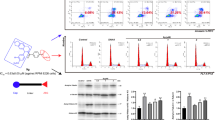
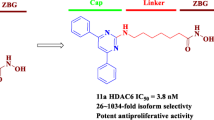
Abstract
In our search for novel small molecules targeting histone deacetylases, we have designed and synthesized a series of novel hydroxamic acids incorporating indole moiety as a cap group (3a–l). Biological evaluation showed that these hydroxamic acids potently inhibited HDAC2 with IC50 values in submicromolar range and up to tenfold (compound 3j) better than that of SAHA (also known as suberoylanilide hydroxamic acid). In four human cancer cell lines [SW620 (colon), PC-3 (prostate), AsPC-1 (pancreatic), NCI-H23 (lung)], the synthesized compounds that exhibited potent cytotoxicity with several compounds (3k, 3l) were found to be 12- to 77-fold more cytotoxic than SAHA. Docking experiments indicated that the compounds tightly bound to HDAC2 at the active binding site with binding affinities much higher than that of SAHA. Our present results demonstrate that these novel hydroxamic acids are potential for further development as anticancer agents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Anticancer drug discovery in the past was often based on random screening or empirical design. As a result, many of the currently used anticancer therapeutics are very toxic, non-specific, and prone to cellular resistance. In order to overcome these disadvantages, today’s anticancer drug design is increasingly based on a molecular target approach. With the advances in cancer molecular biology, dozens of important proteins or genes have been validated as promising targets for anticancer drug design, for example, protein kinases, farnesyl transferases, telomerases, and histone deacetylases, among many others (Nam and Parang 2003).
Histone deacetylases (EC 3.5.1.98, HDACs) constitute a group of enzymes which catalyze removal of the acetyl groups from lysine residues in the acetylated histones (Witt et al. 2009; De Ruijter et al. 2003). In mammalians, at least 18 different HDAC isoforms have been identified. Based on the relative sequence similarity, these isoforms can be categorized into four classes (Witt et al. 2009; De Ruijter et al. 2003; Zwergel et al. 2015). Relating to cancer biology, two classes including class I and class II of HDACs have been comprehensively investigated and demonstrated to be deeply involved in a number of cell-related processes (De Ruijter et al. 2003; Zwergel et al. 2015; West and Johnstone 2014). Especially class I, which has four members (HDAC1, 2, 3 and HDAC8) have been demonstrated to promote cellular proliferation. These HDAC isoforms (HDAC1, 2, and 3) and some isoforms of class II (HDAC4, 5) have also been shown to prevent cellular apoptosis and differentiation. On the other hand, several HDAC isoforms, including HDAC4, 6, 7, and 10, have been demonstrated to promote angiogenesis and cell migration. These two processes are known to be very important for cancer cell metastasis (Zwergel et al. 2015; West and Johnstone 2014). Inhibition of HDAC isoforms has been shown to cause a number of events which led to differentiation, apoptosis, and cell cycle arrest in different types of tumor cells (Zwergel et al. 2015; West and Johnstone 2014). In addition, selective inhibition of the growth of tumor cells as a result from HDAC inhibition has been demonstrated not only in vitro but also in a number of in vivo preclinical models and clinical settings (Hamm and Costa 2015; Glaser 2007). Therefore, design of compounds that appropriately target different HDAC isoforms has became very interesting approach in cancer drug development (Bolden et al. 2006). In the past decades, with the extensive efforts of medicinal chemists worldwide, dozens of HDAC inhibitors have been reported. Structurally, these inhibitors are diverse, from short-chain fatty acids (like butyric or valproic acid) to hydroxamic acids (like suberoylanilide hydroxamic acid, also known as SAHA), or benzamides (Dallavalle et al. 2009; Bracker et al. 2009; Iyer and Foss 2015; Valente and Mai 2014; Li et al. 2013; Ververis et al. 2013; Qiu et al. 2013). Up to 2016, at least 5 HDAC inhibitors have been approved for use in clinical settings. The first HDAC inhibitor approved by the U.S. FDA in October 2006 for the treatment of cutaneous T cell lymphoma (CTCL) was vorinostat (trade name, Zolinza®) (SAHA) (Fig. 1). The second HDAC inhibitor went to the market was romidepsin (trade name, Istodax®), which was approved by the U.S. FDA for the same indication in 2009. The third HDAC inhibitor (belinostat, PXD101) was approved in 2014 in the US for the use against peripheral T cell lymphoma. Meanwhile, panobinostat (LBH-589, trade name Farydak®) was approved by the U.S. FDA for the treatment of multiple myeloma in Feb 2015 (Cheng et al. 2015). In that same year, chidamide (Epidaza®) was approved by the Chinese FDA for relapsed or refractory peripheral T cell lymphoma (Malini 2015). In addition, a number of other HDAC inhibitors such as entinostat (MS-27-527), mocetinostat (MGCD0103), or givinostat (ITF2357) are currently under different phases of clinical trials for several types of cancer (Fig. 1).
In our research program to find novel hydroxamic acids as potential inhibitors of HDACs and anticancer agents, we have previously reported several series of heterocyclic analogs of SAHA, which incorporated benzothiazole, 5-aryl-1,3,4-thiadiazole, or 2-oxoindoline systems (Fig. 2) (Oanh et al. 2011; Tung et al. 2013; Nam et al. 2013 and 2014). These compounds were found to display very potent HDAC inhibitory activity as well as cytotoxicity. Some representative compounds from these series also exhibited significant antitumor activity in PC-3 prostate cancer cells’ xenografted mice model (Tung et al. 2013). Inspired by these results, we expanded our research into new series of hydroxamic acids which incorporated indole moiety as a cap group. The present paper reports the results we obtained from the synthesis, biological evaluation, and docking study on these novel indole-based hydroxamic acids.
