Abstract
Bacteria have evolved unique mechanisms that allow them survive in the presence of strong selection pressures. Included in these mechanisms is the ability to share genetic determinants among and between species of bacteria thus spreading metal or antibiotic resistance traits quickly. The textile industry in response to demand has developed antimicrobial fabrics by the addition of bactericidal compounds during production. Some of these antimicrobials include metal nanoparticles, quaternary ammonia compounds, and broad spectrum compounds like triclosan. Bacteria have already expressed resistance to each of these bactericides. Here we discuss the evolutionary and ecological consequences of antimicrobial textiles in terms of co-selection. We predict that continued use of such materials could result in increased and widespread resistance to specific antimicrobials, especially metals, with an increased resistance to antibiotics. Such increases have the potential to find their way into other bacterial populations of human pathogens leading to serious and unintended public health consequences.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Colonizing all known areas of the biosphere, there are approximately five nonillion (5 × 1030) bacteria on Earth, forming much of the world’s biomass (Whitman et al. 1998). Bacteria arose 3.8 billion years ago and, as a group, have survived longer than all other organisms combined. More importantly, they are still here. During this incredible evolutionary history microbial diversity has evolved to such an extent that, based on 16SRNA and metagenome sequencing, there is significantly far more genetic diversity among bacteria and archaea than among all other organisms.
Undoubtedly the first bacteria evolved in rich toxic metal environments (Silver 1998) or other similarly harsh environments. Survival required the ability to circumvent the toxic nature of these inhospitable extreme conditions. Continued existence and perpetuation of a species is contingent on the genetic repertoire. Consequently, it is difficult to avoid the conclusion that metal resistance genes were among the very first gene systems to evolve, and the genetic basis for metal homeostasis is evident in both recently diverged bacterial phyla as well as ancient crenarchaeota clades (Baker-Austin et al. 2005). Over time as organisms modified their environments, niches arose that did not require metal resistance and such traits were presumably lost or culled from the genome of certain species. However, in metal-rich environments these genes quite certainly persisted (Haefeli et al. 1984).
Bacteria have extraordinary adaptive genetic capacities and these capacities have been primarily shaped by horizontal gene transfer of mobile genetic elements (MGE) (Szczepanowski et al. 2008; O’Brien 2002; Ochman et al. 2000; Yurieva et al. 1997). Interestingly, the genes for regulation, resistance, and biosynthesis are often found linked together on the same continuous strand of DNA (Clardy et al. 2009). Thus, the evolutionary ecological history of bacterial genes is driven, under many circumstances, by horizontal gene transfer events suggesting that the survival capabilities of specific microbes are more dependent on the promiscuity and plasticity of its genome than the genetic characteristics of its ancestors.
The ability of MGE to interact with bacterial genomic DNA across multiple species and environmental barriers creates a unique evolutionary problem. Bacterial cells in ecological proximity to each other may be more important in microbial adaptations and gene exchange than genetic relatedness alone. Microbes that are able to draw from the collective genetic resistome (D’Costa et al. 2007) should have higher chances of survival than those individuals that lack such ability. However because of reproduction by binary fission, bacterial cells closest are often copies of each other—and that fact frequently colors our collective interpretation regarding bacterial evolution.
Bacterial MGE spread not only vertically, by inheritance, but also horizontally by involving phylogenetically distant cells (Davison 1999; Lorenz and Wackernagel 1994; Sobecky 1999). The horizontal dissemination of MGEs can be accomplished via plasmid exchange (conjugation), the uptake of naked DNA from the environment (transformation), or via viral infection mechanisms (transduction). The metagenome of a bacterial community should indicate past and present selective pressures in that environment (Turner et al. 2002). It has been well documented that genes conferring antibiotic resistance are often more plentiful in bacterial communities exposed to antibiotic contamination (Heuer and Smalla 2007; Pei et al. 2006). For example, mercury resistance genes are more abundant in mercury-contaminated sediments (Smalla et al. 2006). However, in contrast to classic selection theory, previous studies have documented that additional genotypes and phenotypes can be co-selected along with traits under direct selection (Alonso et al. 2001; Baker-Austin et al. 2006; Summers et al. 1993).
There are no known examples of extinction of a bacterial lineage; although such may be the case. A number of reasons have been identified for why extinctions in bacteria may be very low (Dykhuizen 1998). Firstly, bacteria rarely starve to death because they can lay dormant until environmental conditions improve. Secondly, bacteria readily share genes between and within “species,” conferring a selective advantage in many instances to just a small subset of the overall population that can thrive even under highly inhospitable conditions. Thirdly, microbes do not require sex to reproduce, eliminating the limiting factor of finding a suitable mate to continue a genetic lineage. Finally, they, as a group, are able to live in a range of environments under extremes of physical and chemical conditions. The premise that bacteria rarely undergo extinction is important in understanding the ability of man to alter bacterial communities and populations through various technologies and methods.
Perhaps more important than low extinction rates are the high speciation rates found among bacteria (Dykhuizen 1998). It is this ability to rapidly adapt and evolve to novel or changing conditions that allows bacteria to exploit extreme habitats and to withstand strong selection that, for more genetically and phenotypically “advanced” organisms, results in mass extinctions. Such exploitation of the environment is primarily facilitated by horizontal gene transfer.
