Abstract
HDAC inhibition has been, for over a decade (and continues to remain), a highly competitive area. Hydroxamic acids represent the largest class of HDAC inhibitors. One product, SAHA is already approved and more than ten different chemical entities are in various clinical stages. A detailed discussion about compounds from various classes like phenyloxopropenyl, amidopropenyl analogues, spiropiperidines, biphenyl/arylamide/styrenyl, tetrahydroisoquinoline-based hydroxamic acid derivatives and N-hydroxyphenylacrylamide derivatives has been included along with the computational studies. It also covers brief details about the HDAC imaging agents. These successes as well as the enormous amount of experiences gained in preclinical and clinical studies may be useful—beyond the HDAC field—to future drug discovery programmes studying hydroxamic acid derivatives.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
1.1 Chromatin
The eukaryotic genome is packaged with histone proteins to form the chromatin, which allows condensing of over a metre of DNA into the small volume of the nucleus. The fundamental repeating unit of the chromatin that occurs generally after every 157–240 base pairs is the nucleosome. The nucleosome core itself is composed of a histone octamer consisting of two copies of H2A, H2B, H3 and H4 histones around which 147 base pairs of DNA are wrapped in 1.65 turns of a flat, left-handed superhelix. The stabilisation of the nucleosome occurs through a series of protein–protein interactions within the histone octamer and by electrostatic and hydrogen bonds between the proteins and the DNA. The core histones are composed of two distinct functional domains: the “histone-fold” motif sufficient for both histone–histone and histone–DNA contacts within the nucleosome, and N and C-terminal tail domains, which remain mostly unresolved in the crystal structures (Luger et al. 1997; Davey et al. 2002). These tails extend away from nucleosomal DNA and are mainly involved in interaction with other nucleosomes or with nuclear factors (Luger and Richmond 1998).
The chromatin itself condenses to more compact structures. Short-range nucleosome–nucleosome interactions result in folded chromatin fibres (“secondary chromatin structure”). Long-range interactions between individual nucleosomes result in fibre–fibre interactions and form tertiary chromatin structures. However, these arrangements are not highly defined structural states. Rather, the current view is to consider them as a continuum of various inter-convertible states at different levels of condensation (Horn and Peterson 2002; Luger et al. 2012).
These structural states of the nucleosomes have obviously a major impact on any process requiring access to genomic DNA, such as transcription, replication and DNA repair. Not surprisingly, a plethora of studies has shown that the chromatin structure plays a crucial role in the regulation of all of these processes. These states and their variation are determined by various factors. The DNA sequence of each nucleosome is unique; and this unique DNA sequence affects the nucleosome structure through its sequence-encoded susceptibility for being distorted into the tight superhelical conformation imposed by the histone octamer. Histones are among the most highly conserved proteins in terms of sequence and structure. However, histone variants have been identified for the histone subtypes, in particular for histone H2A and H3. These variants have an influence on the structures of the nucleosomes (Malik and Henikoff 2003; Henikoff et al. 2004; Brown 2001). Furthermore, several proteins have been found interacting with the chromatin. This adds further complexity in the structural regulations. Posttranslational modifications have been one of the most intensively studied aspects as regulatory factors for structural changes of nucleosomes. Posttranslational modifications are small chemical modifications to amino acid side chains of a protein after its translation. The histone tails which account for almost 30 % of the core histone sequences are, as already mentioned above, unstructured. Thus, they can be the subject of several posttranslational modifications, including acetylation, methylation, phosphorylation, sumoylation or ubiquitination.
1.2 The Histone Acetylase and Deacetylase Machinery
Already in 1964, Allfrey et al. first reported the isolation of acetylated and methylated histones and speculated well ahead on their time about their possible role in the regulation of RNA synthesis (Allfrey and Mirsky 1964; Allfrey et al. 1964). The authors hypothesised that these modifications of the histone structure, particularly acetylation, have an influence on RNA synthesis. Some years later, Riggs et al. (1977) found that the exposure of cultured cells to sodium n-butyrate caused a reversible accumulation of highly acetylated histones. However, the picture became clearer only during the early 1990s, when the first histone deacetylase (HDAC) was cloned in 1996 (Taunton et al. 1996). One year earlier, Kleff et al. (1995) had identified a gene encoding a yeast H4 acetyltransferase (HAT). Currently, a series of HATs and HDACs have been identified to be responsible for the tight control of the acetylation state of histones. HATs are enzymes which transfer an acetyl group from acetyl coenzyme A (acetyl CoA) onto to the ε-amino group of one or more lysine residues contained within the N-terminal tails of the histone proteins. The neutralisation of the basic charge of the tails reduces the electrostatic interactions with the DNA sequence leading to a nucleosome unwrapping (Simon et al. 2011). This relaxation of the chromatin conformation allows the transcriptional factors to access the gene promoter regions and the process of gene expression is facilitated. HATs can be classified into two different classes based on their functional localisation: the nuclear type A HATs and the cytoplasmic type B HATs. The latter ones are involved in the modification of newly synthesised histones before the assembly. Type A HATs can be further divided into five different classes based on structural and functional differences: GNAT, CBP/p300, transcriptional factors such as ATF2, nuclear hormone related (for example SRC4) and the MYST family proteins (Selvi and Kundu 2009; Grant and Berger 1999).
The counteracting histone deacetylases re-establish the positive charge in the N-terminus of the histone tails. This causes a tighter histone–DNA interaction and blocks the binding sites on promoters and thus inhibits gene transcription. The HDAC enzymes can be grouped into two families: the classical HDACs and the silent information regulator (Sir)-related protein (sirtuin) families. HDAC class III proteins (also known as sirtuins, Sirt1–Sirt7) form a structurally and mechanistically distinct class and are defined by their dependency on NAD+ as cofactor. They catalyse the removal of the acetyl group of the acetylated lysines by transferring it to the ribose moiety of NAD+ yielding O-acetyl-ADP-ribose and nicotinamide, which acts as a physiological inhibitor of the sirtuins by means of negative feedback (Huber and Superti-Furga 2011).
Classical HDACs are metalloenzymes harbouring a catalytic pocket with a Zn2+ ion. In humans, classical HDACs are grouped based on the homology to yeast enzymes in four distinct classes that vary in size and function (Grozinger et al. 1999; Gray and Ekstrom 2001; Lin et al. 2006; Ficner 2009). Class I HDACs, which comprise HDACs 1, 2, 3, and 8, share a certain degree of homology to the yeast Rpd3. These enzymes are generally nuclear proteins and are ubiquitously expressed in many human cell lines and tissues. HDAC11 is the sole member of Class IV HDACs and is found to be present in the nucleus. Class II HDACs, which are homologues to yeast Hda1, can be divided into two further sub-classes: class IIa (HDACs 4, 5, 7, 9) and class IIb HDACs (HDACs 6 and 10) (Verdin et al. 2003; Yang and Gregoire 2005). Class IIa HDACs contain a highly conserved C-terminal deacetylase domain with around 420 amino acids, and are homologous to yeast Hda1. However, their N-terminal domain, which has regulatory functions, does not show any similarity to HDACs in other classes. Class IIb HDACs have an additional deacetylase domain, although this duplication is partial in the case of HDAC10. Class II HDACs exhibit nucleocytoplasmic shuffling, suggesting their involvement in the deacetylation of non-histone substrates. For example, deacetylation of acetyl-α-tubulin is mediated by the second deacetylase domain of HDAC6 (L’Hernault and Rosenbaum 1985a, b; Hubbert et al. 2002; Haggarty et al. 2003). Recently, it has become clear that HDACs and not only the class II enzymes are involved in the deacetylation of a series of non-histone proteins (Glozak et al. 2005; Yang and Seto 2008; Singh et al. 2010; Yao and Yang 2011). Proteomic studies allowed to identify a remarkable amount of acetylation sites: Kim et al. (2006) found 388 acetylation sites on 195 proteins in HeLa cells and mouse liver mitochondria by immunoaffinity purification using an anti-acetyl lysine antibody; high-resolution mass spectrometry experiments permitted to identify even more impressive number of 3,600 lysine acetylation sites on 1,750 proteins (Choudhary et al. 2009). These results propose that acetylation/deacetylation is a regulatory modification that rivals phosphorylation in number of substrates (Kouzarides 2000; Choudhary et al. 2009). Furthermore, these acetylation sites were found on proteins involved in diverse cellular processes, such as chromatin remodelling, cell cycle, splicing, nuclear transport, signal transduction and apoptosis.
Several of these proteins were found to be relevant for tumorigenesis and cancer cell proliferation (Buchwald et al. 2009; Yao and Yang 2011). On the other hand, acetylation of histones results in changes of chromatin structures and this has an impact on any process requiring access to genomic DNA. Thus, it is not surprising that abnormal activities of HDACs and HATs have been found involved in the development of several diseases. Indeed, HDACs expression and their activity showed to be altered in many cancers. For example, HDACs are associated with the function of oncogenic-translocation products, such as PML-RARα in acute promyelocytic leukaemia. PML-RAR associate with a corepressor complex containing HDAC activity. This complex is able to inhibit the transcription of genes involved in haematopoietic differentiation, and thus contributing to the differentiation block found in this form of leukaemia (Grignani et al. 1998; Lin et al. 1998; Richon and O’Brien 2002). Subsequently, it has been shown that inhibition of HDACs resulted in a cell-cycle arrest and differentiation through an increased expression of p21WAF1/CIP1. Inhibition also affected tumour survival by blocking angiogenesis through the increased acetylation of HIF-1α and protein degradation through the acetylation of Hsp90. These findings have made HDACs as attractive targets for cancer therapy (Federico and Bagella 2011; Mercurio et al. 2010). The diverse mechanisms through which HDAC inhibitors exert their cytotoxic activity include induction of apoptosis by both intrinsic and extrinsic pathways, selective expression of repressed genes, cell-cycle arrest, DNA damage and repair, inhibition of angiogenesis, cell death due to accumulation of reactive oxygen species (ROS), autophagy and immunomodulatory effects (Bolden et al. 2006).
During the last 20 years, a variety of HDAC inhibitors have been developed based on the structures of the naturally occurring inhibitors, or discovered randomly in HDAC screening assays. The currently available HDAC inhibitors can be classified according to the nature of the metal binding group into different categories, i.e. hydroxamic acid-based, carboxylic acid-based, disulfide-based, epoxide-based, and anilide-based inhibitors. The hydroxamic acid-based HDAC inhibitors can be sub-classified according to the nature of cap structure (Table 1) (Elaut et al. 2007) or broadly classified as small-capped and large-capped hydroxamic acid-based inhibitors.
2 Histone Deacetylase Inhibitors: From n-Butyrate to SAHA
The first example of HDAC inhibition was reported in 1977. Riggs et al. (1977) found an accumulation of acetylated histones in the presence of sodium n-butyrate (Fig. 1). It was then questioned, if the accumulation of highly acetylated histones caused by this carboxylic acid was really linked to the observed induced differentiation in tumoral HeLa and Friend erythroleukemia cells. It took some years until Yoshida et al. (1987) found that Trichostatin A (TSA) (Fig. 1), a hydroxamic acid derivative originally found as fungistatic antibiotic (Tsuji et al. 1976), caused induction of Friend leukaemia cell differentiation at low nanomolar concentration. Three years later, the same authors disclosed that the (R) isomer of TSA was able to cause an accumulation of highly acetylated histones in vivo and to inhibit the activity of the partially purified histone deacetylase in vitro. The (S) isomer as well as the trichostatic acid did not show inhibitory activity and had no effect on the induction of Friend cell differentiation and the inhibition of the cell-cycle progression (Yoshida et al. 1990a). Furthermore, (R)-TSA was found to exhibit a significant lower inhibitory activity in a tumour cell line containing mutated histone deacetylases. These important observations provided the first evidence that the effect of TSA on cell proliferation and differentiation was directly related to the inhibition of histone deacetylases (Yoshida et al. 1987, 1990b). After 16 years later, suberoylanilide hydroxamic acid (SAHA) (Fig. 1) gained approval by FDA as the first HDAC inhibitor to be used for the treatment of cutaneous T cell lymphoma (CTCL) (Marks 2007; Marks and Breslow 2007). SAHA, disclosed in 1993 (Breslow et al. 1993), was the result of two decades of research activities carried out at the Columbia University and the Memorial-Sloan Kettering Cancer Center. In the early 1970s, Friend et al. (1971) found that DMSO acted as differentiating agent—murine erythroleukemia cells (MELC) after being placed in culture with dimethylsulfoxide turned red, suggesting the presence of haemoglobin. The search for new derivatives able to induce cytodifferentiation and growth arrest resulted in the discovery of polar solvent species, such as N-methylformamide or acetamide (Tanaka et al. 1975). While these compounds showed only a slight increase of activity compared to DMSO, dimeric amides proved to be much more potent. They were able to induce differentiation at low millimolar concentrations (Reuben et al. 1976, 1978). One example of this series, hexamethylene bisacetamide (HMBA), was selected for clinical studies. The trials showed that the compound was able to induce minor and partial remission in myelodysplastic syndrome and acute myelogenous leukaemia. However, the remission resulted to be transient and the doses required were not tolerated by cancer patients (Andreeff et al. 1992). Even though the biological target of these compounds was not identified, it was hypothesised that the dimeric amide structures may possibly act as chelating agents of metal ions. Exploration of new chemical entities led to the discovery of hydroxamic acids as well as bishydroxamates (Breslow et al. 1991; Richon et al. 1996). One example, SAHA, (but also other hydroxamic acid derivatives), was approximately 2,000-fold more potent than HMBA and was able to induce differentiation in MELC cells at low micromolar concentrations. Richon et al. (1998) confirmed that SAHA, like TSA, showed an HDAC inhibitory activity and caused accumulation of hyperacetylated histone H4 in cultured cells. One year later, the X-ray crystallographic structures of SAHA and TSA bound to the Aquifex aeolicus HDAC homologue histone deacetylase-like protein (HDLP) revealed that the hydroxamic acid moiety of both compounds is doubly coordinated to a zinc atom at the bottom of a cavity (Finnin et al. 1999). In 2000, SAHA entered clinical studies and, after intravenous and the oral administration, showed a good safety profile and antitumor activity in different malignancies, in particular haematological malignancies (Kelly et al. 2003, 2005). Finally, in October 2006, FDA approved the compound as first HDAC inhibitor for the treatment of CTCL, after that a significant response rate in prior therapy-resistant CTCL patients was observed (Mann et al. 2007).
