Abstract
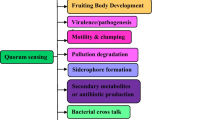
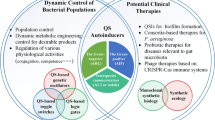
The discovery of antibiotics was a wonderful solution to provide relief to human beings from infectious diseases. However, indiscriminate usage of antibiotics turned out to be counterproductive. It was observed that patients were not getting cured in spite of the systematic use of antibiotics. In fact, microbes had developed resistance to antibiotics. This perturbation has been in operation even with antibiotics subsequently developed during the next 6–7 decades (D’Costa et al. 2006). Pharmaceutical companies are no longer interested in investing money into this business (Spellberg et al. 2004; Courvalin 2008). It obliged scientists to look for alternative drugs and new drug targets. It was realised that more than 80 % of the infectious diseases are caused by microbial pathogens, through specialised structures – biofilms. It enables bacteria to survive the lethal effect of drugs, as they “become” up to 1,000 times more resistant to antibiotics (Kalia 2013; Gui et al. 2014; Kalia et al. 2014a, b). These biofilms are developed by bacteria in a population density-dependent process called quorum sensing (QS) (Dong and Zhang 2005). Most Gram-negative bacteria operate through a QS system termed as LuxR/I-type, where acylated homoserine lactones (AHLs) acts as signals. QS signals consist of the lactone ring with varying acyl chains (Yang et al. 2012; Shang et al. 2014). QS regulates the expression of virulence factors, antibiotic production, nitrogen fixation, sporulation, conjugation, swarming, etc. (Borlee et al. 2008; Kalia and Purohit 2011; Kalia 2013; Wang et al. 2013; Zhang et al. 2013; Kalia et al. 2014a, b). These properties allow such bacteria to dominate the community structure. It is thus no surprise that the competing organisms have also developed mechanisms to interfere with the QSS and degrade these signals – a phenomenon termed as quorum quenching (QQ) (Kalia and Purohit 2011; Annapoorani et al. 2012; Bakkiyaraj et al. 2013; Kalia 2013; Agarwala et al. 2014).
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Horizontal Gene Transfer
- Quorum Sense
- Quorum Sense System
- Acylated Homoserine Lactone
- Indole Acetic Acid Production
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
The discovery of antibiotics was a wonderful solution to provide relief to human beings from infectious diseases. However, indiscriminate usage of antibiotics turned out to be counterproductive. It was observed that patients were not getting cured in spite of the systematic use of antibiotics. In fact, microbes had developed resistance to antibiotics. This perturbation has been in operation even with antibiotics subsequently developed during the next 6–7 decades (D’Costa et al. 2006). Pharmaceutical companies are no longer interested in investing money into this business (Spellberg et al. 2004; Courvalin 2008). It obliged scientists to look for alternative drugs and new drug targets. It was realised that more than 80 % of the infectious diseases are caused by microbial pathogens, through specialised structures – biofilms. It enables bacteria to survive the lethal effect of drugs, as they “become” up to 1,000 times more resistant to antibiotics (Kalia 2013; Gui et al. 2014; Kalia et al. 2014a, b). These biofilms are developed by bacteria in a population density-dependent process called quorum sensing (QS) (Dong and Zhang 2005). Most Gram-negative bacteria operate through a QS system termed as LuxR/I-type, where acylated homoserine lactones (AHLs) acts as signals. QS signals consist of the lactone ring with varying acyl chains (Yang et al. 2012; Shang et al. 2014). QS regulates the expression of virulence factors, antibiotic production, nitrogen fixation, sporulation, conjugation, swarming, etc. (Borlee et al. 2008; Kalia and Purohit 2011; Kalia 2013; Wang et al. 2013; Zhang et al. 2013; Kalia et al. 2014a, b). These properties allow such bacteria to dominate the community structure. It is thus no surprise that the competing organisms have also developed mechanisms to interfere with the QSS and degrade these signals – a phenomenon termed as quorum quenching (QQ) (Kalia and Purohit 2011; Annapoorani et al. 2012; Bakkiyaraj et al. 2013; Kalia 2013; Agarwala et al. 2014).
Signal Degradation
QS signals – AHLs – can be degraded through: (1) chemical, (2) metabolic and (3) enzymatic routes. In the chemical degradation route, alkaline pH leads to the opening of the lactone ring which thus inactivates the AHL signals produced by plant pathogen – Erwinia species (Byers et al. 2002; Yates et al. 2002). This process can be reversed at acidic pH, where the cyclisation of the lactone ring leads to the reformation of an active signal molecule. In addition to chemical inactivation, bacteria, such as Variovorax paradoxus and Pseudomonas aeruginosa PAI-A can metabolise AHLs to use them as an energy source (Leadbetter and Greenberg 2000; Huang et al. 2003). In addition to these mechanisms, QS signals can be degraded enzymatically through AHL-lactonase, AHL-acylase, oxidoreductases and lactonase-like enzymes (paraoxonases) (Tables 1 and 2), which hydrolyze either the lactone ring or the amide bond of the AHL (Dong et al. 2000; Lee et al. 2002; Lin et al. 2003).
Diversity of Organisms Possessing AHL-Lactonase
The distribution of AHL-lactonase has been reported among diverse taxa: Actinobacteria, Bacteroidetes, and Firmicutes. It is interesting to learn that members of Acidobacteria, Planctomycetes, Sphingobacteria, and Spirochaetales possess only AHL-lactonase. However, most of bacteria belonging to these taxa do not show the presence of AHL-acylase (Kalia et al. 2011). AHL-lactonase belonging to the superfamily – metallohydrolase – has been reported to be produced by Bacillus, Arthrobacter, Acidobacteria, Agrobacterium, Klebsiella, Streptomyces, Rhodococcus, Pseudomonas, Comamonas, Shewanella, etc. (Kalia et al. 2011; Chen et al. 2013). The activity of the enzyme, AHL-lactonase, is influenced by a wide range of metal ions and chelating reagents. The variation in enhancing the activity has been observed with certain metal ions such as Mg2+ and Zn2+ at high concentration of 10 mM. However, this activity was observed to decline dramatically at lower metal ion concentration of 1 mM (Chen et al. 2010). In other cases, the enhancement in enzyme activity was similar due to the presence of metal ions such as Na+, K+, Ca2+, Fe3+ and Mn2+ in the range of 1–10 mM. In contrast, quite a few ions at concentrations ranging between 0.2 and 2 mM did not have any effect on the activity of this enzyme: Ca2+, Mn2+, Co2+, Ni2+ and Cd2+ (Wang et al. 2004). There was complete inhibition in AHL-lactonase activity in the presence of heavy metals (1–10 mM): Cu2+, Cr3+, Hg2+ and Ag+ (Chen et al. 2010). Sodium do-decyl sulphate inhibited the enzyme activity whereas reagents such as ethylenediaminetetraacetic acid helped to enhance AHL-lactonase activity at 10 mM (Chen et al. 2010). In previous studies, reagents like 2,2′-bipyridine and o-phenanthroline were found to have no impact on this enzyme (Wang et al. 2004). It is important to know the determinants that regulate the broad substrate specificity of AHL-lactonases. In case of Bacillus thuringiensis AiiA, F107 residue was found to have an important role in the selective interaction preferably for longer acyl-chain substrates (Liu et al. 2013). Identification of such residues may help in the development of highly selective QSIs as potential therapeutics.
