Abstract
The genus Cucumis comprises about 33 species with a major native diverisity center in tropical and southern Africa. It includes some commercially important and widely grown vegetables such as cucumber and melon. The Wild species of Cucumis is a reservoir of potentially useful genes for genetic improvement of cucumber and melon. These Wild species represent a broad genetic base of valuable characters, such as resistance to diseases, pests, and abiotic stress, which are not present in cucumber and melon. In spite of their importance, pace of utilization of this wild group has been rather slow. In addition to their cross-incompatibility problems with cultivated species, there is a lack of information on various aspects of these wild relatives. However, the first successful interpecific hybridization between wild species Cucumis hystrix and cultivated cucumber (Cucumis sativus L.) provided a new opportunity to get insight into the wild species. In this chapter, a comprehensive description on various aspects of wild Cucumis species including taxonomy, morphology, origin, distribution, cytology, evolution and phylogenetics, genomics, as well as the utilization in cucumber improvement are presented for the first time. Future objectives are also proposed for further exploitation and utilization of these important wild relatives.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
6.1 Introduction
The genus Cucumis L. is one of the important genera of flowering plants. It includes cucumber (C. sativus L.) and melon (C. melo L.), two of the most economically important and widely cultivated vegetable crops in the world (Pitrat et al. 1999). However, cucumber and melon suffer from a range of devastating fungal, bacterial, viral, and insect diseases (Whitaker and Davis 1962). The wild Cucumis species are of economic interest because they are a reservoir of potentially useful genes such as biotic stress resistances. A comprehensive knowledge of these wild species is of great importance for conservation and utilization of genetic resources that can be employed for cucumber and melon improvement.
6.2 Basic Botany of the Genus Cucumis
6.2.1 Taxonomy
Cucumis belongs to the family Cucurbitaceae, subfamily Cucurbitoideae, and is currently placed in the tribe Benincaseae (Jeffrey 2005). According to Kirkbride (1993), the genus Cucumis is represented by 32 species, among which two species of Cucumis, C. sativus L. (Cucumber) and C. melo L. (melon), are of great commercial importance. Besides cucumber and melon, the species C. anguria (West Indian gherkin) and C. metuliferus (African horned cucumber) are commercially cultivated in several areas as well (Garcia-Mas et al. 2004). Other wild species originating mostly from arid and/or semi-arid regions of Africa are cultivated as ornamental plants (e.g., C. dipsaceus – “hedgehog gourd” and C. myriocarpus – “gooseberry gourd”) (Rubatzky and Yamaguchi 1997).
Taxonomy of Cucumis was first described by Linnaeus in 1753 (Ghebretinsae et al. 2007). According to this classification, the genus Cucumis contains seven species, all of which were cultivated or economically useful. There have been a number of taxonomic placements of Cucumis since the work of Linnaeus (Pangalo 1950; Jeffrey 1962, 1967, 1980, 1990; Kirkbride 1993; Schaefer 2007).
The most comprehensive placement of Cucumis was proposed by Kirkbride (1993). On the basis of his investigations, the genus Cucumis was divided into two subgenera with different geographical origin and basic chromosome numbers. Subgenus Melo (30 spp., n = 12) was originated in Africa and was partitioned into two sections (Melo and Aculeatosi), whereas subg. Cucumis (two spp., n = 7) was originated in Asia. A detailed taxonomic depiction of the genus Cucumis elaborated by Kirkbride (1993) is given in Table 6.1. However, this taxonomic treatment was challenged by the rediscovery of C. hystrix, a wild Cucumis species of Asian origin possessing 24 chromosomes. C. hystrix (2n = 24) was successfully crossed with cucumber (C. sativus, 2n = 14) (Chen et al. 1997). A new species C. × hytivus Chen and Kirkbride was proposed in 2000 followed by chromosome doubling of the F1 hybrid (Chen and Kirkbride 2000).