Resistance to antibiotics is a prime example of this rapid evolution. The word antibiotic was first used as noun by Selman Waksman in 1941 as a description of molecules produced by one microbe that has a negative effect on the growth of another (Clardy et al. 2009). Large-scale production of antibiotics began in the early 1940s with the expectation that infectious diseases might become relegated to the past. In fact the use of antibiotics dropped the rate of death from infectious disease from nearly 800/100,000 in 1900 to <40/100,000 in 1980 (Walsh and Wright 2005). Unfortunately, by the mid-1940s antibiotic-resistant bacteria had emerged. It is this ever expanding resistance to antibiotics, both old and new generation, that has motivated the emergence of novel delivery or bactericidal methods. The discovery of antibiotics has been considered one of the defining events in medicine and science during the twentieth century (Davies and Davies 2010). However, the ever increasing levels of antibiotic resistance in agriculture, hospitals, and in environmental reservoirs seemingly in step with their use are equally or of greater importance than their discovery. Microbes relying on the strong selection imposed by the misuse and overuse of antibiotics have been able to exploit mechanisms of gene exchange to spread every source of resistance genes among close and unrelated taxa. In the process they have developed multiple mechanisms of resistance and many species harbor multiple resistance genes. Some species have been screened that are resistant to almost every class of antibiotic, including recently developed drugs (Stepanauskas et al. 2005, 2006).
Genotypes may persist in a bacterial community even after a given selective pressure no longer exists (Enne et al. 2001). This observation is counter intuitive as there should be a fitness cost associated with maintaining extraneous DNA when there is no immediate benefit. However, several studies have shown that bacteria carrying these extra-genetic elements do not seem to show decreased fitness when compared to strains of the same species without the traits (Andersson and Levin 1999; Schrag et al. 1997). Therefore, various gene combinations can and do remain in the bacterial community and are available for transfer via horizontal gene transfer mechanisms or selective increase under changing environmental conditions.
2 Use of Metals as Bactericides
Although various metals are necessary for bacterial survival as micronutrients, e.g., Cr, Co, Cu, Mn, Mo, Ni, Se, W, V, Zn, and Fe, many of these elements are toxic at higher concentrations. The antibacterial effects of some of these and certain other metals have been known since antiquity (Silver and Phung 1996) prompting their use in dentistry and medicine (Kim et al. 2007; Catauro et al. 2004; Crabtree et al. 2003). The efficacy of a metal biocide is dependent on the concentrations used and the exposure time. While many studies report dramatic decreases in bacterial numbers (see discussion below) very few studies have shown complete elimination of bacteria. This observation is critical and drives the remaining discussion.
3 Nanoparticles
The bioavailability of particles is enhanced as the size decreases. Indeed the science of nanotechnology is focused on making materials with significantly improved physical, chemical, and biological properties (Wang 2000) with increased functionality because of the nanosize. However, there are a number of problems with bringing nanotechnology into large-scale commercial use (Mazzola 2003; Serov et al. 2003; Ohshima 2003) not the least of which is significant differences between batches using the same protocols to produce standard reference materials for use in experiments.
One area where nanotechnology may have an impact is drug delivery and therapeutics. The use of nanoparticles as an antimicrobial allows the delivery of metals to the actual cell. This increase in the efficacy of the metal as a biocide has brought about what has been called a “new generation of antimicrobials” (Rai et al. 2009). Shrivastava et al. (2008) presented what they described as the enhanced antimicrobial effect of novel silver nanoparticles. Interestingly, their methods were more pronounced against gram-negative bacteria than gram-positive organisms. In fact none of the doses of silver nanoparticles they used were effective against Staphylococcus aureus.
There is a general consensus regarding the modes of antimicrobial activity of metal and carbon nanoparticles. Most studies indicate that nanoparticles cause disruption of bacterial cell membranes in probable response to various oxygen species (Neal 2008). For this to occur there needs to be contact between the bacteria and the nanoparticle with interfacial process such as electrostatic interactions. Toxicity of the nanoparticle may also be important and while unlikely, accumulation does in some instances occur after disruption of the membrane. With the discovery of the mode of action of nanoparticles there seems to be a similar sense of wonder to that expressed when the first “wonder drugs” were developed, i.e., an assurance that we can beat the processes of bacterial evolution. We will discuss the evolutionary arms race between man and bacteria below.
4 Antimicrobial Textiles
Textiles, both natural fiber and engineered fibers, provide surfaces for microbial growth. Given optimal conditions of temperature, water, minimal nutrients, electron donors and acceptors, and a carbon source, microbes associated with textiles can multiply rapidly. In general, engineered fibers are more resistant to microbial degradation than naturally derived fibers. Textiles thus become an excellent media for microbial growth with the resulting consequences such as odor, discoloration, and increased likelihood of infections, etc. Cultural evolution of human societies, especially in developed countries, has placed an ever increasing demand for clothing that is hygienic. Gao and Cranston (2008) indicate that sportswear, socks, shoe linings, and lingerie account for 85% of the total production of antimicrobial textiles. Production of these textiles in Western Europe has increased more than 15% per year between 2001 and 2005 (Gao and Cranston 2008). Other industries have increased the demand for antimicrobial fibers for use in air filters, automotive and outdoor textiles, home furnishings, and textiles used in medicine.
Manufacturers have met this increasing demand imposed by a public concern about comfort, hygiene, and physical well-being by incorporating a wide range of broad spectrum biocides. Various antimicrobials have been used including silver, quaternary ammonium compounds, and triclosan on most of the products of the textile industry (Table 1). These substances are applied at various stages of the textile process such as finishing or through incorporation into synthetic fibers during extrusion.
