Abstract
The detection of human cancer at an early stage presents a daunting challenge due to the lack of a specific non-invasive marker. The discovery of microRNAs (miRNAs), a class of non-coding RNAs that regulate target gene expression at the post-transcriptional level, has opened a new avenue for tumor diagnosis. Once thought to be unstable RNA molecules, miRNAs are now shown to be stably expressed in serum, plasma, urine, saliva, milk, and other body fluids. Results of the last 2 years establish the quantification of circulating miRNAs in serum/plasma as an extremely promising approach for cancer diagnosis, especially at an early stage. The aim of this chapter is to review the recently reported studies on the circulating miRNAs in cancer patients and to estimate their great potential as a class of highly specific and sensitive blood-based biomarkers for tumor classification and prognostication. Meanwhile, this chapter also addresses certain critical issues that hinder the wide application of the new approach. Some potential challenges for the transition of circulating miRNAs from a research setting to a clinical application are also highlighted. As a blood-based biomarker of cancer, circulating miRNAs have the potential to uncover cancer at an early stage and greatly reduce the worldwide health burden of cancer.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
21.1 Introduction
Identifying the sensitive, specific, and non-invasive biomarkers for detecting tumors at an early stage has been regarded as a key point in tumor prevention and therapy. In general, human cancers, like other diseases, are easier to treat and control when detected at an early stage of disease. For example, 65% of non-small cell lung cancer (NSCLC) patients have advanced disease at the time of diagnosis (Brambilla et al. 2003; Patz et al. 2000). Most NSCLC cases, particularly at stages I and II, rarely show symptoms and are difficult to be detected (Brambilla et al. 2003; Patz et al. 2000; Rossi et al. 2005). The 5-year survival rate following surgical resection is approximately 70% for patients with stage I NSCLC, but that rate drops to only 30% for patients with stage III NSCLC (Dominioni et al. 2000). Thus, the earlier detection of NSCLC will greatly facilitate more effective management of the disease. Likewise, early detection of cancer is highly demanded for patients with other cancers, including colon cancer, cervical cancer, ovarian cancer, breast cancer, etc. The development of non-invasive or minimally invasive tests capable to uncover cancer at an early stage can greatly reduce the worldwide health burden of cancers.
Unfortunately, few effective tests are available so far. To date, the reference gold standard in diagnosing human cancers, such as NSCLC, is pathologic evidence of malignant cells, which typically requires an invasive strategy, such as surgery, biopsy, needle aspiration, bronchoscopy, or thoracotomy. Chest X-ray and computed tomography screening can detect some cancers at an early stage (Henschke et al. 2006; Oken et al. 2005), but the diagnostic procedures and the hazards of the associated radiation may outweigh the potential benefits (Mascalchi et al. 2006; Taplin et al. 2008). For breast cancer, mammography is currently the gold standard diagnostic tool; however, it also has limitations, including ionizing radiation and a false positive rate of 8–10% (Taplin et al. 2008). These concerns have led researchers to seek novel biomarkers and new diagnostic assays to non-invasively assess tumors.
Blood-based biomarkers are appealing because they can be used in screening asymptomatic patients. It is generally believed that disease-related biomarkers, including antigens, enzymes, and lipid components, could shed from the tissues or cells with ongoing diseases and be present in blood. Thus, a variety of protein markers, genetic markers, and epigenetic markers have been extensively investigated in blood in an attempt to seek the markers for early stage tumors. For example, several currently available serum/plasma biomarkers, such as carcinoembryonic antigen (CEA), neuron-specific enolase (NSE), chromogranin A (CgA), carbohydrate antigen 15-3 (CA 15-3), carbohydrate antigen 19-9 (CA19-9), alpha-fetoprotein (AFP), and human chorionic gonadotropin (hCG), offer some promise of analyzing tumors comprehensively without the need to carry out a biopsy or a surgical procedure (Dbouk et al. 2007; Huber et al. 1992; Schneider et al. 2000; Sorio et al. 2006; Tarro et al. 2005). However, up to date, only limited success has been achieved in identifying markers those are clinically effective in diagnostic tests. The major concern of these markers is their low sensitivities and specificities (Harris et al. 2007). Even for prostate-specific antigen (PSA), which has been shown to be sensitive for advanced prostate cancer, the specificity is still limited, as evidenced by a high frequency of falsely elevated values in men with benign prostatic hyperplasia (Stenman et al. 1999). Conventional methodologies for the detection and quantification of antigens have also been limited in clinical use due to their low-throughput nature. Recent studies in proteomics have led to the development and broad use of mass spectrometry, a technology that shows the potential to identify unknown proteins in plasma (Gerszten et al. 2010; Gygi et al. 2002; Makawita and Diamandis 2010; Ward et al. 2006; Zhou et al. 2002). With the new technologies and better sample preparation protocols, the sensitivity of mass spectrometry is extremely high, with a detection limit of 1 pg/mL. However, such type of assay is expensive and labor-intensive, which makes it difficult for widespread application in clinical diagnosis. Although the enzyme-linked immunosorbent assay (ELISA) can be performed in a high-throughput manner and is suitable for the detection of diagnostic antigens, its detection sensitivity is greatly dependent on antibody affinities. Therefore, it is important to develop new methods and novel diagnostic biomarkers for the detection of early events of cancer.
Recently, a new class of RNA regulatory genes known as microRNAs (miRNAs) has been found to introduce a whole new layer of gene regulation in eukaryotes (Bartel 2004; He and Hannon 2004). MiRNAs are endogenous non-coding RNAs, consisting of 19–24 nucleotides in length (Bartel 2004; He and Hannon 2004). They play an important role in regulating gene expression by base-pairing to the complementary sites on the target mRNAs, thus blocking the translation or triggering the degradation of the target mRNAs (Bartel 2004; He and Hannon 2004). Since the discovery of the first miRNA lin-4 in C. elegans, thousands of miRNAs have been identified experimentally or computationally from a variety of species (Bartel 2004; He and Hannon 2004). MiRNAs are currently estimated to comprise 1–5% of animal genes and collectively regulate up to 30% of genes, making them one of the most abundant classes of regulators (Bartel 2004; He and Hannon 2004). Over the past several years, it has been well demonstrated that alteration of the miRNA expression contributes to the pathogenesis of many human malignancies (Calin and Croce 2006; Cho 2010; Esquela-Kerscher and Slack 2006). A number of miRNAs, including miR-15a/miR-16-1 (Calin et al. 2002), miR-21 (Meng et al. 2007; Ribas et al. 2009; Selaru et al. 2009), miR-155 (Eis et al. 2005; Gironella et al. 2007), miR-200 family (Gregory et al. 2008), miR-210 (Camps et al. 2008; Mathew and Simon 2009), etc, have been found to be differentially expressed in various tumor tissues and cells. Based on their differential roles in carcinogenesis, miRNAs are categorized as either oncogenes or tumor suppressors. These miRNAs not only serve as specific indicators of various aspects of carcinogenesis and tumor development, but also provide important sources for the development of future miRNA-based cancer treatment and therapies (Calin and Croce 2006; Cho 2010; Esquela-Kerscher and Slack 2006).