3 Histone Deacetylase Inhibitors: Orally Available Hydroxamic Acid Derivatives
While SAHA showed a considerable efficacy in different haematological malignancies in clinical trials, patient response in other cancers remained much more uncertain and often rather limited (Graham et al. 2009; Mercurio et al. 2010). SAHA had demonstrated some activity in patients with advanced solid tumours in phase I trials. However, phase II studies in patients with breast, colorectal or non-small cell lung cancer showed a limited drug exposure, which did not allow a reliable efficacy analysis (Vansteenkiste et al. 2008). SAHA was well tolerated, but exhibited only medium potency (Marks and Breslow 2007) and was cleared rapidly (Kelly et al. 2003). Short half-lives in vitro as well as in vivo are quite common to several first generation hydroxamic acid inhibitors (Elaut et al. 2007), and thus hydroxamates are often considered as poor drugs. They are frequently down prioritised in drug discovery programmes because of their poor physicochemical and ADME properties, despite their good in vitro potency (Elaut et al. 2007; Flipo et al. 2009).
However, the overall impressive in vitro and in vivo data of the first drugs made HDACs and their inhibition an attractive target for several drug discovery programmes. Research efforts were directed to highly potent compounds with a prolonged in vivo exposure and generally a better pharmacokinetic (PK) profile. In fact, we can find a significant number of research groups, both from industry and academia, involved in the exploration of novel chemical entities with HDAC inhibitory properties. Almost 400 PCT patent applications claiming new HDAC inhibiting agents is just one indicator of the impressive amount of efforts done in this field during the last decade. It is noticeable that more than 50 % of the applications are related to hydroxamic acid derivatives (Thaler 2012). Another sign is that the significant number of molecules have entered clinical studies. Around 20 chemical entities, either alone or in combination, have entered clinical studies for the treatment of several diseases, mainly tumours. Among them there is a considerable amount of hydroxamic acid derivatives e.g. LBH589 or panobinostat (Novartis) (Atadja 2009; Neri et al. 2012), and PXD101 or belinostat (Topotarget) (Steele et al. 2008) are currently in phase III studies, ITF-2357 or givinostat (Italfarmaco) (Rambaldi et al. 2012), PCI-24781 or abexinostat (Servier, Pharmacyclics) (Buggy et al. 2006), SB939 or pracinostat (S*BIO Pte Ltd) (Novotny-Diermayr et al. 2010; Wang et al. 2011), JNJ-26481585 or quisinostat (Johnson & Johnson) (Arts et al. 2009; Tong et al. 2010) and 4SC-201 or resminostat (4SC) (Mandl-Weber et al. 2010; Brunetto et al. 2009) have reached clinical phase II (Fig. 2). Examples in phase I can be given as AR-42 or OSU-HDAC-42 (Arno Therapeutics) (Kulp et al. 2006; Lu et al. 2005), CG-200745 (CrystalGenomics Inc) (Hwang et al. 2012), CHR-3996 (Chroma Therapeutics) (Donald et al. 2010; Banerji et al. 2010), the HDAC6 selective ACY-1215 (Acetylon Pharmaceuticals Inc) (Santo et al. 2012) and the dual kinase and HDAC inhibitor CUDC-101 (Curis and Ligand Pharmaceuticals) (Cai et al. 2010; Lai et al. 2010; Shimizu et al. 2010) (Fig. 3).
In the following sections, the research efforts directed towards the development of hydroxamic acid derivatives with improved PK properties are discussed.
3.1 From SB639 to Pracinostat
Scientists from the Singaporean biotech company S*Bio Pte explored different fused heterocyclic rings such as benzimidazoles as new linkers for low molecular weight HDAC inhibiting agents. This research led to the discovery of the first lead compound, SB639 (Fig. 4) (Wang et al. 2009). The compound showed good in vitro potency with an IC50 value of 0.035 ± 0.016 μM against HDAC1 and antiproliferative activity against the human colon cancer cell line COLO205 in the submicromolar range (IC50 = 0.14 ± 0.05 μM). The compound was stable in human and dog liver microsomes with a half-life of around 1 h. However, the half-life dropped in rat and mouse microsomes to 6 and 3 min, respectively. The compound showed high clearance rates in mice (15.8 L/h/kg) and in rats (3.84 L/h/kg). The oral bioavailability was 13 % (mice) and 10.5 % (rats), respectively (Venkatesh et al. 2007). Nevertheless, the plasma concentration of the compound reached levels above the HDAC IC50 value and the compound showed antitumor activity in HCT116 tumour-bearing nude mice after oral administration (Wang et al. 2009).
Metabolic profiling experiments in rat liver hepatocytes showed that the major metabolic paths of SB639 were the oxidation of the pyrrolidinylethyl group and the reduction of the hydroxamic acid to amide (Wang et al. 2011). Further development of this series was carried out considering particularly the metabolic stability of the compounds. This expansion resulted in several compounds. Among them, compound SB939 (Fig. 4) was found to be potent against HDAC1 with an IC50 value of 0.077 ± 0.014 μM and showed antiproliferative activity against a series of cancer cell lines in the submicromolar range. For example, the compound with an IC50 of 0.48 ± 0.27 μM was around six times more active than SAHA (IC50 = 2.85 ± 0.27 μM) against HCT116 cells. An interesting observation was that the microsomal liability in human and mouse microsomes generally increased with higher lipophilicity of the molecules. SB939 as well as several other derivatives were relatively stable in human and dog microsomes. On the other hand, a major variability was found in rodents. Consistent with the in vitro microsomal data, SB939 exhibited an oral bioavailability of 65, 34 and 3.1 % in beagle dogs, nude mice and Wistar rats, respectively. Furthermore, the compound had high systemic clearance (relative to liver blood flow) of 1.5, 9.2 and 4.5 L/h/kg and high volume of distribution at steady state of 4.2, 3.5 and 1.7 l/kg in dog, mouse and rat, respectively (Jayaraman et al. 2011). The superior in vitro activities and PK properties compared to SAHA translated into a higher antitumor efficacy in a HCT116 tumour xenograft model. SB939 was approximately twice as efficacious at the maximum tolerated dose as SAHA: the tumour growth inhibition for SAHA was 48 % at 200 mg/kg compared to 94 % for SB939 at 100 mg/kg. In addition, an accumulation of the compound was observed in tumour tissues (Novotny-Diermayr et al. 2010). This was not found after administering the reference of SAHA. In addition, SB939 has good aqueous solubility and high permeability in human Caco-2 cells with low efflux. These characteristics are indicative of a high-intestinal absorption in vivo (Jayaraman et al. 2011). In fact, phase I studies conducted in patients with advanced solid malignancies showed that the pharmacologically active concentrations were achieved already at the lowest dose of 10 mg (Jayaraman et al. 2011). The drug was rapidly absorbed (t max = 1–3 h). The mean elimination half-life was around 7 h and oral clearance was 53 ± 8.5 L/h. C max and AUC were dose proportional between 10 and 60 mg doses. Furthermore, no substantial accumulation of SB939 on day 15 following repeated dosing was found. At the 60 mg dose high acetylation levels was found in all patients indicating sustained target inhibition. Two patients experienced prolonged disease stabilisation (Yong et al. 2009). An interesting aspect is that the human PK was successfully predicted based on the in vitro ADME data using an ADME simulator and allometric scaling (Jayaraman et al. 2011).
3.2 CHR-3996
The HDAC project at the British biotech company, Chroma Therapeutics, was focused on the synthesis of pyrimidine hydroxamate derivatives bound to bicyclic hexahydropyrrolo[3,4-c]pyrrole and azabicyclo[3,1,0]hexane linkers (Moffat et al. 2010). The inhibitory activities of around 25 described azabicyclo[3,1,0]hexanes varied only slightly and were in the low nanomolar range against HDACs from nuclear HeLa extracts. This narrow range in activity despite substantial variations of the linker structures is not uncommon; for example, as already described above, a similar behaviour was experienced in house for several examples within the amidopropenyl and the spiropiperidine series (cf. Sects. 3.4 and 3.5). The compounds showed good antiproliferative activity in tumoral HCT116 cells with GI50 values in the micro- or submicromolar range. In particular, it was found that diminishing the polar surface area was beneficial for the cell growth inhibition. PK studies using oral cassette dosing allowed identifying CHR-3996 as one lead compound (Fig. 5). The compound was potent against HDACs 1, 2 and 3 with IC50 values of 3, 4 and 7 nM, respectively. On the other hand, it was significantly less active against the class II members HDACs 5 and 6. CHR-3996 showed good oral bioavailability in rat and dog of 27 and 40 %, respectively. Furthermore, the compound exhibited high systemic clearance (relative to liver blood flow) of 4.2 (dog) and 8.0 L/h/kg (rat), respectively. In vivo HCT116 xenograft experiments showed that the compound had a significant antitumor activity: administration of 50 mg/kg once daily orally resulted in almost complete inhibition of the tumour. CHR-3996 advanced into clinical studies. Phase I clinical studies showed that the compound was rapidly absorbed, had a mean elimination plasma half-life of 3 h and mean AUC values exceeding the levels effective in xenograft studies at doses ≥40 mg. Five patients demonstrated stable disease for at least two cycles (Banerji et al. 2010, 2012).
3.3 Phenylhydroxamic Acid Derivatives (AR-42)
The phenylhydroxamic acid derivative AR-42 (known also as OSU-HDAC-42 or (S)-HDAC-42) (Fig. 6) was originated at the Ohio State University. The compound resulted from an exploration of new HDAC inhibitors composed of short-chain fatty acids linked to a hydroxamic acid group as Zn2+-chelating motif (Lu et al. 2005). The compound showed good HDAC inhibitory potency with an IC50 value of 16 nM and was around five times more potent than its corresponding R-isomer. Antiproliferative activities in three myeloma cell lines, IM-9, RPMI-8226 and U266, were in the submicromolar range. The compound was around four to seven times more active than SAHA (Bai et al. 2011). This good potency, but even more its pharamacokinetic behaviour in rodents, has made the compound an attractive agent for further studies. Indeed, Ar-42 displayed a good oral bioavailability in rats (Cheng et al. 2006a) and in mice (Cheng et al. 2006b): the oral bioavailability was ~100 % in rats and 27.4 % in mice. Total body clearance was 1.40 L/h/kg (rats) and 1.47 L/h/kg (mice). The good in vitro potency as well as the PK profile translated in good antitumor activity in different in vivo efficacy models. For example, the compound administered 25 mg/kg daily and 50 mg/kg every other day significantly inhibited tumour growth in mouse PC-3 tumour xenografts by 52 and 67 %, respectively (Kulp et al. 2006). The compound was licensed to Arno Therapeutics, which is currently conducting phase I/IIa trials in adult patients with relapsed or recurrent haematological malignancies and solid tumours (http://clinicaltrials.gov/ct2/show/NCT01129193?term=AR-42&rank=1).
3.4 Phenyloxopropenyl and Amidopropenyl Analogues
The HDAC inhibitory programme in DAC was initiated in collaboration with Antonello Mai from the University of Rome, who had intensively studied aroyl-pyrrolyl hydroxyamides (APHAs) as histone deacetylase inhibitors (Massa et al. 2001; Mai et al. 2005a, b, 2006) and identified several class II selective compounds. It is noticeable that these research efforts have origins in programmes carried out by the same group in the late 1980s, when pyrrole-containing analogues of trichostatin were explored for their antifungal, antibacterial and antiviral activities (Massa et al. 1990) but ended with HDAC inhibitory (Massa et al. 2001) activity.
A small library of aryl- and heteroaryl-hydroxamic acid fragments was designed and subjected for biological characterisation (Thaler et al. 2010a). The HDAC inhibitory activity was performed using HeLa nuclear extract as enzyme source and the percentage of inhibition of the enzymes was assessed at 0.1, 1.0 and 10 μM concentration of the inhibitors. As shown in Table 2, the phenyl hydroxamate (1) was almost inactive under the experimental conditions, whereas 2 was moderately active with 36 % enzyme inhibition at 1 μM. Introduction of an acrylic group (3) led to a further increase in activity. Within the pyridine-acrylhydroxamic acid series, the pyridine-2-yl derivative (4) demonstrated to be the most potent compound and the activity decreased from the pyridine-2-yl to the pyridine-4-yl analogue (6). The electron-rich pyrrolyl derivative (7), present in the APHA series, was less potent than the other acryl hydroxamates.
The heteroaryl-hydroxamate 4 and the phenylacryl-hydroxamate 3 were selected for further expansion. In the first step, a scaffold with a phenyloxopropenyl moiety (Mai et al. 2005a) positioned in meta or para position with respect to the acrylhydroxamate substituent was prepared. The meta phenyl analogue 11 (Table 3), with an IC50 value of 0.085 μM, was almost 20 times more potent than the corresponding para derivative 8 (IC50 = 1.48 μM). A slightly different behaviour was found in the pyridinyl series: the 2,5 di-substituted derivative 10 and the 2,6 di-substituted analogue 12 exhibited similar IC50 value (~0.02 μM) and were around 15 times more active than the 2,4-substituted derivative 9. Further, the compounds and in particular the pyridinyl derivatives showed good antiproliferative effects in the chronic myelogenous leukaemia cell line K562. In specific, the cellular IC50 values for 10, 12 and 9 ranged between 0.5 and 1 μM. The phenylacrylates 8 and 11 exhibited IC50 values of 2.3–3.2 μM, respectively.