The effectiveness of these treatments has been varied. For example, Tomšič et al. (2009) examined the antimicrobial activity of AgCl that had been embedded in a silica matrix on cotton and found that the biocidal effects were strongly influenced by the concentration of the silver present in the coatings. Saengkiettiyut et al. (2008) demonstrated that silver nanoparticles on cotton fabrics have excellent antibacterial activity against Staphylococcus aureus, Staphylococcus aureus methicillin resistance strain (MRSA), Escherichia coli and Pseudomonas aeruginosa. In another study (Dubas et al. 2006) antimicrobial silver nanoparticles were immobilized on nylon and silk fibers using a layer-by-layer deposition method. This procedure showed an 80% reduction in Staphylococcus aureus on silk fibers and a 50% reduction on the nylon fibers. While this procedure did demonstrate a decrease in bacterial activity there were significant numbers of bacteria remaining on both fibers.
Conversely, a study (Li et al. 2006) designed to assess the antimicrobial activity of silver nitrate and titanium dioxide nanoparticle coated facemasks in an effort to protect against infectious agents found unequivocal results. After 48 h of exposure and incubation the authors found a 100% reduction in viable Escherichia coli and S. aureus whereas there was a 20% and 50% respective increase in the viable counts of these two species on untreated masks. A study that compared the effect of Ag nanoparticles on microbial activity on cotton and wool showed variable results for both fibers and species of microorganism examined (Falletta et al. 2008). When applied to cotton and polyester fibers there was complete inhibition of bacterial and fungal growth; however, when applied to wool there was only inhibition against fungi.
While numerous other studies can be reviewed similar results are obtained, i.e., high variability between and within studies on the effectiveness of a nanoparticle treatment. This variability ranges from complete inhibition to no inhibition and seems to be dependent on dose, exposure, and type of fiber under consideration. Most importantly is the observation that the efficacy of this treatment regime is, in many cases, not completely successful against a wide variety of microorganisms.
5 Cross-Selection and Co-selection of Metal Resistance Traits in Nature
Strikingly, the structural basis of microbial resistance to metals and antibiotics is highly similar (Fig. 1). Bacteria use just a handful of mechanisms to detoxify metals and antimicrobials, such as active extrusion from the cell, sequestration, and exclusion of toxin by the cell membrane, among others. Of further evidence of this symmetry, recent studies have implicated the role metal contamination has as a selective agent in the proliferation of antibiotic resistance (See Baker-Austin et al. 2006 for review). These studies have documented that various associations between types and levels of metal contamination and environmental patterns of antibiotic resistance exist in nature. Such patterns suggest that various mechanisms underlie the co-selection of metal and antibiotic resistance. There are at least two co-selection mechanisms that include co-resistance and cross-resistance. Co-resistance occurs when different resistance genes are found on the same genetic element, e.g., resistance to aminoglycosides and Hg resistance genes on the same mobile genetic element such as a plasmid or integrase. In cross-resistance the same genetic determinant is responsible for more than one type of resistance, e.g., Cd and tetracycline efflux pumps where the same pump is used to rid the cell of either Cd or tetracycline.
Metal contamination of the environment is different from the release of antibiotics because antibiotics can, over time, be degraded. Metals cannot be degraded although their mobility through soil matrices can be altered by changes in valence state and the chemical composition of soil particles. Depending on the valence state of the metal the bioavailability similarly changes. If the metal is tightly bound to a surface it is less likely to have an impact on the microbial communities than the same metal that is in solution. The toxic impact of metals on bacteria is dependent on how much the microorganism interacts with the metal as well as the chemical state of the metal upon contact (e.g., reduced, oxidized, methylated, etc.).
In controlled laboratory experiments, it has been shown that exposure of bacteria to either metals or antibiotics results in an increase in the proportion of bacteria resistant to both metals and antibiotics. In these experiments, naïve bacteria in river water were exposed to either Cd or Ni chloride at increasing concentrations or to the antibiotics tetracycline or ampicillin at increasing concentrations. Both the metals and antibiotics had significant effects on the fraction of cultivable bacteria in the microcosms. Total numbers of bacteria decreased nearly one order of magnitude after being exposed to the metals. Of those bacteria remaining the proportions of resistant bacteria increased dramatically. Where less than 20% of cultivable bacteria obtained from the control microcosms (river water) were resistant to any of the tested antibiotics between 50 and 100% of isolates from the microcosms amended with either Cd or Ni were resistant to at least one antibiotic (Stepanauskas et al. 2006). Of equal interest while only 3% of isolates obtained from the control microcosms were resistant to Cd or Ni, 50–60% of isolates obtained from the microcosms amended with tetracycline and ampicillin were resistant to at least one of the two metals. Of greater concern when river water microbial assemblages were individually exposed to each of the four toxicants, the frequency of microorganisms with multiple resistances (metal and antibiotic resistances) increased. This study is important for at least three reasons:
-
1.
It demonstrated that bacteria exposed to metals increase the frequency of antibiotic-resistant bacteria.
-
2.
It demonstrated that bacteria exposed to antibiotics increase the frequency of metal-resistant bacteria.
-
3.
It provided the first experimental evidence that the exposure of freshwater microbial assemblages to individual metals and antibiotics selects for multiresistant microorganisms (Stepanauskas et al. 2006).
These laboratory studies as well as numerous observational studies from the environment confirm that exposure of bacteria to metals increases the proportion of bacteria resistant to the metals but more importantly the number of bacteria resistant to antibiotics. Even without sequencing it is apparent that co-selection is occurring. Whether the mechanisms in these multiresistant phenotypes are cross-resistance or co-resistance is interesting from a basic science perspective but the most important outcomes have significant practical importance and may directly affect public health.
