Abstract
In this chapter we will describe recent developments in MEG network analysis, where we will focus on the rationale behind, and application in clinical cohorts of, an atlas-based beamforming approach. This approach contains 3 main components, namely (i) the reconstruction of time-series of neuronal activation through beamforming; (ii) the use of a standard atlas, which enables comparisons across studies and modalities; (iii) the estimation of functional connectivity using the Phase Lag Index (PLI), a measure that is insensitive to the effects of field spread/volume conduction. Moreover, we will discuss the use of the minimum spanning tree (MST), which allows for a bias-free characterization of the topology of the reconstructed functional networks. Application of this approach will be illustrated through examples from recent studies in patients with gliomas, Parkinson’s disease, and Multiple Sclerosis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Resting-state
- Network analysis
- Graph theory
- Minimum spanning tree
- Atlas-based beamformer
- Phase lag index (PLI)
- Clinical applications
1 Functional Brain Networks
The brain consists of billions of interconnecting neurons, forming an extremely complex system (Tononi et al. 1998; Tononi and Edelman 1998) in which clusters of neurons are organized as functional units with more-or-less specific information processing capabilities (e.g. Born and Bradley 2005; Grodzinsky 2000). Yet, cognitive functions require the coordinated activity of these spatially separated units, where the oscillatory nature of neuronal activity may provide a possible mechanism (Buzsaki and Wang 2012; Engel et al. 2001; Fries 2005; Singer 1999; Varela et al. 2001). These interacting units form a large-scale complex network (Bullmore and Sporns 2012; Schnitzler and Gross 2005). The organization of such complex brain networks can be characterized using concepts from graph theory (Bullmore and Sporns 2009; Reijneveld et al. 2007; Stam and Reijneveld 2007; Watts and Strogatz 1998). Application of graph theoretical tools to human brain networks has shown that the brain is organized according to a highly efficient topology that combines a high level of local integration (i.e. dense local clustering of connections) with a high level of global efficiency (i.e. critical long-distance connections), forming a so-called small-world organization (Bassett and Bullmore 2006; Stam and van Straaten 2012b; Watts and Strogatz 1998). In addition, brain networks in healthy subjects contain a subset of relatively highly connected regions (‘hubs’) (Achard et al. 2006; Barabasi and Albert 1999). These hubs seem to be mutually and densely interconnected, forming a connectivity backbone or “rich club” crucial for efficient brain communication (van den Heuvel et al. 2012; van den Heuvel and Sporns 2011).
It has been shown that network topology is highly heritable (Smit et al. 2008, 2010), that the network configuration changes during the life span (Smit et al. 2012) and that there are gender differences (Smit et al. 2008; Tian et al. 2011). Moreover, an increasing number of studies has shown that various brain disorders disturb the optimal organization of the functional brain networks (for reviews see Reijneveld et al. 2007; Stam and van Straaten 2012b; van Straaten and Stam 2013), and that these network alterations correlate with cognitive performance, as well as with parameters of disease severity and/or progression.
2 Source-Space Analysis
Magnetoencephalography, with its high temporal resolution, can be used to characterize the functional brain networks that are formed by interacting sources of oscillatory activity. Although such an analysis can be performed directly at the sensor-level, there are several factors that should be considered. Firstly, multiple sensors pick up the signals from a single source due to the nature of the electromagnetic signals (Sarvas 1987), known as field spread, as well as due to volume conduction.Footnote 1 Both these phenomena may lead to erroneous estimates of functional connectivity. It is important to realize though that projection of the signals to source-level in itself does not eliminate these effects (Hillebrand et al. 2012). Secondly, the mixture of signals originating from spatially separated brain areas can result in under- or overestimation of functional connectivity (Schoffelen and Gross 2009). Demixing the contribution from spatially separate sources, as well as enabling a more straightforward interpretation of the functional data in relation to its underlying structure, are therefore the main reasons to perform an analysis in source-space. This requires the solution of the inverse problem, i.e. the problem of estimating the electrical current distribution that produced the recorded magnetic flux. This is an ill-posed problem, meaning that there is no unique solution, unless prior knowledge (or constraints) is added. We know, for example, that the cortical current density is small (the moment per unit area is typically of the order of 50 pAm/mm2; Lü and Williamson 1991), and solutions with estimated source strengths of several Ampere-meter can therefore safely be ignored. Different source reconstruction techniques exist (Baillet et al. 2001), and they vary in the type and number of constraints that are imposed (Hillebrand and Barnes 2005; Wipf and Nagarajan 2009). Constraints might be that there are only a small number of sources active at a specific instant in time (multi-dipole solutions; Supek and Aine 1993), that the whole cortex is active to some degree but with the minimum energy necessary to describe the measured data (minimum norm solutions; Hamalainen and Ilmoniemi 1994), or that there are no perfectly linearly correlated areas of activation within the brain (beamformers; Robinson and Vrba 1999; Sekihara and Nagarajan 2008; van Veen et al. 1997).
In recent years, beamforming has become one of the main source reconstruction approaches for MEG. It has been argued that the uncorrelated-source assumption may be realistic for many empirical datasets (Hillebrand and Barnes 2005), and violations of this assumption can be tolerated to some extent (Hadjipapas et al. 2005). For those cases where strongly correlated sources are encountered, for example during auditory stimulation or parallel processing of visual stimuli, the beamformer formulism can be adapted (Brookes et al. 2007; Dalal et al. 2006; Diwakar et al. 2011; Hui et al. 2010; Quraan and Cheyne 2010). From a practical point of view, there are few parameters to set when performing beamformer analysis, the main ones being the time-frequency window(s) in the data for which to perform the source reconstruction (Dalal et al. 2008). Source reconstruction is achieved in a sequential manner, where for each target location in the brain (typically a grid consisting of 5 × 5 × 5 mm voxels is used; Barnes et al. 2004) neuronal activity is estimated using an optimal set of beamformer weights, W:
where \({\hat{\text{Q}}}\)is the estimated source strength in nAm for a source at a given target voxel, and with a certain orientation; B is a vector containing the recorded magnetic flux at a given latency.