Experimental
Chemistry
All reagents and solvents were purchased from Aldrich, Fluka Chemical Corp. (Milwaukee, WI, USA), or Merck unless noted otherwise. Solvents were used directly as purchased unless otherwise indicated. Thin-layer chromatography which was performed using Whatman® 250 μm Silica Gel GF Uniplates and visualized under UV light at 254 and 365 nm was used to check the progress of reactions and preliminary evaluation of compounds’ homogeneity. In all cases, the compounds achieved purity of 97% or above, as determined by HPLC. Melting points were measured using a Gallenkamp Melting Point Apparatus (LabMerchant, London, United Kingdom) and are uncorrected. Purification of compounds was carried out using crystallization methods and/or open silica gel column flash chromatography employing Merck silica gel 60 (240–400 mesh) as a stationary phase. Nuclear magnetic resonance spectra (1H NMR) were recorded on a Bruker 500 MHz spectrometer with DMSO-d 6 as solvent unless otherwise indicated. Tetramethylsilane was used as an internal standard. Chemical shifts are reported in parts per million (ppm), downfield from tetramethylsilane. Mass spectra with different ionization modes including electron ionization (EI) and Electrospray ionization (ESI) were recorded using PE Biosystems API2000 (Perkin Elmer, Palo Alto, CA, USA) and Mariner® (Azco Biotech, Inc. Oceanside, CA, USA) mass spectrometers, respectively.
The synthesis of a series of novel hydroxamic acids incorporating indole moiety (3a–l) was carried out as illustrated in Scheme 1. One indoline-incorporated hydroxamic acid (5) was synthesized according to Scheme 2. Details are described as follows.
General procedures for the synthesis of compounds 3a–l
Compounds 1a–l (2.00 mmol) were dissolved in DMF (abs) (3 mL). To the resulting solutions, NaH (120 mg, 5.00 mmol) was added. The reaction mixtures were stirred at 80 °C for 2 h and then KI (33.2 mg, 2.00 mmol) was added. After stirring for further 10 min, 0.20 mL of a solution of methyl bromoalkanoates (2.00 mmol) in DMF (abs) (0.2 mL) was dropped slowly into the mixtures. The reaction mixtures were stirred at room temperature for 12–24 h. Upon completion, the resultant mixtures were cooled, poured into ice-cold water, and acidified to pH ~ 6. Then, the mixtures were extracted by dichloromethane and dried over anhydrous sodium sulfate, filtered, and dried to give 2a–l, which were used for the next step without further purification.
A solution of each intermediate 2a–l (~2.00 mmol) was dissolved in a sufficient volume of methanol (10 mL). The mixture was cooled to −5 ÷ 0 °C. Then, hydroxylamine. HCl (1.40 g, 20.0 mmol) was added and stirred for 15 min, followed by dropwise addition of a solution of NaOH to pH ~ 12. The mixture was stirred until the reaction completed. The resultant mixture was poured into ice-cold water and neutralized to pH ~ 7 by dropwise addition of a solution of HCl 5%. The mixture was extracted by ethyl acetate and dried over anhydrous sodium sulfate. After evaporation under vacuum to give a black or umber viscous liquid (3a–l), the compounds 3a–l were purified by flash column chromatography (SiO2). Elution was carried out using a mobile phase consisting of DCM/MeOH (25/1).
N-Hydroxy-5-(1H-indol-1-yl)pentanamide (3a)
Viscous liquid, black; Yield: 52%. R f = 0.46 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3233 (NH), 3195 (OH), 3015 (C–H, arene), 2845 (CH, CH2), 1725, 1655 (C=O), 1612 (C=C), 1455 (C–N). ESI–MS (m/z): 230.9 [M − H]–. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.34 (1H, s, −OH); 8.67 (1H, s, −NH); 7.53 (1H, d, J = 7.5 Hz, H-4′); 7.45 (1H, d, J = 7.5 Hz, H-7′); 7.35 (1H, s, H-2′); 7.11 (1H, t, J = 6.5 Hz, H-6′); 7.00 (1H, t, J = 7 Hz, H-5′); 6.42 (1H, s, H-3′); 4.17 (2H, s, H-5); 1.97 (2H, s, H-2); 1.81–1.66 (2H, m, H-4); 1.46 (2H, s, H-3). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 168.82; 135.61; 128.54; 128.06; 120.88; 120.37; 118.77; 109.71; 100.35; 45.11; 31.83; 29.44; 22.54. Anal. Calcd. For C13H16N2O2 (232.27): C, 67.22; H, 6.94; N, 12.06. Found: C, 67.25; H, 6.97; N, 12.09.
5-(6-Chloro-1H-indol-1-yl)-N-hydroxypentanamide (3b)
Viscous liquid, black; Yield: 48%. R f = 0.42 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3445 (NH), 3185 (OH), 3045 (C–H, arene), 2855 (CH, CH2), 1715, 1650 (C=O), 1615 (C=C), 1455 (C–N). ESI–MS (m/z): 264.9 [M − H]–. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.36 (1H, s, –OH); 8.69 (1H, s, –NH); 7.61 (1H, s, H-7′); 7.53 (1H, d, J = 8.5 Hz, H-4′); 7.39 (1H, d, J = 3 Hz, H-2′); 7.01 (1H, q, J = 1.5, 8.5 Hz, H-5′); 6.45 (1H, d, J = 3 Hz, H-3′); 4.15 (2H, t, J = 7 Hz, H-5); 1.96 (2H, t, J = 7.5 Hz, H-2); 1.69 (2H, quintet, J = 7.5 Hz, H-4); 1.44 (2H, quintet, J = 7.5 Hz, H-3). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 168.83; 136.09; 129.70; 126.78; 125.90; 121.68; 119.15; 109.65; 100.79; 45.22; 31.83; 29.40; 22.49. Anal. Calcd. For C13H15ClN2O2 (266.72): C, 58.54; H, 5.67; N, 10.50. Found: C, 58.56; H, 5.65; N, 10.53.
5-(5-Chloro-1H-indol-1-yl)-N-hydroxypentanamide (3c)
Viscous liquid, black; Yield: 50%. R f = 0.41 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3435 (NH), 3180 (OH), 3035 (C–H, arene), 2850 (CH, CH2), 1720, 1645 (C=O), 1620 (C=C), 1450 (C–N). ESI–MS (m/z): 264.8 [M − H]–. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.35 (1H, s, –OH); 8.68 (1H, s, –NH); 7.58 (1H, d, J = 1.5 Hz, H-4′); 7.49 (1H, d, J = 8.5 Hz, H-7′); 7.42 (1H, d, J = 2.5 Hz, H-2′); 7.11 (1H, q, J = 1.5, 8.5 Hz, H-6′); 6.42 (1H, d, J = 3 Hz, H-3′); 4.15 (2H, t, J = 7 Hz, H-5); 1.95 (2H, t, J = 7.5 Hz, H-2); 1.69 (2H, quintet, J = 7.5 Hz, H-4); 1.43 (2H, quintet, J = 7.5 Hz, H-3). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 168.79; 134.16; 130.28; 129.16; 123.56; 120.82; 119.50; 111.33; 100.23; 45.32; 31.80; 29.41; 22.48. Anal. Calcd. For C13H15ClN2O2 (266.72): C, 58.54; H, 5.67; N, 10.50. Found: C, 58.55; H, 5.66; N, 10.52.