6 Resistance to Ag and Other Metals Used in Microbial Textiles
Silver and colleagues (Silver 2003; Silver and Phung 1996; Silver et al. 2006) have for some time been warning of increased resistance to metals used in biocides and to silver specifically. Basing their predictions on models explaining the rapid and widespread increase in antibiotic resistance that has been carefully documented, they suggest that horizontal gene transfer, under strong selection of metal exposure, will move metal resistance traits quickly through microbial assemblages and populations. A restrained word of caution has been offered by Percival et al. (2005) who suggest that the risks of antibacterial resistance developing from the use of metal containing biocides have been overstated. They feel that there have been too few studies that have effectively documented the prevalence of resistance. Where Silver (2006) argues the possibility that widespread use of Ag may promote the development of resistance, Percival et al. (2005) citing numerous studies to support their conjecture, counter that the probability of transferring silver resistance genes is low, unstable, difficult to maintain and to transfer. Interestingly, Percival et al. (2005) suggest that the occurrence of silver resistance can be variable under different conditions of exposure and that it is difficult to distinguish between sensitive and resistant bacteria. They then provide a list of different bacteria that have been shown to be silver resistant or from which silver-resistance conferring plasmids have been isolated. While the list is not exhaustive it is suggestive that the trait can occur in many different types of bacteria and should be considered indicative of the possibility of many other bacteria showing the trait.
The greatest possibility of finding silver-resistant bacteria is in environments where usage or occurrence of metals is highest such as dentistry where amalgams contain 35% silver (Brunne 1986), in hospital burn units where silver is used in dressings (Klasen 2000), or on catheters (Sampath et al. 1995). Several controlled laboratory studies have shown a relationship between bacterial resistance to biocides and cross-resistance to antibiotics. While these studies create various levels of contention among researchers the results seem to be following the same paths trod in the early days of studies on antibiotic resistance.
Following Percival et al. (2005) publication, Silver et al. (2006) provided a new review of silver resistance studies. From this review we find that bacterial resistance to Ag+ has been observed repeatedly (Silver et al. 2006) but that until recently the genetic mechanisms were not understood. Genes located on plasmids frequently confer resistance to toxic metals including silver (Davis et al. 2005; Gupta et al. 2001). The plasmid pMG101, which was originally found in Salmonella encodes resistance to several antibiotics as well as Hg2+, Ag+, and tellurite (Gupta et al. 1999, 2001; McHugh et al. 1975). However, genes conferring resistance to Ag are not restricted to plasmids. In Salmonella and Escherichia coli, a chromosomal Ag+ resistance determinant has been described (Gupta et al. 2001; Franke et al. 2001; Silver 2003). Selection imposed by the widespread, often indiscriminate use of antiseptics, has resulted in increased resistance to Ag+ in clinical and hospital settings (Davis et al. 2005). For example, (Silver 2003) found in a random sampling of enteric bacteria from a Chicago hospital that more than 10% had resistance genes for Ag+ resistance. These observations suggest that the potential for wide-scale horizontal gene transfer of metal resistance genes, similar to antibiotic resistance genes, could occur under such strong selection. Such strong selection is clearly employed in hospital settings but is also used in the application to textiles. However, Chopra (2007) followed with a review which concluded the variability in results from various studies is probably due to a failure to establish standard procedures for determining MIC values, a lack of recognized breakpoints, differences in the way products and media release silver, and finally that there is no standard way to measure inhibitory activity. Chopra (2007) suggests that because the clinical incidence of silver resistance remains low the emergence of resistance can be contained or minimized if silver ion concentrations are high and bactericidal activity rapid. Gao and Cranston (2008) counter that because large quantities of biocides are frequently required on textiles to achieve an adequate durability and effect there should be increased concern. Most biocides used in textiles can and do induce bacterial resistance to the biocide (Table 1) but more importantly this selection increases resistance to antibiotics through co-selection. Gao and Cranston (2008) suggest that the long-term benefits and potential problems of antimicrobial textiles need to be closely monitored.
What is curious about these review articles is that many fail to discuss the other side of resistance, i.e., naturally occurring resistance and the use of bacteria to synthesize nanoparticles of various metals. Nanoparticle scientists rely on the ability of various bacteria to not only resist metal toxicity but to sequester metals within the cells (Mandal et al. 2006). As early as 1979, bioaccumulation of silver in bacteria had been described in a multispecies community of bacteria (Charley and Bull 1979). In this study, an isolated stable community of three species had an unusually high tolerance to silver where tolerance is defined as remaining viable under the imposed conditions but not necessarily reproducing. The community was composed of Pseudomonas maltophilia, Staphylococcus aureus and an unidentified species. The effect of exposure to Ag was a change in the relative abundance of the three species. Tolerance of high silver concentrations was reduced when the community was grown in the absence of silver suggesting that these resistance-bearing bacteria were less able to compete in the nonselective environment. Considering that in several recent studies silver did not affect or had minimal effect of S. aureus it seems surprising that these earlier studies on environmental bacteria did not help design experiments or to define the hypotheses and predictions regarding antimicrobial textile studies.
The most important observation that needs to be made in studies on the efficacy of antimicrobial textiles is how many of the bacteria that remain are metal resistant. If we decrease the total viable bacteria to ranges found in various studies (92–99.99%) that means that there are still between 1,000 and hundreds of thousands of bacteria remaining. It is likely that the remaining bacteria are simply tolerant and not necessarily resistant. The same is true of bacteria exposed to an initial dose of antibiotics, i.e., the initial low dose does not kill all bacteria at once and hence the need to take the full regimen of antibiotics to increase the likelihood that all disease-causing bacteria are eliminated. Failure to do so means that these tolerant or minimally resistant bacteria are free to proliferate either in the host or in the environment resulting in an increase in both resistant pathogens and their genes that can be transferred to other species of bacteria. Since bacterial growth rates are high these remaining bacteria can increase in density or pass their genes to bacteria that are adapted for the ambient conditions and thus increase the number of resistant genes in the assemblage.