These weights are optimal in the sense that the values of the weights are chosen such that activity would be fully reconstructed for a target location, if this target location happens to be active (this is called the unit-gain constraint), whilst rejecting the contribution from all other sources, be it within or outside the brain. For a mathematical description we refer the reader to Robinson and Vrba 1999; Sekihara and Nagarajan 2008; van Veen et al. 1997, and for a review see Hillebrand et al. 2005).
Although other source reconstruction approaches also require accurate MEG/MRI co-registration and modeling of the volume conductor, beamforming is particularly sensitive to inaccuracies in the forward solution (Hillebrand and Barnes 2011, 2003; Vrba 2002): the unit-gain constraint described above results in a suppression of source activity if there is a deviation from the correct forward solution.
Equation 1 assumes that the orientation of a source is known. In practice, this is not the case, and the orientation can be set to the one that gives the maximum beamformer output (scalar beamformer; Robinson and Vrba 1999; Sekihara et al. 2004), the orientation of the cortical surface could be used (but see Hillebrand and Barnes 2003), or one could estimate the beamformer output for three orthogonal directions (vector beamformer; van Veen et al. 1997). Finally, the decrease in sensitivity for deeper sources (Hillebrand and Barnes 2002) results in an increase in the (norm of) the beamformer weights with source depth, and a disproportionate amplification of white sensor noise for deeper sources. To compensate for this depth bias, the beamformer weights, or equivalently the reconstructed beamformer image (Cheyne et al. 2006), are typically rescaled using a (projection of) the sensor noise. An estimate of the sensor noise therefore has to be provided. The effects of noise can further be reduced through regularization (Vrba 2002).
Once the beamformer weights have been estimated, one can reconstruct a three-dimensional volumetric image of activity (or of a change in activity in case experimental conditions are contrasted; see also Brookes et al. 2005). The statistical significance of these individual images is difficult to determine (but see Barnes and Hillebrand 2003), yet one can readily perform group-level statistics using tools developed for functional Magnetic Resonance Imaging (fMRI; Singh et al. 2002, 2003). We have recently introduced an atlas-based approach that also enables the comparison of beamformer results across individuals (see Fig. 1; Hillebrand et al. 2012). For each individual, the neuronal activity is reconstructed for a limited set of regions-of-interest (ROIs) that covers almost the entire brain, where the ROIs can be obtained from a standard atlas (Collins et al. 1995; Evans et al. 2012; Lancaster et al. 1997, 2000; Tzourio-Mazoyer et al., 2002). This approach has two main advantages: i) it enables the comparison between different modalities (Bullmore and Sporns 2009); ii) the number of ROIs is always the same across individuals, such that functional networks can more readily be compared (but see below).
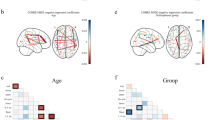
Examples of recent applications of an atlas-based beamformer in combination with functional network analysis. Panel a shows data from a group of 13 healthy controls. The mean alpha band PLI, also known as the weighted degree or node strength in terms of graph theory, for each ROI, is displayed as a color-coded map (thresholded at p = 0.05) on a schematic of the parcellated template brain (modified from Hillebrand et al. 2012). Note that the regions in the occipital lobe are most strongly connected, as can be expected for the alpha band. Panel b shows the connections (PLI) from each ROI to all other ROIs, using an arbitrary threshold. Again, there is a clear pattern of strong connections between regions in the occipital lobe, with additional connections to areas in the temporal and frontal lobes. The upper panel in c shows a similar patterns for data from 17 healthy controls from a different study (Tewarie et al. 2014). This figure displays only the connections that formed part of the MST in the alpha2 band (the colorbar indicates PLI values). Interestingly, there seems to be a shift from an occipital to frontal pattern in healthy controls to a more diffuse pattern in 21 patients with Multiple Sclerosis (lower panel) (modified from Tewarie et al. 2014). Moreover, Tewarie and colleagues showed that this change in network topology correlated with reduced overall cognitive performance
3 Functional Connectivity in Source-Space
The recorded MEG data can be projected through the estimated beamformer weights in order to obtain the time-series for each voxel in the brain (Eq. 1), which are often referred to as virtual electrodes. In order to obtain a single time-series for each ROI, we subsequently select the voxel with maximum power as representative for the ROI. These ROI time-series can then be used as input for functional connectivity analysis. A wide range of functional connectivity estimators are available (Pereda et al. 2005), yet most of these measures are sensitive to the effects of volume conduction and field spread. One could remove these biases before performing connectivity analysis (Brookes et al. 2012; Hipp et al. 2012), or estimate the extent of the bias through simulations (Brookes et al. 2011a). Perhaps more straightforward is the use of measures such as the imaginary part of coherency (Nolte et al. 2004), phase-slope index (Nolte et al. 2008), the Phase Lag Index (PLI; Stam et al. 2007) and related lagged phase synchronization (Pascual-Marqui 2007), as these are inherently insensitive to these biases, where the PLI has the additional advantage that it does not directly depend on the amplitude of the signals (but see Muthukumaraswamy and Singh 2011). These measures have therefore gained popularity in recent years (Canuet et al. 2011, 2012; Guggisberg et al. 2008; Ioannides et al. 2012; Martino et al. 2011; Nolte and Muller 2010; Ponsen et al. 2013; Sekihara et al. 2011; Shahbazi et al. 2012; Tarapore et al. 2012).