Hydrolysis of the lactone ring inactivates the QS signal and thus influences the QS-mediated characteristics (Dong et al. 2000, 2002, 2004; Lee et al. 2002; Park et al. 2003; Ulrich 2004; Dong and Zhang 2005; Liu et al. 2005; Thomas et al. 2005; Bai et al. 2008; Riaz et al. 2008; Uroz et al. 2009; Chan et al. 2010; Han et al. 2010; Deng et al. 2011; Yin et al. 2012). Among the different organisms known to produce lactonase, Bacillus has a broad substrate specificity with a preference for signals with (s)-configuration (Fuqua et al. 2001; Thomas et al. 2005). Bacillus spp. could degrade AHL signal HAI-1 of V. harveyi (Dong et al. 2002; Bai et al. 2008). Many homologues of this AHL-lactonase have been identified (Ulrich 2004) (Table 1). In a recent effort to look for diversity of AHL-lactonases, screening of 800 different bacteria revealed 42 strains of different Bacillus spp. to have aiiA gene (Huma et al. 2011). Other Bacillus species reported to express AHL-lactonase activity belonged to B. amyloliquefaciens, B. subtilis, B. mycoides and B. microestinctum (Dong et al. 2002; Pan et al. 2008; Han et al. 2010; Yin et al. 2010). Comparative genomics (in silico) study has also proved helpful in providing insights into genetic variability of genes for AHL-lactonase: B. weihenstephanensis KBAB4, B. licheniformis ATCC 14580, Geobacillus sp. WCH70, Dorea longicatena DSM 13814, Pelotomaculum thermopropionicum SI, Moorella thermoacetica ATCC 39073, Clostridium beijerinckii NCIMB 8052, Lysinibacillus sphaericus C3-41, Bacillus sp. B14905, Staphylococcus saprophyticus subsp. saprophyticus ATCC 15305, Bacillus cereus W, Bacillus subtilis, B. thuringiensis, B. thuringiensis serovar israelensis ATCC 35646, B. thuringiensis str. Al Hakam, Clostridium kluyveri DSM 555, Staphylococcus aureus RF122, S. aureus subsp. aureus MSSA476, S. aureus Mu50, S. aureus MW2, Thermosinus carboxydivorans Nor1, Caldicellulosiruptor saccharolyticus DSM 8903 and Clostridium scindens ATCC 35704 (Huma et al. 2011; Kalia et al. 2011).
Eukaryotic organisms are also known to exhibit activities similar to AHL-lactonases. Phialocephala, Ascomycetes and Meliniomyces are fungi which have lactonase activities for QS signals C6-HSL and 3OC6-HSL (Uroz and Heinon 2008). Human epithelial cells show activities for enzymes paraoxonases – PON1, PON2 and PON3 (Ng et al. 2005; Dong et al. 2007). These enzymes inactivate QS signals 3OC12-HSL produced by P. aeruginosa infecting human respiratory system (Chun et al. 2004; Hastings 2004). Human paraoxonases (PON2) hydrolyze and inactivate QS signals – 3OC6-HSL of pathogenic bacteria. PON lactonases are specific to six-member ring lactones than 5-member ring analogs (Draganov et al. 2005).
Diversity of Organisms Possessing AHL-acylase
The enzyme AHL-acylase, present in Gram-positive and Gram-negative bacteria, acts by cleaving the side chain of the signal molecule – AHL. The activity of AHL-acylase depends on acyl-chain lengths (Table 1). AHL-acylase reported from Ralstonia eutropha although quite specific, however has preference for long-chain AHLs more than 8 carbons (Lin et al. 2003). AHL-acylase from Streptomyces sp. M664 can be exploited for degrading AHLs with less than 8 carbons (Park et al. 2005). P. aeruginosa PAO1 can also degrade long-chain AHLs (Lamont and Martin 2003; Zhang and Dong 2004; Huang et al. 2006; Sio et al. 2006). The diversity of organisms showing AHL-acylase is reflected by their presence in Ralstonia solanacearum GMI1000 (Chen et al. 2009) and Shewanella sp. (Morohoshi et al. 2005). Comamonas can degrade acyl-chain lengths between 4 and 16 carbons with varying substitutions (Uroz et al. 2003, 2007). Homology to the acylase was seen in two diverse organisms such as nitrogen-fixing cyanobacterium Anabaena (Nostoc) sp. PCC7120 and QuiP of P. aeruginosa PAO1 (Romero et al. 2008). In spite of a wide diversity of taxa having organisms possessing AHL-lactonase or AHL-acylase, however, cyanobacterial members have been reported to possess only AHL-acylase (Kalia et al. 2011).
Organisms Possessing Multiple AHL-Degrading Enzymes
Rhodococcus erythropolis is unique with a wide range of QQ abilities (Table 1). R. erythropolis W2 is one of those strains which possess activities for AHL-lactonase, AHL-acylase and oxidoreductase (Uroz et al. 2005; Park et al. 2006). In silico studies have also reported the presence of organisms with multiple AHL hydrolytic enzymes: (1) Deinococcus radiodurans, (2) Hyphomonas neptunium and (3) Photorhabdus luminescens (Kalia et al. 2011).
Expression of Prokaryotic Genes for AHL-Lactonase in Different Hosts
Attempts to enhance the activity of AHL-lactonase have been made by expressing the genes in different organisms (Table 1). Most of the studies have been targeted to express aiiA of the Bacillus species (Kumar et al. 2013; Tinh et al. 2013). These heterologous expressions have proved effective in manipulating a wide range of QS-mediated characteristics in different organisms.
Expression of Bacillus aiiA
In Escherichia
Gene aiiA from Bacillus subtilis subsp. endophyticus BS1 expressed in Escherichia coli BL21 (DE3) pLysS proved effective in attenuating the soft rot symptoms caused by plant pathogen Erwinia carotovora (Pan et al. 2008). Using the same host, the expression of genes aiiA from (1) B. thuringiensis and (2) Bacillus amyloliquefaciens interfered with the pathogenicity caused by E. carotovora by inhibiting the activities of the QS signals – 3OC6-HSL, C6-HSL and C8-HSL (Lee et al. 2002; Yin et al. 2010), whereas (3) Bacillus sp. BC6 could inhibit biofilm formation abilities of Vibrio cholerae (Augustine et al. 2010). The expression of aiiA from B. thuringiensis subsp. morrisoni was weak in comparison to that of B. thuringiensis subsp. kyushuensis, as observed by their AHL degradation capacity. However, when E. coli BL21 (DE3) was used as a host, these differences were no longer evident with different QS signals: 3OC6-HSL, C6-HSL and C8-HSL (Lee et al. 2002).