Recent studies by our group and others have showed that miRNAs are stably present in many body fluids, including serum (Chen et al. 2008; Gilad et al. 2008; Mitchell et al. 2008), plasma (Chen et al. 2008; Mitchell et al. 2008), saliva (Park et al. 2009), urine (Hanke et al. 2010), milk (Kosaka et al. 2010), and cell culture supernatants (Skog et al. 2008; Valadi et al. 2007). Once thought as unstable RNA molecules, circulating miRNAs are in fact highly stable and readily detected in serum and plasma. Biochemical analyses indicate that they are resistant to RNase digestion, as well as extreme pH and temperature (Chen et al. 2008; Mitchell et al. 2008). More importantly, the expression patterns of circulating miRNAs are tightly correlated with various diseases including cancer, diabetes, and tissue injury. These results firmly establish the quantification of the tumor-derived miRNAs in serum/plasma as an extremely promising approach for cancer diagnostics.
In this chapter, we provide an update and overview of the new findings that reveal the potential use of circulating miRNAs as clinically diagnostic biomarkers of various cancers and other diseases. Additionally, we summarize the approaches used to detect circulating miRNAs and address some critical issues in the quantification of these molecules. Finally, we estimate their impact on making the ongoing research closer to clinical application.
21.2 Detection and Characterization of Circulating MiRNAs in Serum and Plasma
21.2.1 MiRNAs and Human Cancer
MiRNAs have been studied intensively in the field of oncology research, and emerging evidence suggests that miRNAs are involved in the pathogenesis of cancers, mainly by regulating the translation of oncogenes and tumor suppressors (Calin and Croce 2006; Cho 2010; Esquela-Kerscher and Slack 2006). By employing the techniques including miRNA cloning, quantitative reverse transcription-polymerase chain reaction (qRT-PCR), microarray, and bead-based profiling method, changes in the expression profiles of miRNAs have been detected in a broad spectrum of hematological malignancies and solid tumors (Cummins et al. 2006; Iorio et al. 2005; Lu et al. 2005; Volinia et al. 2006; Yanaihara et al. 2006). Some miRNAs are frequently found to be up-regulated or down-regulated in tumors compared with normal tissues, in agreement with their dual role in carcinogenesis as either “oncogenes” or “tumor suppressors” (Calin and Croce 2006; Cho 2010; Esquela-Kerscher and Slack 2006). Although expression differences may not necessarily reflect causal events of carcinogenesis, such changes may, nevertheless, be useful for classifying tumors and predicting their outcomes (Lu et al. 2005; Rosenfeld et al. 2008; Yanaihara et al. 2006). Other reports also showed that miRNAs are remarkably stable in formalin-fixed, paraffin-embedded tissues, as well as fresh snap-frozen specimens (Li et al. 2007; Xi et al. 2007). Such information is particularly useful for applying tumor miRNA expression profiles as biomarkers to diminish the diagnostic uncertainty. Since the biopsy for pathologic analysis is a common routine procedure to diagnose solid tumors, the expression profile of tissue miRNAs from biopsy samples become a novel biomarker in early tumor detection. Many studies have demonstrated the differential expression of miRNAs between cancer and the related normal tissue and indicated the potential use of these molecules as biomarkers. However, for developing a non-invasive or minimally invasive human cancer diagnostic tool, more intriguing question is whether the tumor-derived miRNAs are presented and detectable in serum, plasma, and other human body fluids.
21.2.2 Existence of MiRNAs in Serum and Plasma
Dysregulated expression of miRNAs in various tissues has been associated with a variety of human cancers. More recently, miRNAs’ occurrence in the serum and plasma of humans has been repeatedly observed. The first description of the presence of circulating miRNAs in serum and their potential as cancer markers was reported by Lawrie et al. (2008) (Table 21.1). By comparing the levels of several tumor-associated miRNAs, such as miR-155, miR-210, and miR-21, in the serum of diffuse large B cell lymphoma (DLBCL) patients with those of healthy controls, Lawrie et al. (2008) determined that patients with DLBCL had high serum levels of miR-21. More interestingly, miR-21 levels were found to be associated with the relapse-free survival of these patients (Lawrie et al. 2008).
Around the same time, Mitchell et al. (2008) showed that miRNAs were present in human plasma in a remarkably stable form and were protected from endogenous RNase activity (Table 21.1). Using a mouse xenograft model where a human prostate cancer cell line was implanted into mice, they showed that there were tumor-derived miRNAs circulating in blood (Mitchell et al. 2008). By filtering plasma samples through a 0.22 μm filter or subjecting these samples to a series of centrifugations followed by RNA extraction and miRNA detection, they showed that those plasma miRNAs were not associated with cells or the broken cellular parts (Mitchell et al. 2008). Finally, the expression of certain miRNAs in cancer patient plasma, particularly the prostate cancer-associated miR-141, was found to be significantly different from that in control plasma (Mitchell et al. 2008). These data indicate that tumor-derived miRNAs in human plasma could serve as important biomarkers for the blood-based detection of human cancer.
In our study, we give the first comprehensive tally of miRNAs in blood serum and identify patterns of miRNAs that distinguish patients with two kinds of cancer and diabetes from healthy subjects (Chen et al. 2008) (Table 21.1). Firstly, we detected miRNAs in the sera of various animal species including human, mouse, rat, bovine, and horse by employing a stem-loop qRT-PCR assay (Chen et al. 2008). Secondly, we showed that serum miRNAs were highly stable even in the harsh conditions such as extreme temperature and pH, or RNase digestion, and serum miRNAs can be almost fully recovered from boiling, freeze-thaw cycles or long-time storage in frozen condition (Chen et al. 2008). Moreover, we showed that the levels of miRNAs in serum were reproducible and consistent among individuals of the same species (Chen et al. 2008). In order to obtain the complete expression profile of miRNAs in serum, we further performed high-throughput Solexa sequencing using various serum samples pooled from healthy controls or cancer patients. As a high-throughput sequencing technology producing highly accurate, reproducible, and quantitative readouts of small RNAs, Solexa can detect and define all small RNA molecules including miRNA with various expression levels, and can unambiguously distinguish miRNAs from other small RNAs (Chen et al. 2008, 2009). By this method, we investigated the types and levels of miRNAs in the serum samples pooled from healthy volunteers (male and female) or patients with various diseases including NSCLC, colorectal cancer (CRC), and diabetes. For each disease, we identified a unique expression pattern of miRNAs that differed from that of healthy people (Chen et al. 2008). Our study presented the first genome-wide expression profile of miRNAs in human serum. Interestingly, we also observed striking differences between the miRNAs in the serum and blood cells of cancer patients, while there were no differences between the serum and blood cells of healthy individuals (Chen et al. 2008). The data suggest that some serum miRNAs of cancer patients cannot be derived from circulating blood cells but likely from tumor cells or tissues that are associated with tumorigenesis. It is of interest to note that NSCLC patients shared a large number of serum miRNAs with CRC patients, and Pearson correlation further indicated that the levels of miRNAs in serum from NSCLC patients and CRC patients were consistent (Chen et al. 2008), suggesting that there are some “common” tumor-related miRNAs in serum from cancer patients. More specifically, our results determined that miR-25 and miR-223, two miRNAs previously shown to be involved in tumorigenesis, were highly expressed in NSCLC serum compared to those in normal serum (Chen et al. 2008). Therefore, these two serum-miRNAs could serve as novel blood-based biomarkers of NSCLC.
Taken together, these early reports by us and other groups clearly showed that miRNAs circulate in a stable, cell-free form in the bloodstream and that the abundance of specific miRNAs in plasma or serum can serve as biomarkers of cancers.