The PLI is defined as (Stam et al. 2007):
where Δϕ is the difference between the instantaneous phases for two time-series defined in the interval [−π, π], tk are discrete time-steps, and < > denotes the mean. In short, PLI is a measure for the asymmetry of the distribution of phase differences between two signals, and ranges between 0 and 1. A PLI value of 0 indicates no coupling, coupling with a phase difference of 0 ± nπ radians (with n an integer), or an equal distribution of positive and negative phase differences. Common sources lead to a phase difference of 0 ± nπ radians between two signals, hence the PLI is insensitive to the influence of common sources (Stam et al. 2007). A PLI > 0 is obtained when the distribution of phase differences is asymmetric and is indicative of functional coupling between two signals. Note that PLI does not indicate which of the signals is leading in phase (but see Stam and van Straaten 2012a) and that it potentially discards true interactions with zero-phase lag. Moreover, a value of 0 for uncoupled sources is only achieved for (infinitely) long time-series, hence the PLI is affected by the length of the time-series. PLI also underestimates connectivity between sources with small-lag interactions. A modification of PLI addresses this issue, albeit at the expense of introducing an arbitrary bias favoring large phase differences and mixing of the estimation of consistency of phase differences with the estimation of the magnitude of the phase difference (Vinck et al. 2011).
4 Topology of the Functional Network
Graph theory provides the mathematical framework to characterize the topology of the functional network that is formed by the interacting sources. For this purpose, each ROI is denoted as a vertex (node) and each connection (e.g. the PLI value) is denoted as an edge between the vertices (see Fig. 1). Various graph-theoretical measures can subsequently be used to characterize the network (e.g. Rubinov and Sporns 2010). Two such measures, the clustering coefficientFootnote 2 and the (average shortest) path length,Footnote 3 can be used to explain how the brain can fulfill two seemingly contradictory requirements, namely the processing of information in local functional units on the one hand (‘segregation’) and simultaneous coordination of activity in and between these spatially separated units (‘integration’) (Sporns et al. 2002, 2004). Watts and Strogatz (1998) famously demonstrated with a simple rewiring model that adding a few long distance connections to a network with many local interconnections results in a high clustering yet small average path length. Many large networks, including the brain, have such a so-called small-world configuration (Bassett and Bullmore 2006; Stam 2004). However, this model does not provide a completely satisfactory description of functional brain networks, since it can not explain the occurrence of hubs (Eguiluz et al. 2005). Similarly, the scale-free growth model by Barabasi and Albert (1999), which explains the occurrence of hubs, does not capture the high level of clustering and (hierarchical) modularity observed in experimental data (Meunier et al. 2009). Obviously, we currently lack a model that integrates small-world and scale-free models and fully and elegantly explains the observed functional brain network characteristics (Bullmore and Bassett 2011; Clune et al. 2013; Stam and van Straaten 2012b).
From a practical point of view, although the application of graph theory at the source-level already aids the interpretation of results and the comparability across studies, it is not trivial to compare network topology across individuals, groups, studies or modalities, as was elegantly shown by van Wijk et al. (2010). At the heart of the problem lies the observation that many network properties depend on the size, sparsity (percentage of all possible edges that are present), and the average degree (i.e. the average number of connections per node) of the network. Fixing the number of nodes and average degree in the network (by setting a threshold) does eliminate size effects but may introduce spurious connections or ignore strong connections in the network, and using random surrogates for normalization does not solve this problem either (and may even exuberate it; van Wijk et al. 2010).
A novel approach is to construct the minimum spanning tree (MST) of the original graphs (Boersma et al. 2013; Jackson and Read 2010a, b; Wang et al. 2008). A tree is a sub-graph that does not contain circles or loops and connects all nodes in the original graph, and the MST is the tree that has the minimum total weight (i.e. the sum of all edge valuesFootnote 4) of all possible spanning trees of the original graph. If the original graph contains N nodes than the MST always has N nodes and M = N−1 edges, therefore enabling direct comparison of MSTs between groups and avoiding aforementioned methodological difficulties. Furthermore, if the original network can be interpreted as a kind of transport network, and if edge weights in the original graph possess strong fluctuations, also called the strong disorder limit, then all transport in the original graph flows over the MST (van Mieghem and van Langen 2005), forming the critical backbone of the original graph (van Mieghem and Magdalena 2005; Wang et al. 2008). Interestingly, it seems that for source-reconstructed MEG data for patients with Multiple Sclerosis, as well as for healthy controls, there is a tendency of the weight distribution towards the strong disorder limit (Tewarie et al. 2014). This implies that there is a high probability that the MSTs for both patients and healthy controls can be considered as the critical backbone of the original functional brain networks. Hence, analysis of the minimum spanning tree not only provides a bias free approach to network analysis, but also captures important properties of the original network.