The good biochemical and cellular inhibitory activity of this series, however, was compromised with an overall low metabolic stability in microsomes. It was observed that the compounds were almost completely degraded in mouse microsomes after 30 min incubation: the percentage of the recovery ranged from just 2 % (8) to 14 % (9) (Table 3). Similar data were obtained in human microsome preparations with the exception of compound 12, which was more stable than the other examples. Metabolic profiling experiments of 8 in mouse microsomes showed that the major sites of metabolism were the hydroxamic acid moiety and the unsaturated ketone (Thaler et al. 2009). While the major metabolite of the hydroxamic acid moiety was the acrylic acid (8a), α,β-unsaturated ketone was either reduced to the corresponding alcohol (8b) or to the saturated ketone derivative (8c) (Fig. 7).
A similar outcome was found for the pyridine derivative 9. Since the hydroxamic acid was essential for the biological activity, research efforts were directed towards new chemical entities devoid of the metabolically unstable α,β-unsaturated ketone. One approach was the replacement of the phenyl by 4-phenylpiperazine group (Thaler et al. 2010b) (Fig. 8).
Similar to the phenyloxopropenyl series, 4-phenylpiperazinyl analogues of the following four linkers were prepared: phenylacrylate, meta and para substituted, and pyridin-2-ylacrylate, substituted in positions 5 or 6. As summarised in Table 4, replacement of the phenyl led to a minor decrease in the HDAC inhibitory activity. However, the trends within both series were quite comparable: the meta substituted phenylacrylate derivative 15 exhibited an IC50 value of 0.354 μM, and was about 15 times more potent than the corresponding para substituted (13). In the pyridinyl series, the differences between 2,5 and 2,6 di-substituted pyridin-2-ylacrylates were smaller: compound 16 exhibited an IC50 value of 0.069 μM and was the most potent example in this series, around four times more than 14. Comparison of the compounds containing a central phenyl ring with those with the pyridinyl ring showed the inhibitors with the pyridinyl ring were generally more potent. Compounds 14, 15 and 16 exhibited comparable antiproliferative potency in leukemic K562 and human colon cancer HCT116 cells with IC50 values of around 1 μM. Hydroxamate 13, in line with the lower enzymatic inhibition, had IC50 values in K562 and HCT116 cells of 3.77 and 5.77 μM, respectively.

The major difference between the phenyloxopropenyl and the 4-phenylpiperazinyl series was their microsomal stability: all four amidopropenyl compounds were metabolised <50 % in the tested conditions and were remarkably more stable than the ketone analogues. The only exception was 13 with a recovery of just 27 % in mouse microsomal preparation. These results confirmed that the α,β-unsaturated ketone moiety was a major site of metabolic liability and replacement with an amide group resulted in stable compounds in microsomal preparations.
These encouraging data prompted to explore this series in a greater depth and the hydroxamic acid 16 was selected as starting point for this expansion. The synthetic efforts were concentrated on a series of cyclic amine derivatives substituted by phenyl. This selection was based on the fact that hydrophobic aromatic groups represent a common surface recognition domain in several HDAC inhibitors (Miller et al. 2003). As shown in Table 5, IC50 values ranged between 0.025 μM (19) and 0.294 μM (17). These differences in activities are quite small compared to the substantial structural variations. These findings can be ascribed to the fact that the surface region of the HDAC enzyme is highly flexible and able to accommodate inhibitors with different capping groups (Somoza et al. 2004; Wang et al. 2005). The compounds exhibited comparable antiproliferative activities in K562 and HCT116 cells; compound 19 (IC50 = 0.445 μM) was around four-fold more active than the least potent analogue 20 (IC50 = 1.730 μM) in K562 cells. Furthermore, hydroxamic acid 19 was around five times more active than 20 (IC50 = 1.470 μM) in HCT116 cells.

All compounds exhibited a good stability in mouse and human microsomes with <50 % of the inhibitors metabolised. The only outlier was compound 21, with 29 and 46 % remaining in mouse and human microsomes, respectively.
However, the remarkable stability of the compounds in microsomes was not confirmed by in vivo PK experiments in mice. Three selected derivatives, the aryl-hydroxamate 15 and the two heteroaryl-hydroxamates 16 and 19, displayed a low oral bioavailability with F ≤ 10 % (Table 6). The compounds also showed very high clearance rates. In particular, inhibitors 15 and 16 showed clearance rates, which exceed the hepatic blood flow of 5.4 L/h/kg in mice (Davies and Morris 1993).
Further investigations revealed that the three representative compounds had high clearance rates in rat and human hepatocytes. These observations indicated a metabolic degradation of these hydroxamic acid derivatives catalysed preferentially by non-microsomal enzymes. Reduction of the hydroxamic acid group to its corresponding amide was identified for the three compounds as one relevant biotransformation pathway. The same outcome was found for SAHA, which was studied as reference compound. Similar results had been previously reported for TSA (Elaut et al. 2002). These observations suggest that this metabolic behaviour is likely to be common to further hydroxamic acids.
These results are in agreement with the observed in vivo PK behaviour in mice. They show that the metabolic stability of these hydroxamates in microsomes was neither found in hepatocytes nor in vivo. ADME experiments employing microsomes are widely used and they are often very suitable indicators for the in vivo PK behaviour. However, our experiences as well as other cases have shown that microsomes may not be sufficient for predicting in vivo clearances of structures containing hydroxamic acid moieties. Hepatocyte preparations, even though more expensive, demonstrated to be a more valid alternative for predicting the in vivo clearance behaviour of the studied hydroxamates.
3.5 Spiropiperidines
The phenyloxopropenyl and the amidopropenyl series resulted in the discovery of several compounds with good in vitro biochemical and antiproliferative activity. However, none of them exhibited an oral bioavailability superior to 20 % in mice (Thaler et al. 2010a, b). Therefore, various other scaffolds were evaluated. Major focus was particularly on privileged structures, which already had provided derivatives with good oral bioavailability. Privileged structures with their inherent affinity for diverse biological targets represent an ideal source of core scaffolds for the design of molecules able to target various receptors (DeSimone et al. 2004; Costantino and Barlocco 2006). One of them is the conformationally constraint 4-oxospiro[chroman-2,4′-piperidine] ring system, which is, for example, present in antiarrhythmic agents (Elliott et al. 1992).
Some 4-oxospirochromanes linked to an acrylhydroxamic acid group as zinc binding motif were prepared based on the results obtained during the exploration of different hydroxamic acid fragments (Fig. 9) (Thaler et al. 2010a; Varasi et al. 2011). In the first run, unsubstituted and the N-methyl, N-acetyl and N-benzyl substituted piperidine analogues (22–25, Table 7) were synthesised. As shown in Table 7, all compounds inhibited the HDAC activity in the submicromolar range. No major differences in potencies were found: the unsubstituted spiropiperidine (22) with an IC50 value of 0.082 μM was around eight-fold more active than the least potent one within this series, the acetyl analogue (24). Compound 24 was also the least potent compound within this group against leukemic K562 and colon cancer HCT116 cells, while the other analogues showed good antiproliferative activity in the micromolar or submicromolar range. Next, synthetic efforts were focused on modifications of the central spirochromane core structure. Shift of the acrylamide moiety to position 7 and reduction of the chromane ring size to spirobenzofuran furnished compounds, which were generally less active (Varasi et al. 2011). Then, the importance of the ketone group was assessed. The hydroxy derivative 25a (Fig. 10) and the chromene 25b exhibited IC50 values of 1.21 and 1.19 μM, respectively, and were around ten-fold less active than the 4-oxo-chromane 25. Also the chromane 25c (IC50 = 9.0 μM) and the N-acetyl derivatives 25d (IC50 = 5.15 μM) were significantly less active. In addition, these four compounds showed a lower antiproliferative activity in K562 and HCT116 cells confirming the importance of the carbonyl group for the biochemical and cellular potency (Varasi et al. 2011).

Analogues of the 4-Oxospirochromane 25 (Table 1)
These explorations consolidated the 4-oxospiro[chromane-2,4′-piperidine]-6-yl series as the most promising scaffold for a further expansion. For this purpose, different N-piperidine substituents and a number of compounds were made around the benzyl derivative 25 (Thaler et al. 2012). The few representatives 22–25, prepared in the first run, showed no major differences in potencies. This indicated that variations on this site were likely well tolerated. But at that stage, one of the major objectives was to obtain inhibitors with a good ADME profile with an acceptable activity.
Not surprisingly, no major differences in HDAC inhibitory activities were found. As shown in Table 7, the phenyl spiropiperidine (26) exhibited an IC50 value of 0.41 μM, whereas all other tested compounds had IC50 values in the range of 0.10–0.28 μM. All compounds showed good antiproliferative activities in K562 and HCT116 cells with IC50 values in the submicromolar to low micromolar range. The 2-phenylethyl analogue (27) was the most potent and the spirocycle (26) the least potent compound among the unsubstituted derivatives. Further elongation of the alkyl chain did not result in an increase of activity. Compound 32 with IC50 value of 0.15 μM was the most potent compound against K562 and HCT116 cells among the substituted benzyl analogues.
But as already mentioned earlier, the main selection criteria were set by the stability experiments. In fact, the tested compounds already showed different metabolic behaviours in microsomal preparations. In specific, the phenyl analogue (26) was rapidly metabolised both in mouse and human microsomes, with only 23 and 4 % of the product recovered unmodified after 30 min of incubation. The benzyl spirocycle (25) demonstrated a better stability with 32 % remaining in the mouse and 55 % in the human preparation. The 2-phenylethyl analogue (27) was even more stable with 57 % (mouse) and 66 % (human) remaining, respectively. However, this trend did not continue, when the alkyl chain was further increased: the 3-phenylpropyl spirocycle (28) had a microsomal stability similar or even lower than the benzyl derivative (25). The 4-fluorobenzyl derivative (31) emerged as the most stable derivative among the substituted benzyl analogues with 47 and 63 % of the compound recovered in mouse and human microsomes, respectively. Within the fluoro series, the 2-fluorobenzyl derivative (29) had the lowest stability compared to the 3- and 4-fluoro analogues (30) and (31) with 23 and 36 % remaining in mouse and human microsomes, respectively. On the other hand, no major differences were observed among the ortho, meta and para methoxybenzyl inhibitors (32–34).
Three representative examples 25, 27 and 31 displayed good oral bioavailability in mouse PK experiments with an oral bioavailability F of 31.5, 51.1 and 27.6 %, respectively (Table 8). Spirocycle 25 showed a high systemic plasma clearance with 5.88 mL/h/kg, similar to the hepatic flood flow of 5.4 mL/h/kg in mice (Davies and Morris 1993). The other two examples, 27 and 31, exhibited lower clearance rates with 4.79 and 3.35 mL/h/kg, respectively. This outcome is in line with their superior stability in human and mouse microsomes compared to 25. Estimated elimination half-lives for the three examples ranged between 8 and 13 h. All three examples had very high estimated steady-state volume of distribution (V ss), suggesting substantial distribution into tissues. The V ss of 27 with 8.6 L/kg was in the same range of the compound 25 (8.9 L/kg). The 4-fluorobenzyl derivative (31) had a V ss of 5.9 L/kg, lower than the other two spirocycles, 25 and 27, but still ~8 times the total body water in mice (Table 8) (Davies and Morris 1993).
The three spirocycles also showed good in vivo antitumor activity in an HCT116 xenograft model after oral administration. Consistent with the PK experiments, the 2-phenylethyl and the para fluorobenzyl analogues (27 and 31) were slightly more potent than 25. In specific, the calculated T/C were 0.2 at 75 mg/kg for 31 and 0.2 and 0.34 at 150 mg/kg for 27 and 25, respectively (Varasi et al. 2011; Thaler et al. 2012).
3.6 Biphenyl/Arylamide/Styrenyl Hydroxamic Acid Analogues
In mid 1999, the Abbott Medicinal Chemistry started to work on HDAC programme with the preparation and evaluation of several series of hydroxamic acids. Two HDAC inhibitors, the macrocyclic hydroxamate (35) and the α-ketoamide (36), were evaluated for their selectivity profile in comparison to SAHA and MS-275, but no selectivity was observed for a particular class of HDAC.


The biphenyl ether series was among the first examined families leading to the identification of 37a (Table 9) as a screening hit and having complimentary fit to the proposed general structure for HDAC inhibitors (Jung et al. 1999; Jung 2001). The SAR studies concluded that the 6-methylene linker was optimal and a longer/shorter tether showed loss of potency (37a, 37b–d) (Table 9). Reduction of double bond to saturated alkyl ether (37f) showed loss of inhibitory activity. Similar results were observed for compounds 38a–b (Woo et al. 2002).
The nitrile group on the biphenyl was not important for activity (37f and 37g), but replacement of the biphenyl moiety with phenyl resulted in a significant loss of potency (37h). Replacement of the tether ether of 37i with alkenyl (37j) and alkyl (37k) showed similar trends as with compounds 37g, 37e and 37h. All compounds had modest cellular potency (>1 μM) against HT1080 and MDA435 cell lines and were not considered for further studies.
The poor cellular activity profile of SAHA and the ‘reverse-amide’ series stimulated the evaluation of some biphenyl analogues (39a–i, Table 10). The ‘SAHA-like’ amides, the para-biphenyl and meta-biphenyl isomers (39a and 39b) were found equipotent and had shown submicromolar cellular activity against both cell lines (Remiszewski et al. 2002). The series has shown beneficial effect by introducing a 4-phenylthiazol-2-yl moiety (39c) as compared to 39a or 39b. The meta-biphenyl analogue (39e) was more potent than the para-biphenyl isomer (39d); the reverse-amide 4-phenyl thiazole (39f) was equipotent with the meta-biphenyl analogue (39e), but exhibited a better cellular activity. Some of the favourable replacements include the 5-phenyl oxazol-2-yl (39g), 5-phenyl fur-2-yl (39h) and 5-phenylthiophen-2-yl (39i) (Curtin et al. 2002).
