Abstract
Label-free Raman imaging is a noninvasive spectroscopic method for investigating the nature and distribution of molecular species within a sample. In this chapter, we describe the applications of conventional Raman imaging, as well as the related techniques of coherent anti-Stokes Raman scattering (CARS) and stimulated Raman scattering (SRS) imaging for medical, life sciences, and other biological applications.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
1 Definition of the Topic
Label-free Raman imaging is a noninvasive spectroscopic method for investigating the nature and distribution of molecular species within a sample. In this chapter, we describe the applications of conventional Raman imaging, as well as the related techniques of coherent anti-Stokes Raman scattering (CARS) and stimulated Raman scattering (SRS) imaging for medical, life sciences, and other biological applications.
2 Overview
The application of Raman imaging to a wide range of biological materials has shown very promising results and is now providing biological insights that are not available to other methods. In this chapter, we first discuss typical experimental considerations for Raman, CARS, and SRS biological imaging in terms of sample preparation, instrumentation and measurement parameters, data processing, and data analysis approaches. Secondly, we discuss the recent applications of Raman, CARS, and SRS imaging of biological materials ranging from human cells and tissues for disease diagnosis to nonmammalian samples including plants, shellfish, algae, and nematode worms. The three techniques have been used to investigate wide-ranging topics from organism development, disease progression, drug uptake and metabolism, the effects of environmental pollution and toxin effects on cells, all the way through to testing the quality of food products, drugs, and even biologically derived charcoal. Finally, information obtained from Raman, CARS, and SRS images can be complimented by other techniques, and we discuss various multimodal imaging approaches that have been employed for biological analysis.
3 Introduction
Raman spectroscopy is based on the inelastic scattering of light, i.e., where incident and scattered photons have different energies, or frequencies, after interaction with matter. The frequency shift between the incident and scattered photons is indicative of the vibrational level of a molecule [1]. Therefore, a Raman experiment gives rise to a spectrum containing information about the chemical bonds present in a sample and can be thought of as a “fingerprint” of a sample. Raman spectroscopy has been used to characterize many different sample types in fields as wide ranging as materials science, organometallic chemistry, and geology [2]. However, key instrumentation advances such as the advent of laser-based Raman spectrometers in the 1960s [3] and their combination with optical microscopes in the 1970s opened up applications in fields that had often proved challenging with weaker excitation sources [4], including the analysis of biological materials.
Subsequent developments in instrumentation have improved the speed of collection to such levels where, in the last few years, it has been possible to measure Raman images, where each pixel contains a Raman spectrum with sufficient signal to noise for subsequent data analysis, within a few minutes. This has facilitated the application of Raman imaging to many types of biological samples, including live cells and tissues, providing information on the spatial distribution as well as the nature of the chemical components of the sample. Some studies have also shown that time-lapse Raman imaging is possible (for examples, see [5,6,7]) providing additional temporal information on changes in chemical distribution and composition.
Raman spectroscopy has a number of advantages over other analytical imaging techniques such as fluorescence imaging. In general, Raman spectroscopy is nondestructive, meaning that a sample can be measured multiple times, for Raman-based time-lapse imaging or for subsequent measurements with other analytical techniques. In contrast to many biological assays, this allows Raman measurement while the sample continues to carry out its normal function, e.g., cells can be reimplanted and continue to develop. Raman spectra can be collected from samples in many forms such as liquids, including aqueous solutions, soft and hard tissues, solids, and powders. Samples can be relatively simple compositions, as may be found in a pharmaceutical tablet, or more complex mixtures of components that may be present in tissue sections or whole organisms. This flexibility in sample format means that sample preparation steps are often simple and can be kept to a minimum to avoid inducing noticeable changes in the sample under investigation. In addition, it is possible to measure samples in conditions that mimic, or are close to, the normal sample environment, particularly useful for in vitro cell or in vivo tissue analyses, for example.
One of the main advantages of Raman spectroscopy is that it does not require the use of labels to identify sample components, which is particularly beneficial where the presence of a label may perturb the sample. This means that no prior knowledge of the sample composition is needed to perform a Raman experiment and Raman spectroscopy therefore has the potential to measure previously unknown components in a sample. (It is worth noting that although Raman spectroscopy can be performed on labeled samples, including Raman-active dyes, deuterium-labeled samples, and molecular tags that produce bands in the silent region (e.g., [8]), these are not routinely applied in Raman experiments. ) The imaging studies outlined in this chapter have all been restricted to those that do not use labels, including deuterium labeling, or the use of signal enhancement, e.g., from surface plasmons, utilized in techniques such as tip-enhanced Raman scattering (TERS) or surface-enhanced Raman scattering (SERS). Readers interested in biological applications of these enhanced Raman techniques are referred to several recent reviews on the topic (e.g., [9,10,11]). In addition, the other imaging techniques described in the multimodal Raman section have been limited to those that also do not require any sample labeling.
With improvements in the speed of Raman imaging measurements in recent years, there has been a corresponding increase in the number of published studies using Raman imaging, particularly for biological applications. While there are a number of recent reviews detailing the current status of Raman imaging for particular topics within bioanalysis, such as focusing on plant biology, hard tissue applications, and so on, this chapter aims to showcase the wide range of biological and medical-related applications utilizing Raman imaging. Although this chapter primarily focuses on spontaneous Raman imaging, there are several other label-free Raman techniques available that can vastly improve imaging speed, but at a trade-off of spectral resolution and range. CARS and SRS are the most well-known approaches and use coherent nonlinear effects to boost the measured signal. As such, an overview of the implementation and types of results obtained with CARS and SRS is also included. Raman, CARS, and SRS can all, potentially, be combined with other techniques to provide additional information on the nature of the sample under investigation. Therefore, the final section of this chapter focuses on recent developments in combining Raman-based imaging modalities with other imaging techniques including infrared spectroscopy, mass spectrometry, digital holographic microscopy, and Brillouin imaging.
In the following chapters, we will outline some of the main considerations Raman, CARS, and SRS imaging including sample preparation (Chap. 2.1), instrumentation (Chap. 2.2), and data analysis (Chap. 2.3). In Chap. 3, we summarize the key research findings in the last few years, primarily from 2010 onward, in the fields of Raman (Chap. 3.1), CARS and SRS (Chap. 3.2), and combined imaging approaches (Chap. 3.3) for applications in biosensing and medical diagnosis.
4 Experimental and Instrumental Methodology
4.1 Sample Preparation
Raman spectroscopy can be performed on a wide range of different sample types from solutions such as tears, blood, plasma, and other bodily fluids (see Baker et al. [12] for a recent review) to solid materials including teeth and bones (see Gamsjaeger et al. [13] for a review). In terms of imaging studies, Raman has been used to study a wide range of both soft and hard tissues in organisms ranging from humans and other mammals to plants, fungi, and bacteria . Each of these sample types has their own preparation constraints, and some example protocols have been outlined in a recent review [14]. Some general considerations for both soft and hard tissues are discussed below.
4.1.1 Sample Considerations
While sample preparation for Raman imaging is usually straightforward, there are some factors that should be considered to ensure good-quality images. In general, it is more challenging to perform Raman spectroscopy on dark-colored samples than it is for lighter-colored or transparent samples, as darker colors can absorb the excitation laser light resulting in damage to the sample. For example, Zhang et al. noted Raman spectroscopy of black hair was difficult due to the melanin content which absorbed the laser light resulting in sample destruction and so restricted their analyses to lighter-colored hair [15]. Therefore, on darker pigmented samples, it may be necessary to limit the excitation laser power used to prevent sample damage or shift the wavelength to a nonabsorbing region, both of which can have an effect on the signal-to-noise quality of the generated Raman spectra.
In many exploratory Raman studies, it can be useful to compare the Raman images obtained with other imaging techniques, one of the most established being fluorescence imaging. Fluorescence imaging (unless limited to autofluorescence) requires the use of labels, either through the use of dyes, immunostains, or expression of fluorescent proteins such as green fluorescent protein (GFP) in the cells or tissues of interest. In general, the presence of these labels will generate a broad fluorescence signal that completely obscures the Raman bands in a spectrum. This precludes the use of pre-stained samples for Raman, i.e., any fluorescence labels should be added after the Raman imaging has been performed, and means that Raman imaging cannot routinely be applied to cells or tissues from transgenic animals expressing fluorophores.
Currently, most Raman imaging of cells has been based on those in 2D cell culture, i.e., individual cells or monolayers of cells . However, in the last few years the possibilities of using 3D cell cultures have emerged. With continuing advancements in fields such as scaffold engineering and tissue fabrication [16], or tumor tissue architecture [17], for example, the ability to measure biomolecules, cells, and tissues in 3D will be a key tool for future research in these areas. While the practicalities of successful 3D cell culture preparation are beyond the scope of this chapter, it is worth noting that it has already been shown that Raman spectra of sufficient quality for analysis can be obtained from within both spheroidal 3D cultures, approximately 300 μm depth, and cells embedded in a matrix, at approximately 120 μm depth when using 785 nm excitation [18]. The effects of the laser excitation wavelength on the penetration depth achievable are discussed in more detail in the Raman instrumentation section. In a similar manner, imaging of both hard and soft tissue has often required sectioning the sample to produce thinner, and often flatter, subsections of the sample for analysis. Depending on the strength of the sample, sectioning can result in tearing, fracturing, or other distortions to the sample. As such, developments in imaging of 3D cell cultures may also be useful for imaging of tissues by reducing the need for sectioning or may have an impact on the applications of Raman imaging in surgery, for example, where tissue sectioning prior to measurement is not possible.
4.1.2 Live Versus Preserved Samples
While it can be argued that imaging live samples, in conditions close to their natural environment, is most desirable, it is not always possible to do so. Examples where imaging live samples is not feasible include the use of nonsimultaneous multiple imaging techniques where movement in the sample between measurements will affect image registration, where the phenomenon under investigation occurs faster than the time taken to collect an image, or where samples must be collected in a different location to the imaging measurement, e.g., tissues collected during surgery or from tissue banks. In circumstances where samples need to be preserved prior to measurement, determining which fixation method to use can be a challenging question. A number of studies have investigated fixation and cryopreservation methods for cells [19, 20] including oocytes [21] and tissues [22,23,24] in terms of their effects on sample morphology and biochemical composition as well as on Raman spectra and image quality. Chemical fixation methods, such as formalin fixation, and cytocentrifugation have been shown to preserve the cell in a state close to that of live cells, while air-drying results in more distinct changes [19]. When using chemical-based fixation methods for cells, aldehyde-based fixation protocols perform better than alcohol-based methods, and parameters such as the temperature of the fixation media also influence the quality of both the Raman images and spectra [20]. In tissues, common fixation, embedding, and dewaxing procedures can lead to a loss of lipid content, with the consequence that other cellular components such as proteins are relatively enriched in the resulting spectra [23]. Depending on the tissue, xylene removal may not be completely effective, resulting in some paraffin signal present throughout the sample, and may be particularly prevalent in some tissue types. This paraffin residue can also affect the interpretation of molecules in the tissue section that have a spectral overlap with paraffin, such as phospholipids [22]. However, the spectra are still detailed enough to allow for clear determination of different tissue layers in skin [24] and colon tissue [22], for example.
Although not commonly analyzed by Raman imaging, it is also possible to measure biofluids, and for imaging, this would generally mean that the biofluid would be dried out on a suitable sample carrier so that variations in sample composition can be measured. When using this approach, it is important to consider the concentration of the sample deposited onto the sample carrier as greater homogeneity in the distribution of components is seen for more dilute samples, as was shown for proteins in synovial fluid [25].
4.1.3 Sample Substrates and Imaging Environment
The substrate that a sample is placed on for measurement can influence the Raman spectrum, especially if the sample is relatively thin, for low numerical apertures, or if Raman spectra from z-positions close to the substrate are needed . Many plastics, such as those used for cell culture dishes, are not suitable for Raman imaging as they give rise to very strong Raman spectra. Glass microscope slides and coverslips can be a suitable option, particularly when using longer excitation wavelengths, although most glass slides do produce some broad bands below approximately 1000 cm−1 which can be particularly noticeable in spectra collected close to the glass surface. Quartz microscope slides and coverslips also give rise to relatively weak, broad Raman bands, but these tend to cover a more limited wavenumber range than those observed with glass substrates, making quartz a good option. In terms of cell imaging, cell culture dishes with either glass or quartz bottoms are available, allowing cell culture and Raman measurements to be performed on the same substrate, minimizing the environmental change experienced by the cells. Good-quality CaF2 exhibits a strong Raman band at approximately 321 cm−1 but does not give rise to strong bands in the fingerprint or high wavenumber regions, particularly when using visible excitation [26, 27]. Therefore CaF2 slides can also be used for Raman imaging in cases where the low wavenumber regions are not required. Other common sample carriers may not always be suitable for Raman imaging, depending on the nature of the sample. For example, cells do not appear to proliferate well on ZnSe [28], while other substrates may alter the Raman signal obtained, e.g., MirrIR Low-E glass slides can increase the Raman signal via reflections but require measurements to be performed in a transflection orientation [29] which may not always be desirable.
Depending on the sample, the substrate may need to be treated in order that the sample adheres to the substrate . This is common for cell imaging where the substrate is coated with compounds, often proteins such as gelatine [30] or poly-L-lysine, which cells often adhere to better than the bare substrate. For other samples, such as nonadherent cells, samples may have to be embedded in a matrix prior to measurement. Example matrix materials include cellulose, methylcellulose, collagen, glycerol, and complex matrices such as Matrigel. However, it should be noted that all of these matrix compounds produce Raman spectra of their own, and so careful selection of the right matrix material should be based on minimizing the spectral overlap with the sample of interest.
Hard tissues and many preserved samples can be measured in air. However, for live samples, and indeed some preserved samples, they should be immersed in liquid during the Raman imaging experiment in order to prevent the sample drying out . Raman spectra are not strongly affected by the presence of water, allowing the use of aqueous solutions for sample immersion. Although water may be suitable for some preserved samples, for most live sample imaging and particularly for longer-term imaging, additional salts and sugars are needed to maintain the health of the sample. Simple buffers such as phosphate-buffered saline supplemented with an energy source such as glucose do not generally make a significant contribution to the Raman spectrum of the immersed sample. It is also possible to measure samples in common culture media such as RPMI 1640, DMEM, MEM, etc. as long as they do not contain colored indicators such as phenol red, which create significant background contributions to the Raman spectra, often completely swamping any Raman signals from the sample. Supplemented culture media, e.g., with fetal bovine serum added, can still be used during Raman imaging experiments, but depending on the concentration of the supplemented compounds, they may contribute to the Raman spectrum.
4.1.4 Sample Number and Statistical Significance of Results
Finally, when designing any experiment that uses human or animal samples, or other samples that are in limited supply, it is important to consider the number of samples that should be analyzed . For exploratory studies, the sample number may not be as important, but for any studies from which conclusions about human health, disease markers, purity assessments, and so on are drawn, it is important to ensure that a sufficient number of samples are analyzed to ensure the conclusions drawn are robust and unbiased. The number of samples that will be required for a particular study is dependent on several factors including the type of samples, the hypothesis or scientific question under investigation, and the confidence limits required for reporting the data. More information on the parameters affecting the appropriate number of samples for a study and methods to determine appropriate samples sizes have been outlined previously (e.g., [31,32,33]). With spectroscopic data in mind, detailed information on the selection of the appropriate number of statistically independent samples for use in classification models has been provided by Beleites et al. While they focus on classifier training and testing, the points covered are also applicable to other experiments (i.e., where classifier training is not used) including where limited sample numbers are available [34].
4.2 Instrumentation
4.2.1 Raman Instrumentation
The components of typical Raman microscopes have been comprehensively described in recent reviews by Butler et al., Krafft et al., and Pence and Mahadevan-Jansen [14, 35, 36]. Therefore, in this chapter we have concentrated on those elements of particular importance for the analysis of biological specimens and the developments in instrumentation that are specific for imaging microscopes.
Raman spectroscopy can be achieved using excitation wavelengths from the UV, through the visible range, and into the infrared with the wavelength employed influencing several aspects of the Raman spectra obtained from biological samples. Using near-infrared (NIR) excitation wavelengths avoids the problems of fluorescence, partly due to the fact that the excitation photon energy is lower and cannot drive the same transitions to fluorescing excited states that occur with visible photon energies. Additionally the Raman spectrum and fluorescence do not overlap significantly [37] and NIR light penetrates deeper into tissues such as skin [38], compared with visible and UV light. However, as the efficiency of most CCD detectors is lower in the NIR region, and the Raman intensity is inversely proportional to the 4th power of the excitation wavelength, acquiring Raman spectra with good signal to noise in the NIR can require relatively long collection times [37]. Measurements taken with excitation lasers operating in the visible region, therefore, benefit from increased Raman intensities and better detector efficiencies; however, this is at the cost of increased overlap with fluorescence contributions to the Raman spectrum. Despite this, many of the studies discussed in 3.1 have been performed with visible excitation, both in the green and red regions of the electromagnetic spectrum. The UV region of the spectrum generally suffers less from fluorescence contributions [37] but, due to the high energy of the excitation light, can cause photodamage and photodegradation of the sample. Deep UV imaging is not commonly carried out due to the risk of sample damage. However, under carefully controlled conditions, it is possible and generates Raman images rich in information on nucleic acid bases and aromatic amino acids. Adding Lanthanide ions can also significantly extend the sample lifetime under deep UV imaging conditions [39]. The choice of excitation laser may also depend on the sample under investigation as resonance enhancement of Raman signals can occur when the excitation wavelength is close to an electronic absorption band in the sample [40]. As examples of molecules of biological interest, resonance Raman spectra of nucleic acid bases and aromatic amino acids can be recorded with UV excitation, while carotenoids such as β-carotene and heme-based molecules generate resonance Raman spectra at visible wavelengths [40, 41].
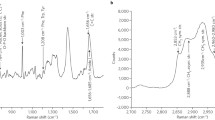
Raman measurements were initially performed as point-based measurements (see Fig. 7.1a for a basic overview of a Raman microscope) , which eventually led to the ability to capture high-quality spectra from distinct points in biological samples using confocal detection around the early 1990s, where the techniques were referred to as “microspectroscopy” rather than “imaging” [42]. The approach was initially very limited in terms of speed: if the detector is only one dimensional and the acquisition time for a single Raman spectra is in the order of seconds, then modest sized Raman images could take hours to acquire (e.g., 100 × 100 pixels, 1 s per pixel, ~3 h). Especially for biological samples, where higher power may result in sample damage, and even fixed samples that may degrade over long measurement times, there has been a strong need for faster Raman imaging. Significantly faster biological images can be obtained by using a line-shaped focus with a two-dimensional detector, modality [43], allowing the observation of biological dynamics with a time scale of minutes, such as apoptosis [44]. The 2D detector allows for the parallel acquisition of all spectra from each point in the line-shaped focus. For the current state of the art, high-end deep-cooled CCDs can collect more than 90% of the incoming light, the laser excitation can be applied up to the limits of sample damage, and the numerical aperture (capture angle) of the microscope objectives is effectively maximized; the imaging speed for diffraction limited images by spontaneous Raman can be on the order of minutes per cell and not substantially faster unless some other factors are exploited. One intuitively obvious way to improve imaging speed is to reduce the sampling points. Interestingly, this does not always result in a significant degradation of the final image. Spectra taken from points in cells separated by much more than the diffraction limit can show notably different features [45], but in general, the oversampling in order to maximize resolution includes a large amount of redundant spectral measurements that are not significantly different from their adjacent pixels. This depends of course on the spatial variability of the sample but also results from the physics of the microscope itself, where the point spread function blurs each measured point. To avoid excessive blur, oversampling is common, but for more uniform samples, where the differences from region to region are more gradual, such requirements can be relaxed in order to speed up the measurement time. Even for highly heterogeneous samples, intentionally undersampling and using compressed sensing reconstruction techniques can be very powerful in optimizing the total information acquired for a given amount of measurement time and laser power [46].