N-Hydroxy-5-(5-methoxy-1H-indol-1-yl)pentanamide (3d)
Viscous liquid, black; Yield: 48%. R f = 0.39 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3445 (NH), 3175 (OH), 3040 (C–H, arene), 2850 (CH, CH2), 1705, 1605 (C=O), 1615 (C=C), 1450 (C–N). ESI–MS (m/z): 260.9 [M − H]–. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.37 (1H, s, –OH); 8.70 (1H, s, –NH); 7.33 (1H, d, J = 9 Hz, H-7′); 7.28 (1H, d, J = 3 Hz, H-4′); 7.05 (1H, d, J = 2,5 Hz, H-2′); 6.76 (1H, q, J = 2.0, 8.5 Hz, H-6′); 6.33 (1H, d, J = 2.5 Hz, H-3′); 4.09 (2H, t, J = 7 Hz, H-5); 3.75 (3H, s, –OCH3); 1.96 (2H, t, J = 7.5 Hz, H-2); 1.69 (2H, quintet, J = 7.5 Hz, H-4); 1.44 (2H, quintet, J = 7.5 Hz, H-3). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 168.90; 153.38; 130.99; 128.98; 128.50; 111.07; 110.42; 102.22; 100.03; 55.34; 45.30; 31.88; 29.54; 22.58. Anal. Calcd. For C14H18N2O3 (262.30): C, 64.10; H, 6.92; N, 10.68. Found: C, 64.12; H, 6.90; N, 10.70.
N-Hydroxy-6-(1H-indol-1-yl)hexanamide (3e)
Viscous liquid, black; Yield: 54%. R f = 0.45 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3440 (NH), 3190 (OH), 3040 (C–H, arene), 2856 (CH, CH2), 1714, 1655 (C=O), 1608 (C=C), 1463 (C–N). ESI–MS (m/z): 245 [M − H]–. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.43 (1H, s, –OH); 8.69 (1H, s, –NH); 7.53 (1H, s, H-5′); 7.43 (1H, s, H-8′); 7.33 (1H, s, H-2′); 7.12 (1H, s, H-7′); 7.01 (1H, s, H-6′); 6.41 (1H, s, H-3′); 4.13 (2H, s, H-6); 1.93 (2H, s, H-2); 1.74 (2H, s, H-5); 1.52 (2H, s, H-3); 1.22 (2H, s, H-4). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 169.06; 135.62; 128.52; 128.09; 120.92; 120.41; 118.80; 109.70; 100.36; 45.33; 32.16; 29.56; 25.88; 24.73. Anal. Calcd. For C14H18N2O2 (246.31): C, 68.27; H, 7.37; N, 11.37. Found: C, 68.29; H, 7.35; N, 11.38.
6-(6-Chloro-1H-indol-1-yl)-N-hydroxyhexanamide (3f)
Viscous liquid, black; Yield: 50%. R f = 0.40 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3430 (NH), 3191 (OH), 3043 (C–H, arene), 2850 (CH, CH2), 1714, 1655 (C=O), 1608 (C=C), 1460 (C–N). ESI–MS (m/z): 281.10 [M + H]+. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.29 (1H, s, –OH); 8.63 (1H, s, –NH); 7.59 (1H, s, H-7′); 7.53 (1H, d, J = 8.5 Hz, H-4′); 7.39 (1H, d, J = 3 Hz, H-2′); 7.01 (1H, q, J = 2.0, 8.5 Hz, H-5′); 6.44 (1H, d, J = 3 Hz, H-3′); 4.13 (2H, t, J = 7 Hz, H-6); 1.90 (2H, t, J = 7 Hz, H-2); 1.69 (2H, quintet, J = 7.5 Hz, H-5); 1.48 (2H, quintet, J = 7.5 Hz, H-3); 1.20 (2H, m, H-4). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 168.90; 136.04; 129.67; 126.74; 125.83; 121.64; 119.09; 109.60; 100.72; 45.37; 32.10; 29.47; 25.76; 24.65. Anal. Calcd. For C14H17ClN2O2 (280.75): C, 59.89; H, 6.10; N, 9.98. Found: C, 59.91; H, 6.12; N, 10.01.
6-(5-Chloro-1H-indol-1-yl)-N-hydroxyhexanamide (3g)
Viscous liquid, black; Yield: 48%. R f = 0.37 (DCM:MeOH = 10:1). IR (KBr, cm−1): 33,350 (NH), 3175 (OH), 3125 (C–H, arene), 2885 (CH, CH2), 1700, 1650 (C=O), 1618 (C=C), 1443 (C–N). ESI–MS (m/z): 315.31 [M + NH4OH]–. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.31 (1H, s, –OH); 8.66 (1H, s, –NH); 7.57 (1H, s, H-7′); 7.48 (1H, s, H-4′); 7.43 (1H, s, H-2′); 7.11 (1H, s, H-6′); 6.41 (1H, s, H-3′); 4.14 (2H, s, H-6); 1.91 (2H, s, H-2); 1.72 (2H, s, H-5); 1.50 (2H, s, H-3); 1.21 (2H, s, H-4). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 168.96; 134.14; 130.26; 129.14; 123.51; 120.80; 119.48; 111.31; 100.20; 45.51; 32.10; 29.50; 25.78; 24.65. Anal. Calcd. For C14H17ClN2O2 (280.75): C, 59.89; H, 6.10; N, 9.98. Found: C, 59.90; H, 6.14; N, 10.00.
N-Hydroxy-6-(5-methoxy-1H-indol-1-yl)hexanamide(3h)
Viscous liquid, black; Yield: 46%. R f = 0.35 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3435 (NH), 3165 (OH), 3024 (C–H, arene), 2843 (CH, CH2), 1724, 1642 (C=O), 1612 (C=C), 1435 (C–N). ESI–MS (m/z): 311.33 [M + NH4OH]–. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.33 (1H, s, –OH); 8.68 (1H, s, –NH); 7.32 (1H, d, J = 9.0 Hz, H-7′); 7.28 (1H, d, J = 2.5 Hz, H-4′); 7.03 (1H, d, J = 2 Hz, H-2′); 6.75 (1H, q, J = 2.0, 9.0 Hz, H-6′); 6.31 (1H, d, J = 2.0 Hz, H-3′); 4.07 (2H, t, J = 7 Hz, H-6); 3.74 (3H, s, –OCH3); 1.89 (2H, t, J = 7 Hz, H-2); 1.68 (2H, quintet, J = 7.5 Hz, H-5); 1.47 (2H, quintet, J = 7.5 Hz, H-3); 1.16 (2H, m, H-4). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 169.02; 153.34; 130.96; 129.00; 128.47; 111.08; 110.45; 102.14; 100.01; 55.33; 45.50; 32.18; 29.67; 25.89; 24.76. Anal. Calcd. For C15H20N2O3 (276.33): C, 65.20; H, 7.30; N, 10.14. Found: C, 65.23; H, 7.32; N, 10.11.