Recently, on the basis of molecular phylogenetic studies, 19 species of five genera including Cucumella, Dicoelospermum, Mukia, Myrmecosicyos, and Oreosyce have been transferred to Cucumis, resulting in 14 new combinations, two changes in status, and three new names (Cucumis indicus, C. kirkbrideana, and C. oreosyce). A complete morphological key to all these species now included in Cucumis was provided in Schaefer (2007).
6.2.2 Morphology
The genus Cucumis includes annual and perennial taxa, and the fruit morphology is the most important character within the genus (Fig. 6.1). Following morphological descriptions are given by combining the data from Kirkbride (1993), Rubatzky and Yamaguchi (1997), and Kristkova et al. (2003).
6.2.2.1 Plants
Herbs, exceptionally semi-shrubs, usually having a trailing or climbing growth habit, are monoecious, or rarely dioecious or andromonoecious; root systems are rarely woody (C. trigonus) and extensive, but usually shallow and rarely tuberous (C. kalahariensis); stems are angled, sulcate, not aculeate or rarely aculeate (C. aculeatuc and C. ficifolius), and variously pubescent or rarely glabrous, with non-breakaway hairs or rarely breakaway hairs (C. sacleuxii); nodes are geniculate or not geniculate. Each node has a single leaf and a simple tendril (sometimes curling), except that of C. humifructus, which has a fasicicle of five to eight tendrils, and that of C. rigidus, which lacks them; tendrils of C. insignis are either simple or bifid. Tendrils are variously pubescent, rarely glabrous, or rarely aculeate.
6.2.2.2 Leaves
Simple and petiolate. Petioles vary in length (with regard to the length of a leaf blade). They are not aculeate or rarely aculeate and variously pubescent or rarely glabrous, with non-breakaway hairs or rarely breakaway hairs. The majority of species have a uniform type of pubescence on the petioles. C. sagittatus and C. thulinianus have two pubescence types uniformly mixed over the entire petiole, and C. myriocarpus has three types separated into distinct zones on the petioles, retrose–strigose on the base, hirsute in the middle, and antrorse–strigose at apex; leaf blades are 3- or 5-palmately lobed, trilobite, pentalobate, heptalobate, or entire; Central leaf lobe is symmetrical, entire, or sometimes pinnatifid; lateral leaf lobes are asymmetrical, or sometimes symmetrical, entire, or sometimes pinnatifid.
6.2.2.3 Inflorescences and Flowers
The inflorescence is unisexual and most species are monoecious. C. humifructus has only androgynous inflorescences (i.e., inflorescence with both male and female flowers and the female flower below the male ones), and C. metuliferus has mainly unisexual inflorescence and a few gynecandrous ones (i.e., inflorescence with both female and male flowers and the female flower above the male ones).
Male inflorescence consists of solitary flowers, or fasciculate, racemose, paniculate, or rarely modified compound dichasial from 1 to 18 flowered, sessile, or rarely pedunculate. Male inflorescences are often multiflowered and rarely branched. When the inflorescences are branched, the male flowers are always pedicellate. Male flowers are 5-merous; pedicel is terete or rarely sulcate in cross section, variously pubescent or rarely glabrous, and without bracteoles or rarely subtended by a bracteole (C. heptadactylus). Calyx consists of five or rarely four lobes, linear to oblong, or narrowly to broadly triangular in outline, acute to narrow in the apex, and variously pubescent or rarely glabrous. Corolla is yellow, infundibular, or rarely campanulate and is variously pubescent or rarely glabrous. Corolla is fused into a basal tube. Corolla leaves are elliptic to broad, ovate to shallow, obovate to narrowly, or rarely oblong or broadly triangular in outline, narrowly to broadly acute or obtuse, and sometimes also mucronate at the apex. Three stamens are free, with separation from the free portion of the hypanthium above the ovary. Two of them are 2-thecate and one is 1-thecate. Filaments are terete or radially compressed in cross section and are glabrous or with basal puberculence and glabrous apically. Anther theceae is sigmoid and glabrous with the edges shortly pubescent. Anther connective is extended, obovate, oblong to narrow, transversely broadly oblong, or ovate, unilobate or rarely bilobate, obtuse or rarely acute in the apex, minutely papillate, sometimes smooth, or rarely glabrous, fimbriate, or crenulate at the apex. Disk is cylindrical or rarely consisting of three papillae and is glabrous.