21.2.3 Circulating MiRNAs as a Biomarker for Cancer Diagnosis
More recently, miRNAs’ occurrence in the serum and plasma of humans has been repeatedly observed (Table 21.1). By comparing the plasma samples of patients with CRC to those of healthy controls, Ng et al. (2009) found that both miR-17-3p and miR-92 are significantly elevated in the plasma of CRC patients. Interestingly, the levels of these miRNAs in plasma were significantly reduced after surgery in 10 patients with CRC (Ng et al. 2009), suggesting that plasma miRNAs may serve as indicators for monitoring cancer patients. Further validation with an independent set of plasma samples indicated that miR-92 differentiated CRC not only from normal subjects but also from gastric cancer and inflammatory bowel disease (Ng et al. 2009). In discriminating CRC from control subjects, this marker yielded an ROC curve area of 88.5% (the sensitivity was 89% and the specificity was 70%) (Ng et al. 2009). Thus, miR-92 has reasonable sensitivity for CRC detection and can be employed as a potential non-invasive blood-based biomarker for CRC screening. In accordance with this observation, Huang et al. (2010) also measured the levels of 12 miRNAs in plasma samples from patients with advanced colorectal neoplasia and healthy controls using qRT-PCR and found that plasma miR-29a and miR-92a have significant diagnostic value for advanced neoplasia.
Detection of the expression of individual or a panel of circulating miRNAs in serum and plasma from cancer patients has also been widely reported by other investigators (Table 21.1). To determine the utility of serum miRNAs as biomarkers for epithelial ovarian cancer, Resnick et al. (2009) compared the serum miRNA levels in cancer specimens to normal specimens with qRT-PCR assay. They identified eight miRNAs that were significantly differentially expressed between cancer and normal specimens, of which miR-21, miR-92, miR-93, miR-126, and miR-29a were over-expressed while miR-155, miR-127, and miR-99b were under-expressed (Resnick et al. 2009). Employing a pan-human miRNA, high density microarray, Lodes et al. (2009) evaluated miRNA expression patterns in human serum for five types of human cancer, and found that miRNAs present in 1 mL of serum were sufficient to detect miRNA expression patterns, without the need for amplification techniques. Their findings provide a clue to use these expression patterns in correctly discriminating between normal and cancer patient samples. Moreover, in the process of identifying differentially-expressed circulating miRNAs in pancreatic cancer, Wang et al. (2009a) revealed that four pancreatic cancer-specific miRNAs, including miR-21, miR-210, miR-155, and miR-196a, had elevated expression levels in the serum of pancreatic cancer patients compared to that of normal controls. By both microarray and qRT-PCR analysis, Tanaka et al. (2009) revealed that miR-638 is stably present in human plasma while miR-92a dramatically decreased in the plasma of acute leukemia patients, and the ratio of miR-92a/miR-638 in plasma was useful for distinguishing leukemia patients from healthy controls. In sum, the miRNA level in serum and plasma has strong potential for clinical application as a novel biomarker for detection of various human cancers.
21.2.4 Circulating MiRNAs as a Biomarker for Tumor Classification and Prognostication
MiRNAs in serum and plasma may also be used as a minimally invasive method for tumor classification and prognostication (Table 21.1). After confirming that RNA species can be detected in archived serum samples, Zhu et al. (2009) reported that the level of miR-155 was elevated in the serum of progesterone receptor positive patients compared to those with progesterone receptor negative breast cancer. Tsujiura et al. (2010) showed that the plasma concentrations of four miRNAs including miR-17-5p, miR-21, miR-106a, and miR-106b were significantly higher in gastric cancer patients than controls, whereas let-7a was lower in gastric cancer patients. Interestingly, they found that the levels of miR-21 and miR-106a were significantly reduced in post-operative samples (Tsujiura et al. 2010). Liu et al. (2010) showed that miR-31 in plasma was significantly elevated in oral squamous cell carcinoma patients relative to age and sex-matched control individuals. They also showed that the plasma miR-31 in patients was remarkably reduced after tumor resection (Liu et al. 2010). In order to investigate the potential relevance of circulating miRNAs with tumor progression in prostate cancer, Brase et al. (2011) screened miRNA expression in serum samples from patients with metastatic and localized prostate cancer and found that various miRNAs were highly abundant in the sera of patients with metastatic disease. After selecting and validating five up-regulated miRNAs in two independent sample sets, they demonstrated that miR-375 and miR-141 were pronounced markers for high-risk tumors (Brase et al. 2011). In sum, their study indicates that circulating miRNAs offer promise as non-invasive biomarkers for tumor progression. In a recent study, Hu et al. (2010) investigate the role of serum miRNA in predicting the prognosis of NSCLC by a genome-wide serum miRNA expression analysis. Specifically, they used Solexa sequencing followed by individual qRT-PCR assays to test the difference in levels of serum miRNAs between patients with longer survival and patients with shorter survival. Levels of four miRNAs including miR-486, miR-30d, miR-1, and miR-499 were found to be significantly associated with overall survival. The results revealed a great potential of serum miRNA signature as disease fingerprints to predict survival.
21.2.5 Circulating MiRNAs as an Indicator to Reflect Other Diseases
The profile of circulating miRNAs could also be used as an indicator to reflect other diseases or even subtle physiological condition changes (Table 21.2). For instance, some findings suggest the potential of tissue-specific circulating miRNAs as sensitive and informative biomarkers of tissue injury (Ji et al. 2009; Laterza et al. 2009; Wang et al. 2009b). To identify reliable and predictive markers to detect the early signs of drug-induced injury to the liver, one of the most vulnerable organs in the body, Wang et al. (2009b) used acetaminophen overdose-induced liver injury in the mouse as a model system and observed significant differences in the spectrum and levels of miRNAs in both liver tissues and in plasma between control and overdosed mice. A number of elevated circulating miRNAs in plasma collected from acetaminophen-overdosed animals are highly expressed in the liver (Wang et al. 2009b). In particular, miR-122 and miR-192, which are enriched in the liver tissue, exhibit dose- and exposure duration-dependent changes in the plasma that parallel serum aminotransferase levels and the histopathology of liver degeneration, but their changes can be detected significantly earlier (Wang et al. 2009b). The results suggest the potential of using specific circulating miRNAs as sensitive and informative biomarkers for drug-induced liver injury. Another study by Laterza et al. (2009) investigated the use of liver-, muscle-, and brain-specific miRNAs as circulating biomarkers of tissue injury. Tissue-specific miRNAs, miR-122, miR-133a, and miR-124, in plasma samples from rats treated with liver or muscle toxicants and from a rat surgical model of stroke, were determined by a highly sensitive qRT-PCR assay (Laterza et al. 2009). Increases in plasma concentrations of miR-122, miR-133a, and miR-124 corresponding to injuries in liver, muscle, and brain, respectively, were observed. Using isoproterenol-induced myocardial injury in rats as a model, Ji et al. (2009) showed that plasma concentrations of miR-208 increased significantly after myocardial injury and showed a similar time course to the concentration of plasma cardiac troponin I (cTnI), a classic biomarker of myocardial injury. Interestingly, Wang et al. (2010b) also found that elevated cardiac-specific miR-208a in plasma could be a novel biomarker for early detection of acute myocardial infarction (AMI) in humans. Likewise, Ai et al. (2010) also showed that muscle-specific miR-1 was significantly higher in plasma from AMI patients compared with non-AMI subjects and its level dropped to normal on discharge following medication. Moreover, Tijsen et al. (2010) identify 6 miRNAs that are elevated in patients with heart failure (HF), among which miR-423-5p is most strongly related to the clinical diagnosis of heart failure. Wang et al. (2010a) showed that serum miR-146a and miR-223 were significantly reduced in septic patients compared with systemic inflammatory response syndrome (SIRS) patients and healthy controls, suggesting that serum miR-146a and miR-223 might serve as new biomarkers for sepsis. Furthermore, Gilad et al. (2008) reported that healthy individuals shared a similar serum miRNA profile, while human placenta-associated miRNAs such as miR-527 and miR-526a were significantly elevated in pregnant women compared to those in non-pregnant women. Chim and co-workers (2008) also reported that placental miRNAs can be detected in serum and plasma, and the levels of these miRNAs correlate with the stages of pregnancy.