Basic outline of instrumentation for (a) Raman, (b) CARS, and (c) SRS microscopes. In all cases, many optical components such as lenses and filters have been omitted for simplicity. In addition, only the laser path is represented, and indications of laser beam width, polarization, etc., are not shown. (a) Raman microscopy. Laser light is directed to the sample on the microscope stage via a microscope objective, illuminating a point on the sample. Backscattered light is directed via a bandpass filter through a notch filter (rejecting light that is the same wavelength as the laser excitation) and slit (controlling the amount and angle of the light entering the spectrograph) before dispersion of the light, via gratings, and collection on the CCD camera. The addition of cylindrical lenses in the system, in conjunction with a 2D detector, results in parallel collection of spectra from all points along a line illumination on the sample. (b) CARS microscopy. Many different instrumentation approaches have been used for CARS microscopy, and so only a stylized version is given here. Two laser wavelengths, a pump beam and Stokes beam, are directed to the sample via a microscope objective. The output light is collected via a second objective and detected via a spectrograph and CCD camera. Generation of the two laser wavelengths is often done using a pulsed laser in conjunction with an optical parametric oscillator (OPO) or optical parametric amplifier (OPA) or in conjunction with a photonic crystal fiber. (c) SRS microscopy. Similarly to CARS, SRS employs two laser wavelengths, a pump beam and a Stokes beam, with the intensity of one beam modulated. These beams are directed to the sample via a microscope objective where (in the presence of a resonant vibrational mode) some degree of modulation is transferred between the two beams. The light is collected via a second microscope objective and the change in modulation of the beams detected via photodiodes attached to a lock-in amplifier
For larger samples, particularly tissue sections, even with improvements mentioned above, measurement times with full imaging can still be prohibitive. Various approaches to better “target” the Raman measurements to specific regions of interest have been suggested in order to further reduce measurement times required. The majority of these approaches involve combining Raman imaging into a multimodal system where other analysis techniques provide an overview of the sample (discussed in more detail in the multimodal imaging section) allowing Raman measurements to be taken from the regions of interest identified by the other modality. By using scanning mirrors, it is possible to select regions in a cell and determine how much spatial information to average from the sample, which can result in improved Raman spectra compared to single point or even full imaging [47]. Another approach is to use the Raman data collected as a means of determining the next measurement points. Rowlands et al. implemented this approach by first taking two spectra at random. Next they calculated a cubic spline interpolant and a Kriging interpolant between these two points. Depending on the result from the Kriging interpolant calculation, the next measurement point is taken as either the position furthest from those already measured or the position with the largest difference between the two interpolants. The process is repeated with the new measurement points until the difference between the two interpolants reaches an acceptably small level. The increase in overall measurement speed is based on the assumption that these two interpolant values will converge in areas where there is little variation in sample features. This should result in fewer measurement points taken in these regions while concentrating the measurements in areas with higher variability [48]. An alternative method for reducing the number of spectral measurements required to analyze a sample is to employ compressed sensing approaches, where a signal can be reconstructed from a sparse subset of data points. This approach has recently been applied to hyperspectral Raman images, without the need for microscope modifications. High-resolution and relatively low-noise Raman images could be reconstructed from five times less data point sampling than would be used in conventional image Raman imaging, increasing the speed of measurement. The reduced exposure time for the sample has added benefits of reducing photobleaching on sensitive samples [46].
4.2.2 Coherent Anti-Stokes Raman Scattering (CARS) Instrumentation
The discussion so far has been concentrated on spontaneous Raman scattering, whereby an incident photon is scattered according to the vibrational bonds in the sample. The excitation light does not need to be coherent, although in practice, lasers provide the most efficient narrow bandwidth and easily focused excitation fields. The low Raman scattering cross section for most biological materials means that the techniques require large photon flux, optimized collection geometries (i.e., high NA lenses), and sensitive detectors. With excitation powers approaching the limits of sample damage, one approach to improve signal is to tune the laser excitation to resonate with a vibrational band of interest. This then requires coherent excitation and is usually implemented with pulsed lasers. Pulsed lasers have the additional advantages of having high peak power, which is necessary to drive the nonlinear interactions, and are usually tunable. Coherent Raman techniques have limitations in the ability to measure and quantitatively interpret rich spectral information, but rather than competing directly with spontaneous Raman spectroscopy in terms of spectral range and resolution, they are usually exploited for their strengths, which include the ability to provide high-speed, high-contrast spectral imaging. Until now, the most common way to use nonlinear coherent effects to boost the Raman signal has been to use coherent anti-Stokes Raman scattering (CARS) that can be used as an imaging mode [49]. It is also possible to combine CARS in a complementary manner with spontaneous Raman spectroscopy to provide detailed spectra from one mode, with high signals from the CARS mode [50].
CARS imaging is implemented by using two wavelengths of laser excitation, one is the pump beam which drives a molecular excitation to a virtual state, which can then be driven, by stimulated excitation, down to a vibrational state by a Stokes beam (Fig. 7.1b). The molecule can then be excited to a higher virtual state and driven by stimulated emission down to the ground state. When the frequency difference between the two incident beams is resonant with a vibrational transition in the sample, the output signal increases. With the pathways involving virtual states, the process occurs essentially instantaneously, following a four-wave mixing process, rather than following actual excitation and de-excitation transitions which occur in fluorescence. This means that the output CARS light is coherent and directional. This has implications for detection methods including the possibility of gating the detection to reduce background signals, as well as control of the wave front and optimizing detection efficiency, while maintaining polarization. It also makes the process highly nonlinear with regard to the incident laser intensities. These points allow a degree of optical sectioning which, in conjugation with the near-infrared wavelengths, allows imaging in tissue samples that are difficult to image by spontaneous Raman imaging. For CARS, the fact that the anti-Stokes shift is used means that the output signal is also far away from the Stokes fluorescence typically generated from the sample.
While CARS imaging is a relatively complex imaging mode, in its most basic form, it provides only the spatial contrast resulting from a single vibrational mode. In practice this can be useful, but the ability to record multiple bands, or an entire spectrum, remains a key strength of spontaneous Raman imaging. CARS research has therefore pushed toward acquiring hyperspectral information. There are two instrumental approaches to achieving hyperspectral CARS imaging. One is to change the laser wavelength to achieve the signals for each vibrational modes of interest. The second is to use a broadband source to produce multiple excitations of vibrational modes [51].
The rapid pace of development in CARS techniques means that there are a large amount of different reports using different variations in instrumentation, and a full discussion is beyond the scope of this current work. However, key features of CARS include the spectral selectivity and high-speed imaging capability [52]. The inherent nonlinearity in CARS can also be exploited to increase both spatial and spectral resolution by measuring the onset of saturation in the CARS signals [53].
4.2.3 Stimulated Raman Scattering (SRS) Instrumentation
There are multiple ways in which coherent excitation and nonlinearity can be exploited to achieve high signals in Raman measurements. Aside from CARS techniques, discussed above, pulsed laser excitation can be used to stimulate Raman scattering, known as SRS. SRS can be used as a rapid imaging modality, with tunable selectivity for bands of interest. A good overview of the technique, with an explanation of the differences between SRS, CARS, and spontaneous Raman scattering, is available [52]. A significant advantage of SRS over CARS is the reduced background signal contribution. In terms of implementation, SRS uses two pulsed laser beams at pump and Stokes frequencies (Fig. 7.1c), and the intensity of one of the pulsed beams is modulated. Due the nonlinear interaction of the two laser beams, which occurs in the presence of a resonant vibrational mode at the laser focus, a degree of modulation is transferred from one beam to the other. This can then be detected using a lock-in amplifier and is related to the amount and type of vibrational modes in the sample, producing imaging contrast. SRS shares some of the overall limitations of CARS in the sense that it is challenging to acquire full spectra, but a number of different approaches have been made toward this goal [54]. High frame rate imaging, while stepping the spectral collection range, has already been demonstrated at video rates and above [55].
4.3 Data Analysis
4.3.1 Data Pre-processing
Interpretation of Raman spectra, especially Raman spectra acquired during imaging (which tend to have lower signal-to-noise levels than point spectra), usually requires some level of pre-processing before analysis, either by visual inspection of the spectra or by subsequent analysis by chemometric algorithms. Many of the points for consideration have also been discussed in a recent review by Byrne et al. [56], and particular attention to how experimental parameters affect Raman peak intensities, and hence Raman quantification, has been addressed by Kumar et al. [57]. Ideally, when analyzing Raman spectra, there should be no contribution to the spectra from any source other than the sample. While it is not always feasible to remove all other influences to the spectra, pre-processing steps aim to remove non-sample contributions as much as possible. Roughly, the sources of influence on the spectra arise from the instrument, the substrate, and the sample itself.
The exact performance of Raman instruments will vary slightly from machine to machine and will also change over time for an individual instrument particularly where maintenance procedures such as laser alignment or replacement of components are performed. Therefore, in order to allow comparisons between measurements, both those taken on the same instrument and those taken on different instruments, it is advisable to regularly asses the response of the Raman instrument(s) used and, as and when necessary, correct parameters such as data point spacing and scattering intensities [56]. There are various approaches to calibrate Raman instruments , for example, Slater et al. have outlined a calibration procedure with the following steps: (1) wavelength calibration using a neon standard, (2) intensity calibration using a tungsten-halogen standard, (3) laser wavelength calibration using a Raman standard, and (4) validation of the calibration using the Raman standard. They suggest cyclohexane as a good Raman standard, using the 801.1 cm−1 Raman band [58], but other materials with a well-characterized Raman spectrum (particularly in terms of band position(s) and intensities) such as silicon are often used as well. It should be noted that using a homogenous sample to calibrate a Raman imaging system may obfuscate optical aberrations that appear when a spatially variant sample is then used after calibration/alignment. Therefore the quality of both spatially resolved components in the image and their spectral features should always be carefully evaluated in acquired data, even when alignment and calibration metrics appear correct. Where it is necessary to compare Raman spectra that have been measured with different excitation wavelengths, calibration procedures such as that suggested by Bocklitz et al. [59] can be used.
Other sources of spectral variation in the Raman spectrum originating from the instrument response include electronic sources of noise such as detector sensor variations, cosmic rays, dark current, and readout noise, which are always present to some degree but are often unpredictable contributions to the spectra. In order to reduce the influence of these sources of noise to the data, Raman images can be filtered or smoothed . Cosmic rays often strike the detector during long measurement times, causing errors that are large but usually very localized. They can then be removed by a local application of a median filter, with a threshold (typically several standard deviations) so that the majority of the data (i.e., the locations where no cosmic rays exist) is not altered in any way. As a 3D hyperstack, Raman images can be smoothed in the spectral domain, the spatial (x-y) domain, or both. Smoothing in the spatial domain is of limited use, since noise will be present throughout the spectral vector represented by one pixel, but smoothing in the spectral domain is commonly used. Depending on the spectrometer resolution, slit width, and laser wavelength, the spectrometer resolution is often less than would be implied by simply looking at the output spectra, and a more realistic (and less noisy) spectra can be produced by smoothing appropriately, considering all of these parameters. Loess fitting and other techniques are commonly employed but are beyond the scope of this current work. In general, however, it should be noted that the output from smoothing procedures should result in removal of noise, without the removal of features interest. Oversmoothing in the spectral domain will result in a loss of biochemical information, while oversmoothing in the spatial domain will lead to blurring of images.
Even with careful selection of a sample carrier, there will usually be some contribution to the Raman spectrum from the sample substrate. There are several ways to remove the background contributions in the spectrum . One method is to select a region of the image where no sample is present, extract an average Raman spectrum from this region, and then subtract this average from each pixel in the image. However, depending on the exact proportion of contributions from the sample and background to each pixel in the image, this may require careful implementation of baseline correction and normalization prior to background subtraction. It can also easily force some pixel values to become negative, which can be problematic for analysis since no real optical signal should be measured as negative. Another method to remove substrate influence in the Raman spectrum is to use chemometric algorithms such as independent component analysis (ICA ) that identify the background contributions as one spectral component [56]. While subtraction approaches can be useful when samples are imaged in simple media or matrices, it should be noted that background subtraction becomes more difficult when samples are imaged in more complex media. This is due to the mismatch between the sample and the surrounding media in terms of composition, particularly where substances are present in the media that are not present in the sample, as subtraction will result in negative contributions to the Raman spectrum of the sample for these substances. It is worth considering that even for simple media, the water “concentration” will be higher outside the sample, and so background subtraction to remove media contributions to the sample spectrum should also be normalized carefully before subtraction.
Some distortions to the Raman spectra can originate from the sample itself and also need to be addressed during spectral pre-processing. Autofluorescence often results in an increased background signal, particularly in the fingerprint region of the Raman spectrum. As most fluorophores have distinct excitation and emission ranges, autofluorescence can be avoided by the choice of excitation laser used. However, this is not always practical, especially if multiple fluorophores are present in the same sample, and the influence of the autofluorescence has to be dealt with once the spectrum is recorded. If the autofluorescence is relatively uniform across an image, for example, if it originates from the sample substrate, it is usually possible to remove the bulk of its influence by background subtraction. However, many compounds within the cell generate some level of autofluorescence. Examples include NAD(P)H, collagen, elastin, and flavins [60]. As these will not be uniformly distributed throughout the cell, it is harder to remove their influence, and baseline correction methods are perhaps the most effective solution. However, as the autofluorescence can be a signature of the compounds in the cell, it is not always desirable to remove it, since it can provide meaningful biological information [61].
One of the issues with analysis of tissue sections, and to a certain extent cells, particularly when using chemometric analysis methods, is the effect arising from the edges of the cell/tissue [30, 62]. Spectra originating from these regions can suffer from a number of problems including baseline distortions and widely varying proportions of sample versus background/substrate from pixel to pixel [62]. The high spectral variability in these regions, as compared to the bulk of the tissue, results in a large number of clusters/components that have little biological meaning. Strategies to overcome this and to focus on the relevant differences in the sample include controlling the number of clusters or components so that the edge of the tissue is described by distinct clusters that do not influence the interpretation of the bulk of the sample [30, 62]. Sample thickness can also affect Raman spectra, either through different volumes of sample being probed at each pixel or through out-of-focus effects [63]. A few studies have overcome the out-of-focus effects by using optical profilometry methods to assess the 3D topography of the sample, directing the measurement of the Raman spectra to always be taken from the top layer of the sample [63, 64]. Normalization of spectra, either through the use of an internal standard or through vector normalization , can reduce the spectral differences induced by sample thickness differences but should always be applied after other corrections (background subtraction, baseline correction, and so on) [56].
4.3.2 Data Visualization and Analysis
Raman images of biological samples contain a wealth of information in terms of both chemical contributions and spatial distribution, and visualizing the data is an important step in interpreting this information. A common way to represent the data contained in a Raman hyperspectral dataset involves producing false color images based on the spectral information contained at each pixel in the image, e.g., the area under a Raman band, Raman band intensity at a given wavelength, the ratio between two Raman band intensities, or similar characteristics of the Raman spectra. Multiple false-colored images can be overlaid to compare the distribution of molecules represented by Raman bands at different wavelengths.
Ashton, Hollywood, and Goodacre [65] have shown, using a “rainbow” lookup table (also known as “jet”) that is often the default color lookup table (LUT) for many image plotting software packages, the importance of the way in which the data is plotted in avoiding over-interpreting the data or missing features of interest. Using the simulated data shown in Fig. 7.2, they show that four different images can be obtained from the same dataset depending on how the false color is implemented. As the authors point out, there is no “right” or “wrong” image here; rather each image is able to answer different questions about what is contained within the data. The example, shown in Fig. 7.2, shows this. Image (a) with a significantly truncated range can answer the question “is this an animal?” (b) with a slightly increased maximum can answer “what animal is this?” (c) covering the full range of counts can answer “is there more than one animal here?” as a bird-like shape is also present on the zebra. And finally (D) with discrete blocks of color for the scaling, rather than the continuous scaling in the other three images, shows two bird shapes and so is able to answer the question “how many animals are in the image?” [65]. This illustrates that the appearance of “objects” in the data can be highly dependent on not only the type of analysis performed but on something as simple as the false color LUT applied when showing the image. Although the data shown in this example is analogous to that of a single peak area/intensity/ratio plot, the points raised are also pertinent to overlaid images and to any scale-based images produced during chemometric analyses, for example, PCA scores images.
A comparison of rainbow-shaded images constructed from a simulated Raman map with an intensity range of (a) 500–2000 counts, (b) 500–3500 counts, (c) minimum to maximum intensity, and (d) minimum to maximum intensity but using discrete colors to form the color ramp, (i) and (ii) identify the position of the bird shapes discussed in the text. Color bars are shown to the left of each image (Figure reproduced from [65] published by the Royal Society of Chemistry)
4.3.3 Chemometric Analysis Methods
Chemometric analysis methods aim to clarify components in the data and separate them from noise, as well as provide other statistical tools such as classification of features into groups. There are many chemometric analysis methods that can be applied to Raman spectroscopy, many of which can also be adapted for the analysis of Raman images. Chemometric algorithms are often classified in several ways. They can be univariate, based on one measurement point (wavelength/Raman shift), or can be multivariate, incorporating multiple measurement points in the analysis. They can either be hard classification methods, where a spectrum (or image pixel) can only belong to one class, or they can be soft classification methods where a spectrum or pixel can belong to more than one class. Finally, they can be unsupervised, where no a priori information is used, or can be supervised, where information, for example, in the form of a reference spectrum, is required for the analysis. There are numerous articles that compare and contrast the various methods that have been applied to Raman spectroscopy (e.g., [56, 66,67,68,69,70,71]) to which the reader is referred to for further details. In this chapter, discussion has been limited to a brief overview of those analysis methods most commonly applied to Raman imaging as evidenced by the studies discussed in Sect. 3.
Principal component analysis (PCA ) looks for a fundamental set of independent variables in a dataset [72], thereby reducing the dimensionality of a dataset. PCA produces a set of principal components (PCs) where PC1 describes the largest variation in the dataset, PC2 describes the second largest variation in the dataset, and so on. In terms of Raman imaging, PCA produces a score for each pixel/spectrum along with a loading vector, containing spectral information on the variance in the dataset, for each PC. The loading vectors contain both negative and positive peaks and, depending on the complexity of the sample, can be challenging to interpret [56]. The score values for each pixel can be used to create false color multivariate images showing the distribution of each of the PCs in a similar manner to the univariate plots produced by Raman band intensities/peak areas, etc.
Hierarchical cluster analysis (HCA ) is a method for grouping spectra together into clusters based on their similarities to each other often by calculating Euclidian or Mahalanobis distances [68]. In terms of Raman imaging, each pixel can be assigned to a particular color based on which cluster the spectrum belongs to, creating a false color map showing the location of spectra in each cluster. Selection of the number of clusters is normally user defined, meaning that determining the number of clusters can be somewhat subjective, although consideration of the spectral differences between average spectra for each cluster and the standard deviation for spectra described by one cluster can help determine how many clusters are appropriate.
Vertex component analysis (VCA ) is based on unmixing hyperspectral datasets to find their reference components, also known as end-members. VCA is based on the end-members being vertices of a simplex in 3D space, meaning that all other data points can be described in terms of these end-members [73]. For Raman imaging, this means identifying the spectra that are the most “different” from each other and producing false-colored images (one per end-member) that then describe the abundance of that end-member at each pixel in the image. The end-member images can be overlaid to produce a composite image. As for HCA , many of the implementations of VCA require the number of end-members to be user defined.