N-Hydroxy-7-(1H-indol-1-yl)heptanamide (3i)
Viscous liquid, black; Yield: 55%. R f = 0.45 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3455 (NH), 3163 (OH), 3025 (C–H, arene), 2874 (CH, CH2), 1706, 1648 (C=O), 1617 (C=C), 1424 (C–N). ESI–MS (m/z): 261.0 [M + H]−+, 243.0 [M–OH]+. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.38 (1H, s, –OH); 8.68 (1H, s, –NH); 7.52 (1H, d, J = 7.5 Hz, H-4′); 7.43 (1H, d, J = 7.0 Hz, H-7′); 7.33 (1H, s, H-2′); 7.10 (1H, t, J = 6.5 Hz, H-6′); 6.98 (1H, t, J = 6.5 Hz, H-5′); 6.40 (1H, s, H-3′); 4.13 (2H, s, H-7); 1.92 (2H, s, H-2); 1.72 (2H, m, H-6); 1.44–1.23 (6H, m, H-3, H-4, H-5). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 169.22; 135.64; 128.53; 128.10; 120.91; 120.40; 118.78; 109.69; 100.35; 45.40; 32.19; 29.70; 28.17; 25.98; 25.02. Anal. Calcd. For C15H20N2O2 (260.15): C, 69.20; H, 7.74; N, 10.76. Found: C, 69.23; H, 7.75; N, 10.72.
7-(6-Chloro-1H-indol-1-yl)-N-hydroxyheptanamide (3j)
Viscous liquid, umber; Yield: 53%. R f = 0.42 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3444 (NH), 3194 (OH), 3043 (C–H, arene), 2850 (CH, CH2), 1712, 1655 (C=O), 1618 (C=C), 1463 (C–N). ESI–MS (m/z): 295.11 [M + H]−+. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.30 (1H, s, –OH); 8.62 (1H, s, –NH); 7.59 (1H, s, H-7′); 7.53 (1H, d, J = 8 Hz, H-4′); 7.40 (1H, d, J = 2.5 Hz, H-2′); 7.00 (1H, d, J = 7.5 Hz, H-5′); 6.44 (1H, d, J = 2 Hz, H-3′); 4.13 (2H, t, J = 7.0 Hz, H-7); 1.90 (2H, t, J = 7.0 Hz, H-2); 1.70 (2H, quintet, J = 7.5 Hz, H-6); 1.43 (2H, quintet, J = 7.5 Hz, H-3); 1.23 (4H, m, H-4, H-5). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 169.03; 136.06; 129.69; 126.75; 125.83; 121.66; 119.09; 109.60; 100.73; 45.46; 32.14; 29.63; 28.13; 25.89; 24.98. Anal. Calcd. For C15H19ClN2O2 (294.78): C, 61.12; H, 6.50; N, 9.50. Found: C, 61.15; H, 6.52; N, 9.52.
7-(5-Chloro-1H-indol-1-yl)-N-hydroxyheptanamide (3k)
Viscous liquid, umber; Yield: 50%. R f = 0.39 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3441 (NH), 3197 (OH), 3049 (C–H, arene), 2856 (CH, CH2), 1714, 1655 (C=O), 1608 (C=C), 1463 (C–N). ESI–MS (m/z): 295.44 [M + H]−+. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.33 (1H, s, –OH); 8.67 (1H, s, –NH); 7.57 (1H, d, J = 1.5 Hz, H-4′); 7.48 (1H, d, J = 9.0 Hz, H-7′); 7.43 (1H, d, J = 3.0 Hz, H-2′); 7.10 (1H, q, J = 2.0, 8.5 Hz, H-6′); 6.41 (1H, d, J = 2.5 Hz, H-3′); 4.12 (2H, t, J = 7.0 Hz, H-7); 1.89 (2H, t, J = 7.0 Hz, H-2); 1.69 (2H, quintet, J = 7.5 Hz, H-6); 1.40 (2H, quintet, J = 7.5 Hz, H-3); 1.19 (4H, m, H-4, H-5). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 169.14; 134.21; 130.35; 129.19; 123.56; 120.86; 119.55; 111.39; 100.25; 45.63; 32.20; 29.72; 28.17; 25.96; 25.04. Anal. Calcd. For C15H19ClN2O2 (294.78): C, 61.12; H, 6.50; N, 9.50. Found: C, 61.10; H, 6.54; N, 9.50.
N-Hydroxy-7-(5-methoxy-1H-indol-1-yl)heptanamide (3l)
Viscous liquid, umber; Yield: 47%. R f = 0.41 (DCM:MeOH = 10:1). IR (KBr, cm−1): 3435 (NH), 3185 (OH), 3035 (C–H, arene), 2846 (CH, CH2), 1711, 1654 (C=O), 1607 (C=C), 1463 (C–N). ESI–MS (m/z): 289.0 [M − H]−. 1H-NMR (500 MHz, DMSO-d 6, ppm): δ 10.30 (1H, s, –OH); 8.62 (1H, s, –NH); 7.32 (1H, d, J = 8.5 Hz, H-7′); 7.28 (1H, d, J = 3 Hz, H-4′); 7.03 (1H, d, J = 2.5 Hz, H-2′); 6.75 (1H, q, J = 9.0 Hz, J = 2.5 Hz, H-6′); 6.31 (1H, d, J = 2.5 Hz, H-3′); 4.08 (2H, t, J = 7 Hz, H-7); 3.74 (3H, s, –OCH3), 1.89 (2H, t, J = 7.0 Hz, H-2); 1.69 (2H, quintet, J = 7.5 Hz, H-6); 1.42 (2H, quintet, J = 7.5 Hz, H-3); 1.20 (4H, m, H-4, H-5). 13C NMR (125 MHz, DMSO-d 6, ppm): δ 169.05; 153.29; 130.93; 128.92; 128.42; 111.00; 110.34; 102.15; 99.92; 55.30; 45.50; 32.14; 29.72; 28.13; 25.93; 24.97. Anal. Calcd. For C16H22N2O3 (290.36): C, 66.18; H, 7.64; N, 9.65. Found: C, 66.21; H, 7.66; N, 9.63.
