Abstract
Haloalkaliphiles differ from natronophiles by their requirement for chloride ions in addition to high alkalinity. Natronophilic bacteria grow optimally in soda medium buffered at alkaline pH by a combination of NaHCO3 and Na2CO3. The majority of known haloalkaliphilic and natronophilic prokaryotes are isolated from saline–alkaline ecosystems such as soda lakes and saline–alkaline soils. A great taxonomic and metabolic biodiversity is found in soda systems, enabling the functioning of all the cycles of the essential elements. In spite of the increasing number of haloalkaliphilic and natronophilic isolates, scarce biochemical and functional information on simultaneous adaptation at high salinity and alkalinity is reported. Most of the available data on haloalkaline adaptation can be inferred from the functional characterization of alkaliphilic and halophilic bacterial models as well as from a few haloalkaliphilic and natronophilic genome sequences deposited in databases. At the level of cell envelopes (cell wall and cytoplasmic membrane), the salt and alkaline adaptation strategies are different and relatively conserved between Gram-positive and Gram-negative bacteria. The cell wall of the former group is characterized by the excessive presence of acidic polymers, while cell membranes abound in phospholipids with branched fatty acids. Cell membranes of salt- and alkaline-adapted Gram-negatives contain a large variety of fatty acids as well as significant amounts of nonpolar lipids. Osmotic adaptation mostly depends on the accumulation of organic compatible solutes either by active solute uptake or by combined strategies of importing osmolytes or osmolyte precursors and de novo synthesis of organic compatible solutes. Aerobic and anaerobic haloalkaliphiles are distinguished from each other by very different bioenergetics. Energy conservation in aerobic alkaliphiles and haloalkaliphiles is mainly based on functioning of H+-driven F-type ATP synthase. In spite of the low transmembrane electrochemical proton gradient (equivalent to proton-motive force, pmf) encountered in the alkali-exposed membrane, the energy metabolism remains highly efficient, supporting high growth rate and yield in many aerobic alkaliphiles and haloalkaliphiles. The energetics of haloalkaliphilic anaerobes is less understood, but it seems to involve a greater deal of Na dependency than in their aerobic counterpart. Na+-dependent ATPase activity is reported in a few anaerobic haloalkaliphiles and its role probably deals with active Na+ ejection from the cytoplasm. In haloalkaliphiles and natronophiles, the sodium-motive force (smf) is mainly driving the flagellar movement and sodium/solute symport. Cytoplasmic pH and ion homeostasis in haloalkaliphiles and natronophiles are most probably achieved by a concerted activity of a constellation of alkaline-activated ion transporters, among which Na+/H+ and Mrp-like antiporters have a major contribution.
The online version of the erratum chapter can be found at http://dx.doi.org/10.1007/978-94-007-6488-0_30
An erratum to this chapter can be found at http://dx.doi.org/10.1007/978-94-007-6488-0_30
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Studies of the past decades revealed an impressive diversity of organisms that inhabit highly saline and alkaline environments (Duckworth et al., 1996; Oren, 2002, 2012; Grant and Sorokin, 2011). Most of the prokaryotes and even a few eukaryotic species inhabiting saline–alkaline systems are well adapted to high salinity/alkalinity and high pH conditions. In general, obligate or true halophiles are defined as organisms that require at least 0.2 M or 15 g/l of sodium salts (specifically NaCl) in their milieu, with optimal growth between 0.2 and 5.2 M of total Na+ (Kushner and Kamekura, 1988; Ventosa et al., 1998). Obligate (true) alkaliphilic organisms sensu lato are those living optimally at pH values of at least 9, mostly growing best between pH 9.5 and 10.5 (Krulwich, 2006). Based on their minimal salt or pH requirements, both categories of extremophiles could be further divided into slight, moderate, and extreme halophiles or alkaliphiles, respectively (Table 1) (Grant et al., 1998; Hamamoto and Horikoshi, 1992; Ventosa et al., 1998; Mesbah and Wiegel, 2008). Interestingly, while most of true halophiles are flourishing at neutral or slightly acidic pH (6–7), many true alkaliphiles are also halophiles in their requirements, being termed haloalkaliphiles and qualifying as polyextremophiles. On the opposite, non-halophilic alkaliphiles require high pH and low salt in their medium (Hamamoto and Horikoshi, 1992; Horikoshi, 1999). The term “natronophile” (“natron-loving” organisms or “chloride-independent sodaphile”) has been proposed for the alkaliphilic organisms that have an absolute requirement for Na2CO3/NaHCO3 (Banciu, 2004; Sorokin et al., 2011a; Muyzer et al., 2011a). However, some natronophiles still require low concentrations of chloride ions for the optimal growth in soda brines. Life at haloalkaline conditions is energetically costly, and therefore, extreme haloalkaliphiles could hardly exhibit other extremophilic features (Oren, 2011). Moderate halophilic and thermoalkaliphilic anaerobic bacteria with validly published names belong to the orders Halanaerobiales (e.g., Halonatronum saccharophilum) and Natranaerobiales (e.g., Natranaerobius spp. and Natronovirga spp.). Triple extremophiles are apparently exceptions to the above-mentioned statement, but they are not extreme in any of their optimal growth requirements (Zhilina et al., 2001; Mesbah et al., 2007; Mesbah and Wiegel, 2009). A more frequent situation is that of moderate haloalkaliphiles that are either thermotolerant (e.g., strains of the sulfur-oxidizing bacterium Thioalkalivibrio versutus, some actinomycetes isolated from saline soils) (Sorokin and Kuenen, 2005; Ningthoujam et al., 2009) or psychrotolerant (e.g., Planococcus spp. strain ZD22 isolated from a saline–alkaline soil in the vicinity of an oil field) (Li et al., 2006).
Besides the main metabolic types listed in Table 1, there are different degrees of mixed halophily and alkaliphily observed in various microbes. For example, strains of Bacillus halodurans are facultative alkaliphiles and moderately halotolerants, with optimal growth at pH 9–10 and a salt tolerance ranging from 0 up to 12 % NaCl (Nielsen et al., 1995). Oceanobacillus iheyensis HTE831 is facultatively alkaliphilic and extremely halotolerant and is closely related to Bacillus spp. It grows optimally at pH between 7.5 and 9.5, tolerating up to 21 % NaCl at pH 7.5. Isolated from deep-sea sediment, O. iheyensis can withstand pressures up to 30 MPa (Lu et al., 2001).
Often, saline–alkaline environments hosting haloalkaliphilic prokaryotes are exposed not only to relatively elevated temperatures but also to intense sunlight irradiation and, consequently, to a high rate of water evaporation during dry seasons (Grant and Sorokin, 2011). In these harsh conditions, survival of salt- and alkaline-loving microorganisms depends on a palette of adaptive mechanisms that aim to remain alive during prolonged periods of low water activity. Acute and long-term adaptation as a response to changing salinity and/or alkalinity include modulation of membrane fluidity, acquisition of osmoprotectants, adjustments of energy metabolism concomitant with fine-tuning of stress-sensing, chemotaxis, and nutrients transport at the membrane level. Model halophilic and alkaliphilic organisms have for a long time been used to test for phenotypic and genotypic response to variation of external salinity and pH, respectively. A few examples of detailed physiological investigations concerning the double-stress adaptation at high salinity and alkalinity are known, and they are mostly focused on natronophilic lithoautotrophic bacteria and haloalkaliphilic archaea. In this chapter, we endeavor to discuss up-to-date relevant findings in the haloalkaliphilic adaptative strategies as they were directly observed or as extrapolation of well-known alkaliphilic and halophilic settings. Accumulating information on genome sequences aids to reveal even more insights in the mosaic of various adaptive strategies to the haloalkaline environment.
2 Habitats and Diversity of Haloalkaliphilic and Natronophilic Bacteria
The majority of known obligate haloalkaliphilic and natronophilic bacteria and archaea are isolated from soda lakes and saline–alkaline soils. In the past, detailed investigations on the biota living in soda systems have been carried out mainly in soda lakes from the East African Rift Valley and western USA. Currently, Central Asian (Chinese, Mongolian, and Siberian) soda lakes are under intense screening for microbial diversity.
2.1 Saline–Alkaline Habitats
Saline–alkaline environments are characterized by increased salinity (>60 g/l total salinity) and high actual (soluble) alkalinity (up to molar concentrations) and pH (pH >9.5–11). Because in such environments salinity and alkalinity are simultaneously provided by a high concentration of sodium carbonate and sodium bicarbonate, they are also known as soda systems. Saline–alkaline (soda) lakes or soils are located mostly in arid and semiarid regions where particular climate, geochemical, hydrological, and topological factors interact. Differing from neutral saline lakes (pH 6–8, with low buffering capacity and high NaCl, Na2SO4, K+, and Mg2+ concentration), soda lakes are strong natural buffering systems at pH >10, containing high concentrations of Na2CO3, NaHCO3, and, in some cases, a significant concentration of NaCl. One of the major chemical characteristics of soda lakes is the lack of solubilized divalent cations (Mg2+, Ca2+) that precipitate as carbonates under alkaline conditions. The early removal of divalent cation carbonates results in the accumulation of sodium or potassium carbonates, thus increasing the concentration of monovalent cations. By repeated cycles of concentration and evaporation, salts crystallize and brines are formed. In some parts of the world, the shallow lakes may end as a layer of solid rock termed trona (crystalline sodium sesquihydrate, Na2CO3 . NaHCO3 . 2H2O) by evaporation during dry seasons (Grant and Sorokin, 2011). A broad range of intermediate saline and/or alkaline lakes occurs by the mixing of the minerals in various ratios. Other important consequences of the soda brine chemistry, including the presence of high levels of soluble inorganic phosphorus, low toxicity of sulfide and nitrite, and high toxicity of ammonia, have important implications on the microbial system.
Salt lakes originating from sea are called thalassohaline, while those with land origin are termed athalassohaline (Oren, 2002). Most of neutral salt lakes and salterns (pH 7–8) are thalassohaline, while soda lakes are obviously of continental (athalassic) origin. Bodies of water with a significantly higher salinity (>80 g/l, w/v) than that of the seawater (35 g/l, w/v) are categorized as highly saline or hypersaline lakes (Poehls and Smith, 2009). Many soda lakes are also hypersaline, reaching a salinity of over 300 g/l. Well-known saline–alkaline lakes are located in eastern Africa (along the East African Rift Valley), the western and northern USA, the Middle East (Turkey, Armenia), and Central Asia (southern Siberia, northeastern and Inner Mongolia, China). Remote soda lakes are also located in other parts of the world, such as Australia, Central and South America (Chile, Venezuela), and southern Asia (India). Low-salt alkaline ponds and soils are described in Europe (Austria, Hungary, Serbia) (Tindall, 1988).
Saline–alkaline soils are either natural or man-made, formed upon intensive irrigation/evaporation cycles. Natural sodic (soda or soda solonchak) soils and deserts are formed in dry areas located in Europe (e.g., the Pannonian Basin), Asia (SW Siberia, NE Mongolia, Central China, India), Africa (Egypt), and North America. In comparison with soda lakes, the composition and diversity of microbial community living in soda soils is scarcely characterized (Lysak et al., 1994; Zenova et al., 2005; Sorokin et al., 2008e).
Saline–alkaline ecosystems have attracted the attention of investigators due to their unique chemical and biological features. Soda lakes are considered as models of ancient Martian or Archean terrestrial aquatic ecosystems (Kempe and Degens, 1985; Kempe and Kazmierczak, 1997; Zavarzin, 1993).
In spite of the apparently unfavorable physical and chemical conditions (high pH, high salinity, low water activity, paucity of divalent cations, high irradiation, elevated temperature, etc.), saline–alkaline environments display a full and balanced recycling of chemical elements (C, O, H, N, S, P, metals, etc.) and often a spectacularly high primary biomass production. In this category of extreme habitats, an almost complete trophic web assures the biological utilization and transfer of elements. Prokaryotes, represented by archaea and bacteria, as well as microscopic and macroscopic eukaryotes (protozoa, algae, brine shrimps, insect larvae) thrive in saline–alkaline lakes. Prokaryotes are the most diverse and numerous group of soda lake inhabitants. Recent advances in molecular biology have the potential to reveal complex metacommunities that play key roles in the ecological status of such habitats (Riesenfeld et al., 2004; Grant and Heaphy, 2010).
2.2 Biodiversity of Soda Environments
As a general observation, in saline–alkaline environments, species diversity increases as the salinity and/or alkalinity decreases (Grant and Tindall, 1986). In diluted saline–alkaline lakes of the East African Rift, representatives of macrofauna, such as fishes (Tilapia grahami), are found. Brine shrimp (Artemia sp.), commonly found throughout the world’s saline lakes, is documented in saline–alkaline Mono Lake (USA) (Dana and Lenz, 1986) as well as in hypersaline soda lakes in the Kulunda Steppe (Altai, Russia) (Sorokin et al., 2012d). In soda lakes with moderate salinity (<90 g/l) and pH <10, from East Africa and the western USA, zooplankton species including rotifers (e.g., Brachionus plicatilis), cladocerans (Moina hutchinsoni), and copepods (Diaptomus sicilis) were recorded (Grant and Tindall, 1986; Bozek, 1989). These microscopic invertebrates feed on phytoplankton such as microscopic algae and cyanobacteria. Haloalkaliphilic photoautotrophic cyanobacteria from the genera Arthrospira spp., Cyanospira spp., Synechococcus spp., and Synechocystis spp. consistently contribute to the high primary productivity in hyposaline and moderate soda lakes (Grant et al., 1990). Cyanobacteria are essential both for inorganic C fixation and O2 production in the ecosystem of saline–alkaline lakes. In addition to cyanobacterial populations, hypersaline soda lakes such as Lake Magadi in Kenya and Wadi Natrun haloalkaline lakes in Egypt, host significant communities of anoxygenic phototrophic purple bacteria belonging to Ectothiorhodospira and Halorhodospira, which also contribute to the primary biomass production (Imhoff et al., 1979; Grant and Tindall, 1986). Photolithoautotrophic haloalkaliphilic bacteria such as Thiorhodospira sibirica and Thioalkalicoccus limnaeus are the sole species of their genera that have been isolated from the low saline soda lakes from the Siberian steppe (Bryantseva et al., 1999a, 2000). Anoxygenic, photoheterotrophic heliobacteria such as alkaliphilic, low-salt-tolerant members of Heliorestis spp. were identified in low saline soda lakes from Siberia and Wadi Natrun (Egypt) (Bryantseva et al., 1999b; Asao et al., 2006). A relatively large nutritional and ecological spectrum of aerobic and anaerobic prokaryotes, bacteria and archaea, assures the natural cycling of the elements in the soda environments. Basically, all major metabolic groups are represented among the soda lakes inhabitants (Jones et al., 1998; Zavarzin et al., 1999; Grant and Sorokin, 2011).
Chemoheterotophic aerobes thriving in soda lakes are well represented by archaea, as well as Gram-negative and Gram-positive bacteria. Haloalkaliphilic members of the family Halomonadaceae (e.g., Halomonas magadiensis, H. kenyensis, H. mongoliensis) have been isolated from soda lakes around the world, being an important part of the easily culturable aerobic chemoheterotrophic communities thriving in these ecosystems (Duckworth et al., 1996, 2000; Boltyanskaya et al., 2007; Shapovalova et al., 2008). Other haloalkalitolerant and haloalkaliphilic Halomonas spp. have been discovered in neutral saline lakes (H. alkaliantarctica), salt pools (H. campaniensis, H. alkaliphila), and saline soils (H. campisalis, H. boliviensis) (Poli et al., 2007; Romano et al., 2005a, 2006a; Mormile et al., 1999; Quillaguamán et al., 2004). Soda systems are the sources for many aerobic Gram-positive heterotrophs belonging to the phyla Firmicutes and Actinobacteria. Most Bacillus isolates from soda lakes or soils are alkaliphilic and moderately halophilic or halotolerant (Bacillus alkalisediminis, B. aurantiacus, B. bogoriensis, B. daliensis), and only a few of them have been categorized as true, although moderately, haloalkaliphiles, such as B. locisalis (Borsodi et al., 2008, 2011; Vargas et al., 2005a; Zhai et al., 2012; Márquez et al., 2011). A wealth of other Bacillus-related species with various degrees of mixed halo- and alkaliphily have been recently and continuously isolated from soda lakes: the halophilic and alkalitolerant Salsuginibacillus kocurii and S. halophilus (Carrasco et al., 2007; Cao et al., 2010), the moderately halophilic and alkalitolerant Halolactibacillus alkaliphilus (Cao et al., 2008), and the haloalkalitolerant Amphibacillus jilinensis (Wu et al., 2009).
The important question of polymer degradation at soda lake conditions has barely been touched. So far a possibility of anaerobic utilization of cellulose and pectin as growth substrates was demonstrated in pure cultures of fermentative haloalkaliphilic members of the Clostridia and the Bacteroidetes (Zhilina et al., 2005; Sorokin et al., 2011d, 2012c). Many fermentative and anaerobic respirers from soda lakes belong to the orders Clostridiales, Halanaerobiales, and Natranaerobiales. From the Halanaerobiales, two natronophilic species of Natroniella have been described: the chemoorganotrophic homoacetogenic Natroniella acetigena (Zhilina et al., 1996) and the chemolithoautotrophic sulfidogenic N. sulfidigena (Sorokin et al., 2011b). Both have been isolated from anoxygenic sediments of soda lakes in Kenya and SW Siberia, respectively. Natroniella sulfidigena is capable of acetate-dependent sulfur respiration, being responsible of hydrogen sulfide formation at extremely haloalkaline conditions (pH >10; salinity >2 M Na+). Sulfate-reducing bacteria (SRB) thriving in the anoxic sediments of various soda lakes are represented by extremely or moderately natronophilic lithotrophs belonging to the order Desulfovibrionales (genera Desulfonatronovibrio, Desulfonatronum, and Desulfonatronospira) (Zhilina et al., 1997; Sorokin et al., 2008b, 2011c, 2012b; for a review, see Sorokin et al., 2011a). Moderately natronophilic heterotrophic SRB capable of utilizing propionate or volatile fatty acids as carbon and energy source have been recently assigned to the order Desulfobacterales and described as Desulfonatronobacter acidivorans and Desulfobulbus alkaliphilus (Sorokin et al., 2012a). The most recent example of an extraordinary sulfate reducer described as Desulfohalophilus alkaliarsenatis (Switzer Blum et al., 2012) was isolated from hypersaline–alkaline Searles Lake in California. This new member of the family Desulfohalobiaceae can grow by either sulfate or arsenate respiration at salt-saturating condition and high pH. The oxidative part of sulfur cycle in soda lakes is performed by natronophilic chemolithoautotrophic sulfur-oxidizing bacteria (SOB) belonging to four recognized genera: Thioalkalimicrobium, Thioalkalivibrio, Thioalkalispira, and Thioalkalibacter. Representatives of the former two genera are widely distributed in various soda lakes all over the world, while the latter two were only found occasionally (Sorokin and Kuenen, 2005; Banciu et al., 2008; Sorokin et al., 2011a). Haloalkaliphilic members of versatile fermentative spirochaetes (Spirochaeta alkalica, S. africana, S. asiatica, S. americana, and S. dissipatitropha) were isolated from brine sediments of Lake Magadi in Kenya, from sulfide-saturated mud of Lake Khatyn (Tuva, Siberia), and from sediments of the saline–alkaline Mono and Owens Lakes (California) (Zhilina et al., 1996; Hoover et al., 2003; Pikuta et al., 2009).
Examples of haloalkaliphilic and natronophilic Gram-positive anaerobes isolated from soda environments are the acetogenic Natronoincola histidinovorans (Zhilina et al., 1998) and members of the genus Tindallia (Pikuta et al., 2003; Alazard et al., 2007), halophilic and alkalithermophilic Natranaerobius thermophilus, N. trueperi, and Natronovirga wadinatrunensis (Mesbah et al., 2007, Mesbah and Wiegel, 2009). Haloalkalitolerant and haloalkaliphilic methanotrophs such as Methylomicrobium buryatense (Kaluzhnaya et al., 2001), M. alcaliphilum, and M. kenyense (Kaluzhnaya et al., 2008; Sorokin et al., 2000) identified in the sediments of moderate or highly saline soda lakes are Gram-negative, aerobic bacteria responsible of methane oxidation and formation of CO2 and various organic C1 compounds. Methanol, formaldehyde, and formate formed as products of partial methane oxidation by aerobic methanotrophs, as well as methylamines produced by the breakdown of organic osmolytes, are used as carbon source by moderately haloalkaliphilic aerobic methylobacteria such as Methylophaga alcalica, M. natronica, M. lonarensis, and Methylonatronum kenyense (Doronina et al., 2003a, b; Antony et al., 2012; Sorokin et al., 2007; for review, see Trotsenko et al., 2007).
In addition to bacterial populations, the prokaryotic communities of soda lakes comprise a variety of haloalkaliphilic archaea, many of them belonging to the Halobacteriaceae family. Haloalkaliphilic archaea isolated from soda lakes in East Africa, Central Asia, and the USA are found within the genera Natrialba, Natronomonas, Natronococcus, Natronobacterium, and Natronorubrum (Grant and Sorokin, 2011).
Other haloalkaliphilic bacteria possessing interesting metabolic characteristics or promising biotechnological applications have been isolated from saline–alkaline soils (e.g., the N2-fixing Gram-positive anaerobic Natronobacillus azotifigens) (Sorokin et al., 2008f), soda lakes sediments (Nitriliruptor alcaliphilus capable of growing on nitriles) (Sorokin et al., 2009), soda lake water (arsenite-oxidizing Alkalilimnicola ehrlichii) (Hoeft et al., 2007), coastal lagoon mud (Gram-positive anaerobe fermentative bacterium Halonatronum saccharophilum) (Zhilina et al., 2001), etc.
3 Mechanisms of High-Salt and Alkaline Adaptation in Bacteria
3.1 Adaptation of the Bacterial Cell Envelope to Haloalkaline Conditions
Cell envelopes (cell wall and cytoplasmic membrane) are the first line of defense and, implicitly, of adaptation to changing of environmental factors. The main constituents of cell wall are polysaccharides and proteins, while membranes consist of mainly lipids and proteins. In bacteria, membrane lipids are typically represented by polar phospholipids such as phosphatidylglycerol (PG), phosphatidylethanolamine (PE), and diphosphatidylglycerol or cardiolipin (CL) and nonpolar lipids such as isoprenoids (e.g., quinones, squalene, pigments). Ester phospholipids are constituted from a glycerophosphate backbone and two fatty acids esterified at the C2 and C3 positions. Therefore, the bulk of cell-extracted fatty acids originates from the plasma membrane. In this regard, it is not surprising that a significant part of the halophilic and alkaliphilic adaptation kit in bacteria might include more or less subtle changes of the membrane fatty acid structure and composition. As can be seen from the following evidences, besides particular ways of adapting the lipid composition, there are some general rules that apply in most high-salt and high-pH adaptation strategies.