In Pseudomonas
Expression of aiiA gene from Bacillus sp. A24 has been tested in a wide range of host organisms. There was a direct inhibitory impact on QS-mediated functions of P. aeruginosa PAO1, which include accumulation of QS signals (3OC12-HSL to approximately 0.10 μM), and expression of properties like swarming, motility and secretion of virulence factors (Reimmann et al. 2002). Similarly, soft rot disease of potatoes and eggplants caused by Pectobacterium carotovorum and crown gall disease of tomatoes caused by Agrobacterium tumefaciens could be prevented by expressing aiiA in Pseudomonas fluorescens (Dong et al. 2000; Molina et al. 2003). Vascular wilt of tomato plants caused by Fusarium oxysporum could be controlled when co-inoculated with Pseudomonas chlororaphis. However, this advantage was lost in the presence of AHL-lactonase producing Bacillus sp. A24 (Molina et al. 2003).
In Burkholderia
AHL-lactonase gene aiiA from Bacillus anthracis and B. cereus was effective in degrading AHL signal molecules when expressed in Burkholderia thailandensis. AHL signal molecules, such as C6- to C10-HSL, were significantly reduced from 2.4 to 300 pmol to undetectable levels. Subsequently, this gene was shown to be instrumental in retarding the growth rate such that generation time of B. thailandensis increased from 48 min in the wild-type to 243 min in the genetically modified strain. In addition, this genetically engineered strain also affected the swarming and twitching motility of this pathogen. It has been envisaged that this genetic modification can prove effective in developing vaccine against Gram-negative pathogenic bacteria (Ulrich 2004).
In Erwinia
E. carotovora SCG1 (E7-R3) expressing aiiA gene of Bacillus sp. 240 B1 could effectively inhibit the release of AI signals into the milieu. The activities of extracellular pectolytic enzymes such as pectate lyase, pectin lyase and polygalacturonase were 3–10-fold lower than the wild type. This genetic change in E. carotovora was pivotal for reducing its ability to cause disease on eggplant, potato, celery and leafy vegetables such as cabbage, Chinese cabbage, carrot and cauliflower (Dong et al. 2000).
In Serratia and Lysobacter
Endophytic bacteria, such as Serratia plymuthica, are closely associated with plant rhizosphere and phyllosphere. It has been proposed to be a potential biocontrol agent against fungal diseases. The influence of Bacillus A24 aiiA gene on QSS of S. plymuthica G3 was recorded in terms of the modified adhesion and biofilm-forming abilities. QS signals were completely degraded, which diminished its antifungal activity but augmented its indole acetic acid biosynthesis (Liu et al. 2011). Similar impact on AHL signal concentration and consequent depletion in production of chitinase and pyrrolnitrin were evident in S. plymuthica HRO-C48 expressing aiiA gene (Müller et al. 2009). aiiA gene of Bacillus has been expressed in Lysobacter enzymogenes. Here it affected pathogenicity caused by E. carotovora on Chinese cabbage (Qian et al. 2010).
In Eukaryotes
Aeromonas spp. are known to cause infections in fishes through biofilm formation and efforts are being made to control it (Chu et al. 2013; Mahanty et al. 2013). A unique approach of overproducing AHL-lactonase using eukaryotic high yielding expression system led to many advantages. Expression of aiiA in Pichia pastoris made the fish less susceptible to infection by the bacterial pathogen Aeromonas hydrophila (Chen et al. 2010; Chu et al. 2013). The secreted lactonase was found to be stable and active on a wide pH of 6.5–8.9, having thermal stability at 70 °C, and most importantly was protease resistant. This gave an advantage of having a QSI that can work strongly and efficiently on field trials, without the need to clone genes using biocontrol agents (Chen et al. 2010). Bacillus spp. with AHL-lactonase could protect Macrobrachium rosenbergii, a giant freshwater prawn, from infection caused by Vibrio harveyi infection (Nhan et al. 2010). Tobacco and potato plants modified by the introduction of aiiA gene were resistant to E. carotovora infection (Dong et al. 2001).
In Bacillus spp.
In general, aiiA from Bacillus spp. shows higher expression in a heterologous host such as E. coli. However, certain mutants of the B. thuringiensis BMB171 were found to perform better, but the mutant strains were slow in sporulation process. This property was linked to modifications in its membrane channel. This scenario can be envisaged to allow sustainable commercial production of AHL-lactonase (Zhou et al. 2006).
Expression of Homologues of aiiA in Different Hosts
In Gram-positive bacterium R. erythropolis W2 a unique AHL-lactonase encoded by gene qsdA belonging to phosphotriesterases was found. Like other classes of lactonases, it also has broad substrate specificity for acyl-chain length upto C14. All the clones (E. coli) harbouring qsdA alleles efficiently inactivated AHL signals. P. fluorescens 1855–344 expressing qsdAW2 also conferred resistance to infection against P. carotovorum PCC797. It completely abolished the QS-mediated Ti-plasmid conjugal transfer ability of A. tumefaciens expressing qsdA (Uroz et al. 2008). Similarly, another lactonase variant aidH was found in Gram-negative bacterium Ochrobactrum sp. T63. The encoded protein is a metal-dependent (Mn2+) hydrolase belonging to α/β hydrolase family. Cloning of aidH into P. fluorescens 2P24 and P. carotovorumZ3-3 curtailed biofilm formation and abolished pathogenicity in the tested plants (Mei et al. 2010). The absence of AHL-type substrate specificity makes AidH a unique AHL-lactonase to be used against QS-mediated harmful phenotypes. Similarly, expression of qsdH gene from Pseudoalteromonas byunsanensis in E. coli BL21 (DE3) was effective in inhibiting the QS-regulated functions and pathogenicity caused by E. carotovora (Huang et al. 2012). Heterologous expression of attM (a homologue of aiiA) from A. tumefaciens was able to reduce the QS-mediated pectinase enzyme activity of Azospirillum lipoferum. It also influenced the siderophore and indole acetic acid production (Boyer et al. 2008). An attM paralogous gene (aiiB) was also found in A. tumefaciens, having potent activity against AHLs that can be exploited (Carlier et al. 2003).
Recent developments in metagenomic approaches to look for AHL-lactonase from uncultured bacteria are expected to add to the limited diversity of this enzyme (Williamson et al. 2005). Metagenome of a bacterial community from soil is another genetic resource of great potential to be exploited. Screening of various metagenomic clones allowed the identification of genes such as qlcA, bpiB01 and bpiB04, which had very low or no similarities to any known AHL-lactonases. Expression of bpiB01 in P. aeruginosa PAO1 inhibited its biofilm-forming ability (Riaz et al. 2008; Schipper et al. 2009).
Hetreologous Expression of Eukaryotic Lactonase
Expression of genes for three mouse paraoxonases (PONs) in mammalian cells led to the degradation of AHL in a manner which was quite similar to that of lactonases (Yang et al. 2005). Expression of human paraoxonase in Drosophila melanogaster allowed the organism to survive the onslaught of the lethal action of P. aeruginosa infection (Stoltz et al. 2008).