5 Applications in Neurology
5.1 Glioma
In a recent MEG study we revealed a relationship between resting-state functional network properties and protein expression patterns in tumor tissue collected during neurosurgery (Douw et al. 2013). In particular, between-module connectivity was selectively associated with two epilepsy-related proteins, namely synaptic vesicle protein 2A (SV2A) and poly-glycoprotein (P-gp), yet only for the ROIs that contained tumor tissue. Moreover, receiver operator characteristic (ROC) analysis revealed that SV2A expression could be classified with 100 % accuracy on the basis of the between-module connectivity, indicating that the role of the tumor area in the brain network may be an excellent marker for molecular features of brain tissue, which may be used clinically to monitor the efficacy of the anti-epileptic drug levetiracetam (de Groot et al. 2011). Moreover, lower between-module connectivity in the tumor area and higher number of seizures significantly predicted higher P-gp expression, which is in line with previous research showing that high seizure proneness is related to increased P-gp expression (Miller et al. 2008), and suggests that local network topology is an intermediate level between molecular tissue features and clinical patient status. A separate study (van Dellen et al. 2014) examined the link between functional network organization and seizure status further in a longitudinal study. Resting-state MEG recordings were obtained for 20 lesional epilepsy patients at baseline (preoperatively; T0), and at 3–7 (T1) and 9–15 months after resection (T2). Functional connectivity in the lower alpha band correlated positively with seizure frequency at baseline, especially in regions where lesions were located. MST leaf fraction, a measure of integration of information in the network, was significantly increased between T0 and T2, yet only for the seizure-free patients. Moreover, MST-based eccentricity and betweenness centrality, which are measures of node importance and hub-status, decreased between T0 and T2 in seizure free patients, also in regions that were anatomically close to lesion locations and resection cavities. These results demonstrate that there is a link between successful epilepsy surgery and changes in functional network topology. These insights may eventually be utilized for optimization of neurosurgical approaches.
5.2 Parkinson’s Disease
A longitudinal study involving patients with Parkinson’s disease (PD) also revealed a relationship between disease progression and functional brain network topology (Olde Dubbelink et al. 2014). MST analysis revealed a decentralized and less integrated network configuration in early stage untreated PD, which progressed over time. Conventional analysis of clustering and path length also revealed an initial impaired local efficiency, which continued to progress over time, together with reductions in global efficiency. Importantly, these longitudinal changes in network topology were associated with deteriorating motor function and cognitive performance.
6 Future Developments
Excitingly, network analysis, particularly in combination with a standard parcellation of the brain (e.g. through the use of an anatomical atlas), provides a principled way to compare results across different modalities (Bullmore and Sporns 2009). For example, in recent years there has been an insurgence of research into the functional and cognitive relevance of resting-state functional connectivity as determined using fMRI (van den Heuvel and Hulshoff Pol 2010). Although it is already becoming clear that there is a close link between resting-state networks based on hemodynamic phenomena and the underlying electrophysiological networks (e.g. Brookes et al. 2011b; Niu et al. 2012), we envisage that a bias-free network approach allows for an even more accurate integration of these modalities, leading to a better understanding of brain function. Similarly, this approach enables us to directly link the properties of, and dynamics on, functional networks to the topology of the underlying structural network (Guye et al. 2008; Honey et al. 2007). An interesting direction for future work is the study of the interaction between these two types of networks, i.e. to study how functional plasticity affects the structural network, and vice versa (Assenza et al. 2011). Additionally, the same framework can be used to create anatomically and functionally realistic models that can simulate MEG signals. That is, neural mass models can be placed at each location of the anatomical parcellation scheme, where the anatomical connections between the neural masses can be based on experimental DTI data that were obtained for the same atlas. The parameters in these simulated structural/functional networks can subsequently be adjusted in order to test hypotheses (based on observations in experimental data) about disease mechanisms, or to generate new hypotheses about disease effects that we should be able to observe in experimental studies (de Haan et al. 2012; van Dellen et al. 2013).
The atlas-based beamforming approach itself may be developed further in several aspects. We have proposed to use the voxel with maximum power as representative for a ROI, which can introduce some biases, for example for ROIs that cover a large area of cortex. Indeed, the spatial resolution that is obtainable with MEG varies from millimeters to centimeters across the brain (Hillebrand and Barnes 2002), and depends on factors such as location of the neuronal activity, orientation of the cortex and signal-to-noise ratio (Barnes et al. 2004; Hillebrand and Barnes 2002). Our current hypothesis is that the AAL atlas has a resolution that matches the spatial resolution of MEG resting-state data. However, future research should test whether this hypothesis is valid for all cortical regions, for example through the use of atlases with higher spatial resolution (Evans et al. 2012; Seibert and Brewer 2011). In addition, selection of a single representative voxel might be prone to noise and outliers. However, the optimal method of dealing with multiple voxels within a ROI has not been defined yet, and using for instance an averaging method presents other biases, such as introducing artificial differences in signal-to-noise ratios for different sized ROIs. Similarly, one could argue that a priori selection of a target location within a ROI would speed up the computations. However, beamformer reconstructions vary most around peak activations (Barnes and Hillebrand 2003) and as a consequence, the a priori selection of a target voxel could have the effect that the activity for a ROI is completely missed (Barnes et al. 2004).
Another interesting direction for new research is to study the dynamics of functional networks in more detail (de Pasquale et al. 2012), thereby taking advantage of the strongest attribute of MEG, namely its high temporal resolution. A prerequisite is the development of measures of functional connectivity that have high temporal resolution, yet are insensitive to the effects of volume conduction. This would allow us to study functional networks in more detail, and examine the importance of the evolution of functional networks on short time-scales. For example, it was described above that functional brain networks can be divided into modules; are these modules stable over time (Bassett et al. 2013)? And hubs play an important role in the network, is this also reflected in their dynamics, i.e., do hubs evolve differently than non-hubs? Similarly, how does the formation and re-configuration over time of functional networks relate to cognitive performance? Are these dynamics altered in the diseased brain? And if so, is there a phase-transition that distinguishes the healthy from the diseased brain?