Synthesis of N-hydroxy-4-(indolin-1-yl)heptanamide (6)
Indoline (4, 0.24 g, 2.00 mmol) was dissolved in DMF (abs) (2 mL) in a 50-mL round-bottomed flask containing potassium carbonate (0.55 g, 4.00 mmol) and a catalytic amount of potassium iodide (30 mg). The mixture was stirred at room temperature for 45 min, and then a solution of methyl-7-bromoheptanoate (0.45 g, 2 mmol) in DMF (abs) (1 mL) was added dropwise. The reaction mixture was further stirred for 24 h at room temperature. After that, the whole reaction mixture was transferred into 20 mL of cold water. It was neutralized by a solution of HCl 5% and extracted by dichloromethane. The extract was dried over anhydrous sodium sulfate and evaporated under vacuum to give a bright yellow oil (5). The oil obtained was dissolved directly into 10 mL of methanol in a 50-mL round-bottomed flask. To the resulting solution was added NH2OH.HCl (1.40 g, 20.0 mmol). The mixture was cooled to −5 ÷ 0 °C and stirred at this temperature for about 30 min, and then a solution of NaOH (1.20 g, 30.0 mmol) in water (3 mL) was slowly added. The reaction was continued for 60 min at −5 ÷ 0 °C, then transferred to 20 mL of ice-water, and acidified to pH 5 by a solution of HCl 5% to give precipitates. The precipitates were recrystallized from ethanol/water to give compound 6 as yellow solids. Yield: 48%; mp: 132–133 °C; R f = 0,43 (DCM:MeOH:AcOH, 90:8:1). IR (KBr, cm−1): 3047 (νOH), 2931 (νas-CH2), 2857 (νs-CH2), 1648 (νC=O), 1488, 1470, 1459 (νC…C). CI-MS: m/z 262.9 [M + H]+; 244.9 [M–OH]+. 1H-NMR (500 MHz, DMSO-d 6, ppm):δ 10.35 (1H, s, OH), 8.68 (1H, s, NH), 8.07 (1H, d, J = 8 Hz, H-7′, indoline), 7.20 (1H, d, J = 7.0 Hz, H-4′, indoline); 7.12 (1H, t, J = 7.5 Hz, H-6′, indoline); 6.96 (1H, t, J = 7.5 Hz, H-5′, indoline); 3.99–3.95 (2H, m, CH2, indoline); 3.39–3.37 (2H, m, CH2), 3.10–3.07 (2H, m, CH2, indoline); 2.17–2.15 (2H, m, CH2), 1.78–1.72 (4H, m, CH2), 1.33–1.29 (4H, m, CH2); 13C NMR (125 MHz, DMSO-d 6, ppm): δ 169.22; 135.64; 128.53; 128.10; 120.91; 120.40; 118.78; 109.69; 100.35; 45.40; 32.19; 29.70; 28.17; 25.98; 25.02. Anal. Calcd. For C15H22N2O2 (262.17): C, 68.67; H, 8.45; N, 10.68. Found: C, 68.71; H, 8.48; N, 10.71.
Cytotoxicity assay
The cytotoxicity of the synthesized compounds was evaluated against four human cancer cell lines, including SW620 (colon cancer), PC3 (prostate cancer), AsPC-1 (pancreatic cancer), and NCI-H23 (lung cancer). The cell lines were purchased from a Cancer Cell Bank at the Korea Research Institute of Bioscience and Biotechnology (KRIBB). The media, sera, and other reagents that were used for cell culture in this assay were obtained from GIBCO Co. Ltd. (Grand Island, New York, USA). The cells were cultured in DMEM (Dulbecco’s Modified Eagle Medium) until confluence. The cells were then trypsinized and suspended at 4 × 104 cells/mL of cell culture medium for SW620 or 5 × 104 cells/mL of cell culture medium for NCI-H23, PC-3, and AsPC-1. On day 0, each well of the 96-well plates was seeded with 180 µL of cell suspension. The plates were then incubated in a 5% CO2 incubator at 37 °C for 24 h. Compounds were initially dissolved in dimethyl sulfoxide (DMSO) and diluted to appropriate concentrations by culture medium. Then, 20 µL of each compound’s samples, which were prepared as described above, was added to each well of the 96-well plates, which had been seeded with cell suspension and incubated for 24 h at various concentrations. The plates were further incubated for 48 h. Cytotoxicity of the compounds was measured by the colorimetric method, as described previously (Skehan et al. 1990) with slight modifications (Nam et al. 2003; You et al. 2003; Ye et al. 2007). The IC50 values were calculated using a Probits method (Wu et al. 1992) and computed averages of three independent determinations (SD ≤ 10%).
Enzyme assay
The HDAC2 enzyme was purchased from BPS Bioscience (San Diego, CA, USA). The HDAC enzymatic assay was performed using a Fluorogenic HDAC Assay Kit (BPS Bioscience) according to the manufacturer’s instructions. Briefly, HDAC2 enzymes were incubated with vehicle or various concentrations of the assayed samples or SAHA for 30 min at 37 °C in the presence of an HDAC fluorimetric substrate. The HDAC assay developer (which produces a fluorophore in reaction mixture) was added, and the fluorescence was measured using VICTOR3 (PerkinElmer, Waltham, MA, USA) with excitation at 360 nm and emission at 460 nm. The measured activities were subtracted by the vehicle-treated control enzyme activities and IC50 values were calculated using GraphPad Prism (GraphPad Software, San Diego, CA, USA).
Docking studies
An AutoDock Vina program (Trott and Olson 2010) (The Scripps Research Institute, CA, USA) was used for docking. The structure of HDAC2 protein in complex with SAHA (Lauffer et al. 2013) was obtained from the Protein Data Bank (PDB ID: 4LXZ). The coordinates of the compounds were generated by using the GlycoBioChem PRODRG2 Server (http://davapc1.bioch.dundee.ac.uk/prodrg/) (Schuttelkopf and van Aalten 2004). For the docking studies, the grid maps were centered on the SAHA-binding site and comprised 26 × 26 × 22 points with 1.0 Å spacing after SAHA was removed from the complex structure, as described in the previous works (Oanh et al. 2011; Tung et al. 2013; Nam et al. 2014). As the ionization state of hydroxamic acids in complex with Zn2+ was widely suggested as negative hydroxamate coordination (Wu et al. 2011), herein their hydroxyl groups were deprotonated. The AutoDock Vina program was performed using eight-way multithreading and the other parameters had default settings.