7 Problems with Nanoparticles on Textiles
One very important observation regarding antimicrobial textiles is that the effective particles or applications do not necessarily remain on the textile but can be released by repeated washings and enter into the environment. Unfortunately, there has not been sufficient research to mimic natural systems in laboratory settings or to actual controlled releases into complex natural systems (Neal 2008). Failure to perform carefully designed experiments has resulted in a poor understanding of likely consequences from the release of these substances into the environment. Neal (2008) in reviewing what the then current studies were, states that there is a general failure to identify any significant effects at the microbial level of nanoparticles in complex systems. Does this means that there are no effects or that we have not yet observed or measured the component of the system where an effect has occurred?
There is a cause for concern about the toxicity and bioavailability of these substances to bacteria involved in critical ecosystem level processes especially biogeochemical cycles such as carbon and nitrogen cycling. Chronic low level release of these antimicrobials into the environment will increase tolerance and over time result in increased numbers of diverse species carry resistance genes. More importantly, the chronic input of metal-rich nanoparticles into the environment may have unforeseen consequences. A clear understanding of effects on critical players in important biogeochemical processes must be explored under various natural conditions. Only then can we make intelligent predictions of future impacts or lack thereof.
The pathways to increased exposure of environmental bacteria are similar between antibiotics in the environment and nanoparticles. Washing of metal-coated textiles will distribute released substances into waterways. Increased antibiotic resistance traits can be found at much higher levels below water treatment facilities and below intensive agricultural areas. We would predict similar increases in metal-resistant bacteria below such facilities due to co-selection caused by increased selection by the antibiotics present in the water column (Baquero et al. 2008). Because nanoparticles and antibiotics will have the same vectors of inputs into waterways it will be difficult to ascertain whether increased metal resistance found downstream is due to direct selection or via co-selection mechanisms. The same problem is true in clinical situations. Because of the strong selection from the various biocidal agents used it may be difficult to determine the exact mechanism of increased metal and antibiotic resistance. Nevertheless, such studies are imperative.
8 Evolutionary Arms Races
An evolutionary arms race occurs between two organisms when one is antagonistic or parasitic towards the other. Selection should favor traits in the host or antagonized species that prevent or decrease the incidence of infection or antagonism. However, this change in the host then provides selection for traits in the antagonizing species which can overcome the acquired traits of the host. This tit-for-tat cycle continues as a spiral of ever increasing point and counter-point between the two species with each evolving to unique characteristics that promote survival.
The limits of this spiral are determined by the genetic resources of the two organisms. Researchers have investigated such an arms race between bacteria and bacteriophage (Weitz et al. 2005). In this system the bacteria evolve to change the binding site(s) that are used by the phage to gain entry into the bacterial cell. This in turn results in selection of phage that are able to bind to novel sites. It is clear that these arms races never eliminate either the host or the infectious agent but the interaction controls densities.
Given that bacteria evolved in a metal-rich environment we should expect that various gene combinations can be found that confer resistance to metals. Indeed, Silver (1998) has stated that there are genes for all metals. In other words, there are no toxic metals for which there are no genes or gene systems that can confer resistance. Furthermore, there are genes that actually allow bacteria to accumulate or sequester metals within the cytoplasm, such as metallochaperones. As man continues to attempt to modify his environment, especially in the realm of clinical, strong selection is imposed on potential disease-causing organisms. The result is a general decrease in bacterial numbers and in many cases what appears as an initial elimination of the disease-causing bacterial agents. However, over time resistant bacteria emerge with traits that allow them to survive in ever increasing concentrations of the bactericidal substances. Thus, an evolutionary arms race is occurring between the capacity of bacteria to evolve and the creative ability of man to make new substances that are effective against disease-causing bacteria. This is a new area of evolution that, although the mechanisms used by bacteria have been in existence for millennia, is not between one species evolving in response to the evolution of another species. Rather the selection is imposed by the technological innovation of man and the genetic evolution of bacteria. The only question is which species has the greatest probability to counter the other. Given the evolutionary history of microorganisms and their impressive ability to develop resistances to all known classes of naturally occurring and synthetic drug agents, bacteria represent formidable opponents.
Novel or what seem to be novel methods to treat bacterial infections, regardless of whether the substance used is newly synthesized and thus never been found before in nature, has only a limited life expectancy or efficacy. Time and again, bacteria have overcome these barriers and continue to infect man with decreasing options for effective clinical intervention.