Multivariate curve resolution (MCR ) is another method for elucidating the pure signals from a data matrix of mixed measurements, based on the idea that the mixture is a bilinear model of pure signal contributions, or at least the variation in the dataset can be described by a bilinear model. Raman images must be unwrapped prior to MCR analysis and the results reformed into images [74]. Thus MCR produces distribution images and corresponding spectra for each of the pure components identified during the MCR analysis.
K-means clustering is another clustering method that aims to reduce a data set into a defined number of groups. The process starts with “k” number of groups consisting of a random data point. Subsequent data points are added to the group whose mean is closest to that of the data point, and the mean of each group is adjusted as new data points are added [75]. For Raman imaging, k-means clustering can be used to produce false-colored maps, with each pixel assigned to a single cluster, as well as also generating an average spectrum for each cluster.
5 Key Research Findings
5.1 Key Research Findings in Raman Imaging
5.1.1 Cell State and Metabolism
The use of Raman imaging for analysis of cells and tissues, particularly when used for many of the applications to human health and disease discussed later, is dependent on understanding the origins of the Raman signals generated. The simplest way to do this is to compare cell and tissue spectra to the spectra of individual cellular components. This is relatively straightforward when the sample has a simple composition but can be challenging where a large number of different molecular species are all contributing to the overall signal. One way to tackle this complexity, particularly for cellular imaging, although the same can be applied to tissue imaging, is to identify marker bands or spectral profiles for classes of molecules, such as a lipid or protein signature. An alternative approach, depicted in Fig. 7.3, is to compare Raman and fluorescence images in order to identify spectral profiles for different cellular compartments and organelles [76]. Ramoji et al. have also shown that false color Raman images can identify some of the same cellular features as classical staining methods in leukocytes [77]. In another example, this time exploiting the resonance Raman effect, the colocalization of cytochromes B and C has been used to pinpoint the location of mitochondria in yeast cells [78]. A more recent study has taken hierarchical cluster analysis (HCA ) of Raman images and matched the distribution of each of the components generated in terms of the best overlap they have with corresponding fluorescence images. In doing so, Raman spectra that are related to each of the fluorescent stains used can be generated and can then be used as training data/reference spectra for further analysis of Raman images, without the need for the fluorescent staining of later samples [79].
Comparison of immunofluorescence images (a–d) and Raman-based images (e–h) measured by Klein et al. The red channels reflect actin, green channels reflect Golgi, and the blue channels reflect the nucleus. There are differences in distribution between the Raman signals and that of the fluorescence images, particularly in terms of actin distribution (correlation between a and e = 0.36, b and f = 0.74, c and g = 0.71). However, the combination of the components can provide a multichannel Raman image with a high degree of similarity to the fluorescence images [76] (Reproduced from Elsevier [76])
One of the main applications of resonance Raman imaging in a biological context has been for the study of cytochromes. As key components of the electron transport chain, cytochrome molecules have been used as indicators of living cells, particularly with regard to their redox states. Okada et al. followed the diffusion and oxidative state of cytochrome C during apoptosis and showed that these changes are independent of cell morphology [6]. The redox state of mitochondrial cytochromes in cardiomyocytes has also been estimated, based on the Raman intensities originating from cytochromes c, c1, and b, with reduced forms providing more intense Raman signals than the oxidized forms [80].
Several groups showed that with careful consideration of experimental conditions, it is possible to image live cells multiple times, allowing changes in the cells to be recorded at different time points (e.g., [5,6,7]). Pully et al. showed that changes in carotenoids induced by exposure to light could be detected before other changes in cell health, such as blebbing – a process that was accompanied by spectral changes in DNA and proteins. They also noted that some of the main constraints on time-lapse imaging studies are the control of nutrients, temperature, and pH of the cellular environment [5].
Raman spectroscopy has commonly been employed to discriminate between different cell types. Raman images characterizing the chemical composition (primarily heme, vitamin A and lipid content) of several types of cells originating from the liver (liver sinusoidal endothelial cells, hepatocytes, and hepatic stellate cells) have been used to identify each cell type [81]. Spectral differences were also used to classify different cell types/classes that may typically be present in blood from cancer patients, such as leukocytes , leukemic cells, and tumor cells [82]. The biochemical differences in Raman spectra of lymphocytes have been used to discriminate between T and B lymphocytes [83], and Raman imaging of murine stem cell colonies was able to identify nuclei and cytoplasm within the cells, as well as the extracellular matrix between the cells [84]. Raman imaging is also capable of distinguishing between different microorganisms , as shown by the identification of Bacillus subtilis, Staphylococcus epidermidis, and Saccharomyces cerevisiae from a mixture of all three species [85].
Identification of the current metabolic or activation states of cells can be a key step in understanding the course of diseases occurring when these normal processes go wrong. Raman imaging can determine the activation state of B lymphocytes, with a decrease in phospholipid content associated with activation [86]. Ghita et al. proposed that the differentiation status of neural stem cells could also be determined by nucleic acid Raman bands, specifically those indicating the levels of cytoplasmic RNA present, with higher concentrations seen in neural stem cells compared with glial cells [87]. Raman imaging has been used to assess the quality of human sperm, with healthy sperm exhibiting higher-intensity Raman bands associated with DNA and mitochondria, compared with sperm that had undergone UV-induced damage [88]. Chinese hamster ovary cells can be used for the production of proteins and monoclonal antibodies with Raman imaging able to discriminate between the different cell lines, as well as potentially able to indicate high-producing cell lines via the presence of a protein -rich signal that the authors attribute to the endoplasmic reticulum (ER ) [89]. Raman imaging has also been used to characterize the communication between different cell types, with actin and DNA content and distribution shown to be important in the immunological synapse that forms between dendritic cells and T lymphocytes [90].
Raman imaging of the cell cycle provides information on the different states of a cell during normal cellular processes. An early study was able to identify the cell cycle stage based on the Raman spectra obtained. However, the study suffered from problems in identifying the nuclear boundary which, as the authors pointed out, could be addressed by increasing spatial resolution of the Raman images but at the cost of the time taken for the measurements [91]. Shulze et al. were able to study the nucleus in detail, particularly in terms of the RNA and DNA distributions [92]. Combining Raman measurements with a second modality to identify the nucleus prior to Raman imaging has been suggested as a method for speeding up the Raman imaging [91]. Hsu et al. used such an approach, combining Raman measurements with autofluorescence imaging to follow the division of cells during cytokinesis and interphase [93].
5.1.2 Cancer Studies
Many cancer studies are performed on tissue sections, meaning that there is a mix of cell types included within each sample. As Raman imaging provides information on biochemical composition and spatial distribution, it can be used to identify the presence of different cell types within a tissue section. One such study showed that, in colon cancer tissue sections, Raman spectra could be used to identify erythrocytes (rich in heme but lacking other protein or lipid bands), lymphocytes (rich in lipids), connective tissue, and carcinoma (distinguished by differences in protein and DNA content). Although some researchers tend to avoid using 532 nm excitation for biological tissues in order to try and avoid the increased fluorescent background signal, this study found that the regions with this increased background also provided useful biological information as they correlated with areas where p53, a tumor suppressor protein , was active [61].
A different approach, focusing on the spatial distribution of molecules within breast tissue, has been used to assess a number of cell parameters including the size and dimensions of the cell and nucleus, distance between neighboring cells and Voronoi tessellation in order to characterize breast cancer cell types [94]. Larraona-Puy et al. have looked at the different distribution of key molecules in different tissue types, using a classification model based on bands associated with collagen, proline, protein backbone, and DNA to identify regions of the skin. Not only were they able to classify regions of normal tissue such as hair follicles, epidermis, and dermis, closely matching that of histologically stained tissue sections, but they could also identify regions of basal cell carcinoma [95].
Many cancer studies using Raman imaging have focused on the strength of, or lack of, particular signals in the Raman spectra. For example, in a study of cancerous and noncancerous tissue from salivary glands, the noncancerous tissue did not reflect the β-sheet protein content seen in the cancerous samples [96]. The presence of cancer cells in tissue, based on the acetylation and methylation of lysine residues in proteins contained in the cancer cells, has also been proposed [97]. On a cellular level, the presence and distribution of myeloperoxidase in both normal and abnormal promyelocytes, along with its absence in myeloblasts, could be used to identify each cell type as shown in Fig. 7.4. The numbers of each cell type found in bone marrow and peripheral blood smears can then be used for the diagnosis of myelodysplastic syndrome and acute myeloid leukemia [98].
Raman images from acute myeloid leukemia cells analyzed by Vanna et al. (a–d) Cells stained with May-Grünwald-Giemsa stain. (e–h) Bright field images before Raman measurement. (i–l) Hierarchical cluster analysis images. (m–p) multivariate images created from specific bands relating to nucleus (blue) and cytoplasm (green). For images (m–o), the presence of myeloperoxidase is shown in red. For image (p) the presence of hemoglobin is shown in red. Scale bar represents 10 μm (Figure reproduced from [98] with permission from the Royal Society of Chemistry)
Surmaki et al. noted that while many studies of cancer identification or development have concentrated on protein -based changes, lipids and carotenoid molecules are also significant players in the cell; in particular with regard to cancer development, they both have antioxidant properties that are important in the control of reactive oxygen species. In their study, Surmaki et al. used Raman maps to identify regions of carotenoids, lipids, and proteins with spectral differences seen for ductal and lobular carcenomas. They also noted that Raman images will incorporate information from the extracellular matrix, as well as the cells, providing additional information on the cellular environment [99]. Other studies have also focused on carotenoids and lipids in breast cancer, particularly looking at the nature of the lipids present in terms of saturation [100]. The ratio of the intensity of the nucleic acid peak at 785 cm−1 and the tryptophan peak at 752 cm−1 has also been proposed as a simple method for identifying skin tumors [101].
As cancer development is a multistep process, biochemical changes detected by Raman imaging can appear in areas that appear to be normal in morphological terms, with such changes potential markers for the early stages of the disease [102]. For example, an increased amount of lipid droplets in breast cancer cells has been correlated with the aggressiveness of cancer, suggesting that fatty acid synthesis could be an indicator of cancer development, particularly in the early stages of the disease [103]. An increase in lipid droplets has also been implicated in the development of colorectal cancer [104]. Raman imaging has also shown increased lipid content, lipid saturation, and phospholipid content in cells expressing high levels of the Her2/neu receptor, a proto-oncogene implicated in breast cancer, leading Hartsuiker et al. to suggest that the fatty acid content may be an indicator of metastatic potential in cells [105]. The metastasis of cancer cells from the primary tumor to secondary sites around the body is often associated with the later stages of cancer progression and can reflect poorer outcomes for the patient. The presence of metastatic cancer cells can also cause complications at the secondary site. Raman imaging has been used to monitor the effects of metastatic breast cancer cells on endothelial cells in the aorta of mice, where Raman analysis indicated an 18% increase in proteins and a 4% decrease in lipids for metastatic cancer cells. This change in endothelial cell composition may be a factor in the alteration of vasodilation that the authors note is associated with this disease stage [106].
There is currently a push to take Raman spectroscopy into the clinic, particularly to determine if tissue is cancerous to ensure all cancerous tissue is removed, while saving as much healthy tissue as possible, during surgery [107]. One of the main limitations for implementing Raman imaging in this context is the time required for measurements. However, advances such as the selective sampling approach outlined by Rowlands et al. [48] have been applied to basal cell carcinoma [107] and breast cancer tissue [108] potentially bringing the measurement time for large tissue sections, on the order of 1 × 1 cm2, down to as little as 20 min [107].
5.1.3 Lifestyle Diseases and Aging
Lifestyle diseases, including diabetes, atherosclerosis, hypertension, strokes, and heart attacks, are often linked to unhealthy lifestyle choices such as a high-calorie diet rich in fats and sugars, lack of exercise, as well as factors such as smoking and alcohol consumption. These diseases tend to manifest in later life and are becoming more prevalent, particularly in industrialized societies. The risk factors for many of these diseases are shared, and the presence of some of these diseases themselves can also be additional risk factors for other lifestyle diseases. For example, diabetes itself, along with the accumulation of lipids or cholesterol and endothelial damage, is an added risk factor for many other lifestyle diseases. Czamara et al. exploited the sensitivity of Raman to lipids to investigate the calcification process associated with aortic stenosis, finding colocalization between lipid and calcium (hydroxyapatite) even for the very small mineral deposits [109].
The relatively strong Raman signals generated by lipids in cells and tissues have been particularly beneficial for studies of atherosclerosis in both mouse models and human tissues. Kochan et al. [110] were able to show that while the livers of normal mice showed only small areas that were rich in lipids, the livers of atherosclerotic mice were rich in lipids, as were the livers of diabetic mice. In addition, even though the spectral overlap between vitamin A, essential for cell health [111, 112], and heme complicated matters, they were also able to visualize the vitamin A content in the mouse livers, highlighting a decrease in vitamin A content in early stages of disease [110]. Similar results were observed in the brains of mice where lipid to protein ratios were increased in animals in advanced stages of the disease [113]. A change in the structure of proteins contained within endothelial cells has been indicated to contribute to the stiffness of these cells during hypertension. Raman imaging has also identified changes in overall protein structure, α-helix and β-sheet [114], and amino acid composition [115], in mouse models of hypertension and atherosclerosis.
Raman imaging has been used to investigate the effects of sunlight on skin tissue. In a study by Ali et al., the lack of DNA bands in the basal layer was taken as evidence of DNA damage as a result of the solar radiation. The use of chemometric analysis methods also showed irradiation-induced changes in lipid content in the stratum corneum and the dermis [62].
Early identification of the complications of aging, such as the onset of Alzheimer’s or Parkinson’s diseases, is important in managing the disease progression and, consequently, the patient’s quality of life. Raman imaging has been used with the aim of resolving controversy surrounding the potential presence of amyloid-beta (Aβ) protein , characteristic of Alzheimer’s disease, in the cornea of affected patients. If Aβ was present in the eye, this could be an easily accessible tissue to monitor, in contrast to the brain. Techniques, such as staining, had given both positive and negative results for different researchers. Michael et al. then used Raman imaging to characterize the β-sheet to protein ratio for regions in the hippocampus with Aβ plaques or tangles, unaffected regions of the hippocampus, and the cornea. They found that the ratio was elevated in the regions where plaques and tangles were present, but unaffected areas of both the brain and the cornea had lower ratios, indicating no Aβ is present in the cornea [116].
5.1.4 Infectious Diseases
Identifying the presence of infective agents, such as bacteria or parasites, is important for implementing appropriate treatment, while analyzing the corresponding immune response of the infected cells and tissues provides information that may be beneficial for designing drug treatments or vaccines. Raman imaging has been used in a number of studies investigating the presence of bacteria in mammalian cells. For example, the growth of Staphylococcus aureus has been tracked over a 24-h time period, with internalization of the bacteria confirmed via multiple images taken at different planes through the cell to create a 3D image [117]. Both 2D and 3D imaging approaches were used to identify cellular changes associated with Mycobacterium gordonae, with the lipid response in the mammalian cells suggested to originate from lipid -filled phagosome or lipid bodies [118].
Raman imaging of Neospora caninum parasite infection in human brain microvascular endothelial cells was able to indicate the location of the parasite within cells through identification of the parasite nucleus and lipid -rich regions thought to be the parasite membrane. In addition, the presence of a Raman band at 1554 cm−1 was attributed to the presence of lysozyme, indicating Raman imaging can also identify a step in the immune response of the infected cells [119]. Toxoplasma gondii is another protozoan parasite that is known to infect a wide range of hosts and cell types. Cells infected with T. gondii showed increased protein and lipid content, suggesting these molecules are part of the cellular response to the parasite . However, increases in nucleic acid content in the Raman spectra were noted to be proportional to the parasite proliferation, therefore originating from the parasite itself [120]. In malaria, Plasmodium parasites produce hemozoin, a crystalline form of heme, during catabolisis of hemoglobin in infected red blood cells . Hemozoin gives rise to resonance Raman signals when using green excitation wavelengths, allowing low concentrations to be detected within cells. Imaging the presence of hemozoin in erythrocytes is possible, even though erythrocytes also contain high concentrations of hemoglobin, due to the small spectral differences between the two substances [121]. Macrophages, forming part of the immune defense, are known to engulf hemozoin particles released into the bloodstream during malaria, and Raman imaging has been used to identify the presence of hemozoin inside macrophages, along with characterizing the corresponding cellular rearrangements and changes in biochemical composition of the macrophage cells [122].
5.1.5 Drug Treatments and Interactions
The effectiveness of drug treatment is dependent on the chemicals reaching the target site in cells, tissues, and/or organs. Once in the appropriate location, the action of the drug will induce changes, potentially both chemical and morphological, in the surrounding cells and tissues. Conversely, if drugs accumulate in the wrong site, the induced changes in the surrounding cells and tissues will manifest as undesirable side effects. As Raman spectroscopy does not require a priori knowledge of the sample, it is an ideal method to monitor cellular changes induced by drug interactions. Chloroquine is used as a drug treatment for Plasmodium infections that cause malaria. Raman imaging of infected erythrocytes has been used to probe the oxygenation state of hemoglobin as a marker for the effects of the chloroquine exposure [123]. Oxygenation of heme-based molecules has also been used as a marker for the presence of Clostridium difficile infection. Infection also results in altered protein and threonine contributions to the Raman spectra, attributed to the presence of inflammation, with the threonine signal reduced after antibiotic treatment [124]. Raman spectroscopy has been used to monitor the effects of drug treatments in models of disease by monitoring vitamin A levels in hepatic stellate cells [125].
Other studies have noted that Raman imaging can monitor the effects of cancer drugs such as panitumumab on cells, allowing an assessment of their efficacy, even when the mode of action of the drug is not fully understood [126]. Anthracycline compounds are also potential therapies for cancer, as they inhibit topoisomerase, and have been shown to induce changes in the spectra originating from the cell nucleus in a concentration-dependent manner [127]. Two structurally related compounds, nobiletin and 5-demethlynobiletin, have been shown to have different effects, suggesting different modes of action, on cancer cell lines. Raman imaging studies showed that nobiletin induced changes in Raman bands originating from nucleic acids, while 5-demethylnobiletin affected localized lipids [128]. Nucleic acids were also the target for the action of indinavir and lopinavir in cervical cancer cells [129], and both nucleic acid and protein secondary structure changes were seen in myeloma cells after treatment with bortezomib, a proteasome inhibitor [130].
Photodynamic therapies, where a drug taken up into a cell or tissue is activated by light exposure to produce free radicals or reactive oxygen species to induce cell death in nearby cells, are used for cancer treatments. Brozek-Pluska et al. have used Raman imaging to identify the cancer cell composition, track the uptake of hematoporphyrin, and assess the effects on the surrounding tissue after photoactivation. They found cancerous regions were richer in saturated acids and proteins such as mammaglobin A and that cancerous cells took up larger amounts of hematoporphyrin compared to noncancerous regions of the tissue. Additionally, they showed that the Raman intensity of proteins and lipids in the surrounding regions did not alter significantly after photoactivation [131]. Raman imaging has also been able to show that cells with oncogenic mutations do not respond to drug treatments such as erlotinib, in the same way as normal cells do [132], indicating Raman analyses can provide insights into drug resistance as well as drug responses.
Depending on the chemical structure and concentration of a drug , it can be possible to directly measure the presence of the drug in the cell, facilitating simultaneous measurement of the drug position and the response of the surrounding regions of the cell. In drugs containing C≡O bonds, by integrating across 1945–1965 cm−1, the drug location can be visualized, since that spectral region does not overlap with signals from a typical cell. The drug location can be an important step in determining its mode of action, and, in this case, the authors were able to show that the drug accumulates both inside and directly surrounding the nucleus [133]. Although exhibiting some overlap with cell-based vibrations, the C=O bond in the cancer drug paclitaxel has also been used as an effective marker to track the number of cells taking up the drug and its location at different exposure times [134]. Another Raman imaging study, based on understanding the metabolism of erlotinib which contains a C≡C bond, have indicated that it is possible to assess how drugs are modified within the cell, after desmethyl-erlotinib was detected inside exposed cells [135].