3.1.1 Phospholipid and Fatty Acid Composition of Halophilic, Alkaliphilic, and Haloalkaliphilic Membranes
Based on their hydrocarbon chain structure, fatty acids can be divided into two main classes: straight-chain and branched-chain fatty acids (BFAs). Alicyclic (cyclopropane) fatty acids (CFAs) contain a cyclic structure within their molecule and are characteristic of Gram-negative bacteria (e.g., C19:0 cyclo ω7c). In general, many Gram-positive bacteria have significant amounts of BFAs (Kaneda, 1977). The most common straight-chained fatty acids found in bacteria are the saturated C14:0 (myristic acid), C16:0 (palmitic acid), C18:0 (stearic acid), C16:1 (palmitoleic acid), and the monounsaturated C18:1 (oleic acid). Branching of fatty acids generally occurs as iso and anteiso positioning of methyl residue at the second or at the third to the last carbon in the chain. Examples of iso-branched and anteiso-branched fatty acids that are widespread in Gram-positives are iso-C17:0 (15-methylpalmitic acid) and anteiso-C17:0 (14-methylpalmitic acid). Fatty acids are presently used as important chemotaxonomic markers in differentiation of bacterial species (Oren, 2012).
The major functional difference between the straight and branched chains, as well as between saturated and unsaturated fatty acids, resides in their contribution to the membrane fluidity (Kaneda, 1991). Variations of saturation, length, and branching in fatty acids are, therefore, one of the key factors changing the membrane fluidity as a short- and long-term response to environmental stress. Nevertheless, adjustments of the fatty acid composition are adding value to other adaptive changes in membrane lipids, such as modification of the zwitterionic to anionic lipid ratios (i.e., PE to PG and/or CL) or varying the production of isoprenoid derivatives.
The effects of varying salt concentration on membrane lipid composition are relatively well documented in halophilic bacteria with a broad salt tolerance such as Vibrio spp., Halomonas spp., and Chromohalobacter spp. (Adams et al., 1990; Adams and Russell, 1992; Russell, 1993; Vreeland et al., 1984; Arahal et al., 2001; Vargas et al., 2005b). As general conclusions drawn from experiments of salt stress effect on lipid composition in bacteria, one can affirm that the membrane fluidity is finely modulated by a complex interplay between environmental factors (temperature and salinity) on one side and cell response in terms of different phospholipids and fatty acids production on the other side. In Gram-negative halophilic bacteria (e.g., Halomonas spp.), growth at elevated salt concentration is correlated with high contents of straight chain, more unsaturated and lesser branched-chain fatty acids in the membranes (Monteoliva-Sanchez et al., 1988; Adams et al., 1990). At increasing salinity or at close to optimal growth salinity, the relative amount of long-chain acids (with C16 < C18) increases, whereas that of short-chain acids (C14 and/or C15) decreases (Imhoff and Thiemann, 1991; Valderrama et al., 1998). At high salt (as NaCl) concentration, the anionic phospholipids (PG) predominate over zwitterionic ones (PE). Additionally, CL and CFA increase at high salt concentration as observed in various moderately halophilic bacteria (for a review, see Ventosa et al., 1998). In the extremely halophilic bacterium Salinibacter ruber, CL is making up to 20 % of total polar lipids, a significantly higher proportion than that found in other halophilic bacteria (Lattanzio et al., 2009). The role of CL in the energetic membranes is well known. Cardiolipin is a major membrane lipid that binds c-type cytochromes and cytochrome c oxidase (Choi and Swanson, 1995; Robinson, 1993; Schägger, 2002) maintaining their optimal activity. In archaeal membranes CL derivatives are associated with terminal oxidases and are considered essential for haloarchaea adaptation at osmotic stress (Corcelli, 2009). Significant amounts of CL are correlated with high cytochrome c contents and cytochrome c oxidase activity in the membranes of alkaliphilic Bacillus strains (Clejan et al., 1986; Hicks and Krulwich, 1995) as well as in the natronophilic SOB Thioalkalivibrio spp. (Banciu et al., 2005).
Adaptation to high pH is similarly reflected in the membrane composition. In the alkalitolerant Listeria monocytogenes, the cells develop a higher proportion of BFAs of the anteiso form at pH 9. It has been suggested that the balance between anteiso- and iso-fatty acids may be more important than changes in the amounts and/or degrees of saturation of fatty acids in pH adaptation (Giotis et al., 2007). The facultatively alkaliphilic Bacillus pseudofirmus OF4 proved an excellent and intensively studied model organism to follow the effects of varying the external pH at the phenotype and genotype level. Like many Bacillus species, B. pseudofirmus OF4 possesses branched-chain fatty acids as major acids (90 % of total fatty acids). The obligate alkaliphilic Bacillus strains contain unusually high proportions of unsaturated fatty acids (20 % of the total fatty acids and most common being C16:1). In addition, obligate alkaliphilic strains of Bacillus have appreciable amounts of PG, CL, long-chain fatty acids, squalene, and C40 isoprenoids as compared to facultative strains (Clejan et al., 1986). Interestingly, the mixture of a high concentration of BFAs and a substantial fraction of unsaturated fatty acids (UFAs) may make the membrane leaky at a suboptimal (neutral) pH in the obligate alkaliphiles, as implied from the experiments performed by Dunkley et al. (1991).
In many Gram-positive moderately halophilic, alkaliphilic, or alkalitolerant species related to genus Bacillus (e.g., Alkalibacillus spp., Halalkalibacillus spp., Oceanobacillus spp.), the prevailing cell fatty acids are the BFAs (see also Table 2). However, dominance of BFA in cell membrane seems to be a general feature of Gram-positives (Kaneda, 1991) suggesting that the distinctive haloalkaline adaptations might be accommodated within the cell wall structure. It is considered that the branched anteiso-C15 fatty acid has a comparable function to that of UFAs in bacteria with a straight-chain lipid type, mainly because it has the lowest phase transition temperature (Tm = −16.5 °C) of all BFAs (Kaneda, 1991). The biosynthesis of UFAs occurs either aerobically or anaerobically (Aguilar and de Mendoza, 2006). UFAs, mostly as C16:1ω7 and C18:1ω7, are relatively common among anaerobes such as members of the family Clostridiaceae (Zhu et al., 2009). Representatives of haloalkaliphilic anaerobic bacteria (e.g., Tindallia spp., Natroniella spp., Natronincola spp.) contain significant amounts of UFAs. UFAs have a lower transition phase temperatures than their saturated counterparts, and consequently, they may profoundly influence the membrane fluidity (Silvius, 1982; Aguilar and de Mendoza, 2006).
Cyclopropane fatty acids (CFAs) are synthetized by the transfer of a methylene group from S-adenosyl-l-methionine to a double bond of unsaturated fatty acid chains of membrane phospholipids. This conversion is catalyzed by cyclopropane fatty acid (CFA) synthase. CFA synthase is present in many bacteria and it is recognized to regulate the membrane lipid composition and fluidity, thus playing an essential part in the adaptation of bacteria in response to a drastic perturbation of the environment (To et al., 2011). The extremely salt-tolerant natronophilic Thioalkalivibrio strain ALJ15 has a very similar fatty acids composition to the closely related Ectothiorhodospira spp. and Halorhodospira spp. (Imhoff and Thiemann, 1991) and Halomonas spp. (Valderrama et al., 1998; Romano et al., 2001), in which C16:0, C18:1, and cyc-C19 are the dominant fatty acids. The concentration of monounsaturated C18:1 is, however, two times higher in haloalkaliphilic SOB than in the halophilic Halomonas salina. Moreover, the concentration of cyc-C19 is significantly (5 times) higher in the natronophilic Thioalkalivibrio spp. than that measured in Ectothiorhodospira and Halomonas salina. Increasing salt concentration induced the decrease of monounsaturated C18:1 and concomitant increase of cyc-C19 in Thioalkalivibrio spp. (Banciu et al., 2005), Halomonas spp. (H. salina and H. halophila) (Valderrama et al., 1998; Monteoliva-Sanchez et al., 1988), and Lactobacillus strains (Gilarova et al., 1994). This is consistent with the knowledge that cis-monounsaturated fatty acids like C18:1 can be converted into cyc-C19 and vice versa. This may suggest a salt-dependent activation of CFA synthase. The raised CFA content at the expense of unsaturated fatty acids would contribute to the increasing of membrane rigidity (Russell, 1993). A stimulation of CFA synthase activity by addition of organic compatible solutes (e.g., glycine betaine) was observed in the moderately halophilic bacterium Pseudomonas halosaccarolytica (Monteoliva-Sanchez et al., 1993). CFAs such as cyc-C19 are considered as biomarkers for Gram-negative bacteria in a similar way as the BFAs for the Gram-positives. They are common in the membrane of anaerobic SRB and have been associated with the low redox potential in soils (Dowling et al., 1986; Zelles, 1999).
The less common aldehydes and dimethylacetal (DMA) fatty acids are specifically obtained during extraction of cellular fatty acids from anaerobes, and they are often used to differentiate between species. Aldehydes and 1,1-dimethylacetals result from plasmalogens by chemical lysis during lipid analysis procedures (Mayberry and Lane, 1993). Bacterial plasmalogens are proven to have a substantial effect on membrane fluidity. In the plasmalogen-deficient mutants of Megasphaera elsdenii, Kaufman et al. (1990) have observed the prevalence of a more stable lamellar phase. Studies on the psychrophilic actinobacteria Subtercola boreus and Subtercola frigoramans, with high contents of BFAs and DMA, have revealed that lowering of the growth temperature favored the production of DMAs with shorter carbon chain lengths and the increase of anteiso-branched DMA at the expense of iso-branched congeners (Månnistö et al., 2000). Generally it is believed that the presence of plasmalogens and their glycerol acetals could be responsible for substantial plasticity of bacterial membranes, especially for cells growing at a wide range of temperatures, salinity, and the presence of solutes such as hydrocarbons and solvents that have the potential of perturbing the bilayer arrangement of the cell membrane. Another possible role of plasmalogens in anaerobic bacteria is to protect the cell against oxidative stress (Goldfine, 2010).
Polyunsaturated fatty acids (PUFAs) are typical for cyanobacterial membranes where they are believed to play important roles in growth, respiration, and photosynthesis. The γ-linolenic acid (GLA, γC18:3) has an extremely low phase transition temperature (−60 °C) and abounds in membranes of halotoalkalitolerant Arthrospira (Spirulina) spp. Its occurrence in Arthrospira may be important in protecting the photosynthetic machinery from photoinhibition at low temperatures in a similar fashion as shown in Synechocystis strains (Mühling et al., 2005; Tasaka et al., 1996).
Squalene (C30 isoprenoid) is a nonpolar lipid, likely to be located between the lipid monolayers, in the hydrophobic core of the membranes (Hauss et al., 2002). Squalene can act as a barrier decreasing the membrane permeability for ions (Clejan and Krulwich, 1988). In the eukaryotic membranes, squalenes are precursors of cholesterol synthesis, a lipid that increases the membrane rigidity.
The alkaliphilic bacteria were shown to contain high concentrations of squalene and anionic phospholipids, especially cardiolipin (Clejan et al., 1986). Squalene and its synthesizing enzyme have been detected in natronophilic SOB Thioalkalivibrio spp., suggesting a reinforced structure of cell membrane (Banciu et al., 2005; Muyzer et al., 2011a).
Recently, a surprisingly high amount of the triterpene compound lanosterol has been detected in Thioalkalivibrio paradoxus, comprising 50 % from the total membrane lipids (D. Sorokin, unpublished data). This finding indicates a possible important structural function of sterols in the alkaliphily and warrants the necessity of closer attention. One of the possibilities is that these compounds may help the cell membrane to lower the membrane permeability to ions (i.e., protons, sodium ions) and also to cope with water availability in the surroundings. In this regard, the change of cell surface composition should also involve a change in hydrophilicity (and hydrophobicity) in order to control the water flux across cell envelopes. High salt is usually associated with low water activity and danger of desiccation, while lower salt than the optimal concentration correlates with higher water activity and excessive water inflow that may result in cell lysis. In this aspect it has been demonstrated that the cell surface of Gram-negative halophilic Halomonas elongata increased its hydrophilic nature with the elevation of external NaCl concentration (Hart and Vreeland, 1988).
3.1.2 The Acidic Cell Wall of Halo- and Alkaliphiles
In Gram-positive alkaliphilic and halophilic bacteria, the cell wall is clearly playing an important role in protecting the cell against the stress caused by high alkalinity and salinity.
The cell wall of facultatively alkaliphilic Bacillus halodurans C-125 is built from peptidoglycan and two acidic polymers: teichuronic acid (TUA) and teichuronopeptide (TUP). TUA consists of galacturonic acid, glucuronic acid, and N-acetylglucosamine, while TUP is a complex of polyglucuronic acid and poly-γ-l-glutamic acid (Aono, 1990). Cell wall-defective mutants of B. halodurans C-125 lost their ability to grow at alkaline pH, indicating the essential role of the cell wall in survival at high pH (Aono and Ohtani, 1990). Although not obligatory, the presence of negatively charged polymers in the structure of the alkaliphilic cell wall may favor the attachment of cations (such as sodium and hydronium ions) and repel hydroxide ions (Aono, 1990; Horikoshi, 1999). The presence of TUP seems restricted to cell wall of a few alkaliphilic Bacillus species. Only recently, the comparative genome analysis of Oceanobacillus iheyensis has revealed the presence of a putative protein showing significant similarity to the tupA gene product involved in TUP biosynthesis in B. halodurans (Takami et al., 2002). Proteomic and genomic analyses in the extreme facultative alkaliphile Bacillus pseudofirmus OF4 have shown that TUP is lacking and is substituted by an S-layer of protein nature (e.g., SlpA) (Gilmour et al., 2000; Janto et al., 2011). Additionally, a capsule made of negatively charged poly-γ-d-glutamate is anchored to the outer surface of the S-layer. Poly-γ-d-glutamate (PGA) occurs in the cell envelope of several non-extremophilic (e.g., Bacillus amyloliquefaciens) and facultative halophilic bacteria such as Planococcus halophilus and the moderately halophilic Halobacillus halophilus (Kandler et al., 1983; Geng et al., 2011). An α-linked l-glutamate polypeptide is present as complex exopolymers in some representatives of the archaea, including the haloalkaliphilic archaeon Natronococcus occultus and the extreme halophilic Natrialba aegyptiaca (Niemetz et al., 1997; Hezayen et al., 2001).
3.2 Osmoadaptation in Haloalkaliphilic and Natronophilic Bacteria
Presence of high sodium chloride and/or sodium (bi)carbonate concentration in the external milieu of living cells results in a high osmotic pressure and low water activity. This situation, often called osmotic stress, is also analogous to a desiccation stress. Cells that are not capable of response to such unusual physical and chemical environmental parameters will lose cytoplasmic water which tends to diffuse outward the cell. Non-halophilic organisms shrink and perish in highly saline conditions. On the other hand, halotolerants and halophiles developed several strategies to counterbalance the high external osmotic pressure. Extreme halophiles are narrowly specialized to live exclusively at high osmolarity and they are harmed and lysed in hypotonic conditions, the presence of too much water becoming a “water stress.” Besides the mechanical support of the cell wall (in Gram-positive bacteria) or a strengthened cell membrane (in archaea and Gram-negative bacteria), adaptation to high salinity/high osmotic pressure (or to variations of such parameters) is aiming to an isosmotic cytoplasm. Osmoadaptation of all living cells has same principle: synthesis and/or accumulation of osmotic compatible solutes (osmoprotectants or osmolytes). Based on their chemical nature, two main categories of osmolytes are met in nature: inorganic and organic osmolytes. Inorganic osmolytes (i.e., KCl) are imported and accumulated in molar concentrations mainly by neutrophilic halophilic and natronophilic archaea as the so-called “salt-in” strategy of osmoadaptation. Some bacteria within the orders Natranaerobiales and Halanaerobiales and the renowned example of halophilic bacterium Salinibacter ruber, a member of the Bacteroidetes, also use KCl as prevailing osmolyte (Oren et al., 2002). Interestingly, short-term osmoadaptation in Escherichia coli and Salmonella typhimurium involves an initial quick import of potassium ions followed by synthesis of glutamate as counterion. This response is enough only to assure the survival of the bacterial cell. The long-term osmoadaptation in such halotolerants, however, is based on the accumulation and/or synthesis of organic compatible solutes (e.g., trehalose, proline, or glycine betaine) which are less harmful to the cytoplasmic constituents (Oren, 1998).
Almost all known haloalkaliphilic or natronophilic bacteria have adapted to withstand a high osmotic pressure either by taking up or by de novo synthesis of organic compatible solutes (“salt-out” strategy). Unlike inorganic osmolytes, the presence of organic compatible solutes in the cytoplasm is more “friendly” to cell components, especially lipids and proteins. The most preferred organic osmolytes of bacteria are ectoine (1,4,5,6-tetrahydro-2-methyl-4-pyrimidinecarboxylic acid) and glycine betaine (N,N,N-trimethylglycine), followed by certain amino acids (glutamate, proline) and disaccharides (sucrose, trehalose). Accumulation of osmolytes is part of salt stress adaptation in many known halotolerant nonpathogenic and pathogenic bacteria (e.g., strains of Escherichia coli, Bacillus spp., Corynebacterium glutamicum, Listeria monocytogenes). They also may act as thermoprotectants in thermophilic prokaryotes. Organic osmolytes have a wide occurrence in algae (e.g., glycerol in Dunaliella spp.) and invertebrates (e.g., trehalose in the brine shrimp Artemia salina) living in saline ecosystems.
3.2.1 The Universal Compatible Solute, Glycine Betaine
Glycine betaine, or simply betaine, is a trimethylated derivative of glycine: a highly polar, low molecular weight, and chemically inert molecule. Betaine is broadly used as osmoprotectant in all three domains of life: bacteria, archaea, and eukaryotes (Oren, 2008; Tuteja, 2007; Lim et al., 2007). In bacteria, glycine betaine is the main osmoticum of high-salt-tolerant cyanobacteria, of halophilic anoxygenic phototrophic bacteria, and of salt-tolerant non-halophilic and halophilic heterotrophic bacteria (Imhoff and Rodriguez-Valera, 1984; Oren, 2002).
In halophilic archaea, de novo biosynthesis of glycine betaine is a rarity; it was assigned to one representative of halophilic methanogens, Methanohalophilus portucalensis both by physiological and genetic evidences (Robertson et al., 1990; Lai and Lai, 2011). Additionally, osmoadaptation in M. portucalensis involves betaine import as well as accumulation of potassium ions up to 1.1 M concentration (Lai et al., 1991). The analysis of the haloarchaeal genomes known by 2011, however, has revealed that 9 out of 10 genomes encode members of the betaine–choline–carnitine (BCC) transporter family (Anderson et al., 2011). Glycine betaine accumulation is a more preferred strategy in many halophilic bacteria and in some halophilic archaea (e.g., Methanosarcina mazei, Halococcus hamelinensis) (Mackay et al., 1984; Spanheimer and Müller, 2008; Burns et al., 2012). Several moderately halophilic/halotolerant and alkaliphilic/alkalitolerant Gram-positive bacteria such as Alkalibacillus filiformis and Oceanobacillus oncorhynchi accumulate glycine betaine and glutamate (Romano et al., 2005b, 2006b). In the halotolerant alkaliphilic O. iheyensis, a large number of solute transport proteins have been inferred from genome analysis. Several genes encode OpuD-like ABC transporters that facilitate betaine and choline uptake, as well as putative BCC transporters and Na+/proline or pantothenate symporters (OpuE transporters). It seems that the organism, which can grow between 0 and 18 % (3.6 M) NaCl, is employing an unusual high number of osmoprotectant transporters, especially when it must cope with a salinity of at least 3 M NaCl (Takami et al., 2002). Similar genes, although in a lesser number, are also present in genomes of the facultatively alkaliphilic and slightly halotolerant Bacillus halodurans C-125 and in the facultatively alkaliphilic B. pseudofirmus OF4 (Takami et al., 2000; Janto et al., 2011). Overall it seems that Gram-positive bacteria respond to osmotic stress by activating and using an important number of ABC and BCC transporters and proteins from the sodium/solute symporter (SSS) family. They possess the ability to import betaine when available in the growth medium or to import betaine precursors (i.e., choline) for cytoplasmic synthesis of this compatible solute.
De novo synthesis of betaine in halophiles is restricted to several autotrophic members of the cyanobacteria and the Ectothiorhodospiraceae (Oren, 2008). Halorhodospira halochloris, Ectothiorhodospira marismortui, and Ect. haloalkaliphila are well-known examples of glycine betaine accumulation in anoxygenic halophilic and haloalkaliphilic phototrophs (Galinski and Trüper, 1982; Imhoff, 1993). Unusually, in the haloalkalitolerant cyanobacterium Aphanothece halophytica, Laloknam et al. (2006) have detected a Na+/betaine transporter, BetT A. halophytica . This transporter is a member of BCC transporter family and is related to OpuD in B. subtilis and ProU in E. coli. It has a high affinity for betaine and differs from Gram-positive OpuD by its low isoelectric point (4.58, in comparison with the basic value pI 9.54 of OpuD from B. subtilis). This is the first and only description of a betaine transport system in cyanobacteria to date. Computational analysis of the Halorhodospira halophila (strain DSM 244/SL1) genome has revealed the existence of putative BCC transporters (Hhal_2364, Hhal_1851, Hhal_1384) as well as of a glycine betaine/l-proline ABC transporter (Hhal_0233).
Obligately chemolithoautotrophic sulfur-oxidizing bacteria (SOB) of the genus Thioalkalivibrio are closely related to Ectothiorhodospira spp. and Halorhodospira spp., being the dominant sulfur-oxidizing bacteria in soda lakes. Most of the Thioalkalivibrio species are true natronophiles, with optimal growth at pH 10 and at 2 M of total Na+. Experimental evidence has shown that in Thioalkalivibrio versutus strain ALJ15 grown in substrate-limited continuous culture at optimal salinity and pH, glycine betaine was the main organic compatible solute. When grown at different salt concentrations, strain ALJ15 accumulated betaine in a salt-dependent fashion: 1.5, 7.5, and 9 % of total dry weight at 0.6, 2, and 4 M Na+, respectively. Sucrose was produced as a minor secondary organic compatible solute (0.3–2.5 % of the total dry weight) in this organism, and its concentration was highest in cells grown at 2 M Na+. It was clear that Thioalkalivibrio spp. grown exclusively on organic-free medium is capable of de novo synthesis of glycine betaine as osmoadaptation strategy (Banciu et al., 2004a, b, 2005). Genome analysis in the Thioalkalivibrio spp. K90mix (phenotypically similar to strain ALJ15) and Thioalkalivibrio sulfidophilus HL-EbGr7 (phenotypically different from strain ALJ15) has indicated the presence of the genes for glycine sarcosine N-methyltransferase and sarcosine dimethylglycine methyltransferase. In addition, Thioalkalivibrio spp. genomes contain the gene for sucrose phosphate synthase needed for the production of sucrose as a minor compatible solute (Muyzer et al., 2011a, b). Biosynthesis of glycine betaine starting from glycine is a three-step methylation process: glycine → sarcosine → dimethylglycine → betaine, and is an energetically expensive pathway impairing growth at extreme salinity conditions (Nyyssölä et al., 2001; Oren, 1999). Other chemolithoautotrophic SOB that produce glycine betaine as main compatible solute include the extremely halophilic Thiohalorhabdus denitrificans and Thiohalospira halophila and the moderately halophilic and facultatively alkaliphilic Thiohalospira alkaliphila, all gammaproteobacteria. The first two species are among the most extreme halophilic lithoautotrophs known, with optimal salinity at 18 % w/v NaCl. They were isolated from surface sediments of inland hypersaline lakes in Russia, Crimea, and Central Asia and from a sea saltern at the shore of the Adriatic Sea. Thiohalospira alkaliphila was isolated from a hypersaline–alkaline lake in the Wadi Natrun and grows up to pH 10 and within a range of 0.5–4 M NaCl (optimum at 2 M NaCl). All these obligately chemolithoautotrophic SOB were grown on mineral media at 4 M and accumulated betaine up to one third of the total cell mass which was clearly much higher than in their natronophilic counterpart Thioalkalivibrio ALJ15 (Sorokin et al., 2008a, c). The reason for this is explained below.