21.3 Detection and Characterization of Circulating MiRNAs in Other Body Fluids
MiRNAs have also been shown to be stably expressed in body fluids including urine (Hanke et al. 2010), saliva (Park et al. 2009), milk (Kosaka et al. 2010), and cell culture supernatants (Skog et al. 2008; Valadi et al. 2007) (Table 21.3). In a recent study, Hanke et al. (2010) develop a clinically applicable and sensitive protocol for the preparation and molecular analysis of miRNAs from urine samples obtained from bladder cancer (BCa) patients or healthy volunteers. They showed that many miRNAs had higher abundance in the urine samples from BCa patients compared to those from healthy volunteers (Hanke et al. 2010). Among 157 miRNA species being analyzed, the differential expression of miR-126 and miR-182 in urine was found to be able to identify BCa (Hanke et al. 2010). Several studies have shown the presence of miRNA in saliva and their potential as an additional set of oral cancer biomarkers. Park and co-workers (2009) detected approximately 50 miRNAs in both the whole and supernatant saliva, and showed that endogenous saliva miRNA degraded much slower compared with exogenous miRNA. By comparing the expressions of miRNAs in saliva of 50 oral squamous cell carcinoma patients and 50 healthy matched control subjects by qRT-PCR, they identified two miRNAs, miR-125a and miR-200a, whose expression levels were significantly lower in the saliva of oral squamous cell carcinoma patients than in control subjects (Park et al. 2009). In an attempt to identify putative body fluid-specific miRNAs, Hanson et al. (2009) reported the first comprehensive evaluation of miRNA expression in dried, forensically relevant biological fluids including blood, semen, saliva, vaginal secretions, and menstrual blood. They identified a panel of nine miRNAs (miR-451, miR-16, miR-135b, miR-10b, miR-658, miR-205, miR-124a, miR-372, and miR-412) that can be used as sensitive biomarkers to identify the body fluid origin of forensic biological stain. The results of this study provide an initial indication that miRNA profiling may provide a promising alternative approach for identification of body fluid in forensic casework.
Based on the findings discussed above, we conclude that: (i) miRNAs are stably expressed in human serum, plasma, and other body fluids and can be readily detected by various techniques including Solexa sequencing, miRNA microarray, and qRT-PCR; and (ii) the expression levels and patterns of circulating miRNAs are correlated with various human dysfunctional conditions. Given their size, abundance, tissue specificity, and relative stability, circulating miRNAs in serum, plasma, and other body fluids hold a great promise as non-invasive or minimally invasive biomarkers of various diseases including cancers.
21.4 Experimental Techniques and Issues in Analysis of Circulating MiRNAs
Quantification of circulating miRNAs requires effective tools. The explosion of interest in miRNAs over the recent years results in the development of various technique platforms. High-throughput profiling techniques such as Solexa sequencing, miRNA microarray, and bead-based miRNA profiling are effective tools for obtaining expression profiles of circulating miRNAs. These high-throughput techniques greatly facilitate the process of circulating miRNA expression profiling; in this sense, they are superior to the existing low-throughput techniques such as northern blotting and cloning. However, since high-throughput techniques generally require a relative large volume of serum/plasma to enrich total RNAs, they are usually used in initial screening processes. qRT-PCR methodologies have been widely applied in miRNA research, especially in assessing the low level of certain serum miRNAs in individual samples. To date, the most widely-used and successful approach in terms of specificity and sensitivity is a two-step approach using looped miRNA-specific reverse transcription primers and TaqMan probes (Chen et al. 2005; Tang et al. 2006). So far, both high and low-throughput techniques are extremely useful in miRNA study and are responsible for the majority of studies regarding to miRNA research. To make the detection of circulating miRNAs more simple, some new techniques have recently been developed (Lusi et al. 2009). Using an electrochemical genosensor, Lusi et al. (2009) were able to directly detect miRNAs without the need of PCR and a labeling reaction, and the test is simple, very fast and ultrasensitive, with a detection limit of 0.1 pmol. This approach, as well as other revolutionary technologies, may greatly facilitate the application of circulating miRNAs as blood-based biomarkers for the diagnosis of various human cancers.
The lack of a house-keeping gene for normalization is a major technique issue in the quantification of circulating miRNAs. In contrast to tissue or cellular miRNAs, of which the expression levels of certain miRNAs can be normalized against U6 snRNA or house-keeping miRNAs (Peltier and Latham 2008), circulating miRNAs have no such internal control for miRNA normalization. Although miRNAs are stable in serum and plasma, their levels may alter under various conditions. Moreover, there is considerable sample-to-sample variability in both protein and lipid content of plasma and serum samples, which could affect efficiency of RNA extraction. Therefore, how to normalize the level of circulating miRNA is an important issue the investigators concerning from the beginning. Since there is no common circulating miRNA that can be used as a standard, normalizing the level of circulating miRNAs by the volume of serum or plasma samples might be currently the most feasible way to solve the problem. Furthermore, since the yield of RNA from small volume of plasma or serum samples (i.e. 100 μL) is obviously below the limit of accurate quantification by spectrophotometry, it is reasonable to use a fixed volume of serum/plasma sample as input for qRT-PCR rather than to use a fixed mass of input RNA. However, this approach cannot rule out the contamination by technical variations. A more ideal approach is to normalize circulating miRNA levels using exogenous non-human (e.g. C. elegans or plants) miRNAs, which were usually spiked-in after the initial denaturation of plasma or serum samples (Mitchell et al. 2008; Hu et al. 2010; Kroh et al. 2010). The inclusion of the spiked-in miRNAs is important for adjusting for technical variations in RNA extraction. An effective normalization strategy for biological variability, however, is currently not well-developed. The use of so-called “invariant” miRNAs as endogenous controls, such as miR-16 (Lawrie et al. 2008) and miR-638 (Tanaka et al. 2009), has been proposed by some investigators. However, future studies should be performed to confirm that these miRNAs are indeed expressed at high levels in plasma and serum and are relatively invariant across large numbers of samples.