Female flowers are solitary or rarely in fasciclate inflorescences; sessile flowers arise from leaf axils, very often from secondary branches. They are pedicellate and 5-merous. Pedicel is terete or sulcate in cross section and is variously pubescent, with non-breakaway hairs or rarely with breakaway hairs. Hypanthium is hourglass-shaped. The constricted portion and the lower bulge fused to the ovary. The upper bulge of hypanthium is free from the ovary. Free portion of hypanthium is campanulate. Ovary has three to five placentas with numerous horizontal ovules. Calyx has five, occasionally four or six lobes of the same shape as male flowers. Corolla is yellow and infundibular, with the same shape and types of pubescence as male flower. Corolla tube is present or absent. Three staminodes are present or rarely absent, separating from the free portion of hypanthium above the ovary. Style is terete in cross section, glabrous, subtented by a circular disk, or rarely lacking one. Stigma is copular, lobate, or sometimes entire or sublobate, with one to six or rarely nine finger-like projections on the margin.
6.2.2.4 Fruits and Seeds
Fruits are pendulous; fruit is spherical, oval, oblong, elongated, or blocky in shape and variable in size; fruit surface varies in the number and size of scattered spiny tubercles (warts), or sharp soft hairs. It can be smooth and glabrous, sometimes deeply ridged or covered with a corky (reticulate) netting (e.g., for C. melo); skin color varies from pale to very dark green, sometimes with longitudinal indentations or stripes. In maturity, the skin color is white cream to orange brown. Inferior flesh color can be white, green, pink, or orange. The fruit stalk is referred to as a pedicel. The pedicel is sulcate or sometimes terete in cross section and is variously pubescent or rarely glabrous.
Mature seeds have white, cream to yellow color. They are smooth, compressed, ovoid to elliptic, immarginate, with an acute edge, and unwinged or rarely apically winged. C. humifructus develops its fruits below ground.
6.2.3 Cytology
Cytologically, the genus Cucumis, like all other Cucurbits, is a less studied genus (Ramachandran and Narayan 1985). Most Cucumis species are diploid with 12 pairs of chromosomes (2n = 24): C. africanus, C. anguria, C. dipsaceus, C. ficifolius, C. hirsutus, C. humifructus, C. metiluferus, C. myriocarpus, C. melo, C. prophetarum, C. pustulatus, C. sagittatus, C. sacleuxii, C. zeyheri, and C. hystrix. Among these species, three have also been reported to be polyploid: C. ficifolius, 2n = 48 (Dane and Tsuchiya 1979; den Nijs and Visser 1985), C. pustulatus, 2n = 48 (Ramachandran 1984; den Nijs and Visser 1985; Ramachandran and Narayan 1985) or 2n = 72 (Dane and Tsuchiya 1979), and C. zeyheri, 2n = 48 (Dane and Tsuchiya 1976, 1979; Varekamp et al. 1982; Ramachandran 1984; den Nijs and Visser 1985; Ramachandran and Narayan 1985). C. sativus is the only species of Cucumis reported to have a chromosome count of 2n = 14 (Fig. 6.2).
There are two base chromosome numbers in Cucumis: x = 7 and x = 12. Two different hypotheses have been put forward to explain the relationship between the two basic chromosome numbers. The fragmentation hypothesis suggests that x = 12 has derived from x = 7 by fragmentation of particular chromosomes followed by de novo regeneration of centromeres (Bhaduri and Bose 1947; Ayyangar 1967). The fusion hypothesis, on the other hand, says that the basic number x = 7 might have arisen from x = 12 possibly by unequal translocation or fusion of non-homologous chromosomes (Trivedi and Roy 1970). Comparative genomics between C. melo and C. sativus may clarify the phylogeny of these species (Danin-Poleg et al. 2001).
