Abstract
Orthodontics developed as a specialty of dentistry almost a century ago. Historical proofs clearly demonstrate that the history of orthodontics extends back to the ancient years. It seems that man very early realized the need for orthodontic treatment in order to accomplish correct function and to improve esthetics of the stomatognathic system and more importantly of the whole face.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Periodontal Ligament
- Metabolic Bone Disease
- Osteoclast Precursor
- Mature Osteoclast
- Orthodontic Tooth Movement
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Orthodontics developed as a specialty of dentistry almost a century ago. Historical proofs clearly demonstrate that the history of orthodontics extends back to the ancient years. It seems that man very early realized the need for orthodontic treatment in order to accomplish correct function and to improve esthetics of the stomatognathic system and more importantly of the whole face.
The great Edward Angle, the father of modern orthodontics, categorized malocclusion and introduced his treatment principles. These principles were subsequently improved by other great figures in orthodontics, Charles Tweed, Raymond Begg, Joseph Jarabak, and Robert Strang, just to mention a few.
Orthodontic treatment at the beginning was more or less empirical, focusing mainly on the technicalities of tooth movement. Scientific evidence in the form of histological findings and tissue reactions to orthodontic tooth movement was developed later with the classic work of Kaare Reitan from Norway, who introduced the tension and compression theory based on histological findings [1, 2].
In Europe, orthodontics took a different route, that of dentofacial orthopedics. Later, the concept of orthopedic treatment of dentofacial anomalies found substantiation in Moss’s functional matrix theory [3]. Function affects the structure and form of the jaws and face, and treatment based on a functional approach helps correct certain forms of skeletal anomalies. This orthopedic approach found its scientific support in the classical histologic work of James McNamara. The scientific impact of this work was extreme, and its publication was the turning point for orthodontics worldwide.
One of the most important researchers, teachers, and visionaries in the history of orthodontics and at that time the editor of the prestigious American Journal of Orthodontics, Thomas M. Graber, to whom this book is dedicated, added the words Dentofacial Orthopedics to the title of the journal, putting orthodontics on a route toward the twenty-first century.
In recent years, the field of orthodontics and dentofacial orthopedics follows closely the scientific advances in medical biology, mainly bone biology. Many complicated biochemical techniques are now being used in order to identify specific tissue reactions to orthodontic tooth movement, or more clearly to force-induced alveolar bone remodeling. Moreover, histology and molecular biology provided us with the tools to identify the biological events that follow the application of external mechanical stimulation/loading to alveolar bone and cartilage tissue. Complete elucidation of the biochemical bone tissue response will greatly improve our diagnostics, treatment planning, and outcome.
In the following sections, the main histological and histochemical protocols, as well as the major osteoblast and osteoclast cell tissue techniques, are presented. In addition, techniques and systems for external mechanical force application are described.
2 Histological Methods
2.1 Decalcification
The study of bone and cartilage cell morphology is of paramount importance for understanding the function of these cells. However, the physical rigidity of these tissues poses significant difficulties for the cutting of sections. These difficulties are mainly due to the intimate mixture of hard (bone and teeth) and soft tissues (osteoid, cartilage, and fat) within the same biopsy sample [4]. In order to obtain adequate sections, the embedded material should undergo a specific softening procedure, called decalcification. A more accurate term for this process should be demineralization since, except for calcium, other minerals are also removed. In general, decalcification methods are divided into two categories: acid (mainly nitric, hydrochloric, and formic) and neutral (ethylenediaminetetraacetic acid—EDTA). Mineral (nitric and hydrochloric) and organic (formic) acids are preferable for routine decalcification because they remove large quantities of calcium at a rapid rate. However, they may damage cellular morphology, and therefore they are not recommended for small samples of hard tissue. On the other hand, neutral decalcification is the method of choice for small quantities of tissue since it preserves perfectly the cellular characteristics. Nevertheless, it penetrates tissue very slowly and is comparatively expensive when large amounts are used. A series of studies have shown that EDTA decalcification preserves proteins and nucleic acids for immunohistochemical, FISH, ISH, and CGH analyses [4–6]. However, some investigators believe that this procedure reduces enzyme activity and affects DNA and RNA function [7–9]. Therefore, they suggest that fresh-frozen specimens must be processed in an undecalcified way and sectioned with technologically advanced cryotomes (such as the CryoJane® Tape-Transfer System), which unfortunately are not available in most histology laboratories [10, 11].
2.2 Histochemical Methods
Hematoxylin and eosin (H&E) stain is an excellent method for visualizing the cell nucleus and cytoplasm, especially after tissue decalcification. The hematoxylin solution highlights nuclei, whereas eosin is bound on proteins and thus stains primarily the cytoplasm (Fig. 8.1). However, the study of bone biology requires more accurate detection and characterization of the cells involved in bone metabolism. Therefore, special histochemical stains have been developed.
Cells of mononuclear origin express the band 5 isoenzymes of tartrate-resistant acid phosphatase (TRAP) [12, 13]. This enzyme is characterized by cathodal electrophoretic mobility at pH 4 and by resistance to inhibition by L(+)-tartate [14]. In mammals, TRAP has been detected in several tissue systems as a minor acid phosphatase isoenzyme [15, 16]. Nonetheless, it is primarily expressed in bone-resorbing multinucleated osteoclasts of the skeleton [12, 17]. The function of TRAP remains obscure. Several studies have proposed that in resorbing osteoclasts, TRAP is localized in the ruffled border or in the Howship lacunae [18]. However, in vitro and immunoelectron microscopy studies have documented that TRAP is also located in large transcytotic vesicles [19, 20]. Therefore, degradation of extracellular matrix proteins (namely, bone sialoproteins, osteopontin, and osteonectin) occurs in both the Howship lacunae and the intracellular transcytotic vesicles [21]. TRAP can function as an excellent, highly specific osteoclast marker, which can be easily detected by commercially available histochemical kits. The histochemical TRAP staining results can be evaluated under light microscopy. The TRAP-positive osteoclast surface and the TRAP-positive osteoclast number can be calculated either manually or with the use of proper software [22, 23]. This stain can be applied on both decalcified and non-decalcified tissues [12, 17, 23]. The development of more sophisticated TRAP protocols, such as fluorescent-based TRAP stains, holds promise for better visual results and can be combined with other immunofluorescent, as well as immunohistochemical methods [24]. Example of TRAP labeling is shown in Figs. 8.2 and 8.3.
Higher-magnification micrograph of area shown in Fig. 8.1, highlighting TRAP-positive osteoclasts (original magnification ×20)
A commonly used marker of osteogenic development is alkaline phosphatase, an enzyme that was discovered by Robison in 1923 [25]. The term ALP was subsequently introduced in 1979 by McComb and colleagues [26]. Four distinct genes that encode for 4 different ALP isoenzymes have been discovered in humans: intestinal, placental germ-like, and tissue-non-specific (TNS). TNSALP is expressed in liver, bone, and kidney [27, 28]. The biological role of this protein is largely unknown. Within bone tissue and teeth, ALP is produced by osteoblasts and is involved in the process of osteoid mineralization [29]. This function is facilitated by local elevation of inorganic phosphate and destruction of inhibitors of hydroxyapatite crystal growth, phosphate transportation, and ATPase or tyrosine-specific phosphoprotein phosphatase activity [30]. Numerous studies of bone and cartilage development have highlighted the importance of measurement of ALP for the evaluation of osteoblastic activity. The identification of ALP activity in tissue sections is made primarily by histochemical methods [31, 32]. These methods are applied mainly to non-decalcified, frozen bone, and cartilage tissues. Indeed, ALP is sensitive to decalcification procedures since they remove the zinc and magnesium ions that are essential for ALP activity [31]. ALP is expressed in stimulated osteoblasts, bone-lining cells, and some newly formed osteocytes as well as in pre-apoptotic chondroblasts. Recently, histochemical methods that can be applied to decalcified, paraffin-embedded skeletal tissue have been developed [33, 34]. These methods are relatively easy, cheap, and reproducible and can be used in conventional pathology/histology laboratories.
In addition to histochemisty, immunohistochemical methods have been developed for the detection of ALP activity and localization [32, 35], using polyclonal antibodies against TNSAP or tissue-specific monoclonal antibodies against the bone isoform [32, 36, 37]. Histochemical and immunohistochemical approaches provide an in situ estimation of ALP localization and function, as they reveal differential localization of the examined enzyme during the different steps of bone and cartilage development and maturation.
Another commonly used assay for the study of bone maturation and mineralization was described in 1901 by von Kossa [38]. An example of this assay is shown in Fig. 8.4.
The von Kossa assay is an excellent method for the detection of calcium depositions. Notably, this stain does not react with calcium but with phosphate and carbonate ions in the presence of acid material [39]. More specifically, this method is based upon the principle that cationic silver ions can be removed from solution by carbonate or phosphate ions because of their respective positions in the electrochemical series. When undecalcified tissue sections are treated with 5 % silver nitrate solution, cationic silver replaces calcium in the original salt and forms a silver salt that can be displayed by a reduction to metallic silver. This reaction is photochemical, and the activation energy is supplied from strong violet or ultraviolet light [40, 41]. Silver ions are associated with phosphate ions, and therefore they are considered to indirectly uncover calcium deposits. After the treatment with silver nitrate and aqueous sodium thiosulfate, counterstaining is required. For this purpose, van Gieson and hematoxylin-eosin stains are recommended. It can be seen in Fig. 8.4 that when the von Kossa assay is completed, sites of calcium deposition are stained black, and the osteoid and collagen are stained red, whereas fibrous tissue and red blood cells are stained yellow.