Heterologous Expression of AHL-Acylase Gene from Ralstonia
Cloning and expression of gene aiiD from Ralstonia sp. into E. coli was very effective in abolishing the 3 QS signals: 3OC8-HSL, 3OC10-HSL and 3OC12-HSL, while its overexpression within Brucella melitensis affected the QS-mediated process and clumping phenotypes (Godefroid et al. 2010) (Table 2). This inactivation was quite rapid as it was observed within 3 h of incubation. Wild-type P. aeruginosa PAO1 possessing plasmid pUCM9-PAO1 showed accumulation of QS signal in normal concentrations. In contrast, introduction of pUCaiiD encoding AHL-acylase gene into P. aeruginosa PAO1 affected a wide range of QS regulated expressions. The pathogenic bacterium could not accumulate AHL signals: C4-HSL and 3OC12-HSL. The genetically engineered P. aeruginosa was not able to produce elastase and pyocyanin in normal quantities, and its ability to swarm was significantly restricted. The pathogen could no longer paralyse the nematode, Caenorhabditis elegans, which had more than 80 % survival rate (Lin et al. 2003). Subsequently, an aculeacin A acylase was reported from Ralstonia solanacearum GMI1000. The aac gene had 83 % identity with acylase gene aiiD, but no significant resemblance with lactonase gene aiiA sequence (Chen et al. 2009). This enzyme is active against AHLs having side chains more than 6 carbons. Expression of aac gene in C. violaceum CV026 could inhibit AHL-mediated activities such as production of violacein and chitinase enzyme (Chen et al. 2009).
Over Expression of AHL-Acylase Gene from Pseudomonas
P. aeruginosa genes coding for AHL-acylase have the potential to be exploited for developing novel therapies against the infections caused by this pathogen. Over expression of gene PA2385 also known as qsc112 and pvdQ (Whiteley et al. 1999; Lamont and Martin 2003) has differential behaviour against AHL signals. It could not hydrolyse QS signals: C10-HSL, C12-HSL and C14-HSL (Sio et al. 2006). This gene was first cloned into the P. aeruginosa-E. coli shuttle vector pME6032 and electroporated in P. aeruginosa PAO1. Introduction of gene PA2385 absolutely extripated 3OC12-HSL, but there was no impact on the level of C4-HSL. Over expression of PA2385 abolished activities of enzymes, LasB, and also minimised the expression of lecA gene. It, however, did not affect production of pyoverdin (Sio et al. 2006).
A novel AHL-acylase coded by aiiC (all3924) was reported from filamentous nitrogen fixing cyanobacterium, Anabaena sp. PCC7120. It showed homology to quiP of P. aeruginosa PAO1. This gene was also cloned into shuttle vector pME6032 and expressed in E. coli. AiiC has broad specificity with respect to the acyl-chain length, but did not degrade short chain AHLs (Romero et al. 2008). Although P. aeruginosa PAO1 has been established to possess pvdQ and quiP, however, two more genes, pa1893 and pa0305 belonging to the Ntn hydrolase superfamily, have been predicted to encode for penicillin acylase (Wahjudi et al. 2011). Biosensor assays showed their ability to degrade C6 to C14-HSLs. Pa0305 was found to be 56 times more efficient in degrading 3OC12-HSL than C8-HSL. The enzyme was effective in killing C. elegans as well (Table 2).
Gene ahlM from Streptomyces sp. M664 was able to express and degrade AHLs in Streptomyces lividans (Park et al. 2005). AhlM enzyme had higher deacylation activity towards long acyl-chain AHLs. It could also degrade penicillin by the same mechanism. It had high activity, even at low concentration of 2 μg/ml. It could significantly affect the production level of virulence factors such as elastase, protease and LasA of P. aeruginosa (Park et al. 2005).
Horizontal Gene Transfer
Screening for AHL-degrading proteins has revealed the presence of aiiA gene in a large number of organisms. These organisms belong to taxonomically diverse genera. AHL-lactonases from different sources may share homology among themselves. Such genetic similarity can be explained on the basis of horizontal gene transfer (HGT). The occurrence of HGT among bacterial kingdom and its impact of genetic and phenotypic changes has been well reported (Lal et al. 2008). The unique AHL-lactonase of R. erythropolis W2 seems to be a case of HGT (Uroz et al. 2008).
In silico studies have also highlighted the occurrence of HGT involving AHL-degrading enzymes (Kalia et al. 2011). Genetic variability in AHL-lactonase coding gene aiiA was reported on the basis of their phylogenetic relationships and the restriction digestion patterns (Huma et al. 2011; Kalia et al. 2011). Taxonomically diverse organisms were found to show discrepancies in their phylogenetic relationships for aiiA. Genes for AHL-lactonase from Firmicutes, Moorella thermoacetica ATCC 39073, and β-Proteobacteria,Burkholderia graminis C4D1M, were very close to each other with a bootstrap value, BV of 1000. Similar correlations with high BVs were also recorded among: (i) Actinobacteria such as Mycobacterium sp. MCS and Rubrobacter xylanophilus DSM9941 on one hand and members of α-Proteobacteria (Granulibacter bethesdensis CGDNIH1, Acidiphilium cryptum JF-5 and A. tumefaciens) on the other and (ii) also between D. radiodurans R1 (Deinococcus-Thermus) and Xylella fastidiosa (γ-Proteobacteria).
Taxonomic discrepancies in the phylogenetic tree of the gene for AHL-acylase were reported between different groups: (1) D. radiodurans R1 and Ralstonia spp. and (2) δ-Proteobacteria, Plesiocystis pacifica SIR-1, and Acidobacteria, Solibacter usitatus. This emphasises that in nature, organisms live in symbiotic relationships and share these genes through HGTs for evolving better survival strategies (Kalia et al. 2011).
Conclusion
Heterologous expression of QS signal degrading enzymes can be exploited for producing them on a commercial scale. The immediate application is to use them as potential drugs against biofilm-forming bacteria. Their usage in a range of biotechnological applications has been discussed in a subsequent chapter.