The composition of drugs can be measured via Raman imaging, providing information on both the nature and the distribution of active ingredients, other tablet ingredients such as binding or dilution agents, and the nature of the polymer coating (if present). Vukosavljevic et al. used this approach to look at the composition of tablets containing verapamil hydrochloride, a calcium channel blocker, and were able to determine that drug release occurs through pores that form in the polymer coating of the tablet [64]. In order to understand the process of drug transfer from topologically applied drugs, e.g., those contained in lotions or creams, Goto, Morita, and Terada have used Raman imaging to follow the crystallization of components from creams containing urea as the active ingredient [136].
Modeling the uptake of nanoparticles is another potential application where Raman imaging can provide insight into the cellular response to particulate matter. Nanoparticles are sometimes employed as vehicles for drug delivery into cells, and Chernenko et al. have used Raman imaging to follow the uptake and metabolism of two nanocarriers (poly-(-caprolactone) (PCL) and poly(lactic-co-glycolic acid) (PLGA)) tracking their location within HeLa cells via Raman bands characteristic for the two molecules [137]. A significant impact of nanoparticles on human health is the potential detrimental effect on our cells and organs from environmental pollutants. Particulate matter in air, for example, has been shown to contribute to lung cancer [138]. Taking the subcellular environment and induced changes in metabolism as potential indicators of a toxic response, Dorney et al. investigated the local environment of polystyrene nanoparticles that were taken up into the cytoplasm of human lung adenocarcinoma cells [139]. Raman imaging studies of fibroblasts and macrophages showed differences in nanoparticle uptake when exposed to TiO2 nanoparticles , with fibroblasts storing the nanoparticles in small vesicles, whereas the macrophages, with a higher uptake of nanoparticles , distributed them in much larger vesicles [140]. Other studies have suggested that nanoparticles such as TiO2 and α-FeO(OH) can pass across the nuclear membrane and accumulate in the nucleus [141]. Extending these studies into multicellular organisms, cyanobacteria have been shown to internalize TiO2 nanoparticles [142]. The uptake of TiO2 nanoparticles by Caenorhabditis elegans has shown that despite the increase in sample complexity, Raman imaging can still provide information on the position of the nanoparticles as well as the changes they induce in the nematode itself, particularly with respect to proteins [143].
5.1.6 Hard Tissues
Raman imaging has been used to study various aspects of tooth structure in humans and primates. A recent study concentrated on the ratios of carbonate and phosphate in the dentine – enamel junction, as well as determining the presence of collagen as part of the organic component of teeth [144]. The composition of enamel in teeth determines tooth strength with hypomineralization linked to increased porosity, faster wear, and increased susceptibility to decay. Healthy enamel was shown to have strong spectral features attributed to phosphate-rich minerals, whereas lesions exhibited both carbonate substitution in the mineral components and an increase in protein content [145]. Similar mineral substitution has also been shown in sea urchin teeth where a simple band shift was sufficient to determine the presence of Mg2+ within the CaCO3 crystal structure [146]. In direct clinical applications, Raman imaging has identified the presence of pyrophosphate an atypical femoral fracture, with no pyrophosphate observed for control samples [147].
A particularly intriguing study used Raman imaging to assess structural defects created during tooth formation, based on the idea that these defects could be used as an assessment of illnesses and other stressors occurring during the early years of child development. The authors correlated the information from Raman imaging and other analytical techniques, such as mass spectrometry , with that of medical and behavioral data that noted the occurrences of stressful events in the lives of primates. Raman imaging was able to identify small features, primarily changes in protein and biomineral structure, that were not visible via light microscopy, and the resolution provided by the Raman images meant the size of the defect, and therefore the length of time the stressful event lasted could be estimated [148].
The repair of teeth, using resins, has also been studied by Raman imaging. Toledano et al. investigated dentin adhesive systems in terms of their ability to induce remineralization at the dentin interface [149]. In a further study, they found mechanical stresses (modeling the effect of chewing or grinding on the remineralization process) resulted in alterations of the mineral/matrix ratio [150].
Several studies have investigated the process of mineralization during bone cell differentiation. Hashimoto et al. followed the mineralization process during the first 72-h of cell differentiation, using Raman imaging to locate the presence of hydroxyapatite [151]. In a further study, they showed that the β-carotene concentration was proportional to the degree of mineralization, suggesting the presence of β-carotene could be used as an indicator for the initiation of mineralization . In addition, they also monitored cytochrome C as an indicator of apoptosis, with the onset of apoptosis in different cells linked to the fluctuations in cytochrome C intensity seen in the Raman spectra [7]. In another study of the mineralization process of bone nodules, Raman imaging identified the transition of hydroxyapatite from amorphous to crystalline during the 28-day monitoring period [152]. Extending this to more direct medical applications, Gao et al. monitored human mesenchymal stem cells, placed on a scaffold that would be implanted during bone regeneration therapies, to assess the effects of such a scaffold on osteogenic differentiation. They observed two stages of mineralization over a 28-day period and also noted that the mineralization process in cells on the scaffold was slightly delayed compared to that of cells on culture plates [153]. Of consequence for bioengineering, Raman imaging has shown that other substrates can also affect stem cells including strontium ion incorporation in bioactive glass resulting in increases in lipid rafts and cellular and membrane cholesterol [154].
Hair is an easily accessible sample for analysis. Wei et al. have investigated the potential for hair samples to provide information on the health status of an individual, namely, those suffering from rectal cancer. They found that different sections of the hair may be more useful than others for such studies in that there were little differences in cuticle between healthy people and cancer patients. The cortex and medulla, on the other hand, both showed changes in proteins, with cancer patient’s hair reflecting lower protein content in the cortex, less α-helical, and more β-sheet proteins in the medulla [155]. Other hair studies have focused on the distribution of particular compounds such as squalene, a lipid produced by skin cells in humans, noting it is primarily in the cortex [156]. Franzen et al. have also used Raman imaging to study the hair follicle, as it allows easy penetration into the skin for drugs, etc. They characterized the follicle in terms of the hair, sebum, and skin dermis and epidermis structures in both humans and pigs, as porcine skin is often used as a model for the human skin [63].
5.1.7 Plants, Algae, Molluscs, Bacteria, Fungi, and Small Multicellular Organisms
In plants, Raman imaging has primarily been used to investigate the composition of plant cell walls and other structural elements, with a focus on molecules including lignin and polysaccharides such as pectin and cellulose. The lignin and total carbohydrate content of cell walls in poplar trees has been described by Pererra et al. [157]. Analysis of Norway spruce and Scots pine showed similar lignin and cellulose distributions between both species, but spruce trees showed higher amounts of lignin-coniferyl aldehyde/alcohol groups in the secondary cell wall compared to the middle lamella and primary cell wall. Pine, on the other hand, showed comparable lignin-coniferyl aldehyde/alcohol content in the secondary cell wall and middle lamella, while the primary cell wall reflected lower levels [158]. The cell wall architecture in different regions of Miscanthus sinensis cv. has also been investigated with Raman imaging, providing information on the orientation of cellulose fibers with respect to the fiber axis [159]. The orientation of cellulose fibers was also important in brittle culm rice plant mutants, where cellulose fibers are arranged in a random orientation, as opposed to the parallel orientation found in wild-type plants [160]. Cellulose is an important source of ethanol for fuel during fermentation, and Chu et al. used Raman imaging to show that NaOH treatment of Miscanthus × giganteus samples removed lignin while leaving the cellulose behind [161]. Vermaak et al. have also imaged the distribution of molecules within plants, focusing on the steroidal glycoside p57 which has potential appetite suppressant properties, and found that p57 is primarily found in the inner fleshy part of the plant with low amounts in the outer layers [162]. In addition to the main plant, pollen grains are also amenable to Raman imaging studies. Analysis of intact grains provides information on the composition of the pollen grain walls, and molecular information on the substructure of the grains can be obtained from Raman images of sectioned grains [163].
Using tomatoes as a model system, Chylinska et al. identified the presence and distribution of pectin, cellulose, and polysaccharide molecules [164]. Tomatoes were also the focus for a recent study into ester cross-linking in cutin [165]. The distribution of pectins, cellulose, and hemicellulose in apples has been determined at different time points during the growth and storage of the fruit. These components are relatively evenly distributed throughout the young fruit. As the fruit matures, the pectins become more concentrated in the edges of the cell walls. During storage, the pectin distribution, once again, becomes more evenly distributed [166]. The distribution of plant cell wall polymers, namely, pectin and lignin, in different parts of the plant Phormium tenax is shown in Fig. 7.5. Higher pectin content in the lower regions of the plant (Fig. 7.5 P1) was attributed to higher water content. This was not found in higher regions of the plant (Fig. 7.5 P3 and P4), which show a higher lignin content, reduced water content, and increased mechanical stability [167]. The distribution of lipids and lignin has also been used to understand the waterproofing characteristics of Arabidopsis plants [168]. The rhizosphere bacterium Pantoea sp. YR343 has been studied in terms of its co-culture with Arabidopsis thaliana, with Raman imaging able to distinguish wild-type and mutant bacteria via the resonance Raman contributions originating from carotenoids [169].
Raman images obtained from different regions of the Phormium tenax plant. P1 is in the lower region of the stem, P3 from the upper region of the stem, and P4 from the leaf. ALL: intensities calculated by integrating between 2769 and 3087 cm−1 representing all cell wall polymers. CEL: intensities calculated by integrating between 1075 and 1114 cm−1 reflecting changes in cellulose orientation. PEC: intensities calculated by integrating between 842 and 872 cm−1 reflecting pectin content. LIG: intensities calculated by integrating between 1641 and 1546 cm−1 reflecting lignin content (Reproduced from Springer [167])
Raman imaging has been used to study algae, for example, to determine the proportions of silica, inorganic and organic components found in the cell walls of diatoms [170]. Monitoring the ratio of biomolecules in crops intended for producing biofuels is an important step in ensuring a good-quality fuel, for example, with a high calorific content, is produced. Chiu et al. used Raman imaging to assess the lipid and carbohydrate content in algal cells when subjected to stress. They showed that the lipid -to-carbohydrate ratio is affected by the nature of the stress applied, with nitrogen depletion resulting in increased carbohydrate content, but nitrogen depletion in combination with an increase in salinity increased the lipid content [171]. Polysaccharides are also a key component of fungi as well as plants and algae, and a recent study illustrated the ability of Raman imaging to simultaneously measure multiple polysaccharide components, including α- and β glucans and mannan, from fungal spores [172].
Raman imaging has also been used to investigate the nature and distribution of biochemicals within multicellular, living organisms. The connections between hard and soft tissues in molluscs have also been visualized by Raman imaging to investigate the interface between the living tissue and shell [173]. Raman imaging can also be used to study the whole organism as shown by Nakamura, Hotta, and Oka who followed the development of the Ciona intestinalis (sea squirt) embryo during development from the two-cell stage through to the tail bud stage. They were able to identify muscle and ectoderm at multiple stages throughout the development and to discriminate these cells from others that had divided from the same parent cell but that had different developmental fates [174]. Nematodes are particularly accessible samples for Raman imaging as they are usually semitransparent. Raman imaging has also been employed to assess the changes in Radopholus similis nematodes, a banana plant parasite , to the anigorufone compounds produced by the banana plants in response to a nematode infection. Exposed nematodes showed an increase in lipid -rich droplets which the authors suggest may be a defense mechanism to limit the toxic effects of the anigorufone on the nematode [175].
5.1.8 Food and Drug Quality and Safety
Raman imaging has been used in a number of studies investigating the distribution of components within the sample. Smith et al. have imaged the distribution of fat, protein, and water in processed cheese , along with additives such as trisodium citrate, starch, and paprika. [176]. Carotenoids are responsible for the color in fruit and vegetables such as tomatoes and peppers and are an important part of our diets as they have protective capabilities including antioxidant properties and vitamin A activity. Raman imaging has been used to study the nature and subcellular location of carotenoids in chili peppers , as the subcellular location has been proposed to affect the dietary availability of the carotenoids. Carotenoids were located in lipid bodies that were outside of the chromoblast. The pepper varieties studied (four red and one orange when ripe) were found to contain different carotenoid compounds. For example, lutein was the main carotenoid component of Costeno-Amarillo, whereas NuMex Heritage 6-4 was rich in β-carotene [177].
The incidence of several food scandals in recent years, such as melamine in powdered milk in China and horse meat in meat products in the UK and Europe, has raised questions about the origins and quality of our food products. A number of studies have shown that Raman imaging can be a useful tool for quality control and the identification of contaminants in a number of food items. Quin et al. tested the potential for Raman imaging to determine the presence of contaminants such as ammonium sulfate, dicyandiamide, urea, and melamine in dry milk. Taking advantage of the well-defined Raman spectrum of the contaminants, compared to the broader-featured milk powder spectrum, they could simultaneously detect these contaminants at concentrations between 0.1% and 5.0% [178]. Extending this, Dhakal et al. tested the potential for Raman imaging to determine the presence of melamine in spiked samples of skimmed milk powder . Using a thresholding approach on the images, they were able to detect melamine at concentrations as low as 0.005% [179].
The compound benzylisothiocyanate (BITC) has antimicrobial properties. Clement et al. used Raman imaging to investigate the action of BITC when incorporated into food packaging, with a focus on its effects on the food-borne mold Aspergillus ochraceus. They showed BITC-induced spectral changes associated with most classes of macromolecule – saccharides, proteins, lipids, amino acids, and so on – indicating BITC disrupts multiple cellular processes in A. ochraceus, which will ultimately lead to the death of the microbe [180].
Not all food adulterations are dangerous to human health such as those that involve the substitution of one food product with another, usually cheaper, food product. Eksi-Kocak et al. have noted that pistachio nuts, as a more expensive crop, are often bulked out by the addition of green peas which exhibit similar green-colored kernels. They were able to use spectral feature differences observed across small areas of Raman images to identify the presence of both pistachio and green pea in mixtures of different proportions [181].
Raman imaging has also been applied to the analysis of drug mixtures in both genuine and counterfeit tablets. Kwok and Taylor compared genuine Cialis tablets, containing the active ingredient tadalafil and binding agents lactose monohydrate and microcrystalline cellulose, to two fake tablets. These fake tablets were found to contain talk, starch, calcium sulfate, and several unknown ingredients in addition to varying amounts of the active ingredient. They were also able to show that the genuine tablets had a much more even distribution of the ingredients compared to the counterfeit tablets [182]. Sacré et al. showed that the spatial differences in the chemical distribution of sildenafil in counterfeit and genuine Viagra tablets were not sufficient to discriminate between the two groups; however, the presence of lactose was a clear marker for counterfeit tablets [183].
5.2 Key Research Findings in Nonlinear Raman Imaging
5.2.1 Coherent Anti-Stokes Raman Scattering (CARS) Imaging
The sensitivity of CARS to lipids has been shown to visualize the accumulation of lipids in adipocytes with higher definition and at an earlier stage to oil red O staining commonly used for marking lipids [184]. This sensitivity has meant that CARS imaging has predominantly focused on areas where the presence of lipids or lipid metabolism is a key component of the sample. Bradley et al. have used CARS to monitor the early stages of early embryo development. They tracked the distribution of lipid droplets showing that the early embryo, at the two and four cell stages, exhibit small lipid droplets. As development continues, cells exhibit fewer droplets but these are larger in size. They also noted that the composition of the lipid droplets varied, with different ratios of saturated and unsaturated lipids found in droplets from the same embryo [185]. Unsaturated fatty acids were the focus of another study that investigated the nature of the fatty acids that were involved in the transport of yolk lipoprotein into C. elegans oocytes. Through the analysis of mutants deficient in polyunsaturated fatty acid synthesis, the authors were able to determine that omega-6 fatty acids were a key regulator of this fatty acid delivery process [186].
Jüngst et al. exploited the sensitivity of CARS to lipids in order to monitor the fusion process of lipid droplets in adipocytes, observing the cells over a 120-h period. They found that the increase in size of lipid droplets was mainly due to fusion of smaller droplets, rather than being due to synthesis of new lipids. They showed that the transfer of material was from the smaller droplets to the larger droplets, with the rate of transfer dependent on a number of parameters including the relative size difference and the diameter of the smaller droplet [187]. A similar study also monitored lipid droplets during the development of adipocytes, this time illustrating that the nature of fatty acids present in the cell culture medium can influence the fatty acid composition of the lipid droplets. The maturation stage of the adipocyte was also shown to be a determining factor in the speed of the cellular response to the change in fatty acid composition of the extracellular environment [188]. The manifestation of many lifestyle diseases is accompanied with changes in lipid signals. CARS imaging has identified increased lipid signals in brain samples from mice suffering from Alzheimer’s disease [189].
Lipid droplets are also important in algae, and Cavonius et al. were able to identify three phases in the synthesis process. They employed time gating in order to measure the lipid signals despite the strong pigmentation from the chloroplasts, but they did note that, when measuring time-lapse images of live algal systems, the extended exposure to the CARS excitation light is likely to have an effect on the light harvesting of the chloroplasts [190].
The myelin sheath surrounding nerves also gives rise to strong CARS signals due to its fat content. Berlanger et al. used CARS to image the myelin content of the spinal cord in live mice [191]. Turcotte et al. used polarization-resolved CARS to study myelination in live zebra fish and to assess promyelination drugs [192]. CARS imaging of lipids had also been used to indirectly measure the sweating process by covering a small area of skin with an exogenous oil, thereby filling the sweat pores. The oil can then be imaged as it is flushed out by the discharge of sweat [193]. Although CARS-based research is dominated by lipid imaging, a few studies have been engaged in determining the distribution of other molecular classes. Wang et al. looked at the distribution of pheomelanin, responsible for red hair pigmentation, in melanocytes, amelanotic melanomas, and a red-haired mouse model [194].
As with conventional Raman imaging, CARS can be used as a method to assess the impact of nanoparticle uptake on tissues. Johnston et al. were able to locate the presence of both TiO2 and Au nanoparticles in macrophages, hepatocytes, and live animals with the CARS signals able to visualize both the cells and the position of the nanoparticles [195]. An investigation into the uptake of environmental microplastics by crabs has also used CARS to locate the particles within the crab [196].
Some CARS studies have also used spontaneous Raman measurements (usually point spectra) to provide spectral information while still retaining the speed of CARS measurement. This approach has been used to identify brain tumors, with tumor boarders and regions of infiltration imaged at the cellular level. The CARS signal was moderately reduced in metastatic tumors and markedly reduced in the glioblastoma regions, with Raman spectroscopy showing that this reduction in CARS signal was due to a lower lipid content in the tumors [197]. FAST (femtosecond adaptive spectroscopic techniques) CARS imaging and Raman spectroscopy have been used in conjunction to distinguish between polarized and non-polarized epithelia in breast tissue, based on the lipid ordering present [17]. In studies of breast and prostate cancer cells, CARS images showed an increase in the number and size of lipid droplets, while Raman spectroscopy indicated an increase in saturation of the lipids present, after treatment with the drugs medroxyprogesterone acetate or synthetic androgen R1881 [198].