In addition, the H&E stain highlights the cellular components of the examined sections, providing important information regarding the histology of the examined tissues. The von Kossa assay is a very simple, accurate, and inexpensive technique for the study of bone maturation and mineralization. Furthermore, von Kossa-stained sections can be used for histomorphometric analyses. The purpose of histomorphometry is the evaluation of the structural integrity of the skeleton, the degree of bone formation and mineralization, and the rate of bone resorption. The tested parameters that reflect skeleton structural integrity are the total bone volume, the volume of cancellous bone, and the amount of trabecular osteoid. The parameters that are associated with bone formation and mineralization are the surface of the trabecular osteoid, the surface of mineralization, the distance between two tetracycline-pulse labels per day (see Sect. 8.2.3), and the mineralization lag time. Finally, the factors that indicate osteoclast-resorption function are: the trabecular, cortical, and periosteal resorptive surfaces; the trabecular osteoclast count (number of osteoclasts per area); and the cortical porosity (percentage of the cortex that contains pores without osteoclasts) [42, 43]. Histomorphometry is a method of choice for the study of conditions such as metabolic bone diseases, neoplasias, bone remodeling, and fracture repair, as well as bone-cartilage response to biomechanical stress [43].
2.3 Fluorescent Labeling
Bone tissue continually undergoes shape and structure changes. Accretions in length and thickness, modeling, and drift activities lead to morphologic alterations that determine the relative position between various skeletal parts. Metabolic bone diseases, bone repair processes, mechanical stress, and aging create new functional demands and are responsible for the subsequent structural adaptations. An accurate method for the detection of such structural/functional modifications is fluorescent labeling [44]. Fluorescent double-labeling is used to calculate kinetic data on bone turnover. The fluorochromes are administrated systematically and form long-lasting chelate complexes with apatite, via their active iminodiacetic acid groups. Hence, they can serve as markers that allow the identification of mineralized tissues [45]. Different types of fluorochromes, such as yellow tetracyclines, xylenol orange, alizarin red derivatives, or green fluorescein derivatives like calcein (Fig. 8.5) or DCAF, which produce different colors, are available [46, 47]. The first dose of fluorescent dye is incorporated in the newly formed bone at the bone-osteoid interface, where it appears as a linear fluorescence under UV light microscopy. The second dose is administrated 3–14 days after the first. The amount of bone that has been synthesized during this time period can be calculated by measuring the width between the two lines of fluorescence. Dosing of the tetracycline is dependent upon the model individual, such as human, rat, or rabbit.
2.4 Immunohistochemistry (IHC)
Immunohistochemistry is used in everyday practice at pathology and histology laboratories. It is a relatively simple method for the in situ detection of proteins. IHC is based on the principle that specific intra- or extracellular antigens are bound to monoclonal or polyclonal antibodies that are associated with specific enzymes. The detection of the antigen-antibody complex is achieved with the use of chromogens. The principal enzyme that facilitates the antibody detection is peroxydase, and the chromogen that is most commonly used is 3,3′-diaminobenzidine tetrahydrochloride (DAB). There are two major categories of IHC methods: direct and indirect. Indirect IHC methods include peroxydase-antiperoxydase (PAP), avidin-biotin complex (ABC), and biotin-streptavidin assay (B-SA), which is the most popular [48]. The B-SA IHC method relies on the non-immunologic binding of biotin to streptavidin (a 60-kD protein), produced by Streptomyces avidinii. Three reagents are used: (a) the primary antibody, which is specific for the antigen of interest; (b) the secondary (biotinylated) antibody that binds the first one; and (c) the streptavidin-peroxidase reagent that is associated to the secondary antibody. IHC provides an in situ approach to investigate the expression and activation status of the examined proteins.
Furthermore, this is a useful method for bone biology studies since it detects the expression levels of several proteins implicated in bone development and growth. Among them, osteonectin (ON) and osteocalcin (OC) are the most significant. Examples are shown in Figs. 8.6 and 8.7. ON is a 35–45-kD protein that has the ability to bind to Ca+2, hydroxyapatite, and collagen [49]. Among its structural features, the two EF-hand high-affinity calcium-binding sites are functionally the most significant. ON mediates the deposition of hydroxyapatite and is involved in the regulation of the osteoblastic cell cycle and bone maturation. OC (also called bone gla protein) is a 5-kD protein that belongs to a family of extracellular matrix proteins named gla proteins. OC possesses one disulfide bridge, and the gla residues are located in α helical region. A large volume of in vitro and in vivo studies have documented that OC plays a central role in bone remodeling and skeletal development. More specifically, it activates osteoclasts and recruits their precursors, determining the transition from bone resorption to bone formation [50–53]. Immunohistochemical detection of ON characterizes early stages of bone development, whereas OC typically determines later steps of skeletal growth and osteoid mineralization. Other proteins such as Cbfa1/Runx2 and AP-1 transcription factors have also been found to participate in chondroblastic/osteoblastic differentiation and maturation [54, 55]. Very recent IHC data on rat TMJs have shown that these proteins are selectively expressed in bone and cartilage tissue and that their differential expression highlights different maturation levels during the process of chondro-osteogenesis [56, 57]. Therefore, they support the notion that Runx2 and AP-1 (c-Jun/c-Fos heterodimer) can be used as bone/cartilage markers and indicators of chondro-osteoblastic maturation.
Same tumor shown in Fig. 8.6, now immunostained for bone-specific marker osteocalcin (original magnification ×10)
2.5 In Situ Hybridization
In situ hybridization (ISH) is a valuable molecular method in the field of bone research and diagnosis since it detects the localization of specific nucleic acids at the level of individual cells or complex tissue sections, combining histochemistry with recombinant DNA technology [58–60]. It was first described in 1969 and is based on the specific binding of a labeled nucleotide probe to target DNA or RNA sequences [61, 62]. Probes for ISH (double-stranded DNA, single-stranded antisense RNA, single-stranded DNA probes generated by polymerase chain reaction procedure, synthetic oligodeoxynucleotides, or oligoprobes) are usually 50–300 bases long. Originally, they were labeled with radioisotopes that limited ISH utility for research and diagnostic purposes [60]. Nonetheless, the introduction of non-isotopic labels and development of detection methods based on classical histochemical and immunohistochemical assays reduced the background and improved the signal resolution, expanding the range of ISH applications. In the fields of orthopedic and orthodontic research, ISH can be applicable for both cytogenetic and archival preparations [23, 63–67]. A key advantage of histologic sections is that the examined cells are evaluated in their native architecture and localization. This is often essential, for example, in the study of conditions such as response to stress or metabolic bone diseases, where differential localization of cells with distinct molecular and biochemical properties determines the degree and the quality of bone growth. Furthermore, since locus-specific ISH can be detected by nonfluorescence reagents, it can be easily visualized with bright-field microscopy. The role of fixation is of great importance for ISH since it ensures the integrity of the nucleic acids and the preservation of tissue morphology. Cross-linking fixatives such as formalin and glutaraldehyde seem to provide the best results [68–70]. The degree of DNA/RNA damage caused by decalcification procedures is controversial [64, 65]. However, the combination of EDTA decalcification with formalin fixation seems to be efficient. After fixation and decalcification, unstained sections are transferred onto coated glass slides with advanced adherence properties that prevent tissue floating during the ISH process. Afterward, tissues are treated with pepsin and proteinase K that digest cellular proteins and facilitate probe access to targeted nucleic acids. Optimization of tissue preparation is a major technical challenge. The optimal protein digestion conditions are basically determined empirically for each tissue and probe combination. In order to evaluate whether the signal visualized by ISH is specific for the target DNA or RNA sequence, the use of controls (hybridization of samples from the same tissue with a probe complementary to the assessed probe and identical to the targeted sequence that does not generate any signal) is essential.
The applications of ISH in the areas of bone and dental research are numerous. For instance, it can be performed in order to determine the spatial and temporal distributions of specific mRNA sequences, especially in cases where the gene products are below the threshold of IHC detection. Indeed, by using ISH, the role of genes (namely, Ihh, PTHrP, TRAP, OC, ON, OPN, collagen type II, and Runx1, -2, -3) that are involved in cartilage-bone growth, development, and remodeling has been investigated. Furthermore, ISH has proved to be a valuable tool in the study of pathologic conditions such as metabolic and inflammatory bone and teeth diseases and neoplasias [23, 63, 66, 67, 71, 72]. In situ end-labeling (ISEL) is an ISH-related method that is performed on formalin-fixed, paraffin-embedded tissues for the identification of cells that undergo programmed cell death. ISEL detects the presence of DNA strand breaks that are generated by activated endogenous nucleases during apoptosis [73–75]. More specifically, in the presence of DNA polymerase, the DNA strand breaks are hybridized with non-isotopic reporter molecules, which can be detected with IHC methods. The ISEL assay can be applied as a corollary to the TdT-mediated dUTP-dioxigenin nick-end-labeling (TUNEL) method, which specifically labels the 3′-hydroxyl terminal of DNA strand breaks. Apoptotic cells are recognized by their dark nuclei (TUNEL-positive reaction). During the process of endochondral bone formation, chondrocyts and osteocytes progressively mature and undergo programmed cell death. Osteoclasts are also susceptible to apoptosis in the absence of trophic and growth-stimulating factors, such as M-CSF and RANKL [75, 76]. Obviously, skeletal conditions that induce apoptosis (such as mechanical or biochemical stress and inflammatory or neoplastic diseases) are under intense scrutiny. ISEL and TUNEL are the methods of choice for the investigation of these apoptotic phenomena.