As for the karyotype of Cucumis, most studies have focused on the cultivated species: cucumber and melon (Figs. 6.3 and 6.4). However, discriminatory information from karyotype analysis for detailing relationships in Cucumis has been difficult to access due to the small chromosome size and poor stainability. Ramachandran and Seshadri (1986) used C-banding and pachytene analysis to compare the genomes of cucumber and muskmelon (C. melo L.), but their study did not differentiate chromosomes by measurement, and their description of the chromosome morphology and C-banding figures are equivocal.
Karyotypes (A1, A2) and their ideograms (B1, B2) of Jiashi and Huangjin melon according to Zhang et al. (2005)
Chromosomal DNA amounts varied in different species of Cucumis (Ramachandran and Narayan 1985). The DNA amounts varied from 1.373 to 2.483 pg in diploids and from 2.846 to 3.886 pg in tetraploids. DNA amount was not correlated with chromosome number and periodicity. Tetraploids were found to have double the quantity of nuclear DNA of diploids.
6.2.4 Origin and Distribution
The center of origin for Cucumis species is likely Africa for the most wild species with chromosome 2n = 24, while the Middle East and southern Asia has been considered an important center of diversification for melon and cucumber, respectively (Dane et al. 1980; McCreight et al. 1993; Staub et al. 1999).
Cucumis species occurred in a large scale from 38°N to 37°S of the Old World, with more species in the southern hemisphere than in the northern hemisphere. With latitude rising, Cucumis species sharply decreased in the northern hemisphere. There were abundant species near the equator. Cucumis species occurred in about 75 countries, but 90.6% of them were from Ethiopia, Kenya, Somalia, South Africa, and Tanzania. Somalia had more rare species in absolute and relative. Thirty-three of 75 countries had only one species (Liu 2007). Table 6.2 presents the details of the distribution of Cucumis species.
6.3 Germplasm Conservation
There are some centers, which are engaged in the conservation of Cucumis germplasm worldwide. In the United States, plant germplasm is maintained and evaluated by the US National Plant Germplasm System (NPGS). In Europe, the International Plant Genetic Resources Institute (IPGRI) coordinates institutional germplasm holdings. In China, the Crop Germplasm Resources Institute of Chinese Academy of Agricultural Sciences (CAAS) is responsible for the germplasm conservation.
Germplasm information can be found in some good germplasm resources information network web servers, such as Germplasm Resources Information network of United States (http://www.ars-grin.gov), Chinese Crop Germplasm Resources Information System (http://icgr.caas.net.cn), N.I. Vavilov Research Institute of Plant Industry of Russia (http://www.vir.nw.ru), and the European Central Cucurbits Database (http://www.comav.upv.es).
According to the American germplasm resources information network, the regional plant introduction (PI) station of NPGS at Ames, Iowa, houses about 1,486 C. sativus accessions of worldwide origin and currently lists 3,074 accessions in its melon inventory. The collection of wild species of Cucumis includes 48 C. africanus L. f. accessions, 50 C. anguria L. accessions, 10 C. anguria var. anguria accessions, 17 C. anguria var. longaculeatus J. H. Kirkbr. accessions, one C. asper Cogn. accession, one C. canoxyi Thulin & Al-Gifri accession, six C. dipsaceus Ehrenb. ex Spach accessions, seven C. ficifolius A. Rich. accessions, one C. heptadactylus Naudin accession, three C. hirsutus Sond. accessions, one C. meeusei C. Jeffrey accession, 42 C. metulifer E. Mey. ex Naudin accessions, 21 C. myriocarpus Naudin accessions, two C. myriocarpus subsp. leptodermis (Schweick.) C. Jeffrey & P. Halliday accessions, three C. myriocarpus subsp. myriocarpus accessions, three C. prophetarum L. accessions, seven C. pustulatus Hook. f. accessions, four C. sagittatus Peyr. accessions, one C. subsericeus Hook. f. accession, eight C. zambianus Widrlechner et al. accessions, and nine C. zeyheri Sond. accessions. The collection also has 88 accessions labeled Cucumis sp., which may include some C. melo or C. metuliferous accessions (http://www.ars-grin.gov/cgi-bin/npgs/html/genform.pl; Table 6.3).