Abbreviations
- AHL:
-
Acyl homoserine lactone
- C4-HSL:
-
N-butanoyl-L-HSL
- C6-HSL:
-
N-hexanoyl HSL
- C8-HSL:
-
N-octanoyl HSL
- C10-HSL:
-
N-decanoyl HSL
- C12-HSL:
-
N-dodecanoyl HSL
- C14-HSL:
-
N-tetradecanoyl-HSL
- 3OC6-HSL:
-
3-oxo-N-hexanoyl-HSL
- 3OC8-HSL:
-
3-oxo-N-octanoyl-HSL
- 3OC10-HSL:
-
3-oxo-N-decanoyl-HSL
- 3OC12-HSL:
-
3-oxo-N-dodecanoyl- HSL
- HSL:
-
Homoserine lactone
- QS:
-
Quorum sensing
- QSS:
-
Quorum sensing systems
- QQ:
-
Quorum quenching
References
Agarwala M, Choudhury B, Yadav RNS (2014) Comparative study of antibiofilm activity of copper oxide and iron oxide nanoparticles against multidrug resistant biofilm forming uropathogens. Indian J Microbiol. doi:10.1007/s12088-014-0462-z
Annapoorani A, Jabbar AKKA, Musthafa SKS, Pandian SK, Ravi AV (2012) Inhibition of quorum sensing mediated virulence factors production in urinary pathogen Serratia marcescens PS1 by marine sponges. Indian J Microbiol 52:160–166. doi:10.1007/s12088-012-0272-0
Augustine N, Kumar P, Thomas S (2010) Inhibition of Vibrio cholerae biofilm by AiiA enzyme produced from Bacillus spp. Arch Microbiol 192:1019–1022. doi:10.1007/s00203-010-0633-1
Bai F, Han Y, Chen J, Zhang X-H (2008) Disruption of quorum sensing in Vibrio harveyi by the AiiA protein of Bacillus thuringiensis. Aquaculture 274:36–40. doi:10.1016/j.aquaculture.2007.11.024
Bakkiyaraj D, Sivasankar C, Pandian SK (2013) Anti-pathogenic potential of coral associated bacteria isolated from Gulf of Mannar against Pseudomonas aeruginosa. Indian J Microbiol 53:111–113. doi:10.1007/s12088-012-0342-3
Bijtenhoorn P, Mayerhofer H, Müller-Dieckmann J, Utpatel C, Schipper C, Hornung C, Szesny M, Grond S, Thürmer A, Brzuszkiewicz E, Daniel R, Dierking K, Schulenburg H, Streit WR (2011) A novel metagenomic short-chain dehydrogenase/reductase attenuates Pseudomonas aeruginosa biofilm formation and virulence on Caenorhabditis elegans. PLoS One 6:e26278. doi:10.1371/journal.pone.0026278
Borlee BR, Geske GD, Robinson CJ, Blackwell HE, Handelsman J (2008) Quorum-sensing signals in the microbial community of the cabbage white butterfly larval midgut. ISME J 2:1101–1111. doi:10.1038/ismej.2008.70
Boyer M, Bally R, Perrotto S, Chaintreuil C, Wisniewski-Dyé F (2008) A quorum-quenching approach to identify quorum-sensing-regulated functions in Azospirillum lipoferum. Res Microbiol 159:699–708. doi:10.1016/j.resmic.2008.08.003
Byers JT, Lucas C, Salmond GP, Welch M (2002) Nonenzymatic turnover of an Erwinia carotovora quorum–sensing signaling molecule. J Bacteriol 184:1163–1171. doi:10.1128/jb.184.4.1163-1171.2002
Carlier A, Uroz S, Smadja B, Fray R, Latour X, Dessaux Y, Faure D (2003) The Ti plasmid of Agrobacterium tumefaciens harbors an attM-paralogous gene, aiiB, also encoding N-acyl homoserine lactonase activity. Appl Environ Microbiol 69:4989–4993. doi:10.1128/AEM.69.8.4989-4993.2003
Chan KG, Wong CS, Yin WF, Sam CK, Koh CL (2010) Rapid degradation of N-3-oxo-acylhomoserine lactones by a Bacillus cereus isolate from Malaysian rainforest soil. Antonie Van Leeuwenhoek 98:299–305. doi:10.1007/s10482-010-9438-0
Chen C-N, Chen C-J, Liao C-T, Lee C-Y (2009) A probable aculeacin A acylase from the Ralstonia solanacearum GMI1000 is N-acyl-homoserine lactone acylase with quorum-quenching activity. BMC Microbiol 9:89. doi:10.1186/1471-2180-9-89
Chen R, Zhou Z, Cao Y, Yao B (2010) High yield expression of an AHL-lactonase from Bacillus sp. B546 in Pichia pastoris and its application to reduce Aeromonas hydrophila mortality in aquaculture. Microb Cell Fact 9:39
Chen F, Gao Y, Chen X, Yu Z, Li X (2013) Quorum quenching enzymes and their application in degrading signal molecules to block quorum sensing-dependent infection. Int J Mol Sci 14:17477–17500. doi:10.3390/ijms140917477
Chu W, Liu Y, Jiang Y, Zhu W, Zhuang X (2013) Production of N-acyl homoserine lactones and virulence factors of waterborne Aeromonas hydrophila. Indian J Microbiol 53:264–268. doi:10.1007/s12088-013-0381-4
Chun CK, Ozer EA, Welsh MJ, Zabner J, Greenberg EP (2004) Inactivation of a Pseudomonas aeruginosa quorum-sensing signal by human airway epithelia. Proc Natl Acad Sci USA 101:3587–3590. doi:10.1073/pnas.0308750101
Courvalin P (2008) Predictable and unpredictable evolution of antibiotic resistance. J Intern Med 264:4–16. doi:10.1111/j.1365-2796.2008.01940.x
Czajkowski R, Krzyżanowska D, Karczewska J, Atkinson S, Przysowa J, Lojkowska E, Williams P, Jafra S (2011) Inactivation of AHLs by Ochrobactrum sp. A44 depends on the activity of a novel class of AHL acylase. Environ Microbiol Rep 3:59–68. doi:10.1111/j.1758-2229.2010.00188.x
D’Costa VM, McGrann KM, Hughes DW, Wright GD (2006) Sampling the antibiotic resistome. Science 311:374–377. doi:10.1126/science.1120800
Deng Y, Zhu Y, Wang P, Zhu L, Zheng J, Li R, Ruan L, Peng D, Sun M (2011) Complete genome sequence of Bacillus subtilis BSn5, an endophytic bacterium of Amorphophallus konjac with antimicrobial activity for the plant pathogen Erwinia carotovora subsp. carotovora. J Bacteriol 193:2070–2071. doi:10.1128/JB.00129-11
Dong YH, Zhang LH (2005) Quorum sensing and quorum quenching enzymes. J Microbiol 43:101–109
Dong YH, Xu JL, Li XZ, Zhang LH (2000) AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora. Proc Natl Acad Sci USA 97:3526–3531. doi:10.1073/pnas.97.7.3526
Dong YH, Wang LH, Xu JL, Zhang HB, Zhang XF, Zhang LH (2001) Quenching quorum-sensing dependent bacterial infection by an N-acyl homoserine lactonase. Nature 411:813–817. doi:10.1038/35081101
Dong YH, Gusti AR, Zhang Q, Xu JL, Zhang LH (2002) Identification of quorum-quenching N-acyl homoserine lactonases from Bacillus species. Appl Environ Microbiol 68:1754–1759. doi:10.1128/AEM.68.4.1754-1759.2002