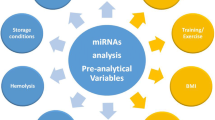
Furthermore, the clinical effectiveness of circulating miRNAs as biomarkers is likely to be affected by pre-analytic factors. For example, when generating plasma, there is a risk of contamination of the plasma supernatant by cells from the cellular pellet when aspirating (Kroh et al. 2010). Moreover, the type of anti-coagulant used in plasma collection tubes is also important to consider. Whereas EDTA and citrate are both acceptable anti-coagulants for downstream qRT-PCR, the use of heparin as the anti-coagulant potently inhibits subsequent PCR (Kroh et al. 2010). Furthermore, there are other pre-analytic variables that have yet to be studied carefully. Although miRNAs appear to be stable to extended room temperature incubation of plasma (Chen et al. 2008; Mitchell et al. 2008), it is not yet known whether the duration of time taken between collecting and processing of plasma or serum affects miRNA levels (Kroh et al. 2010). In the absence of data on this pre-analytic variable, it is prudent in designing studies to standardize conditions as much as possible with respect to time elapsed between collecting blood and processing for plasma or serum. These are some other factors, such as the diurnal variation in miRNA levels, fasting vs. non-fasting state at blood collection, etc, that may plausibly affect either the amount of miRNA present in a given plasma or serum sample (Kroh et al. 2010). It is prudent to try to match as many variables as possible in the collection and/or selection of case and control samples for research studies.
21.5 Circulating MiRNAs Serving as Novel Potential Biomarkers for Early Tumor Detection, Diagnosis, and Prognosis
21.5.1 A Panel of Circulating MiRNAs Instead of Individual Circulating MiRNA as a Biomarker for Tumor Detection
Ideal blood-based biomarkers of tumors must fit two criteria: (a) specificity, i.e. their presence is associated with the occurrence of only a specific type of tumor; and (b) proportionality, i.e., their levels in serum and plasma should be correlated with the extent of tumor development. Early studies clearly demonstrated that circulating miRNAs perfectly satisfied these standards. Tremendous efforts have been devoted to identify circulating miRNA-based novel non-invasive biomarkers for early tumor detection, diagnosis, and prognosis.
Early reports on circulating miRNAs as disease fingerprints mainly focused on individual or several tumor-related miRNAs. This approach has its advantage because detecting a single miRNA molecule or a few target miRNAs in circulation is much simpler and more straightforward than comprehensively detecting all circulating miRNAs. Successful examples of using this approach include the first two studies in this field, conducted by Lawrie et al. (2008) and Mitchell et al. (2008), respectively. However, in general, the specificity of biomarkers based on a single miRNA or several individual miRNAs is relatively poor. For instance, elevated liver-specific miR-122 in plasma or serum could result from not only hepatocellular carcinoma but also hepatitis B virus infection, cirrhosis, and general liver injury (Wang et al. 2009b). The molecular basis for the limitation of individual miRNA as a tumor biomarker is the diverse, complex nature of malignancy. For a normal cell transforming into a tumor cell, many different genes would be involved. Accordingly, there should be many miRNAs that target these genes contributing to this complex process. In other word, there are many miRNAs that are dysregulated in each type of cancer. For hepatocellular carcinoma, the altered miRNAs can be derived not only from liver but also immune system. To identify circulating miRNA-based biomarkers for various types of cancer, it is necessary to screen the serum/plasma miRNAs in a genome-wide manner and find all differentially-expressed miRNAs in serum/plasma of patients with that specific type of cancer. From this regard, employing a miRNA expression panel instead of an individual miRNA as a biomarker represents a rational option to circumvent the limitation in miRNAs utilization in tumor diagnosis.
21.5.2 A Working Model for Identifying Circulating MiRNA-based Biomarkers for Diseases
In the process of searching circulating miRNA-based cancer biomarker, the scientists have developed a working model to identify and refine differentially expressed miRNAs in cancer serum/plasma samples compared to control samples (Fig. 21.1). The analysis was separated into three steps: (i) initial screening by high-throughput techniques such as Solexa sequencing, microarray, or miRNA cloning using pooled or several serum samples; (ii) qRT-PCR validation in a large number of individual serum samples arranged in multiple training and testing sets; and (iii) statistical evaluation of the diagnostic or prognostic value of the circulating miRNA profiling system by algorithms such as ROC curve, Risk score analysis, and cluster analysis. Briefly, after the initial screening, a panel of differentially-expressed miRNAs could be derived by comparing the serum/plasma miRNA profile in healthy volunteers and cancer patients. Since the sample sizes used in the initial screening stage were small, the miRNAs selected had to be further validated by qRT-PCR assay at an individual level. This model has been proven to be greatly successful in identifying serum miRNA-based biomarkers for lung cancer classification and predicting the survival of lung cancer patients (Hu et al. 2010).
In general, initial screening by high-throughput techniques, such as Solexa and microarray, is to identify circulating miRNAs that are differentially expressed in patients’ serum, plasma, and other body fluids. However, if we know the specific miRNAs that are linked to certain type of diseases, we can directly determine their levels using miRNA qRT-PCR assay. One of alternative approaches is to use multiplex qRT-PCR assay to identify a panel of specific circulating miRNAs, which will be faster and simpler than sequencing or microarray studies.
21.6 Some Key Issues Regarding the Sources and Biological Functions of Circulating MiRNAs
21.6.1 Sources of Circulating MiRNAs
To establish circulating miRNAs as novel disease biomarkers, a key issue is to clarify their sources. Theoretically, there are two major pathways that have been proposed for miRNAs to enter the circulation: (i) passive leakage from broken cells and (ii) active secretion via cell-derived microvesicles (MVs) (Fig. 21.2).
Schematic description of the sources of circulating miRNAs. There are two major pathways that have been proposed for miRNAs to enter the circulation: (i) passive leakage from broken cells and (ii) active secretion via MVs. In the first pathway, tissues or cells at the primary site may break up under the conditions of tissue damage or cell apoptosis, and the miRNAs may leak into the circulating blood from these cells. In the second pathway, miRNAs are first loaded into MVs (exosomes and microparticles) inside the cells, and then enter the circulation accompanying the secretion of MVs. MiRNA, MicroRNA; MVs, Microvesicles
Passive leakage from broken cells: Although direct leakage of cellular miRNAs into the circulating is not common under normal circumstance, it still occurs at the time of tissue damage or cell apoptosis. Under the circumstances of tumor metastasis or chronic inflammation, tissues or cells at the primary site may break up; therefore, miRNAs may leak into the circulating blood from these cells. This type of miRNA release may also happen in certain blood cells, such as platelets and monocytes, because of their relatively short half-lives. In general, passive leakage from broken cells and tissues is a process that does not require energy.
Active secretion via cell-derived MVs: MVs are small vesicles that are shed from almost all cell types under both normal and pathological conditions (Cocucci et al. 2009; Thery et al. 2002). They generally include exosomes and microparticles, whose size and vesicular structures are quite different from each other. These secreted MVs bear surface receptors/ligands of the original cells and have the potential to selectively interact with specific target cells and mediate intercellular communication by transporting bioactive lipids, mRNA, or proteins between cells (Thery et al. 2002). MVs have been identified from many cellular sources including monocytes, macrophages, endothelial cells, leukocytes, and tumor cells and are now thought to have pivotal roles in tumor invasion and metastases, inflammation, coagulation, and stem-cell renewal and expansion (Cocucci et al. 2009; Thery et al. 2002). Recent studies clearly showed that MVs from cultured cells and human peripheral blood contain miRNAs (Hunter et al. 2008; Skog et al. 2008; Valadi et al. 2007; Zhang et al. 2010). These findings support that MVs might serve as efficient carriers for delivery of circulating miRNAs. In this pathway, miRNAs are first loaded into small secretory vesicles or granules inside the cells, and then enter the circulation accompanying the secretion of MVs. In contrast to leakage of miRNAs from broken cells, this process is active and ATP-dependent.