3 Polymerase Chain Reaction (PCR) and Reverse Transcriptase PCR (RT-PCR)
The polymerase chain reaction (PCR) assay is a recently developed molecular method for the detection, amplification, and quantization of nucleic acids [77]. Karry Mullis first described it in 1985 [78, 79], winning a Nobel Prize for his achievement. The reagents that are required for PCR include: (a) the dsDNA (template DNA); (b) two PCR primers, which are oligonucleotide sequences of single-stranded DNA that match the sequences at either end of the targeted DNA segment; (c) a thermostable enzyme to synthesize DNA copies; (The most commonly used enzyme is Taq polymerase from the heat-resistant bacterium, Thermus aquaticus.); (d) a pool of four deoxynucleotides (dATP, dCTP, dGTP, dTTP) that will be consumed by polymerase for the synthesis of the new DNA; and (e) buffers containing magnesium that are necessary for the function of Taq polymerase. A single PCR cycle is composed of three sequential steps: denaturation of the dsDNA at 92–96°C, primer annealing or hybridization at 55–72°C, and the synthesis or extension step at 72°C. During the denaturation step, the high temperature facilitates the breaking of hydrogen bonds between complementary bases, resulting in the separation of the dsDNA into two single strands. During the annealing step, the temperature drops, allowing the primers to hybridize to the complementary sequences on the template strands. Usually the primers are 20–30 bases long. The longer the primer, the more specific the binding on the target sequence. During the final step, new DNA strands are produced by Taq polymerase at 72°C. The elongation of the new DNA strands begins by using the oligonucleotide primers as starting points. DNA synthesis progresses from 5′ to 3′ for both new strands. Taq polymerase has the ability to synthesize approximately 1,000 base pairs per minute. The aforementioned three-step procedure is repeated from 30 to 50 times, leading to the synthesis of more than 1 × 109 copies of the original DNA template sequence.
The use of RNA, such as messenger RNA (mRNA), as a template for PCR amplification is accomplished by a modified PCR assay, named reverse transcriptase PCR (RT-PCR) [60, 80]. In a typical RT-PCR, mRNA is extracted from tissue samples or cells and then is copied into DNA (complementary DNA—cDNA) via reverse transcription, which is facilitated by the function of an enzyme called reverse transcriptase. This step is fundamental since Taq polymerase cannot use RNA as a template for the synthesis of PCR products. Several primers (such as random hexamers and oligo-dT) can be used for the reverse transcription step of PCR. The cDNA that is produced serves as the substrate for a classic PCR, as described earlier. In contrast to classic PCR that amplifies genomic DNA (which contains introns and exons), RT-PCR amplifies cDNA (which contains only exons) and therefore only useful genetic information.
The evaluation and analysis of PCR and RT-PCR products are usually made by gel electrophoresis. Electrophoresis is used for the separation of negatively charged nucleic acids, which are mobilized through a liquid or solid matrix by an electric field. Separation is based either on their molecule size or on their three-dimensional conformation. Agarose gel permits the separation of large DNA fragments (1–20 kb), whereas acrylamide gel is optimal for smaller (up to several base pairs) fragments. DNA molecules are visualized by ethidium bromide. The application of PCR in the fields of orthopedics and orthodontics is numerous. More specifically, in the area of bone pathology and oncology, this method can be used for the detection of mutations, polymorphisms, and other genomic alterations in oncogenes and tumor suppressor genes that are involved in tumorigenesis (such as p53, Rb1, p16, HER2/neu, EGF-R, and EXT2) [60]. Regarding skeletal biology, PCR can be used for the detection and quantitative assessment of genes and growth factors that are implicated in several molecular processes, including metabolic bone diseases, tumors, and cellular response to stress factors [81, 82]. Technologic advantages, such as real-time PCR and real-time quantitative TaqMan RT-PCR, are very sensitive, accurate, and highly reproducible methods for the study of gene expression and precise quantification of PCR products [60, 83, 84]. In addition, the development of the in situ PCR assay is an ideal combination of PCR and in situ hybridization that permits the selective amplification and evaluation of specific genetic loci within intact cells [85]. This method can be applied to cells, frozen sections, and sections from archived paraffin-embedded material and has the unique advantage of detecting specific genes within their native environment.
4 Microdissection Techniques
Formalin-fixed, paraffin-embedded tissues (FFPET) are some of the most widely available and quality-controlled materials for clinical and basic science studies. However, FFPET are complex, three-dimensional structures, composed of different cell populations with distinct functions. In bone biology, a large volume of studies is focused on the morphologic characteristics and the functional interactions between different cell populations (osteoblasts—osteocytes—chondroblasts). Profoundly, cellular heterogeneity constitutes a major drawback for molecular genetic analyses. Tissue microdissection (TM) represents a reliable method to isolate morphologically well-defined cells and obtain relatively pure cellular populations [81, 83, 84, 86]. DNA or RNA extracted from these populations can be amplified by PCR and then undergo molecular studies for the detection of genetic characteristics and the quantification of genomic alterations. TM can be performed by several different techniques ranging from simple, inexpensive manual TM to more sophisticated (but significantly more expensive) methods such as laser-captured microdissection (LCM).
Manual TM is performed under direct optical visualization of the tissue sample with the use of a stereomicroscope. The target cells are identified on 5-μm thick tissue sections and then dissected with sharp and accurate instruments such as a 30-gauge needle or surgical blade. For better results, tissue sections should undergo deparaffinization prior to microdissection [86]. Tissue fragments are collected in tubes and prepared for nucleic acid extraction and PCR analysis. The primers should be designed to generate small PCR products (<200 bp), since PCR with larger targets may fail. MTM is a fast, cost-effective method that can be applied to any tissue and does not require expensive instrumentation. Nonetheless, neighboring tissues and cells, such as lymphocytes and red blood cells, can very easily contaminate the cell population of interest. In order to obtain uncontaminated cell populations, LCM is the method of choice. LCM was first described by Emmert-Buck in 1999 [87, 88], and several commercially available microdissection systems that use laser technology were developed soon after. The major components of an LCM system are an inverted microscope, an infrared laser, a control unit for the laser, a control mechanism for the microscope stage, a digital camera, and a monitor [89]. LCM can be applied on both frozen and FFPE tissues. Importantly, deparaffinization is required prior to microdissection [90]. The major disadvantage of LCM is the prerequisite of very expensive equipment and well-trained technical stuff. TM is one of the most promising FFPET-based techniques that bridge the gap between morphology and molecular/genetic characteristics.
5 Culture of Osteoclasts and Osteoblasts
5.1 Osteoclastic Cell Lineage
Bone resorption and bone synthesis are fundamental processes that determine normal bone morphology, skeletal mass, and calcium homeostasis. Any disturbances of this finely tuned interplay result in pathologic conditions such as osteoporosis, osteopetrosis, metabolic bone diseases, fractures, and malignant hypercalcemia. The cells that are specialized to carry out bone resorption are the osteoclasts. Osteoclasts are derived from the pluripotential hematopoietic progenitor CFU-GM (colony-forming unit—granulocyte and macrophage), which also gives rise to monocytes and macrophage-committed precursors [91, 92]. The human mononuclear osteoclast precursor circulates in the monocyte fraction of the peripheral blood. It expresses the monocyte/macrophage integrins CD11b-c and the lipopolysaccharide receptor antigen CD14 [93, 94], as well as the macrophage-associated phenotypes NSE, Mac-1, and Mac-2. In contrast, they are negative for osteoclast-specific markers, namely, tartrate-resistant acid phosphatase (TRAP), vitronectin, and calcitonin receptors [95, 96]. Osteoclast activity is directly and specifically inhibited by calcitonin [97], and therefore receptors that bind calcitonin are considered to be reliable and highly specific markers of mammalian osteoclasts [98]. However, only a small fraction (approximately 2–5 %) of the monocyte/macrophage phenotype cells will eventually differentiate to mature osteoclasts [94]. Under the influence of the transcription factors PU-1 and MiTf, stem cells are committed into the myeloid lineage. In order to progress to the monocyte lineage and express the RANK receptor, M-CSF (macrophage colony-stimulating factor) is required. M-CSF is produced by mesenchymal/stromal cells, including osteoblasts, and is an absolute requirement for the proliferation and differentiation of osteoclast progenitors [99]. M-CSF acts via a tyrosine kinase receptor, named c-fms [100]. Precursors need the presence of RANKL to truly commit to the osteoclast lineage and begin the differentiation program. RANKL is a member of the tumor necrosis factors (TNF) family, which is expressed on the surface of osteoblasts/stromal cells and released by activated T-cells. It binds to RANK receptors on osteoclast precursors and induces their maturation through the nuclear factor-κΒ (NFκΒ) and Jun N-terminal kinase pathways [101, 102]. A member of the tumor necrosis factor receptors superfamily called osteoprotegerin is a decoy receptor for RANKL that inhibits the differentiation and function of osteoclasts [103, 104]. The transition from mononuclear precursor cell to mature osteoclast involves a stepwise loss of macrophage markers and gradual acquisition of phenotypic characteristics specific for osteoclasts. More specifically, since postmitotic osteoclast precursors begin to differentiate into committed osteoclast precursors, they express osteoclast-associated phenotypes, such as TRAP and calcitonin receptors [105]. In contrast, some of the macrophage-related markers, namely, NSE and Mac-1, disappear during osteoclast maturation. Furthermore, they respond to hormones, including 1,25-dihydroxyvitamin D3, parathyroid hormone, and certain cytokines such as IL-1, IL-6, prostaglandins, and colony-stimulating factors. When differentiation of the precursors into pre-osteoclasts is completed, these mononuclear cells begin to fuse, giving genesis to the multinucleated fully mature osteoclasts. Recent evidence suggests that mature osteoclasts undergo apoptosis after a cycle of resorption, a process augmented by estrogens [106].