Dong YH, Zhang XF, Xu JL, Zhang LH (2004) Insecticidal Bacillus thuringiensis silences Erwinia carotovora virulence by a new form of microbial antagonism, signal interference. Appl Environ Microbiol 70:954–960. doi:10.1128/AEM.70.2.954-960.2004
Dong YH, Wang LH, Zhang LH (2007) Quorum-quenching microbial infections: mechanisms and implications. Phil Trans R Soc Lond B Biol Sci 362:1201–1211. doi:10.1098/rstb.2007.2045
Draganov DI, Teiber JF, Speelman A, Osawa Y, Sunahara R, LaDu BN (2005) Human paraoxonases (PON1, PON2 and PON3) are lactonases with overlapping and distinct substrate specificities. J Lipid Res 46:1239–1247. doi:10.1194/jlr.M400511-JLR200
Fuqua C, Parsek MR, Greenberg EP (2001) Regulation of gene expression by cell-to-cell communication: acyl-homoserine lactone quorum sensing. Annu Rev Genet 35:439–468. doi:10.1146/annurev.genet.35.102401.090913
Godefroid M, Svensson MV, Cambier P, Uzureau S, Mirabella A, De Bolle X, Van Cutsem P, Widmalm G, Letesson JJ (2010) Brucella melitensis 16M produces a mannan and other extracellular matrix components typical of a biofilm. FEMS Immunol Med Microbiol 59:364–377. doi:10.1111/j.1574-695X.2010.00689.x
Gui Z, Wang H, Ding T, Zhu W, Zhuang X, Chu W (2014) Azithromycin reduces the production of α-hemolysin and biofilm formation in Staphylococcus aureus. Indian J Microbiol 54:114–117. doi:10.1007/s12088-013-0438-4
Han Y, Chen F, Li N, Zhu B, Li X (2010) Bacillus marcorestinctum sp. nov., a novel soil acylhomoserine lactone quorum-sensing signal quenching bacterium. Int J Mol Sci 11:507–520. doi:10.3390/ijms11020507
Hastings JW (2004) Bacterial quorum–sensing signals are inactivated by mammalian cells. Proc Natl Acad Sci U S A 101:3993–3994. doi:10.1073/pnas.0400874101
Huang JJ, Han J-I, Zhang L-H, Leadbetter JR (2003) Utilization of acylhomoserine lactone quorum signals for growth by a soil pseudomonad and Pseudomonas aeruginosa PAO1. Appl Environ Microbiol 69:5941–5949. doi:10.1128/AEM.69.10.5941-5949.2003
Huang JJ, Petersen A, Whiteley M, Leadbetter JR (2006) Identification of QuiP, the product of gene PA1032, as the second acyl-homoserine lactone acylase of Pseudomonas aeruginosa PAO1. Appl Environ Microbiol 72:1190–1197. doi:10.1128/AEM.72.2.1190-1197.2006
Huang W, Lin Y, Yi S, Liu P, Shen J, Shao Z, Liu Z (2012) QsdH, a novel AHL lactonase in the RND-type inner membrane of marine Pseudoalteromonas byunsanensis strain 1A01261. PLoS One 7:e46587. doi:10.1371/journal.pone.0046587.t001
Huma N, Shankar P, Kushwah J, Bhushan A, Joshi J, Mukherjee T, Raju SC, Purohit HJ, Kalia VC (2011) Diversity and polymorphism in AHL-lactonase gene (aiiA) of Bacillus. J Microbiol Biotechnol 21:1001–1011. doi:10.4014/jmb.1105.05056
Kalia VC (2013) Quorum sensing inhibitors: an overview. Biotechnol Adv 31:224–245. doi:10.1016/j.biotechadv.2012.10.004
Kalia VC, Purohit HJ (2011) Quenching the quorum sensing system: potential antibacterial drug targets. Crit Rev Microbiol 37:121–140. doi:10.3109/1040841X.2010.532479
Kalia VC, Raju SC, Purohit HJ (2011) Genomic analysis reveals versatile organisms for quorum quenching enzymes: acyl-homoserine lactone-acylase and -lactonase. Open Microbiol J 5:1–13. doi:10.2174/187428580110501000
Kalia VC, Kumar P, Pandian SK, Sharma P (2014a) Biofouling control by quorum quenching. In: Kim SK (ed) Handbook of marine biotechnology. Springer
Kalia VC, Wood TK, Kumar P (2014b) Evolution of resistance to quorum-sensing inhibitors. Microb Ecol 68:13–23. doi:10.1007/s00248-013-0316-y
Kumar P, Patel SKS, Lee JK, Kalia VC (2013) Extending the limits of Bacillus for novel biotechnological applications. Biotechnol Adv 31:1543–1561. doi:10.1016/j.biotechadv.2013.08.007
Lal S, Cheema S, Kalia VC (2008) Phylogeny vs genome reshuffling: horizontal gene transfer. Indian J Microbiol 48:228–242. doi:10.1007/s12088-008-0034-1
Lamont IL, Martin LW (2003) Identification and characterization of novel pyoverdine synthesis genes in Pseudomonas aeruginosa. Microbiology 149:833–842. doi:10.1099/mic.0.26085-0
Leadbetter JR, Greenberg EP (2000) Metabolism of acylhomoserine lactone quorum–sensing signals by Variovorax paradoxus. J Bacteriol 182:6921–6926. doi:10.1128/JB.182.24.6921-6926.2000
Lee SJ, Park SY, Lee JJ, Yum DY, Koo BT, Lee JK (2002) Genes encoding the N–acyl homoserine lactone–degrading enzyme are widespread in many subspecies of Bacillus thuringiensis. Appl Environ Microbiol 68:3919–3924. doi:10.1128/AEM.68.8.3919-3924.2002
Lin YH, Xu JL, Hu J, Wang LH, Ong SL, Leadbetter JR, Zhang LH (2003) Acyl–homoserine lactone acylase from Ralstonia strain XJ12B represents a novel and potent class of quorum–quenching enzymes. Mol Microbiol 47:849–860. doi:10.1046/j.1365-2958.2003.03351.x
Liu D, Lepore BW, Petsko GA, Thomas PW, Stone EM, Fast W, Ringe D (2005) Three–dimensional structure of the quorum quenching N-acyl homoserine lactone hydrolase from Bacillus thuringiensis. Proc Natl Acad Sci USA 102:11882–11887. doi:10.1073/pnas.0505255102
Liu X, Jia J, Popat R, Ortori CA, Li J, Diggle SP, Gao K, Cámara M (2011) Characterization of two quorum sensing systems in the endophytic Serratia plymuthica strain G3: differential control of motility and biofilm formation according to life style. BMC Microbiol 11:26. doi:10.1186/1471-2180-11-26
Liu CF, Liu D, Momb J, Thomas PW, Lajoie A, Petsko GA, Fast W, Ringe D (2013) A distal phenylalanine clamp in a hydrophobic channel controls the substrate specificity in the quorum-quenching metallo-γ-lactonase (AiiA) from Bacillus thuringiensis. Biochemistry 52:1603–1610. doi:10.1021/bi400050j
Mahanty A, Mishra S, Bosu R, Maurya UK, Netam SP, Sarkar B (2013) Phytoextracts-synthesized silver nanoparticles inhibit bacterial fish pathogen Aeromonas hydrophila. Indian J Microbiol 53:438–446. doi:10.1007/s12088-013-0409-9