Chemolithoautotrophic SOB such as Thioalkalivibrio halophilus HL17T and Thioalkalibacter halophilus ALCO1T grow well both at high NaCl and Na2CO3/NaHCO3 concentration (up to 4 M of total Na+), equally at pH 7.5 and 10. Such versatile, facultatively alkaliphilic high-salt-tolerant halophilic and facultatively natronophilic strains are ideal candidates for a direct study of the physiological effects of chloride versus carbonate/bicarbonate anions at same (high) sodium concentration, as well as at of variable pH. The NaCl-grown biomass of Tv. halophilus contained 19.8 % (w/w) glycine betaine, while the soda-grown biomass contained 12.4 % glycine betaine at the same concentration (4 M) of total Na+ (Banciu et al., 2004b). A comparable trend has been observed in haloalkaliphilic Thioalkalibacter halophilus. Unlike Tv. halophilus, strain ALCO1 is producing ectoine as the main compatible solute and betaine as a minor one. The specific concentration of osmolytes (ectoine and glycine betaine) measured in strain ALCO1 cells grown at 3 M NaCl (pH 7.5) was approximately two times higher than in cells grown at 3 M Na-soda, pH 10 (Banciu et al., 2008). Dissimilarities in the production of compatible solutes in NaCl- and soda-grown cells could be at least partially explained by physicochemical differences between NaCl and Na2CO3/NaHCO3 solutions. A decisive parameter that may influence the accumulation of osmolytes in halophiles and natronophiles could be the osmotic pressure of the chloride- and soda-rich media which is determined primarily by the electrolytic features of two types of sodium salts (see Table 3 in Banciu et al., 2004b). The calculated osmotic pressure in the 4 M NaCl solution was 2.8 times higher than that of 2 M Na2CO3 (= 4 M Na+). This result is reasonably close to the experimentally measured osmotic pressure in growth media with 4 M NaCl and 4 M Na+ soda (as a mixture of Na2CO3/NaHCO3), at 30 °C, a value which is 1.8 times higher in chloride medium than in carbonate medium. This clearly indicates that apart from the many negative effects of high alkalinity pH, the life in soda brines also has some benefits as compared to the life in NaCl-rich environment. Such physiological experiments on model organisms, complemented by theoretical simulations, could add more hard evidence to our suggestion of using the term natronophily as a special, yet different, form of halophily. As sodium sulfate brines have essentially the same electrolytic properties as sodium carbonate, one may expect that, except for the high pH, the osmoadaptation pattern of natronophiles put into Na2SO4 brines might also be similar.
3.2.2 Ectoine Is a Multivalent Compatible Solute
Ectoine (1,4,5,6-tetrahydro-2-methyl-4-pyrimidine carboxylic acid) is a derivative of aspartate. It was discovered in the phototrophic bacterium Halorhodospira halochloris (Galinski et al., 1985). In a similar manner as betaine, ectoine is accumulated by import and/or synthesis in salt-stressed cells. Unlike glycine betaine which is ubiquitous among bacteria, ectoine is the preferred compatible solute in many aerobic chemoheterotrophic eubacteria. The hydroxylated derivative of ectoine, β- (or 5)-hydroxyectoine, is more frequently found among halophilic and halotolerant Gram-positive bacteria (Severin et al., 1992; Detkova and Boltyanskaya, 2007). Ectoine alone is a minor compatible solute in the anoxygenic phototrophic Halorhodospira spp., while together with hydroxyectoine, ectoine is the dominant osmolyte in Halomonas spp., Salinivibrio spp. among the Proteobacteria, and in Marinococcus spp. and Virgibacillus spp., members of the Firmicutes (Severin et al., 1992; Grant, 2004).
By using 13C-NMR spectroscopy and HPLC analysis, Kuhlmann and Bremer (2002) have proven de novo synthesis of ectoine in a variety of Bacillus and Bacillus-related species grown under salt stress conditions. The obligate alkaliphilic B. alcalophilus synthesized ectoine, the halophilic and alkalitolerant Virgibacillus salexigens produced both ectoine and hydroxyectoine, whereas Virgibacillus pantothenticus synthesized both ectoine and proline. The ability to synthesize ectoine from l-aspartate-semialdehyde in a three-step pathway is widespread within the genus Bacillus and closely related taxa. In these Gram-positive microbes, ectoine biosynthetic genes, ectABC, encode diaminobutyric acid (DABA) acetyltransferase (EctA), DABA aminotransferase (EctB), and ectoine synthase (EctC). Expression of ect genes is controlled by salinity or osmotic pressure outside the cell (Louis and Galinski, 1997). The ectABC genes and the proteins for ectoine synthesis are highly conserved among the Proteobacteria and the Actinobacteria that synthesize ectoine (Lo et al., 2009).
In certain species (e.g., Halomonas elongata), stress conditions such as elevated temperature trigger hydroxylation of some ectoine to form hydroxyectoine. In species capable of hydroxyectoine formation, two different genes, ectD and ectE, coding for ectoine hydroxylase have been identified (ectD in Virgibacillus salexigens and ectD/ectE in the halotolerant gammaproteobacterium Chromohalobacter salexigens) (Bursy et al., 2007; García-Estepa et al., 2006).
Most ectoine synthesizing species have also developed ectoine import systems. In V. pantothenticus, another gene, ectT, encodes a protein (EctT) that is a member of the BCC carriers. The EctT transporter specifically mediates the import of ectoine and hydroxyectoine but also possesses minor uptake activities for proline and glycine betaine (Kuhlmann et al., 2011).
In the extremely halotolerant Halomonas elongata DSM 2581T, Grammann et al. (2002) discovered a novel solute transporter, termed TeaABC, that belongs to the TRAP (tripartite ATP-independent periplasmic) transporters family. The osmoregulated transporter TeaABC is encoded by three genes (teaABC), and it mediates the uptake of ectoine and hydroxyectoine. Its assumed physiological role in H. elongata would be the recovery of ectoine that is excreted or lost in the surroundings of the cell by a yet unknown mechanism. Active recovery of ectoine (and/or other organic osmolytes) after solute excretion upon osmotic downshock might be an energy-saving strategy for salt tolerance in organisms that must cope with shifts of salinity in their environment (Oren, 1999). The amino acid sequences of the small, TeaB (YP_003899344.1) and large, TeaC (YP_003899345.1) transmembrane components of TeaABC carrier from H. elongata are significantly similar (40 and 56 %, respectively) to those of the small (NP_244258.1) and large (NP_244257.1) subunits of TRAP transporter deduced from the B. halodurans genome.
In aerobic halophilic and halotolerant methylobacteria from soda lakes, the main compatible solute is ectoine, followed by glutamate and sucrose. The total pool of compatible solutes is enlarged by increasing NaCl concentration in Methylomicrobium alcaliphilum, Methylophaga natronica, M. alcalica, and M. lonarensis (Trotsenko and Khmelenina, 2002; Doronina et al., 2003a, b; Trotsenko et al., 2007; Antony et al., 2012). The gene cluster ectABC has been identified in Methylobacter marinus and the natronophilic Methylomicrobium kenyense AMO1T. In other C1-utilizing bacteria (e.g., the haloalkaliphilic Methylomicrobium alcaliphilum ML1 and Methylophaga alcalica M8, and the moderate halophilic Methylophaga thalassica 33146T and Methylarcula marina h1T) the ectABC–ask operon is additionally present, encoding aspartate kinase (Ask). Upstream the gene cluster ectABC–ask, another gene, ectR, encodes the MarR-like transcriptional regulator named EctR. In methano/methylotrophic bacteria hydroxyectoine is lacking, a fact proven by the absence of ectD-like genes (Reshetnikov et al., 2011).
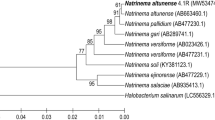
Chemolithoautotrophic natronophilic and haloalkaliphilic SOB from the genera Thioalkalimicrobium and Thioalkalibacter also produce ectoine as the prevailing compatible solute (Banciu et al., 2005, 2008). The production of ectoine in Thioalkalimicrobium aerophilum AL3T is clearly stimulated by increasing the total Na+ concentration from 0.2 to 1.2 M in the growth medium. At the highest salinity tolerated by the organism (1.2 M of total Na+), ectoine accounted for 8.7 % of the total dry weight. Another organic osmolyte detected in strain AL3T in minor amounts was glutamate. The glutamate production decreased with increasing salt concentration (Banciu et al., 2005). Apparently, species of Thioalkalimicrobium are also capable of ectoine uptake by the TRAP system. A BLASTP search for the 388 amino acid sequence of a putative extracellular solute-binding protein, family 7, from Thioalkalimicrobium aerophilum AL3T (Accession number ZP_08933362.1) has retrieved 71 and 57 % identity with the TRAP dicarboxylate transporter – DctP subunit from Thiomicrospira crunogena XCL-2 (YP_392275.1) and Marinomonas sp. MWYL1 (YP_001343266.1), respectively, and 50 % identity score with the TRAP transporter substrate-binding protein from Halomonas elongata DSM 2581 (YP_003899494.1). The phylogenetic relationships of the DctM subunit of TRAP-like proteins found in several alkaliphilic, haloalkaliphilic, and halophilic bacteria are presented in Fig. 1a. Moreover, genome sequences of Thioalkalimicrobium spp. have also indicated the possible presence of BCC transporters in these SOB.
Phylogeny of four transporters essential for haloalkaliphilic adaptation: DctM (a), RnfA/NqrE (b), MrpA/MnhA (c), and TrkH/TrkH-like proteins (d) from different haloalkaliphilic (filled circles), halophilic or halotolerant (open circles), and alkaliphilic/alkalitolerant bacteria (filled triangles/open triangles). Proteins of non-extremophilic bacterial species were used as reference sequences. Gene products or proteins were selected after BLASTP analysis. Subsequently, the selected protein sequences were aligned by ClustalW. Evolutionary analyses were conducted in MEGA5 by using the neighbor joining method. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates) is shown next to the branches. The bar indicates 10 % sequence difference. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the p-distance method and are in units of the number of amino acid differences per site. All ambiguous positions in amino acid sequences were removed for each sequence pair.
Although ectoine and betaine require comparable energy investments from the cell metabolism, there are a number of advantages of using ectoine over glycine betaine. Ectoine and hydroxyectoine are not only involved in osmoprotection of salt-stressed cells, but they are required for heat- and cold-shock adaptation of bacterial cells (Bursy et al., 2008; Kuhlmann et al., 2011). Hydroxyectoine has protective and stabilizing effects on proteins, including enzymes, when cells are exposed to thermal stress or desiccation, apparently playing same role as mannosylglycerate compounds in archaea (Lippert and Galinski, 1992; Borges et al., 2002). Ectoines might enhance the fluidity of the cell membrane through increasing the hydration of the surface and the mobility of the lipid head groups (Harishchandra et al., 2010).
Besides its prevalent osmoprotective role, ectoine can act as a nitrogen storage compound, as carbon and/or energy source (Galinski and Herzog, 1990; Khmelenina et al., 1999; Vargas et al., 2006). It can be degraded and reused by the cells. Ectoine degradation proceeds via hydrolysis to Nα-acetyl-l-2,4-diaminobutyric acid, followed by deacetylation to diaminobutyric acid. In Halomonas elongata, diaminobutyric acid can either generate aspartate or reenter the ectoine synthesis pathway, forming a cycle of ectoine synthesis and degradation. However, a comparison of the available bacterial genomes indicated that the ectoine degradation pathway exists predominantly in non-halophilic bacteria unable to synthesize ectoine. Energetic considerations indicated that ectoine turnover might be fast and it could be finely tuned by the cell’s necessity to respond promptly to changing conditions (Schwibbert et al., 2011). On the other hand, occurrence of ectoine as the main compatible solute in halotolerant and halophilic alkaliphilic aerobic methane-oxidizing bacteria as well as in low-salt-tolerant natronophilic Thioalkalimicrobium spp. is correlated with the limitation of their tolerance to high salt (<15 % w/v NaCl). At the same time, most of haloalkaliphilic phototrophic SOB (e.g., Halorhodospira spp.), natronophilic SOB (e.g., Thioalkalivibrio spp.) and SRB (e.g., Desulfonatronospira spp.), and extremely halophilic SOB (i.e., Thiohalorhabdus spp., Thiohalospira spp.) thrive under a broad range of salinity up to extreme salinity (>20 % w/v NaCl), and this ability is often associated with the use of betaine as the main compatible solute (Oren, 1999; Banciu et al., 2005; Sorokin et al., 2008b; Muyzer et al., 2011a, b). These observations allow us to conclude that the glycine betaine is a more efficient osmolyte at extreme salt concentration.
3.2.3 Glutamate as an Additional Anionic Osmolyte
Glutamate is an acidic compatible solute usually associated with low-salt response in halotolerant non-halophilic marine bacteria, moderately halophilic bacteria from the Firmicutes and the Gammaproteobacteria, and methanogenic archaea. In most cases, glutamate is accompanied by various other compatible solutes (potassium ions, glycine betaine, glutamine, ectoine, proline). As already stated, some aerobic methylotrophic and chemolithoautotrophic SOB bacteria from soda lakes are capable of salt-dependent glutamate production.
In the moderately haloalkaliphilic O. iheyensis and B. halodurans, glutamate synthesis from the branched-chain amino acids (i.e., leucine, isoleucine, and valine) is thought to play a notable role in alkaline adaptation. The putative ABC transporters for branched-chain amino acids identified in their genomes could facilitate the uptake of glutamate precursors. Since l-glutamic acid is negatively charged at pH values above its pKa (3.9 or 4.3), the converted l-glutamic acid and its accompanying proton could contribute to the cytoplasmic pH which is about two units lower than the external pH of around 10.5 (Krulwich et al., 2007; Takami, 2011).
3.2.4 Sucrose and Trehalose: Minor Osmolytes with Stabilizing Roles
Nonreducing disaccharides such as sucrose and trehalose are energy costly compounds involved in osmotic stress adaptation of many freshwater and low-salt-tolerant cyanobacteria (e.g., Synechococcus spp., Phormidium spp.) and non-halophilic and slightly halophilic bacteria when they need to adapt to elevated salt concentrations (Mackay et al., 1984; Oren, 1999). Unlike polyols and amino acid derivatives used as osmolytes, sucrose and trehalose have a lesser stabilizing effect on enzymes at high salt and have lower water solubility. Disaccharides and their derivatives are the most energetically expensive compatible solutes, and therefore, their synthesis should be carefully managed in salt-stressed cells. As main compatible solute (5 % w/w, at 2.5 M Na+), sucrose is present in the moderately natronophilic SRB Desulfonatronovibrio, together with the rare osmolyte compound N-acetylglutaminylglutamine amide (N-AGGN) found at an approximate concentration of 2 % w/w (Sorokin et al., 2011c). Sucrose, in a mixture with the amino acid glutamate, is used as a secondary compatible solute in aerobic methylotrophs from soda lakes, where sucrose increased significantly at the higher growth limit of salinity (Doronina et al., 2003a, b). As a minor osmolyte, sucrose is also present in chemolithoautotrophic SOB of the genus Thioalkalivibrio, where it reaches a maximum concentration (2.5 % of cell dry weight) at the optimal total sodium concentration (2 M) (Banciu et al., 2005).
Trehalose has multiple uses in osmoprotection, as thermolyte and as anti-drying agent in low-salt-tolerant cyanobacteria, halophilic actinomycetes, moderately halophiles, and thermophilic bacteria (Mackay et al., 1984; Alarico et al., 2005; Roberts, 2000). In the extremely halophilic actinomycete Actinopolyspora halophila grown at 24 % NaCl, Nyyssölä and Leisola (2001) showed that main compatible solute was betaine (33 % of the cellular dry weight), followed by a significant amount of trehalose (9.7 % w/w). In the moderately halophilic sulfate-reducer Desulfovibrio halophilus grown at 15 % NaCl, trehalose is accumulated as principal osmolyte in the absence of exogenous betaine (Welsh et al., 1996). In combination with ectoine, trehalose is implicated in the thermal tolerance of halophilic Chromohalobacter salexigens (Reina-Bueno et al., 2012). Recent progress on deciphering prokaryotic genomes has allowed identification of genes for trehalose synthesis in halotolerant cyanobacteria and in halophilic and haloalkaliphilic bacteria and archaea. The widely distributed pathway of trehalose synthesis that involves trehalose-6-phosphate synthase (TPS) and trehalose-phosphatase (TPP) could be inferred from genomes of Synechococcus spp., Synechocystis spp., Halalkalicoccus jeotgali, Haladaptatus paucihalophilus, Haloterrigena turkmenica, Natronobacterium gregoryi, Natrialba magadii, etc., as well as from those of the halophilic bacterium Salinibacter ruber and the haloalkaliphilic “Halanaerobium hydrogenoformans.” Another pathway converts maltodextrins (maltooligosaccharides, glycogen, and starch) to trehalose in two enzymatic steps catalyzed by maltooligosyl trehalose synthase (TreY) and maltooligosyl trehalose trehalohydrolase (TreZ), respectively (Elbein et al., 2003). Trehalose synthesis by TreY and TreZ has been observed in archaea belonging to Sulfolobus but can also be assumed from the genome sequence of the haloalkaliphilic methanotrophic bacterium Methylomicrobium alcaliphilum (Avonce et al., 2006; personal BLASTP search). It is worth mentioning that exogenous trehalose can be taken up by specialized import systems (e.g., phosphotransferase system – PTS – trehalose-specific enzyme II, and trehalose/maltose binding protein, TMBP) found in a large number of heterotrophic bacteria and archaea, as concluded from a BLASTP analysis of available amino acid sequences. Following its uptake, trehalose is phosphorylated to trehalose-6-phosphate which can be further degraded by trehalose-6-phosphate hydrolase (TreA in B. subtilis and TreC in E. coli) to provide the cells with glucose (Helfert et al., 1995; Rimmele and Boos, 1994). TreA-like proteins are present in halophilic and alkaliphilic Bacillus species (Bacillus halodurans, B. clausii, B. selenitireducens, and B. pseudofirmus OF4), as well as in Oceanobacillus iheyensis and Halobacillus halophilus.
3.2.5 K+ Is the Main Osmolyte in Anaerobic Extreme Haloalkaliphiles
The natronophilic acetogenic Natroniella acetigena from the order Halanaerobiales as well as halophilic thermoalkaliphiles belonging to the Natranaerobiales (Natranaerobius spp. and Natronovirga spp.) have adopted the salt-in strategy to withstand high salinity conditions. In N. acetigena, the measured intracellular concentrations of K+ (ca. 0.9 M) were almost two orders of magnitude higher than the K+ level in the medium. At the same time, intracellular Na+ and Cl− concentrations were close to the extracellular values (Detkova and Boltyanskaya, 2007). Intracellular K+ concentration in representatives of the Natranaerobiales is reliably kept at constant values of 0.2–0.3 M over a wide range of external K+ (8–400 mM), at optimal or suboptimal pH and salinity. Apparently, in halophilic alkalithermophiles of the order Natranaerobiales, potassium homeostasis is tightly regulated. Accumulation of K+ is accompanied by that of Cl- ions, which reach molar concentration. However, the sum of both ions does not equilibrate the cytoplasm osmolarity with that of the outer environment. The genome analysis of Natranaerobius thermophilus has revealed the ability of this organism to import and even to synthesize glycine betaine. Apparently, this finding was confirmed experimentally. Intracellular concentrations of glycine betaine and glutamate have been detected in molar concentrations in Natranaerobius thermophilus. Such an observation, if confirmed by a detailed experimental study, would constitute a first known case of a fermentative extreme halophile capable of producing organic compatible solutes and use the “salt-in” strategy concomitantly (Mesbah and Wiegel, 2012).
3.3 Adaptation of the Oxidative Phosphorylation (OXPHOS) Machinery to Overcome Low PMF
In highly saline and alkaline environments, the burden due to elevated osmotic pressure is doubled by an extremely low level of H+ concentration. The bulk medium where haloalkaliphiles and natronophiles live is weakly supporting a favorable proton-motive force (pmf) to drive ATP synthesis by oxidative phosphorylation pathway as postulated by Peter Mitchell. Despite the apparently adverse pH gradient, alkaliphilic and, implicitly, the haloalkaliphilic prokaryotes flourish at alkaline conditions. Moreover, haloalkaliphiles are splendidly facing the challenge of expensive life in a hyperosmotic medium by synthesizing energetically costly compatible solutes. To respond to the multiple extreme factors, which additionally include high irradiation and, sometimes, relatively high temperatures (45–60 °C), true haloalkaliphilic and natronophilic microbes must have an efficient energy metabolism. The adaptative modifications are met not only in the core of the ATP synthesis process but are also found in the respiratory chain components and in the overall organization of the cytoplasmic enzyme machinery. In the following we will briefly discuss the alkaliphilic adaptations of the respiratory chain components as it is best known from several model organisms.
3.3.1 Features of Alkaliphilic F-Type ATP Synthase
Three major ways of generating the ATP are known in various haloalkaliphiles and natronophiles: (i) membrane-bound oxidative phosphorylation in aerobic chemotrophs, (ii) photophosphorylation linked to photosynthetic inner membranes of aerobic and anaerobic phototrophs, and (iii) substrate-level phosphorylation of anaerobic (or fermentative) bacteria. Mixed sources of ATP could be found in non-obligately metabolic situations such as facultatively aerobes and in anaerobic phototrophs that rely on either (i) and (iii) or (ii) and (iii). One of the most intriguing types of energy metabolism occurs in obligately aerobic chemolithoautotrophic alkaliphilic bacteria, where anabolic maintenance and active transport processes rely almost exclusively on oxidative phosphorylation which is directly affected by the apparently suboptimal proton electrochemical gradient.