21.6.2 Molecular Basis of the High Stability of Circulating MiRNAs
The remarkable stability of miRNAs in serum and plasma samples raises important and intriguing questions regarding the mechanism by which miRNAs are resistant to RNase digestion and harsh conditions. To date, the molecular basis of the high stability of circulating miRNAs remains largely unknown. Nevertheless, several hypotheses have been proposed: (i) circulating miRNAs are protected by packaging inside MVs such as exosomes and microparticles; (ii) circulating miRNAs are protected via association with other molecules (e.g. in a RNA-protein complex); and (iii) circulating miRNAs are modified (Fig. 21.3).
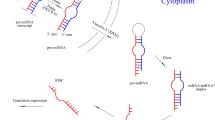
The molecular basis of the stability of circulating miRNAs. In contrast to naked miRNAs, circulating miRNAs are well preserved in harsh conditions including RNase digestion, extreme temperature and pH, extended storage, freeze-thaw cycles, etc. Several hypotheses regarding the molecular basis of the high stability of circulating miRNAs have been proposed: (i) circulating miRNAs are encapsulated in MVs such as exosomes and microparticles, and the membrane structures of MVs protect miRNAs from degradation; (ii) circulating miRNAs are embedded in RISC and are therefore stabilized via association with these miRNA-binding proteins; and (iii) circulating miRNAs are protected by modifications including methylation, adenylation, and uridylation. MiRNA, MicroRNA; MVs, Microvesicles; RISC, RNA-induced silencing complex
Potential protection of circulating miRNAs by MVs: Based on the fact that a number of circulating miRNAs in bloodstream are encapsulated in cell-derived MVs, the membrane structures of MVs may protect miRNAs from degradation. Although there is no direct evidence so far to demonstrate that this mechanism is indeed responsible for the stability of circulating miRNAs in serum and plasma, our recent study found that miRNA expression levels in MVs treated with RNase were unchanged compared to those in untreated MVs, while miRNA levels in MVs treated with both RNase and Triton X-100 (destroy the double layer membrane of MVs) significantly reduced (Zhang et al. 2010). The results suggested that circulating miRNAs are, at least in part, protected by the membrane structures of MVs.
Possible stabilization of circulating miRNAs by miRNA-binding proteins: Since miRNAs are shown to bind to proteins such as Argonaute 2 (AGO2) family in forming the RNA-induced silencing complex (RISC) (Gregory et al. 2005; Lingel et al. 2003), it is possible that these proteins in turn stabilize miRNAs in harsh conditions including RNase digestion, extreme temperature and pH, etc. However, the nature of circulating miRNA-binding proteins remains to be identified, and additional studies will be needed to explore the hypothesis.
Possible modification of circulating miRNAs: The hypothesis of circulating miRNA modification is mainly based on the comparison between circulating miRNAs and tissue/cellular miRNAs. Since the same individual miRNA is more stable in serum or plasma than that in tissue/cell, in terms of the resistance to RNase digestion, it is reasonable to suggest that these circulating miRNAs may be modified in certain ways. General modifications of miRNAs include methylation (Yu et al. 2005), adenylation (Katoh et al. 2009; Lu et al. 2009), and uridylation (Jones et al. 2009), and plant miRNA modifications play critical roles in stabilizing miRNAs and regulating miRNA functions (Lu et al. 2005; Yu et al. 2005). A recent report by Katoh et al. (2009) also found that mammalian miRNAs can be selectively stabilized by 3′ adenylation mediated by the cytoplasmic poly(A) polymerase GLD-2. However, it is currently unknown whether circulating miRNAs are methylated, adenylated, or uridylated.
21.6.3 Correlation Between Tissue MiRNAs and Circulating MiRNAs
Identification of the correlation between circulating miRNAs and tissue miRNAs also supports the hypothesis that circulating miRNAs can serve as ideal biomarkers for cancers. It is conceivable that there is a connection between tissue miRNAs and circulating miRNAs. Indeed, many miRNAs show the same trend of alteration, either increase or decrease, in the plasma/serum and tumor tissues of patients with various types of cancer. For examples, the expression of miR-25 and miR-223 was found increased in both the serum of lung cancer patients (Chen et al. 2008) and their lung tumor tissues (Volinia et al. 2006). The level of miR-155 was also found to be elevated in the tumor tissues/cells (Eis et al. 2005) and plasma of lymphoma patients (Lawrie et al. 2008). The correlation between tissue and circulating miRNAs provides evidence for the hypothesis that circulating miRNAs can reflect various aspects of human physiological status and serve as fingerprints for disease diagnosis.
21.6.4 Potential Biological Functions of Circulating MiRNAs
Although the detection of circulating miRNAs in serum, plasma, and other body fluids has been widely reported, the biological and physiological functions of circulating miRNAs are largely unknown. Recent studies have shown that miRNAs can be secreted and delivered into target cells via MVs, and that these exogenous miRNAs can regulate the expression of target genes and cellular functions in recipient cells (Pegtel et al. 2010; Skog et al. 2008; Valadi et al. 2007; Yuan et al. 2009). These findings provide a mechanism for transport and exchange of miRNAs among non-adjacent cells, and open up the possibility that miRNAs may serve as signaling molecules allowing for coordinated intercellular regulation of gene expression.
The ability of MVs to transfer miRNA raises very exciting possibilities for therapeutic uses. Cells engineered to express miRNA may be capable of delivering these small molecules to local cellular environments via MVs. These engineered cells can be encapsulated to provide sustained local delivery. Since current techniques for gene transfer use viral or synthetic agents as delivery carriers, their replacement by MVs released from autologous transplants of engineered cells will offer the advantage of a virus-free approach and make the prospects of gene therapy safer.
From this point of view, secreted miRNAs may represent a class of signaling molecules that play an important role in mediating intercellular communication. Moreover, the secretion and targeting of miRNAs among the different cells establish a highly regulated complex network under various physiological and pathological conditions. The research in secreted miRNAs will not only provide valuable information for developing biomarkers for disease diagnosis, but also shed light on our understanding of the highly complex cell communication network under various physiological and pathological conditions.
21.7 Concluding Remarks and Future Directions
Since the association of miRNAs with tumorigenesis was discovered several years ago, many attempts have been made to develop the sensitive and robust miRNA-based tests for early tumor diagnostics. Both miRNAs derived from tumor cells and affected tissues have been extensively evaluated as a diagnostic and prognostic tool to monitor cancers (Cummins et al. 2006; Iorio et al. 2005; Lu et al. 2005; Volinia et al. 2006; Yanaihara et al. 2006). Recently, we and other groups have found that human serum/plasma contained a large amount of stable miRNAs, and that the expression pattern of serum/plasma miRNA altered in reflection of various disease conditions, including cancers. The enormous potential of circulating miRNAs as a class of ideal cancer biomarkers is based on the following facts: (i) they are remarkably stable molecules, well preserved in harsh conditions, and resistant to RNase activity; (ii) their expression profiles are specifically correlated with certain type of cancer; and (iii) they are easily accessible and can be detected in a relatively non-invasive manner by various techniques.