Notes
- 1.
In a spherically symmetric volume conductor the magnetic fields produced by the volume currents cancel out exactly (Sarvas 1987), but in a realistically shaped volume conductor there are observable effects of volume conduction.
- 2.
The (unweighted) clustering coefficient denotes the likelihood that neighbours of a node are also connected to each other, and characterizes the tendency of nodes to form local clusters.
- 3.
The average shortest path length is a measure for global integration of the network. It is defined as the harmonic mean of shortest paths between all possible node pairs in the network.
- 4.
For the construction of the MST, the edge weight is defined as 1/(functional connectivity estimate), e.g. 1/PLI.
References
Achard S, Salvador R, Whitcher B, Suckling J, Bullmore E (2006) A resilient, low-frequency, small-world human brain functional network with highly connected association cortical hubs. J Neurosci 26:63–72
Assenza S, Gutierrez R, Gomez-Gardenes J, Latora V, Boccaletti S (2011) Emergence of structural patterns out of synchronization in networks with competitive interactions. Sci Rep 1:99
Baillet S, Mosher JC, Leahy RM (2001) Electromagnetic brain mapping. IEEE Signal Process Mag 18:14–30
Barabasi AL, Albert R (1999) Emergence of scaling in random networks. Science 286:509–512
Barnes GR, Hillebrand A (2003) Statistical flattening of MEG beamformer images. Hum Brain Mapp 18:1–12
Barnes GR, Hillebrand A, Fawcett IP, Singh KD (2004) Realistic spatial sampling for MEG beamformer images. Hum Brain Mapp 23:120–127
Bassett DS, Bullmore E (2006) Small-world brain networks. Neuroscientist 12:512–523
Bassett DS, Porter MA, Wymbs NF, Grafton ST, Carlson JM, Mucha PJ (2013) Robust detection of dynamic community structure in networks. Chaos 23:013142
Boersma M, Smit DJ, Boomsma DI, De Geus EJ, Delemarre-van de Waal HA, Stam CJ (2013) Growing trees in child brains: graph theoretical analysis of electroencephalography-derived minimum spanning tree in 5- and 7-year-old children reflects brain maturation. Brain Connectivity 3: 50–60
Born RT, Bradley DC (2005) Structure and function of visual area MT. Annu Rev Neurosci 28:157–189
Brookes MJ, Hale JR, Zumer JM, Stevenson CM, Francis ST, Barnes GR et al (2011a) Measuring functional connectivity using MEG: methodology and comparison with fcMRI. NeuroImage 56:1082–1104
Brookes MJ, Stevenson CM, Barnes GR, Hillebrand A, Simpson MI, Francis ST et al (2007) Beamformer reconstruction of correlated sources using a modified source model. NeuroImage 34:1454–1465
Brookes MJ, Woolrich M, Luckhoo H, Price D, Hale JR, Stephenson MC et al (2011b) Investigating the electrophysiological basis of resting state networks using magnetoencephalography. Proc Natl Acad Sci USA 108:16783–16788
Brookes MJ, Woolrich MW, Barnes GR (2012) Measuring functional connectivity in MEG: a multivariate approach insensitive to linear source leakage. NeuroImage 63:910–920
Brookes MJ, Gibson AM, Hall SD, Furlong PL, Barnes GR, Hillebrand A et al (2005) GLM-beamformer method demonstrates stationary field, alpha ERD and gamma ERS co-localisation with fMRI BOLD response in visual cortex. NeuroImage 26:302–308
Bullmore E, Sporns O (2009) Complex brain networks: graph theoretical analysis of structural and functional systems. Nat Rev Neurosci 10:186–198
Bullmore E, Sporns O (2012) The economy of brain network organization. Nat Rev Neurosci 13:336–349
Bullmore ET, Bassett DS (2011) Brain graphs: graphical models of the human brain connectome. Annu Rev Clin Psychol 7:113–140
Buzsaki G, Wang XJ (2012) Mechanisms of gamma oscillations. Annu Rev Neurosci 35:203–225
Canuet L, Ishii R, Pascual-Marqui RD, Iwase M, Kurimoto R, Aoki Y et al (2011) Resting-state EEG source localization and functional connectivity in schizophrenia-like psychosis of epilepsy. PLoS ONE 6:e27863
Canuet L, Tellado I, Couceiro V, Fraile C, Fernandez-Novoa L, Ishii R et al (2012) Resting-state network disruption and APOE genotype in Alzheimer’s disease: a lagged functional connectivity study. PLoS ONE 7:e46289
Cheyne D, Bakhtazad L, Gaetz W (2006) Spatiotemporal mapping of cortical activity accompanying voluntary movements using an event-related beamforming approach. Hum Brain Mapp 27:213–229
Clune J, Mouret JB, Lipson H (2013) The evolutionary origins of modularity. Proc R Soc B: Biol Sci 280:20122863
Collins DL, Holmes CJ, Peters TM, Evans AC (1995) Automatic 3-D model-based neuroanatomical segmentation. Hum Brain Mapp 3:190–208
Dalal SS, Guggisberg AG, Edwards E, Sekihara K, Findlay AM, Canolty RT et al (2008) Five-dimensional neuroimaging: localization of the time-frequency dynamics of cortical activity. Neuroimage 40:1686–1700
Dalal SS, Sekihara K, Nagarajan SS (2006) Modified beamformers for coherent source region suppression. IEEE Trans Biomed Eng 53:1357–1363