Oxidative phosphorylation (OXPHOS) is a membrane-localized metabolic pathway that couples the generation of an electrochemical gradient with ADP phosphorylation to form ATP. In prokaryotes the transmembrane electrochemical gradient, usually based on protons, is the result of the respiration process. Respiratory chain components perform simultaneous electron transfer from an internal or external donor to an oxygen molecule as the final electron acceptor, coupled with proton pumping across the membrane toward the outer or positive (P) side of the cell. The free Gibbs energy stored in the electrochemical gradient is transferred to chemical bonds within the ATP molecule by the ATP synthase activity of F-type primary pump. Universally found in all three domains of life, F-ATP synthase is a reversible enzyme that catalyzes both ATP synthesis and hydrolysis. The hydrolytic activity of the enzyme is mainly observed in anaerobes, while in many other bacteria, this function is low or latent. The equilibrium between the hydrolytic and the synthetic function of F-type ATP synthase is carefully controlled to maintain appropriate cytoplasmic ATP and ADP concentrations. The catalytic domain of the ATP synthase complex (also termed F1) is located in the cytoplasm. The F1 portion of ATP synthase consists of α3β3γδε subunits and is relatively well conserved among all organisms. The transporting part of the F-ATP synthase (F0) is a transmembrane complex made of a variable number of subunits (ab2c10–15 in bacteria) facing the external milieu of the cell, as well as the hydrophobic membrane. The F0 factor of ATP synthase is highly variable among different prokaryotic species. The critical issue that functioning of ATP synthase toward ATP synthesis must address is the apparently low electrochemical gradient. Therefore, it is expected that specific structural adaptations of this key enzyme to be found in the a-, b-, and c-subunits of the F0 part.
The alkaliphilic adaptations of the F-ATP synthase and OXPHOS both at the physiological and the genetic levels have been documented in-depth in the aerobic, facultatively alkaliphilic bacterium Bacillus pseudofirmus OF4 (for review, see Hicks et al., 2010). As previously mentioned in this review, the facultative alkaliphilic and/or halophilic organisms are appropriate models to study the changes and adaptations at the genotype and phenotype levels.
There are certain key features of the alkaliphilic F1F0-ATP synthase from B. pseudofirmus OF4 that allow ATP synthesis under conditions of low pmf, and they are summarized below together with general and particular considerations regarding other alkaliphiles.
1. The functioning of the alkaliphilic F-ATP synthase toward ATP synthesis is H + -coupled. So far, all F-type ATP synthases with a high rate of ATP synthesis from true and facultative alkaliphiles are proton-coupled. The recent examples of proton-driven F1Fo-ATP synthase deduced from genome analysis are the aerobic lithoautotrophic extreme natronophilic Thioalkalivibrio spp. that demands high sodium concentration for optimal respiratory activity and growth (Banciu et al., 2004a; Muyzer et al., 2011a, b) and the anaerobic natronophiles with respiratory metabolism Desulfurivibrio alkaliphilus and Desulfonatronospira thiodismutans (Sorokin et al., 2008b, d; Hicks et al., 2010). In anaerobic alkaliphilic, alkalitolerant, or haloalkaliphilic bacteria, ATP is almost entirely formed by substrate-level phosphorylation. However, Na+-driven F-type ATP synthase is present in several species with various degrees of alkaliphily (e.g., the alkalithermophilic Clostridium paradoxum and the halophilic thermoalkaliphilic Natranaerobius thermophilus) (Ferguson et al., 2006; Mesbah and Wiegel, 2011). The only cyanobacterium with Na+-translocating ATPase activity reported so far is the alkaliphilic and halotolerant Aphanothece halophytica (Soontharapirakkul and Incharoensakdi, 2010). In these bacteria, a membrane-bound V-type ATPase works at the expense of ATP to translocate Na+ outwardly, thus contributing to sodium homeostasis in cytoplasm but not to the energy generation. Same type of the sodium-coupled ATPase has been detected in the genomes of a natronophilic clostridium Dethiobacter alkaliphilus (Sorokin et al., 2008d; Hicks et al., 2010) and a fermentative low G+C firmicutes Amphibacillus (Kaieda et al., 1998; Satoh and Koyama, 2005).
2. The intimate construction of alkaliphilic F-ATP synthase favors the catalysis of ATP synthesis and blocks the hydrolytic activity. The regulation of bacterial F-ATP synthase activity is achieved by two major mechanisms. Tight but reversible binding of ADP as magnesium salt is an effective inhibitory process that slowers the ATPase activity in all known F-ATP synthases (Minkov et al., 1979; Feldman and Boyer, 1985; Konno et al., 2011). In bacterial and chloroplast F-ATP synthase, the ε subunit is an “endogenous inhibitor.” Fine-tuning of conformational changes in the ε subunit is controlled by the pmf and the ATP/ADP ratio (Kato-Yamada et al., 1999; Suzuki et al., 2003). It is very likely that haloalkaliphilic ATP synthase is not exceptional in this regard. For the Na+-translocating ATP synthase from Natranaerobius thermophilus, with a high hydrolytic activity and a low capacity for ATP synthesis, Mesbah and Wiegel (2011) have proposed a regulatory mechanism mediated by the ε subunit.
3. Most of adaptive modifications (as compared with the neutralophilic ATP synthase) have been identified in the F 0 factor.
In the alkaliphilic Bacillus or Bacillus-related species (i.e., B. alcalophilus, B. halodurans, B. pseudofirmus OF4, B. selenitireducens, and Caldalkalibacillus thermarum TA2.A1), the c-ring is made up of 13 subunits, a slightly higher number than that of decameric c-ring of most neutrophilic cells. Accordingly, the H+ to ATP ratio in these alkaliphiles would be 4.3, higher than the more usual 3.3 proton equivalents for one mol of synthesized ATP as calculated for neutrophilic ATP synthase (Hicks et al., 2010). Comparative analysis of genomic sequences revealed alkaliphilic-specific sequences in the genes coding for a- and c-subunits as well as a different pattern of alkaliphilic motifs in the Gram-positive alkaliphiles as compared with Gram-negative ones. Moreover, different levels of alkaliphily are accordingly reflected in these sequences (Hicks et al., 2010). Two specific amino acid sequences related to alkaliphily in Bacillus species were observed in the primary structure of the c-subunit: AxAxAxA – or at least three alanine residues, and PxxExxP – two proline residues flanking the glutamic acid with a critical carboxylate residue. The first motif is apparently contributing to spatial ordination of the c-ring, while the second is responsible for ion binding (Wang et al., 2004; Liu et al., 2009).
Another peculiarity of the primary structure of alkaliphilic c-subunits is the presence of a threonine residue (Thr33) near the loop edge of the hairpin-like structure, which may influence the dynamics of the rotor part of the ATP synthase edifice. Recently, a high resolution crystal structure of the rotor ring in B. pseudofirmus OF4 unveiled a rather “tulip beer glass” shape of the c-subunit ring, different from the “hour-glass” shape of the c-rotors in other bacteria. At the same time, the alkaliphilic motifs in c-subunits promote tight proton binding in the ion-binding site in the form of a tetra-coordinated hydronium ion (H3O+) (Preiss et al., 2010).
A brief comparative analysis by multiple alignments of available sequences for ATP synthase c-subunits from several haloalkaliphilic species (the aerobic Gram-negative Thioalkalivibrio spp., Thioalkalimicrobium spp., and Methylomicrobium alcaliphilum; anaerobic Gram-negative Thiorhodospira sibirica and Ectothiorhodospira spp.; anaerobic Gram-positive Alkaliphilus spp.) indicated the presence of “non-alkaliphilic” motif GxA/GxGxG, the same motif as found in c-subunit of Escherichia coli or Oceanobacillus iheyensis (see also Hicks et al., 2010). When searching for the second motif of c-subunit, analogous to alkaliphilic PxxExxP, same observation applied: non-alkaliphilic GxxD/ExxP/T motif has been detected in all surveyed haloalkaliphiles. Thus, it seems that minute changes in the last amino acid of the N-terminus motif would influence the kinetics of ion binding at the nearby glutamate/aspartate residue. Thr33 was not found, but all analyzed extremophiles contain the nonpolar Ala33 residue instead.
In the stator part of F1F0-ATP synthase, the a-subunit of true alkaliphilic Bacillus spp. contains at least two crucial amino acids, Lys180 and Gly212, that are not found in the neutrophilic counterpart. Situated at the interface between the a- and the c-ring, Lys180 and Gly212 are essential for ATP synthesis at elevated pH, being probably implicated in proton capturing and transferring to the rotor ring of ATP synthase (Fujisawa et al., 2010). Our brief search within the sequence databases has indicated the substitution of Lys180 and Gly212 in the haloalkaliphilic Thioalkalivibrio spp., Thioalkalimicrobium spp., Methylomicrobium alcaliphilum, and Thiorhodospira sibirica in an E. coli-like pattern (Lys180/Gly212 being replaced with Gly/His or Lys). These substitutions in places that face each other in transmembrane helices 4 and 5, respectively, could ensure similar microenvironment for transferring the protons as it does in the a-subunit of alkaliphilic Bacillus spp. Additionally, the external surface of the a-subunit in alkaliphilic Bacillus spp. is somewhat more polar than the analogous region of non-alkaliphilic Bacillus species or E. coli. This region may have an enhanced capacity of retaining H+ on the outer surface of the membrane (Wang et al., 2004).
The molecular adaptations of ATP synthase from alkaliphilic, aerobic, and Gram-positive species only partially explain the ability of such organism to withstand a broad range of pH and/or extreme alkalinity. In particular, the phosphorylation potential (DGp), the parameter that reflects the energy required to sustain the [ATP]/[ADP][Pi] ratio, is unexpectedly high under conditions in which the pmf is relatively low. As discussed by Hicks et al. (2010), in B. pseudofirmus OF4 grown at pH 10.5, the measured pmf is –50 mV, while ΔGp is 478 mV. Similarly, in the thermophilic Caldalkalibacillus thermarum grown at pH 10, the observed pmf was –78 mV, while ΔGp was around 430 mV. Close values (pmf = –58 to –89 mV, and ΔGp = 477–491 mV) were measured in a sucrose-energized cell suspension of Natranaerobius thermophilus (Mesbah et al., 2009). In all situations, the observed ΔGp is close to that measured in the neutrophiles. However, in the obligately anaerobic fermentative haloalkaliphile, the measured ΔGp does not necessarily reflect the capacity of ATP synthesis by OXPHOS. Based on the relation [H+]/[ATP] = ΔGp/pmf and assuming the value of 3.3 for [H+]/[ATP], the theoretical pmf value that supports the observed ΔGp would be around two times higher. A slightly increased [H+] to [ATP] ratio of 4.3, corresponding to tridecameric c-rings of B. pseudofirmus and C. thermarum, does not suffice to obtain such a phosphorylation potential (Hicks et al., 2010). In this light, the hypotheses of localized proton gradients or local proton microcircuits that boost the efficiency of proton transfer, sequestration, and transport from respiratory chain components to the ATP synthase complex have been now revived since Williams proposed the proton circuits theory in 1961 (Williams, 1988; Guffanti and Krulwich, 1992; Hicks et al., 2010). In support of this theory, some evidence has risen. Localized microcircuits of protons and fast proton diffusion alongside the membrane surface have been directly measured in liposomes by using a fluorescence correlation spectroscopy (FCS)-based approach (Brändén et al., 2006). Several other reports exist on the possibility of faster diffusion of protons along the lipid and/or protein surface of membranes than the expelling of protons out into the bulk phase (Heberle et al., 1994). Protons extruded during respiration were apparently retained close to the outer surface of the cell membrane in alkaliphilic B. clarkii K24-1U, but this observation needs further confirmation (Yoshimune et al., 2010).
3.3.2 Adaptation of Respiratory Chain Components at Alkaline pH
The aerobic facultatively alkaliphilic Bacillus pseudofirmus OF4 as well as B. halodurans C-125 and Oceanobacillus iheyensis have a primary H+-extrusion respiratory chain with caa3- and bd-type terminal oxidases (Krulwich et al., 1998; Takami et al., 2000, 2002). Additionally, cytochrome bo3 oxidase is inferred from the O. iheyensis and B. halodurans genomes (Takami et al., 2000, 2002). Cytochrome caa3 oxidase is induced during growth at alkaline pH. Liu et al. (2007) observed close physical interaction between caa3 oxidase and F1F0-ATPsynthase that might increase the efficiency of proton transfer to the F-ATP synthase. Cytochrome bd is a non-proton-pumping quinol oxidase that is upregulated during the alkaline pH adaptation in E. coli K-12 concomitantly with a decrease in expression of proton-extruding complexes (cytochrome o and types I and II NADH dehydrogenases) of the respiratory chain. Such a concerted response would probably diminish the export of protons from the cytoplasm during generation of pmf (Maurer et al., 2005).
The midpoint redox potential of cytochrome c, one of the main components of the respiratory chain, is considerably lower (< +100 mV) in alkaliphilic than in neutrophilic bacteria (approx. +220 mV). At the same time, the cytochrome caa3 oxidase (complex IV) that accepts electrons from cytochrome c has an apparently normal redox potential (approx. +250 mV) (Muntyan and Bloch, 2008). Moreover, cytochrome c is highly abundant in alkaliphilic Bacillus when grown at alkaline pH (for reviews, see Hicks and Krulwich, 1995; Krulwich et al., 1998). It is suggested that the overall construction of the respiratory chain permits a quicker and more efficient H+ translocation and electron flow along the membrane in alkaliphiles than in neutralophiles (Goto et al., 2005).
In the aerobic natronophilic SOB, dominant cytochromes are both of the c- and b-type, in Thioalkalivibrio spp., and c-type only in Thioalkalimicrobium spp. Terminal oxidases have been identified as caa3-, cbb3- and o-type in the first genus, and cytochrome cbb3 type in the latter. Presence of cytochrome cbb3 oxidase, a low efficient proton pump (H+/e− = 0.2–0.75) with high oxygen affinity (K m ∼ 7 nM) and a relatively low midpoint potential of its active site heme (E m,7 ∼ −60 mV), is indicative of microaerobic adaptation (Pitcher et al., 2002; Rauhamäki et al., 2009). Microoxic conditions are likely in the saline environment where oxygen solubility is affected by presence of salts. In E. coli cytochrome o oxidase is produced under oxygen-rich conditions, does not extrude protons, and has a relatively low oxygen affinity (K m = 1.4–2.9 μM) (Tseng et al., 1996). Cytochrome o oxidase oxidizes ubiquinol but not cytochrome c, representing an alternative route for electron transfer. Its presence in Thioalkalivibrio spp. with two altogether different terminal oxidases and low-potential cytochrome b may suggest a highly controlled, multiple-way electron flow and versatility of growing under variable oxic/microoxic conditions.
The high sodium concentration in the external medium of haloalkaliphilic bacteria would have a dramatic effect on their internal milieu unless they possess extruding systems to adjust their internal sodium level. One of such regulators of Na+-homeostasis could be the Na+-dependent NADH:quinone oxidoreductase (Nqr or type III NAD dehydrogenase). Nqr is a respiration-dependent primary sodium pump that couples sodium extrusion with electron transfer from NADH to ubiquinone to generate smf. Nqr is present in many Gram-negative marine bacteria such as Vibrio spp. and Shewanella spp. where it is related to the absolute sodium requirement for growth (Oh et al., 1991). Coexistence of proton-translocating NADH dehydrogenase (NDH-1) and Nqr is predicted from the genomes of pathogens (e.g., Pseudomonas spp., Neisseria spp., Yersinia spp.), obligate chemolithoautotrophs (Nitrosomonas europaea), and euryarchaeota (Methanosarcina acetivorans) (Melo et al., 2004). Homologous nqr genes (rnfABCDGE) that code for the Rnf complex were recently identified in the genome of the natronophilic Thioalkalivibrio spp. and Thioalkalimicrobium spp., as well as in other soda-loving bacteria (Fig. 1b). In Thioalkalivibrio spp., the rnf operon coexists with the nuo operon (nuoABCDEFGHIJKLMN) encoding type I proton-pumping NADH dehydrogenase (NDH-1). Rnf are bacterial redox-driven ion pumps with flavin-binding subunits (D and G) homologous to Na+-Nqr B and C subunits (Steuber, 2001; Backiel et al., 2008). Genes coding for Nqr as well as for NDH-1 (nuo genes) were also discovered in the genome of haloalkaliphilic SRB Desulfurivibrio alkaliphilus (Pereira et al., 2011). Coexistence of both types of NADH dehydrogenases suggests a finely tuned redox metabolism in association with sodium extrusion from the cytoplasm.
Type II NADH dehydrogenase (energy-uncoupled NADH dehydrogenase or NDH-2) capable of sodium export as well as of Na+/H+ antiport (NapA) activity was described in the moderate halophile Halobacillus dabanensis D-8 (Yang et al., 2006). Genes for NDH-2 exist in the Gram-positive alkaliphilic B. halodurans C-125, B. clausii, and B. pseudofirmus OF4. Their predicted products are two types of NHA-2. NHA-2A found to have high sequence similarity (>70 %) among alkaliphilic Bacillus, while the gene for NHA-2B of Bacillus pseudofirmus OF is 56 % similar to that of putative “Na+-NADH dehydrogenase (NapA)” from Halobacillus dabanensis. None of the NHA-2 expressed proteins has Na+/H+ antiport activity in B. pseudofirmus OF4, and they are supposed to play a gating role for electron entry in the respiratory chain (Liu et al., 2008). Interestingly, the facultatively alkaliphilic and halotolerant marine bacterium Oceanobacillus iheyensis seems to rely only on NDH-2, lacking other types of respiratory NADH dehydrogenases (Melo et al., 2004).
Isoprenoid (respiratory) quinones function as electron and proton carriers in respiratory and photosynthetic chains. Quinones are also known as antioxidants. Ubiquinones (high-potential quinones, E m = +70 to +112 mV) are found in eukaryotes and are widespread in aerobic alpha-, beta-, and gammaproteobacteria. Menaquinones (low-potential quinones, with E m7.5 between –110 and –70 mV) are typical for prokaryotes and are often related to anaerobic metabolism (Nowicka and Kruk, 2010). In many haloalkaliphilic, alkalitolerant halophilic, and moderately halophilic Gram-positive bacteria, the major respiratory quinone is menaquinone (MK)-7 (e.g., in Bacillus locisalis, B. qingdaonensis, B. chagannorensis, Salsuginibacillus kocurii) (Márquez et al., 2011; Wang et al., 2007; Carrasco et al., 2007). MK-7 is also dominant in the membranes of alkalitolerant halophilic, moderately halophilic, and alkaliphilic halotolerant Gram-negative bacteria belonging to Virgibacillus spp., Mongoliicoccus spp., and Mongoliitalea spp. (Chen et al., 2009; Liu et al., 2012; Yang et al., 2012). Haloalkaliphilic species of Halomonas (H. campaniensis, H. desiderata, H. campisalis) contain MK-9 and MK-8 as dominant quinones. In aerobic natronophilic Thioalkalivibrio spp. and Thioalkalimicrobium spp. that belong to the Gammaproteobacteria, the dominant respiratory quinone is the high midpoint potential ubiquinone-8 (UQ-8) (Sorokin and Kuenen, 2005). A combination of ubiquinones and menaquinones is characteristic of phototrophic sulfur bacteria of the genera Ectothiorhodospira and Halorhodospira. MK-7 and UQ-7 are major respiratory quinones in Ect. magna, Ect. shaposhnikovii, and Ect. vacuolata (Bryantseva et al., 2010). The closely related haloalkaliphilic Halorhodospira halophila distinctively utilizes two menaquinones, MK-8 and MK-7, for the photosynthetic electron transfer reaction and UQ-8 for the respiratory reaction (Schoepp-Cothenet et al., 2009).
3.4 NA+-Dependent Flagellar Movement
Naturally, the concentration of sodium ions is higher in the surrounding medium than within the cell. When speaking of alkaliphilic, halophilic, and haloalkaliphilic bacteria, the favorable (inwardly oriented) sodium gradient in the form of an electrochemical sodium gradient or sodium-motive force (smf) is exploited in several ways. Flagellar movement driven by smf has been well documented in haloalkalitolerant Vibrio spp. and in alkaliphilic Bacillus spp. and haloalkaliphilic bacteria (Atsumi et al., 1992; Fujinami et al., 2009; Krulwich et al., 2007). A Na+-dependent flagellar motor, similar to PomAB of Vibrio spp. and MotPS of Bacillus spp., was predicted from the genomes of Thioalkalivibrio sulfidophilus HL-EbGr7 and Thioalkalivibrio K90mix. These motor proteins are also closely related to a similar protein of Halorhodospira spp. Remarkably, the sodium-dependent movement coexists with proton-driven MotB-like flagellar motors in Thioalkalivibrio spp., suggesting that the chemotaxis of these bacteria is finely regulated through a complex relationship between osmo- and pH-sensing (Muyzer et al., 2011b). In low-salt-tolerant natronophilic Thioalkalimicrobium spp. and Methylomicrobium alcaliphilum, the genome sequences indicated the presence of H+-dependent flagellar motility, both putative OmpA/MotB complex and MotA-like flagellar stator protein being present. In M. alcaliphilum, however, MotA (YP_004918404.1) and PomA (YP_004917195.1) gene products coexist.
3.5 pH and Ion Homeostasis in the Cytoplasm of Haloalkaliphiles
The most important strategy of regulating cytoplasmic pH in bacteria depends on the activity of primary and secondary proton transporters. Primary H+-pumps comprise proton-extruding complexes of the respiratory chain and H+-driven F-ATP synthase, while secondary proton transporters are antiporters that exchange monovalent cations for protons (Padan et al., 2005; Krulwich et al., 2011). The key antiport system of halo- and alkaliphiles introduces H+ while extruding Na+ that inherently accumulates within the cell living at high salinity and/or alkalinity. The alkaliphilic and/or haloalkaliphilic Na+/H+ antiporters are part of a constellation of many types of transporting proteins that seems to have minute particular adaptations in almost every species. The concerted result of membrane transport activity is the accomplishment of ionic (i.e., H+, Na+) and volume homeostasis.
3.5.1 Na+/H+ Antiporters
Presently, Na+/H+ antiporters are classified into five families (NhaA to NhaE) in the monovalent cation–proton antiporter (CPA) superfamily (http://www.tcdb.org/). Nha’s differ in their structure, topology, substrate affinity, stoichiometry, and kinetics of ion exchange.
Members of the Na+/H+ antiporter NhaA family are commonly found among all living cells and make an essential contribution to the Na+ and pH homeostasis during alkaline adaptation of non-extremophilic and extremophilic organisms (Padan et al., 2001). The activity of electrogenic NhaA is regulated by the cytoplasmic pH, and it catalyzes the inward translocation of 2H+ in exchange with 1Na+ (or Li+) that is extruded, resulting in a net positive charge accumulation inside the cytoplasm. The secondary transport achieved by NhaA is energized by the electric potential (∆Ψ) component of pmf. The prototype member of this family is NhaA from E. coli (Ec-NhaA). Ec-NhaA has a cytoplasmic pH-sensing domain in loop VIII-IX. It has a high affinity for sodium with apparent K m = 0.2 mM Na+ at pH 8 (Gerchman et al., 1993; Tzubery et al., 2004). Expression of the nhA gene is modulated by Na+ at the transcriptional level, while the pH-dependent activity is an intrinsic function of the translated protein. The growth of E. coli at alkaline pH supported by the combined action of two transporters, NhaA and a multidrug transporter MdfA. Apparently, NhaA is aiding non-alkaliphilic cells to withstand alkalinity up to pH 9, while MdfA, which has Na+/H+ as well as K+/H+ antiport activity in addition to efflux of multiple drug substrates in exchange for H+, takes over at higher pH (Padan et al., 2005). Ec-NhaA has maximal activity at pH 8.5, and growth of E. coli cells at pH >9 would raise the cytoplasmic pH at inhibitory value for NhaA activity. Ec-NhaA has many homologs and paralogs in bacteria and archaea (for review, see Padan, 2008). A survey of the database (BLASTP) for available protein sequences or gene products closely related to NhaA has revealed few homologs in halophilic Halomonas spp. and Chromohalobacter salexigens.