According to the AD HOC meeting on Cucurbit genetic resources held in Turkey in 2002, the number of accessions of Cucumis species, including landraces, breeding material, and wild relatives, maintained in European collections was 14,333. Among them, there are 33 accessions of C. anguria, 31 accessions of C. dipsaceus, 11 accessions of C. ficifolius, 7,553 accessions of C. melo, 11 accessions of C. metuliferus, 12 accessions of C. myriocarpus, 5,896 accessions of C. sativus, 10 accessions of C. zeyheri, and 776 accessions of C. spp. (Table 6.4).
In China, it is reported that there are 1,506 accessions of C. sativus (Sheng et al. 2006) and 1,003 accessions of C. melo (Ma et al. 2003); however, the collection of wild species is not clear.
Although Cucumis accessions are held by numerous collections around the World, the most wild species are less formal collections for research purposes and through personal exchanges among scientists throughout the world. Those wild species are often not documented or represented in the NPGS base collection or IPGRI collection, so samples acquired through personal contact could be important.
6.4 Evolution and Phylogenetic Relationships
Knowing the closest relatives and natural composition of the genus Cucumis L. is important simply because of the ongoing efforts by plant breeders worldwide to improve melon and cucumber with traits from wild relatives (Renner and Schaefer 2008). Quite a few studies, using morphological, cytology, and molecular characters such as isozymes, random amplified polymorphic DNA (RAPD), chloroplast simple sequence repeat (cpSSR), and internal transcribed spacer (ITS), have been carried out to determine the Cucumis phylogeny.
Early studies on karyomorphological investigations of 13 species in the genus Cucumis L. indicated that South African annual species are the primitive and identified five distinct groups in taxa with 2n = 24. They are (1) C. leptodermis and C. africanus; (2) C. ficifolius, C. hookeri, and C. dipsaceus; (3) C. myriocarpus, C. zeyheri, C. prophetarum, Cucumis species CUCU44/74, and C. anguria; (4) C. metuliferus; and (5) C. sagittatus and C. melo (Singh and Yadava 1984).
The crossability between species, chromosome pairing, and pollen fertility in F1 hybrids were also investigated for assessing species relationships and Cucumis phylogeny. Deakin et al. (1971) were the first to produce a comprehensive and monographic account on relative cross-compatibility between Cucumis species and pollen fertility of their F1 hybrids. Their studies involved 14 species including cultivated C. melo L. On the basis of these data obtained, they grouped Cucumis species into four major groups. Singh and Yadava (1984) had investigations on interspecific crossability in eight Cucumis species (2n = 24, C. melo, C. dipsaceus, C. anguria var. anguria, C. anguria var. longipus, C. myriocarpus, C. zeyheri, C. prophetarum, and C. species). Information on chromosome pairing and pollen fertility of the hybrids from 15 combinations had been utilized for tracing the phylogenetic relationships among these taxa.
Esquinas-Alcazar (1977) studied the alloenzyme variation and relationships in the genus Cucumis and divided the genus Cucumis into four groups. (1) Ser. angurioidei: C. aculeatus, C. africanus, C. anguria, C. dipsaceus, C. ficifolius, C. heptactylus, C. myriocarpus, C. pustulatus, and C. zeyheri; (2) Ser. metuliferi: C. aculeatus, C. metuliferus, and C. sagittatus (some accessions); (3) Ser melo: C. melo and C. sagittatus (some accessions); and (4) Ser. sativus: C. sativus.