Mei GY, Yan XX, Turak A, Luo ZQ, Zhang LQ (2010) AidH, an alpha/beta-hydrolase fold family member from an Ochrobactrum sp. strain, is a novel N-acylhomoserine lactonase. Appl Environ Microbiol 76:4933–4942. doi:10.1128/AEM.00477-10
Molina L, Constantinescu F, Michel L, Reimmann C, Duffy B, Défago G (2003) Degradation of pathogen quorum-sensing molecules by soil bacteria: a preventive and curative biological control mechanism. FEMS Microbiol Ecol 45:71–81. doi:10.1016/S0168-6496(03)00125-9
Morohoshi T, Ebata A, Nakazawa S, Kato N, Ikeda T (2005) N-acyl homoserine lactone–producing or –degrading bacteria isolated from the intestinal microbial flora of Ayu fish (Plecoglossus altivelis). Microb Environ 20:264–268. doi:10.1264/jsme2.20.264
Morohoshi T, Nakazawa S, Ebata A, Kato N, Ikeda T (2008) Identification and characterization of N-acylhomoserine lactone-acylase from the fish intestinal Shewanella sp. strain MIB015. Biosci Biotechnol Biochem 72:1887–1893. doi:10.1271/bbb.80139
Müller H, Westendorf C, Leitner E, Chernin L, Riedel K, Schmidt S, Eberl L, Berg G (2009) Quorum‐sensing effects in the antagonistic rhizosphere bacterium Serratia plymuthica HRO‐C48. FEMS Microbiol Ecol 67:468–478. doi:10.1111/j.1574-6941.2008.00635.x
Ng CJ, Shih DM, Hama SY, Villa N, Navab M, Reddy ST (2005) The paraoxonase gene family and atherosclerosis. Free Radical Biol Med 38:153–163. doi:10.1016/j.freeradbiomed.2004.09.035
Nhan DT, Cam DTV, Wille M, Defoirdt T, Bossier P, Sorgeloos P (2010) Quorum quenching bacteria protect Macrobrachium rosenbergii larvae from Vibrio harveyi infection. J Appl Microbiol 109:1007–1016. doi:10.1111/j.1365-2672.2010.04728.x
Pan J, Huang T, Yao F, Huang Z, Powell CA, Qiu S, Guan X (2008) Expression and characterization of aiiA gene from Bacillus subtilis BS-1. Microbiol Res 163:711–716. doi:10.1016/j.micres.2007.12.002
Papaioannou E, Wahjudi M, Nadal-Jimenez P, Koch G, Setroikromo R, Quax WJ (2009) Quorum-quenching acylase reduces the virulence of Pseudomonas aeruginosa in a Caenorhabditis elegans infection model. Antimicrob Agents Chemother 53:4891–4897. doi:10.1128/AAC.00380-09
Park SY, Lee SJ, Oh TK, Oh JW, Koo BT, Yum DY, Lee JK (2003) AhlD, an N–acylhomoserine lactonase in Arthrobacter sp., predicted homologues in other bacteria. Microbiology 149:1541–1550. doi:10.1099/mic.0.26269-0
Park SY, Kang HO, Jang HS, Lee JK, Koo BT, Yum DY (2005) Identification of extracellular N–acylhomoserine lactone acylase from a Streptomyces sp. and its application to quorum quenching. Appl Environ Microbiol 71:2632–2641. doi:10.1128/AEM.71.5.2632-2641.2005
Park SY, Hwang BJ, Shin MH, Kim JA, Kim HK, Lee JK (2006) N-acyl homoserine lactone producing Rhodococcus sp. with different AHL-degrading activities. FEMS Microbiol Lett 261:102–108. doi:10.1111/j.1574-6968.2006.00336.x
Qian GL, Fan JQ, Chen DF, Kang YJ, Han B, Hu BS, Liu FQ (2010) Reducing Pectobacterium virulence by expression of an N-acyl homoserine lactonase gene P lpp -aiiA in Lysobacter enzymogenes strain OH11. Biol Control 52:17–23. doi:10.1016/j.biocontrol.2009.05.007
Reimmann C, Ginet N, Michel L, Keel C, Michaux P, Krishnapillai V, Zala M, Heurlier K, Triandafillu K, Harms H, Défago G, Haas D (2002) Genetically programmed autoinducer destruction reduces virulence gene expression and swarming motility in Pseudomonas aeruginosa PAO1. Microbiology 148:923–932
Riaz K, Elmerich C, Moreira D, Raffoux A, Dessaux Y, Faure D (2008) A metagenomic analysis of soil bacteria extends the diversity of quorum–quenching lactonases. Environ Microbiol 10:560–570. doi:10.1111/j.1462-2920.2007.01475.x
Romero M, Diggle SP, Heeb S, Cámara C, Otero A (2008) Quorum quenching activity in Anabaena sp. PCC7120: identification of AiiC, a novel AHL–acylase. FEMS Microbiol Lett 280:73–80. doi:10.1111/j.1574-6968.2007.01046.x
Schipper C, Hornung C, Bijtenhoorn P, Quitschau M, Grond S, Streit WR (2009) Metagenome-derived clones encoding two novel lactonase family proteins involved in biofilm inhibition in Pseudomonas aeruginosa. Appl Environ Microbiol 75:224–233. doi:10.1128/AEM.01389-08
Shang Z, Wang H, Zhou S, Chu W (2014) Characterization of N-Acyl-homoserine lactones (AHLs)-deficient clinical isolates of Pseudomonas aeruginosa. Indian J Microbiol 54:158–162. doi:10.1007/s12088-014-0449-9
Shepherd RW, Lindow SE (2009) Two dissimilar N-acyl-homoserine lactone acylases of Pseudomonas syringae influence colony and biofilm morphology. Appl Environ Microbiol 75:45–53. doi:10.1128/AEM.01723-08
Sio CF, Otten LG, Cool RH, Diggle SP, Braun PG, Bos R, Daykin M, Cámara M, Williams P, Quax WJ (2006) Quorum Quenching by an N-Acyl-Homoserine lactone acylase from Pseudomonas aeruginosa PAO1. Infect Immun 74:1673–1682. doi:10.1128/IAI.74.3.1673-1682.2006
Spellberg B, Powers JH, Brass EP, Miller LG, Edwards JE Jr (2004) Trends in antimicrobial drug development: implications for the future. Clin Infect Dis 38:1279–1286. doi:10.1086/420937
Stoltz DA, Ozer EA, Taft PJ, Barry M, Liu L, Kiss PJ, Moninger TO, Parsek MR, Zabner J (2008) Drosophila are protected from Pseudomonas aeruginosa lethality by transgenic expression of paraoxonase-1. J Clin Invest 118:3123–3131. doi:10.1172/JCI35147
Thomas PW, Stone EM, Costello AL, Tierney DL, Fast W (2005) The quorum-quenching lactonase from Bacillus thuringiensis is a metalloprotein. Biochemistry 44:7559–7569. doi:10.1021/bi050050m
Tinh NTN, Dung NV, Trung CT (2013) In vitro characterization of a recombinant AHL-Lactonase from Bacillus cereus isolated from a striped catfish (Pangasianodon hypophthalmus) pond. Indian J Microbiol 53:485–487. doi:10.1007/s12088-013-0415-y