Two members of the NhaC family of Na+/H+ antiporters have been originally uncovered in Bacillus pseudofirmus OF4 and Bacillus subtilis, respectively. One member (also designed as Bp-NhaC) is a relatively high-affinity, electrogenic Na+/H+ antiporter with a minor role in alkaliphily (Ito et al., 1997); the other is a paralog described as mediating an electroneutral malate-H+/Na+-lactate antiport important in malate uptake at low pmf (Wei et al., 2000). Bp-NhaC of B. pseudofirmus remains the only member of the NhaC family that is functionally characterized. Genes coding for NhaCs exist in both Gram-negative bacteria and Gram-positive bacteria as well as archaea (i.e., Haloferax spp.). An NhaC homolog is predicted from the genome of the halophilic thermoalkaliphilic Natranaerobius thermophilus JW/NM-WN-LF (Zhao et al., 2011). The nhaC gene product of N. thermophilus showed high amino acid identity (42–44 %) with similar products in the genomes of the alkaliphilic Bacillus clausii KSM-K16, B. halodurans C-125, the obligately anaerobic, non-halophilic and moderately alkaliphilic Alkaliphilus metalliredigens QYMF and A. oremlandii OhILAs, as well as with a partial protein sequence from Halomonas boliviensis LC1 (acc. no. ZP_09188066).
Since the first reports on Na+/H+ antiport activity in the moderately halophilic Salinivibrio costicola and the slightly halophilic V. parahaemolyticus (Hamaide et al., 1983; MacLeod et al., 1988), evidence arose for the simultaneous presence of NhaA-, NhaB-, and NhaD-type antiporters in halophilic bacteria of Vibrio spp. Genes coding for NhaA and NhaB from V. parahaemolyticus complemented mutants unable to grow at neutral pH and high salinity (0.5 M LiCl) or at alkaline pH (8.5) in the presence of low salt concentration (0.05 M LiCl). Expression of nhaA and nhaB was proposed to be pH-regulated (Kuroda et al., 2005). Genes for the latter antiporter, NhaD (or Vp-NhaD), identified for the first time in V. parahaemolyticus by Nozaki et al. (1998), were also isolated and their product functionally characterized in Vibrio cholerae (Vc-NhaD) and the haloalkaliphilic Alkalimonas amylolytica (Aa-NhaD) (Dzioba et al., 2002; Ostroumov et al., 2002; Liu et al., 2005).
NhaD proteins catalyze Na+(Li+)/H+ antiport at a yet unknown stoichiometry. Homologs of nhaD gene products are found among many proteobacteria, including truly halophilic Halomonas spp. (Kurz et al., 2006). Activity of NhaD is optimal at alkaline pH (pHopt = 8 for Vc-NhaD; 8.5–9 for Vp-NhaD, and 9.5 for Aa-NhaD). The activity of NhaD from Alkalimonas amylolytica has a relatively low Na+ affinity (K m = 0.5 mM Na+) and it is optimal at a concentration of 600 mM Na+ (Liu et al., 2005). These characteristics of haloalkaliphilic NhaD (Aa-NhaD) seem in accordance with the supposed chemical parameters of the cytoplasm during growth at saline and alkaline conditions. However, pH homeostasis in Alkalimonas amylolytica, capable of growing optimally at pH 10–10.5 and 2–3 % (w/v) NaCl, is achieved upon combined activities of at least three other ion transporters: the K+/H+ antiporter (Aa-NhaP), the K+ efflux pump (Aa-KefB, homolog of glutathione-regulated K+ efflux protein B, KefB from E. coli), and, to a lesser extent, the calcium/cation antiporter (Aa-CaxA) (Wei et al., 2007).
3.5.2 Mrp and Mrp-Like Antiporters
In aerobic alkaliphilic Bacillus strains, growth at alkaline pH is strictly dependent on the presence of external Na+. Therefore, in these bacteria, pH homeostasis is mainly accomplished by a variety of Na+/H+ antiporters (Krulwich et al., 2011). A large hetero-oligomeric Mrp antiporter that extrudes Na+ (K+ or Li+) in exchange for H+ has a crucial role for survival at high pH (Ito et al., 2001; Morino et al., 2008, 2010). Unlike Ec-NhaA that is a single-gene product, the multidrug resistance efflux pump Mrp from Bacillus spp. is a seven-component system (MrpA–MrpG) encoded by an operon containing seven genes. Originally described in Bacillus halodurans C-125 by Hamamoto et al. (1994), it turned out that Mrp genes are virtually spread in many Gram-negative and Gram-positive bacteria and archaea, with vital contribution to cytoplasmic pH regulation and general physiology of microorganisms (Swartz et al., 2005). Mrp-like clusters from cyanobacteria have been demonstrated to be involved in salt stress tolerance and CO2 deficiency-induced expression (Blanco-Rivero et al., 2005). A full Mrp-like complex (Ap–MrpA–G) with Na+/H+ and Li+/H+ antiport activity was functionally described in the halophilic and alkalitolerant cyanobacterium Aphanothece halophytica (Fukaya et al., 2009). An interesting tripartite clustering of the genes bicA, napA1-2, and an mrp-like gene was described in A. halophytica. The first gene codes for Na+-dependent HCO3 − uptake, the second for a pH-regulated Na+/H+ antiporter, active at alkaline pH (Price, 2011; Wutipraditkul et al., 2005). A model of cooperation between the three transporters was proposed in A. halophytica. At alkaline and saline conditions, when CO2 becomes limiting, the Na+ extruded by Na+/H+ antiport catalyzed by NapA1-2 and the Mrp-like complex drives the inward flux of HCO3 −. The pH of the cytoplasm is maintained at a tolerable level by simultaneous proton import (Fukaya et al., 2009). Genes of the mrp-like operon (mnhA-G), together with those for other cation/proton antiporters (nhaP), have been reported in the natronophilic Thioalkalivibrio spp. (Muyzer et al., 2011a, b) as well as in the genomes of the arsenite-oxidizing Alkalilimnicola ehrlichii and Halorhodospira halophila (see also Fig. 1c).
3.5.3 K+ Transport and Intracellular Homeostasis
It is expected that the alkaliphiles retain an outwardly directed K+ gradient. Therefore, K+/H+ antiporters might be energized by the electrochemical potassium gradient to drive protons inside the cell. However, K+/H+ antiporters are not expected to play central roles in cytoplasmic acidification in alkaliphiles as the removal of intracellular K+ is not desirable. Accumulation of (monovalent) cations in the halo/alkaliphilic cytoplasm and particularly of K+ is a strategy either to neutralize the charge of anionic osmolytes (i.e., glutamate, chloride ions) or to compensate for the risk of Na+ toxicity at high intracellular pH (Padan and Krulwich, 2000).
Potassium transporters participate in the overall pH, K+, and volume homeostasis in non-halophilic or slightly halophilic bacteria (e.g., E. coli, Vibrio alginolyticus) (Radchenko et al., 2006; for a review, see Epstein, 2003), as well as in osmoregulation of Halanaerobiales and haloarchaea (Oren, 2002). K+ transporters that contribute to pH and K+ homeostasis of halo- and/or alkaliphiles are NhaPs belonging to the monovalent cation/proton antiporter-1 (CPA1) family and members of the K+ transporter (Trk) family. The Aa-NhaP – K+(NH4 +)/H+ antiporter was characterized in Alkalimonas amylolytica (see above) (Wei et al., 2007). The Ap-NhaP – Na+(Ca2+)/H+ antiporter has been functionally described in the halotolerant cyanobacterium Aphanothece halophytica. It exhibits Na+/H+ antiporter activity over a wide pH range (5–9) and shows high Ca2+/H+ antiporter activity at alkaline pH (Waditee et al., 2001).
The K+ uptake protein TrkH could be inferred from the genome sequences of natronophilic and haloalkaliphilic Thioalkalimicrobium spp., Thioalkalivibrio spp., Methylomicrobium alcaliphilum, and Desulfovibrio alkaliphilus (Fig. 1d). It was also characterized in Alkalimonas amylolytica (Guo et al., 2009). Kinetics experiments revealed that the Aa–TrkAH complex has an optimal activity at pH >8.5 and it is salt tolerant. The TrkAH complex has a channel-like shape and a large acidic extracellular region. It catalyzes NAD+-regulated K+ uptake possibly coupled with H+ import (Domene and Furini, 2012).
3.5.4 Sodium Import Closes the Sodium Cycle in Salt- and Alkaline-Adapted Cells
Two major routes for Na+ entry in prokaryotic cells are known: Na+/solute symport and Na+ diffusion mediated by channels (including the flagellar Na+ channel).
Na+-dependent solute uptake is essential in providing sodium for antiport activity as well as nutrients (i.e., sugars, amino acids) or precursors (such as choline) for the purpose of general metabolism including osmoadaptation. Na+/solute symporters are ubiquitously spread in many bacteria and archaea as well as in eukaryotes. They belong to the sodium/solute symporter (SSS) family according to the Transporter Classification system (Saier et al., 2009). Activity of Na+/solute symporters is energized by the electrochemical Na+ gradient (smf) (Jung, 2001). Na+-mediated import of l-proline by homologs of the PutP symporter from E. coli and Vibrio parahaemolyticus, respectively, is largely found in pathogens, slightly halophilic marine bacteria, extreme halophilic bacteria (Salinibacter ruber, Halomonas elongata), and archaea (Halobacterium spp., Haloferax spp.) (Jung et al., 2012).
Na+-coupled import of a nonmetabolizable solute (α-aminoisobutyric acid) has been reported in alkaliphilic Bacillus as early as 1985 by Krulwich et al. Glutamate and sucrose uptake are dependent on the sodium gradient in Caldalkalibacillus thermarum TA2.A1. Apparently the electrical component of the smf is the driving force for solute uptake in this thermoalkaliphile (Peddie et al., 1999, 2000). Among the anaerobic halophilic and thermoalkaliphilic bacteria, Na+-coupled solute symport was detected in Natranaerobius thermophilus (Zhao et al., 2011). In comparative physiological experiments with anaerobic alkaliphilic bacteria from soda lakes, Pitriuk et al. (2004) concluded that metabolism and substrate-level energy formation during growth at alkaline pH in the saccharolytic fermenters Spirochaeta alkalica, Amphibacillus tropicus, and the Halanaerobiales representative Halonatronum saccharophilum heavily depend on transmembrane H+ and Na+ gradients.
The bacterial Na+-selective channels NavBac belong to the voltage-gated ion channel (VIC) superfamily. The prototype protein of NavBac is the NaChBac channel discovered in the alkaliphilic Bacillus halodurans C-125 (Ren et al., 2001). The sodium-selective NaChBac is a channel protein with a high specificity for Na+, although its activation and inactivation kinetics are slower than that of NaV’s (Ren et al., 2001; for a review, see Charalambous and Wallace, 2011). NaChBac homologues are encountered in diverse marine bacteria and/or bacteria living in alkaline niches (Koishi et al., 2004; Ito et al., 2004). A recent BLASTP search reported by Irie et al. (2010) yielded 26 homologs of NaChBAC, including those from Oceanobacillus iheyensis, the anaerobic Alkalilimnicola ehrlichii, and the haloalkaliphilic Halorhodospira halophila. However, our updated BLASTP evaluation of NaChBAC homologs indicated their presence in Halomonas spp. (37 % amino acid identity), Ectothiorhodospira spp. (39 %), and Thioalkalivibrio spp. (40 %).
A Na+-channel potentiated at elevated pH has been described as playing roles in pH homeostasis and pH-sensing in the facultatively alkaliphilic Bacillus pseudofirmus OF4. Deletion of the gene (ncbA) coding for the voltage-gated Na+ channel NaVBP impaired growth at alkaline pH, indicating its significant contribution to the cytoplasmic ion homeostasis. Moreover, NaVBP plays an active role in the motility and chemotaxis only during growth at pH 10 (Ito et al., 2004; Fujinami et al., 2007).
3.5.5 Bicarbonate Transport in Lithotrophic Haloalkaliphiles and Natronophiles
Uptake of carbon dioxide (CO2) and/or bicarbonate (HCO3 −) by obligately autotrophic organisms is essential for providing the inorganic carbon needed in assimilation. Soda lakes are virtually unlimited sources for carbonate (CO3 2−), but, since it is not transported, the main source of inorganic carbon in soda lakes seems to be HCO3 − which becomes less accessible at very high pH (>10). Therefore, the soda lake autotrophs have to rely on active transport of bicarbonate ion. However, hypersaline soda lakes host a high abundance and diversity of photo- and chemolithoautotrophic SOB from the family Ectothiorhodospiraceae (Thioalkalivibrio spp., Halorhodospira spp.) (Sorokin and Kuenen, 2005; Kovaleva et al., 2011). Low to moderately saline soda lakes, instead, have an impressive seasonal biomass production by oxygenic phototrophic cyanobacteria.
In cyanobacteria, the CO2-concentrating mechanism (CCM) involving the uptake of CO2 and HCO3 − is driven by light energy (Kaplan et al., 2008). To date, at least two CO2 uptake systems bound to thylakoids, and three HCO3 − transporters localized in plasma membrane covering the entire range of inorganic carbon availability are known in cyanobacteria. By this CCM, inorganic carbon is captured in the cytoplasm and delivered as HCO3 − through thylakoid membranes to the RuBisCO active site (Price, 2011). Two membrane transporters for bicarbonate are sodium-dependent: the inducible, high-affinity HCO3 −/SO4 2− transporter, SbtA (Shibata et al., 2002), and the low-affinity, high-flux HCO3 − transporter, BicA, identified in marine cyanobacteria (Price, 2011). The HCO3 −/Na+ symporter SbtA is a multi-pass membrane protein that is induced at low dissolved CO2. SbtA, identified in Synechocystis PCC 6803, has a high affinity for bicarbonate (K0.5 < 5 μM HCO3 −), and its function is dependent on the smf generated by the activity of the Na+/H+ antiporter (Shibata et al., 2002). SbtA is particularly interesting as it works at low bicarbonate and high sodium concentration. Genes related to the sbtA gene from Synechocystis PCC 6803 are present in the genome sequences of the natronophilic SOB Thioalkalivibrio K90mix, Thioalkalivibrio thiocyanoxidans ARh4, Thioalkalimicrobium aerophilum AL3, and Thioalkalimicrobium cyclicum ALM1, but not in Thioalkalivibrio sulfidophilus HL-EbGr7. In all natronophilic chemolithoautotrophic SOB, however, a gene for low-affinity high-flux HCO3 − transporter, SulP, is present (Muyzer et al., 2011a,b; personal BLASTP search). A comparative analysis of gene products also revealed homologous sequences of sbtA in haloalkalitolerant cyanobacteria (Synechococcus spp., Anabaena spp., and Cyanothece spp.) (Muyzer et al., 2011b). Apparently, SbtA bicarbonate transporters are not restricted to autotrophic bacteria. From a survey of the protein sequence database, homologous genes for SbtA are identified in alkaliphilic Bacillus and Bacillus-related species (B. pseudofirmus OF4 – acc. nr. YP_003425900.1, B. halodurans C-125 – acc. nr. NP_244736.1, and Caldalkalibacillus thermarum – acc. nr. ZP_08531624.1). In a complementation experiment, a gene fragment that encodes a homolog to the SulP transporter family member, BicA, from the haloalkaliphilic, strictly heterotrophic Alkalimonas amylolytica N10 allowed an alkali-sensitive E. coli mutant to grow at high pH (Wei et al., 2007). Interestingly, the closest homolog of the BicA protein from Alkalimonas amylolytica (acc. nr. ABG37985.1) is an uncharacterized anion transporter from haloalkaliphilic methanotroph Methylomicrobium alcaliphilum (acc. nr. YP_004916439.1) with 61 % amino acid identity. It is tempting to speculate that the active uptake of HCO3 − in heterotrophic haloalkaliphilic bacteria could be an indicative of inorganic C fixation by anaplerotic pathways that alternatively replenish the Krebs cycle during starvation periods (Ashworth and Kornberg, 1966).
3.6 Adaptation to High-Light and Oxidative Stress by Specific Membrane Pigments
Apart from their specific roles in photosynthesis (i.e., bacteriochlorophylls and carotenoids) and light-driven transport processes (rhodopsins), pigments produced by several halophilic and haloalkaliphilic prokaryotes may intervene in high-light/UV protection and prevent oxidative-stress damages as antioxidants. Such biological functions may be attributed to carotenoids from archaeal membranes and halophilic bacteria (e.g., bacterioruberin from Halobacterium salinarum, salinixanthin of Salinibacter ruber) (Oren, 2002). Natronochrome and chloronatronochrome, two yellow pigments composed of a C15-polyene chain attached to a phenyl group, were chemically characterized from the membranes of non-photosynthetic natronophile Thioalkalivibrio versutus ALJ15 (Takaichi et al., 2004). Yellow-membrane-bound pigments have been extracted from the chemolithoautotrophic SOB Thiohalospira spp. (Sorokin et al., 2008a). No functional characterization has been done on these yellow polyene pigments, but their possible involvement in light- and oxidative-stress protection was suggested, as well as a function as an additional membrane barrier to H+ and Na+ leakage in a similar manner as squalene (Haines, 2001; Hauss et al., 2002). Xanthomonadin, a structurally related yellow pigment isolated from the outer membrane of the phytopathogenic gammaproteobacterium Xanthomonas spp., was demonstrated to improve the survival rate of cells under high light conditions and to protect lipids against peroxidation in vitro (Poplawsky et al., 2000; Rajagopal et al., 1997).
4 Conclusions
Most moderate and extreme haloalkaliphilic and natronophilic prokaryotes are mainly distributed within the groups of aerobic Gram-negative bacteria, anaerobic, low G+C, Gram-positive bacteria, and haloarchaea. Haloalkaliphiles and natronophiles take up and/or produce compatible solutes in a similar manner as halotolerant and halophilic prokaryotes. The unique feature, however, is met in natronophiles that demand high soda medium and must apparently face a lower osmotic pressure at the same sodium concentration than the true halophiles. This situation has presumably important consequences to the bioenergetics of natronophiles. One such consequence is the energy saved out of synthesizing and/or importing high amounts of organic compatible solutes, energy that could be further invested in a multitude of low-cost/high-benefit strategies to overcome the low pmf. H+-dependent ATP synthesis is universally found among aerobic alkaliphilic, haloalkaliphilic, and natronophilic bacteria. High production of low-potential cytochromes of the c-type, point mutations in the H+-translocating subunits of F-ATP synthase are complemented by a bifurcated electron transfer chain. Such adaptations build a highly efficient OXPHOS machinery under the conditions of an unfavorable electrochemical proton gradient. Last but not least, a broad variety of Na+- and pH-regulated transporters are interconnected and differently expressed to maximize the exploitation of the favorable electrochemical sodium gradient (smf). Smf is the driving force for flagellar movement and, indirectly, for chemotaxis, as well as for acidification of cytoplasm, a critical process to resist the alkaline surrounding. Other adaptations that have not been discussed in this chapter are met in the structures and kinetics of intracellular and extracellular enzymes of haloalkaliphiles. Overall, the physiology and genetic evidences indicate a conserved, general pattern of salt and alkaline adaptation over a broad range of alkaline pH (9–11) and salinity (up to saturation) in haloalkaliphiles/natronophiles. Genomes of salt- and alkaline-tolerant strains of Bacillus spp. and E. coli, the most common experimental models, apparently encode the basic survival kits to respond the moderate osmotic and alkali shocks. However, those kits are not sufficient to cross to the regions where true haloalkaliphiles are flourishing. Further in-depth analyses of alkaliphilic and haloalkaliphilic genomes completed by integration with other “omics” and physiological evidences would definitively enlighten us on the bases as well as on the peculiarities of haloalkaline adaptation in prokaryotes.