Some heteroaromatic phenyl replacements on the ‘reverse-amide’ template (40a) are shown in Table 11. The 2-pyridyl and 2-pyrrole substituents (40b and 40c) had given equipotent inhibitors and the polycyclic heterocycles (40d and 40e) had shown improved potency. N-substitution on the indole moiety of 40d with phenyl (40h) or substituents on the indole phenyl ring (40i) led to an increase of potency, while the N-methyl analogue (40g) was less active (Wada et al. 2003).

Examples of expected improvements of inhibitory activity by using a highly rigid, lipophilic tether like analogues are shown in Table 12. Submicromolar activity was observed with the styrenyl tethers (41a, 41b) and the potency was diminished with saturation of double bond (41c). The para substituted phenyl derivative 41a resulted to be slightly more active than the corresponding 1,3-phenyl analogue (41d). The 2-anilino-2-oxo-ethyl analogue of 41a resulted to be almost three times more active in the HDAC inhibitory assay and showed an antiproliferative activity with IC50 values in the submicromolar range. Further modifications of 41e as in meta-biphenyl analogue (41g) and 4-phenylthiazol-2-yl-hydroxamate (41h) had shown excellent inhibitory and cellular activity (Curtin et al. 2002).
In a further study, the linking amide functionality in the arylamide 39c was removed giving hydroxamates 42a and 42b. Both compounds showed a lower HDAC inhibitory activity with IC50 values of 19 nM (42a) and 9 nM (42b), respectively, as well as a reduced cellular activity (Dai et al. 2003).


As illustrated in the previous examples, variations of the cap group are in general well tolerated. Even significant structural variation, e.g. by introducing a macrocyclic cap group resulted in derivatives with potent HDAC inhibitory activity (Sternson et al. 2001). For example, the hydroxamic acid (43) having a cyclic tetrapeptide terminal group has shown reversible HDAC inhibition (HDAC1 (IC50 = 1.9 nM) with a slight selectivity over HDAC6 (IC50 = 19 nM).

Modifications of the peptidic analogue 43 by replacing the cyclic tetrapeptide with the peptidomimetic core of Abbott’s macrocyclic matrix metalloproteinase inhibitor (Steinman et al. 1998) resulted in derivatives, which had reduced HDAC inhibitory activity. For example, the macrocyclic hydroxamic acid inhibitors 44 and 45a (Fig. 11) exhibited HDAC IC50 values of 2.1 μM and 38 nM, respectively (Curtin et al. 2002).
The studies concluded that the parent macrocycle succinimide 45a had significant antiproliferative activity, but any further modifications did not make any significant improvement in the HDAC and cellular growth inhibitory activity (Table 13).
The nature of the amino acidic side chain was important for activity. Compound 45g, devoid of benzyl moiety, resulted to be a weak HDAC inhibitor with no measureable antiproliferative potency at the tested conditions. Compound 45h, which comprises an L-cyclohexylalanine residue, regained the potency and the para methoxy-Ph analogue 45i exhibited potencies similar to the macrocyclic inhibitor 45a. The penta- and hexa-methylene tethered analogues of the disubstituted succinimide (45e and 45j) had similar activity, while the tetra-methylene analogue (45k) was much less potent (Curtin et al. 2002; Curtin and Glaser 2003).
3.7 Tetrahydroisoquinoline-Based Hydroxamic Acid Derivatives
A novel class of tetrahydroisoquinoline-based hydroxamic acid analogues was evaluated in vitro and in vivo as potential anticancer and HDAC inhibitor agents at the Shandong University (Zhang et al. 2010, 2011a, b). The designing was based on the common pharmacophore having three parts, the Zn2+ binding group and a linker and the surface recognition domain (Miller et al. 2003). The 1,2,3,4-tetrahydroisoquinoline-3-carboxylic acid having distinct geometrical conformation and biological activity (Klutchko et al. 1986) was used to design novel HDAC inhibitors (46, 47, 48, Table 14) (Zhang et al. 2010). Most of the target compounds showed an inhibitory activity against HDAC8 comparable to SAHA, some of them were more potent than the reference compound. Modifications of the R 1, both in the BOC series 46 and well as for the free amine analogues 47, were well tolerated with IC50 values ranging from 1.06 (47l) to 8.21 μM (47a). Compounds with aromatic R 1 groups (46a–m, 47a–m) were slightly more potent than the compounds bearing aliphatic R 1 groups (46n–p, 47n–p).


A further expansion led to compound 48a, which resulted to be more potent than the derivatives in the 46 or 47 series, such as 46d. On the other hand, the rigid compounds 48b and 48c were found almost inactive. These results indicated that variations of the substituents on the secondary amine influence the inhibitory activity more than the R1 groups in the 46 or 47 series. Some of the compounds showed also better antiproliferative activity against HCT116, SKOV3 and HL60 cell lines.






Based on the lead structure 46d, further extensive structure–activity relationship studies were performed in order to optimise anticancer activities of this series (Zhang et al. 2011b). For this purpose, the R1 moiety (4-OCH3–Ph) was kept fixed and the Boc group was replaced with other functional groups as shown in Table 15; the less polar BOC protected compounds (50a–h) were more potent than the corresponding deprotected analogues (51a–h). The reduction of the amide group to a tertiary amine group was found detrimental (50l–n). The antiproliferative activities of potent compounds were found similar to that of SAHA in A549 (lung cancer) and MDA-MB-231 (breast cancer) cell lines (Zhang et al. 2011b). Furthermore, compound 50e showed in vivo antitumor activity comparable to SAHA.

As further exploration, the synthetic efforts were directed towards compounds (examples 52a–c) having the tertiary butyloxycarbonyl group and a better water solubility (Fig. 12), but the in vitro results were inferior to those of 50e. A further exploration culminated in compounds 53a, b (Fig. 13) ,which exhibited mid-nMIC50 values against HDAC8 and potent growth inhibition in multiple tumour cell lines. Compounds 50e and 52a–c were selected for in vivo activity experiments and showed good anticancer potencies comparable to SAHA in a human breast carcinoma (MDA-MB-231) xenograft model after ip administration.
3.8 N-Hydroxy Phenylacrylamide Derivatives
Research activities at the Novartis Institutes for Biomedical Research resulted in a series of potent HDAC inhibitors, among them the clinical candidates LAQ824 and LBH589 in the early 2000 (Remiszewski et al. 2003). The development of second-generation HDAC inhibitors was aimed towards improving in vitro potency and in vivo efficacy of hydroxamate-based HDAC inhibitors and simultaneous elimination of interactions with cardiac ion channels linked to QT prolongation (Shultz et al. 2011). The N-hydroxy phenylacrymide pharmacophore explored also in their first-generation HDAC inhibitors was modified using structure-based drug design, physicochemical property modulation and a matched molecular pair approaches and a series of data were prepared (Shultz et al. 2011, see Table 16). The cardiac safety profile was measured using the iCSI (in vitro cardiac safety index) parameter. Compounds 54a, 54b and 54c were found to be highly potent, efficacious, and having greater in vitro cardiac safety as compared to several HDAC inhibitors in clinical trials. Steric effect and amphiphilicity were identified as the contributing parameters for the activity (Table 16). Compounds 54a and 54c showed significant in vivo antitumor activity in an HCT116 xenograft model. The tumour growth regression in mice treated with 10 mg/kg of 54a, administrated iv, qd for eight total doses, or with 54c, administered 50 mg/kg iv, qd for 13 total doses, resulted to be 22 or 9 %, respectively.

4 Isoform Selective HDAC Inhibitors
Sodium n-butyrate, trichostatin and SAHA have in common that their biological targets were unknown at the time when their antitumor activity in different cell lines was assessed. Their activity against HDACs has been only ascertained afterwards. Most of HDAC inhibitors are natural products or derivatives, such as the cyclic tetrapeptide trapoxin, which contains an epoxide group capable of binding irreversibly with histone deacetylase enzymes (Itazaki et al. 1990; Kijima et al. 1993), and the thiol FK228. FK228 was the second HDAC inhibitor reaching the market after gaining approval by the FDA in November 2009 (Ueda et al. 1994; Nakajima et al. 1998; Bertino and Otterson 2011; Grant et al. 2010; Jain and Zain 2011). After the discovery of the above-mentioned inhibitors, the knowledge on HDACs and their biological functions has grown rapidly. New assays have been developed and crystal structures of several HDACs have been disclosed after the structure of Aquifex aeolicus HDAC homologue HDLP (Finnin et al. 1999). Thus, with the advances of the biological knowledge on HDACs, there is no surprise that the selectivity profile for TSA and SAHA was found to be typical for “pan-inhibitors”. The compounds were able to inhibit all enzymes with similar potency, though this belief has been recently challenged (Bradner et al. 2010).
Currently, the more advanced compounds can be classified either as pan-HDAC inhibitors, such as the majority of the hydroxamic acids, or as class I selective inhibitors, such as the pyrimidinhydroxamates CHR-3996 or JNJ-26481585 (Arts et al. 2009) as well as some non-hydroxamic acid derivatives like aminoanilide MGCD0103 (Garcia-Manero et al. 2008; Younes et al. 2011). In addition, there are compounds, like the thiol FK228, which can be described as a potent class I inhibitor and a moderate class II inhibitor. The clinical studies of these HDAC inhibitors have shown a similar toxicity profile, which includes gastrointestinal disturbances, fatigue and in several cases cardiotoxicity (Marsoni et al. 2008; Hymes 2010; Lynch et al. 2012). Toxicities were similar in pan-HDAC inhibitors, class I selective inhibitors and FK228. Thus, the clinical outcome has left open the question, whether selective inhibitors of HDACs would be less toxic and overall more advantageous compared to the more advanced HDAC inhibitors as anticancer agents.
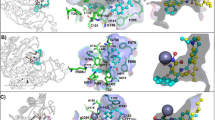
As mentioned above, the growing understanding of HDACs and their biological functions provided new tools or knowledge, which is crucial for the design of new selective HDAC inhibitors. Recombinant HDAC proteins as enzymes for the assays have become available. Furthermore, class IIa and IV specific substrates have been identified, which allow a precise profiling of compounds against biochemically active HDAC1-11 (Hauser and Jung 2009; Lahm et al. 2007; Bradner et al. 2010; Madsen and Olsen 2012). Interestingly, it has been recently found that HDAC3 showed in vitro decrotonylase activity (Madsen and Olsen 2012). Thus, the knowledge on HDACs continues to grow and it won’t be surprising to find that these enzymes may be possibly involved in further activities in cellular environments. In this context, it should be remembered that HDACs are components of multiprotein complexes (frequently having multiple enzymatic activities); and testing isolated recombinant HDACs in biochemical assays may not necessarily represent the best mode to reveal their cellular enzymatic functions. The crystal structures of human histone deacetylases HDAC2 (Cronin et al. 2009), HDAC3 (Watson et al. 2012), HDAC4 (Bottomley et al. 2008), HDAC7 (Schuetz et al. 2008) and HDAC8 (Somoza et al. 2004; Vannini et al. 2004) revealed the differences in the active enzymatic sites. These discoveries have provided new tools for the design of isoform selective compounds or compounds with a different selectivity profile than the older ones. These compounds may clarify if the toxicities seen in the more advanced HDAC inhibitors could be associated to one or more HDAC isoform(s).
At present, there are some examples, which exhibit isoform selectivity different from that of the above described HDAC inhibitors (Fig. 14). One example is the HDAC8 selective inhibitor PCI-34051 (Pharmacyclics) (Balasubramanian et al. 2009). PCI-34051 was found to be over 200-fold selective for HDAC8 compared to HDAC1-3, HDAC6 and HDAC10. The compound selectively induced apoptosis in cell lines derived from T cell lymphomas and leukaemias, but not in other haematopoietic or solid tumour cells. A further example is compound 55, presented as A8B4 (Tang et al. 2011). The compound showed IC50 values of 0.023 μM against HDAC8, 3.6 μM against HDAC2, and 15 μM against HDAC3/nCoR2. No further data are yet available. Other examples comprise some ortho substituted N-hydroxycinnamates, e.g., 56 (Huang et al. 2012). Compound 56 exhibited IC50 values of 27.2 ± 3.1 nM against HDAC8, 3 μM against HDACs 1 and 3 and ≥20 μM against HDACs 2, 4, 6, 10 and 11.
More examples can be found in the field of HDAC6 selective compounds (Fig. 15). The first example, tubacin, was already discovered in 2003, but its development was hampered by its poor drug properties (Haggarty et al. 2003). A homology model of HDAC6 revealed that the dimensions of the outer rim of the catalytic channel differ greatly between HDAC6 and HDAC1 (17.5 Å vs. 12.5 Å) (Butler et al. 2010). This structural diversity allowed designing compounds containing a cap moiety large and rigid enough to accommodate the rim region of HDAC6 but not of HDAC1. The resulting lead compound Tubastatin A was more than 1,000-fold selective versus all isoforms excluding HDAC8, where it had approximately 57-fold selectivity. The same laboratory disclosed recently the urea derivative Nexturastat A, which—compared to Tubastatin A—exhibited an improved inhibition of HDAC6, while maintaining the selectivity of approximately 600-fold relative to the inhibition of HDAC1 (Bergmann et al. 2012). The compound showed also a superior antiproliferative activity against B16 melanoma cells (GI50 = 14.3 ± 1.15 μM) than Tubastatin (40.5 ± 1.21 μM), but lower than LBH589 (0.150 ± 0.001 μM).
Also the HDAC6 selectivity has been found by exploring different hydroxamic acids linked to a macrocyclic structure (Auzzas et al. 2010). Representative compound (S)-9 inhibited HDAC6 with an IC50 value of 0.84 nM. The inhibitor was around 30 times more selective over HDAC1 and HDAC7 and even more over the other isoforms. HDAC6 selective inhibitors have been also described by scientists from Millennium Pharmaceuticals. For example, representative compound 57 (exemplified as I-126 in the in WO2012/054332) blocked HDAC6 activity by 95.6 % at 0.37 μM (England et al. 2012).