A method for increasing the spectral information provided by CARS is to measure broadband CARS, which produces spectra that are more akin to conventional Raman spectra, with the first report showing a vibrational spectrum covering the region from 600 to 3200 cm−1 [199]. This opens up the technique for more detailed investigation of protein structure, as the peak shifts associated with α-helix, β-sheet, and other protein structural forms can be measured. Bito et al. found that the cortex of hair is mainly α-helical, while the cuticle is richer in β-sheet and random coil structures, with an α-helix to β-sheet transition observed after chemical and mechanical treatments [200]. Several recent studies have begun to explore the possibilities of combining CARS with chemometric analysis techniques. Lin et al. applied PCA to CARS images of the meibum lipids produced by meibomian glands in the eyelid, using the lipid composition to identify the different functional regions within the gland [201]. El-Mashtoly et al. focused on the 2700–3000 cm−1 region and used a random forest approach to automate identification of common cellular organelles such as the endoplasmic reticulum, lipid droplets, nucleus, and nucleolus [202].
5.2.2 Stimulated Raman Scattering (SRS) Imaging
SRS has been used to visualize different biomolecules within food products . Roeffaers et al. were able to image the two phases in mayonnaise, fat and water, as an example of a simple emulsion. SRS images could also be obtained from cheese and a soy-based drink which have extra components, proteins and carbohydrates, as well as fat and water [203]. In plants , SRS has been shown to discriminate between different wax components, cellulose and pectins [204], and to assess effects of maleic acid treatment on lignins in cell walls of poplars [205]. SRS has also been used to investigate the uptake of the fungicides azoxystrobin and chlorothalonil in plants, both of which contain C≡N bonds allowing clear identification of the peaks in the “silent region” of the spectrum [206].
Egawa et al. have used SRS to study pig and human skin with a particular focus on the various stages of differentiation in keratinocytes [207]. SRS has also been used to monitor the changes occurring in mammalian tissues during disease manifestation. Tian et al. studied the nerve degeneration occurring in amyotrophic lateral sclerosis (ALS) in mouse models of the disease. They found changes in the peripheral myelin, with lipid depositions appearing in ALS-affected individuals. In addition, they monitored the effects of treatment with minocycline, a drug known to slow the progression of ALS in mice, which appeared to reduce the number of lipid deposits observed, but did not reduce them to the levels seen in unaffected individuals [208]. SRS has also been used to investigate the lipids present in ovarian cancer stem cells, with cells such as ALDH+/CD133+ cells that have high tumorigenic and metastatic potential [209] found to have a high degree of unsaturated lipids [209]. Li et al. also noted that the degree of saturation in lipids could be indicative of the metabolism after noting COV362 cells grown in a monolayer had lower levels of lipid unsaturation compared to those grown as spheres [210]. Many molecules give rise to Raman signals between 2800 and 3000 cm−1, but Lu et al. have exploited the small shifts in position for each of the main classes of molecules to measure signals from DNA with SRS. They showed that it was possible to measure SRS signals from DNA in a sample of mixed components, within a cell, and within skin samples, as well as using SRS time-lapse imaging to monitor chromosome dynamics [211].
As with CARS, a few recent studies have been extending the analysis of SRS signals through the application of chemometric analyses. Satoh et al. have used PCA , concentrating on the 2800–3100 cm−1 region, to visualize acetaminophen-induced liver injury in mice [212]. Datta et al. used VCA to identify lipid droplets within a protein matrix when investigating metabolism and oxidative stress in cardiomyocytes derived from pluripotent stem cells [213].
5.3 Multimodal Imaging
Although the speed at which Raman imaging can be performed has increased with developments such as line scanning and improvements in detector sensitivity, it is still slow in comparison to many cellular processes. One way to compensate for this is to combine Raman with other modalities that have faster imaging capabilities and provide additional information on the nature of the sample. Multimodal approaches to Raman measurement can ideally combine two or more techniques that are applied to the sample simultaneously, so that the information between the different modes can be most easily compared. The instrumental challenges of that approach can be significant [214], but highly useful measurements can still come from multimodal techniques carried out sequentially on a sample [215]. Indeed, by relaxing the constraint of measuring simultaneous multimodal information, a wider variety of modalities may be used. For example, fluorescent staining measured following Raman can be a very useful validation technique.
One of the big challenges with multimodal imaging is the image registration between different measurement techniques. This is particularly problematic where one modality is used to provide information on what another modality is measuring as differences in the z-resolution, and probed volume will mean that there will be differences in the morphological and, potentially, molecular species measured by each technique [79, 214].
5.3.1 Raman and Infrared (IR) Spectroscopy
Raman and IR spectroscopies are both forms of vibrational spectroscopy, but as Raman imaging is based on a scattering process and IR imaging is based on absorption, they can be used to provide complimentary information on a sample. IR imaging is much more sensitive to the presence of water, meaning that it can be difficult to measure aqueous samples, restricting many of the studies to tissue sections or other samples that can be dried prior to measurement.
Garip et al. have used both Raman and IR imaging to investigate the changes in rat liver during treatment with simvastatin . Both Raman and IR images showed varying decreases in lipids and proteins, leading to an overall increased lipid :protein ratio, and Raman analysis also suggested a relative increase in DNA to proteins. The authors concluded that simvastatin causes notable damage to the liver, possibly through oxidative stress mechanisms [216]. Lau et al. used Raman and IR in conjunction to study the distribution of biomolecules within Steinernema kraussei nematode worms. Due to the different measurement configurations, confocal Raman images provided high spatial resolution images in a single plane through the nematode . IR images performed in transmission mode, on the other hand, provided complementary information in terms of the biochemical composition of the whole thickness of the worm [217].
In terms of hard tissues, Raman and IR imaging have been used in conjunction to study untreated human hair with both techniques providing information on the protein secondary structure and lipid composition of hair fibers. In addition, due to the fact that Raman is sensitive to disulfide linkages, Raman images also provided information on keratin, which is rich in cysteine amino acids [15]. Bone tissue engineering was the focus of a study concentrating on determining the biocompatibility of a composite made from hydroxyapatite and β-glucan. Here, Raman imaging was used in conjunction with infrared spectroscopic imaging to investigate the differences between the composite and decalcified bone, identifying subtle changes in the mineral composition and a decrease in phosphate contributions with increasing age of the sample [218]. Chia et al. have also used Raman and IR imaging to study the formation of biochar (produced by heating biomass in limited oxygen environments) used as a soil additive. FTIR was able to show the effects of mineral additives on heat-induced chemical changes in the biochar, while Raman data provided information on the carbon structure in biochar and mineral-enriched biochar [219].
Raman and IR imaging have also been combined with additional modalities . Caine et al. added CARS and fluorescence imaging in order to investigate the alterations to the brain caused by a stroke. Raman provided high spatial resolution imaging of the lipid acyl groups revealing that the neuropil within in the peri-infarct zone showed reduced amount of lipid acyl groups. CARS showed that both the ischemic infarct and the surrounding peri-infarct zone were mainly devoid of myelinated axons. This information was complimented by IR and fluorescence imaging of the same areas [220]. Palombo et al. combined the chemical information obtained from Raman and IR imaging with Brillouin imaging, providing information on the mechanical properties, primarily the viscoelasticity, of Barrett’s esophagus samples [221].
5.3.2 Raman and Mass Spectrometry (MS) Imaging
In recent years, mass spectrometry imaging has been applied to a number of biological samples (for a recent review, see Spengler et al. [222]) with a small number of studies looking to combine the mass-based sample information from MS with the vibrational information from Raman imaging. Lanni et al. combined secondary ion mass spectrometry (SIMS) with Raman in order to image bacterial biofilms of Pseudomonas aeruginosa, identifying components such as quinolones in the sample [223]. Ahlf et al. used an alternative MS modality, matrix-assisted laser desorption ionization (MALDI), combined with Raman for the analysis of 3D cell cultures. In this case, unlike Lanni et al., Ahlf et al. did not directly correlate the Raman and MS images, instead analyzing both modalities with PCA and then correlating the components obtained. For both techniques, the main variation, identified by the first principal component, identified the necrotic core of the culture, while the second component was associated with the proliferating edge [224]. Bocklitz et al. have proposed a workflow for combining Raman imaging with MALDI-based MS imaging, using a larynx carcinoma sample as their target. In this workflow, Raman images were compared to the H&E-stained images, with Raman identifying two layers in the epithelium attributed to differences in metabolic state. MALDI images were then taken from regions where the Raman and pathologist classifications were in agreement and from regions where the two did not agree, with m/z values from the MALDI imaging providing additional information to aid in classifying the tissue [225].
5.3.3 Raman and Atomic Force Microscopy (AFM)
AFM probes the mechanical properties of a sample such as viscoelasticity, stiffness, and topography [226]. Recently, the information provided by AFM imaging has been used to try and decouple the molecular concentration of components within cells from the topography of the sample in Raman images, as a move toward quantitative Raman measurements that are completely independent of sample thickness. In this case, correlated Raman and AFM images were used to produce quantitative maps of the cellular distribution of components such as proteins and RNA [227]. Correlated AFM and Raman images have also been used to study the effects of ex vivo treatment methods, particularly those commonly used in cell fixation protocols, on human erythrocytes [228]. Raman and AFM have also been used to assess the oxidation state of heme in erythrocytes [229].
Raman and AFM studies have often included IR imaging analyses as well . One such study used all three techniques to investigate the content and distribution of usnic acid, which has potential antibiotic properties, in three different species of lichen [230]. Raman imaging and AFM have also been used in conjunction with IR images to study mycobacterium and gram-negative bacteria , Pseudomonas putida and Escherichia coli in co-culture [231].
5.3.4 Raman and Digital Holographic Microscopy (DHM)/Quantitative Phase Imaging
Digital holographic microscopy (DHM) is a quantitative , label-free, and noninvasive technique that provides information in terms of the phase shifts induced by a sample [232]. DHM can be performed at high frame rates, making it an ideal complement to the relatively slow collection times required for Raman imaging. The implementation of a dual DHM and Raman system has been achieved using the same light source for sequential measurements [233] and by using two light sources and spectral separation allowing for simultaneous measurements from the two modalities [232]. The quantitative phase image information provided by DHM has been used to screen red blood cells infected with malaria, identifying cells of interest which can then be further analyzed by Raman to provide chemical and spatial distribution of hemozoin, for example [233]. Simultaneous measurements have provided information on the classes of biomolecules that give rise to the phase signals. Raman spectra of cells are often lipid -rich, with moderate-intensity bands from proteins and smaller contributions from other molecules such as nucleic acids. On the other hand, proteins and nucleic acids generate a significant proportion of the contrast in phase images, while the contribution from lipids is less intense. This highlights an additional aspect of the complementarity between the two imaging modes [232]. The combination of Raman and DHM has also been used to assess the health of bovine sperm cells during selection for artificial insemination. In this case, the morphology of the cells, provided by DHM, gave insight into the health of the sperm, and the Raman images and spectra were used to identify the X-carrying and Y-carrying sperm [234].
5.3.5 CARS, Second Harmonic Generation (SHG), and Two-Photon Emission Fluorescence (TPEF)
CARS has often been used in conjunction with second harmonic generation (SHG) and two-photon emission fluorescence (TPEF) imaging . This triple combined approach has been used to study the muscle tissue changes induced by cold shock in crustaceans, with CARS providing information on the ER , cisternae, and T-tubules [235]. All three techniques are able to identify the epidermis, dermis, and subcutis in the human skin, but Heuke et al. demonstrated that an overlay of all three images combined provided clear delineation between the tissue types, illustrating this with example regions including a sweat gland, sebaceous gland, hair bulb, and blood vessels [60]. In cancer applications, the three techniques have been used to discriminate between normal, desmoplastic tissue and cancerous lesions in the lung [236] and to identify normal tissue, tumor, and necrotic tissue in the kidney [237]. In atherosclerosis studies, CARS has been used in conjunction with TPEF and SHG to study the plaque deposits [238, 239]. The vitamin A content of pig liver has also been investigated with all three modalities where CARS produced images of the vitamin A droplets, fluorescence imaging highlighted the hepatocytes and liver sinusoids, and SHG showed the location of connective tissues [240]. Chernovskaia et al. combined CARS at two different wavelengths, with TPEF also at two different wavelengths and SHG to provide in total five image channels for the investigation of inflammation in diseases such as Crohn’s disease and inflammatory bowel syndrome as shown in Fig. 7.6. The authors used the information from these combined modalities to predict different levels of chronicity, architecture, and activity associated with the samples [241].
Multimodal images (a, c, d, f, h, i) and corresponding H&E-stained tissue sections (b, e, g, j) of normal (a–e) and diseased (f–j) colon mucosa. Sub-figures (a) and (f) show multimodal images acquired with CARS at 2930 cm−1, TPEF at 503–548 nm, and SHG. Sub-figures (c) and (h) show details from (a) and (f) represented by the white boxes. Sub-figures (d) and (i) show multimodal images corresponding to sub-figures (c) and (h), respectively, that were acquired with the combination of CARS at 2850 cm−1, TPEF at 426–490 nm, and SHG. Red, CARS; green, TPEF; blue, SHG. White scale bar represents 1 mm and black scale bar 250 μm. Arrows: 1, lymphocytes, increase overall fluorescence, loss of crypt density; 2, crypt branching; 3, irregularities of crypts’ shape; 4, flattened epithelial layer facing the lumen (Figure reproduced from Nature Publishing Group [241])
CARS and TPEF have been used together to study the reorganization of the major classes of biomolecules in cells during the cell cycle. Along with quantitative analysis of concentrations of these molecules at particular points in the cell provided by Raman spectra, the study showed that proteins are inhomogenously distributed throughout the cell and that lipids tend to be more abundant at the edges of the cell. In addition, only the local concentration of DNA was found to fluctuate during the mitosis and G1 phases of the cell cycle [242].
CARS and SHG have been used to provide complimentary information on fibrosis in nonalcoholic fatty liver disease with SHG providing information on the nature of the fibrosis and CARS identifying the liver fat [243]. In a mouse model of obesity, both techniques were used to visualize the increase in size of adipocytes induced by high fat and fructose diets [244]. The combination of CARS and SHG has also been employed in several studies centered on proteins. Fibrin hydrogels are one type of substrate used as a scaffold in regenerative medicine, and Mortati et al. used SHG and CARS to follow the collagen production of mesenchymal stem cells, as an indicator of cell differentiation, and corneal fibroblasts in 3D cultures on these scaffolds [245]. Brackmann et al. used CARS and SHG in combination to investigate orange fruits/vegetables such as sweet potatoes, carrots, and mangos , with CARS used to visualize the carotenoids and SHG highlighting the starch granules in each sample [246]. Brackmann et al. also looked at the interaction of cells with a matrix, this time investigating the interactions of smooth muscle cells with a cellulose matrix in terms of how they compress the matrix [247].
Mansfield et al. have compared CARS and SRS for imaging of superficial zone chondrocytes showing that SRS provides a spectrum closer to that of the Raman and also gives rise to a clearer image. They then went on to combine TPEF, SHG, and SRS imaging to investigate the structure of cartilage, with three SRS channels providing images of carbonate, phosphate, and CH stretching. The TPEF images provided information on autofluorescence, while the SHG images highlighted the collagen fibers [248].
6 Conclusions and Future Perspectives
Raman imaging has been applied to a wide range of sample types and fields of research that all have an impact on human health. That includes a growing variety of applications including fundamental studies of biological reactions, through the assessment and quantification of disease states and the impact of environmental toxins on biological systems or more indirectly through determining the quality of foodstuffs and drugs. Future developments in Raman imaging will, no doubt, continue to expand the applications of Raman analyses to other fields of biochemical analysis.
In terms of applications for medical diagnosis, many studies have shown the potential for Raman in characterizing changes in cells and tissues that can be thought of as markers for diseases as wide ranging as cancer, atherosclerosis, dental lesions, and parasitic infections. The next stage will be to expand these small-scale studies to larger sample sizes and more complex sample groups (e.g., samples from patients who exhibit symptoms from multiple medical conditions) with a view to applying Raman-based diagnostics in clinics and surgeries. In many cases, this will involve the development of smaller, dedicated instruments and the development of automated software for determining the status of a sample. While much of the instrumentation currently being developed for use in surgery or other clinical settings are primarily concerned with recording single point spectra, the information provided by Raman mapping and imaging studies is useful when developing such probes [249], particularly in terms of the spatial distributions and concentrations of the key molecules of interest in a given sample.
In terms of instrumentation, it is challenging to directly improve the amounts of acquired Raman signal, since for many setups, detector sensitivity, grating efficiencies, and so on are relatively close to what might be expected theoretically. Instead, improvements in Raman imaging are likely to come from improving the speed of measurement via measures such as increased degree of parallel acquisition (i.e., in the same way that line scanning drastically improved Raman imaging speed, other improvements might be implemented) or by increasing resolution [250]. Additionally, improvements are already underway by use of smart selective sampling, compressed sensing, or other techniques to reduce the number of spectra required to construct an image. In essence, these directions involve the optimization of the desired information content that can be measured in a constrained set of parameters, where laser power, sample damage, and measurement time are all crucial considerations. The optimum measurement would then provide the best answer to a given hypothesis, by optimizing the measurement protocol according to the physical constraints. Most current work does not take this approach, partly because the optimum measurement may require a flexible and customizable imaging system. There is then a trade-off between the best-case measurement approaches and practical considerations. Nevertheless, we can expect significant improvements in Raman imaging, with increased attention to exploiting these factors. In addition, combining Raman imaging with other modalities, providing correlated but nonredundant information, will be important in future studies, particularly where the other modalities measure different characteristics of the sample, compared to the Raman spectra. In terms of live sample imaging, complimentary modalities with high frame rates will be particularly useful characterizing sample changes taking place during the Raman imaging measurement, currently on the order of a few minutes for a typical cell.
The complimentary information obtained from multimodal imaging, particularly when measurements are taken simultaneously, will also further our understanding of what biochemical information is contained within the Raman images. As Raman spectra contain contributions from all Raman-active molecules in a sample, they are rich in information but complex to interpret. Chemometric algorithms are very useful for decomposing the spectra into various types of components, but these component spectra are also composed of contributions from mixtures of molecules. Therefore, correlating Raman images with those obtained from other techniques will allow us to more accurately identify the nature of the molecules contributing to particular Raman signals. This knowledge can be transferred back to single modality Raman studies, allowing more detailed interpretation of the biochemical composition of samples.