References
Adams RL, Russell NJ (1992) Interactive effects of salt concentration and temperature on growth and lipid composition in the moderately halophilic bacterium Vibrio costicola. Can J Microbiol 38:823–827
Adams RL, Kogut M, Russell NJ (1990) The effect of salinity on growth and lipid composition of a moderately halophilic Gram-negative bacterium HX. Biochem Cell Biol 68:249–254
Aguilar PS, de Mendoza D (2006) Control of fatty acid desaturation, a mechanism conserved from bacteria to humans. Mol Microbiol 62:1507–1514
Alarico S, Empadinhas N, Simões C, Silva Z, Henne A, Mingote A, Santos H, da Costa MS (2005) Distribution of genes for synthesis of trehalose and mannosylglycerate in Thermus spp. and direct correlation of these genes with halotolerance. Appl Environ Microbiol 71:2460–2466
Alazard D, Badillo C, Fardeau ML, Cayol JL, Thomas P, Roldan T, Tholozan JL, Ollivier B (2007) Tindallia texcoconensis sp. nov., a new haloalkaliphilic bacterium isolated from lake Texcoco, Mexico. Extremophiles 11:33–39
Anderson I, Scheuner C, Göker M, Mavromatis K, Hooper SD, Porat I, Klenk HP, Ivanova N, Kyrpides N (2011) Novel insights into the diversity of catabolic metabolism from ten haloarchaeal genomes. PLoS One 6:e20237
Anil Kumar P, Srinivas TN, Madhu S, Manorama R, Shivaji S (2010a) Indibacter alkaliphilus gen. nov., sp. nov., an alkaliphilic bacterium isolated from a haloalkaline lake. Int J Syst Evol Microbiol 60:721–726
Anil Kumar P, Srinivas TN, Pavan Kumar P, Madhu S, Shivaji S (2010b) Nitritalea halalkaliphila gen. nov., sp. nov., an alkaliphilic bacterium of the family ‘Cyclobacteriaceae’, phylum Bacteroidetes. Int J Syst Evol Microbiol 60:2320–2325
Antony CP, Doronina NV, Boden R, Trotsenko YA, Shouche YS, Murrell JC (2012) Methylophaga lonarensis sp. nov., a moderately haloalkaliphilic methylotroph isolated from the soda lake sediments of a meteorite impact crater. Int J Syst Evol Microbiol 62:1613–1618
Aono R (1990) The poly-α- and -β-1,4-glucuronic acid moiety of teichuronopeptide from the cell wall of the alkalophilic Bacillus strain C-125. Biochem J 270:363–367
Aono R, Ohtani M (1990) Loss of alkalophily in cell-wall-component-defective mutants derived from alkalophilic Bacillus C-125. Isolation and partial characterization of the mutants. Biochem J 266:933–936
Arahal DR, García MT, Vargas C, Cánovas D, Nieto JJ, Ventosa A (2001) Chromohalobacter salexigens sp. nov., a moderately halophilic species that includes Halomonas elongata DSM 3043 and ATCC33174. Int J Syst Evol Microbiol 51:1457–1462
Asao M, Jung DO, Achenbach LA, Madigan MT (2006) Heliorestis convoluta sp. nov., a coiled, alkaliphilic heliobacterium from the Wadi El Natroun, Egypt. Extremophiles 10:403–410
Ashworth JM, Kornberg HL (1966) The anaplerotic fixation of carbon dioxide by Escherichia coli. Proc R Soc Lond B Biol Sci 165:179–188
Atsumi T, McCarter L, Imae Y (1992) Polar and lateral flagellar motors of marine Vibrio are driven by different ion-motive forces. Nature 355:182–184
Avonce N, Mendoza-Vargas A, Morett E, Iturriaga G (2006) Insights on the evolution of trehalose biosynthesis. BMC Evol Biol 6:109
Backiel J, Juárez O, Zagorevski DV, Wang Z, Nilges MJ, Barquera B (2008) Covalent binding of flavins to RnfG and RnfD in the Rnf complex from Vibrio cholerae. Biochemistry 47:11273–11284
Banciu HL (2004) Physiology of alkaliphilic sulfur-oxidizing bacteria from soda lakes. PhD thesis, Delft University, Optima BV, Rotterdam
Banciu H, Sorokin DY, Kleerebezem R, Galinski EA, Muyzer G, Kuenen JG (2004a) Growth kinetics of haloalkaliphilic sulfur-oxidizing bacterium Thioalkalivibrio versutus strain ALJ15 in continuous culture. Extremophiles 8:185–192
Banciu H, Sorokin DY, Galinski EA, Muyzer G, Kleerebezem R, Kuenen JG (2004b) Thioalkalivibrio halophilus sp. nov., a novel obligately chemolithoautotrophic, facultatively alkaliphilic, and extremely salt-tolerant, sulfur-oxidizing bacterium from a hypersaline alkaline lake. Extremophiles 8:325–334
Banciu H, Sorokin DY, Rijpstra WIC, Sinninghe Damsté JS, Galinski EA, Takaichi S, Muyzer G, Kuenen JG (2005) Fatty acid, compatible solute and pigment composition of obligately chemolithoautotrophic alkaliphilic sulfur-oxidizing bacteria from soda lakes. FEMS Microbiol Lett 243:181–187
Banciu HL, Sorokin DY, Tourova TP, Galinski EA, Muntyan MS, Kuenen JG, Muyzer G (2008) Influence of salts and pH on growth and activity of a novel facultatively alkaliphilic, extremely salt-tolerant, obligately chemolithoautotrophic sulfur-oxidizing Gammaproteobacterium Thioalkalibacter halophilus gen. nov., sp. nov. from South-Western Siberian soda lakes. Extremophiles 12:391–404
Blanco-Rivero A, Legans F, Fernandez-Valiente E, Calle P, Fernandez-Pinas F (2005) mrpA, a gene with roles in resistance to Na+ and adaptation to alkaline pH in cyanobacterium Anabaena sp. PCC7120. Microbiology (UK) 151:1671–1682
Boltyanskaya YV, Kevbrin VV, Lysenko AM, Kolganova TV, Turova TP, Osipov GA, Zhilina TN (2007) Halomonas mongoliensis sp. nov. and Halomonas kenyensis sp. nov., new haloalkaliphilic denitrifiers capable of N2O reduction, isolated from soda lakes. Microbiology (Moscow) 76:739–747
Borges N, Ramos A, Raven ND, Sharp RJ, Santos H (2002) Comparative study of the thermostabilizing properties of mannosylglycerate and other compatible solutes on model enzymes. Extremophiles 6:209–216
Borsodi AK, Márialigeti K, Szabó G, Palatinszky M, Pollák B, Kéki Z, Kovács AL, Schumann P, Tóth EM (2008) Bacillus aurantiacus sp. nov., an alkaliphilic and moderately halophilic bacterium isolated from Hungarian soda lakes. Int J Syst Evol Microbiol 58:845–851
Borsodi AK, Pollák B, Kéki Z, Rusznyák A, Kovács AL, Spröer C, Schumann P, Márialigeti K, Tóth EM (2011) Bacillus alkalisediminis sp. nov., an alkaliphilic and moderately halophilic bacterium isolated from sediment of extremely shallow soda ponds. Int J Syst Evol Microbiol 61:1880–1886
Bozek MA (1989) Orientation of zooplankton to the oxycline in Big Soda Lake, Nevada. West North Am Nat 49:535–539
Brändén M, Sandén T, Brzezinski P, Widengren J (2006) Localized proton microcircuits at the biological membrane-water interface. Proc Natl Acad Sci U S A 103:19766–19770
Bryantseva IA, Gorlenko VM, Kompantseva EL, Imhoff JF, Süling J, Mityushina L (1999a) Thiorhodospira sibirica gen. nov., sp. nov., a new alkaliphilic purple sulfur bacterium from a Siberian soda lake. Int J Syst Bacteriol 49:697–703
Bryantseva IA, Gorlenko VM, Kompantseva EL, Achenbach LA, Madigan MT (1999b) Heliorestis daurensis, gen. nov. sp. nov., an alkaliphilic rod-to-coiled-shaped phototrophic heliobacterium from a Siberian soda lake. Arch Microbiol 172:167–174
Bryantseva IA, Gorlenko VM, Kompantseva EL, Imhoff JF (2000) Thioalkalicoccus limnaeus gen. nov., sp. nov., a new alkaliphilic purple sulfur bacterium with bacteriochlorophyll b. Int J Syst Evol Microbiol 50:2157–2163
Bryantseva IA, Turova TP, Kovaleva OL, Kostrikina NA, Gorlenko VM (2010) Ectothiorhodospira magna sp. nov., a new large alkaliphilic purple sulfur bacterium. Microbiology (Moscow) 79:780–790
Burns BP, Gudhka RK, Neilan BA (2012) Genome sequence of the halophilic archaeon Halococcus hamelinensis. J Bacteriol 194:2100–2101
Bursy J, Pierik AJ, Pica N, Bremer E (2007) Osmotically induced synthesis of the compatible solute hydroxyectoine is mediated by an evolutionarily conserved ectoine hydroxylase. J Biol Chem 282:31147–31155
Bursy J, Kuhlmann AU, Pittelkow M, Hartmann H, Jebbar M, Pierik AJ, Bremer E (2008) Synthesis and uptake of the compatible solutes ectoine and 5-hydroxyectoine by Streptomyces coelicolor A3(2) in response to salt and heat stresses. Appl Environ Microbiol 74:7286–72896
Cao SJ, Qu JH, Yang JS, Sun Q, Yuan HL (2008) Halolactibacillus alkaliphilus sp. nov., a moderately alkaliphilic and halophilic bacterium isolated from a soda lake in Inner Mongolia, China, and emended description of the genus Halolactibacillus. Int J Syst Evol Microbiol 58:2169–2173
Cao SJ, Qu JH, Yuan HL, Li BZ (2010) Salsuginibacillus halophilus sp. nov., a halophilic bacterium isolated from a soda lake. Int J Syst Evol Microbiol 60:1339–1343
Carrasco IJ, Márquez MC, Xue Y, Ma Y, Cowan DA, Jones BE, Grant WD, Ventosa A (2007) Salsuginibacillus kocurii gen. nov., sp. nov., a moderately halophilic bacterium from soda-lake sediment. Int J Syst Evol Microbiol 57:2381–2386
Charalambous K, Wallace BA (2011) NaChBac, the long lost sodium channel ancestor. Biochemistry 50:6742–6752
Chen YG, Cui XL, Wang YX, Zhang YQ, Tang SK, Li WJ, Liu ZX, Wen ML, Peng Q (2009) Virgibacillus sediminis sp. nov., a moderately halophilic bacterium isolated from a salt lake in China. Int J Syst Evol Microbiol 59:2058–2063
Choi S, Swanson JM (1995) Interaction of cytochrome c with cardiolipin: an infrared spectroscopic study. Biophys Chem 54:271–278
Clejan S, Krulwich TA (1988) Permeability studies of lipid vesicles from alkalophilic Bacillus firmus showing opposing effects of membrane isoprenoid and diacylglycerol fractions and suggesting a possible basis for obligate alkalophily. Biochim Biophys Acta 946:40–48
Clejan S, Krulwich TA, Mondrus KR, Seto-Young D (1986) Membrane lipid composition of obligately and facultatively alkalophilic strains of Bacillus spp. J Bacteriol 168:334–340
Corcelli A (2009) The cardiolipin analogues of Archaea. Biochim Biophys Acta 1788:2101–2106
Dana GL, Lenz PH (1986) Effects of increasing salinity on an Artemia population from Mono Lake, California. Oecologia 68:428–436
Detkova EN, Boltyanskaya YV (2007) Osmoadaptation of haloalkaliphilic bacteria, role of osmoregulators and their possible practical application. Microbiology (Moscow) 76:511–522
Domene C, Furini S (2012) Molecular dynamics simulations of the TrkH membrane protein. Biochemistry 51:1559–1565
Doronina NV, Darmaeva TD, Trotsenko YA (2003a) Methylophaga alcalica sp. nov., a new alkaliphilic and moderately halophilic, obligately methylotrophic bacterium from an East Mongolian saline soda lake. Int J Syst Evol Microbiol 53:223–229
Doronina NV, Darmaeva TD, Trotsenko YA (2003b) Methylophaga natronica sp.nov., a new alkaliphilic and moderately halophilic, restricted-facultatively methylotrophic bacterium from Soda Lake of the Southern Transbaikal Region. Syst Appl Microbiol 26:382–389
Dowling NJE, Widdel F, White DC (1986) Phospholipid ester-linked fatty acid biomarkers of acetate-oxidizing sulphate-reducers and other sulphide-forming bacteria. Microbiology (UK) 132:1815–1825
Duckworth AW, Grant WD, Jones BE, van Steenbergen R (1996) Phylogenetic diversity of soda lakes alkaliphiles. FEMS Microbiol Ecol 19:181–191
Duckworth AW, Grant WD, Jones BE, Meijer D, Márquez MC, Ventosa A (2000) Halomonas magadii sp. nov., a new member of the genus Halomonas, isolated from a soda lake of the East African Rift Valley. Extremophiles 4:53–60
Dunkley EA Jr, Guffanti AA, Clejan S, Krulwich TA (1991) Facultative alkaliphiles lack fatty acid desaturase activity and lose the ability to grow at near-neutral pH when supplemented with an unsaturated fatty acid. J Bacteriol 173:1331–1334
Dzioba J, Ostroumov E, Winogrodzki A, Dibrov P (2002) Cloning, functional expression in Escherichia coli and primary characterization of a new Na+/H+ antiporter, NhaD, of Vibrio cholerae. Mol Cell Biochem 229:119–124
Elbein AD, Pan YT, Pastuszak I, Carroll D (2003) New insights on trehalose, a multifunctional molecule. Glycobiology 13:17R–27R
Epstein W (2003) The roles and regulation of potassium in bacteria. Prog Nucleic Acid Res Mol Biol 75:293–320
Feldman RI, Boyer PD (1985) The role of tightly bound ADP on chloroplast ATPase. J Biol Chem 260:13088–13094
Ferguson SA, Keis S, Cook GM (2006) Biochemical and molecular characterization of a Na+-translocating F1F0-ATPase from the thermoalkaliphilic bacterium Clostridium paradoxum. J Bacteriol 188:5045–5054
Fujinami S, Sato T, Trimmer JS, Spiller BW, Clapham DE, Krulwich TA, Kawagishi I, Ito M (2007) The voltage-gated Na+ channel NaVBP co-localizes with methyl-accepting chemotaxis protein at cell poles of alkaliphilic Bacillus pseudofirmus OF4. Microbiology (UK) 153:4027–4038
Fujinami S, Terahara N, Krulwich TA, Ito M (2009) Motility and chemotaxis in alkaliphilic Bacillus species. Future Microbiol 4:1137–1149
Fujisawa M, Fackelmayer OJ, Liu J, Krulwich TA, Hicks DB (2010) The ATP synthase a-subunit of extreme alkaliphiles is a distinct variant, mutations in the critical alkaliphile-specific residue Lys-180 and other residues that support alkaliphile oxidative phosphorylation. J Biol Chem 285:32105–32115
Fukaya F, Promden W, Hibino T, Tanaka Y, Nakamura T, Takabe T (2009) An Mrp-like cluster in the halotolerant cyanobacterium Aphanothece halophytica functions as a Na+/H+ antiporter. Appl Environ Microbiol 75:6626–6629
Galinski EA, Herzog RM (1990) The role of trehalose as a substitute for nitrogen-containing compatible solutes (Ectothiorhodospira halochloris). Arch Microbiol 153:607–613
Galinski EA, Trüper HG (1982) Betaine, a compatible solute in the extremely halophilic phototrophic bacterium Ectothiorhodospira halochloris. FEMS Microbiol Lett 13:357–360
Galinski EA, Pfeiffer HP, Trüper HG (1985) 1,4,5,6-Tetrahydro-2-methyl-4-pyrimidinecarboxylic acid – a novel cyclic amino acid from halophilic phototrophic bacteria of the genus Ectothiorhodospira. Eur J Biochem 149:135–139
García-Estepa R, Argandoña M, Reina-Bueno M, Capote N, Iglesias-Guerra F, Nieto JJ, Vargas C (2006) The ectD gene, which is involved in the synthesis of the compatible solute hydroxyectoine, is essential for thermoprotection of the halophilic bacterium Chromohalobacter salexigens. J Bacteriol 188:3774–3784
Geng W, Cao M, Song C, Xie H, Liu L, Yang C, Feng J, Zhang W, Jin Y, Du Y, Wang S (2011) Complete genome sequence of Bacillus amyloliquefaciens LL3, which exhibits glutamic acid-independent production of poly-γ-glutamic acid. J Bacteriol 193:3393–3394
Gerchman Y, Olami Y, Romon A, Taglicht D, Schuldiner S, Padan E (1993) Histidine-226 is part of the pH sensor of NhaA, a Na+, H+ antiporter in Escherichia coli. Proc Natl Acad Sci U S A 90:1212–1216
Gilmour R, Messner P, Guffanti AA, Kent R, Scheberl A, Kendrick N, Krulwich TA (2000) Two-dimensional gel electrophoresis analyses of pH-dependent protein expression in facultatively alkaliphilic Bacillus pseudofirmus OF4 lead to characterization of an S-layer protein with a role in alkaliphily. J Bacteriol 182:5969–5981
Giotis ES, McDowell DA, Blair IS, Wilkinson BJ (2007) Role of branched-chain fatty acids in pH stress tolerance in Listeria monocytogenes. Appl Environ Microbiol 73:997–1001
Goldfine H (2010) The appearance, disappearance and reappearance of plasmalogens in evolution. Prog Lipid Res 49:493–498
Goto T, Matsuno T, Hishinuma-Narisawa M, Yamazaki K, Matsuyama H, Inoue N, Yumoto I (2005) Cytochrome c and bioenergetic hypothetical model for alkaliphilic Bacillus spp. J Biosci Bioeng 100:365–379
Grammann K, Volke A, Kunte HJ (2002) New type of osmoregulated solute transporter identified in halophilic members of the bacteria domain, TRAP transporter TeaABC mediates uptake of ectoine and hydroxyectoine in Halomonas elongata DSM 2581T. J Bacteriol 184:3078–30785
Grant WD (2004) Life at low water activity. Philos Trans R Soc Lond B Biol Sci 359:1249–1267
Grant WD, Heaphy S (2010) Metagenomics and recovery of enzyme genes from alkaline saline environments. Environ Technol 31:1135–1143
Grant WD, Sorokin DY (2011) Distribution and diversity of soda lake alkaliphiles. In: Horikoshi K (ed) Extremophiles handbook. Part 2. Springer, Tokyo, pp 27–54
Grant WD, Tindall BJ (1986) Alkaline saline environment. In: Herbert RA, Codd GA (eds) Microbes in extreme environments. Academic, London, pp 25–54
Grant WD, Mwatha WE, Jones BE (1990) Alkaliphiles, ecology, diversity and applications. FEMS Microbiol Rev 75:255–270
Grant WD, Gemmell RT, McGenity TJ (1998) Halophiles. In: Horikoshi K, Grant WD (eds) Extremophiles–microbial life in extreme environments. Wiley, New York, pp 93–132
Guffanti AA, Krulwich TA (1992) Features of apparent nonchemiosmotic energization of oxidative phosphorylation by alkaliphilic Bacillus firmus OF4. J Biol Chem 267:9580–9588
Guo Y, Xue Y, Liu J, Wang Q, Ma Y (2009) Characterization and function analysis of a halo-alkaline-adaptable Trk K+ uptake system in Alkalimonas amylolytica strain N10. Sci China C Life Sci 52:949–957
Haines TH (2001) Do sterols reduce proton and sodium leaks through lipid bilayers? Prog Lipid Res 40:299–324
Hamaide F, Kushner DJ, Sprott GD (1983) Proton motive force and Na+/H+ antiport in a moderate halophile. J Bacteriol 156:537–544
Hamamoto T, Horikoshi K (1992) Alkaliphiles. In: Lederberg J (ed) Encyclopedia of microbiology. Academic, San Diego, pp 81–87
Hamamoto T, Hashimoto M, Hino M, Kitada M, Seto Y, Kudo T, Horikoshi K (1994) Characterization of a gene responsible for the Na+/H+ antiporter system of alkalophilic Bacillus species strain C-125. Mol Microbiol 14:939–946
Harishchandra RK, Wulff S, Lentzen G, Neuhaus T, Galla HJ (2010) The effect of compatible solute ectoines on the structural organization of lipid monolayer and bilayer membranes. Biophys Chem 150:37–46
Hart DJ, Vreeland RH (1988) Changes in the hydrophobic-hydrophilic cell surface character of Halomonas elongata in response to NaCl. J Bacteriol 170:132–135
Hauss T, Dante S, Dencher NA, Haines TH (2002) Squalane is in the midplane of the lipid bilayer, implications for its function as a proton permeability barrier. Biochim Biophys Acta 1556:149–154
Heberle J, Riesle J, Thiedemann G, Oesterhelt D, Dencher NA (1994) Proton migration along the membrane surface and retarded surface to bulk transfer. Nature 370:379–382
Helfert C, Gotsche S, Dahl MK (1995) Cleavage of trehalose-phosphate in Bacillus subtilis is catalysed by a phospho-α-(1-1)-glucosidase encoded by the treA gene. Mol Microbiol 16:111–120
Hezayen FF, Rehm BH, Tindall BJ, Steinbüchel A (2001) Transfer of Natrialba asiatica B1T to Natrialba taiwanensis sp. nov. and description of Natrialba aegyptiaca sp. nov., a novel extremely halophilic, aerobic, non-pigmented member of the Archaea from Egypt that produces extracellular poly(glutamic acid). Int J Syst Evol Microbiol 51:1133–1142
Hicks DB, Krulwich TA (1995) The respiratory chain of alkaliphilic bacteria. Biochim Biophys Acta 1229:303–314
Hicks DB, Liu J, Fujisawa M, Krulwich TA (2010) F1F0-ATP synthases of alkaliphilic bacteria, lessons from their adaptations. Biochim Biophys Acta 1797:1362–1377
Hoeft SE, Switzer Blum J, Stolz JF, Tabita FR, Witte B, King GM, Santini JM, Oremland RS (2007) Alkalilimnicola ehrlichii sp. nov., a novel, arsenite-oxidizing haloalkaliphilic gammaproteobacterium capable of chemoautotrophic or heterotrophic growth with nitrate or oxygen as the electron acceptor. Int J Syst Evol Microbiol 57:504–512
Hoover RB, Pikuta EV, Bej AK, Marsic D, Whitman WB, Tang J, Krader P (2003) Spirochaeta americana sp. nov., a new haloalkaliphilic, obligately anaerobic spirochaete isolated from soda Mono Lake in California. Int J Syst Evol Microbiol 53:815–821
Horikoshi K (1999) Alkaliphiles. Haward Academic, Amsterdam
Imhoff JF (1993) Osmotic adaptation in halophilic and halotolerant microorganisms. In: Vreeland RH, Hochstein LI (eds) The biology of halophilic bacteria. CRC Press, Boca Raton, pp 211–253
Imhoff JF, Rodriguez-Valera F (1984) Betaine is the main compatible solute of halophilic eubacteria. J Bacteriol 160:478–479
Imhoff JF, Thiemann B (1991) Influence of salt concentration and temperature on the fatty acid compositions of Ectothiorhodospira and other halophilic phototrophic purple bacteria. Arch Microbiol 156:370–375
Imhoff JF, Sahl HG, Soliman GSH, Trüper HG (1979) The Wadi Natrun, chemical composition and microbial mass developments in alkaline brines of eutrophic desert lakes. Geomicrobiol J 1:219–234