5.2 Culture of Osteoclasts
Since osteoclasts originate from hemopoietic stem cells, bone marrow culture can be used for the study of osteoclast formation. Indeed, Testa and colleagues first demonstrated that multinucleated osteoclasts can be developed in long-term cultures of feline marrow cells [107]. Traditionally, osteoclasts have been generated in cocultures of osteoblasts or stromal cells and hematopoietic cells derived from spleen or bone marrow. For these studies, murine bone marrow cells can be aseptically extracted from long bones of 6–9-day-old mice, following removal of the adhering soft tissues [108]. Afterward, the ends of the bones are removed with scissors, and the bone marrow cells are extracted by slow injection of α-minimum essential medium (α-MEM) into one end of the bone. Bone marrow cells are collected and washed, suspended in α-MEM, and evaluated for viability. Approximately 1 × 107 bone marrow cells can be obtain from a tibia. Coculture methods rely upon the principle that osteoblasts secrete M-CSF and express RANKL after stimulation by 1,25-dihydroxyvitamin D3 and dexamethasone. RANKL binds RANK receptors to monocytic osteoclast precursors, promoting their fusion and thus synthesis of mature multinucleated osteoclasts [101]. Most of the coculture systems occupy UMR-106 rat osteosarcoma cell lines [109].
More recently, following the discovery of RANKL in 1998 [110], it has become possible to generate bone-resorbing osteoclasts without the requirement of osteoblasts in the culture. RANKL ligand and M-CSF can be added directly to osteoclast-precursor cultures, driving the formation of multinucleated, active bone-resorbing cells. This method is easier and more reliable than the coculture method since it employs cells from one single lineage. For osteoclastogenesis experiments that use bone marrow, the extracted cells are washed and cultured in α-MEM with fetal bovine serum (FBS) (10 %), M-CSF (5 ng/mL), and penicillin/streptomycin (1 %) for 48 h. Non-adherent hematopoietic stem cell precursors can be purified with Ficoll-Paque Plus (Amersham Biotech). The interfacial cell layer is isolated for culture in α-MEM with FBS (10 %), M-CSF (30–50 ng/mL), and RANKL (30–100 ng/mL). A TRAP kit can be used for osteoclast staining and counting. Other studies have described the development of osteoclasts from peripheral blood mononuclear cells (PBMNC) [105, 111, 112]. Briefly, 15 mL of blood is mixed with 15 mL of phosphate-buffered saline (PBS) (37°C), purified with 15 mL of Ficoll-Paque, and then centrifuged. Overlying cells are isolated, resuspended in 10 % PBS, diluted with 40 mL PBS, and centrifuged. Isolated PBMNC are placed in a 96-well plate and prewetted by soaking in 100 μL complete α-MEM containing 25 ng/mL M-CSF and 30 ng/mL recombinant RANKL at 37°C. The complete medium should be replaced every 2–3 days. The culture duration for both TRAP staining and pit assays is usually 2–3 weeks.
The multinucleated cells generated by cell cultures can be identified by the presence of certain osteoclast-specific markers: cathepsin K, calcitonin receptor, TRAP, type II carbonic anhydrase, and vitronectin receptor. However, the hallmark of osteoclast identification is the presence of resorption areas on calcified substrates, as defined by osteoclast-resorption lacunae (pit) assays [105, 109, 112].
5.3 Osteoblastic Cell Lineage
Osteoblasts arise from pluripotent mesenchymal stem cells, the colony-forming units—fibroblasts (CFU-Fs), which under appropriate stimulation can also give genesis to lipoblasts, chondroblasts, myoblasts, and fibroblasts [113, 114]. The bone morphogenic proteins (BMP) 2–7 and TGF-beta induce the upregulation of transcription factors that mediate the commitment of CFU-Fs toward the osteogenic lineage. The runt homology domain Runx2/Cbfa1 and the zinc finger protein osterix [54, 115, 116] are transcriptional regulators (Fig. 8.8) that facilitate the expression of genes (collagen type I, osteopontin, and alkaline phosphatase) that define the phenotypic features of bone-forming cells. Therefore, these proteins are referred as master regulators of osteoblast morphology. In vivo experiments have documented that knock-out mice lacking Runx2/Cbfa1 and osterix genes do not produce bone [116, 117]. Locally acting proteins, such as platelet-derived growth factor (PDGF), fibroblast growth factor (FGF), insulin-like growth factor (IGF), and activator protein-1 (AP-1), as well as systemic, blood-circulating molecules, namely, corticosteroids, 1,25 dihydroxyvitamin D3, parathyroid hormone, prostaglandins, and cytokines, are also implicated in specific steps of osteoblastic proliferation and maturation [118].
5.4 Osteoblastic Cell Cultures
The vast majority of the osteoblastic cell culture protocols use two complementary approaches for isolation of bone-forming cells. The first one utilizes freshly isolated short-term cultures of tissue-derived cells (primary or early passage cultures), whereas the second (permanent cell cultures) uses permanent cell lines, either from osteosarcoma tumors [119, 120] or from osteoblastic cell clones selected from primary cultures [121, 122].
5.5 Primary and Early Passage Cultures
Short-term cultures are carried out on non-transformed cells that have not undergone genomic alterations mutations, and therefore they retain most of their native phenotypic characteristics [123]. In primary and early passage cultures, bone-forming cells of different levels of differentiation can be encountered. Selective separation of these cells gives the investigators the opportunity to study the specific features of each subgroup, as well as the biological interactions between the bone-forming cells subpopulations [124]. Enzymatic digestion of bone matrix facilitated by proteases like collagenase or trypsin is the most commonly used method for the isolation of osteo-producing cells. Enzymatic digestion has been performed on fetal, neonatal, and adult calvariae, as well on long bones from mice [125], rats [126], chickens [127], cattle [128, 129], and humans [130, 131]. Osteoblastic cells are collected at different times as the digestion proceeds. Cells from the later digests (after 40–120 min) express the most “osteoblast-like” phenotype [126]. Nevertheless, primary culture systems have some disadvantages. Enzymatic isolation may have cytotoxic effects on the cells, whereas proteases may digest several cell surface proteins, affecting the phenotype of osteoblasts. Additionally, studies on mouse bone-forming cells have shown that in short-term cultures osteoblasts preserve their phenotypic features for a short period of time [125, 132].
Bone-forming cells can also be obtained from culture systems that use periosteum [133], bone marrow stroma [134], or periodontal ligaments [135–137]. These tissues contain mesenchymal/osteoblast precursors, which can lead to the genesis of cells that occupy bone-forming features. Dexamethasone [135], retinoic acid [138], and BMP-2 [134] have been shown to augment the aforementioned phenomena. This method provides significant information regarding the biochemical and molecular events that are implicated in the process of osteoblastic differentiation and maturation.
5.6 Permanent Cultures
Most of the permanent cell lines are derived from cells that have undergone malignant transformation and become immortalized. Immortalized osteoblastic cells have not acquired all the genetic and morphologic characteristics of fully transformed, osteosarcoma cells, and they maintain their osteoblastic phenotype on a continuing basis, providing large amounts of stable cell populations that are ideal for biochemical studies. The most popular osteosarcoma cell lines are UMR-6 (rat) and ROS 17–2 (rat). Each exhibits different features [124, 132] and serves different purposes. Osteoblast-like cell lines (namely, SaOS, TE-85, MG-63, OHS-4) have also originated from human osteosarcomas [119, 120]. The greatest disadvantage of osteosarcoma cell lines is that the process of immortalization-transformation may have affected the genotype and phenotype of the osteoblastic cells. In order to overcome this caveat, permanent osteoblastic cell lines, such as MC3T3-E1, have been developed from normal mouse calvarias [139]. Osteoblast characterization in cell cultures is based upon biochemical and morphological elements. Osteoblast differentiation/maturation occupies distinct phases, identified by the expression of different sets of genes.
More specifically, during the proliferation phase, osteoblasts express collagen type 1 and histone proteins, growth factors (TGF-beta), specific transcription factors (c-Fos, c-Myc, Fra-1, c-Jun, JunD), and the osteoblastic master regulator Cbfa1/Runx2 [140–143]. As the differentiation of the progenitor cells proceeds and extracellular matrix begins to mature, proteins such as collagen type 1, alkaline phosphatase, osteopontin, osteonectin, bone sialoproteins, and PTH/RTHrP are upregulated. During the period of matrix mineralization, mature osteoblasts are characterized by the expression of osteocalcin, osteopontin, osteocalcin, and collagenase. Apoptotic cells are observed during the mineralization phase associated with the formation of bone nodules and the expression of the apoptosis-related factors Bax and the cell cycle regulator/tumor suppressor gene P53 [106]. Both in vivo and in vitro studies have demonstrated that histologically similar osteoblasts in different proliferation and maturation stages display heterogeneous profiles in proteins and mRNA levels [140–142, 144]. Notably, a recent immunohistochemical in situ hybridization study conducted on fetal rat calvaria has shown that ALP and PTHrP-R are globally expressed by all osteoblasts irrespective of their maturation status [140]. Ultimately, osteoblasts are identified by their histological configuration, histochemical properties (i.e., ALP positive staining), and, most importantly, by their ability to synthesize bone matrix.