From the evolutionary and systematic point of view, Perl-Treves and Galun (1985) compared the phylogenies of Cucumis based on cpDNA and nuclear-coded isozymes. The comparison was carried out for 21 Cucumis species and the two phylogenies were found to share the main dendrogram features, which also agreed well with most taxonomic data available on Cucumis. Accordingly, most of the African Cucumis species form a close group (“Anguria group,” C. africanus, C. anguria, C. dipsaceus, C. ficifolius, C. heptactylus, C. meesusei, C. myriocarpus, C. prophetarum, C. pustulatus, and C. zeyheri), which was distant from the melon (C. melo), and from a few other distant species (C. humifructus, C. metuliferus, and C. sagittatus), all of which were far apart from each other. The cucumber (C. sativus) was the most distant species within the genus.
In 1989, C. hystrix Chakr., a wild Cucumis species, was rediscovered in Yunnan, China, by Jinfeng Chen (Chen et al. 1994). Subsequent research revealed that C. hystrix has 2n = 24 instead of 2n = 14 as in the Asian members. This finding challenges the basic chromosome number theory that African Cucumis have n = 12 and that Asian Cucumis have n = 7. Isozyme patterns (Chen et al. 1995) suggested that C. hystrix has closer genetic affinities with C. sativus than with C. melo, even though C. hystrix and C. melo possess the same number of chromosomes.
Chung et al. (2006) used nine chloroplast SSR (cpSSR) markers to investigate the phylogenetic relationships among African Cucumis species (x = 12) accessions, C. melo accessions, C. sativus accessions, and C. hystrix accessions. Sequence variation analysis identified a group of African Cucumis species and a group composed of C. melo, C. sativus, and C. hystrix species leading to the conclusion that C. hystrix is the progenitor species of C. sativus, or that they at least share a common ancestral lineage.
Zhuang et al. (2006) investigated the phylogenetic relationships in Cucumis species using RAPD. Their focus was mainly on the analysis of genetic relationship among C. hystrix, C. sativus, and C. melon and C. × hytivus, a new synthetic species. On the basis of results, a modified taxonomic system was proposed that C. hystrix should remain in subgen. Cucumis, although it had a chromosome number different from that of C. sativus. With the interspecific hybrids C. × hytivus as the third species, subgen. Cucumis was thus made up by three species. Although the basic chromosome number and geographic location theorized were challenged by the proposed system, the use of it will likely assist in the exploitation of the wild Cucumis species in Asia.
Using sequences of the internal transcribed spacer (ITS) 1 and 2 regions of the nuclear ribosomal RNA genes, Jobst et al. (1998) evaluated the phylogenetic relationships among different members of the family Cucurbitaceae. Six Cucumis species along with C. melo and C. sativus were analyzed in the study and the results obtained by ITS sequence data were highly congruent with isoenzyme data of Puchalski and Robinson (1990). Garcia-Mas et al. (2004) defined phylogenetic relationships among Cucumis species also using the nuclear ribosomal DNA ITS region and microsatellite markers. In their study, the genus Cucumis was splited into five groups: cucumbers, melons, C. metuliferus, a group containing 12 wild species of Cucumis, and Oreosyce Africana, and a fifth group comprising C. sagittatus and C. globosus.
Kocyan et al. (2007) presented a multilocus chloroplast phylogeny for the Cucurbitaceae that included all putative close relatives of Cucumis. Their results support a paraphyletic Cucumis, with Cucumela, Dicaelospermum, Mukia, Myrmecosicyos, and Oreosyce nested among species of this genus. Although evolutionary relationships are not completely resolved, following the discovery by Koeyan and coworkers that Cucumis as traditionally circumscribed (Kirkbride 1993) was highly unnatural, two molecular phylogenetic studies reinvestigated species relationships in a much more broadly circumscribed Cucumis (Ghebretinsae et al. 2007; Renner et al. 2007).