Results and discussion
Chemistry
The target hydroxamic acids (3a–l) were synthesized via two-step pathway, as illustrated in Scheme 1. The first step was a nucleophilic substitution (SN2) between indole (1) and methyl bromoalkanoates (including methyl 5-bromopentanoate, methyl 6-bromohexanoate, and methyl 7-bromoheptanoate) using sodium hydride (NaH) as a base and a catalytic amount of potassium iodide in anhydrous dimethylformamide (DMF, abs). The second step was also a nucleophilic acyl substitution between hydroxylamine, which was generated from hydroxylamine hydrochloride, and the esters (2). This reaction occurred under alkaline conditions in methanol at −5 °C. The overall yields of compounds 3a–l were moderate (from 45 to 55%).
One compound bearing an indoline moiety instead of indole (compound 6) was also synthesized by a similar strategy via a two-step pathway (Scheme 2). The overall yield was 48% calculated from the indoline starting material 4. The structures of the synthesized compounds (3a–l, 6) were determined straightforwardly by analysis of spectroscopic data, including IR, MS, 1H-NMR, and 13C-NMR.
Bioactivity
It has been demonstrated that HDAC2 deacetylates a number of histone proteins and plays very important roles in different events relating to cancer cell proliferation. HDAC2 is also one of the key proteins that cause cell apoptosis arrest (Pelzel et al. 2010). Therefore, in this study, we firstly screened the synthesized compounds for inhibitory effects against this type of enzyme. The HDAC2 inhibitory activities of compounds 3a–l and compound 6 are summarized in Table 1. Table 1 shows that the HDAC2 inhibitory potency of the compounds increased with the length of the carbon bridge between the indole and hydroxamic acid moieties. For example, the IC50 of compounds 3a (4C-bridge), 3e (5C-bridge), and 3i (6C bridge) were 53.16, 26.20, and 0.93 μM, respectively. Thus, compound 3i (6C bridge) was found to be more than 28- and 57-fold more potent than compounds 3e and 3a, respectively. Similar trends were also observed for almost all cases (e.g., compounds 3j–l vs. 3f–h; 3g–h vs. 3c–d). Thus, 6C-bridge was found to be the best among different length linkers investigated. Substitution of either Cl or –OCH3 groups at positions 5 or 6 on the indole ring was found to enhance the HDAC2 inhibitory effects significantly (e.g., compounds 3c-d vs. compound 3a; compounds 3f–h vs. compound 3e; compounds 3j–l vs. compound 3i). It was found that in spite of the truncation of the amide moiety, the HDAC2 inhibition of the indole-based hydroxamic acids 3i–l was still more potent than SAHA. Especially, compounds 3j, 3k, and 3l were ten-, three- and fivefold more potent than SAHA in terms of HDAC2 inhibition. Thus, the indole ring could be more favorable as a cap group compared to the phenyl ring in SAHA. In contrast, the indoline heterocycle seemed to be less favorable, as manifested by the inhibitory effects of compound 6 towards HDAC2 activity (Table 1).

All compounds were then evaluated for their cytotoxicity against four human cancer cell lines, including SW620 (human colon cancer), PC-3 (prostate cancer), AsPC-1 (pancreas cancer), and NCI-H23 (lung cancer). In this evaluation, we used an SRB method (Skehan et al. 1990). The IC50 values of compounds were calculated using a Probits method (Wu et al. 1992) and expressed as averages of three independent determinations (SD ≤ 10%) (Table 1). The IC50 values of compounds shown in Table 1 clearly demonstrate a very good correlation between the cytotoxicity of the compounds in four cancer cell lines with their inhibition of HDAC2. Six compounds, including 3g–3l, were more cytotoxic than SAHA. Two compounds 3k and 3l were the most potent ones in the series. Especially compound 3l was approximately 12-fold more cytotoxic than SAHA in PC3 cells, and up to 77-fold more potent than SAHA in terms of cytotoxicity towards NCI-H23 cell line.
Preliminary assessment of drug-like properties of these compounds showed that all compounds met criteria of Lipinski’s rule of five. The logP values of the compounds in the series were in the range of 1.96–3.58, generally favorable for cellular penetration while retaining acceptable water solubility.
Docking studies
The crystal structure of HDAC2 in complex with SAHA (PDB ID: 4LXZ) has been reported by Lauffer and co-workers (Lauffer et al. 2013), so we decided to utilize this crystal structure in docking experiments to study the interaction between these hydroxamic acids and HDAC. A control docking with co-crystal SAHA to the crystal structures of HDAC2 using AutoDock Vina program (Trott and Olson 2010) was firstly carried out, taking into account the RMSD values and interactions with the enzyme (Oanh et al. 2011; Tung et al. 2013; Nam et al. 2014; Huong et al. 2017). Accordingly, the re-docked SAHA displayed low deviation (RMSD = 0.627 Å), docking score of −7.4 kcal/mol and similar interaction pattern to the co-crystal SAHA compound (key binding site residues include Asp104, His145, His146, and Tyr308). Considering the suitable results obtained, our protocol is further applied for the docking studies of synthesized hydroxamic derivatives.
From docking experiments, it was found that all of the hydroxamic acids synthesized were well located in the active site of the enzyme with binding affinities comparable or slightly higher than that of SAHA, as indicated by docking score energies ranging from −7.2 to −7.7 kcal/mol. Importantly, all the compounds exhibited chelate interaction with the catalytic Zn2+ ion in a similar manner as SAHA did (Fig. 3). It was found that the linker-chelator motif is the principal component of HDAC inhibitors (Wu et al. 2011), and the Zn2+ ion formed octahedral coordination geometry with the catalytic residues (Asp181, His183, and Asp269) and the docked ligand. The coordination distances from Zn2+ to carbonyl and hydroxyl oxygen atoms of hydroxamic acids were in the range of 2.3–2.8 Å (those of SAHA range 2.3–2.4 Å). With the exception of 3a, 3c, 3d, and 3f, all the compounds formed two H-bonds with His145 and His146 in the binding site. These interactions are typical in pharmacophore model of SAHA-like inhibitors proposed by Vanommeslaeghea et al. (2005). Compounds with extensive hydrogen-bonding interactions are 3b, 3g, 3h, and 3i–l, whose hydroxamic groups participated in four H-bonds with His145, His146, Asp181, and/or Glu208. Additionally, the indole part was found to enhance hydrophobic interactions compared with aniline ring of SAHA at the rim of the pocket (Fig. 4). Residues involved in hydrophobic interactions namely, Leu276, His183, and Tyr209 were quite consistent. The aliphatic linkers were positioned to appropriately fit in narrow hydrophobic tunnel of the active pocket of HDAC2 enzyme. Interestingly, the activity of this series correlated to some degree with the length of the linker. Accordingly, compounds 3i–l displayed suitable flexibility and better superposition with SAHA than other analogs.