5.7 Mechanical Stretching of Cell Cultures
Mechanical forces are essential physiological factors that regulate the structural properties of bone tissue. Mechanical loading stimulates the osteoblastic function and plays a fundamental role in bone remodeling and skeletal homoestasis [145–148]. In cell cultures, osteoblasts display significantly similar phenotypical and genotypic features with fibroblasts. Therefore, osteoblasts have been characterized as sophisticated fibroblasts [143]. Human periodontal ligament (hPDL) is connective tissue that lies between the tooth root and the alveolar bone [149]. PDL fibroblasts comprise an osteoblast-like population, which may undergo osteoblastic differentiation under the influence of a variety of extracellular stimuli, including mechanical loading in vivo and in vitro [136, 137, 150, 151]. This fact generated the notion that the development of a PDL fibroblast stretch application device might have considerable contribution toward understanding the molecular events that underlie mechanical sensing, biochemical coupling, and the response to mechano-transduction within the periodontal ligament tissue [152].
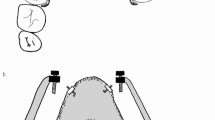
The stretch devices, similarly to those used for stretch application to other tissues [153], are mainly based on culturing cells in dishes with a flexible bottom. The culture surface can be stretched so that the cells attached to this surface are stretched also. HPDL fibroblasts are obtained from explant cultures of PDL tissues dissected from roots of healthy teeth [152, 154, 155]. The explant is cultured in Dulbecco’s modified Eagle’s medium (DMEM) enhanced with 10 % (volume per volume) FCS, nonessential amino acids, and antibiotics (100 IU/mL penicillin, 100 μg/mL streptomycin, and 0.25 μg/mL amphotericin). Cultures are maintained at 37°C in a 5 % CO2 environment and fed every 2 or 3 days. Fibroblastic cells from the explants start outgrowing 8–10 days after the culture initiation. The cells are trypsinized with 0.15 % trypsin/0.5 mM EDTA, harvested by centrifugation, washed in PBS, transferred into 75 cm2 flasks containing the complete medium, and cultured to confluence (Passage 0). Following trypsin digestion, cells are subcultivated at a 1:4 split ratio on tissue culture dishes that carry a flexible, hydrophilic growth surface. PDL cells from third to sixth passages can be used for the experiments. The flexible bottom dishes can be altered from a flat position to a convex configuration by placing a Plexiglas template with a convex surface underneath and applying a weight on the top of the dish cover, thus forcing the membrane to adapt to the convex surface. The strain level can be measured by calculating the percentage of membrane expansion. The same effect can be also achieved by using more complicated systems with elaborate vacuums controlled by sophisticated software [156]. After continuous stretch application for the appropriate time intervals, the medium is removed, the cells are lysed, and proteins are extracted for further biochemical analyses [157, 158].
5.7.1
The future of orthodontics sounds bright and flourishing! In an era of cells, molecules, and targeted pharmaceutical intervention, orthodontics cannot stay behind. Basic research focusing on the tissue and cells reactions within the periodontal ligament and the surrounding alveolar bone slowly but steadily unravels the inside biological phenomena responsible for restructuring the architecture of the area and the occurring orthodontic tooth movement. Surely, it is a very specific research area that requires deep biological knowledge and dexterity with complex techniques far beyond the clinical interest of the everyday clinical orthodontist.
The techniques provided in this section focus on presenting the special tips and hinds required for those orthodontics who are involved in basic research, since general chapters do not covered such a specific material. Main histological and histochemical protocols, as well as specific osteoblast and osteoclast cell tissue techniques, are pivotal tools in order to understand in depth the remodeling alveolar background. More important, the external force application systems presented here show the variety of the parameters that should be taken into account when force application in biological systems is studied! One part is the cells and the molecules, but the second one, also decisive for the clinical outcome, is the force. And both parts up to date do not appear clear cut!
It is rather obvious that we, orthodontists, have reached our limits in terms of clinical intervention the classical way. It is the basic research that will give us that quantum leap that we need in optimizing our treatment: reduce the extent of treatment, abolish the cooperation, eliminate the invasive pin approaches, secure retention and treatment outcome, and last but not least teach how to use force magnitude and duration in a scientific way! Furthermore, this will provide information to the industry for the development of even more efficient orthodontic materials!
The time of pharmacological intervention within the periodontal ligament is not far. As targeted drugs develop in medicine for other conditions related to force application and bone remodeling such as in osteoporosis, orthodontics will follow. Studies appear more and more often examining the reaction of periodontal ligament tissue to external force application in the presence of injected drugs within the periodontal space. This is surely the future!
And since everything is in the periodontal ligament, its cells and fibers, and without this magnificent apparatus orthodontic tooth movement is not at all possible, we better be informed and well educated! The benefits are rather easy to imagine.
References
McNamara JA Jr (1985) The role of functional appliances in contemporary orthodontics. In: Johnston LE (ed) New vistas in orthodontics. Lea and Febiger, Philadelphia, pp 38–75
Reitan K (1962) Bone formation and resorption during reversed tooth movement. In: Kraus BS, Riedel RA (eds) Vistas in orthodontics. Lea and Febiger, Philadelphia, pp 69–84
Moss ML (1962) The functional matrix. In: Kraus BS, Riedel RA (eds) Vistas in orthodontics. Lea and Febiger, Philadelphia, pp 85–98
Akiyama T, Miyazaki T, Buillet P, Nakamura K, Strasser A, Tanaka S (2005) In vitro and in vivo assays for osteoclast apoptosis. Biol Proced Online 7:48–59
Alers JC, Krijtenburg PJ, Vissers KJ, van Dekken H (1999) Effect of bone decalcification procedures on DNA in situ hybridization and comparative genomic hybridization: EDTA is highly preferable to a routinely used acid decalcifier. J Histochem Cytochem 47:703–710
Alvarez JI, Teitelbaum SL, Blair HC, Greenfield EM, Athanasou NA, Ross FP (1991) Generation of avian cells with the osteoclast phenotype from mononuclear phagocytes. Endocrinology 128:2324–2335
Anderson GN, Marks SC Jr (1989) Tartrate-resistant acid ATPase as a cytochemical marker for osteoclasts. J Histochem Cytochem 37:115–117
Ansari B, Coates PJ, Greenstein BD, Hall PA (1993) In situ end labeling detects DNA strand breaks in apoptosis and other physiological and pathological states. J Pathol 170:1–8
Arceo N, Sauk JJ, Moehring J, Foster RA, Somerman MJ (1991) Human periodontal cells initiate mineral-like nodules in vitro. J Periodontol 62:499–503
Athanasou NA, Quinn J (1990) Immunophenotypic differences between osteoclasts and macrophage polykaryons: immunohistological distinction and implications for osteoclast ontogeny and function. J Clin Pathol 43:997–1003
Athanasou NA (1996) Cellular biology of bone-resorbing cells. J Bone Joint Surg Am 78:1096–1113
Aubin JE, Candeliere GA, Bonnelye E (1999) The heterogeneity of the osteoblast phenotype. Endocrinologist 31:25–31
Aubin JE, Heersche JNM, Merrilees MJ, Sodek J (1982) Isolation of bone cell clones with differences in growth, hormone responses and extracellular matrix production. J Cell Biol 92:452–461
Aubin JE (1998) Bone stem cells. J Cell Biol 30–31:73–82
Auf’mkolk B, Hauschka P, Schwartz E (1985) Characterization of human bone cells in culture. Calcif Tissue Int 37:228–235
Balanti P, Minisola S, Pacitti MT, Scarnecchia L, Rosso R, Mazzuoli GF, Bonucci E (1997) Tartrate-resistant acid phosphate activity as osteoclastic marker: sensitivity of cytochemical assessment and serum assay in comparison with standardized osteoclast histomorphometry. Osteoporos Int 7:39–43
Basdra EK, Komposch G (1997) Osteoblast-like properties of human periodontal ligament cells: an in vitro analysis. Eur J Orthod 19:615–621