References
Smith E, Dent G (2005) Modern Raman spectroscopy a practical approach. Wiley, Chichester
Grasselli JG, Snavely MK, Bulkin BJ (1980) Applications of Raman spectroscopy. Phys Rep 65(4):231–344
Mantsch HH (2013) The road to medical vibrational spectroscopy – a history. Analyst 138(14):3863–3870
Edwards HGM (2009) Raman spectroscopy of inorganic materials in art and archaeology: spectroscopic analysis of historical mysteries. In: Yarwood J, Douthwaite R, Duckett SB (eds) Spectroscopic properties of inorganic and organometallic compounds. Techniques, materials and applications. RSC Publishing, Cambridge, UK
Pully VV, Lenferink ATM, Otto C (2011) Time lapse Raman imaging of single live lymphocytes. J Raman Spectrosc 42(2):167–173
Okada M, Smith NI, Palonpon AF, Endo H, Kawata S, Sodeoka M, Fujita K (2012) Label-free Raman observation of cytochrome C dynamics during apoptosis. Proc Natl Acad Sci U S A 109(1):28–32
Hashimoto A, Yamaguchi Y, Chiu L-d, Morimoto C, Fujita K, Takedachi M, Kawata S, Murakami S, Tamiya E (2015) Time-lapse imaging of osteoblast differentiation. Sci Rep 5:12529
Yamakoshi H, Dodo K, Palonpon A, Ando J, Fujita K, Kawata S, Sodeoka M (2012) Alkene-tag Raman imaging for visualisation of mobile small molecules in living cells. J Am Chem Soc 134(51):20681–20689
Ando J, Fujita K (2013) Metallic nanoparticles as SERS agents for biomolecular imaging. Curr Pharm Biotechnol 14(2):141–149
Laing S, Gracie K, Faulds K (2016) Multiplex in vitro detection using SERS. Chem Soc Rev 45(7):1901–1918
Lane LA, Qian X, Nie S (2015) SERS nanoparticles in medicine: from label-free detection to spectroscopic tagging. Chem Rev 115(19):10489–10529
Baker MJ, Hussain SR, Lovergne L, Untereiner V, Hughes C, Lukaszewski RA, Thiėfin G, Sockalingum GD (2016) Developing and understanding biofluid vibrational spectroscopy: a critical review. Chem Soc Rev 143:2238–2247
Gamsjaeger S, Mendelsohn R, Boskey AL, Gourion-Arsiquaud S, Klaushofer K, Paschalis EP (2014) Vibrational spectroscopic imaging for the evaluation of matrix and mineral chemistry. Curr Osteoporos Rep 12(4):454–464
Butler HJ, Ashton L, Bird B, Cinque G, Curtis K, Dorney J, Esmonde-White K, Fullwood NJ, Gardner B, Martin-Hirsch PL, Walsh MJ, McAinsh MR, Stone N, Martin FL (2016) Using Raman spectroscopy to characterise biological materials. Nat Protoc 11:664–687
Zhang G, Senak L, Moore DJ (2011) Measuring changes in chemistry, composition and molecular structure within hair fibres by infrared and Raman spectroscopic imaging. J Biomed Opt 16(5):056009
Teodori L, Crupi A, Costa A, Diaspro A, Melzer S, Tarnok A (2017) Three-dimensional imaging technologies: a priority for the advancement of tissue engineering and a challenge for the imaging community. J Biophotonics 10(1):24–45
Yue S, Cárdenas-Mora JM, Chaboub LS, Lelièvre SA, Cheng J-X (2012) Label-free analysis of breast tissue polarity by Raman imaging of lipid phase. Biophys J 102(5):1215–1223
Charwat V, Schütze K, Holnthoner W, Lavrentieva A, Gangnus R, Hofbauer P, Hoffmann C, Angres B, Kasper C (2015) Potential and limitations of microscopy and Raman spectroscopy for live cell analysis of 3D cultures. J Biotechnol 205:70–81
Draux F, Gobinet C, Sulé-Suso J, Trussardi A, Manfait M, Jeannesson P, Sockalingum GD (2010) Raman spectral imaging of single cancer cells: probing the impact of sample fixation methods. Anal Bioanal Chem 397(7):2727–2737
Hobro AJ, Smith NI (2017) An evaluation of fixation methods: spatial and compositional cellular changes observed by Raman imaging. Vib Spectrosc 91:31–45
Bogliolo L, Murrone O, Piccinini M, Ariu F, Ledda S, Tilocca S, Albertini DF (2015) Evaluation of the impact of vitrification on the actin cytoskeleton of in vitro matured ovine oocytes by means of Raman microspectroscopy. J Assist Reprod Genet 32(2):185–193
Gaifulina R, Maher AT, Kendall C, Nelson J, Rodriguez-Justo M, Lau K, Thomas GM (2016) Label-free Raman spectroscopic imaging to extract morphological and chemical information from a formalin-fixed, paraffin-embedded rat colon tissue section. Int J Exp Pathol 97(4):337–350
Ali SM, Bonnier F, Tfayli A, Lambkin H, Flynn K, McDonagh V, Healy C, Lee TC, Lyng FM, Byrne HJ (2013) Raman spectroscopic analysis of human skin tissue sections ex-vivo: evaluating the effects of tissue processing and dewaxing. J Biomed Opt 18(6):061202
Ali SM, Bonnier F, Lambkin H, Flynn K, McDonagh V, Healy C, Lee TC, Lyng FM, Byrne HJ (2013) A comparison of Raman, FTIR and ATR-FTIR microspectroscopy for imaging human skin tissue sections. Anal Methods 5:2281–2291
Esmonde-White KA, Esmonde-White FWL, Morris MD, Roessler BJ (2014) Characterisation of biofluids prepared by sessile drop formation. Analyst 139(11):2734–2741
Palonpon AF, Ando J, Yamakoshi K, Dodo K, Sodeoka M, Kawata S, Fujita K (2013) Raman and SERS microscopy for molecular imaging of live cells. Nat Protoc 8:677–692
Gasparov L, Jegorel T, Loetgering L, Middey S, Chakhalian J (2014) Thin film substrates from the Raman spectroscopy point of view. J Raman Spectrosc 45(6):465–469
Draux F, Jeannesson P, Beljebbar A, Tfayli A, Fourre N, Manfait M, Sule-Suso J, Sockalingum GD (2009) Raman spectral imaging of single living cancer cells: a preliminary study. Analyst 134(3):542–548
Fogarty SW, Patel II, Martin FL, Fullwood NJ (2014) Surface-enhanced Raman spectroscopy of the endothelial cell membrane. PLoS One 9(9):e106283
Bonnier F, Knief P, Lim B, Meade AD, Dorney J, Bhattacharya K, Lyng FM, Byrne HJ (2010) Imaging live cells grown on a 3D collagen matrix using Raman microspectroscopy. Analyst 135(12):3169–3177
Lachin JM (1981) Introduction to sample size determination and power analysis for clinical trials. Control Clin Trials 2(2):93–113
Lenith RV (2001) Some practical guidelines for effective sample size determination. Am Stat 55(3):187–193
Dell RB, Holleran S, Ramakrishnan R (2002) Sample size determination. ILAR J 43(4):207–213
Beletes C, Neugebauer U, Bocklitz T, Krafft C, Popp J (2013) Sample size planning for classification models. Anal Chim Acta 760:25–33
Krafft C, Schie IW, Meyer T, Schmitt M, Popp J (2016) Developments in spontaneous and coherent Raman scattering microscopic imaging for biomedical applications. Chem Soc Rev 45(7):1819–1849
Pence I, Mahadevan-Jansen A (2016) Clinical instrumentation and applications of Raman spectroscopy. Chem Soc Rev 45(7):1958–1979
Thorley FC, Baldwin KJ, Lee DC, Batchelder DN (2006) Dependence of the Raman spectra of drug substances upon laser excitation wavelength. Journal of Raman Spectroscopy 37(1–3):335–341
Meesters AA, Pitassi LHU, Campos V, Wolkerstorfer A, Dierickx CC (2014) Transcutaneous laser treatment of leg veins. Lasers Med Sci 29(2):481–492
Kumamoto Y, Fujita K, Smith NI, Kawata S (2016) Deep-UV biological imaging by lanthanide ion molecular protection. Biomed Opt Express 7(1):158–170
Efremov EV, Ariese F, Gooijer C (2008) Achievements in resonance Raman spectroscopy review of a technique with a distinct analytical chemistry potential. Anal Chim Acta 606(2):119–134
Spiro TG, Strekas TC (1974) Resonance Raman spectra of heme proteins. Effects of oxidation and spin state. J Am Chem Soc 96(2):338–345
Puppels GJ, de Mul FFM, Otto C, Greve J, Robert-Nicoud M, Arndt-Jovin DJ, Jovin TB (1990) Studying single living cells and chromosomes by confocal Raman microspectroscopy. Nature 347(6290):301–303
De Grauw CJ, Otto C, Greve J (1997) Line-scan Raman microspectrometry for biological applications. Appl Spectrosc 51(11):1607–1612
Hamada K, Fujita K, Smith NI, Kobayashi M, Inouye Y, Kawata S (2008) Raman microscopy for dynamic molecular imaging of living cells. J Biomed Opt 13(4):044027
Pascut FC, Goh HT, Welch N, Buttery LD, Denning C, Notingher I (2011) Non-invasive detection and imaging of molecular markers in live cardiomyocytes derived from human embryonic stem cells. Biophys J 100(1):251–259
Pavillon N, Smith NI (2016) Compressed sensing laser scanning microscopy. Opt Express 24(26):30038–30052
Pavillon N, Smith NI (2015) Maximizing throughput in label-free microspectroscopy with hybrid Raman imaging. J Biomed Opt 20(1):016007
Rowlands CJ, Varma S, Perkins W, Leach I, Williams H, Notingher I (2012) Rapid acquisition of Raman spectral maps through minimal sampling: applications in tissue imaging. J Biophotonics 5(3):200–229
Zumbusch A, Holtom GR, Xie XS (1999) Three-dimensional vibrational imaging by coherent anti-stokes Raman scattering. Physics Review Letters 82:4142–4145
Becker K, Kiefer J (2016) Combined spontaneous stokes and coherent anti-stokes Raman scattering spectroscopy. Applied Physics B 122:127
Kano H, Segawa H, Okuno M, Leproux P, Couderc V (2016) Hyperspectral coherent Raman imaging – principle, theory, instrumentation and applications to life sciences. J Raman Spectrosc 47(1):116–123
Camp CH Jr, Cicerone MT (2015) Chemically sensitive bioimaging with coherent Raman scattering. Nat Photonics 9:295–305
Yonemaru Y, Palonpon AF, Kawano S, Smith NI, Kawata S, Fujita K (2015) Super-spatial- and -spectral-resolution in vibrational imaging via saturated coherent anti-stokes Raman scattering. Phys Rev Appl 4:014010
Prince RC, Frontiera RR, Potma EO (2017) Stimulated Raman scattering: from bulk to nano. Chem Rev 117(7):5070–5094
Ozeki Y, Umemura W, Otsuka Y, Satoh S, Hashimoto H, Sumimura K, Nishizawa N, Fukui K, Itoh K (2012) High-speed molecular spectral imaging of tissue with stimulated Raman scattering. Nat Photonics 6:845–851
Byrne HJ, Knief P, Keating ME, Bonnier F (2016) Spectral pre and post processing for infrared and Raman spectroscopy of biological tissues and cells. Chem Soc Rev 45(7):1865–1878
Kumar S, Verma T, Mukherjee R, Ariese F, Somasundaram K (2016) Umapathy Raman and infrared microscopy: towards quantitative evaluation for clinical research by ratiometric analysis. Chem Soc Rev 45(7):1879–1900
Slater JB, Tedesco JM, Fairchild RC, Lewis IR (2001) Raman spectrometry and its adaptation to the industrial environment. In: Lewis IR, Edwards HGM (eds) Handbook of Raman spectroscopy from the research laboratory to the process line. CRC Press, Boca Raton
Bocklitz TW, Dörfer T, Heinke R, Schmitt M, Popp J (2015) Spectrometer calibration protocol for Raman spectra recorded with different excitation wavelengths. Spectrochim Acta A Mol Biomol Spectrosc 149:544–549
Heuke S, Vogler N, Meyer T, Akimov D, Kluschke F, Röwert-Huber H-J, Lademann J, Dietzekand B, Popp J (2013) Multimodal mapping of human skin. Br J Dermatol 169(4):794–803
Mavarani L, Petersen D, El-Mashtoly SF, Mosig A, Tannapfel A, Kötting C, Gerwert K (2013) Spectral histopathology of colon cancer tissue sections by Raman imaging with 532 nm excitation provides label-free annotation of lymphocytes, erythrocytes and proliferating nuclei of cancer cells. Analyst 138(14):4035–4039
Ali SM, Bonnier F, Ptasinski K, Lambkin H, Flynn K, Lyng FM, Byrne HJ (2013) Raman spectroscopic mapping for the analysis of solar radiation induced skin damage. Analyst 138(14):3946–3956
Franzen L, Mathes C, Hansen S, Windbergs M (2013) Advanced chemical imaging and comparisons of human and porcine hair follicles for drug delivery by confocal Raman microscopy. J Biomed Opt 18(6):061210
Vukosavljevic B, De Kinder L, Siepmann J, Muschert S, Windbergs M (2016) Novel insights into controlled drug release from coated pellets by confocal Raman microscopy. J Raman Spectrosc 47(7):757–762
Ashton L, Hollywood KA, Goodacre R (2015) Making colourful sense of Raman images of single cells. Analyst 140(6):1852–1858
Shinzawa H, Awa K, Kanematsu W, Ozaki Y (2009) Multivariate data analysis for Raman spectroscopic imaging. J Raman Spectrosc 40(12):1720–1725
Bonifacio A, Beleites C, Vittur F, Marsich E, Semeraro S, Paoletti S, Sergo V (2010) Chemical mapping of articular cartilage sections with Raman mapping employing uni- and multi-variate methods for data analysis. Analyst 135(12):3193–3204
Matthäus C, Chernenko T, Quintero L, Miljković M, Milane L, Kale M, Amiji M, Torchilin V, Diem M (2010) Raman micro-spectral imaging of cells and intracellular drug delivery using nanocarrier systems. In: Dieing T, Hollrichter O, Toporski J (eds) Confocal Raman microscopy. Springer, Berlin
Miljković M, Chernenko T, Romeo MJ, Bird B, Matthäus C, Diem M (2010) Label-free imaging of human cells: algorithms for image reconstruction of Raman hyperspectral datasets. Analyst 135(8):2002–2013
Hedegaard M, Matthäus C, Hassing S, Krafft C, Diem M, Popp J (2011) Spectral unmixing and clustering algorithms for assessment of single cells by Raman microscopic imaging. Theor Chem Accounts 130(4):1249–1260
Vajna B, Patyi G, Nagy Z, Bódis A, Farkas A, Marosi G (2011) Comparison of chemometric methods in the analysis of pharmaceuticals with hyperspectral Raman imaging. J Raman Spectrosc 42(11):1977–1986
Hotelling H (1933) Analysis of a complex of statistical variables into principal components. Warwick & York, Baltimore
Nascimento JMP, Bioucas Dias JM (2005) Vertex component analysis: a fast algorithm to unmix hyperspectral data. IEEE Trans Geosci Remote Sens 43(4):898–910
De Juan A, Jaumot J, Tauler R (2014) Multivariate curve resolution (MCR). Solving the mixture analysis problem. Anal Methods 6(14):4964–4976
MacQueen J (1967) Some methods for classification and analysis of multivariate observations. In: Proceedings of the fifth berkeley symposium on mathematical statistics and probability. Statistics, vol 1. University of California Press, Berkeley, pp 281–297
Klein K, Gigler AM, Aschenbrenner T, Monetti R, Bunk W, Jamitzky F, Morfill G, Stark RW, Schlegel J (2012) Label-free live cell imaging with confocal microscopy. Biophys J 102(2):360–368
Ramoji A, Neugebauer U, Bocklitz T, Foerster M, Kiehntopf M, Bauer M, Popp J (2012) Towards a spectroscopic hemogram: Raman spectroscopic differentiation of the two most abundant leukocytes from peripheral blood. Anal Chem 84(12):533–5342
Kakita M, Kaliaperumal V, Hamaguchi H (2012) Resonance Raman quantification of the redox state of cytochromes B and C in vivo and in vitro. J Biophotonics 5(1):20–24
Krauß SD, Petersen D, Niedieker D, Fricke I, Freier E, El-Mashtoly SF, Gerwert K, Mosig A (2015) Colocalization of fluorescence and Raman microscopic images for the identification of subcellular compartments: a validation study. Analyst 140(7):2360–2368
Brazhe NA, Treiman M, Brazhe AR, Find NL, Maksimov GV, Sosnovtseva OV (2012) Mapping of redox state of mitochondrial cytochromes in live cardiomyocytes using Raman microspectroscopy. PLoS One 7(9):e41190
Kochan K, Kus E, Filipek A, Szafrańska K, Chlopicki S, Baranska M (2017) Label-free spectroscopic characterisation of live liver sinusoidal endothelial cells (LSECs) isolated from the murine liver. Analyst 142(8):1308–1319
Neugebauer U, Clement JH, Bocklitz T, Krafft C, Popp J (2010) Identification and differentiation of single cells from peripheral blood by Raman spectroscopic imaging. J Biophotonics 3(8–9):579–587
Hobro AJ, Kumagai Y, Akira S, Smith NI (2016) Raman spectroscopy as a tool for label-free lymphocyte cell line discrimination. Analyst 141(12):3756–3764
Zuser E, Chernenko T, Newmark J, Miljković M, Diem M (2010) Confocal Raman microspectral imaging (CRMI) of murine stem cell colonies. Analyst 135:3030–3033
Ashton L, Lau K, Winder CL, Goodacre R (2011) Raman spectroscopy: lighting up the future of microbial identification. Future Microbiol 6(9):991–997
Mazur AI, Monahan JL, Miljkovic M, Laver N, Diem M, Bird B (2013) Vibrational spectroscopic changes of B-lymphocytes upon activation. J Biophotonics 6(1):101–109
Ghita A, Pascut FC, Mather M, Sottile V, Notingher I (2012) Cytoplasmic RNA in undifferentiated neural stem cells: a potential label-free Raman spectral marker for assessing the undifferentiated status. Anal Chem 84(7):3155–3162
Meister K, Schmidt DA, Bründermann E, Havenith M (2010) Confocal Raman microspectroscopy as an analytical tool to assess the mitochondrial status in human spermatozoa. Analyst 135(6):1370–1374
Mateu BP, Harreither E, Schosserer M, Puxbaum V, Gludovacz E, Borth N, Gierlinger N, Grillari J (2016) Label-free live cell imaging by confocal Raman spectroscopy identifies CHO host and producer cell lines. Biotechnol J 12(1):1600037
Zoladek AB, Johal RK, Garcia-Nieto S, Pascut F, Shakesheff KM, Ghaemmaghami AM, Notingher I (2010) Label free molecular imaging of the immunological synapse between dendritic and T-cells by Raman micro-spectroscopy. Analyst 135(12):3205–3212
Konorov SO, Schulze HG, Piret JM, Blades MW, Turner RFB (2013) Label-free determination of the cell cycle phase in human embryonic stem cells by Raman microspectroscopy. Anal Chem 85(19):8996–9002
Schulze HG, Konorov SO, Piret JM, Blades MW, Turner RFB (2013) Label-free imaging of mammalian cell nucleoli by Raman microspectroscopy. Analyst 138(12):3416–3423
Hsu J-F, Hsieh P-Y, Hsu H-Y, Shigeto S (2015) When cells divide: label-free multimodal spectral imaging for exploratory molecular investigation of living cells during cytokinesis. Sci Rep 5:17541
Yang Y, Li F, Gao L, Wang Z, Thrall MJ, Shen SS, Wong KK, Wong STC (2011) Differential diagnosis of breast cancer using quantitative, label-free and molecular vibrational imaging. Biomed Opt Express 2(8):2160
Larraona-Puy M, Ghita A, Zoladek A, Perkins W, Varma S, Leach IH, Koloydenko AA, Williams H, Notingher I (2011) Discrimination between basal cell carcinoma and hair follicles in skin tissue sections by Raman microspectroscopy. J Mol Struct 993(1–3):57–61