From docking score evaluation however, very little variance in the binding affinities among the compounds with different substituted groups was observed. For example, the docking score energies of predicted binding modes on HDAC2 of compounds 3i (no substituent on the indole moiety) and 3l (with a methoxy substituent at position 5 on the indole moiety) were found to be −7.7 and −7.6 kcal/mol, respectively. These values were not much different from that of compounds 3j and 3k (−7.4 and −7.3 kcal/mol, respectively). Thus, the difference in the docking score could not explain for the three- to ninefold difference in the IC50 value of compound 3i with that of compounds 3j–l in the HDAC2 enzyme inhibition assay (Table 1). Unsurprisingly, the reasons of unmatched results between docking scores and IC50 values have been widely discussed in the literature (Huang and Zou 2010; Cheng 2015; Ramírez and Caballero 2016), including suboptimal docking tool and/or setup, imperfection of the docking scoring algorithm, missing structural information related to solvation, protonation, tautomerism, isomerism during the docking procedures, and even unsuitable IC50 value assessment, to name a few. In order to better correlate the binding affinities of the compounds towards HDACs and their HDAC inhibitory effects, a more realistic binding energy estimation approach should be considered, such as QM/MM-GBSA, MM-PBSA, or MM-GBSA (Khandelwal et al. 2015).
Conclusions
In conclusion, we have reported that a series of hydroxamic acids incorporated indole moiety as a cap group and different linker lengths with significant HDAC2 inhibitory effects and potent cytotoxicity against several human cancer cell lines, including SW620 (human colon cancer), PC-3 (prostate cancer), AsPC-1 (pancreas cancer), and NCI-H23 (lung cancer). The results we obtained from this study again confirm that the indole moiety could well serve as a cap group for hydroxamic acid HDAC inhibitors. Also, different substituents on the indole system substantially enhanced both HDAC inhibition and cytotoxicity of the resulting compounds. From this study, several potent HDAC inhibitors with strong cytotoxicity against human cancer cells, such as compounds 3k, 3l, have emerged.
References
Bolden JE, Peart MJ, Johnstone RW (2006) Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov 5:769–784. doi:10.1038/nrd2133
Bracker TU, Sommer A, Fichtner I, Faus H, Haendler B, Hess-Stumpp H (2009) Efficacy of MS-275, a selective inhibitor of class I histone deacetylases, in human colon cancer models. Int J Oncol 35:909–920. doi:10.3892/ijo_00000406
Cheng YC (2015) Beware of docking! Trends Pharmacol Sci 36:78–95. doi:10.1016/j.tips.2014.12.001
Cheng T, Grasse L, Shah J, Chandra J (2015) Panobinostat, a pan-histone deacetylase inhibitor: rationale for and application to treatment of multiple myeloma. Drugs Today (Barc) 51:491–504. doi:10.1358/dot.2015.51.8.2362311
Dallavalle S, Cincinelli R, Nannei R, Merlini L, Morini G, Penco S, Pisano C, Vesci L, Barbarino M, Zuco V, De Cesare M, Zunino F (2009) Design, synthesis, and evaluation of biphenyl-4-yl-acrylohydroxamic acid derivatives as histone deacetylase (HDAC) inhibitors. Eur J Med Chem 44:1900–1912. doi:10.1016/j.ejmech.2008.11.005
De Ruijter AJM, Gennip AHV, Caron HN, Kemp S, Kuilenburg ABPV (2003) Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem J 370:737–739. doi:10.1042/BJ20021321
Glaser KB (2007) HDAC inhibitors: clinical update and mechanism-based potential. Biochem Pharmacol 74:659–671. doi:10.1016/j.bcp.2007.04.007
Hamm CA, Costa FF (2015) Epinomes as therapeutic targets. Pharmacol Ther 151:72–86. doi:10.1016/j.pharmthera.2015.03.003
Huang S-Y, Zou X (2010) Advances and challenges in protein-ligand docking. Int J Mol Sci 11:3016–3034. doi:10.3390/ijms11083016
Huong TTL, Dung DTM, Huan NV, Cuong LV, Hai PT, Huong LTT, Kim J, Kim YG, Han SB, Nam NH (2017) Novel N-hydroxybenzamides incorporating 2-oxoindoline with unexpected potent histone deacetylase inhibitory effects and antitumor cytotoxicity. Bioorg Chem. doi: 10.1016/j.bioorg.2017.02.002 (in press)
Iyer SP, Foss FF (2015) Romidepsin for the treatment of peripheral T-cell lymphoma. Oncologist 20:1084–1091. doi:10.1634/theoncologist.2015-0043
Khandelwal A, Lukacova V, Comez D, Kroll DM, Raha S, Balaz S (2015) A combination of docking, QM/MM methods, and MD simulation for binding affinity estimation of metalloprotein ligands. J Med Chem 48:437–447. doi:10.1021/jm049050v
Lauffer BE, Mintzer R, Fong R, Mukund S, Tam C, Zilberleyb I, Flicke B, Ritscher A, Fedorowicz G, Vallero R, Ortwine DF, Gunzner J, Modrusan Z, Neumann L, Koth CM, Lupardus PJ, Kaminker JS, Heise CE, Steiner P (2013) Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. J Biol Chem 288:26926–26943. doi:10.1074/jbc.M113.490706
Li J, Li G, Xu X (2013) Histone deacetylase inhibitors: an attractive strategy for cancer therapy. Curr Med Chem 20:1858–1886. doi:10.1074/jbc.M113.490706
Malini G (2015) HDAC inhibitors still need a home run, despite recent approval. Nat Rev Drug Discov 14:225–226. doi:10.1038/nrd4583
Nam NH, Parang K (2003) Current targets for anticancer drugs discovery. Curr Drug Targets 4:159–179. doi:10.2174/1389450033346966