Basdra EK, Komposch G (1999) Transmission and scanning electron microscopic analysis of mineralized loci formed by human periodontal ligament cells in vitro. J Orofac Orthop 60:77–86
Basdra EK, Papavassiliou AG, Huber LA (1995) Rab and rho GTPases are involved in specific response of periodontal ligament fibroblasts to mechanical stretching. Biochim Biophys Acta 1268:209–213
Beck GR Jr, Zerler B, Moran E (2001) Gene array analysis of osteoblast differentiation. Cell Growth Differ 12:61–83
Bianco P, Riminucci M, Kusnetsov S, Robey PG (1999) Multipotential cells in the bone marrow stroma: regulation in the context of organ physiology. Crit Rev Eukaryot Gene Expr 9:159–173
Blair HC, Athanasou NA (2004) Recent advances in osteoclast biology and pathological bone resorption. Histol Histopathol 19:189–199
Brigati DJ, Myerson D, Leary JJ, Spalholz V, Travis SZ, Fong CK, Hsiung GD, Ward DC (1983) Detection of viral genomes in cultured cells and paraffin-embedded tissue sections using biotin-labeled hybridization probes. Virology 126:32–50
Brown RSD, Edwards J, Barlett JW, Dogan A (2002) Routine acid decalcification of bone marrow samples can preserve DNA for FISH and CGH studies in metastatic prostate cancer. J Histochem Cytochem 50:113–115
Buckley KA, Chan BYY, Fraser WD, Gallager JA (2004) Human osteoclast culture from peripheral blood monocytes: phenotypic characterization and quantitation of resorption. In: Picot J (ed) Methods in molecular medicine, vol 107, 2nd edn, Human cell culture protocols. Humana Press, Totowa, pp 55–68
Candeliere GA, Liu F, Aubin JE (2001) Individual osteoblasts in the developing calvaria express different gene repertoires. Bone 28:351–361
Carnes DL, Maeder CL, Craves DT (1997) Cells with osteoblastic phenotypes can be explanted from human gingival and periodontal ligament. J Periodontol 68:701–707
Chambers TJ, Magnus CJ (1982) Calcitonin alters the behaviour of isolated osteoclasts. J Pathol 136:27–39
Chenu C, Colucci S, Grano M, Zigrino P, Barattolo R, Zambonin G, Baldini N, Vergnaud P, Delmas PD, Zallone AZ (1994) Osteocalcin induces chemotaxis, secretion of matrix gla proteins and calcium-mediated intracellular signaling in human osteoclast-like cells. J Cell Biol 127:1149–1158
Curran S, McKay JA, McLeod HL, Murray JI (2000) Laser capture microscopy. Mol Pathol 53:64–68
D’Errico JA, Macneil RL, Takata T, Berry J, Strayhorn C, Somerman MJ (1997) Expression of bone associated markers by tooth root lining cells, in situ and in vitro. Bone 20:117–126
Defranco D, Lian J, Glowacki J (1992) Differential effects of glucocorticoid on recruitment and activity of osteoclasts induced by normal and osteocalcin-deficient bone implanted in rats. Endocrinology 131:114–121
Devlin CJ, Brickell PM, Taylor ER, Hornbruch A, Craig RK (1988) In situ hybridization reveals differentiated spatial distribution of mRNAs for type I and type II collagen in chick limb bud. Development 103:111–118
Dougall WC, Glaccum M, Charrier K, Rohrbach K, Brasel K, De Smedt T, Daro E, Smith J, Tometsko ME, Maliszewski CR, Armstrong A, Shen V, Bain S, Cosman D, Anderson D, Morrisey PJ, Peschon JJ, Schuh J (1990) RANK is essential for osteoclast and lymph node development. Genes Dev 13:2412–2424
Ducy P, Schinke T, Karsenty G (2000) The osteoblast: a sophisticated fibroblast under central surveillance. Science 289:1501–1504
Ducy P, Zhang R, Geoffroy V, Ridall AL, Karsenty G (1997) Osf2/Cbfa1: a transcriptional activator of osteoblast differentiation. Cell 89:747–754
Egelberg J (1987) Regeneration and repair of periodontal tissues. J Periodontol Res 22:233–242
Ek-Rylander B, Flores M, Wendal M, Heinegard D, Andersson G (1994) Dephosphorylation of osteopontin and bone sialoprotein by osteoclastic tartrate-resistant acid phosphatase. Modulation of osteoclast adhesion in vitro. J Biol Chem 269:14853–14856
Emmert-Buck MR, Bonner RF, Smith PD, Chuaqui RF, Zhuang Z, Goldstein SR, Weia RA, Liotta LA (1996) Laser capture microdissection. Science 274:998–1001
Evans CE, Galasko CS, Ward C (1990) Effects of donor age on the growth in vitro of cells obtained from human trabecular bone. J Orthop Res 8:234–237
Faust J, Lacey DL, Hunt P, Burgess TL, Scully S, Van G, Eli A, Qian Y, Shalhoub V (1999) Osteoclast markers accumulate on cells developing from human peripheral blood mononuclear precursors. J Cell Biochem 72:67–80
Fedarko NS, Vetter UK, Gehron Robey P (1995) Age-related changes in bone matrix structure in vitro. Calcif Tissue Int 56:S41–S43
Fend F, Kremer M, Quantilla-Martinez L (2006) Laser capture microdissection: methodical aspects and applications with emphasis on immune-laser capture microdissection. Pathobiology 68:209–214
Filgueira L (2004) Fluorescence-based staining for tartrate-resistant acid phosphatase (TRAP) in osteoclasts combined with other fluorescent dyes and protocols. J Histochem Cytochem 52:411–414
Fournier B, Price P (1991) Characterization of a new human osteosarcoma cell line OHS-4. J Cell Biol 114:577–583
Frost HM, Villneueva AR, Roth H, Stanisavljevic S (1961) Tetracycline bone labeling. J Clin Pharmacol 1:206–216
Fujikawa Y, Quinn JM, Sabokbar A, McGee JO, Athanasou NA (1996) The human osteoclast precursor circulates in the monocyte fraction. Endocrinology 137:4058–4060
Gaffney EF, O’Neil AJ, Staunton MJ (1995) In situ end labeling, light microscopic assessment and ultrastructure of apoptosis in lung carcinoma. J Clin Pathol 48:1017–1021
Gall JG, Pardue ML (1969) Formation and detection of RNA–DNA hybrid molecules in cytological preparations. Proc Natl Acad Sci U S A 63:378–383
Gaulier A, Fourcade C, Szekeres G, Pulik M (1994) Bone marrow one step fixation decalcification in Lowy FMA solution – an immunohistologic and in situ hybridization study. Pathol Res Pract 190:1149–1161
Glowaski J, Lian B (1987) Impaired recruitment and differentiation of osteoclast progenitors by osteocalcin-depleted bone implants. Cell Differ 21:247–254
Granet C, Boutahar N, Vico L, Alexandre C, Lafage-Proust MH (2001) MAPK and SRC-kinases control EGR-1 and NF-kappa B inductions by changes in mechanical environment in osteoblasts. Biochem Biophys Res Commun 284:622–631
Halleen JM, Raisanen S, Sallo JJ, Roodman GD, Hentunen TA, Lahenkari PP, Kaija H, Vihko P, Vaananen HK (1999) Intracellular fragmentation of bone resorption products by reactive oxygen species generated by osteoclastic tartrate-resistant acid phosphatase. J Biol Chem 274:22907–22910
Harris H (1990) The human alkaline phosphatase: what we know and what we don’t know. Clin Chim Acta 186:133–150
Hattersley G, Chambers TJ (1989) Calcitonin receptors as markers for osteoclastic differentiation: correlation between generation of bone-resorptive cells that express calcitonin receptors in mouse bone marrow cultures. Endocrinology 125:1606–1612
Hayman AR, Bune AJ, Bradley JR, Rashbass J, Cox TM (2000) Osteoclastic tartrate-resistant acid phosphatase (Acp 5): its localization to dendritic cells and diverse murine tissues. J Histochem Cytochem 48:219–227
Hofler H, Childers H, Montminy MR, Lechan RM, Goodman RH, Wolfe HJ (1986) In situ hybridization methods for the detection of somatostatin mRNA in tissue sections using antisense RNA probes. Histochem J 18:597–604
Hoshi K, Amizuka N, Oda K, Ikehara Y, Ozawa H (1997) Immunolocalization of tissue non-specific alkaline phosphatase in mice. Histochem Cell Biol 107:183–191
Hsu SM, Raine L, Fanger H (1981) Use of avidin-biotin-peroxidase complex (ABC) in immunoperoxidase techniques: a comparison between ABC and unlabeled antibody (PAP) procedures. J Histochem Cytochem 29:577–580
Hughes DE, Wright KR, Uy HL, Sasaki A, Yoneda T, Roodman GD, Mundy GR, Boyce BF (1995) Bisphosphonates promote apoptosis on murine osteoclasts in vitro and in vivo. J Bone Miner Res 10:1478–1487
Hunt JL, Finkelstein SD (2004) Microdissection techniques for molecular testing in surgical pathology. Arch Pathol Lab Med 128:1372–1378
Ikeda T, Nomura S, Yamaguchi A, Suda T, Yoshiki S (1992) In situ hybridization of bone matrix proteins in undecalcified adult rat bone sections. J Histochem Cytochem 40: 1079–1088
Jacquet R, Hillyer J, Landis WJ (2005) Analysis of connective tissues by laser capture microdissection and reverse transcriptase – polymerase chain reaction. Anal Biochem 337:22–34
John HA, Birnstiel ML, Jones KW (1969) RNA–DNA hybrids at the cytological level. Nature 223:582–587
Kadkol SS, Gage WR, Pasternack GR (1999) In situ hybridization – theory and practice. Mol Diagn 4:169–183
Kawamoto T, Shimizu M (2000) A method for preparing 2- to 50-μm thick fresh-frozen sections of large samples and undecalcified hard tissues. Histochem Cell Biol 113:331–339