Currently, ACY-1215 has attracted great interest. The HDAC6 selective inhibitor initiated phase I/II clinical studies in July 2011 (www.clinicaltrials.gov/ct2/show/NCT01323751). The compound exhibited an IC50 value of 5 nM against HDAC6 and was around ten times less active against HDACs 1, 2 and 3. The activities against the class II enzymes HDAC 4, 5, 7 9 and against HDAC11 were in the micromolar range (Santo et al. 2012). Antiproliferative activities in Multiple Myeloma (MM) cell lines resulted with IC50 values ranging from 2 to 8 μM. A synergistic effect was found, when the compound was dosed in combination with bortezomib. These synergistic effects were seen both in vitro and in vivo in a human MM xenograft mouse model.
Inhibitors selective for other enzymes are even less explored. A series of diphenylmethylene hydroxamic acids as class IIa selective HDAC inhibitors have been described by scientists from Methylgene (Fig. 16) (Tessier et al. 2009). One of the described compounds, N-hydroxy-9H-xanthene-9-carboxamide (58), was somewhat more selective for HDAC7 with an IC50 of 0.05 μM. Other class II selective compounds have been disclosed in WO2012/103008 (CHDI Foundation Inc. 2012). Representative compound 59 had IC50 values of 0.02 μM against HDAC4 and 0.22 μM in human T cell lymphocyte Jurkat E6.1 cells. These structures are quite different from the hydroxamic acids described so far. In fact, the X-ray structure of HDAC7 (Schuetz et al. 2008) revealed that the catalytic domain of class IIa HDACs is significantly different from that of the previously reported class I and class IIb-like HDACs. The presence of a hydrophobic pocket in the enlarged active site of HDAC7 is absent in class I and IIb and allows to accommodate compounds with bulky moieties adjacent to the hydroxamic acid as Zn2+-chelating motif.
5 Computational Studies
In 1999, the histone deacetylase-like protein (HDLP) in complex with TSA and SAHA has been elucidated (Finnin et al. 1999). Later, various structure-based and ligand-based computational studies were carried out towards development of HDAC inhibitors (Wang 2009) and the results of various studies are as summarised below:
-
Very few isoforms selective HDAC inhibitors have been studied and rarely any compound having high selectivity has been found.
-
The zinc-binding groups are present in the majority of HDAC inhibitors. The ionisation states of the HDAC enzymes and the bound ligands need further study.
-
Future consideration is required towards the flexibility of the HDAC8’s pocket and surface mobility.
-
For rapid virtual screening of libraries, a more effective scoring function with predictive power is required.
Very few QSAR studies on HDAC inhibitors have been reported until now. The first QSAR model was developed using TSA- and SAHA-like hydroxamic acids (Lan-Hargest et al. 2002) suggesting the importance of shape and area of the molecules for biological activity (Wang et al. 2004). Further, on a data set of 124 compounds, a QSAR study has been reported. The model has shown role of van der Waals surface area and hydrophobicity towards biological activity (Xie et al. 2004). A series of substituted biaryl hydroxamates and mercaptoacetamides as HDAC inhibitors against pancreatic cancer cell growth with nanomolar potency have been designed and synthesised (Kozikowski et al. 2008) and QSAR derived equations have shown significant correlations between different HDAC isoforms (Wang 2009).
The binding mode of indole amide analogues in human HDAC1 catalytic core has been explored by Guo et al. (2005). Further, 3D-QSAR model using CoMFA and CoMSIA has been established for 29 substituted hydroxamic-based HDAC inhibitors with an indole amide residue at the terminus. A comparison between both studies concluded good correlation between the two analyses. On the same set of compounds, a multiple regression analysis by Bajpai et al. (2013) revealed that the inhibition of the histone deacetylase by this series of compounds might involve the dispersion interaction with the receptor where charge transfer between pairs of atoms might greatly help to polarise the molecule. The results obtained by this multiple regression analysis were in good agreement with those obtained by CoMFA and CoMSIA.
Juvale et al. (2006) performed CoMFA and CoMSIA studies on a data set of 40 hydroxamate analogues reported by Jung et al. (1999), Remiszewski et al. (2002) and Woo et al. (2002). The derived models have shown significance of steric and electronic fields along with lipophilicity as contributing parameters. A 3D-QSAR pharmacophore model has been developed using 30 known HDAC inhibitors (Chen et al. 2008). It concluded the essential ligand features, i.e. hydrogen-bond acceptor and hydrogen-bond donor features corresponding to the metal-binding function and coordination to the Zn(II) ion. Along with this, the hydrophobic/π-π stacking interactions between ligand and enzyme play a critical role for the inhibitory activity.
6 Development of HDAC Imaging Agents
As outlined above, a significant number of HDAC inhibitors have entered various clinical phases. In vivo imaging of HDACs and their inhibition is important and hence development of appropriate imaging agents has become a major challenge.
The conventional anatomic imaging modalities used to monitor clinical response can only help to identify tumour size or rudimentary physiologic changes and both of them occur relatively late after treatment onset. The ex vivo techniques can be utilised to analyse readily accessible tissue specimens. Thus, imaging probes are needed to investigate the efficiency of existing novel HDAC inhibitors with the hope that they may provide basic scientific insights which could lead to novel clinical applications.
Very few compounds for in vivo HDAC imaging have been reported. Some examples include 6-([18F]fluoroacetamide)-1-hexanoicanilide (FAHA) (60) (Reid et al. 2009) and [11C]MS-275, a carbon-11-labeled version of the benzamide class HDAC inhibitor MS-275 (entinostat) (61) for cerebral imaging (Hooker et al. 2010). These efforts were terminated because the short half-life of carbon-11- (~20 min) compared to fluorine-18 (~110 min) presents technical challenges and limits potential applications. Ronen and co-workers studied Boc-lysine trifluoroacetic acid 6 (BLT) as a substrate for HDACs using 19F magnetic resonance spectroscopy (Sankaranarayanapillai et al. 2008). Hendricks et al. reported development of 18F-suberoylanilide hydroxamic acid (18F-SAHA) (62), a close analogue of the clinically relevant SAHA. This compound results to be one of the first 18F-position emission tomography (PET) HDAC imaging agent (Hendricks et al. 2011).



7 Conclusion
Hydroxamic acids demonstrated to be very potent inhibitors of several enzymes, such as matrix metalloproteinases, carbonic anhydrases, lipoxygenases, ureases and many others (Muri et al. 2002; Gupta and Sharma 2013). However, hydroxamic acid derivatives are often considered as poor drugs and are down prioritised in drug discovery programmes despite their good in vitro potency, the reasons being their poor physicochemical and ADME properties. Drug discovery programmes related to hydroxamic acids as HDAC inhibitors faced the same issues; nevertheless, the research efforts in this field have been appreciable and several successes achieved such as discovery of acetohydroxamic acid (Marwick 1983), adrafinil (Siwak et al. 2003) and SAHA (Grant et al. 2007). SAHA was considerably efficacious in different haematological malignancies, even though having just medium potency and being cleared rapidly (Kelly et al. 2003; Marks and Breslow 2007), while the clinical outcome in other cancers remained much more uncertain and often rather limited (Graham et al. 2009; Mercurio et al. 2010). However, the notable in vitro and in vivo data of these first inhibitors made HDACs an attractive target for drug discovery programmes. Intensive research efforts resulted in a remarkable number of clinical candidates—over ten hydroxamic acid derivatives—for tumours and some other diseases. This chapter presents a case history of the drug discovery for the treatment of cancers based on HDAC inhibition by hydroxamic acids with a particular attention for the PK properties of the drugs discovered. Extensive in vitro ADME assays, for example by studying the stability of the inhibitors in hepatocytes, or PK cassette experiments as well as computational tools for simulating the ADME properties have been proved fundamental for overcoming the poor drug properties.
The toxicity profile of SAHA and of other more “mature” HDAC inhibitors are not strictly related to the hydroxamic acid groups, but can also be related to 2-aminoanilides, such as MGCD0103, or to thiols such as FK228. To date, a major topic in the HDAC inhibitory field is the development of selective HDAC inhibitors with a different selectivity profile than those already in advanced clinical studies. These compounds may clarify if the toxicities observed in the more advanced HDAC inhibitors might be associated with one or several HDACs. For future drug discovery programmes based on hydroxamic acid derivatives, the experiences gained from these preclinical and clinical studies may be of great value.
Abbreviations
- AUC:
-
Area under curve
- B16:
-
Murine melanoma cell line
- Caco-2:
-
Human epithelial colorectal adenocarcinoma cells
- COLO205:
-
Human colon cancer cell line
- CTCL:
-
Cutaneous T cell lymphoma
- F :
-
Dose-corrected area under curve (AUC) non-intravenous divided by AUC intravenous
- HAT:
-
Histone acetyltransferase
- HCT116:
-
Human colon cancer cell line
- HDAC:
-
Histone deacetylase
- HDLP:
-
Histone deacetylase-like protein
- ip:
-
Intraperitoneal
- iv:
-
Intravenous
- K562:
-
Chronic myelogenous leukaemia cell line
- MM:
-
Multiple myeloma
- qd:
-
Every day
- PK:
-
Pharmacokinetic
- t 1/2 :
-
Half-life
- T/C:
-
Ratio mean relative tumour volume of the treated tumours/mean relative volume of control group
- V ss :
-
Steady-state volume of distribution
References
Allfrey VG, Faulkner R, Mirsky AE (1964) Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci U S A 51:786–794
Allfrey VG, Mirsky AE (1964) Structural modifications of histones and their possible role in the regulation of RNA synthesis. Science 144:559
Andreeff M, Stone R, Michaeli J, Young CW, Tong WP, Sogoloff H, Ervin T, Kufe D, Rifkind RA, Marks PA (1992) Hexamethylene bisacetamide in myelodysplastic syndrome and acute myelogenous leukemia: a phase II clinical trial with a differentiation-inducing agent. Blood 80:2604–2609
Arts J, King P, Marien A, Floren W, Belien A, Janssen L, Pilatte I, Roux B, Decrane L, Gilissen R, Hickson I, Vreys V, Cox E, Bol K, Talloen W, Goris I, Andries L, Du Jardin M, Janicot M, Page M, van Emelen K, Angibaud P (2009) JNJ-26481585, a novel “second-generation” oral histone deacetylase inhibitor, shows broad-spectrum preclinical antitumoral activity. Clin Cancer Res 15:6841–6851
Atadja P (2009) Development of the pan-DAC inhibitor panobinostat (LBH589): successes and challenges. Cancer Lett 280:233–241
Auzzas L, Larsson A, Matera R, Baraldi A, Deschenes-Simard B, Giannini G, Cabri W, Battistuzzi G, Gallo G, Ciacci A, Vesci L, Pisano C, Hanessian S (2010) Non-natural macrocyclic inhibitors of histone deacetylases: design, synthesis, and activity. J Med Chem 53:8387–8399
Bai LY, Omar HA, Chiu CF, Chi ZP, Hu JL, Weng JR (2011) Antitumor effects of (S)-HDAC42, a phenylbutyrate-derived histone deacetylase inhibitor, in multiple myeloma cells. Cancer Chemother Pharmacol 68:489–496
Bajpai A, Agarwal N, Srivastava V, Mishra A, Gupta SP (2013) A comparative 2D QSAR study on a series of hydroxamic acid-based histone deacetylase inhibitors vis-à-vis comparative molecular field analysis and comparative molecular similarity indices analysis. Personal Communication.