Nam NH, Lee C-W, Hong D-H, Kim H-M, Bae K-H, Ahn B-Z (2003) Antiinvasive, antiangiogenic and antitumour activity of Ephedra sinica extract. Phytother Res 17:70–76. doi:10.1002/ptr.901
Nam NH, Huong TL, Dung DTM, Oanh DTK, Dung PTP, Quyen D, Kim KR, Han BW, Kim YS, Hong JT, Han SB (2013) Novel isatin-based hydroxamic acids as histone deacetylase inhibitors and antitumor agents. Eur J Med Chem 70:477–486. doi:10.1016/j.ejmech.2013.10.045
Nam NH, Huong TL, Dung DTM, Oanh DTK, Dung PTP, Kim KR, Han BW, Kim YS, Hong JT, Han SB (2014) Synthesis, bioevaluation and docking study of 5-substitutedphenyl-1,3,4-thiadiazole-based hydroxamic acids as histone deacetylase inhibitors and antitumor agents. J Enzyme Inhib Med Chem 29:611–618. doi:10.1016/j.ejmech.2013.10.045
Oanh DTK, Hai HV, Hue VTM, Park SH, Kim HJ, Han BW, Kim HS, Hong JT, Han SB, Nam NH (2011) Benzothiazole-containing hydroxamic acids as histone deacetylase inhibitors and antitumor agents. Bioorg Med Chem Lett 21:7509–7512. doi:10.1016/j.bmcl.2011.07.124
Pelzel HR, Schlamp CL, Nickells RW (2010) Histone H4 deacetylation plays a critical role in early gene silencing during neuronal apoptosis. BMC Neurosci 11:62. doi:10.1186/1471-2202-11-62
Qiu T, Zhou L, Zhu W, Wang T, Wang J, Shu Y, Liu P (2013) Effects of treatment with histone deacetylase inhibitors in solid tumors: a review based on 30 clinical trials. Future Oncol 9:255–269. doi:10.2217/fon.12.173
Ramírez D, Caballero J (2016) Is it reliable to use common molecular docking methods for comparing the binding affinities of enantiomer pairs for their protein target? Int J Mol Sci 17:E525. doi:10.3390/ijms17040525
Schuttelkopf AW, van Aalten DM (2004) PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr D Biol Crystallogr 60:1355–1363. doi:10.1107/S0907444904011679
Skehan P, Storeng R, Scudiero D, Monk A, MacMahon J, Vistica D, Warren JT, Bokesch H, Kenney S, Boyd MR (1990) New colorimetric cytotoxicity assay for anticancer drug screening. J Natl Cancer Inst 82:1107–1112. doi:10.1093/jnci/82.13.1107
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461. doi:10.1002/jcc.21334
Tung TT, Oanh DT, Dung PT, Hue VT, Park SH, Han BW, Kim Y, Hong JT, Han SB, Nam NH (2013) New benzothiazole/thiazole-containing hydroxamic acids as potent histone deacetylase inhibitors and antitumor agents. Med Chem 9:1051–1057. doi:10.2174/15734064113099990027
Valente S, Mai A (2014) Small-molecule inhibitors of histone deacetylase for the treatment of cancer and non-cancer diseases: a patent review (2011–2013). Expert Opin Ther Pat 24:401–415. doi:10.1517/13543776.2014.877446
Vanommeslaeghea K, Loverixb S, Geerlings P, Tourwéa D (2005) DFT-based ranking of zinc-binding groups in histone deacetylase inhibitors. Bioorg Med Chem 13:6070–7082. doi:10.1016/j.bmc.2005.06.009
Ververis K, Hiong A, Karagiannis TC, Licciardi PV (2013) Histone deacetylase inhibitors (HDACIs): multitargeted anticancer agents. Biologics 7:47–60. doi:10.2147/BTT.S29965
West AC, Johnstone RW (2014) New and emerging HDAC inhibitors for cancer treatment. J Clin Investig 124:30–39. doi:10.1172/JCI69738
Witt O, Deubzer HE, Milde T, Oehme I (2009) HDAC family: what are the cancer relevant targets? Cancer Lett 277:8–21. doi:10.1016/j.canlet.2008.08.016
Wu L, Smythe AM, Stinson SF, Mullendore LA, Monks A, Scudiero DA, Paull KD, Koutsoukos AD, Rubinstein LV, Boyd MR, Shoemaker RH (1992) Multidrug-resistant phenotype of disease-oriented panels of human tumor cell lines used for anticancer drug screening. Can Res 52:3029–3034. doi:10.1016/j.canlet.2008.08.016
Wu R, Lu Z, Cao Z, Zhang Y (2011) Zinc chelation with hydroxamate in histone deacetylases modulated by water access to the linker binding channel. J Am Chem Soc 133:6110–6113. doi:10.1021/ja111104p
Ye G, Nam NH, Kumar A, Saleh A, Shenoy DB, Amiji MM, Lin X, Sun G, Parang K (2007) Synthesis and evaluation of tripodal peptide analogues for cellular delivery of phosphopeptides. J Med Chem 50:3604–3617. doi:10.1021/jm070416o
You YJ, Kim Y, Nam NH, Ahn BZ (2003) Antitumor activity of unsaturated fatty acid esters of 4′-demethyldeoxypodophyllotoxin. Bioorg Med Chem Lett 13:2629–2632. doi:10.1016/S0960-894X(03)00558-4
Zwergel C, Valente S, Jacob C, Mai A (2015) Emerging approaches for histone deacetylase inhibitor drug discovery. Expert Opin Drug Discov 10:599–613. doi:10.1517/17460441.2015.1038236
Acknowledgements
We acknowledge the principal financial supports from the National Foundation for Science and Technology of Vietnam (NAFOSTED, Grant Number 104.01-2015.08). This work was also partly supported by the Grant Number 104.01-2014.55 (from NAFOSTED), the Korean Medical Research Center program (Grant Number 2008-0062275), and a small Grant from Hanoi University of Pharmacy.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors report no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huong, T.T.L., Van Cuong, L., Huong, P.T. et al. Exploration of some indole-based hydroxamic acids as histone deacetylase inhibitors and antitumor agents. Chem. Pap. 71, 1759–1769 (2017). https://doi.org/10.1007/s11696-017-0172-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11696-017-0172-1