Kawamoto T (2003) Use of a new adhesive film for the preparation of multi-purpose fresh-frozen sections from hard tissues, whole-animals, insects and plants. Arch Histol Cytol 66:123–143
Khosla S, Kleerekoper M (2003) Biochemical markers of bone turnover. In: Favus MJ (ed) Primer on the metabolic bone disorders of mineral metabolism, 2003rd edn. American Society for Bone and Mineral Research, Washington, D.C., pp 166–172
Kleerekoper M (1996) Biochemical markers of bone remodeling. Am J Med Sci 312: 270–277
Kletsas D, Basdra EK, Papavassiliou AG (1998) Mechanical stress induces DNA synthesis in PDL fibroblasts by a mechanism unrelated to autocrine growth factor action. FEBS Lett 430:358–362
Kletsas D, Basdra EK, Papavassiliou AG (2002) Effect of protein kinase inhibitors on the stretch-elicited c-Fos and c-Jun up-regulation in human PDL osteoblast-like cells. J Cell Physiol 190:313–321
Kodama H, Amagai Y, Sudo H, Kasai S, Yamamoto S (1981) Establishment of a clonal osteogenic cell line from newborn mouse calvaria. Jpn J Oral Biol 23:899–901
Komminoth P, Long AA (1993) In situ polymerase chain reaction. An overview of methods, applications and limitations of new molecular technique. Virchows Arch B Cell Pathol Incl Mol Pathol 64:67–73
Komori T, Yagi H, Nomura S, Yamaguchi A, Sasaki K, Deguchi K, Shimizu Y, Bronson RT, Gao Y-H, Inada M, Okamoto R, Kitamura Y, Yoshiki S, Kashimoto T (1997) Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 89:755–764
Kurihara N, Chenu C, Miller M, Civin C, Roodman GD (1990) Identification of committed mononuclear precursors for osteoclast-like cells formed in long term human marrow cultures. Endocrinology 126:2733–2741
Lacey DL, Timms E, Tan HL, Kelley MJ, Dunstan CR, Burgess T, Elliott R, Colombero A, Elliott G, Scully S, Hsu H, Sullivan J, Hawkins N, Davy E, Capparelli C, Qian YX, Kaufman S, Sarosi I, Shalhoub V, Senaldi G, Guo J, Delaney J, Boyle WJ (1998) Osteoprotegerin ligand is a cytokine that regulates osteoclast differentiation and activation. Cell 93:165–176
Lang P, Schultzberg M, Andersson G (2001) Expression and distribution of tartrate-resistant purple acid phosphatase in the rat nervous system. J Histochem Cytochem 49:379–396
Li CY, Yam LT, Lam KW (1970) Acid phosphatase isoenzyme in human leukocytes in normal and pathologic conditions. J Histochem Cytochem 18:473–481
Lian JB, Stein GS, Aubin JE (2003) Bone formation: maturation and functional activities of osteoblast lineage. In: Favus MJ (ed) Primer on the metabolic bone diseases and disorders of mineral metabolism, 5th edn. American Society for Bone and Mineral Research, Washington, D.C., pp 13–28
Lillie R, Fuller H (1976) Histopathologic technique and practical histochemistry. McGraw-Hill, New York
Luben RA, Wong GL, Cohn DV (1976) Biochemical characterization with parathormone and calcitonin of isolated bone cells; provisional identification of osteoclasts and osteoblasts. Endocrinology 99:526–534
MacLean HE, Kronenberg HM (2005) Localization of Indian hedgehog and PTH/PTHrP receptor in relation to chondrocyte proliferation during mouse bone development. Dev Growth Differ 47:59–63
Majeska RJ (1996) Culture of osteoblastic cells. In: Bilezikian JP, Raisz LG, Rodan GA (eds) Principles of bone biology. Academic, San Diego, pp 1229–1237
Mallory FB (1938) Pathological Technique: A practical manual for workers in pathological histology, including directions for the performance of autopsies and microphotography Saunders, Philadelphia, pp 143–144
Mankin C (1982) Bone acid phosphatase: tartrate-resistant acid phosphatase as a marker of osteoclast function. Calcif Tissue Int 34:285
Marks SC, Grolman M-L (1987) Tartrate-resistant acid phosphatase in mononuclear and multinuclear cells during the bone resorption of tooth eruption. J Histochem Cytochem 35:1221–1230
Martin RB (1989) Label escape theory revised: the effects of resting periods and section thickness. Bone 10:255–264
Matsuzaki K, Udagawa N, Takahashi N, Yamaguchi K, Yasuda H, Shima N, Morinaga T, Toyama Y, Yabe Y, Higashio K, Suda T (1998) Osteoclast differentiation factor (ODF) induces osteoclast-like cell formation in human peripheral blood mononuclear cell cultures. Biochem Biophys Res Commun 246:199–204
Matthews JB (1982) Influence of decalcifications on immunohistochemical staining of formalin-fixed paraffin-embedded tissue. J Clin Pathol 35:1392–1394
McCabe LR, Kockx M, Lian J, Stein J, Stein G (1995) Selective expression of fos- and jun-related genes during osteoblast proliferation and differentiation. Exp Cell Res 218:255–262
McComb RB, Bowers GN, Posen S (1979) Alkaline phosphatase. Plenum Press, New York
McNicol AM, Farquharson MA (1997) In situ hybridization and its diagnostic applications in pathology. J Pathol 182:250–261
Meloan SN, Puchtler H (1985) Chemical mechanisms of staining methods: von Kossa’s technique. What von Kossa really wrote and a modified reaction for selective demonstration of inorganic phosphate. J Histotechnol 8:11–13
Miao D, Scutt A (2004) Histochemical localization of alkaline phosphatase activity in decalcified bone and cartilage. J Histochem Cytochem 50:333–340
Milch RA, Rall DP, Tobie JE (1958) Fluorescence of tetracycline antibiotics in bone. J Bone Joint Surg 40A:897–910
Moss DW (1992) Perspectives in alkaline phosphatase research. Clin Chem 28:2486–2492
Mukai K, Yoshimura S, Anzai M (1986) Effects of decalcification on immunoperoxidase staining. Am J Surg Pathol 10:413–419
Mullink H, Henzen-Logmans SC, Tadama TM, Mol JJ, Meijer CJ (1985) Influence of fixation and decalcification on the immunohistochemical staining of cell-specific markers in paraffin-embedded human bone biopsies. J Histochem Cytochem 33:1103–1109
Mullis K, Faloona F, Scharf S, Saiki R, Horn G, Erlich H (1986) Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Cold Spring Harb Symp Quant Biol 51:263–273
Mullis KB, Faloona FA (1987) Specific synthesis of DNA in vitro via a polymerase catalyzed chain reaction. Methods Enzymol 155:335–350
Mundy GR, Chen D, Zhao M, Dallas S, Xu C, Harris S (2001) Growth regulatory factors and bone. Rev Endocr Metab Disord 2:105–115
Nakahara H, Bruder SP, Haynesworth SE, Holecek JJ, Baber MA, Goldberg VM, Caplan AI (1990) Bone and cartilage formation in diffusion chambers by subcultured cells derived from the periosteum. Bone 11:181–188
Nakano Y, Toyasawa S, Takano Y (2004) Eccentric localization of osteocytes expressing enzymatic activities, protein and mRNA signals for type 5 tartrate-resistant acid phosphatase (TRAP). J Histochem Cytochem 52:1475–1482
Nakashima K, Zhou X, Kunkel G, Zhang Z, Deng JM, Behringer RR, de Crombrugghe B (2002) The novel zinc finger-containing transcription factor osterix is required for osteoblast differentiation and bone formation. Cell 108:17–29
Nijweide PJ, van der Plas A, Scherf JP (1981) Biochemical and histological studies on various bone cell preparations. Calcif Tissue Int 33:529–540
Papachristou D, Pirttiniemi P, Kantomaa T, Agnantis N, Basdra EK (2006) Fos-and Jun-related transcription factors are involved in the signal transduction pathway of mechanical loading in condylar chondrocytes. Eur J Orthod 28:20–26
Papachristou DJ, Goodman MA, Cieply K, Hunt JL, Rao UMN (2006) Comparison of allelic losses in chondroblastoma and primary chondrosarcoma of bone and correlation with fluorescence in situ hybridization. Human Pathol 37:890–898, Epub 2006 May 22
Papachristou DJ, Pirttiniemi P, Kantomaa T, Papavassiliou AG, Basdra EK (2005) JNK/ERK–AP-1/Runx2 induction “paves the way” to cartilage load-ignited chondroblastic differentiation. Histochem Cell Biol 124:215–223
Parfitt AM, Drenzer MK, Glorieux FH, Kanis JA, Malluche H, Meunier PJ, Ott SM, Recker RR (1987) Bone histomorphometry: standardization of nomenclature, symbols and units. J Bone Miner Res 2:595–610
Parfitt AM (2002) Physiologic and pathogenetic significance of bone histomorphometric data. In: Coe FL, Favus MJ (eds) Disorders of bone and mineral metabolism, 2nd edn. Lippincott, Williams and Wilkins, Philadelphia, pp 469–485
Peverali FA, Basdra EK, Papavassiliou AG (2001) Stretch – mediated activation of selective MAPK subtypes and potentiation of AP-1 binding in human osteoblastic cells. Mol Med 7:68–78
Raab-Cullen DM, Akhter MP, Kimmel DB, Recker RR (1994) Periosteal bone formation stimulated by externally induced bending strains. J Bone Miner Res 9:1143–1152
Rahn BA, Perren SM (1971) Xylenol orange, a fluorochrome useful in polychrome sequential labeling of calcifying tissues. Stain Technol 46:125–129