Brozek-Pluska B, Kopec M, Niedzwiecka I, Morawiec-Sztandera A (2015) Label free determination of lipid composition and secondary protein structure of human salivary noncancerous and cancerous tissues by Raman microspectroscopy. Analyst 140(7):2107–2113
Brozek-Pluska B, Kopec M, Abramczyk H (2016) Development of a new diagnostic Raman method for monitoring epigenetic modifications in the cancer cells of human breast tissue. Anal Methods 8(48):8542–8553
Vanna R, Ronchi P, Lenferink ATM, Tresoldi C, Morasso C, Mehn D, Bedoni M, Picciolini S, Terstappen LWMM, Ciceri F, Otto C, Gramatica F (2015) Label-free imaging and identification of typical cells of acute myeloid leukaemia and myelodysplastic syndrome by Raman microspectroscopy. Analyst 140(4):1054–1064
Surmacki J, Brozek-Pluska B, Kordek R, Abramczyk H (2015) The lipid-reactive oxygen species phenotype of breast cancer. Raman spectroscopy and mapping, PCA and PLSDA for invasive ductal carcinoma and invasive lobular carcinoma. Molecular tumorigenic mechanisms beyond the Warburg effect. Analyst 140(7):2121–2133
Brozek-Pluska B, Kopec M, Surmacki J, Abramczyk H (2015) Raman microspectroscopy of noncancerous and cancerous breast tissues: identification and phase transitions of linoleic and oleic acids by Raman low temperature studies. Analyst 140(7):2134–2143
Piredda P, Berning M, Boukamp P, Volkmer A (2015) Subcellular Raman microspectroscopic imaging of nucleic acids and tryptophan for distinction of normal human skin cells and tumorigenic keratinocytes. Anal Chem 87(13):6778–6785
Kumar P, Bhattacharjee T, Pandey M, Hole A, Ingle A, Krishna CM (2016) Raman spectroscopy in experimental oral carcinogenesis: investigation of abnormal changes in control tissues. J Raman Spectrosc 47(11):1318–1326
Abramczyk H, Surmacki J, Kopeć M, Olejnik AK, Lubecka-Pietruszewska K, Fabianowska-Majewska K (2015) The role of lipid droplets and adipocytes in cancer. Raman imaging of cell cultures: MCF10A, MCF7 and MDA-MB-231 compared to adipocytes in cancerous breast tissue. Analyst 140(7):2224–2235
Tirinato L, Liberale C, Di Franco S, Candeloro P, Benfante A, La Rocca R, Potze L, Marotta R, Ruffilli R, Rajamanickam VP, Malerba M, De Angelis F, Falqui A, Carbone E, Todaro M, Medema JP, Stassi G, Di Fabrizio E (2015) Lipid droplets: a new player in colorectal cancer stem cells unveiled by spectroscopic imaging. Stem Cells 33(1):35–44
Hartsuiker L, Zeijen NJL, Terstappen LWMM, Otto C (2010) A comparison of breast cancer tumor cells with varying expression of the Her2/neu receptor by Raman spectroscopic imaging. Analyst 135(12):3220–3226
Pacia MZ, Buczek E, Blazejczyk A, Gregorius A, Wietrzyk J, Chlopicki S, Baranska M, Kaczorc A (2016) 3D Raman imaging of systemic endothelial dysfunction in the murine model of metastatic breast cancer. Anal Bioanal Chem 408:3381–3387
Kong K, Rowlands CJ, Varma S, Perkins W, Leach IH, Koloydenko AA, Pitiot A, Williams HC, Notingher I (2014) Increasing the speed of tumour diagnosis during surgery with selective scanning Raman microscopy. J Mol Struct 1073:56–65
Kong K, Zaabar F, Rakha E, Ellis I, Koloydenko A, Notingher I (2014) Towards intra-operative diagnosis of tumours during breast conserving surgery by selective-sampling Raman microspectroscopy. Phys Med Biol 59(20):6141–6152
Czamara K, Natorska J, Kapusta P, Baranska M, Kaczor A (2015) Raman microspectroscopy of human aortic valves. Investigation of the local and global biochemical changes associated with calcification in aortic stenosis. Analyst 140(7):2164–2170
Kochan K, Marzec KM, Chruszcz-Lipska K, Jasztal A, Maslak E, Musiolik H, Chłopicki S, Baranska M (2013) Pathological changes in the biochemical profile of the liver in atherosclerosis and diabetes assessed by Raman spectroscopy. Analyst 138(14):3885–3890
Marzec KM, Kochan K, Fedorowicz A, Jasztal A, Chruszcz-Lipska K, Dobrowolski JC, Chlopicki S, Baranska M (2015) Raman microimaging of murine lungs: insight into the vitamin A content. Analyst 140(7):2171–2177
Büntemeyer H, Lehmann J (2001) The role of vitamins in cell culture media in animal cell technology: from target to market. In: ESACT proceedings, vol 1. Springer Netherlands, pp 204–206
Kochan K, Chrabaszcz K, Szczur B, Maslak E, Dybas J, Marzec KM (2016) IR and Raman imaging of murine brains from control and ApoE/LDLR−/− mice with advanced atherosclerosis. Analyst 141(18):5329–5338
Pacia MZ, Mateuszuk L, Chlopicki S, Baranska M, Kaczor A (2015) Biochemical changes of the endothelium in the murine model of NO-deficient hypertension. Analyst 140(7):2178–2184
Rygula A, Pacia MZ, Mateuszuk L, Kaczor A, Kostogrys RB, Chlopickiad S, Baranska M (2015) Identification of a biochemical marker for endothelial dysfunction using Raman spectroscopy. Analyst 140(7):2185–2189
Michael R, Otto C, Lenferink A, Gelpi E, Montenegro GA, Rosandić J, Tresserra F, Barraquer RI, Vrensen GFJM (2014) Absence of amyloid beta in lenses of Alzheimer’s patients: a confocal Raman study. Exp Eye Res 119:44–53
Große C, Bergner N, Dellith J, Heller R, Bauer M, Mellmann A, Popp J, Neugebauer U (2015) Label-free imaging and spectroscopic analysis of intracellular bacterial infections. Anal Chem 87(4):2137–2142
Silge A, Abdou E, Schneider K, Meisel S, Bocklitz T, Lu-Walther H-W, Heintzmann R, Rösch P, Popp J (2015) Shedding light on host niches: label-free in situ detection of Mycobacterium gordonae via carotenoids in macrophages by Raman microspectroscopy. Cell Microbiol 17(6):832–842
Kong K, Rowlands CJ, Elsheikha H, Notingher I (2012) Label-free molecular analysis of live Neospora caninum tachyzoites in host cells by selective scanning microspectroscopy. Analyst 137(18):4119–4112
Naemat A, Elsheikha HM, Al-sandaqchi A, Kong K, Ghita A, Notingher I (2015) Analysis of interaction between the apicomplexan protozoan Toxoplasma gondii and host cells using label-free Raman spectroscopy. Analyst 140(3):756–764
Hobro AJ, Konishi A, Coban C, Smith NI (2013) Raman spectroscopic analysis of malaria disease progression via blood and plasma samples. Analyst 138(14):3927–3933
Hobro AJ, Pavillon N, Fujita K, Ozkan M, Coban C, Smith NI (2015) Label-free Raman imaging of the macrophage response to the malaria pigment hemozoin. Analyst 140(7):2350–2359
Kozicki M, Creek DJ, Sexton A, Morahan BJ, Wesełucha-Birczyńska A, Wood BR (2015) An attenuated total reflection (ATR) and Raman spectroscopic investigation into the effects of chloroquine on Plasmodium falciparum infected red blood cells. Analyst 140(7):2236–2246
Czepiel J, Kozicki M, Panasiuk P, Birczyńska M, Garlicki A, Wesełucha-Birczyńska A (2015) Clostridium difficile the hospital plague. Analyst 140(7):2513–2522
Kochan K, Marzec KM, Maslak E, Chlopicki S, Baranska M (2015) Raman spectroscopic studies of vitamin A content in the liver: a biomarker of a healthy liver. Analyst 140(7):2074–2079
El-Mashtoly SF, Yosef HK, Petersen D, Mavarani L, Maghnouj A, Hahn S, Kötting C, Gerwert K (2015) Label-free Raman spectroscopic imaging monitors the integral physiological relevant drug responses in cancer cells. Anal Chem 87(14):7297–7304
Majzner K, Wojcik T, Szafraniec E, Lukawska M, Oszczapowicz I, Chlopicki S, Baranska M (2015) Nuclear accumulation of anthracyclines in the endothelium studied by bimodal imaging: fluorescence and Raman microscopy. Analyst 140(7):2302–2310
Zhang H, Zheng J, Liu A, Xiao H, He L (2016) Label-free imaging and characterisation of cancer cell responses to polymethoxyflavones using Raman microscopy. J Agric Food Chem 64(51):9708–9713
Kim D-H, Jarvis RM, Allwood JW, Batman G, Moore RE, Marsden-Edwards E, Hampson L, Hampson IN, Goodacre R (2010) Raman chemical mapping reveals site of action of HIV protease inhibitors in HPV16 E6 expressing cervical carcinoma cells. Anal Bioanal Chem 398(7):3051–3061
Kang JW, Singh SP, Nguyen FT, Lue N, Sung Y, So PTC, Dasari RR (2016) Investigating effects of proteasome inhibitor on multiple myeloma cells using confocal Raman microscopy. Sensors 16(12):2133
Brozek-Pluska B, Kopec M (2016) Raman microspectroscopy of hematoporphorins, imaging of the non-cancerous and the cancerous human breast tissues with photosensitisers. Spectrochim Acta A Mol Biomol Spectrosc 169:182–191
Yosef HK, Mavarani L, Maghnouj A, Hahn S, El-Mashtoly SF, Gerwert K (2015) In vitro prediction of the efficacy of molecularly targeted cancer therapy by Raman spectral imaging. Anal Bioanal Chem 407(27):8321–8331
Meister K, Niesel J, Schatzschneider U, Metzler-Nolte N, Schmidt DA, Havenith M (2010) Label-free imaging of metal-carbonyl complexes in live cells by Raman microspectroscopy. Angew Chem Int Ed 49(19):3310–3312
Salehi H, Derely L, Vegh A-G, Durand J-C, Gergely C, Larroque C, Fauroux M-A, Cuisinier FJG (2013) Label free detection of anticancer drug Paclitaxel in living cells by confocal Raman microscopy. Appl Phys Lett 102(11):113701
El-Mashtoly SF, Petersen D, Yosef HK, Mosig A, Reinacher-Schick A, Kötting C, Gerwert K (2014) Label-free imaging of drug distribution and metabolism in colon cancer cells by Raman microscopy. Analyst 139(5):1155–1161
Goto N, Morita Y, Terada K (2016) Deposits from creams containing 20% (w/w) urea and suppression of crystallisation (part 3): novel analytical methods based on Raman spectroscopy for the characterisation of deposits and deposition phenomena of creams containing 20% (w/w) urea. Chem Pharm Bull 64(8):1099–1107
Chernenko T, Matthäus C, Milane L, Quintero L, Amiji M, Diem M (2009) Label-free raman spectral imaging of intracellular delivery and degradation of polymeric nanoparticle systems. ACS Nano 3(11):3552–3559
Raaschou-Nielsen O, Andersen ZJ, Beelen R, Samoli E, Stafoggia M, Weinmayr G, Hoffmann B, Fischer P, Nieuwenhuijsen MJ, Brunekreef B, Xun WW, Katsouyanni K, Dimakopoulou K, Sommar J, Forsberg B, Modig L, Oudin A, Oftedal B, Schwarze PE, Nafstad P, De Faire U, Pedersen NL, Östenson C-G, Fratiglioni L, Penell J, Korek M, Pershagen G, Eriksen KT, Sørensen M, Tjønneland A, Ellermann T, Eeftens M, Peeters PH, Meliefste K, Wang M, Bueno-de-Mesquita B, Key TJ, de Hoogh K, Concin H, Nagel G, Vilier A, Grioni S, Krogh V, Tsai M-Y, Ricceri F, Sacerdote C, Galassi C, Migliore E, Ranzi A, Cesaroni G, Badaloni C, Forastiere F, Tamayo I, Amiano P, Dorronsoro M, Trichopoulou A, Bamia C, Vineis P, Hoek G (2013) Air pollution and lung cancer incidence in 17 European cohorts: prospective analyses from the European Study of Cohorts for Air Pollution Effects (ESCAPE). Lancet Oncol 14(9):813–822
Dorney J, Bonnier F, Garcia A, Casey A, Chambers G, Byrne HJ (2012) Identifying and localising intracellular nanoparticles using Raman spectroscopy. Analyst 137(5):1111–1119
Bräutigam K, Bocklitz T, Silgea A, Dierker C, Ossig R, Schnekenburger J, Cialla D, Rösch P, Popp J (2014) Comparative two- and three-dimensional analysis of nanoparticle localisation in different cell types by Raman spectroscopic imaging. J Mol Struct 1073:44–50
Ahlinder L, Ekstrand-Hammarström B, Geladi P, Osterlund L (2015) Large uptake of titania and iron oxide nanoparticles in the nucleus of lung epithelial cells as measured by Raman imaging and multivariate classification. Biophys J 105(2):310–319
Cherchi C, Chernenko T, Diem M (2011) Impact of nano titanium dioxide exposure on cellular structure of anabaena variabilis and evidence of internalisation. Environ Toxicol Chem 30(4):861–869
Iannarelli L, Giovannozzi AM, Morelli F, Viscotti F, Bigini P, Maurino V, Spoto G, Martra G, Ortel E, Hodoroaba V-D, Rossi AM, Diomede L (2016) Shape engineered TiO2 nanoparticles in C. elegans. A Raman imaging based approach to assist tissue-specific toxicological studies. RSC Adv 6(74):70501–70509
Alebrahim MA, Krafft C, Popp J (2015) Raman imaging to study structural and chemical features of the dentine enamel junction. IOP Conf Ser: Mater Sci Eng 92:012014
Fraser SJ, Natarajan AK, Clark ASS, Drummond BK, Gordon KC (2015) A Raman spectroscopic study of teeth affected with molar – incisor hypomineralisation. J Raman Spectrosc 46(2):202–210
Masic A, Weaver JC (2015) Large area submicron chemical imaging of magnesium in sea urchin teeth. J Struct Biol 189(3):269–275
Shabestari M, Eriksen EF, Paschalis EP, Roschger P, Gamsjaeger S, Klaushofer K, Berzlanovich A, Nogues X, Puig L, Diez-Perez A (2017) Presence of pyrophosphate in bone from an atypical femoral fracture site: a case report. Bone Reports 6:81–86
Austin C, Smith TM, Farahani RM, Hinde K, Carter EA, Lee J, Lay PA, Kennedy BJ, Sarrafpour B, Wright RJ, Wright RO, Arora M (2016) Uncovering system-specific stress signatures in primate teeth with multimodal imaging. Sci Rep 6:18802
Toledano M, Aguilera FS, Osorio E, Cabello I, Osorio R (2014) Microanalysis of thermal-induced changes at the resin-dentin interface. Microsc Microanal 20(4):1218–1233
Toledano M, Cabello I, Aguilera FS, Osorio E, Osorio R (2015) Effect of in vitro chewing and bruxism events on remineralisation at the resin-dentin interface. J Biomech 48(1):14–21
Hashimoto A, Chiu L-d, Sawada K, Ikeuchi T, Fujita K, Takedachi M, Yamaguchi Y, Kawata S, Murakami S, Tamiya E (2014) In situ Raman imaging of osteoblastic mineralisation. J Raman Spectrosc 45(2):157–161
Ghita A, Pascut FC, Sottile V, Notingher I (2014) Monitoring the mineralization of bone nodules in vitro by space- and time-resolved Raman microscopy. Analyst 139(1):55–58
Gao Y, Xu C, Wang L (2016) Non-invasive monitoring of the osteogenic differentiation of human mesenchymal stem cells on a polycaprolactone scaffold using Raman imaging. RSC Adv 6(66):61771
Autefage H, Gentleman E, Littmann E, Hedegaard MA, Von Erlach T, O’Donnell M, Burden FR, Winkler DA, Stevens MM (2015) Sparse feature selection methods identify unexpected glabal cellular response to strontium containing materials. Proc Natl Acad Sci U S A 112(14):4280–4285
Wei X, Wang X, Fang Y, Huang Q (2013) Comparison of hair from rectum cancer patients and from healthy persons by Raman microspectroscopy and imaging. J Mol Struct 1048:83–87
Wu Y, Chen G, Ji C, Hoptroff M, Jones A, Collins LZ, Janssen H-G (2016) Gas chromatography – mass spectrometry and Raman imaging measurement of squalene content and distribution in human hair. Anal Bioanal Chem 408(9):2357–2362
Perera PN, Schmidt M, Chuck PJ, Adams PD (2011) Blind image analysis for the compositional and structural characterisation of plant cell walls. Anal Chim Acta 702(2):172–177
Hänninen T, Kontturi E, Vuonnen T (2011) Distribution of lignin and its coniferyl alcohol and coniferyl aldehyde groups in Picea abies and Pinus sylvestris as observed by Raman imaging. Phytochemistry 72(14–15):1889–1895
Ma J, Lv X, Yang S, Tian G, Liu X (2015) Structural insight into cell wall architecture of Miscanthus sinensis using correlative microscopy approaches. Microsc Microanal 21(5):1304–1313
Sun L, Singh S, Joo M, Vega-Sanchez M, Ronald P, Simmons BA, Adams P, Auer M (2016) Non-invasive imaging of cellulose microfibril orientation within plant cell walls by polarised Raman microspectroscopy. Biotechnol Bioeng 113(1):82–90
Chu L-Q, Masyuko R, Sweedler JV, Bohn PW (2010) Base-induced delignification of miscanthus × giganteus studied by three-dimensional confocal Raman imaging. Bioresour Technol 101(13):4919–4925
Vermaak I, Viljoen AM, Hamman JH, Baranska M (2010) The potential application of FT-Raman spectroscopy for the quantification and mapping of the steroidal glycoside P57 in Hoodia gordonii. Phytochem Lett 3(3):156–160
Zimmermann B, Bağcıoğlu M, Sandt C, Kohler A (2015) Vibrational microspectroscopy enables chemical characterisation of single pollen grains as well as comparative analysis of plant species based on pollen ultrastructure. Planta 242(5):1237–1250
Chylińska M, Szymańska-Chargot M, Zdunek A (2014) Imaging of polysaccharides in the tomato cell wall with Raman microspectroscopy. Plant Methods 10:14
Philippe G, Gaillard C, Petit J, Geneix N, Dalgalarrondo M, Bres C, Mauxion J-P, Franke R, Rothan C, Schreiber L, Marion D, Bakan B (2016) Ester cross-link profiling of the cutin polymer of wild-type and cutin synthase tomato mutants highlights different mechanisms of polymerisation. Plant Physiol 170(2):807–820
Szymańska-Chargot M, Chylińska M, Pieczywek PM, Rösch P, Schmitt M, Popp J, Zdunek A (2016) Raman imaging of changes in polysaccharide distribution in the cell wall during apple fruit development and senescence. Planta 243:935–945
Richter S, Müssig J, Gierlinger N (2011) Functional plant cell wall design revealed by the Raman imaging approach. Planta 233(4):763–772
Mateu BP, Hauser MT, Heredia A, Gierlinger N (2016) Waterproofing in Arabidopsis: following phenolics and lipids in situ by confocal Raman microscopy. Front Chem 4:10
Polisetti S, Bible AN, Morrell-Falvey JL, Bohn PW (2016) Raman chemical imaging of the rhizosphere bacterium Pantoea sp. YR343 and its co-culture with Arabidopsis thaliana. Analyst 141(7):2175
Kammer M, Hedrich R, Ehrlich H, Popp J, Brunner E, Krafft C (2010) Spatially resolved determination of the structure and composition of diatom cell walls by Raman and FTIR imaging. Anal Bioanal Chem 398(1):509–517
Chiu L-d, Ho S-H, Shimada R, Ren N-Q, Ozawa T (2017) Rapid in vitro lipid/carbohydrate quantification of single microalgal cell by Raman spectral imaging to reveal salinity-induced starch-to-lipid shift. Biotechnol Biofuels 10:9