Balasubramanian S, Steggerda S, Sirisawad M, Schreeder M, Doiron L, Buggy JJ (2009) The histone deacetylase-8 (HDAC8) selective inhibitor PCI-34051 decreases interleukin-1 beta secretion in vitro and reduces inflammation in vivo. In: 50th ASH annual meeting and exposition, San Francisco, American society of Hematology, abstr 2581
Banerji U, van Doorn L, Papadatos-Pastos D, Debnam P, Tall M, Toal M, Hooftman L, Verweij J, Eskens F (2010) A phase I pharmacokinetic (PK) and pharmacodynamic (PD) study of CHR-3996, a class 1 selective histone deacetylase inhibitor (HDACi), in patients with advanced solid tumors. J Clin Oncol 28(Suppl):2552
Banerji U, van Doorn L, Papadatos-Pastos D, Kristeleit R, Debnam P, Tall M, Stewart A, Raynaud F, Garrett MD, Toal M, Hooftman L, De Bono JS, Verweij J, Eskens FA (2012) A phase I pharmacokinetic and pharmacodynamic study of CHR-3996, an oral class I selective histone deacetylase inhibitor in refractory solid tumors. Clin Cancer Res 18:2687–2694
Bergman JA, Woan K, Perez-Villarroel P, Villagra A, Sotomayor EM, Kozikowski AP (2012) Selective histone deacetylase 6 inhibitors bearing substituted urea linkers inhibit melanoma cell growth. J Med Chem 55:9891–9899
Bertino EM, Otterson GA (2011) Romidepsin: a novel histone deacetylase inhibitor for cancer. Expert Opin Investig Drugs 20:1151–1158
Bolden JE, Peart MJ, Johnstone RW (2006) Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov 5:769–784
Bottomley MJ, Lo Surdo P, Di Giovine P, Cirillo A, Scarpelli R, Ferrigno F, Jones P, Neddermann P, De Francesco R, Steinkuhler C, Gallinari P, Carfi A (2008) Structural and functional analysis of the human HDAC4 catalytic domain reveals a regulatory structural zinc-binding domain. J Biol Chem 283:26694–26704
Bradner JE, West N, Grachan ML, Greenberg EF, Haggarty SJ, Warnow T, Mazitschek R (2010) Chemical phylogenetics of histone deacetylases. Nat Chem Biol 6:238–243
Breslow R, Jursic B, Yan ZF, Friedman E, Leng L, Ngo L, Rifkind RA, Marks PA (1991) Potent cytodifferentiating agents related to hexamethylenebisacetamide. Proc Natl Acad Sci U S A 88:5542–5546
Breslow R, Marks PA, Rifkind RA, Jursic B (1993) Novel potent inducers of terminal differentiation and methods of use thereof. PTC Int Appl WO199307148
Brown DT (2001) Histone variants: are they functionally heterogeneous? Genome Biol 2: REVIEWS0006
Brunetto AT, Ang JE, Lal R, Olmos D, Frentzas S, Mais A, Hauns B, Mollenhauer M, Lahu G, de Bono JS (2009) A first-in-human phase I study of 4SC-201, an oral histone deacetylase (HDAC) inhibitor, in patients with advanced solid tumors. J Clin Oncol 27:15s (abstr 3530)
Buchwald M, Kramer OH, Heinzel T (2009) HDACi–targets beyond chromatin. Cancer Lett 280:160–167
Buggy JJ, Cao ZA, Bass KE, Verner E, Balasubramanian S, Liu L, Schultz BE, Young PR, Dalrymple SA (2006) CRA-024781: a novel synthetic inhibitor of histone deacetylase enzymes with antitumor activity in vitro and in vivo. Mol Cancer Ther 5:1309–1317
Butler KV, Kalin J, Brochier C, Vistoli G, Langley B, Kozikowski AP (2010) Rational design and simple chemistry yield a superior, neuroprotective HDAC6 inhibitor, tubastatin A. J Am Chem Soc 132:10842–10846
Cai X, Zhai HX, Wang J, Forrester J, Qu H, Yin L, Lai CJ, Bao R, Qian C (2010) Discovery of 7-(4-(3-ethynylphenylamino)-7-methoxyquinazolin-6-yloxy)-N-hydroxyheptanam ide (CUDc-101) as a potent multi-acting HDAC, EGFR, and HER2 inhibitor for the treatment of cancer. J Med Chem 53:2000–2009
Cheng H, Jones W, Wei X, Liu Z, Wang D, Kulp S, Chen C-S, Covey JKC (2006a) Preclinical pharmacokinetics studies of R- and S- enantiomers of the histone deacetylase inhibitor, HDAC-42 (NSC 731438), in the rat. Proc Am Assoc Cancer Res 47 (abstr 686)
Cheng H, Liu Z, Kulp SK, Chen C-S, Covey JM, Chan KK (2006b) Preclinical pharmacokinetic studies with s-HDAC-42 (NSC 736012), an inhibitor of histone deacetylase, by LC-MS/MS Proc Amer Assoc Cancer Res 47 (abstr 3091)
Chen YD, Jiang YJ, Zhou JW, Yu QS, You QD (2008) Identification of ligand features essential for HDACs inhibitors by pharmacophore modeling. J Mol Graph Model 26:1160–1168
Choudhary C, Kumar C, Gnad F, Nielsen ML, Rehman M, Walther TC, Olsen JV, Mann M (2009) Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325:834–840
Costantino L, Barlocco D (2006) Privileged structures as leads in medicinal chemistry. Curr Med Chem 13:65–85
Cronin CN, Hilgers MT, Knuth MW, Navre ME, Sang C, Skene RJ, Tari LW, Wilson KP, Wittmer D, Zou H (2009) Crystallization of histone deacetylase 2. US Patent 7507522
Curtin ML, Garland RB, Heyman HR, Frey RR, Michaelides MR, Li J, Pease LJ, Glaser KB, Marcotte PA, Davidsen SK (2002) Succinimide hydroxamic acids as potent inhibitors of histone deacetylase (HDAC). Bioorg Med Chem Lett 12:2919–2923
Curtin M, Glaser K (2003) Histone deacetylase inhibitors: the abbott experience. Curr Med Chem 10:2373–2392
Dai Y, Guo Y, Curtin ML, Li J, Pease LJ, Guo J, Marcotte PA, Glaser KB, Davidsen SK, Michaelides MR (2003) A novel series of histone deacetylase inhibitors incorporating hetero aromatic ring systems as connection units. Bioorg Med Chem Lett 13:3817–3820
Davey CA, Sargent DF, Luger K, Maeder AW, Richmond TJ (2002) Solvent mediated interactions in the structure of the nucleosome core particle at 1.9 a resolution. J Mol Biol 319:1097–1113
Davies B, Morris T (1993) Physiological parameters in laboratory animals and humans. Pharm Res 10:1093–1095
DeSimone RW, Currie KS, Mitchell SA, Darrow JW, Pippin DA (2004) Privileged structures: applications in drug discovery. Comb Chem High Throughput Screen 7:473–494
Donald A, Belfield A, Day F, Patel S, Clark V, Needham L, Owen J, Bone E, Brotherton D, Bawden L, Rowlands M, Wibata J, Stimson L, Raynaud F, Aherne W, Moffat D (2010) The discovery and anti-tumor activity CHR-3996—a novel, orally available inhibitor of class 1 histone deacetylases. Paper presented at the EFMC-ISMC 2010 XXIst international symposium on medicinal chemistry, Brussels
Elaut G, Rogiers V, Vanhaecke T (2007) The pharmaceutical potential of histone deacetylase inhibitors. Curr Pharm Des 13:2584–2620
Elaut G, Torok G, Vinken M, Laus G, Papeleu P, Tourwe D, Rogiers V (2002) Major phase I biotransformation pathways of trichostatin a in rat hepatocytes and in rat and human liver microsomes. Drug Metab Dispos 30:1320–1328
Elliott JM, Selnick HG, Claremon DA, Baldwin JJ, Buhrow SA, Butcher JW, Habecker CN, King SW, Lynch JJ Jr, Phillips BT et al (1992) 4-Oxospiro[benzopyran-2,4′-piperidines] as class III antiarrhythmic agents. Pharmacological studies on 3,4-dihydro-1′-[2-(benzofurazan-5-yl)-ethyl]-6-methanesulfonamidospiro[(2H)-1-benzopyran-2,4′-piperidin]-4-one(L-691,121). J Med Chem 35:3973–3976
England D, Gigstad KM, Gould AE, Ma L, Xu H (2012) Substituted Hydroxamic Acids and Uses Therof. WO2012/054332
Federico M, Bagella L (2011) Histone deacetylase inhibitors in the treatment of hematological malignancies and solid tumors. J Biomed Biotechnol 2011:475–641
Ficner R (2009) Novel structural insights into class I and II histone deacetylases. Curr Top Med Chem 9:235–240
Finnin MS, Donigian JR, Cohen A, Richon VM, Rifkind RA, Marks PA, Breslow R, Pavletich NP (1999) Structures of a histone deacetylase homologue bound to the TSA and SAHA inhibitors. Nature 401:188–193
Flipo M, Charton J, Hocine A, Dassonneville S, Deprez B, Deprez-Poulain R (2009) Hydroxamates: relationships between structure and plasma stability. J Med Chem 52:6790–6802
Friend C, Scher W, Holland JG, Sato T (1971) Hemoglobin synthesis in murine virus-induced leukemic cells in vitro: stimulation of erythroid differentiation by dimethyl sulfoxide. Proc Natl Acad Sci U S A 68:378–382
Garcia-Manero G, Assouline S, Cortes J, Estrov Z, Kantarjian H, Yang H, Newsome WM, Miller WH Jr, Rousseau C, Kalita A, Bonfils C, Dubay M, Patterson TA, Li Z, Besterman JM, Reid G, Laille E, Martell RE, Minden M (2008) Phase 1 study of the oral isotype specific histone deacetylase inhibitor MGCD0103 in leukemia. Blood 112:981–989
Glozak MA, Sengupta N, Zhang X, Seto E (2005) Acetylation and deacetylation of non-histone proteins. Gene 363:15–23
Graham JS, Kaye SB, Brown R (2009) The promises and pitfalls of epigenetic therapies in solid tumours. Eur J Cancer 45:1129–1136
Grant C, Rahman F, Piekarz R, Peer C, Frye R, Robey RW, Gardner ER, Figg WD, Bates SE (2010) Romidepsin: a new therapy for cutaneous T-cell lymphoma and a potential therapy for solid tumors. Expert Rev Anticancer Ther 10:997–1008
Grant PA, Berger SL (1999) Histone acetyltransferase complexes. Semin Cell Dev Biol 10:169–177
Grant S, Easley C, Kirkpatrick P (2007) Vorinostat. Nat Rev Drug Discov 6:21–22
Gray SG, Ekstrom TJ (2001) The human histone deacetylase family. Exp Cell Res 262:75–83
Grignani F, De Matteis S, Nervi C, Tomassoni L, Gelmetti V, Cioce M, Fanelli M, Ruthardt M, Ferrara FF, Zamir I, Seiser C, Lazar MA, Minucci S, Pelicci PG (1998) Fusion proteins of the retinoic acid receptor-alpha recruit histone deacetylase in promyelocytic leukaemia. Nature 391:815–818
Grozinger CM, Hassig CA, Schreiber SL (1999) Three proteins define a class of human histone deacetylases related to yeast Hda1p. Proc Natl Acad Sci U S A 96:4868–4873
Guo Y, Xiao J, Guo Z, Chu F, Cheng Y, Wu S (2005) Exploration of a binding mode of indole amide analogues as potent histone deacetylase inhibitors and 3D-QSAR analyses. Bioorg Med Chem 13:5424–5434
Gupta SP, Sharma A (2013) Chemistry of hydroxamic acids (Chapter 1 of this book)
Haggarty SJ, Koeller KM, Wong JC, Grozinger CM, Schreiber SL (2003) Domain-selective small-molecule inhibitor of histone deacetylase 6 (HDAC6)-mediated tubulin deacetylation. Proc Natl Acad Sci U S A 100:4389–4394
Hauser AT, Jung M (2009) Assays for histone deacetylases. Curr Top Med Chem 9:227–234
Hendricks JA, Keliher EJ, Marinelli B, Reiner T, Weissleder R, Mazitschek R (2011) In vivo PET imaging of histone deacetylases by 18F-Suberoylanilide hydroxamic Acid (18F-SAHA). J Med Chem 54:5576–5582
Henikoff S, Furuyama T, Ahmad K (2004) Histone variants, nucleosome assembly and epigenetic inheritance. Trends Genet 20:320–326
Hooker JM, Kim SW, Alexoff D, Xu Y, Shea C, Reid A, Volkow N, Fowler JS (2010) Histone deacetylase inhibitor MS-275 exhibits poor brain penetration: pharmacokinetic studies of [11C]MS-275 using positron emission tomography. ACS Chem Neurosci 1:65–73
Horn PJ, Peterson CL (2002) Molecular biology. Chromatin higher order folding—wrapping up transcription. Science 297:1824–1827
Huang WJ, Wang YC, Chao SW, Yang CY, Chen LC, Lin MH, Hou WC, Chen MY, Lee TL, Yang P, Chang CI (2012) Synthesis and biological evaluation of ortho-Aryl N-hydroxycinnamides as potent histone deacetylase (HDAC) 8 isoform-selective inhibitors. Chem Med Chem 7:1815–1824
Hubbert C, Guardiola A, Shao R, Kawaguchi Y, Ito A, Nixon A, Yoshida M, Wang XF, Yao TP (2002) HDAC6 is a microtubule-associated deacetylase. Nature 417:455–458
Huber K, Superti-Furga G (2011) After the grape rush: sirtuins as epigenetic drug targets in neurodegenerative disorders. Bioorg Med Chem 19:3616–3624
Hwang JJ, Kim YS, Kim T, Kim MJ, Jeong IG, Lee JH, Choi J, Jang S, Ro S, Kim CS (2012) A novel histone deacetylase inhibitor, CG200745, potentiates anticancer effect of docetaxel in prostate cancer via decreasing Mcl-1 and Bcl-(XL). Invest New Drugs 30:1434–1442
Hymes KB (2010) The role of histone deacetylase inhibitors in the treatment of patients with cutaneous T-cell lymphoma. Clin Lymphoma Myeloma Leuk 10:98–109
Itazaki H, Nagashima K, Sugita K, Yoshida H, Kawamura Y, Yasuda Y, Matsumoto K, Ishii K, Uotani N, Nakai H et al (1990) Isolation and structural elucidation of new cyclotetrapeptides, trapoxins A and B, having detransformation activities as antitumor agents. J Antibiot (Tokyo) 43:1524–1532
Jain S, Zain J (2011) Romidepsin in the treatment of cutaneous T-cell lymphoma. J Blood Med 2:37–47
