidence. In: Hu, ZM., Fraser, C. (eds) Seaweed Phylogeography. Springer, Dordrecht. https://doi.org/10.1007/978-94-017-7534-2_7
Download citation
DOI: https://doi.org/10.1007/978-94-017-7534-2_7
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-017-7532-8
Online ISBN: 978-94-017-7534-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)