9 Is There a Reason for Concern?
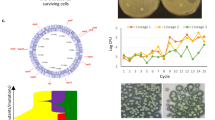
One additional finding first reported in Baker-Austin et al. (2009) raises the specter of unwitting artificial selection for so-called superbugs. They showed that with the increasing number of antibiotics to which a bacterial isolate is resistant, there appeared to be a concomitant increase in the concentration of the antibiotic that was minimally inhibitive to those isolates. We refer to this as the “multiresistance effect.” The bacteria in Baker-Austin et al. (2009) were shellfish pathogens Vibrio vulnificus, collected from two known metal contaminated estuaries and from a third nearly pristine estuary along the southeastern Atlantic coast. Figure 2 shows a similar pattern among Escherichia coli bacteria from the same collection events (unpublished). The obvious implication is that constant exposure to heavy metals, in naturally occurring or anthropogenic elevated concentrations, co-selects for multiple antibiotic resistance traits. More importantly, the more traits acquired, the higher the concentration (on average) of the corresponding antibiotics required to inhibit growth. This is a somewhat puzzling outcome given the prevailing hypothesis that the acquisition of new traits/genes comes with a fitness cost that isolates without such traits do not have to bear. Traditionally, a trade-off is expected between the costs of carrying and replicating new genes in the genome and the conferred fitness benefit in the presence of the selective agent. However, MacLean et al. (2010) point out that
An illustration among estuarine E. Coli isolates of the unexpected “multiresistance effect.” This is a positive and statistically significant (p < 0.05) relationship between the number of antibiotics to which an isolate is resistant and the mean minimum Inhibitory concentration (MIC) among only those traits acquired per isolate, expressed on a standard normal deviate scale
Once a population is dominated by resistant cells, mutants that have second-site compensatory mutations that recover the cost of resistance have higher fitness than resistant genotypes lacking the compensatory mutations, and natural selection ultimately results in the fixation of compensatory mutations,
which benefits remain, they point out, in the absences of antibiotics. Perhaps, this explains the “multiresistance effect,” but to do so completely requires us to acknowledge conveniently high second-site mutation rates given metal contaminant selection pressure in only ecological time. We suspect the more likely explanation still has to do with HGT of those genes which recover fitness costs from carrying antibiotic resistance genes. After all, as we have discussed above, such genes are likely to have been around for a very long time.
This research finding of a “multiresistance effect,” combined with evidence provided earlier in this chapter should be sufficient cause for concern over the widespread use of antimicrobials including metals on textiles. Co-selected bacteria with resistance traits are likely to find their way into public water treatment facilities and become gene nurseries for multiply resistant strains. Furthermore, HGT will only add to this concern from the acquisition of antibiotic resistance genes by human pathogens in such public works settings. Note that our discussion here does not technically qualify as an ugly prediction, but should give pause for the assessment of current research on these potential biological innovations that may have unintended consequences.
10 Where Do We Go from Here?
Modern culture imposes economic incentives for the development of products that meet perceived needs. Included in these needs are fabrics that help to eliminate odors, that resist degradation, and that are capable of killing or greatly reducing potential disease-causing organisms. Prior to the development of broad-spectrum bactericides washing clothes was sufficient for the effective removal or temporary inactivation of bacteria present in clothing. However, medicine and dentistry require materials that can fight off infections. The development of these materials opened the door for the development of textiles to be used by the general public. Many products are available to kill 99.99% of microbes on contact such as hand washes and wipes. Progression towards fabrics and materials with antibacterial substances embedded or attached is a continued step towards trying to rid humans of possible infection-causing agents.
Given the seemingly unlimited capacity of bacteria to respond to and exploit novel environments it is not surprising that resistance to metals, biocides, and antibiotics occurs. Both metals and antibiotic are found in nature although the role of antibiotics in nature is not fully elucidated. It is our contention that the use of bactericidal products will over time select for resistant bacteria. Bacteria resistant to the bactericide probably carry with it other co-selected traits. Bacteria can live in the most inhospitable places on the planet, e.g., high level radioactive waste tanks, thus it is unlikely that even high concentrations of metal nanoparticles would be effective for long periods of time. In addition, the effects of metal nanoparticles, released from antimicrobial textiles into the environment, are not understood and could have serious consequences. Similar concerns have been raised about the long- and short-term effects of human-derived antibiotics on natural systems. While there may have been short-term effects such as the reduction in the abundance of specific groups of bacteria, overall biogeochemical cycles have not been significantly impacted by the release of antibiotics into the environment. However, as noted above, antibiotics can be degraded over time either by microbial activity or through natural weathering whereas metal nanoparticles cannot. Thus, while the effect of antibiotics on natural processes has not been shown to be a major problem the effect of metal nanoparticles is less sure. Regardless of whether metal resistance increases in the community, some species are less capable of obtaining new genetic traits and identifying and characterizing which bacteria are impacted is the question of concern. For example if nitrogen fixing bacteria are negatively impacted we might expect significant changes in nitrogen cycling over time.
The arms race will continue with man and his array of technologies and ability to perceive, conceive, and achieve being matched by the long evolutionary history of microbes. Microbes who first colonized the planet and continue to do so 3.8 billion years later.