Jayaraman R, Pilla Reddy V, Pasha MK, Wang H, Sangthongpitag K, Yeo P, Hu CY, Wu X, Xin L, Goh E, New LS, Ethirajulu K (2011) Preclinical metabolism and disposition of SB939 (Pracinostat), an orally active histone deacetylase inhibitor, and prediction of human pharmacokinetics. Drug Metab Dispos 39:2219–2232
Jung M, Brosch G, Kolle D, Scherf H, Gerhauser C, Loidl P (1999) Amide analogues of trichostatin a as inhibitors of histone deacetylase and inducers of terminal cell differentiation. J Med Chem 42:4669–4679
Jung M (2001) Inhibitors of histone deacetylase as new anticancer agents. Curr Med Chem 8:1501–1511
Juvale DC, Kulkarni VV, Deokar HS, Wagh NK, Padhye SB, Kulkarni VM (2006) 3D-QSAR of histone deacetylase inhibitors: hydroxamate analogues. Org Biomol Chem 15:2858–2868
Kelly WK, O’Connor OA, Krug LM, Chiao JH, Heaney M, Curley T, MacGregore-Cortelli B, Tong W, Secrist JP, Schwartz L, Richardson S, Chu E, Olgac S, Marks PA, Scher H, Richon VM (2005) Phase I study of an oral histone deacetylase inhibitor, suberoylanilide hydroxamic acid, in patients with advanced cancer. J Clin Oncol 23:3923–3931
Kelly WK, Richon VM, O’Connor O, Curley T, MacGregor-Curtelli B, Tong W, Klang M, Schwartz L, Richardson S, Rosa E, Drobnjak M, Cordon-Cordo C, Chiao JH, Rifkind R, Marks PA, Scher H (2003) Phase I clinical trial of histone deacetylase inhibitor: suberoylanilide hydroxamic acid administered intravenously. Clin Cancer Res 9:3578–3588
Kijima M, Yoshida M, Sugita K, Horinouchi S, Beppu T (1993) Trapoxin, an antitumor cyclic tetrapeptide, is an irreversible inhibitor of mammalian histone deacetylase. J Biol Chem 268:22429–22435
Kim SC, Sprung R, Chen Y, Xu Y, Ball H, Pei J, Cheng T, Kho Y, Xiao H, Xiao L, Grishin NV, White M, Yang XJ, Zhao Y (2006) Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol Cell 23:607–618
Kleff S, Andrulis ED, Anderson CW, Sternglanz R (1995) Identification of a gene encoding a yeast histone H4 acetyltransferase. J Biol Chem 270:24674–24677
Klutchko S, Blankley CJ, Fleming RW, Hinkley JM, Werner AE, Nordin I, Holmes A, Hoefle ML, Cohen DM, Essenburg AD, Kaplan HR (1986) Synthesis of novel angiotensin converting enzyme inhibitor quinapril and related compounds. A divergence of structure-activity relationships for non-sulfhydryl types. J Med Chem 29:1953–1961
Kouzarides T (2000) Acetylation: a regulatory modification to rival phosphorylation? EMBO J 19:1176–1179
Kozikowski AP, Chen Y, Gaysin AM, Savoy DN, Billadeau DD, Kim KH (2008) Chemistry, biology, and QSAR studies of substituted biaryl hydroxamates and mercaptoacetamides as HDAC inhibitors-nanomolar-potency inhibitors of pancreatic cancer cell growth. ChemMedChem 3:487–501
Kulp SK, Chen CS, Wang DS, Chen CY (2006) Antitumor effects of a novel phenylbutyrate-based histone deacetylase inhibitor, (S)-HDAC-42, in prostate cancer. Clin Cancer Res 12:5199–5206
L’Hernault SW, Rosenbaum JL (1985a) Chlamydomonas alpha-tubulin is posttranslationally modified by acetylation on the epsilon-amino group of a lysine. Biochemistry 24:473–488
L’Hernault SW, Rosenbaum JL (1985b) Reversal of the posttranslational modification on chlamydomonas flagellar alpha-tubulin occurs during flagellar resorption. J Cell Biol 100:457–462
Lahm A, Paolini C, Pallaoro M, Nardi MC, Jones P, Neddermann P, Sambucini S, Bottomley MJ, Lo Surdo P, Carfi A, Koch U, De Francesco R, Steinkuhler C, Gallinari P (2007) Unraveling the hidden catalytic activity of vertebrate class IIa histone deacetylases. Proc Natl Acad Sci U S A 104:17335–17340
Lai CJ, Bao R, Tao X, Wang J, Atoyan R, Qu H, Wang DG, Yin L, Samson M, Forrester J, Zifcak B, Xu GX, DellaRocca S, Zhai HX, Cai X, Munger WE, Keegan M, Pepicelli CV, Qian C (2010) CUDC-101, a multitargeted inhibitor of histone deacetylase, epidermal growth factor receptor, and human epidermal growth factor receptor 2, exerts potent anticancer activity. Cancer Res 70:3647–3656
Lan-Hargest H-Y, Kaufman RJ, Wiech NL (2002) US Patent Appl US2002143196
Lin HY, Chen CS, Lin SP, Weng JR (2006) Targeting histone deacetylase in cancer therapy. Med Res Rev 26:397–413
Lin RJ, Nagy L, Inoue S, Shao W, Miller WH Jr, Evans RM (1998) Role of the histone deacetylase complex in acute promyelocytic leukaemia. Nature 391:811–814
Lu Q, Wang DS, Chen CS, Hu YD (2005) Structure-based optimization of phenylbutyrate-derived histone deacetylase inhibitors. J Med Chem 48:5530–5535
Luger K, Dechassa ML, Tremethick DJ (2012) New insights into nucleosome and chromatin structure: an ordered state or a disordered affair? Nat Rev Mol Cell Biol 13:436–447
Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 a resolution. Nature 389:251–260
Luger K, Richmond TJ (1998) The histone tails of the nucleosome. Curr Opin Genet Dev 8:140–146
Lynch DR Jr, Washam JB, Newby LK (2012) QT interval prolongation and torsades de pointes in a patient undergoing treatment with vorinostat: a case report and review of the literature. Cardiol J 19:434–438
Madsen AS, Olsen CA (2012) Profiling of substrates for zinc-dependent lysine deacylase enzymes: HDAC3 exhibits decrotonylase activity in vitro. Angew Chem Int Ed Engl 51:9083–9087
Mai A, Massa S, Pezzi R, Simeoni S, Rotili D, Nebbioso A, Scognamiglio A, Altucci L, Loidl P, Brosch G (2005a) Class II (IIa)-selective histone deacetylase inhibitors. 1. Synthesis and biological evaluation of novel (aryloxopropenyl)pyrrolyl hydroxyamides. J Med Chem 48:3344–3353
Mai A, Massa S, Rotili D, Cerbara I, Valente S, Pezzi R, Simeoni S, Ragno R (2005b) Histone deacetylation in epigenetics: an attractive target for anticancer therapy. Med Res Rev 25:261–309
Mai A, Massa S, Valente S, Simeoni S, Ragno R, Bottoni P, Scatena R, Brosch G (2006) Aroyl-pyrrolyl hydroxyamides: influence of pyrrole C4-phenylacetyl substitution on histone deacetylase inhibition. Chem Med Chem 1:225–237
Malik HS, Henikoff S (2003) Phylogenomics of the nucleosome. Nat Struct Biol 10:882–891
Mandl-Weber S, Meinel FG, Jankowsky R, Oduncu F, Schmidmaier R, Baumann P (2010) The novel inhibitor of histone deacetylase resminostat (RAS2410) inhibits proliferation and induces apoptosis in multiple myeloma (MM) cells. Br J Haematol 149:518–528
Mann BS, Johnson JR, Cohen MH, Justice R, Pazdur R (2007) FDA approval summary: vorinostat for treatment of advanced primary cutaneous T-cell lymphoma. Oncologist 12:1247–1252
Marks PA (2007) Discovery and development of SAHA as an anticancer agent. Oncogene 26:1351–1356
Marks PA, Breslow R (2007) Dimethyl sulfoxide to vorinostat: development of this histone deacetylase inhibitor as an anticancer drug. Nat Biotechnol 25:84–90
Marsoni S, Damia G, Camboni G (2008) A work in progress: the clinical development of histone deacetylase inhibitors. Epigenetics 3:164–171
Marwick C (1983) New drugs selectively inhibit kidney stone formation. JAMA 250:321–322
Massa S, Artico M, Corelli F, Mai A, Di Santo R, Cortes S, Marongiu ME, Pani A, La Colla P (1990) Synthesis and antimicrobial and cytotoxic activities of pyrrole-containing analogues of trichostatin A. J Med Chem 33:2845–2849
Massa S, Mai A, Sbardella G, Esposito M, Ragno R, Loidl P, Brosch G (2001) 3-(4-aroyl-1H-pyrrol-2-yl)-N-hydroxy-2-propenamides, a new class of synthetic histone deacetylase inhibitors. J Med Chem 44:2069–2072
Mercurio C, Minucci S, Pelicci PG (2010) Histone deacetylases and epigenetic therapies of hematological malignancies. Pharmacol Res 62:18–34
Miller TA, Witter DJ, Belvedere S (2003) Histone deacetylase inhibitors. J Med Chem 46:5097–5116
Moffat D, Patel S, Day F, Belfield A, Donald A, Rowlands M, Wibawa J, Brotherton D, Stimson L, Clark V, Owen J, Bawden L, Box G, Bone E, Mortenson P, Hardcastle A, van Meurs S, Eccles S, Raynaud F, Aherne W (2010) Discovery of 2-(6-{[(6-fluoroquinolin-2-yl)methyl]amino}bicyclo[3.1.0]hex-3-yl)-N-hydroxypyrim idine-5-carboxamide (CHR-3996), a class I selective orally active histone deacetylase inhibitor. J Med Chem 53:8663–8678
Muri EMF, Nieto MJ, Sindelar RD, Williamson JS (2002) Hydroxamic acids as pharmacological agents. Curr Med Chem 9:1631–1653
Nakajima H, Kim YB, Terano H, Yoshida M, Horinouchi S (1998) FR901228, a potent antitumor antibiotic, is a novel histone deacetylase inhibitor. Exp Cell Res 241:126–133
Neri P, Bahlis NJ, Lonial S (2012) Panobinostat for the treatment of multiple myeloma. Expert Opin Investig Drugs 21:733–747
Novotny-Diermayr V, Sangthongpitag K, Hu CY, Wu X, Sausgruber N, Yeo P, Greicius G, Pettersson S, Liang AL, Loh YK, Bonday Z, Goh KC, Hentze H, Hart S, Wang H, Ethirajulu K, Wood JM (2010) SB939, a novel potent and orally active histone deacetylase inhibitor with high tumor exposure and efficacy in mouse models of colorectal cancer. Mol Cancer Ther 9:642–652
Rambaldi A, Dellacasa CM, Finazzi G, Carobbio A, Ferrari ML, Guglielmelli P, Gattoni E, Salmoiraghi S, Finazzi MC, Di Tollo S, D’Urzo C, Vannucchi AM, Barosi G, Barbui T (2012) A pilot study of the histone-deacetylase inhibitor givinostat in patients with JAK2V617F positive chronic myeloproliferative neoplasms. Br J Haematol 150:446–455
Reid AE, Hooker J, Shumay E, Logan J, Shea C, Kim SW, Collins S, Xu Y, Volkow N, Fowler JS (2009) Evaluation of 6-([(18)F]fluoroacetamido)-1-hexanoicanilide for PET imaging of histone deacetylase in the baboon brain. Nucl Med Biol 36:247–258
Remiszewski S, Sambucetti L, Atadja P, Bair K, Cornell W, Green M, Howell K, Jung M, Kwon P, Trogani N, Walker H (2002) Inhibitors of human histone deacetylase: synthesis and enzyme and cellular activity of straight chain hydroxamates. J Med Chem 45:753–757
Remiszewski SW, Sambucetti LC, Bair KW, Bontempo J, Cesarz D, Chandramouli N, Chen R, Cheung M, Cornell-Kennon S, Dean K, Diamantidis G, France D, Green MA, Howell KL, Kashi R, Kwon P, Lassota P, Martin MS, Mou Y, Perez LB, Sharma S, Smith T, Sorensen E, Taplin F, Trogani N, Versace R, Walker H, Weltchek-Engler S, Wood A, Wu A, Atadja P (2003) N-hydroxy-3-phenyl-2-propenamides as novel inhibitors of human histone deacetylase with in vivo antitumor activity: discovery of (2E)-N-hydroxy-3-[4-[[(2-hydroxyethyl)[2-(1H-indol-3-yl)ethyl]amino]methyl]phenyl]-2-propenamide (NVP-LAQ824). J Med Chem 46:4609–4624
Reuben RC, Khanna PL, Gazitt Y, Breslow R, Rifkind RA, Marks PA (1978) Inducers of erythroleukemic differentiation. Relationship of structure to activity among planar-polar compounds. J Biol Chem 253:4214–4218
Reuben RC, Wife RL, Breslow R, Rifkind RA, Marks PA (1976) A new group of potent inducers of differentiation in murine erythroleukemia cells. Proc Natl Acad Sci U S A 73:862–866
Richon VM, Emiliani S, Verdin E, Webb Y, Breslow R, Rifkind RA, Marks PA (1998) A class of hybrid polar inducers of transformed cell differentiation inhibits histone deacetylases. Proc Natl Acad Sci U S A 95:3003–3007
Richon VM, O’Brien JP (2002) Histone deacetylase inhibitors: a new class of potential therapeutic agents for cancer treatment. Clin Cancer Res 8:662–624
Richon VM, Webb Y, Merger R, Sheppard T, Jursic B, Ngo L, Civoli F, Breslow R, Rifkind RA, Marks PA (1996) Second generation hybrid polar compounds are potent inducers of transformed cell differentiation. Proc Natl Acad Sci U S A 93:5705–5708
Riggs MG, Whittaker RG, Neumann JR, Ingram VM (1977) n-Butyrate causes histone modification in HeLa and Friend erythroleukaemia cells. Nature 268:462–464
Sankaranarayanapillai M, Tong WP, Yuan Q, Bankson JA, Dafni H, Bornmann WG, Soghomonyan S, Pal A, Ramirez MS, Webb D, Kaluarachchi K, Gelovani JG, Ronen SM (2008) Monitoring histone deacetylase inhibition in vivo: noninvasive magnetic resonance spectroscopy method. Mol Imaging 7:92–100
Santo L, Hideshima T, Kung AL, Tseng JC, Tamang D, Yang M, Jarpe M, van Duzer JH, Mazitschek R, Ogier WC, Cirstea D, Rodig S, Eda H, Scullen T, Canavese M, Bradner J, Anderson KC, Jones SS, Raje N (2012) Preclinical activity, pharmacodynamic, and pharmacokinetic properties of a selective HDAC6 inhibitor, ACY-1215, in combination with bortezomib in multiple myeloma. Blood 119:2579–2589