Ramakrishnan PR, Lin W-L, Sodek J, Cho M-H (1995) Synthesis of noncollagenous extracellular matrix proteins during development of mineralized nodules by rat periodontal ligament cells in vitro. Calcif Tissue Int 57:52–59
Recker RR, Berger-Lux MJ (2003) Bone biopsy and histomorphometry in clinical practice. In: Favus MJ (ed) Primer on the metabolic bone disorders of mineral metabolism, 5th edn. American Society for Bone and Mineral Research, Washington, D.C., pp 213–219
Reinhold EG (2003) Bone-labeling techniques. In: Handbook of histology methods for bone and cartilage. Humana Press, Totowa, pp 99–118
Reinholt FP, Mangarelli Widholm S, Ek-Rylander B, Andersson G (1990) Ultrastructural localization of a tartrate-resistant acid ATPase in bone. J Bone Miner Res 5:1055–1061
Rennert H, Leonard BGB (2003) Molecular methods in the diagnostic laboratory. In: Leonard DGB (ed) Diagnostic molecular pathology. Saunders, Philadelphia, pp 25–51
Rennick SL, Fenton TW, Foram DR (2005) The effects of skeletal preparations techniques on DNA from human and non-human bone. J Forensic Sci 50:1–4
Rickard DJ, Sullivan TA, Shenker BJ, Leboy PS, Kazhadan I (1994) Induction of rapid osteoblast differentiation in rat bone marrow stromal cell cultures by dexamethasone and BMP-2. Dev Biol 161:218–228
Ritter N, Farach-Carson M, Butler W (1992) Evidence for the formation of a complex between osteopontin and osteocalcin. J Bone Miner Res 7:877–885
Roach HI (1999) Association of matrix acid alkaline phosphatase with mineralization of cartilage and endochondral bone. Histochem J 31:53–61
Robison R (1923) The possible significance of hexosephosphoric esters in ossification. Biochem J 17:286–293
Rodan SB, Majeska RJ, Rodan GA (1994) Osteosarcoma cells as models for osteoblasts. In: Novak JF, MacMaster JH (eds) Frontiers of osteosarcoma research. Holfeger and Huber, Seattle, pp 193–203
Rubin CT, Gross TS, McLeod KJ, Bain SD (1995) Morphologic stages in lamellar bone formation stimulated by a potent mechanical stimulus. J Bone Miner Res 10:488–495
Saiki RK, Scharf S, Faloona F, Mullis KB, Horn GT, Erlich HA, Arnheim N (1985) Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230:1350–1354
Salomon RN (1995) Introduction to reverse transcriptase polymerase chain reaction. Diagn Mol Pathol 4:2–3
Seth LA, Lee BK, Vary CP (2000) Coordinate expression of novel genes during osteoblast differentiation. J Bone Miner Res 15:1683–1696
Shalhoub V, Faust J, Boyle WJ, Dunstan CR, Kelley M, Kaufman S, Scully S, Van G, Lacey DL (1999) Osteoprotegerin and osteoprotegerin ligand effects on osteoclast formation from human peripheral blood mononuclear cell precursors. J Cell Biochem 72:251–261
Simonet WS, Lacey DL, Dunstan CR, Keley M, Chang MS, Luthy R, Nguyen HQ, Wooden S, Bennett L, Boone T, Shimamato G, DeRose M, Elliott R, Colombero A, Tan HL, Trail G, Sukkivan J, Davy E, Bucay N, Renshaw-Gegg L, Hughes TM, Hill D, Pattison W, Campbell P, Sander S, Van G, Tarpley J, Derby P, Lee R, Boyle WJ (1997) Osteoprotegerin: a novel secreted protein involved in the regulation of bone density. Cell 89:309–319
Specht K, Richter T, Muller U, Walch A, Werner M, Hofler H (2001) Quantitative gene expression analysis in microdissected archival formalin-fixed and paraffin-embedded tumor tissue. Am J Pathol 158:419–429
Speel EJM, Hopman AHN, Komminoth P (1999) Amplification methods to increase the sensitivity of in situ hybridization: play CARD(S). J Histochem Cytochem 47:281–288
Sudo H, Kodama H, Amagai Y, Yamamoto S, Kasai S (1983) In vitro differentiation and calcification in a new clonal osteogenic cell line derived from newborn mouse calvaria. J Cell Biol 96:191–198
Tanaka S, Takahashi N, Udagawa N, Tamura T, Akatsu T, Stanly ER, Kurokawa T, Suda T (1993) Macrophage colony stimulating factor is indispensable for both proliferation and differentiation of osteoclast progenitors. J Clin Invest 91:257–265
Tanner GA, McQuillan PF, Maxwell MR, Keck JK, McAteer JA (1995) An in vitro test of the cell stretch-proliferation hypothesis of renal cyst enlargement. J Am Soc Nephrol 6:1230–1241
Termine JD, Kleinman HK, Whitson SW, Conn KM, McGarney ML, Martin GR (1981) Osteonectin, a bone-specific protein linking mineral to collagen. Cell 26:99–105
Testa N, Allen TD, Lajtha LG, Onions D, Jarret O (1981) Generation of osteoclast in vitro. J Cell Sci 47:127–137
Tulli H, Carlson C, Jayo M (1992) Immunohistochemical method for the simultaneous demonstration of three proteins in EDTA decalcified paraffin embedded bone sections. J Histotechnol 15:93–97
Uehara F, Ohba N, Nakashima Y, Yanagota T, Ozawa M, Muramatsou T (1993) A fixative suitable for in situ hybridization histochemistry. J Histochem Cytochem 41:947–953
Vaananen HK, Zhao H, Mulari M, Halleen JM (2000) The cell biology of osteoclast function. J Cell Sci 113:377–381
von Kossa J (1901) Über die im Organismus kunstlich erzeugbaren Verkalkungen. Beit Path Anat Allg Pathol 29:163–202
Walch A, Specht K, Smida J, Aubele M, Zitzelberger H, Hofler H, Werner M (2001) Tissue microdissection techniques in quantitative genome and gene expression analysis. Histochem Cell Biol 115:269–276
Walsh L, Freemont AJ, Hoyland JA (1993) The effect of tissue decalcification on mRNA retention within bone for in situ hybridization studies. Int J Exp Pathol 74:237–241
Whitson SW, Whitson MA, Bowers DE Jr, Falk MC (1992) Factors influencing synthesis and mineralization of bone matrix from fetal bovine cells grow in vitro. J Bone Miner Res 7:727–741
Whyte MP (1994) Hypophosphatasia and the role of alkaline phosphatase in skeletal mineralization. Endocr Rev 15:439–461
Whyte MP (2008) Hypophosphatasia: nature¢s window on alkaline phosphatase function in humans. In: Bilezikian JP, Raisz LG, Martin TJ, (eds) Principles of Bone Biology. Academic, San Diego
Wong G (1989) Isolation and behavior of isolated bone-forming cells. In: Hall BK (ed) The osteoblast and osteocyte. Telford Press, Boca Raton, pp 171–192
Wong GL, Mg MC (1992) Maturation-associated changes in the cellular composition of mouse calvariae and in the biochemical characteristics of calvarial cells separated into subclasses on percoll density gradients. J Bone Miner Res 7:701–708
Wu XB, Li Y, Schneider A, Yu W, Rajendren G, Iqbal J, Yamamoto M, Alam M, Brunet LJ, Blair HC, Zaidi M, Abe E (2003) Impaired osteoblastic differentiation, reduced bone formation, and severe osteoporosis in noggin-overexpressing mice. J Clin Invest 112:924–934
Yamashiro T, Aberg T, Levanon D, Groner Y, Thesleff I (2002) Expression of Runx1, -2 and -3 during tooth, palate and craniofacial bone development. Mech Dev 119S:S107–S110
Yang X, Karsenty G (2002) Transcription factors in bone: developmental and pathological aspects. Trends Mol Med 8:340–345
Yasuda H, Shima N, Nakagawa N, Yamaguchi K, Kinosaki M, Mochizuki S, Tomoyasu A, Yano K, Goto M, Murakami A, Tsuda E, Morinaga T, Higashio K, Udagawa N, Takahashi N, Suda T (1998) Osteoclast differentiation factor is a ligand for osteoprotegerin/ osteoclastogenesis-inhibitory factor and is identical to TRANCE/RANKL. Proc Natl Acad Sci U S A 95:3597–3602
Yee JA (1983) Properties of osteoblast-like cells isolated from the cortical endosteal bone surfaces of adult rabbits. Calc Tissue Int 35:571–577
Yoshida H, Hayashi S, Kunisada T, Ogawa M, Nishikawa S, Okamura H, Sudo T, Shultz LD, Nishikawa S (1990) The murine mutation osteopetrosis is the coding region of the macrophage colony stimulating factor gene. Nature 1354:442–444
Yoshiki S, Umeda T, Kurahashi Y (1972) An effect reactivation of alkaline phosphatase in hard tissues completely decalcified for light and electron microscopy. Histochemie 29:296–304
Zaidi M, Blair HC, Moonga B, Abe E, Hung L-HC (2003) Osteoclastogenesis, bone resorption, and osteoclast-based therapeutics. J Bone Miner Res 18:599–609
Zhou H, Hammonds RG Jr, Findlay DM, Fuller PJ, Martin TJ, Ng KW (1991) Retinoic acid modulation of mRNA levels in malignant, nontransformed, and immortalized osteoblasts. J Bone Miner Res 6:767–777
Ziros PG, Gil AP, Georgakopoulos T, Habeos I, Kletsas D, Basdra EK, Papavassiliou AG (2002) The bone-specific transcriptional regulator Cbfa1 is a target of mechanical signals in osteoblastic cells. J Biol Chem 277:23934–23941
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Papachristou, D.J., Basdra, E.K. (2013). Basic Science Research Methods in Orthodontics. In: Eliades, T. (eds) Research Methods in Orthodontics. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-31377-6_8
Download citation
DOI: https://doi.org/10.1007/978-3-642-31377-6_8
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-31376-9
Online ISBN: 978-3-642-31377-6
eBook Packages: MedicineMedicine (R0)