References
Alonso A, Sanchez P, Martinez JL (2001) Environmental selection of antibiotic resistance genes. Environ Microbiol 3:1–9
Andersson D, Levin BR (1999) The biological cost of antibiotic resistance. Curr Opin Microbiol 2:489–493
Baker-Austin C, Dopson M, Wexler M, Sawers G, Bond PL (2005) Molecular insight into extreme copper resistance in the acidophilic archaeon ‘Ferroplasma acidarmanus’ Fer1. Microbiology 151:2637–2646
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14:176–182
Baker-Austin C, McArthur JV, Lindell AH, Wright MS, Tuckfield RC, Gooch J, Warner L, Oliver J, Stepanauskas R (2009) Multi-site analysis reveals widespread antibiotic resistance in the marine pathogen Vibrio vulnificus. Microb Ecol 57:151–159
Baquero F, Martinez JL, Canton R (2008) Antibiotics and antibiotic resistance in water environments. Curr Opin Biotechnol 19:260–265
Brunne D (1986) Metal release from dental biomaterials. Biomaterials 7:163–175
Catauro M, Raucci MG, De Gaetano FD, Marotta A (2004) Antibacterial and bioactive silver-containing Na2O × CaO × 2SiO2 glass prepared by sol-gel method. J Mater Sci Mater Med 15:831–837
Charley RC, Bull AT (1979) Bioaccumulation of silver by a multispecies community of bacteria. Arch Microbiol 123:239–244
Chopra I (2007) The increasing use of silver-based products as antimicrobial agents: a useful development or a cause for concern? J Antimicrob Chemother 59:587–59
Clardy J, Fischbach M, Currie C (2009) The natural history of antibiotics. Curr Biol 19:R437–R441
Crabtree JH, Burchette RJ, Siddiqi RA, Huen IT, Handott LL, Fishman A (2003) The efficacy of silver-ion implanted catheters in reducing peritoneal dialysis-related infections. Perit Dial Int 23:368–74
D’Costa VM, Griffiths E, Wright GD (2007) Expanding the soil antibiotic resistome: exploring environmental diversity. Curr Opin Microbiol 2007(10):481–489
Davies J, Davies D (2010) Evolution of antibiotic resistance. Microbiol Mol Biol Rev 74:417–433
Davis IJ, Richard H, Mullany P (2005) Isolation of silver- and antibiotic-resistant Enterobacter cloacae from teeth. Oral Microbiol Immunol 20:191–194
Davison J (1999) Genetic exchange between bacteria in the environment. Plasmid 42:73–91
Dubas ST, Kumlangdudsana P, Potiyaraj P (2006) Layer-by-layer deposition of antimicrobial silver nanoparticles on textile fibers. Colloid Surf A Physicochem Eng Asp 289:105–109
Dykhuizen DE (1998) Santa Rosalia revisited: why are there so many species of bacteria? Antonie Van Leeuwenhoek 73(25–33):1998
Enne VI, Livermore DM, Stephens P, Hall LMC (2001) Persistence of sulphonamide resistance in Escherichia coli in the UK despite national prescribing restriction. Lancet 357:1325–1328
Falletta E, Bonini M, Fratini E, Nostro AL, Pesavento G, Becheri A, Nostro PL, Canton P, Baglioni P (2008) Clusters of poly(acrylates) and silver nanoparticles: structure and applications for antimicrobial fabrics. J Phys Chem 112:11758–11766
Franke S, Gras G, Nies DH (2001) The product of the ybdE gene of the Escherichia coli chromosome is involved in detoxification of silver ions. Microbiology 147:965–972
Gao Y, Cranston R (2008) Recent advances in antimicrobial treatments of textiles. Tex Res J 2008(78):60–72
Gupta A, Matsui K, Lo JF, Silver S (1999) Molecular basis for resistance to silver cations in Salmonella. Nat Med 5:183–188
Gupta A, Phung LT, Taylor DE, Silver S (2001) Silver resistance genes in plasmids of the IncHII incompatibility group and on the Escherichia coli chromosome. Microbiology 147:3393–3402
Haefeli C, Franklin C, Hardy K (1984) Plasmid-determined silver resistance in Pseudomonas stutzeri isolated from a silver mine. J Bacteriol 158:389–392
Heuer H, Smalla K (2007) Manure and sulfadiazine synergistically increased bacterial antibiotic resistance in soil over at least two months. Environ Microbiol 9:657–666
Kim JS, Kuk E, Yu KN, Kim J, Park SJ, Lee HJ, Kim SH, Park YK, Park YH, Hwang C, Kim YK, Lee YS, Jeong DH, Cho M (2007) Antimicrobial effects of silver nanoparticles. Nanomedicine 3:95–101
Klasen HJ (2000) Historical review of the use of silver in the treatment of burns. Part I. Early uses. Burns 26:117–130
Li Y, Leung P, Yao L, Song QW, Newton E (2006) Antimicrobial effect of surgical masks coated with nanoparticles. J Hosp Infect 62:58–63
Lorenz MG, Wackernagel W (1994) Bacterial gene-transfer by natural genetic-transformation in the environment. Microbiol Rev 58:563–602
MacLean RC, Hall AR, Perron GG, Buckling A (2010) The evolution of antibiotic resistance: Insight into the roles of molecular mechanisms of resistance and treatment context. Discovery Medicine. http://www.discoverymedicine.com/R-Craig-MacLean/2010/08/04/the-evolution-of-antibiotic-resistance-insight-into-the-roles-of-molecular-mechanisms-of-resistance-and-treatment-context/
Mahendra Rai M, Yadav A, Gade A (2009) Toxic Silver nanoparticles as a new generation of antimicrobials. Biotechnol Adv 27:76–83
Mandal DME, Bolander D, Mukhopadhyay GS, Mukherjee P (2006) The use of microorganisms for the formation of metal nanoparticles and their application. Appl Microbiol Biotechnol 69:485–492
Mazzola L (2003) Commercializing nanotechnology. Nat Biotechnol 21:1127–1143
McHugh SL, Moellering RC, Hopkins CC, Swartz MN (1975) Salmonella typhimurium resistant to silver nitrate, chloramphenicol, and ampicillin. Lancet 1:235–240