Irie K, Kitagawa K, Nagura H, Imai T, Shimomura T, Fujiyoshi Y (2010) Comparative study of the gating motif and C-type inactivation in prokaryotic voltage-gated sodium channels. J Biol Chem 285:3685–3694
Ito M, Guffanti AA, Zemsky J, Ivey DM, Krulwich TA (1997) Role of the nhaC-encoded Na+/H+ antiporter of alkaliphilic Bacillus firmus OF4. J Bacteriol 179:3851–3857
Ito M, Guffanti AA, Krulwich TA (2001) Mrp-dependent Na+/H+ antiporters of Bacillus exhibit characteristics that are unanticipated for completely secondary active transporters. FEBS Lett 496:117–120
Ito M, Xu H, Guffanti AA, Wei Y, Zvi L, Clapham DE, Krulwich TA (2004) The voltage-gated Na+ channel NaVBP has a role in motility, chemotaxis, and pH homeostasis of an alkaliphilic Bacillus. Proc Natl Acad Sci U S A 101:10566–105671
Janto B, Ahmed A, Ito M, Liu J, Hicks DB, Pagni S, Fackelmayer OJ, Smith TA, Earl J, Elbourne LD, Hassan K, Paulsen IT, Kolstø AB, Tourasse NJ, Ehrlich GD, Boissy R, Ivey DM, Li G, Xue Y, Ma Y, Hu FZ, Krulwich TA (2011) Genome of alkaliphilic Bacillus pseudofirmus OF4 reveals adaptations that support the ability to grow in an external pH range from 7.5 to 11.4. Environ Microbiol 13:3289–3309
Jones BE, Grant WD, Duckworth AW, Owenson GG (1998) Microbial diversity of soda lakes. Extremophiles 2:191–200
Jung H (2001) Towards the molecular mechanism of Na+/solute symport in prokaryotes. Biochim Biophys Acta 1505:131–143
Jung H, Hilger D, Raba M (2012) The Na+/l-proline transporter PutP. Front Biosci 17:745–759
Kaieda N, Wakagi T, Koyama N (1998) Presence of Na+-stimulated V-type ATPase in the membrane of a facultatively anaerobic and halophilic alkaliphile. FEMS Microbiol Lett 167:57–61
Kaluzhnaya MG, Khmelenina V, Eshinimaev B, Suzina N, Nikitin D, Solonin A, Lin JL, McDonald I, Murrell C, Trotsenko Y (2001) Taxonomic characterization of new alkaliphilic and alkalitolerant methanotrophs from soda lakes of the southeastern Transbaikal region and description of Methylomicrobium buryatense sp. nov. Syst Appl Microbiol 24:166–176
Kaluzhnaya MG, Khmelenina V, Eshinimaev B, Sorokin D, Fuse H, Lidstrom M, Trotsenko Y (2008) Classification of halo(alkali)philic and halo(alkali)tolerant methanotrophs provisionally assigned to the genera Methylomicrobium and Methylobacter and emended description of the genus Methylomicrobium. Int J Syst Evol Microbiol 58:591–596
Kandler O, König H, Wiegel J, Claus D (1983) Occurrence of poly-γ-d-glutamic acid and poly-α-l-glutamine in the genera Xanthobacter, Flexithrix, Sporosarcina and Planococcus. Syst Appl Microbiol 4:34–41
Kaneda T (1977) Fatty acids of the genus Bacillus, an example of branched chain preference. Bacteriol Rev 41:391–418
Kaneda T (1991) Iso- and anteiso-fatty acids in bacteria, biosynthesis, function, and taxonomic significance. Microbiol Rev 55:288–302
Kaplan A, Hagemann M, Bauwe H, Kahlon S, Ogawa T (2008) Carbon acquisition by cyanobacteria, mechanisms, comparative genomics, and evolution. In: Herrero A, Flores E (eds) The cyanobacteria, molecular biology, genomics and evolution. Horizon Scientific Press, Norwich, pp 305–334
Kato-Yamada Y, Bald D, Koike M, Motohashi K, Hisabori T, Yoshida M (1999) ε subunit, an endogenous inhibitor of bacterial F1-ATPase, also inhibits F0F1-ATPase. J Biol Chem 274:33991–33994
Kaufman AE, Goldfine H, Narayan O, Gruner SM (1990) Physical studies on the membranes and lipids of plasmalogen deficient Megasphaera elsdenii. Chem Phys Lipids 55:41–48
Kempe S, Degens ET (1985) An early soda ocean? Chem Geol 53:95–108
Kempe S, Kazmierczak J (1997) A terrestrial model for an alkaline Martian hydrosphere. Planet Space Sci 45:1493–1495
Khmelenina VN, Kalyuzhnaya MG, Sakharovsky VG, Suzina NE, Trotsenko YA, Gottschalk G (1999) Osmoadaptation in halophilic and alkaliphilic methanotrophs. Arch Microbiol 172:321–329
Koishi R, Xu H, Ren D, Navarro B, Spiller BW, Shi Q, Clapham DE (2004) A superfamily of voltage-gated sodium channels in bacteria. J Biol Chem 279:9532–9538
Konno H, Isu A, Kim Y, Murakami-Fuse T, Sugano Y, Hisabori T (2011) Characterization of the relationship between ADP- and ε-induced inhibition in cyanobacterial F1-ATPase. J Biol Chem 286:13423–13429
Kovaleva OL, Tourova TP, Muyzer G, Kolganova TV, Sorokin DY (2011) Diversity of RuBisCO and ATP citrate lyase genes in soda lake sediments. FEMS Microbiol Ecol 75:37–47
Krulwich TA (2006) Alkaliphilic prokaryotes. In: Dworkin M, Falkow S, Rosenberg E, Scheifer K-H, Stackebrandt E (eds) The prokaryotes. A handbook on the biology of bacteria, ecophysiology, isolation, identification, applications, vol 2, 3rd edn. Springer, New York, pp 283–308
Krulwich TA, Federbush JG, Guffanti AA (1985) Presence of a nonmetabolizable solute that is translocated with Na+ enhances Na+-dependent pH homeostasis in an alkalophilic Bacillus. J Biol Chem 260:4055–4058
Krulwich TA, Ito M, Gilmour R, Hicks DB, Guffanti AA (1998) Energetics of alkaliphilic Bacillus species, physiology and molecules. Adv Microb Physiol 40:401–438
Krulwich TA, Hicks DB, Swartz TH, Ito M (2007) Bioenergetic adaptations that support alkaliphily. In: Gerday C, Glansdorff N (eds) Physiology and biochemistry of extremophiles. ASM Press, Washington, DC, pp 311–329
Krulwich TA, Sachs G, Padan E (2011) Molecular aspects of bacterial pH sensing and homeostasis. Nat Rev Microbiol 9:330–343
Kuhlmann AU, Bremer E (2002) Osmotically regulated synthesis of the compatible solute ectoine in Bacillus pasteurii and related Bacillus spp. Appl Environ Microbiol 68:772–783
Kuhlmann AU, Hoffmann T, Bursy J, Jebbar M, Bremer E (2011) Ectoine and hydroxyectoine as protectants against osmotic and cold stress, uptake through the SigB-controlled betaine-choline-carnitine transporter-type carrier EctT from Virgibacillus pantothenticus. J Bacteriol 193:4699–4708
Kuroda T, Mizushima T, Tsuchiya T (2005) Physiological roles of three Na+/H+ antiporters in the halophilic bacterium Vibrio parahaemolyticus. Microbiol Immunol 49:711–719
Kurz M, Brünig AN, Galinski EA (2006) NhaD type sodium/proton-antiporter of Halomonas elongata, a salt stress response mechanism in marine habitats? Saline Syst 2:10
Kushner DJ, Kamekura M (1988) Physiology of halophilic eubacteria. In: Rodrıguez-Valera F (ed) Halophilic bacteria, vol 1. CRC Press, Boca Raton, pp 109–138
Lai S-J, Lai M-C (2011) Characterization and regulation of the osmolyte betaine synthesizing enzymes GSMT and SDMT from halophilic methanogen Methanohalophilus portucalensis. PLoS One 6:e25090
Lai M-C, Sowers KR, Robertson DE, Roberts MF, Gunsalus RP (1991) Distribution of compatible solutes in the halophilic methanogenic archaebacteria. J Bacteriol 173:5352–5358
Laloknam S, Tanaka K, Buaboocha T, Waditee R, Incharoensakdi A, Hibino T, Tanaka Y, Takabe T (2006) Halotolerant cyanobacterium Aphanothece halophytica contains a betaine transporter active at alkaline pH and high salinity. Appl Environ Microbiol 72:6018–6026
Lattanzio VM, Baronio M, Oren A, Russell NJ, Corcelli A (2009) Characterization of polar membrane lipids of the extremely halophilic bacterium Salinibacter ruber and possible role of cardiolipin. Biochim Biophys Acta 1791:25–31
Lee JS, Lim JM, Lee KC, Lee JC, Park YH, Kim CJ (2006) Virgibacillus koreensis sp. nov., a novel bacterium from a salt field, and transfer of Virgibacillus picturae to the genus Oceanobacillus as Oceanobacillus picturae comb. nov. with emended descriptions. Int J Syst Evol Microbiol 56:251–257
Li H, Liu YH, Luo N, Zhang XY, Luan TG, Hu JM, Wang ZY, Wu PC, Chen MJ, Lu JQ (2006) Biodegradation of benzene and its derivatives by a psychrotolerant and moderately haloalkaliphilic Planococcus sp. strain ZD22. Res Microbiol 157:629–636
Lim CH, Bot AG, de Jonge HR, Tilly BC (2007) Osmosignaling and volume regulation in intestinal epithelial cells. Methods Enzymol 428:325–342
Lippert K, Galinski EA (1992) Enzyme stabilization by ectoine-type compatible solutes, protection against heating, freezing and drying. Appl Microbiol Biotechnol 37:61–65
Liu J, Xue Y, Wang Q, Wei Y, Swartz TH, Hicks DB, Ito M, Ma Y, Krulwich TA (2005) The activity profile of the NhaD-type Na+(Li+)/H+ antiporter from the soda lake haloalkaliphile Alkalimonas amylolytica is adaptive for the extreme environment. J Bacteriol 187:7589–7595
Liu X, Gong X, Hicks DB, Krulwich TA, Yu L, Yu CA (2007) Interaction between cytochrome caa 3 and F1F0-ATP synthase of alkaliphilic Bacillus pseudofirmus OF4 is demonstrated by saturation transfer electron paramagnetic resonance and differential scanning calorimetry assays. Biochemistry 46:306–313
Liu J, Krulwich TA, Hicks DB (2008) Purification of two putative type II NADH dehydrogenases with different substrate specificities from alkaliphilic Bacillus pseudofirmus OF4. Biochim Biophys Acta 1777:453–461
Liu J, Fujisawa M, Hicks DB, Krulwich TA (2009) Characterization of the functionally critical AXAXAXA and PXXEXXP motifs of the ATP synthase c-subunit from an alkaliphilic Bacillus. J Biol Chem 284:8714–8725
Liu YP, Wang YX, Li YX, Feng FY, Liu HR, Wang J (2012) Mongoliicoccus roseus gen. nov., sp. nov., an alkaliphilic bacterium isolated from a haloalkaline lake. Int J Syst Evol Microbiol 62:2206–2212
Lo CC, Bonner CA, Xie G, D’Souza M, Jensen RA (2009) Cohesion group approach for evolutionary analysis of aspartokinase, an enzyme that feeds a branched network of many biochemical pathways. Microbiol Mol Biol Rev 73:594–651
Louis P, Galinski EA (1997) Characterization of genes for the biosynthesis of the compatible solute ectoine from Marinococcus halophilus and osmoregulated expression in Escherichia coli. Microbiology (UK) 143:1141–1149
Lu J, Nogi Y, Takami H (2001) Oceanobacillus iheyensis gen. nov., sp. nov., a deep-sea extremely halotolerant and alkaliphilic species isolated from a depth of 1050 m on the Iheya Ridge. FEMS Microbiol Lett 205:291–297
Lysak LV, Troshin DV, Chernov IY (1994) Bacterial communities of Solonchaks. Microbiology (Moscow) 63:721–729
Mackay MA, Norton RS, Borowitzka LJ (1984) Organic osmoregulatory solutes in cyanobacteria. J Gen Microbiol 130:2177–2191
MacLeod RA, Wisse GA, Stejskal FL (1988) Sensitivity of some marine bacteria, a moderate halophile, and Escherichia coli to uncouplers at alkaline pH. J Bacteriol 170:4330–4337
Månnistö MK, Schumann P, Rainey FA, Kämpfer P, Tsitko I, Tiirola MA, Salkinoja-Salonen MS (2000) Subtercola boreus gen. nov., sp. nov. and Subtercola frigoramans sp. nov., two new psychrophilic actinobacteria isolated from boreal groundwater. Int J Syst Evol Microbiol 50:1731–1739
Márquez MC, Carrasco IJ, de la Haba RR, Jones BE, Grant WD, Ventosa A (2011) Bacillus locisalis sp. nov., a new haloalkaliphilic species from hypersaline and alkaline lakes of China, Kenya and Tanzania. Syst Appl Microbiol 34:424–428
Maurer LM, Yohannes E, Bondurant SS, Radmacher M, Slonczewski JL (2005) pH regulates genes for flagellar motility, catabolism, and oxidative stress in Escherichia coli K-12. J Bacteriol 187:304–319
Mayberry WR, Lane JR (1993) Sequential alkaline saponification/acid hydrolysis/esterification, a one-tube method with enhanced recovery of both cyclopropane and hydroxylated fatty acids. J Microbiol Methods 18:21–32
Melo AM, Bandeiras TM, Teixeira M (2004) New insights into type II NAD(P)H, quinone oxidoreductases. Microbiol Mol Biol Rev 68:603–616
Mesbah NM, Wiegel J (2008) Life at extreme limits – the anaerobic halophilic alkalithermophiles. Ann N Y Acad Sci 1125:44–57
Mesbah NM, Wiegel J (2009) Natronovirga wadinatrunensis gen. nov., sp. nov. and Natranaerobius trueperi sp. nov., halophilic, alkalithermophilic micro-organisms from soda lakes of the Wadi An Natrun, Egypt. Int J Syst Evol Microbiol 59:2042–2048
Mesbah NM, Cook GM, Wiegel J (2009) The halophilic alkalithermophile Natranaerobius thermophilus adapts to multiple environmental extremes using a large repertoire of Na(K)/H antiporters. Mol Microbiol 74:270–281
Mesbah NM, Wiegel J (2011) The Na+-translocating F1F0-ATPase from the halophilic, alkalithermophile Natranaerobius thermophilus. Biochim Biophys Acta 1807:1133–1142
Mesbah NM, Wiegel J (2012) Life under multiple extreme conditions: diversity and physiology of the halophilic alkalithermophiles. Appl Environ Microbiol 78:4074–4082
Mesbah NM, Hedrick DB, Peacock AD, Rohde M, Wiegel J (2007) Natranaerobius thermophilus gen. nov., sp. nov., a halophilic, alkalithermophilic bacterium from soda lakes of the Wadi An Natrun, Egypt, and proposal of Natranaerobiaceae fam. nov. and Natranaerobiales ord. nov. Int J Syst Evol Microbiol 57:2507–2512
Minkov IB, Fitin AF, Vasilyeva EA, Vinogradov AD (1979) Mg2+-induced ADP-dependent inhibition of the ATPase activity of beef heart mitochondrial coupling factor F1. Biochem Biophys Res Commun 89:1300–1306
Monteoliva-Sanchez M, Ferrer MR, Ramos-Cormenzana A, Quesada E, Monteoliva M (1988) Cellular fatty acid composition of Deleya halophila – effect of growth temperature and salt concentration. J Gen Microbiol 134:199–203
Monteoliva-Sanchez M, Ramos-Cormenzana A, Russell NJ (1993) The effect of salinity and compatible solutes on the biosynthesis of cyclopropane fatty acids in Pseudomonas halosaccharolytica. J Gen Microbiol 139:1877–1884
Morino M, Natsui S, Swartz TH, Krulwich TA, Ito M (2008) Single gene deletions of mrpA to mrpG and mrpE point mutations affect activity of the Mrp Na+/H+ antiporter of alkaliphilic Bacillus and formation of hetero-oligomeric Mrp complexes. J Bacteriol 190:4162–4172
Morino M, Natsui S, Ono T, Swartz TH, Krulwich TA, Ito M (2010) Single site mutations in the hetero-oligomeric Mrp antiporter from alkaliphilic Bacillus pseudofirmus OF4 that affect Na+/H+ antiport activity, sodium exclusion, individual Mrp protein levels, or Mrp complex formation. J Biol Chem 285:30942–30950
Mormile MR, Romine MF, Garcia MT, Ventosa A, Bailey TJ, Peyton BM (1999) Halomonas campisalis sp. nov., a denitrifying, moderately haloalkaliphilic bacterium. Syst Appl Microbiol 22:551–558
Mühling M, Belay A, Whitton BA (2005) Variation in fatty acid composition of Arthrospira (Spirulina) strains. J Appl Phycol 17:137–146
Muntyan MS, Bloch DA (2008) Study of redox potential in cytochrome c covalently bound to terminal oxidase of alkaliphilic Bacillus pseudofirmus FTU. Biochemistry (Moscow) 73:107–111
Muyzer G, Sorokin DY, Mavromatis K, Lapidus A, Clum A, Ivanova N, Pati A, D’Haeseleer P, Woyke T, Kyrpides NC (2011a) Complete genome sequence of “Thioalkalivibrio sulfidophilus” HL-EbGr7. Stand Genomic Sci 4:23–35
Muyzer G, Sorokin DY, Mavromatis K, Lapidus A, Foster B, Sun H, Ivanova N, Pati A, D’haeseleer P, Woyke T, Kyrpides NC (2011b) Complete genome sequence of Thioalkalivibrio sp. K90mix. Stand Genomic Sci 5:341–355
Nielsen P, Fritze D, Priest FG (1995) Phenetic diversity of alkaliphilic Bacillus strains, proposal for nine new species. Microbiology 141:1745–1761
Niemetz R, Kärcher U, Kandler O, Tindall BJ, König H (1997) The cell wall polymer of the extremely halophilic archaeon Natronococcus occultus. Eur J Biochem 249:905–911
Ningthoujam DS, Kshetri P, Samasam S, Nimaichand S (2009) Screening, identification of best producers and optimization of extracellular proteases from moderately halophilic alkalithermotolerant indigenous actinomycetes. World Appl Sci J 7:907–916
Nowicka B, Kruk J (2010) Occurrence, biosynthesis and function of isoprenoid quinones. Biochim Biophys Acta 1797:1587–1605
Nozaki K, Kuroda T, Mizushima T, Tsuchiya T (1998) A new Na+/H+ antiporter, NhaD, of Vibrio parahaemolyticus. Biochim Biophys Acta 1369:213–220
Nyyssölä A, Leisola M (2001) Actinopolyspora halophila has two separate pathways for betaine synthesis. Arch Microbiol 176:294–300
Nyyssölä A, Reinikainen T, Leisola M (2001) Characterization of glycine sarcosine N-methyltransferase and sarcosine dimethylglycine N-methyltransferase. Appl Environ Microbiol 67:2044–2050
Oh S, Kogure K, Ohwada K, Simidu U (1991) Correlation between possession of a respiration-dependent Na+ pump and Na+ requirement for growth of marine bacteria. Appl Environ Microbiol 57:1844–1846
Oren A (1999) Bioenergetic aspects of halophilism. Microbiol Mol Biol Rev 63:334–348
Oren A (2002) Halophilic microorganisms and their environments. Kluwer Academic, Dordrecht
Oren A (2008) Microbial life at high salt concentrations: phylogenetic and metabolic diversity. Saline Syst 4:2
Oren A (2011) Thermodynamic limits to microbial life at high salt concentrations. Environ Microbiol 13:1908–1923
Oren A (2012) Taxonomy of the family Halobacteriaceae, a paradigm for changing concepts in prokaryote systematics. Int J Syst Evol Microbiol 62:263–271
Oren A, Heldal M, Norland S, Galinski EA (2002) Intracellular ion and organic solute concentrations of the extremely halophilic bacterium Salinibacter ruber. Extremophiles 6:491–498
Ostroumov E, Dzioba J, Loewen P, Dibrov P (2002) Asp344 and Thr345 are critical for cation exchange mediated by NhaD, Na+/H+ antiporter of Vibrio cholerae. Biochim Biophys Acta 1564:99–106
Padan E (2008) The enlightening encounter between structure and function in the NhaA Na+-H+ antiporter. Trends Biochem Sci 33:435–443
Padan E, Krulwich TA (2000) Sodium stress. In: Storz G, Hengge-Aronis R (eds) Bacterial stress responses. ASM Press, Washington, DC, pp 117–130
Padan E, Venturi M, Gerchman Y, Dover N (2001) Na+/H+ antiporters. Biochim Biophys Acta 1505:144–157
Padan E, Bibi E, Ito M, Krulwich TA (2005) Alkaline pH homeostasis in bacteria, new insights. Biochim Biophys Acta 1717:67–88
Peddie CJ, Cook GM, Morgan HW (1999) Sodium-dependent glutamate uptake by an alkaliphilic, thermophilic Bacillus strain, TA2.A1. J Bacteriol 181:3172–3177
Peddie CJ, Cook GM, Morgan HW (2000) Sucrose transport by the alkaliphilic, thermophilic Bacillus sp. strain TA2.A1 is dependent on a sodium gradient. Extremophiles 4:291–296
Pereira IA, Ramos AR, Grein F, Marques MC, da Silva SM, Venceslau SS (2011) A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea. Front Microbiol 2:69
Pikuta EV, Hoover RB, Bej AK, Marsic D, Detkova EN, Whitman WB, Krader P (2003) Tindallia californiensis sp. nov., a new anaerobic, haloalkaliphilic, spore-forming acetogen isolated from Mono Lake in California. Extremophiles 7:327–334
Pikuta EV, Hoover RB, Bej AK, Marsic D, Whitman WB, Krader P (2009) Spirochaeta dissipatitropha sp. nov., an alkaliphilic, obligately anaerobic bacterium, and emended description of the genus Spirochaeta Ehrenberg 1835. Int J Syst Evol Microbiol 59:1798–1804
Pitcher RS, Brittain T, Watmough NJ (2002) Cytochrome cbb(3) oxidase and bacterial microaerobic metabolism. Biochem Soc Trans 30:653–658
Pitriuk AV, Detkova EN, Pusheva MA (2004) Comparative study of the energy metabolism of anaerobic alkaliphiles from soda lakes. Microbiology (Moscow) 73:243–248
Poehls DJ, Smith GJ (2009) Encyclopedic dictionary of hydrogeology. Academic, San Diego
Poli A, Esposito E, Orlando P, Lama L, Giordano A, de Appolonia F, Nicolaus B, Gambacorta A (2007) Halomonas alkaliantarctica sp. nov., isolated from saline lake Cape Russell in Antarctica, an alkalophilic moderately halophilic, exopolysaccharide-producing bacterium. Syst Appl Microbiol 30:31–38
Poplawsky AR, Urban SC, Chun W (2000) Biological role of xanthomonadin pigments in Xanthomonas campestris pv. Campestris. Appl Environ Microbiol 66:5123–5127
Preiss L, Yildiz O, Hicks DB, Krulwich TA, Meier T (2010) A new type of proton coordination in an F1F0-ATP synthase rotor ring. PLoS Biol 8:e1000443
Price GD (2011) Inorganic carbon transporters of the cyanobacterial CO2 concentrating mechanism. Photosynth Res 109:47–57
Quillaguamán J, Hatti-Kaul R, Mattiasson B, Alvarez MT, Delgado O (2004) Halomonas boliviensis sp. nov., an alkalitolerant, moderate halophile isolated from soil around a Bolivian hypersaline lake. Int J Syst Evol Microbiol 54:721–725
Radchenko MV, Waditee R, Oshimi S, Fukuhara M, Takabe T, Nakamura T (2006) Cloning, functional expression and primary characterization of Vibrio parahaemolyticus K+/H+ antiporter genes in Escherichia coli. Mol Microbiol 59:651–663