de Groot M, Aronica E, Heimans JJ, Reijneveld JC (2011) Synaptic vesicle protein 2A predicts response to levetiracetam in patients with glioma. Neurology 77:532–539
de Haan W, Mott K, van Straaten EC, Scheltens P, Stam CJ (2012) Activity dependent degeneration explains hub vulnerability in Alzheimer’s disease. PLoS Comput Biol 8:e1002582
de Pasquale F, Della PS, Snyder AZ, Marzetti L, Pizzella V, Romani GL et al (2012) A cortical core for dynamic integration of functional networks in the resting human brain. Neuron 74:753–764
Diwakar M, Tal O, Liu TT, Harrington DL, Srinivasan R, Muzzatti L et al (2011) Accurate reconstruction of temporal correlation for neuronal sources using the enhanced dual-core MEG beamformer. NeuroImage 56:1918–1928
Douw L, de Groot M, van Dellen E, Aronica E, Heimans JJ, Klein M et al (2013) Local MEG networks: the missing link between protein expression and epilepsy in glioma patients? NeuroImage 75:203–211
Eguiluz VM, Chialvo DR, Cecchi GA, Baliki M, Apkarian AV (2005) Scale-free brain functional networks. Phys Rev Lett 94:018102
Engel AK, Fries P, Singer W (2001) Dynamic predictions: oscillations and synchrony in top-down processing. Nat Rev Neurosci 2:704–716
Evans AC, Janke AL, Collins DL, Baillet S (2012) Brain templates and atlases. NeuroImage 62:911–922
Fries P (2005) A mechanism for cognitive dynamics: neuronal communication through neuronal coherence. Trends Cogn Sci 9:474–480
Grodzinsky Y (2000) The neurology of syntax: language use without Broca’s area. Behav Brain Sci 23:1–21
Guggisberg AG, Honma SM, Findlay AM, Dalal SS, Kirsch HE, Berger MS et al (2008) Mapping functional connectivity in patients with brain lesions. Ann Neurol 63:193–203
Guye M, Bartolomei F, Ranjeva JP (2008) Imaging structural and functional connectivity: towards a unified definition of human brain organization? Curr Opin Neurol 21:393–403
Hadjipapas A, Hillebrand A, Holliday IE, Singh KD, Barnes GR (2005) Assessing interactions of linear and nonlinear neuronal sources using MEG beamformers: a proof of concept. Clin Neurophysiol 116:1300–1313
Hamalainen MS, Ilmoniemi RJ (1994) Interpreting magnetic fields of the brain: minimum norm estimates. Med Biol Eng Compu 32:35–42
Hillebrand A, Barnes GR (2011) Practical constraints on estimation of source extent with MEG beamformers. NeuroImage 54:2732–2740
Hillebrand A, Barnes GR (2002) A quantitative assessment of the sensitivity of whole-head meg to activity in the adult human cortex. NeuroImage 16:638–650
Hillebrand A, Barnes GR, Bosboom JL, Berendse HW, Stam CJ (2012) Frequency-dependent functional connectivity within resting-state networks: an atlas-based MEG beamformer solution. NeuroImage 59:3909–3921
Hillebrand A, Barnes GR (2003) The use of anatomical constraints with MEG beamformers. NeuroImage 20:2302–2313
Hillebrand A, Barnes GR (2005) Beamformer analysis of MEG data. Int Rev Neurobiol (Special volume on Magnetoencephalography) 68:149–171
Hillebrand A, Singh KD, Holliday IE, Furlong PL, Barnes GR (2005) A new approach to neuroimaging with magnetoencephalography. Hum Brain Mapp 25:199–211
Hipp JF, Hawellek DJ, Corbetta M, Siegel M, Engel AK (2012) Large-scale cortical correlation structure of spontaneous oscillatory activity. Nat Neurosci 15:884–890
Honey CJ, Kotter R, Breakspear M, Sporns O (2007) Network structure of cerebral cortex shapes functional connectivity on multiple time scales. Proc Natl Acad Sci USA 104:10240–10245
Hui HB, Pantazis D, Bressler SL, Leahy RM (2010) Identifying true cortical interactions in MEG using the nulling beamformer. NeuroImage 49:3161–3174
Ioannides AA, Dimitriadis SI, Saridis GA, Voultsidou M, Poghosyan V, Liu L et al (2012) Source space analysis of event-related dynamic reorganization of brain networks. Comput Math Methods Med 2012:452503
Jackson TS, Read N (2010a) Theory of minimum spanning trees. I. Mean-field theory and strongly disordered spin-glass model. Phys Rev E: Stat, Nonlin, Soft Matter Phys 81:021130
Jackson TS, Read N (2010b) Theory of minimum spanning trees. II. Exact graphical methods and perturbation expansion at the percolation threshold. Phys Rev E: Stat, Nonlin, Soft Matter Phys 81:021131
Lancaster JL, Rainey LH, Summerlin JL, Freitas CS, Fox PT, Evans AC et al (1997) Automated labeling of the human brain: a preliminary report on the development and evaluation of a forward-transform method. Hum Brain Mapp 5:238–242
Lancaster JL, Woldorff MG, Parsons LM, Liotti M, Freitas CS, Rainey L et al (2000) Automated Talairach atlas labels for functional brain mapping. Hum Brain Mapp 10:120–131
Lü ZL, Williamson SJ (1991) Spatial extent of coherent sensory-evoked cortical activity. Exp Brain Res 84:411–416
Martino J, Honma SM, Findlay AM, Guggisberg AG, Owen JP, Kirsch HE et al (2011) Resting functional connectivity in patients with brain tumors in eloquent areas. Ann Neurol 69:521–532
Meunier D, Lambiotte R, Fornito A, Ersche KD, Bullmore ET (2009) Hierarchical modularity in human brain functional networks. Frontiers Neuroinfo 3:37