Neal AL (2008) What can be inferred from bacterium–nanoparticle interactions about the potential consequences of environmental exposure to nanoparticles? Ecotoxicology 17:362–371
O’Brien T (2002) Emergence, spread, and environmental effect of antimicrobial resistance: how use of an antimicrobial anywhere can increase resistance to any antimicrobial anywhere else. Clin Infect Dis 34:S78–84
Ochman H, Lawrence JG, Groisman EA (2000) Lateral gene transfer and the nature of bacterial innovation. Nature 405:299–304
Ohshima M (2003) Control and design problems in material processing—how can process systems engineers contribute to material processing? J Process Cont 7:599–605
Pei RT, Kim SC, Carlson KH, Pruden A (2006) Effect of River Landscape on the sediment concentrations of antibiotics and corresponding antibiotic resistance genes (ARG). Water Res 40:2427–2435
Percival SL, Bowlera PG, Russell D (2005) Bacterial resistance to silver in wound care. J Hosp Infect 60:1–7
Sampath LA, Chowdhury N, Caraos L, Modak SM (1995) Infection resistance of surface modified catheters with either shortlived or prolonged activity. J Hosp Infect 30:201–210
Saengkiettiyut K, Rattanawaleedirojn P, Sangsuk S (2008) A study on the antimicrobial efficacy of nano silver containing textile. CMU J Nat Sci Special Issues on Nanotechnology 7:33–36
Schrag SJ, Perrot VR, Levin BR (1997) Adaptation to the fitness costs of antibiotic resistance in Escherichia coli. Proc R Soc Lond B 264:1287–1291
Serov IN, Zhabrev VA, Margolin VI (2003) Problems of nanotechnology in modern materials science. Glass Phys Chem 29:169–178
Shrivastava S, Bera T, Roy A, Singh G, Ramachandrarao P, Dash D (2008) Characterization of enhanced antibacterial effects of novel silver nanoparticles. Nanotechnology 18:103–112
Silver S, Phung LT (1996) Bacterial heavy metal resistance: new surprises. Annu Rev Microbiol 50:753–789
Silver S (1998) Genes for all metals—a bacterial view of the periodic table: The 1996 Thom Award Lecture. J Ind Microbiol Biotechnol 20:1–12
Silver S (2003) Bacterial silver resistance: molecular biology and uses and misuses of silver compounds. FEMS Microbiol Rev 27:341–353
Silver S, Phung LT (1996) Bacterial heavy metal resistance: new surprises. Annu Rev Microbiol 50:753–89
Silver S, Phung LT, Silver G (2006) Silver as biocides in burn and wound dressings and bacterial resistance to silver compounds. J Ind Microbiol Biotechnol 33:627–634
Smalla K, Haines AS, Jones K, Krogerrecklenfort E, Heuer H, Schloter M, Thomas CM (2006) Increased abundance of IncP-1 beta plasmids and mercury resistance genes in mercury-polluted river sediments: first discovery of IncP-1 beta plasmids with a complex mer transposon as the sole accessory element. Appl Environ Microbiol 72:7253–7259
Sobecky PA (1999) Plasmid ecology of marine sediment microbial communities. Hydrobiologia 401:9–18
Stepanauskas R, Glenn TC, Jagoe CH, Tuckfield RC, Lindell AH, McArthur JV (2005) Elevated microbial tolerance to metals and antibiotics in metal-contaminated industrial environments. Environ Sci Technol 39:3671–3678
Stepanauskas R, Glenn TC, Jagoe CH, Tuckfield RC, Lindell AH, King CJ, McArthur JV (2006) Coselection for microbial resistance to metals and antibiotics in freshwater microcosms. Environ Microbiol 8:1510–1514
Summers A, Wireman J, Vimy MJ, Lorscheider FL, Marshall B, Levy SB, Bennett S, Billard L (1993) Mercury released from dental silver fillings provokes an increase in mercury-resistant and antibiotic-resistant bacteria in oral and intestinal floras of Primates. Antimicrob Agents Chemother 37:825–834
Szczepanowski R, Bekel T, Goesmann A, Krause L, Krömeke H, Kaiser O, Eichler W, Pühler A, Schlüter A (2008) Insight into the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to antimicrobial drugs analysed by the 454-pyrosequencing technology. J Biotechnol 136:54–64
Tomšič B, Simončič B, Orel B, Žerjav M, Schroers H, Simončič A, Samardžija Z (2009) Antimicrobial activity of AgCl embedded in a silica matrix on cotton fabric. Polymers 75:618–626
Turner SL, Bailey MJ, Lilley AK, Thomas CM (2002) Ecological and molecular maintenance strategies of mobile genetic elements. FEMS Microbiol Ecol 42:177–185
Walsh CT, Wright G (2005) Introduction: antibiotic resistance. Chem Rev 105:391–394
Wang ZL (2000) Characterizing the structure and properties of individual wire-like nanoentities. Adv Mater 12:1295–1298
Weitz JS, Hartman H, Levin SA (2005) Coevolutionary arms races between bacteria and bacteriophage. PNAS 102:9535–9540
Whitman WB, Coleman DC, Wiebe WJ (1998) Prokaryotes: the unseen majority. Proc Natl Acad Sci USA 95(12):6578–83
Yurieva O, Kholodii G, Minakhin L, Gorlenko Z, Kalyaeva E, Mindlin S, Nikiforov V (1997) Intercontinental spread of promiscuous mercury-resistance transposons in environmental bacteria. Mol Microbiol 24:321–329
Acknowledgements
Preparation of this chapter was partially supported by the US Department of Energy under Award Number DE-FC09-07SR225056 to the University of Georgia Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
McArthur, J.V., Tuckfield, R.C., Baker-Austin, C. (2012). Antimicrobial Textiles. In: Coates, A. (eds) Antibiotic Resistance. Handbook of Experimental Pharmacology, vol 211. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-28951-4_9
Download citation
DOI: https://doi.org/10.1007/978-3-642-28951-4_9
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-28950-7
Online ISBN: 978-3-642-28951-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)