Schuetz A, Min J, Allali-Hassani A, Schapira M, Shuen M, Loppnau P, Mazitschek R, Kwiatkowski NP, Lewis TA, Maglathin RL, McLean TH, Bochkarev A, Plotnikov AN, Vedadi M, Arrowsmith CH (2008) Human HDAC7 harbors a class IIa histone deacetylase-specific zinc binding motif and cryptic deacetylase activity. J Biol Chem 283:11355–11363
Selvi RB, Kundu TK (2009) Reversible acetylation of chromatin: implication in regulation of gene expression, disease and therapeutics. Biotechnol J 4:375–390
Shimizu T, Tolcher AW, LoRusso P, Papadopoulos K, Patnaik A, Smith L, Keegan M (2010) 364 The first-in-human, first-in-class study of CUDC-101, a multi-targeted inhibitor of HDAC, EGFR, and HER2: A Phase I study in patients with advanced cancer. Eur J Cancer Suppl 8:115
Shultz MD, Cao X, Chen CH, Cho YS, Davis NR, Eckman J, Fan J, Fekete A, Firestone B, Flynn J, Green J, Growney JD, Holmqvist M, Hsu M, Jansson D, Jiang L, Kwon P, Liu G, Lombardo F, Lu Q, Majumdar D, Meta C, Perez L, Pu M, Ramsey T, Remiszewski S, Skolnik S, Traebert M, Urban L, Uttamsingh V, Wang P, Whitebread, Whitehead L, Yan-Neale Y, Yao YM, Zhou L, Atadja P (2011) Optimization of the in vitro cardiac safety of hydroxamate-based histone deacetylase inhibitors. J Med Chem 54:4752–4772
Simon M, North JA, Shimko JC, Forties RA, Ferdinand MB, Manohar M, Zhang M, Fishel R, Ottesen JJ, Poirier MG (2011) Histone fold modifications control nucleosome unwrapping and disassembly. Proc Natl Acad Sci U S A 108:12711–12716
Singh BN, Zhang G, Hwa YL, Li J, Dowdy SC, Jiang SW (2010) Nonhistone protein acetylation as cancer therapy targets. Expert Rev Anticancer Ther 10:935–954
Siwak CT, Tapp PD, Milgram NW (2003) Adrafinil disrupts performance on a delayed nonmatching-to-position task in aged beagle dogs. Pharmacol Biochem Behav 76:161–168
Somoza JR, Skene RJ, Katz BA, Mol C, Ho JD, Jennings AJ, Luong C, Arvai A, Buggy JJ, Chi E, Tang J, Sang BC, Verner E, Wynands R, Leahy EM, Dougan DR, Snell G, Navre M, Knuth MW, Swanson RV, McRee DE, Tari LW (2004) Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure 12:1325–1334
Steele NL, Plumb JA, Vidal L, Tjornelund J, Knoblauch P, Rasmussen A, Ooi CE, Buhl-Jensen P, Brown R, Evans TR, DeBono JS (2008) A phase 1 pharmacokinetic and pharmacodynamic study of the histone deacetylase inhibitor belinostat in patients with advanced solid tumors. Clin Cancer Res 14:804–810
Steinman DH, Curtin ML, Garland RB, Davidsen SK, Heyman HR, Holms JH, Albert DH, Magoc TJ, Nagy IB, Marcotte PA, Li J, Morgan DW, Hutchins C, Summers JB (1998) The design, synthesis, and structure-activity relationships of a series of macrocyclic MMP inhibitors. Bioorg Med Chem Lett 8:2087–2092
Sternson SM, Wong JC, Grozinger CM, Schreiber SL (2001) Synthesis of 7200 small molecules based on a substructural analysis of the histone deacetylase inhibitors trichostatin and trapoxin. Org Lett 3:4239–4242
Tanaka M, Levy J, Terada M, Breslow R, Rifkind RA, Marks PA (1975) Induction of erythroid differentiation in murine virus infected eythroleukemia cells by highly polar compounds. Proc Natl Acad Sci U S A 72:1003–1036
Tang W, Luo T, Greenberg EF, Bradner JE, Schreiber SL (2011) Discovery of histone deacetylase 8 selective inhibitors. Bioorg Med Chem Lett 21:2601–2605
Taunton J, Hassig CA, Schreiber SL (1996) A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 272:408–411
Tessier P, Smil DV, Wahhab A, Leit S, Rahil J, Li Z, Deziel R, Besterman JM (2009) Diphenylmethylene hydroxamic acids as selective class IIa histone deacetylase inhibitors. Bioorg Med Chem Lett 19:5684–5688
Thaler F (2012) Current trends in the development of histone deacetylase inhibitors: a review of recent patent applications. Pharm Pat Analyst 1:75–90
Thaler F, Colombo A, Mai A, Amici R, Bigogno C, Boggio R, Cappa A, Carrara S, Cataudella T, Fusar F, Gianti E, Joppolo di Ventimiglia S, Moroni M, Munari D, Pain G, Regalia N, Sartori L, Vultaggio S, Dondio G, Gagliardi S, Minucci S, Mercurio C, Varasi M (2010a) Synthesis and biological evaluation of N-hydroxyphenylacrylamides and N-hydroxypyridin-2-ylacrylamides as novel histone deacetylase inhibitors. J Med Chem 53:822–829
Thaler F, Colombo A, Mai A, Bigogno C, Boggio R, Carrara S, Joppolo di Ventimiglia S, Munari D, Regalia N, Dondio G, Gagliardi S, Minucci S, Mercurio C, Varasi M (2010b) Synthesis and biological characterization of amidopropenyl-hydroxamates as HDAC inhibitors. ChemMedChem 5:1359–1372
Thaler F, Mai A, Colombo A, Bigogno C, Boggio R, Regalia N, Rozio MG, Vultaggio S, Gagliardi S, Minucci S, Mercurio C, Varasi M (2009) Synthesis and structure-activity relationships of phenyloxopropenyl- and amidopropenyl-hydroxamic acid derivatives as HDAC inhibitors. Paper presented at the abstract paper. XXIII Congresso Nazionale della Società Chimica Italiana, Sorrento
Thaler F, Varasi M, Carenzi G, Colombo A, Abate A, Bigogno C, Boggio R, Carrara S, Cataudella T, Dal Zuffo R, Reali V, Vultaggio S, Dondio G, Gagliardi S, Minucci S, Mercurio C (2012) Spiro[chromane-2,4′-piperidine]-based histone deacetylase inhibitors with improved in vivo activity. Chem Med Chem 7:709–721
Tong WG, Wei Y, Stevenson W, Kuang SQ, Fang Z, Zhang M, Arts J, Garcia-Manero G (2010) Preclinical antileukemia activity of JNJ-26481585, a potent second-generation histone deacetylase inhibitor. Leuk Res 34:221–228
Tsuji N, Kobayashi M, Nagashima K, Wakisaka Y, Koizumi K (1976) A new antifungal antibiotic, trichostatin. J Antibiot (Tokyo) 29:1–6
Ueda H, Nakajima H, Hori Y, Fujita T, Nishimura M, Goto T, Okuhara M (1994) FR901228, a novel antitumor bicyclic depsipeptide produced by chromobacterium violaceum No. 968. I. Taxonomy, fermentation, isolation, physico-chemical and biological properties, and antitumor activity. J Antibiot (Tokyo) 47:301–310
Vannini A, Volpari C, Filocamo G, Casavola EC, Brunetti M, Renzoni D, Chakravarty P, Paolini C, De Francesco R, Gallinari P, Steinkuhler C, Di Marco S (2004) Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proc Natl Acad Sci U S A 101:15064–15069
Vansteenkiste J, Van Cutsem E, Dumez H, Chen C, Ricker JL, Randolph SS, Schoffski P (2008) Early phase II trial of oral vorinostat in relapsed or refractory breast, colorectal, or non-small cell lung cancer. Invest New Drugs 26:483–488
Varasi M, Thaler F, Abate A, Bigogno C, Boggio R, Carenzi G, Cataudella T, Dal Zuffo R, Fulco MC, Rozio MG, Mai A, Dondio G, Minucci S, Mercurio C (2011) Discovery, synthesis, and pharmacological evaluation of spiropiperidine hydroxamic acid based derivatives as structurally novel histone deacetylase (HDAC) inhibitors. J Med Chem 54:3051–3064
Venkatesh PR, Goh E, Zeng P, New LS, Xin L, Pasha MK, Sangthongpitag K, Yeo P, Kantharaj E (2007) In vitro phase I cytochrome P450 metabolism, permeability and pharmacokinetics of SB639, a novel histone deacetylase inhibitor in preclinical species. Biol Pharm Bull 30:1021–1024
Verdin E, Dequiedt F, Kasler HG (2003) Class II histone deacetylases: versatile regulators. Trends Genet 19:286–293
Wada C, Frey R, Ji Z, Curtin M, Garland R, Li J, Pease L, Guo J, Glaser K, Marcotte P, Richardson P, Murphy S, Bouska J, Tapang P, Magoc T, Albert D, Davidsen S, Michaelides M (2003) Alpha-keto amides as inhibitors of histone deacetylase. Bioorg Med Chem Lett 13:3331–3335
Wang D (2009) Computational studies on the histone deacetylases and the design of selective histone deacetylase inhibitors. Curr Top Med Chem 9:241–256
Wang DF, Helquist P, Wiech NL, Wiest O (2005) Toward selective histone deacetylase inhibitor design: homology modeling, docking studies, and molecular dynamics simulations of human class I histone deacetylases. J Med Chem 48:6936–6947
Wang D-F, Wiest O, Helquist P, Lan-Hargestb H-Y, Wiech NL (2004) QSAR Studies of PC-3 cell line inhibition activity of TSA and SAHA-like hydroxamic acids. Bioorg Med Chem Lett 14:707–711
Wang H, Yu N, Chen D, Lee KC, Lye PL, Chang JW, Deng W, Ng MC, Lu T, Khoo ML, Poulsen A, Sangthongpitag K, Wu X, Hu C, Goh KC, Wang X, Fang L, Goh KL, Khng HH, Goh SK, Yeo P, Liu X, Bonday Z, Wood JM, Dymock BW, Kantharaj E, Sun ET (2011) Discovery of (2E)-3-{2-butyl-1-[2-(diethylamino)ethyl]-1H-benzimidazol-5-yl}-N-hydroxyacrylami de (SB939), an orally active histone deacetylase inhibitor with a superior preclinical profile. J Med Chem 54:4694–4720
Wang H, Yu N, Song H, Chen D, Zou Y, Deng W, Lye PL, Chang J, Ng M, Sun ET, Sangthongpitag K, Wang X, Wu X, Khng HH, Fang L, Goh SK, Ong WC, Bonday Z, Stunkel W, Poulsen A, Entzeroth M (2009) N-Hydroxy-1,2-disubstituted-1H-benzimidazol-5-yl acrylamides as novel histone deacetylase inhibitors: design, synthesis, SAR studies, and in vivo antitumor activity. Bioorg Med Chem Lett 19:1403–1408
Watson PJ, Fairall L, Santos GM, Schwabe JW (2012) Structure of HDAC3 bound to co-repressor and inositol tetraphosphate. Nature 481:335–340
Woo SH, Frechette S, Khalil EA, Bouchain G, Vaisburg A, Bernstein N, Moradei O, Leit S, Allan M, Fournel M, Trachy-Bourget M-C, Li Z, Besterman JM, Delorme D (2002) Structurally simple trichostatin A-like staight chain hydroxamates as potent histone deacetylase inhibitors. J Med Chem 45:2877–2885
Yang XJ, Gregoire S (2005) Class II histone deacetylases: from sequence to function, regulation, and clinical implication. Mol Cell Biol 25:2873–2884
Yang XJ, Seto E (2008) Lysine acetylation: codified crosstalk with other posttranslational modifications. Mol Cell 31:449–461
Yao YL, Yang WM (2011) Beyond histone and deacetylase: an overview of cytoplasmic histone deacetylases and their nonhistone substrates. J Biomed Biotechnol 2011:146493
Yong W, Goh B, Toh H, Soo R, Diermayr V, Goh A, Ethirajulu K, Lee S, Seah E, Zhu J (2009) Phase I study of SB939 three times weekly for 3 weeks every 4 weeks in patients with advanced solid malignancies. J Clin Oncol 27 (abstr 2560)
Yoshida M, Hoshikawa Y, Koseki K, Mori K, Beppu T (1990a) Structural specificity for biological activity of trichostatin A, a specific inhibitor of mammalian cell cycle with potent differentiation-inducing activity in Friend leukemia cells. J Antibiot (Tokyo) 43:1101–1106
Yoshida M, Kijima M, Akita M, Beppu T (1990b) Potent and specific inhibition of mammalian histone deacetylase both in vivo and in vitro by trichostatin A. J Biol Chem 265:17174–17179
Yoshida M, Nomura S, Beppu T (1987) Effects of trichostatins on differentiation of murine erythroleukemia cells. Cancer Res 47:3688–3691
Younes A, Oki Y, Bociek RG, Kuruvilla J, Fanale M, Neelapu S, Copeland A, Buglio D, Galal A, Besterman J, Li Z, Drouin M, Patterson T, Ward MR, Paulus JK, Ji Y, Medeiros LJ, Martell RE (2011) Mocetinostat for relapsed classical Hodgkin’s lymphoma: an open-label, single-arm, phase 2 trial. Lancet Oncol 12:1222–1228
Xie A, Liao C, Li Z, Ning Z, Hu W, Lu X, Shi L, Zhou J (2004) Quantitative structure-activity relationship study of histone deacetylase inhibitors. Curr Med Chem Anticancer Agents 4:273–299
Zhang Y, Fang H, Feng J, Jia Y, Wang X, Xu W (2011a) Discovery of a tetrahydroisoquinoline-based hydroxamic acid derivative (ZYJ-34c) as histone deacetylase inhibitor with potent oral antitumor activities. J Med Chem 54:5532–5539
Zhang Y, Feng J, Liu C, Zhang L, Jiao J, Fang H, Su L, Zhang X, Zhang J, Li M, Wang B, Xu W (2010) Design, synthesis and preliminary activity assay of 1,2,3,4-tetrahydroisoquinoline-3-carboxylic acid derivatives as novel histone deacetylases (HDACs) inhibitors. Bioorg Med Chem 18:1761–1772
Zhang Y, Feng J, Jia Y, Wang X, Zhang L, Liu C, Fang H, Xu W (2011b) Development of tetrahydroisoquinoline-based hydroxamic acid serivatives: potent histone deacetylase inhibitors with marked in vitro and in vivo antitumor activities. J Med Chem 54:2823–2838
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Thaler, F., Patil, V.M., Gupta, S.P. (2013). Hydroxamic Acids as Histone Deacetylase Inhibitors. In: Gupta, S. (eds) Hydroxamic Acids. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-38111-9_5
Download citation
DOI: https://doi.org/10.1007/978-3-642-38111-9_5
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-38110-2
Online ISBN: 978-3-642-38111-9
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)