Although circulating miRNAs have triggered much excitement in clinical and scientific communities, this field is only now emerging. Much of the work on circulating miRNAs is still in its infancy and requires further exploration. Since the first identification of serum/plasma miRNAs in 2008 (Chen et al. 2008; Gilad et al. 2008; Lawrie et al. 2008; Mitchell et al. 2008), numerous studies have shown the presence of miRNAs in circulation and their potential use as novel biomarkers for diseases and pathophysiological status, including malignancy, diabetes mellitus, pregnancy, and acute tissue injuries. However, these early studies have been limited by their small sample sizes, their lack of constant standards in quantification, and inconsistencies in methodologies (Chin and Slack 2008). Like many other novel biomarkers at their early stages of research, circulating miRNAs require extensive investigation to validate their great potential. Several areas may need to be focused on in future studies: (i) establish a simple standard assay for the quantification of circulating miRNAs in various body fluids; (ii) test the specificity and sensitivity of circulating miRNA profile-based biomarkers in a large number of samples; in particular, compare the expression of serum/plasma miRNA in different types of cancer to identify the specific biomarkers for specific cancer; and (iii) perform the perspective studies such as cohort studies to determine whether circulating miRNAs can serve as a diagnostic tool to detect cancer at its early stage.
As the functional roles of miRNAs in cancer biology are further uncovered and the methods of circulating miRNA detection and analysis are improved, circulating miRNAs will serve as novel minimally invasive or non-invasive biomarkers for various types of cancer. Their wide applicability and potential importance will probably initiate a revolution in clinical management, including detecting the early stage of tumors, estimating prognosis, predicting therapeutic efficacy, maintaining surveillance, and forecasting disease recurrence.
References
Ai J, Zhang R, Li Y, et al. Circulating microRNA-1 as a potential novel biomarker for acute myocardial infarction. Biochem Biophys Res Commun. 2010;391:73–7.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Brambilla C, Fievet F, Jeanmart M, et al. Early detection of lung cancer: role of biomarkers. Eur Respir J Suppl. 2003;39:36s–44s.
Brase JC, Johannes M, Schlomm T, et al. Circulating miRNAs are correlated with tumor progression in prostate cancer. Int J Cancer. 2011;128:608–16.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–66.
Calin GA, Dumitru CD, Shimizu M, et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA. 2002;99:15524–9.
Camps C, Buffa FM, Colella S, et al. Hsa-miR-210 Is induced by hypoxia and is an independent prognostic factor in breast cancer. Clin Cancer Res. 2008;14:1340–8.
Chen X, Ba Y, Ma L, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18:997–1006.
Chen X, Li Q, Wang J, et al. Identification and characterization of novel amphioxus microRNAs by Solexa sequencing. Genome Biol. 2009;10:R78.
Chen C, Ridzon DA, Broomer AJ, et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005;33:e179.
Chim SS, Shing TK, Hung EC, et al. Detection and characterization of placental microRNAs in maternal plasma. Clin Chem. 2008;54:482–90.
Chin LJ, Slack FJ. A truth serum for cancer-microRNAs have major potential as cancer biomarkers. Cell Res. 2008;18:983–4.
Cho WC. MicroRNAs: potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell Biol. 2010;42:1273–81.
Cocucci E, Racchetti G, Meldolesi J. Shedding microvesicles: artefacts no more. Trends Cell Biol. 2009;19:43–51.
Cummins JM, He Y, Leary RJ, et al. The colorectal microRNAome. Proc Natl Acad Sci USA. 2006;103:3687–92.
Dbouk HA, Tawil A, Nasr F, et al. Significance of CEA and VEGF as diagnostic markers of colorectal cancer in Lebanese patients. Open Clin Cancer J. 2007;1:1–5.
Dominioni L, Imperatori A, Rovera F, et al. Stage I nonsmall cell lung carcinoma: analysis of survival and implications for screening. Cancer. 2000;89:2334–44.
Eis PS, Tam W, Sun L, et al. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc Natl Acad Sci USA. 2005;102:3627–32.
Esquela-Kerscher A, Slack FJ. Oncomirs – microRNAs with a role in cancer. Nature Reviews Cancer. 2006;6:259–69.
Gerszten RE, Carr SA, Sabatine M. Integration of proteomic-based tools for improved biomarkers of myocardial injury. Clin Chem. 2010;56:194–201.
Gilad S, Meiri E, Yogev Y, et al. Serum microRNAs are promising novel biomarkers. PLoS One. 2008;3:e3148.
Gironella M, Seux M, Xie MJ, et al. Tumor protein 53-induced nuclear protein 1 expression is repressed by miR-155, and its restoration inhibits pancreatic tumor development. Proc Natl Acad Sci USA. 2007;104:16170–5.
Gregory PA, Bert AG, Paterson EL, et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol. 2008;10:593–601.
Gregory RI, Chendrimada TP, Cooch N, et al. Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell. 2005;123:631–40.
Gygi SP, Rist B, Griffin TJ, et al. Proteome analysis of low-abundance proteins using multidimensional chromatography and isotope-coded affinity tags. J Proteome Res. 2002;1:47–54.
Hanke M, Hoefig K, Merz H, et al. A robust methodology to study urine microRNA as tumor marker: microRNA-126 and microRNA-182 are related to urinary bladder cancer. Urol Oncol. 2010;28:655–61.
Hanson EK, Lubenow H, Ballantyne J. Identification of forensically relevant body fluids using a panel of differentially expressed microRNAs. Anal Biochem. 2009;387:303–14.
Harris L, Fritsche H, Mennel R, et al. American society of clinical oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J Clin Oncol. 2007;25:5287–312.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Henschke CI, Yankelevitz DF, Libby DM, et al. Survival of patients with stage I lung cancer detected on CT screening. N Engl J Med. 2006;355:1763–71.
Hu Z, Chen X, Zhao Y, et al. Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J Clin Oncol. 2010;28:1721–6.
Huang Z, Huang D, Ni S, et al. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer. 2010;127:118–26.
Huber K, Kirchheimer JC, Ermler D, et al. Determination of plasma urokinase-type plasminogen activator antigen in patients with primary liver cancer: characterization as tumor-associated antigen and comparison with alpha-fetoprotein. Cancer Res. 1992;52:1717–20.
Hunter MP, Ismail N, Zhang X, et al. Detection of microRNA expression in human peripheral blood microvesicles. PLoS One. 2008;3:e3694.
Iorio MV, Ferracin M, Liu CG, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005;65:7065–70.
Ji X, Takahashi R, Hiura Y, et al. Plasma miR-208 as a biomarker of myocardial injury. Clin Chem. 2009;55:1944–9.
Jones MR, Quinton LJ, Blahna MT, et al. Zcchc11-dependent uridylation of microRNA directs cytokine expression. Nat Cell Biol. 2009;11:1157–63.
Katoh T, Sakaguchi Y, Miyauchi K, et al. Selective stabilization of mammalian microRNAs by 3′ adenylation mediated by the cytoplasmic poly(A) polymerase GLD-2. Genes Dev. 2009;23:433–8.
Kosaka N, Izumi H, Sekine K, et al. microRNA as a new immune-regulatory agent in breast milk. Silence. 2010;1:7.
Kroh EM, Parkin RK, Mitchell PS, et al. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods. 2010;50:298–301.
Laterza OF, Lim L, Garrett-Engele PW, et al. Plasma microRNAs as sensitive and specific biomarkers of tissue injury. Clin Chem. 2009;55:1977–83.
Lawrie CH, Gal S, Dunlop HM, et al. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br J Haematol. 2008;141:672–5.
Li J, Smyth P, Flavin R, et al. Comparison of miRNA expression patterns using total RNA extracted from matched samples of formalin-fixed paraffin-embedded (FFPE) cells and snap frozen cells. BMC Biotechnol. 2007;7:36.