Miller DS, Bauer B, Hartz AM (2008) Modulation of P-glycoprotein at the blood-brain barrier: opportunities to improve central nervous system pharmacotherapy. Pharmacol Rev 60:196–209
Muthukumaraswamy SD, Singh KD (2011) A cautionary note on the interpretation of phase-locking estimates with concurrent changes in power. Clin Neurophysiol 122:2324–2325
Niu H, Wang J, Zhao T, Shu N, He Y (2012) Revealing topological organization of human brain functional networks with resting-state functional near infrared spectroscopy. PLoS ONE 7:e45771
Nolte G, Bai O, Wheaton L, Mari Z, Vorbach S, Hallett M (2004) Identifying true brain interaction from EEG data using the imaginary part of coherency. Clin Neurophysiol 115:2292–2307
Nolte G, Muller KR (2010) Localizing and estimating causal relations of interacting brain rhythms. Frontiers Hum Neurosci 4:209
Nolte G, Ziehe A, Nikulin VV, Schlogl A, Kramer N, Brismar T et al (2008) Robustly estimating the flow direction of information in complex physical systems. Phys Rev Lett 100:234101
Olde Dubbelink KTE, Hillebrand A, Stoffers D, Deijen JB, Twisk JWR, Stam CJ et al (2014) Disrupted brain network topology in Parkinson’s disease: a longitudinal magnetoencephalography study. Brain 137:197–207
Pascual-Marqui RD (2007) Instantaneous and lagged measurements of linear and nonlinear dependence between groups of multivariate time series: frequency decomposition. arXiv:0711.1455
Pereda E, Quiroga RQ, Bhattacharya J (2005) Nonlinear multivariate analysis of neurophysiological signals. Prog Neurobiol 77:1–37
Ponsen MM, Stam CJ, Bosboom JLW, Berendse HW, Hillebrand A (2013) A three dimensional anatomical view of oscillatory resting-state activity and functional connectivity in Parkinson’s disease related dementia: an MEG study using atlas-based beamforming. NeuroImage: Clin 2:95–102
Quraan MA, Cheyne D (2010) Reconstruction of correlated brain activity with adaptive spatial filters in MEG. NeuroImage 49:2387–2400
Reijneveld JC, Ponten SC, Berendse HW, Stam CJ (2007) The application of graph theoretical analysis to complex networks in the brain. Clin Neurophysiol 118:2317–2331
Robinson SE, Vrba J (1999) Functional Neuroimaging by Synthetic Aperture Magnetometry (SAM). In: Kotani M, Kuriki S, Karibe H, Nakasato N, Yoshimoto T (eds) Recent advances in biomagnetism. Tohoku Univ Press, Sendai, pp 302–305
Rubinov M, Sporns O (2010) Complex network measures of brain connectivity: uses and interpretations. NeuroImage 52:1059–1069
Sarvas J (1987) Basic mathematical and electromagnetic concepts of the biomagnetic inverse problem. Phys Med Biol 32:11–22
Schnitzler A, Gross J (2005) Normal and pathological oscillatory communication in the brain. Nat Rev Neurosci 6:285–296
Schoffelen JM, Gross J (2009) Source connectivity analysis with MEG and EEG. Hum Brain Mapp 30:1857–1865
Sekihara K, Nagarajan SS, Poeppel D, Marantz A (2004) Asymptotic SNR of scalar and vector minimum-variance beamformers for neuromagnetic source reconstruction. IEEE Trans Biomed Eng 51:1726–1734
Seibert TM, Brewer JB (2011) Default network correlations analyzed on native surfaces. J Neurosci Methods 198:301–311
Sekihara K, Owen JP, Trisno S, Nagarajan SS (2011) Removal of spurious coherence in MEG source-space coherence analysis. IEEE Trans Biomed Eng 58:3121–3129
Sekihara K, Nagarajan SS (2008) Adaptive Spatial Filters for Electromagnetic Brain Imaging. Springer, Heidelberg
Shahbazi AF, Ewald A, Nolte G (2012) Localizing true brain interactions from EEG and MEG data with subspace methods and modified beamformers. Comput Math Methods Med 2012:402341
Singer W (1999) Neuronal synchrony: a versatile code for the definition of relations? Neuron 24:49–25
Singh KD, Barnes GR, Hillebrand A (2003) Group imaging of task-related changes in cortical synchronisation using non-parametric permutation testing. NeuroImage 19:1589–1601
Singh KD, Barnes GR, Hillebrand A, Forde EME, Williams AL (2002) Task-related changes in cortical synchronization are spatially coincident with the hemodynamic response. NeuroImage 16:103–114
Smit DJ, Boersma M, Schnack HG, Micheloyannis S, Boomsma DI, Hulshoff Pol HE et al (2012) The brain matures with stronger functional connectivity and decreased randomness of its network. PLoS ONE 7:e36896
Smit DJ, Boersma M, van Beijsterveldt CE, Posthuma D, Boomsma DI, Stam CJ et al (2010) Endophenotypes in a dynamically connected brain. Behav Genet 40:167–177
Smit DJ, Stam CJ, Posthuma D, Boomsma DI, De Geus EJ (2008) Heritability of “small-world” networks in the brain: a graph theoretical analysis of resting-state EEG functional connectivity. Hum Brain Mapp 29:1368–1378
Sporns O, Chialvo DR, Kaiser M, Hilgetag CC (2004) Organization, development and function of complex brain networks. Trends Cogn Sci 8:418–425
Sporns O, Tononi G, Edelman GM (2002) Theoretical neuroanatomy and the connectivity of the cerebral cortex. Behav Brain Res 135:69–74