Rajagopal L, Sundari CS, Balasubramanian D, Sonti RV (1997) The bacterial pigment xanthomonadin offers protection against photodamage. FEBS Lett 415:125–128
Rauhamäki V, Bloch DA, Verkhovsky MI, Wikström M (2009) Active site of cytochrome cbb3. J Biol Chem 284:11301–11308
Reina-Bueno M, Argandoña M, Salvador M, Rodríguez-Moya J, Iglesias-Guerra F, Csonka LN, Nieto JJ, Vargas C (2012) Role of trehalose in salinity and temperature tolerance in the model halophilic bacterium Chromohalobacter salexigens. PLoS One 7:e33587
Ren D, Navarro B, Xu H, Yue L, Shi Q, Clapham DE (2001) A prokaryotic voltage-gated sodium channel. Science 294:2372–2375
Reshetnikov AS, Khmelenina VN, Mustakhimov II, Trotsenko YA (2011) Genes and enzymes of ectoine biosynthesis in halotolerant methanotrophs. Methods Enzymol 495:15–30
Riesenfeld CS, Schloss PD, Handelsman J (2004) Metagenomics, genomic analysis of microbial communities. Annu Rev Genet 38:525–552
Rimmele M, Boos W (1994) Trehalose-6-phosphate hydrolase of Escherichia coli. J Bacteriol 176:5654–5664
Roberts MF (2000) Osmoadaptation and osmoregulation in archaea. Front Biosci 5:796–812
Robertson DE, Noll D, Roberts MF, Menaia JA, Boone DR (1990) Detection of the osmoregulator betaine in methanogens. Appl Environ Microbiol 56:563–565
Robinson NC (1993) Functional binding of cardiolipin to cytochrome c oxidase. J Bioenerg Biomembr 25:153–163
Romano I, Nicolaus B, Lama L, Trabasso D, Caracciolo G, Gambacorta A (2001) Accumulation of osmoprotectants and lipid pattern modulation in response to growth conditions by Halomonas pantelleriense. Syst Appl Microbiol 24:342–352
Romano I, Giordano A, Lama L, Nicolaus B, Gambacorta A (2005a) Halomonas campaniensis sp. nov., a haloalkaliphilic bacterium isolated from a mineral pool of Campania Region, Italy. Syst Appl Microbiol 28:610–618
Romano I, Lama L, Nicolaus B, Gambacorta A, Giordano A (2005b) Alkalibacillus filiformis sp. nov., isolated from a mineral pool in Campania, Italy. Int J Syst Evol Microbiol 55:2395–2399
Romano I, Lama L, Nicolaus B, Poli A, Gambacorta A, Giordano A (2006a) Halomonas alkaliphila sp. nov., a novel halotolerant alkaliphilic bacterium isolated from a salt pool in Campania (Italy). J Gen Appl Microbiol 52:339–348
Romano I, Lama L, Nicolaus B, Poli A, Gambacorta A, Giordano A (2006b) Oceanobacillus oncorhynchi subsp. incaldanensis subsp. nov., an alkalitolerant halophile isolated from an algal mat collected from a sulfurous spring in Campania (Italy), and emended description of Oceanobacillus oncorhynchi. Int J Syst Evol Microbiol 56:805–810
Russell NJ (1993) Lipids of halophilic and halotolerant microorganisms. In: Vreeland RH, Hochstein LI (eds) The biology of halophilic bacteria. CRC Press, Boca Raton, pp 163–210
Saier MH Jr, Yen MR, Noto K, Tamang DG, Elkan C (2009) The transporter classification database, recent advances. Nucleic Acids Res 37:D274–D278
Satoh M, Koyama N (2005) Cloning and sequencing of the genes for A and B subunits of the V-type Na+-ATPase of a facultatively anaerobic alkaliphile. Anaerobe 11:115–121
Schägger H (2002) Respiratory chain supercomplexes of mitochondria and bacteria. Biochim Biophys Acta 1555:154–159
Schoepp-Cothenet B, Lieutaud C, Baymann F, Verméglio A, Friedrich T, Kramer DM, Nitschke W (2009) Menaquinone as pool quinone in a purple bacterium. Proc Natl Acad Sci U S A 106:8549–8554
Schwibbert K, Marin-Sanguino A, Bagyan I, Heidrich G, Lentzen G, Seitz H, Rampp M, Schuster SC, Klenk HP, Pfeiffer F, Oesterhelt D, Kunte HJ (2011) A blueprint of ectoine metabolism from the genome of the industrial producer Halomonas elongata DSM 2581T. Environ Microbiol 13:1973–1994
Severin J, Wohlfarth A, Galinski EA (1992) The predominant role of recently discovered tetrahydropyrimidines for the osmoadaptation of halophilic eubacteria. J Gen Microbiol 138:1629–1638
Shapovalova AA, Hizhniak TV, Tourova TP, Muyzer G, Sorokin DY (2008) Heterotrophic denitrification at extremely high salt and pH by haloalkaliphilic Gammaproteobacteria from hypersaline soda lakes. Extremophiles 12:619–625
Shibata M, Katoh H, Sonoda M, Ohkawat H, Shimoyama M, Fukuzawa H, Kaplan A, Ogawa T (2002) Genes essential to sodium-dependent bicarbonate transport in cyanobacteria. J Biol Chem 277:18658–18664
Silvius JR (1982) Thermotropic phase transitions of pure lipids in model membranes and their modification by membrane proteins. In: Jost PC, Griffith OH (eds) Lipid-protein interactions, vol 2. Wiley, New York, pp 239–281
Soontharapirakkul K, Incharoensakdi A (2010) A Na+-stimulated ATPase of alkaliphilic halotolerant cyanobacterium Aphanothece halophytica translocates Na+ into proteoliposomes via Na+ uniport mechanism. BMC Biochem 11:30
Sorokin DY, Kuenen JG (2005) Haloalkaliphilic sulfur-oxidizing bacteria in soda lakes. FEMS Microbiol Rev 29:685–702
Sorokin DY, Jones BE, Kuenen JG (2000) An obligate methylotrophic, methane-oxidizing Methylomicrobium species from a highly alkaline environment. Extremophiles 4:145–155
Sorokin DY, Trotsenko YA, Doronina NV, Tourova TP, Muyzer G (2007) Methylohalomonas lacustra gen. nov., sp. nov. and Methylonatronum kenyi gen. nov., sp. nov., new methylotrophic gammaproteobacteria from hypersaline lakes. Int J Syst Evol Microbiol 57:2762–2769
Sorokin DY, Tourova TP, Muyzer G, Kuenen GJ (2008a) Thiohalospira halophila gen. nov., sp. nov. and Thiohalospira alkaliphila sp. nov., novel obligately chemolithoautotrophic, halophilic, sulfur-oxidizing gammaproteobacteria from hypersaline habitats. Int J Syst Evol Microbiol 58:1685–1692
Sorokin DY, Tourova TP, Henstra AM, Stams AJM, Galinski EA, Muyzer G (2008b) Sulfidogenesis at extremely haloalkaline conditions by Desulfonatronospira thiodismutans gen. nov., sp. nov., and Desulfonatronospira delicata sp. nov. – a novel lineage of Deltaproteobacteria from hypersaline soda lakes. Microbiology (UK) 154:1444–1453
Sorokin DY, Tourova TP, Galinski EA, Muyzer G, Kuenen JG (2008c) Thiohalorhabdus denitrificans gen. nov., sp. nov., an extremely halophilic, sulfur-oxidizing, deep-lineage gammaproteobacterium from hypersaline habitats. Int J Syst Evol Microbiol 58:2890–2897
Sorokin DY, Tourova TP, Mussmann M, Muyzer G (2008d) Dethiobacter alkaliphilus gen. nov. sp. nov., and Desulfurivibrio alkaliphilus gen. nov. sp. nov. – two novel representatives of reductive sulfur cycle from soda lakes. Extremophiles 12:431–439
Sorokin ID, Kravchenko IK, Doroshenko EV, Boulygina ES, Zadorina EV, Tourova TP, Sorokin DY (2008e) Haloalkaliphilic diazotrophs in soda solonchak soils. FEMS Microbiol Ecol 65:425–433
Sorokin ID, Zadorina EV, Kravchenko IK, Boulygina ES, Tourova TP, Sorokin DY (2008f) Natronobacillus azotifigens gen. nov., sp. nov., an anaerobic diazotrophic haloalkaliphile from soda-rich habitats. Extremophiles 12:819–827
Sorokin DY, van Pelt S, Tourova TP, Evtushenko LI (2009) Nitriliruptor alkaliphilus gen. nov., sp. nov., a deep-lineage haloalkaliphilic actinobacterium from soda lakes capable of growth on aliphatic nitriles, and proposal of Nitriliruptoraceae fam. nov. and Nitriliruptorales ord. nov. Int J Syst Evol Microbiol 59:248–253
Sorokin DY, Kuenen JG, Muyzer G (2011a) The microbial sulfur cycle at extremely haloalkaline conditions of soda lakes. Front Microbiol 2:44
Sorokin DY, Detkova EN, Muyzer G (2011b) Sulfur-dependent respiration under extremely haloalkaline conditions in soda lake ‘acetogens’ and the description of Natroniella sulfidigena sp. nov. FEMS Microbiol Lett 319:88–95
Sorokin DY, Tourova TP, Kolganova TV, Detkova EN, Galinski EA, Muyzer G (2011c) Culturable diversity of lithotrophic haloalkaliphilic sulfate-reducing bacteria in soda lakes and the description of Desulfonatronum thioautotrophicum sp. nov., Desulfonatronum thiosulfatophilum sp. nov., Desulfonatronovibrio thiodismutans sp. nov., and Desulfonatronovibrio magnus sp. nov. Extremophiles 15:391–401
Sorokin DY, Panteleeva AN, Tourova TP, Kaparullina EN, Muyzer G (2011d) Natronoflexus pectinivorans gen. nov., sp. nov., an obligately anaerobic and alkaliphilic fermentative member of Bacteroidetes from soda lakes. Extremophiles 15:691–696
Sorokin DY, Tourova TP, Panteleeva AN, Muyzer G (2012a) Desulfonatronobacter acidivorans gen. nov., sp. nov., and Desulfobulbus alkaliphilus sp. nov., haloalkaliphilic heterotrophic sulfate-reducing bacteria from soda lakes. Int J Syst Evol Microbiol 62:2107–2113
Sorokin DY, Tourova TP, Abbas B, Suhacheva MV, Muyzer G (2012b) Desulfonatronovibrio halophilus sp. nov., a novel moderately halophilic sulfate-reducing bacterium from hypersaline chloride-sulfate lakes in Central Asia. Extremophiles 16:411–417
Sorokin DY, Tourova TP, Panteleeva AN, Kaparullina EN, Muyzer G (2012c) Anaerobic utilization of pectinous substrates at extremely haloalkaline conditions by Natranaerovirga pectinivora gen. nov., sp. nov., and Natranaerovirga hydrolytica sp. nov., isolated from hypersaline soda lakes. Extremophiles 16:307–315
Sorokin DY, Tourova TP, Sukhacheva MV, Mardanov AV, Ravin NV (2012d) Bacterial chitin utilisation at extremely haloalkaline conditions. Extremophiles 16:883–894
Spanheimer R, Müller V (2008) The molecular basis of salt adaptation in Methanosarcina mazei Gö1. Arch Microbiol 190:271–279
Steuber J (2001) Na+ translocation by bacterial NADH, quinone oxidoreductases, an extension to the complex-I family of primary redox pumps. Biochim Biophys Acta 1505:45–56
Suzuki T, Murakami T, Iino R, Suzuki J, Ono S, Shirakihara Y, Yoshida M (2003) F0F1-ATPase/synthase is geared to the synthesis mode by conformational rearrangement of epsilon subunit in response to proton motive force and ADP/ATP balance. J Biol Chem 278:46840–46846
Swartz TH, Ikewada S, Ishikawa O, Ito M, Krulwich TA (2005) The Mrp system, a giant among monovalent cation/proton antiporters? Extremophiles 9:345–354
Switzer Blum J, Kulp TR, Han H, Lanoil B, Saltikov CW, Stolz JF, Miller LG, Oremland RS (2012) Desulfohalophilus alkaliarsenatis gen. nov., sp. nov., an extremely halophilic sulfate- and arsenate-respiring bacterium from Searles Lake, California. Extremophiles 16:727–742. doi:10.1007/s00792-012-0468-6
Takaichi S, Maoka T, Akimoto N, Sorokin DY, Banciu H, Kuenen JG (2004) Two novel yellow pigments natronochrome and chloronatronochrome from the natrono(alkali)philic sulfur-oxidizing bacterium Thioalkalivibrio versutus ALJ 15. Tetrahedron Lett 45:8303–8305
Takami H (2011) Genomics and evolution of alkaliphilic Bacillus species. In: Horikoshi K (ed) Extremophiles handbook. Springer, Tokyo, pp 184–211
Takami H, Nakasone K, Takaki Y, Maeno G, Sasaki R, Masui N, Fuji F, Hirama C, Nakamura Y, Ogasawara N, Kuhara S, Horikoshi K (2000) Complete genome sequence of the alkaliphilic bacterium Bacillus halodurans and genomic sequence comparison with Bacillus subtilis. Nucleic Acids Res 28:4317–4331
Takami H, Takaki Y, Uchiyama I (2002) Genome sequence of Oceanobacillus iheyensis isolated from the Iheya Ridge and its unexpected adaptive capabilities to extreme environments. Nucleic Acids Res 30:3927–3935
Tasaka Y, Gombos Z, Nishiyama Y, Mohanty P, Ohba T, Ohki K, Murata N (1996) Targeted mutagenesis of acyl-lipid desaturases in Synechocystis, evidence for the important roles of polyunsaturated membrane lipids in growth, respiration and photosynthesis. EMBO J 15:6416–6425
Tindall BJ (1988) Prokaryotic life in the alkaline, saline, athalassic environment. In: Rodriguez-Valera F (ed) Halophilic bacteria, vol 1. CRC Press, Boca Raton, pp 31–70
To TM, Grandvalet C, Tourdot-Maréchal R (2011) Cyclopropanation of membrane unsaturated fatty acids is not essential to the acid stress response of Lactococcus lactis subsp. cremoris. Appl Environ Microbiol 77:3327–3334
Trotsenko YA, Khmelenina VN (2002) Biology of extremophilic and extremotolerant methanotrophs. Arch Microbiol 177:123–131
Trotsenko YA, Doronina NV, Li TD, Reshetnikov AS (2007) Moderately haloalkaliphilic aerobic methylobacteria. Microbiology (Moscow) 76:253–265
Tseng CP, Albrecht J, Gunsalus RP (1996) Effect of microaerophilic cell growth conditions on expression of the aerobic (cyoABCDE and cydAB) and anaerobic (narGHJI, frdABCD, and dmsABC) respiratory pathway genes in Escherichia coli. J Bacteriol 178:1094–1098
Tuteja N (2007) Mechanisms of high salinity tolerance in plants. Methods Enzymol 428:419–438
Tzubery T, Rimon A, Padan E (2004) Mutation E252C increases drastically the Km value for Na+ and causes an alkaline shift of the pH dependence of NhaA Na+/H+ antiporter of Escherichia coli. J Biol Chem 279:3265–7322
Valderrama MJ, Monteoliva-Sanchez M, Quesada E, Ramos-Cormenzana A (1998) Influence of salt concentration on the cellular fatty acid composition of the moderately halophilic bacterium Halomonas salina. Res Microbiol 149:675–679
Vargas VA, Delgado OD, Hatti-Kaul R, Mattiasson B (2005a) Bacillus bogoriensis sp. nov., a novel alkaliphilic, halotolerant bacterium isolated from a Kenyan soda lake. Int J Syst Evol Microbiol 55:899–902
Vargas C, Kallimanis A, Koukkou AI, Calderon MI, Canovas D, Iglesias-Guerra F, Drainas C, Ventosa A, Nieto JJ (2005b) Contribution of chemical changes in membrane lipids to the osmoadaptation of the halophilic bacterium Chromohalobacter salexigens. Syst Appl Microbiol 28:571–581
Vargas C, Jebbar M, Carrasco R, Blanco C, Calderón MI, Iglesias-Guerra F, Nieto JJ (2006) Ectoines as compatible solutes and carbon and energy sources for the halophilic bacterium Chromohalobacter salexigens. J Appl Microbiol 100:98–107
Ventosa A, Nieto JJ, Oren A (1998) Biology of moderately halophilic aerobic bacteria. Microbiol Mol Biol Rev 62:504–544
Vreeland RH, Anderson R, Murray RGE (1984) Cell wall and phospholipid composition and their contribution to the salt tolerance of Halomonas elongata. J Bacteriol 160:879–883
Waditee R, Hibino T, Tanaka Y, Nakamura T, Incharoensakdi A, Takabe T (2001) Halotolerant cyanobacterium Aphanothece halophytica contains an Na+/H+ antiporter, homologous to eukaryotic ones, with novel ion specificity affected by C-terminal tail. J Biol Chem 276:36931–36938
Wang Z, Hicks DB, Guffanti AA, Baldwin K, Krulwich TA (2004) Replacement of amino acid sequence features of a- and c-subunits of ATP synthases of alkaliphilic Bacillus with the Bacillus consensus sequence results in defective oxidative phosphorylation and non-fermentative growth at pH 10.5. J Biol Chem 279:26546–26554
Wang QF, Li W, Liu YL, Cao HH, Li Z, Guo GQ (2007) Bacillus qingdaonensis sp. nov., a moderately haloalkaliphilic bacterium isolated from a crude sea-salt sample collected near Qingdao in eastern China. Int J Syst Evol Microbiol 57:1143–1147
Wei Y, Guffanti AA, Ito M, Krulwich TA (2000) Bacillus subtilis YqkI is a novel malic/Na+-lactate antiporter that enhances growth on malate at low protonmotive force. J Biol Chem 275:30287–30292
Wei Y, Liu J, Ma Y, Krulwich TA (2007) Three putative cation/proton antiporters from the soda lake alkaliphile Alkalimonas amylolytica N10 complement an alkali-sensitive Escherichia coli mutant. Microbiology 153:2168–2179
Welsh DT, Lindsay YE, Caumette P, Herbert RA, Hannan J (1996) Identification of trehalose and glycine betaine as compatible solutes in the moderately halophilic sulfate reducing bacterium, Desulfovibrio halophilus. FEMS Microbiol Lett 140:203–207
Williams RJ (1988) Proton circuits in biological energy interconversions. Annu Rev Biophys Biophys Chem 17:71–97
Wu XY, Zheng G, Zhang WW, Xu XW, Wu M, Zhu XF (2009) Amphibacillus jilinensis sp. nov., a facultatively anaerobic, alkaliphilic Bacillus from a soda lake. Int J Syst Evol Microbiol 60:2540–2543
Wu XY, Shi KL, Xu XW, Wu M, Oren A, Zhu XF (2010) Alkaliphilus halophilus sp. nov., a strictly anaerobic and halophilic bacterium isolated from a saline lake, and emended description of the genus Alkaliphilus. Int J Syst Evol Microbiol 60:2898–2902
Wutipraditkul N, Waditee R, Incharoensakdi A, Hibino T, Tanaka Y, Nakamura T, Shikata M, Takabe T, Takabe T (2005) Halotolerant cyanobacterium Aphanothece halophytica contains NapA-type Na+/H+ antiporters with novel ion specificity that are involved in salt tolerance at alkaline pH. Appl Environ Microbiol 71:4176–4184
Yang L, Jiang J, Zhang B, Zhao B, Wang L, Yang SS (2006) A primary sodium pump gene of the moderate halophile Halobacillus dabanensis exhibits secondary antiporter properties. Biochem Biophys Res Commun 346:612–617
Yang CX, Liu YP, Bao QH, Feng FY, Liu HR, Zhang XJ, Zhao YL (2012) Mongoliitalea lutea gen. nov., sp. nov., an alkaliphilic, halotolerant bacterium isolated from a haloalkaline lake. Int J Syst Evol Microbiol 62:647–653
Yoshimune K, Morimoto H, Hirano Y, Sakamoto J, Matsuyama H, Yumoto I (2010) The obligate alkaliphile Bacillus clarkii K24-1U retains extruded protons at the beginning of respiration. J Bioenerg Biomembr 42:111–116
Zavarzin GA (1993) Epicontinental alkaline water bodies as relict biotopes for the development of terrestrial biota. Microbiology (Moscow) 62:789–800
Zavarzin GA, Zhilina TN, Kevbrin VV (1999) The alkaliphilic microbial community and its functional diversity. Microbiology (Moscow) 68:579–599
Zelles L (1999) Fatty acid patterns of phospholipids and lipopolysaccharides in the characterisation of microbial communities in soil, a review. Biol Fertil Soils 29:111–129
Zenova GM, Oborotov GV, Zvyagintsev DG (2005) Solonchaks, the ecotopes of halophilic and alkalotolerant Streptomycetes. Eur Soil Sci 38:1190–1193
Zhai L, Liao T, Xue Y, Ma Y (2012) Bacillus daliensis sp. nov., an alkaliphilic, Gram-positive bacterium isolated from a soda lake. Int J Syst Evol Microbiol 62:949–953
Zhao B, Mesbah NM, Dalin E, Goodwin L, Nolan M, Pitluck S, Chertkov O, Brettin TS, Han J, Larimer FW, Land ML, Hauser L, Kyrpides N, Wiegel J (2011) Complete genome sequence of the anaerobic, halophilic alkalithermophile Natranaerobius thermophilus JW/NM-WN-LF. J Bacteriol 193:4023–4024
Zhilina TN, Zavarzin GA, Detkova EN, Rainey FA (1996) Natroniella acetigena gen. nov. sp. nov., an extremely haloalkaliphilic, homoacetic bacterium, a new member of Haloanaerobiales. Curr Microbiol 32:320–326
Zhilina TN, Zavarzin GA, Rainey FA, Pikuta EN, Osipov GA, Kostrikina NA (1997) Desulfonatronovibrio hydrogenovorans gen. nov., sp. nov., an alkaliphilic, sulfate-reducing bacterium. Int J Syst Bacteriol 47:144–149
Zhilina TN, Detkova EN, Rainey FA, Osipov GA, Lysenko AM, Kostrikina NA, Zavarzin GA (1998) Natronoincola histidinovorans gen. nov., sp. nov., a new alkaliphilic acetogenic anaerobe. Curr Microbiol 37:177–185
Zhilina TN, Garnova ES, Turova TP, Kostrikina NA, Zavarzin GA (2001) Halonatronum saccharophilum gen. nov., sp. nov., a new haloalkaliphilic bacterium of the order Haloanaerobiales from Lake Magadi. Microbiology (Moscow) 70:64–72
Zhilina TN, Kevbrin VV, Tourova TP, Lysenko AM, Kostrikina NA, Zavarzin GA (2005) Clostridium alkalicellum sp. nov., an obligately alkaliphilic cellulolytic bacterium from a soda lake in the Baikal region. Microbiology (Moscow) 74:557–566
Zhu L, Cheng J, Luo B, Feng S, Lin J, Wang S, Cronan JE, Wang H (2009) Functions of the Clostridium acetobutylicium FabF and FabZ proteins in unsaturated fatty acid biosynthesis. BMC Microbiol 9:119
Acknowledgements
The completion of this review was supported by a grant of the Romanian National Authority for Scientific Research, CNCS – UEFISCDI, project number PN-II-ID-PCE-2011-3-0546.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Banciu, H.L., Sorokin, D.Y. (2013). Adaptation in Haloalkaliphiles and Natronophilic Bacteria. In: Seckbach, J., Oren, A., Stan-Lotter, H. (eds) Polyextremophiles. Cellular Origin, Life in Extreme Habitats and Astrobiology, vol 27. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-6488-0_5
Download citation
DOI: https://doi.org/10.1007/978-94-007-6488-0_5
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-007-6487-3
Online ISBN: 978-94-007-6488-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)