Lingel A, Simon B, Izaurralde E, et al. Structure and nucleic-acid binding of the Drosophila Argonaute 2 PAZ domain. Nature. 2003;426:465–9.
Liu CJ, Kao SY, Tu HF, et al. Increase of microRNA miR-31 level in plasma could be a potential marker of oral cancer. Oral Dis. 2010;16:360–4.
Lodes MJ, Caraballo M, Suciu D, et al. Detection of cancer with serum miRNAs on an oligonucleotide microarray. PLoS One. 2009;4:e6229.
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Lu S, Sun YH, Chiang VL. Adenylation of plant miRNAs. Nucleic Acids Res. 2009;37:1878–85.
Lusi EA, Passamano M, Guarascio P, et al. Innovative electrochemical approach for an early detection of microRNAs. Anal Chem. 2009;81:2819–22.
Makawita S, Diamandis EP. The bottleneck in the cancer biomarker pipeline and protein quantification through mass spectrometry-based approaches: current strategies for candidate verification. Clin Chem. 2010;56:212–22.
Mascalchi M, Belli G, Zappa M, et al. Risk-benefit analysis of X-ray exposure associated with lung cancer screening in the Italung-CT trial. AJR Am J Roentgenol. 2006;187:421–9.
Mathew LK, Simon MC. MiR-210: a sensor for hypoxic stress during tumorigenesis. Mol Cell. 2009;35:737–8.
Meng FY, Henson R, Wehbe-Janek H, et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology. 2007;133:647–58.
Mitchell PS, Parkin RK, Kroh EM, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008;105:10513–8.
Ng EK, Chong WW, Jin H, et al. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut. 2009;58:1375–81.
Oken MM, Marcus PM, Hu P, et al. Baseline chest radiograph for lung cancer detection in the randomized prostate, lung, colorectal and ovarian cancer screening trial. J Natl Cancer Inst. 2005;97:1832–9.
Park NJ, Zhou H, Elashoff D, et al. Salivary microRNA: discovery, characterization, and clinical utility for oral cancer detection. Clin Cancer Res. 2009;15:5473–7.
Patz EF Jr., Goodman PC, Bepler G. Screening for lung cancer. N Engl J Med. 2000;343:1627–33.
Pegtel DM, Cosmopoulos K, Thorley-Lawson DA, et al. Functional delivery of viral miRNAs via exosomes. Proc Natl Acad Sci USA. 2010;107:6328–33.
Peltier HJ, Latham GJ. Normalization of microRNA expression levels in quantitative RT-PCR assays: identification of suitable reference RNA targets in normal and cancerous human solid tissues. RNA. 2008;14:844–52.
Resnick KE, Alder H, Hagan JP, et al. The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol Oncol. 2009;112:55–9.
Ribas J, Ni X, Haffner M, et al. MiR-21: an androgen receptor-regulated microRNA that promotes hormone-dependent and hormone-independent prostate cancer growth. Cancer Res. 2009;69:7165–9.
Rosenfeld N, Aharonov R, Meiri E, et al. MicroRNAs accurately identify cancer tissue origin. Nat Biotechnol. 2008;26:462–9.
Rossi A, Maione P, Colantuoni G, et al. Screening for lung cancer: new horizons? Crit Rev Oncol Hematol. 2005;56:311–20.
Schneider J, Velcovsky HG, Morr H, et al. Comparison of the tumor markers tumor M2-PK, CEA, CYFRA 21-1, NSE and SCC in the diagnosis of lung cancer. Anticancer Res. 2000;20:5053–8.
Selaru FM, Olaru AV, Kan T, et al. MicroRNA-21 is overexpressed in human cholangiocarcinoma and regulates programmed cell death 4 and tissue inhibitor of metalloproteinase 3. Hepatology. 2009;49:1595–601.
Skog J, Wurdinger T, van Rijn S, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol. 2008;10:1470–6.
Sorio C, Mauri P, Pederzoli P, et al. Non-invasive cancer detection: strategies for the identification of novel cancer markers. IUBMB Life. 2006;58:193–8.
Stenman UH, Leinonen J, Zhang WM, et al. Prostate-specific antigen. Semin Cancer Biol. 1999;9:83–93.
Tanaka M, Oikawa K, Takanashi M, et al. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS One. 2009;4:e5532.
Tang F, Hajkova P, Barton SC, et al. MicroRNA expression profiling of single whole embryonic stem cells. Nucleic Acids Res. 2006;34:e9.
Taplin S, Abraham L, Barlow WE, et al. Mammography facility characteristics associated with interpretive accuracy of screening mammography. J Natl Cancer Inst. 2008;100:876–87.
Tarro G, Perna A, Esposito C. Early diagnosis of lung cancer by detection of tumor liberated protein. J Cell Physiol. 2005;203:1–5.
Thery C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2:569–79.
Tijsen AJ, Creemers EE, Moerland PD, et al. MiR423-5p as a circulating biomarker for heart failure. Circ Res. 2010;106:1035–9.
Tsujiura M, Ichikawa D, Komatsu S, et al. Circulating microRNAs in plasma of patients with gastric cancers. Br J Cancer. 2010;102:1174–9.
Valadi H, Ekstrom K, Bossios A, et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007;9:654–9.
Volinia S, Calin GA, Liu CG, et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA. 2006;103:2257–61.
Wang J, Chen J, Chang P, et al. MicroRNAs in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer Prev Res. 2009a;2:807–13.
Wang JF, Yu ML, Yu G, et al. Serum miR-146a and miR-223 as potential new biomarkers for sepsis. Biochem Biophys Res Commun. 2010a;394:184–8.
Wang K, Zhang S, Marzolf B, et al. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc Natl Acad Sci USA. 2009b;106:4402–7.
Wang GK, Zhu JQ, Zhang JT, et al. Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur Heart J. 2010b;31:659–66.
Ward DG, Suggett N, Cheng Y, et al. Identification of serum biomarkers for colon cancer by proteomic analysis. Br J Cancer. 2006;94:1898–905.
Xi Y, Nakajima G, Gavin E, et al. Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. RNA. 2007;13:1668–74.
Yanaihara N, Caplen N, Bowman E, et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006;9:189–98.
Yu B, Yang Z, Li J, et al. Methylation as a crucial step in plant microRNA biogenesis. Science. 2005;307:932–5.
Yuan A, Farber EL, Rapoport AL, et al. Transfer of microRNAs by embryonic stem cell microvesicles. PLoS One. 2009;4:e4722.
Zhang Y, Liu D, Chen X, et al. Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol Cell. 2010;39:133–44.
Zhou Z, Licklider LJ, Gygi SP, et al. Comprehensive proteomic analysis of the human spliceosome. Nature. 2002;419:182–5.
Zhu W, Qin W, Atasoy U, et al. Circulating microRNAs in breast cancer and healthy subjects. BMC Res Notes. 2009;2:89.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer Netherlands
About this chapter
Cite this chapter
Chen, X., Zhang, J., Zen, K., Zhang, CY. (2011). MicroRNAs as Blood-based Biomarkers of Cancer. In: Cho, W. (eds) MicroRNAs in Cancer Translational Research. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-0298-1_21
Download citation
DOI: https://doi.org/10.1007/978-94-007-0298-1_21
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-007-0297-4
Online ISBN: 978-94-007-0298-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)