Stam CJ (2004) Functional connectivity patterns of human magnetoencephalographic recordings: a ‘small-world’ network? Neurosci Lett 355:25–28
Stam CJ, Nolte G, Daffertshofer A (2007) Phase lag index: assessment of functional connectivity from multi channel EEG and MEG with diminished bias from common sources. Hum Brain Mapp 28:1178–1193
Stam CJ, Reijneveld JC (2007) Graph theoretical analysis of complex networks in the brain. Nonlinear Biomed Phys 1:3
Stam CJ, van Straaten EC (2012a) The organization of physiological brain networks. Clin Neurophysiol 123:1067–1087
Stam CJ, van Straaten EC (2012b) Go with the flow: use of a directed phase lag index (dPLI) to characterize patterns of phase relations in a large-scale model of brain dynamics. NeuroImage 62:1415–1428
Supek S, Aine CJ (1993) Simulation studies of multiple dipole neuromagnetic source localization: model order and limits of source resolution. IEEE Trans Biomed Eng 40:529–540
Tarapore PE, Martino J, Guggisberg AG, Owen J, Honma SM, Findlay A et al (2012) Magnetoencephalographic imaging of resting-state functional connectivity predicts postsurgical neurological outcome in brain gliomas. Neurosurgery 71:1012–1022
Tewarie P, Hillebrand A, Schoonheim MM, van Dijk BW, Geurts JJ, Barkhof F et al (2014) Functional brain network analysis using minimum spanning trees in Multiple Sclerosis: an MEG source-space study. NeuroImage 88:308–318
Tian L, Wang J, Yan C, He Y (2011) Hemisphere- and gender-related differences in small-world brain networks: a resting-state functional MRI study. NeuroImage 54:191–202
Tononi G, Edelman GM (1998) Consciousness and complexity. Science 282:1846–1851
Tononi G, Edelman GM, Sporns O (1998) Complexity and coherency: integrating information in the brain. Trends Cogn Sci 2:474–484
Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N et al (2002) Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. NeuroImage 15:273–289
van Dellen E, Hillebrand A, Douw L, Heimans JJ, Reijneveld JC, Stam CJ (2013) Local polymorphic delta activity in cortical lesions causes global decreases in functional connectivity. NeuroImage 83:524–532
van Dellen E, Douw L, Hillebrand A, de Witt Hamer PC, Baayen JC, Heimans JJ et al (2014) Epilepsy surgery outcome and functional network alterations in longitudinal MEG: a minimum spanning tree analysis. NeuroImage 86:354–363
van den Heuvel MP, Hulshoff Pol HE (2010) Exploring the brain network: a review on resting-state fMRI functional connectivity. Eur Neuropsychopharmacol 20:519–534
van den Heuvel MP, Kahn RS, Goni J, Sporns O (2012) High-cost, high-capacity backbone for global brain communication. Proc Natl Acad Sci USA 109:11372–11377
van den Heuvel MP, Sporns O (2011) Rich-club organization of the human connectome. J Neurosci 31:15775–15786
van Mieghem P, Magdalena SM (2005) Phase transition in the link weight structure of networks. Phys Rev E: Stat, Nonlin, Soft Matter Phys 72:056138
van Mieghem P, van Langen S (2005) Influence of the link weight structure on the shortest path. Phys Rev E: Stat, Nonlin, Soft Matter Phys 71:056113
van Straaten EC, Stam CJ (2013) Structure out of chaos: Functional brain network analysis with EEG, MEG, and functional MRI. Eur Neuropsychopharmacol 23:7–18
van Veen BD, van Drongelen W, Yuchtman M, Suzuki A (1997) Localization of brain electrical activity via linearly constrained minimum variance spatial filtering. IEEE Trans Biomed Eng 44:867–880
van Wijk BC, Stam CJ, Daffertshofer A (2010) Comparing brain networks of different size and connectivity density using graph theory. PLoS ONE 5:e13701
Varela F, Lachaux J-P, Rodriguez E, Martinerie J (2001) The brainweb: phase synchronization and large-scale integration. Nat Rev Neurosci 2:229–239
Vinck M, Oostenveld R, van Wingerden M, Battaglia F, Pennartz CM (2011) An improved index of phase-synchronization for electrophysiological data in the presence of volume-conduction, noise and sample-size bias. NeuroImage 55:1548–1565
Vrba J (2002) Magnetoencephalography: the art of finding a needle in a haystack. Physica C 368:1–9
Wang H, Hernandez JM, van Mieghem P (2008) Betweenness centrality in a weighted network. Phys Rev E: Stat, Nonlin, Soft Matter Phys 77:046105
Watts DJ, Strogatz SH (1998) Collective dynamics of ‘small-world’ networks. Nature 393:440–442
Wipf D, Nagarajan S (2009) A unified Bayesian framework for MEG/EEG source imaging. NeuroImage 44:947–966
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Hillebrand, A., Stam, C.J. (2014). Recent Developments in MEG Network Analysis. In: Supek, S., Aine, C. (eds) Magnetoencephalography. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-33045-2_12
Download citation
DOI: https://doi.org/10.1007/978-3-642-33045-2_12
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-33044-5
Online ISBN: 978-3-642-33045-2
eBook Packages: EngineeringEngineering (R0)