Ulrich RL (2004) Quorum quenching: enzymatic disruption of N-acylhomoserine lactone-mediated bacterial communication in Burkholderia thailandensis. Appl Environ Microbiol 70:6173–6180. doi:10.1128/AEM.70.10.6173-6180.2004
Uroz S, Heinon J (2008) Degradation of N-acyl homoserine lactone quorum sensing signal molecules by forest root associated fungi. FEMS Microbiol Ecol 65:271–278. doi:10.1111/j.1574-6941.2008.00477.x
Uroz S, DAngelo–Picard C, Carlier A, Elasri M, Sicot C, Petit A, Oger P, Faure D, Dessaux Y (2003) Novel bacteria degrading N-acylhomoserine lactones and their use as quenchers of quorum-sensing-regulated functions of plant-pathogenic bacteria. Microbiology 149:1981–1989. doi:10.1099/mic.0.26375-0
Uroz S, Chhabra SR, Cámara M, Wiliams P, Oger PM, Dessaux Y (2005) N-Acylhomoserine lactone quorum–sensing molecules are modified and degraded by Rhodococcus erythropolis W2 by both amidolytic and novel oxidoreductase activities. Microbiology 151:3313–3322. doi:10.1099/mic.0.27961-0
Uroz S, Oger P, Chhabra SR, Cámara M, Williams P, Dessaux Y (2007) N-acyl-homoserine lactones are degraded via an amidolytic activity in Comamonas sp. strain D1. Arch Microbiol 187:249–256. doi:10.1007/s00203-006-0186-5
Uroz S, Oger PM, Chapelle E, Adeline MT, Faure D, Dessaux Y (2008) A Rhodococcus qsdA-encoded enzyme defines a novel class of large-spectrum quorum-quenching lactonases. Appl Environ Microbiol 74:1357–1366. doi:10.1128/AEM.02014-07
Uroz S, Dessaux Y, Oger P (2009) Quorum sensing and quorum quenching: the Yin and Yang of bacterial communication. ChemBiochem 10:205–216. doi:10.1002/cbic.200800521
Wahjudi M, Papaioannou E, Hendrawati O, van Assen AH, van Merkerk R, Cool RH, Poelarends GJ, Quax WJ (2011) PA0305 of Pseudomonas aeruginosa is a quorum quenching acylhomoserine lactone acylase belonging to the Ntn hydrolase superfamily. Microbiology 157:2042–2055. doi:10.1099/mic.0.043935-0
Wang LH, Weng LX, Dong YH, Zhang LH (2004) Specificity and enzyme kinetics of the quorum-quenching N-acyl homoserine lactone lactonase (AHL-lactonase). J Biol Chem 279:13645–13651. doi:10.1074/jbc.M311194200
Wang WZ, Morohoshi T, Someya N, Ikeda T (2012) AidC, a novel N-acylhomoserine lactonase from the potato root-associated Cytophaga-Flavobacteria-Bacteroides (CFB) group bacterium Chryseobacterium sp. strain StRB126. Appl Environ Microbiol 78:7985–7992. doi:10.1128/AEM.02188-12
Wang H, Tu F, Gui Z, Lu X, Chu W (2013) Antibiotic resistance profiles and quorum sensing-dependent virulence factors in clinical isolates of Pseudomonas aeruginosa. Indian J Microbiol 53:163–167. doi:10.1007/s12088-013-0370-7
Whiteley M, Lee KM, Greenberg EP (1999) Identification of genes controlled by quorum sensing in Pseudomonas aeruginosa. Proc Natl Acad Sci USA 96:13904–13909. doi:10.1073/pnas.96.24.13904
Williamson LL, Borlee BR, Schloss PD, Guan C, Allen HK, Handelsman J (2005) Intracellular screen to identify metagenomic clones that induce or inhibit a quorum-sensing biosensor. Appl Environ Microbiol 71:6335–6344. doi:10.1128/AEM.71.10.6335-6344.2005
Yang F, Wang LH, Wang J, Dong YH, Hu JY, Zhang LH (2005) Quorum quenching enzyme activity is widely conserved in the sera of mammalian species. FEBS Lett 579:3713–3717. doi:10.1016/j.febslet.2005.05.060
Yang Q, Han Y, Tinh NTN, Hien NT, Bossier P (2012) Detection of quorum sensing signal molecules in Edwardsiella ictaluri Ei-151. Indian J Microbiol 52:581–586. doi:10.1007/s12088-012-0312-9
Yates EA, Philipp B, Buckley C, Atkinson S, Chhabra SR, Sockett RE, Goldner M, Dessaux Y, Cámara M, Smith H, Williams P (2002) N–Acylhomoserine lactones undergo lactonolysis in a pH–, temperature–, acyl chain length–dependent manner during growth of Yersinia pseudotuberculosis and Pseudomonas aeruginosa. Infect Immun 70:5635–5646. doi:10.1128/IAI.70.10.5635-5646.2002
Yin XT, Xu L, Fan SS, Xu LN, Li DC, Liu ZY (2010) Isolation and characterization of an AHL lactonase gene from Bacillus amyloliquefaciens. World J Microbiol Biotechnol 26:1361–1367. doi:10.1007/s11274-010-0308-8
Yin WF, Tung HJ, Sam CK, Koh CL, Chan KG (2012) Quorum quenching Bacillus sonorensis isolated from soya sauce fermentation brine. Sensors 12:4065–4073. doi:10.3390/s120404065
Zhang L-H, Dong Y-H (2004) Quorum sensing and signal interference: diverse implications. Mol Microbiol 53:1563–1571. doi:10.1111/j.1365-2958.2004.04234.x
Zhang Y, Zhang Y, Yang Y, Wang L, Weng L (2013) Identification of a Pseudomonas sp. that inhibits RHL system of quorum sensing. Indian J Microbiol 53:28–35. doi:10.1007/s12088-012-0340-5
Zhou Y, Ye WX, Zhou Y, Zhu CG, Sun M, Yu ZN (2006) Ethanol tolerance, yield of melanin, swarming motility and growth are correlated with the expression levels of aiiA gene in Bacillus thuringiensis. Enzyme Microb Technol 38:967–974. doi:10.1016/j.enzmictec.2005.08.037
Acknowledgement
The authors wish to thank the Director of CSIR-Institute of Genomics and Integrative Biology (IGIB), CSIR-INDEPTH (BSC0111), Government of India for providing the necessary funds and facilities. Authors are also thankful to Academy of Scientific and Innovative Research (AcSIR), New Delhi. PK is thankful to CSIR for granting Senior Research Fellowship.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer India
About this chapter
Cite this chapter
Kumar, P., Koul, S., Patel, S.K.S., Lee, JK., Kalia, V.C. (2015). Heterologous Expression of Quorum Sensing Inhibitory Genes in Diverse Organisms. In: Kalia, V. (eds) Quorum Sensing vs Quorum Quenching: A Battle with No End in Sight. Springer, New Delhi. https://doi.org/10.1007/978-81-322-1982-8_28
Download citation
DOI: https://doi.org/10.1007/978-81-322-1982-8_28
Published:
Publisher Name: Springer, New Delhi
Print ISBN: 978-81-322-1981-1
Online ISBN: 978-81-322-1982-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)