Noothalapati H, Sasaki T, Kaino T, Kawamukai M, Ando M, Hamaguchi H, Yamamoto T (2016) Label-free chemical imaging of fungal spore walls by Raman microscopy and multivariate curve resolution analysis. Sci Rep 6:27789
Yoo HY, Iordachescu M, Huang J, Hennebert E, Kim S, Rho S, Foo M, Flammang P, Zeng H, Hwang D, Waite JH, Hwang DS (2016) Sugary interfaces mitigate contact damage where stiff meets soft. Nat Commun 7:11923
Nakamura MJ, Hotta K, Oka K (2013) Raman spectroscopic imaging of the whole Ciona intestinalis embryo during development. PLoS One 8:e71739
Hölscher D, Dhakshinamoorthy S, Alexandrov T, Becker M, Bretschneider T, Buerkert A, Crecelius AC, De Waele D, Elsen A, Heckel DG, Heklau H, Hertweck C, Kai M, Knop K, Krafft C, Maddula RK, Matthäus C, Popp J, Schneider B, Schubert US, Sikora RA, Svatoš A, Swennen RL (2014) Phenalenon-type phytoalexins mediate resistance of banana plants (musa spp.) to the burrowing nematode Radopholus similis. Proc Natl Acad Sci U S A 111(1):105–110
Smith GPS, Holroyd SE, Reid DCW, Gordon KC (2017) Raman imaging processed cheese and its components. J Raman Spectrosc 48(3):374–383
Kilcrease J, Collins AM, Richins RD, Timlin JA, O’Connell MA (2013) Multiple microscopic approaches demonstrate linkage between chromoplast architecture and carotenoid composition in diverse Capsicum annum fruit. Plant J 76(6):1074–1083
Qin J, Chao K, Kim MS (2013) Simultaneous detection of multiple adulterants in dry milk using macroscale Raman chemical imaging. Food Chem 138(2–3):998–1007
Dhakal S, Chao K, Qin J, Kim M, Chan D (2016) Raman spectral imaging for quantitative contaminant evaluation in skim milk powder. Food Meas 10(2):374–386
Clemente I, Aznar M, Nerín C (2016) Raman imaging spectroscopy as a tool to investigate the cells damage on Aspergillus ochraceus caused by an antimicrobial packaging containing benzylisothiocyanate. Anal Chem 88(9):4772–4779
Eksi-Kocak H, Mentes-Yilmaz O, Boyaci IH (2016) Detection of green pea adulteration in pistachio nut granules by using Raman hyperspectral imaging. Eur Food Res Technol 242(2):271–277
Kwok K, Taylor LS (2012) Analysis of counterfeit Cialis tablets using Raman microscopy and multivariate curve resolution. J Pharm Biomed Anal 66:126–135
Sacré P-Y, Deconinck E, Saerens L, De Beer T, Courselle P, Vancauwenberghe R, Chiap P, Crommen J, De Beer JO (2011) Detection of counterfeit Viagra by Raman microspectroscopy imaging and multivariate analysis. J Pharm Biomed Anal 56(2):454–461
Smus JP, Moura CC, McMorrow E, Tare RS, Oreffo ROC, Mahajan S (2015) Tracking adipogenic differentiation of skeletal stem cells by label-free chemically selective imaging. Chem Sci 6(12):7089–7096
Bradley J, Pope I, Masia F, Sanusi R, Langbein W, Swann K, Paola B (2016) Quantitative imaging of lipids in live mouse oocytes and early embryos using CARS microscopy. Development 143:2238–2247
Chen W-W, Yi Y-H, Chien C-H, Hsiung K-C, Ma T-H, Lin Y-C, Lo SJ, Chang T-C (2016) Specific polyunsaturated fatty acids modulate lipid delivery and oocyte development in C. elegans revealed by molecular-selective label free imaging. Sci Rep 6:32021
Jüngst C, Klein M, Zumbusch A (2013) Long term live cell microscopy studies of lipid droplet fusion dynamics in adipocytes. J Lipid Res 54(12):3419–3429
Di Napoli C, Pope I, Masia F, Langbein W, Watson P, Borri P (2016) Quantitative spatiotemporal chemical profiling of individual lipid droplets by hyperspectral CARS microscopy in living human adipose-derived stem cells. Anal Chem 88(7):3677–3685
Lee JH, Kim DH, Song WK, Oh M-K, Ko D-K (2015) Label-free imaging and quantitative chemical analysis of Alzheimer’s disease brain samples with multimodal multiphoton non-linear optical microscopy. J Biomed Opt 20(5):056013
Cavonius L, Fink H, Kiskis J, Albers E, Undeland I, Enejder A (2015) Imaging of lipids in microalgae with coherent anti-stokes Raman microscopy. Plant Physiol 167(3):603–616
Belanger E, Crépeau J, Laffray S, Vallée R, De Koninck Y, Côté D (2012) Live animal myelin histomorphometry of the spinal cord with video-rate multimodal nonlinear microendoscopy. J Biomed Opt 17(2):021107
Turcotte R, Rutledge DJ, Bélanger E, Dill D, Macklin WB, Côté D (2016) Intravital assessment of myelin molecular order with polarimetric multiphoton microscopy. Sci Rep 6:31685
Chen X, Gasecka P, Formanek F, Galey J-B, Rigneault H (2016) In vivo single human sweat gland activity monitoring using coherent anti-stokes Raman scattering and two-photon excited autofluorescence microscopy. Br J Dermatol 174(4):803–812
Wang H, Osseiran S, Igras V, Nichols AJ, Roider EM, Pruessner J, Tsao H, Fisher DE, Evans CL (2016) Invivo coherent Raman imaging of the melanomagenesis-associated pigment pheomelanin. Sci Rep 6:37986
Johnston HJ, Mouras R, Brown DM, Elfick A, Stone V (2015) Exploring the cellular and tissue uptake of nanomaterials in a range of biological samples using multimodal non-linear optical microscopy. Nanotechnology 26(50):505102
Watts AJR, Lewis C, Goodhead RM, Beckett SJ, Moger J, Tyler CR, Galloway TS (2014) Uptake and retention of microplastics by the shore crab Carcinus maenas. Environ Sci Technol 48(15):8823–8830
Uckermann O, Galli R, Tamosaityte S, Leipnitz E, Geiger KD, Schackert G, Koch E, Steiner G, Kirsch M (2014) Label-free delineation of brain tumours by coherent anti-stokes Raman scattering microscopy in an orthotopic mouse model and human glioblastoma. PLoS One 9(9):e107115
Potcoava MC, Futia GL, Aughenbaugh J, Schlaepfer IR, Gibson EA (2014) Raman and coherent anti-stokes Raman scattering microscopy studies of changes in lipid content and composition in hormone treated breast and prostate cancer cells. J Biomed Opt 19(11):111065
Parekh SH, Lee YJ, Aamer KA, Cicerone MT (2010) Label-free cellular imaging by broadband coherent anti-stokes Raman scattering microscopy. Biophys J 99(8):2695–2704
Bito K, Okuno M, Kano H, Tokuhara S, Naito S, Masukawa Y, Leproux P, Couderc V, Hamaguchi H (2012) Protein secondary structure imaging with ultrabroadband multiplex coherent anti-stokes Raman scattering microspectroscopy. J Phys Chem B 116(4):1425–1457
Lin C-Y, Suhalim JL, Nien CL, Miljković MD, Diem M, Jester JV, Potma EO (2011) Picosecond spectral coherent anti-stokes Raman scattering with principal component analysis of Meibomian glands. J Biomed Opt 16(2):021104
El-Mashtoly SF, Niedieker D, Petersen D, Krauss SD, Freier E, Maghnouj A, Mosig A, Hahn S, Kötting C, Gerwert K (2014) Automated identification of subcellular organelles by coherent anti-stokes Raman scattering. Biophys J 106(9):1910–1920
Roeffaers MBJ, Zhang X, Freudiger CW, Saar BG, van Ruijven M, van Dalen G, Xiao C, Xie XS (2011) Label-free imaging of biomolecules in food products using stimulated Raman microscopy. J Biomed Opt 16(2):021118
Littlejohn GR, Mansfield JC, Parker D, Lind R, Perfect S, Seymour M, Smirnoff N, Love J, Moger J (2015) In vivo chemical and structural analysis of plant cuticle waxes using SRS microscopy. Plant Physiol 168(1):18–28
Zeng Y, Zhao S, Wei H, Tucker MP, Himmel ME, Mosier NS, Meilan R, Ding S-Y (2015) In situ micro-spectroscopic investigation of lignin in poplar cell walls pretreated by maleic acid. Biotechnol Biofuels 8:126
Mansfield JC, Littlejohn GR, Seymour MP, Lind RJ, Perfect S, Moger J (2013) Label-free chemically specific imaging in planta with stimulated Raman scattering microscopy. Anal Chem 85(10):5055–5063
Egawa M, Tokunaga K, Hosoi J, Iwanaga S, Ozeki Y (2016) In situ visualisation of intracellular morphology of epidermal cells using stimulated Raman scattering microscopy. J Biomed Opt 21(8):086017
Tian F, Yang W, Mordes DA, Wang J-Y, Salameh JS, Mok J, Chew J, Sharma A, Leno-Duran E, Suzuki-Uematsu S, Suzuki N, Han SS, Lu F-K, Ji M, Zhang R, Liu Y, Strominger J, Shneider NA, Petrucelli L, Xie XS, Eggan K (2016) Monitoring peripheral nerve degeneration in ALS by label-free stimulated Raman scattering imaging. Nat Commun 7:13283
Conley SJ, Wicha MX (2013) Breast cancer stem cells: from theory to therapy. In: Sell S (ed) Stem cells handbook. Humana Press, Totowa
Li J, Condello S, Thomes-Pepin J, Ma X, Xia Y, Hurley TD, Matei D, Cheng J-X (2017) Lipid desaturation is a metabolic marker and therapeutic target of ovarian cancer stem cells. Cell Stem Cell 20(3):1–12
Lu F-K, Basu S, Igras V, Hoang MP, Ji M, Fu D, Holtom GR, Neel VA, Freudiger CW, Fisher DE, Xie XS (2015) Label-free DNA imaging in vivo with stimulated Raman scattering microscopy. Proc Natl Acad Sci U S A 112(37):11624–11629
Satoh S, Otsuka Y, Ozeki Y, Itoh K, Hashiguchi A, Yamazaki K, Hashimoto H, Sakamoto M (2014) Label-free visualisation of acetaminophen-induced liver injury by high-speed stimulated Raman scattering spectral microscopy and multivariate image analysis. Pathol Int 64(10):518–526
Datta R, Heylman C, George SC, Gratton E (2016) Label-free imaging of metabolism and oxidative stress in human induced pluripotent stem-cell derived cardiomyocytes. Biomed Opt Express 7(5):1690–1701
Pavillon N, Smith NI (2015) Implementation of simultaneous quantitative phase with Raman imaging. EPJ Tech Instrum 2(5):1–11
Petibois C (2010) Imaging methods for elemental, chemical, molecular and morphological analysis of single cells. Anal Bioanal Chem 397:2051–2065
Garip S, Bayari SH, Severcan M, Abbas S, Lednev IK, Severcan F (2016) Structural effects of Simvastatin on rat liver tissue. Fourier transform infrared and Raman microspectroscopic studies. J Biomed Opt 21(2):025008
Lau K, Hobro A, Smith T, Thurston T, Lendl B (2012) Label-free non-destructive in situ biochemical analysis of nematode Steinernema kraussei using FPA-FTIR and Raman spectroscopic imaging. Vib Spectrosc 60:34–42
Sroka-Bartnicka A, Kimber JA, Borkowski L, Pawlowska M, Polkowska I, Kalisz G, Belcarz A, Jozwiak K, Ginalska G, Kazarian SG (2015) The biocompatibility of carbon hydroxyapatite/β-glucan composite for bone tissue engineering studied with Raman and FTIR spectroscopic imaging. Anal Bioanal Chem 407(25):7775–7785
Chia CH, Gong B, Joseph SD, Marjo CE, Munroe P, Rich AM (2012) Imaging of mineral enriched biochar by FTIR, Raman and SEM-EDX. Vib Spectrosc 62:248–257
Caine S, Hackett MJ, Hou H, Kumar S, Maley J, Ivanishvili Z, Suen B, Szmigielski A, Jiang Z, Sylvain NJ, Nichol H (2016) A novel multimodal platform to image molecular and elemental alterations in ischemic stroke. Neurobiol Dis 91:132–142
Palombo F, Madami M, Fioretto D, Nallala J, Barr H, David A, Stone N (2016) Chemico-mechanical imaging of Barratt’s oesophagus. J Biophotonics 9(7):694–700
Spengler B (2015) Mass spectrometry imaging of biomolecular information. Anal Chem 87(1):64–82
Lanni EJ, Masyuko RN, Driscoll CM, Dunham SJB, Shrout JD, Bohn PW, Sweedler JV (2014) Correlated imaging with C60-SIMS and confocal Raman microscopy: visualisation of cell-scale molecular distributions in bacterial biofilms. Anal Chem 86(21):10885–10891
Ahlf DR, Masyuko RN, Hummon AB, Bohn PW (2014) Correlated mass spectrometry imaging and confocal Raman microscopy for studies of three dimensional cell culture sections. Analyst 139(18):4578–4585
Bocklitz TW, Bräutigam K, Urbanek A, Hoffmann F, von Eggeling F, Ernst G, Schmitt M, Schubert U, Guntinas-Lichius O, Popp J (2015) Novel workflow for combining Raman spectroscopy and MALDI-MSI for tissue based studies. Anal Bioanal Chem 407(26):7865–7873
Gavara N (2017) A beginner’s guide to atomic force microscopy probing for cell mechanics. Microsc Res Tech 80(1):75–84
Boifor R, Sinjab F, Strohbuecker S, Sottile V, Notingher I (2016) Towards quantitative molecular mapping of cells by Raman microscopy: using AFM for decoupling molecular concentration and cell topography. Faraday Discuss 187:199–212
Ashgari-Khiavi M, Wood BR, Mechler A, Bambery KR, Buckingham DW, Cooke BM, McNaughton D (2010) Correlation of atomic force microscopy and Raman microspectrscopy to study the effects of ex vivo treatment procedures on human red blood cells. Analyst 135(3):525–530
Marzec KM, Rygula A, Wood BR, Chlopicki S, Baranska High-resolution M (2015) Raman imaging reveals spatial location of heme oxidation sites in single red blood cells of dried smears. J Raman Spectrosc 46(1):76–83
Liao C, Piercey-Normore MD, Sorensen JL, Gough K (2010) In situ imaging of usnic acid in selected Cladonia spp by vibrational spectroscopy. Analyst 135(12):3242–3248
Tang M, McEwen GD, Wu Y, Miller CD, Zhou A (2013) Characterisation and analysis of mycobacteria and gram-negative bacteria and co-culture mixtures by Raman microspectroscopy, FTIR and atomic force microscopy. Anal Bioanal Chem 405(5):1577–1591
Pavillon N, Hobro AJ, Smith NI (2013) Cell optical density and molecular composition revealed by simultaneous multimodal label-free imaging. Biophys J 105(5):1123–1132
Kang JW, Lue N, Kong C-R, Barman I, Dingari NC, Goldfless SJ, Niles JC, Dasari RR, Feld MS (2011) Combined confocal Raman and quantitative phase microscopy system for biomedical diagnosis. Biomed Opt Express 2(9):2484–2492
Ferrara MA, Di Caprio G, Managò S, De Angelis A, Sirleto L, Coppola G, De Luca AC (2015) Label-free imaging and biochemical characterisation of bovine sperm cells. Biosensors 5(2):141–157
Ibrahim A, Hage CH, Souissi A, Leray A, Héliot L, Souissi S, Vandenbunder B (2015) Label-free microscopy and stress responses reveal the functional organisation of Pseudodiaptomus marinus copepod myofibrils. J Struct Biol 191(2):224–235
Xu X, Cheng J, Thrall MJ, Liu Z, Wang X, Wong STC (2013) Multimodal non-linear optical imaging for label-free differentiation of lung cancerous lesions from normal and desmoplastic tissues. Biomed Opt Express 4(12):2855
Galli R, Sablinskas V, Dasevicius D, Laurinavicius A, Jankevicius F, Koch E, Steiner G (2014) Non-linear optical microscopy of kidney tumours. J Biophotonics 7(1–2):23–27
Meyer T, Baumgartl M, Gottschall T, Pascher T, Wuttig A, Matthäus C, Romeike BFM, Brehm BR, Limpert J, Tünnermann A, Guntinas-Lichius O, Dietzek B, Schmitti M, Popp J (2013) A compact microscope setup for multimodal nonlinear imaging in clinics and its application to disease diagnostics. Analyst 138(14):4048–4057
Matthäus C, Cicchi R, Meyer T, Lattermann A, Schmitt M, Romeike BFM, Krafft C, Dietzek B, Brehm BR, Pavone FS, Popp J (2014) Multimodal non-linear imaging of atherosclerotic plaques differentiation of triglyceride and cholesterol deposits. J Innov Opt Health Sci 7(5):1450027
Legesse FB, Heuke S, Galler K, Hoffmann P, Schmitt M, Neugebauer U, Bauer M, Popp J (2016) Hepatic vitamin A content investigation using coherent anti-stokes Raman scattering microscopy. ChemPhysChem 17(24):4043–4051
Chernavskaia O, Heuke S, Vieth M, Friedrich O, Schürmann S, Atreya R, Stallmach A, Neurath MF, Waldner M, Petersen I, Schmitt M, Bocklitz T, Popp J (2016) Beyond endoscopic assessment in inflammatory bowel disease: real-time histology of disease activity by non-linear multimodal imaging. Sci Rep 6:29239
Pliss A, Kuzmin AN, Kachynski AV, Prasad PN (2010) Non-linear optical imaging and microspectrometry of the cell nucleus through the cell cycle. Biophys J 99(10):3483–3491
Pirhonen J, Arola J, Sädevirta S, Luukkonen P, Karppinen S-M, Pihlajaniemi T, Isomäki A, Hukkanen M, Yki-Järvinen H, Ikonen E (2016) Continuous grading of early fibrosis in NAFLD using label free imaging: a proof of concept study. PLoS One 11:e0147804
Haluszka D, Lőrincz K, Kiss N, Szipőcs R, Kuroli E, Gyöngyösi N, Wikonkál NM (2016) Diet-induced obesity skin changes monitored by in vivo SHG and ex vivo CARS microscopy. Biomed Opt Express 7(11):4480–4489
Mortati L, Divieto C, Sassi MP (2012) CARS and SHG microscopy to follow collagen production in living human corneal fibroblasts and mesenchymal stem cells in fibrin hydrogel 3D cultures. J Raman Spectrosc 43(5):675–680
Brackmann C, Bengtsson A, Alminger ML, Svanberg U, Enejder A (2011) Visualisation of β-carotene and starch granules in plant cell using CARS and SHG microscopy. J Raman Spectrosc 42(4):586–592
Brackmann C, Dahlberg J-O, Vrana NE, Lally C, Gatenholm P, Enejder A (2012) Non-linear microscopy of smooth muscle cells in artificial extracellular matrices made of cellulose. J Biophotonics 5(5–6):404–414
Mansfield J, Moger J, Green E, Moger C, Winlove CP (2013) Chemically specific imaging and in-situ chemical analysis of articular cartilage with stimulated Raman scattering. J Biophotonics 6(10):803–814
Kallaway C, Almond LM, Barr H, Wood J, Hutchings J, Kendall C, Stone N (2013) Advances in the clinical application of Raman spectroscopy for cancer diagnosis. Photodiagn Photodyn Ther 10(3):207–219
Watanabe K, Palonpon AF, Smith NI, Chiu L-d, Kasai A, Hashimoto H, Kawano S, Fujita K (2015) Structured line illumination Raman microscopy. Nat Commun 6:10095
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer-Verlag GmbH Germany, part of Springer Nature
About this chapter
Cite this chapter
Hobro, A.J., Smith, N.I. (2018). Label-Free Raman Imaging. In: Kumar, C. (eds) Nanotechnology Characterization Tools for Biosensing and Medical Diagnosis. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-662-56333-5_7
Download citation
DOI: https://doi.org/10.1007/978-3-662-56333-5_7
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-56332-8
Online ISBN: 978-3-662-56333-5
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)