Ghebretinsae et al. (2007) presented a comprehensive molecular phylogeny of Cucumis and the traditionally related genera based on sequences from both nuclear and chloroplast genomes. Their study used a much more complete sampling of species within Cucumis than did previous studies and includes representatives of Cucumella, Dicaelospermum, Mukia, Myrmecosicyos, and Oreosyce. Their combined phylogenetic analyses did not support Kirkbride’s (1993) subdivision of Cucumis into two subgenera based principally upon chromosome number and geographical distribution. Cucumis sensu Kirkbride (1993) is paraphyletic and the Asian Cucumis (subg. Cucumis of Kirkbride) forms a well-supported clade nested within African Cucumis s.s. They identified six clades within the Cucumis complex, designated as Clades I, II, III, IV, V, and VI. Clade I comprised all non-domesticated African Cucumis s.s. after the exclusion of C. hirsutus, C. humifructus, C. metuliferus, C. rostratus, and C. sagittatus; C. hystrix and C. sativus were included in Clade II. Clade III comprised C. melo and C. sagittatus. Clade IV consisted of C. metuliferus and C. rostratus. Clade V comprised four species of Cucumella and Oreosyce. Clade VI consisted of C. hirsutus and C. humifructus.
Based on these two recent molecular phylogenetic studies of Cucumis, Renner and Schaefer (2008) summarize what is now known about phylogenetic relationships in Cucumis. The phylogeny of Cucumis resulting from combined nuclear and chloroplast data (Fig. 6.5) implied that the deepest divergence lies between the common ancestor of C. hirsutus/C. humifructus and the stem lineage of the remainder of the genus. The area of origin of Cucumis cannot be inferred because its sister genus, Muellerargia, has one species in Madagascar and the other one in tropical Australia and Indonesia. The next closest relatives are in African/Asian clades including the genera Coccinia, Zehneria, Neoachmandra, and Peponium, but their exact position is still unresolved. The earliest divergence events in Cucumis likely took place in Africa. However, contrary to the traditional classification (Kirkbride 1993), which grouped C. melo with the African C. hirsutus, C. humifructus, and C. sagittatus, melon instead is closest to an Australian/Asian clade.
Parsimony tree for Cucumis based on combined chloroplast and nuclear DNA sequences according to Renner et al. (2007)
6.5 Interspecific Hybridization Among Cucumis Species
6.5.1 Progress of Interspecific Hybridization
Interspecific hybridization is used to improve crops by transferring specific traits, such as pest and stress resistance, from their wild relatives (Bowley and Taylor 1987). Interspecific hybrids in the Cucurbitaceae have been produced in several genera, including Cucumis (Deakin et al. 1971), Citrullus (Valvilov 1925), Luffa (Singh 1991), and Cucurbita (Weeden and Robinson 1986). In the genus Cucumis, an amphidiploid was reported from the cross of C. anguria L. and C. dipsaceus E. ex S. (Yadava et al. 1986). However, in the Cucurbitaceae only in Cucurbita interspecific hybridization has been successfully utilized for crop improvement (Robinson and Decker-Walters 1997).
Cucumis contains two species of economic importance: melon (C. melo L., 2n = 24) and cucumber (C. sativus L., 2n = 14). The importance of wild Cucumis species has long been recognized because they possess resistance to pathogens, such as powdery mildew, downy mildew, anthracnose, and Fusarium wilt (Leppick 1966; Lower and Edwards 1986; Kirkbride 1993). Genetic variation is relatively limited in cucumber; thus, efforts to create interspecific hybrids have become more critical and meaningful.
The first recorded attempt to make crosses between cucurbits by removing the male flowers and transferring pollen by hand was made by Naudin. He was unsuccessful in obtaining a cross between C. melo and C. myriocarpus (Naudin 1859). Many more studies have also been attempted to make interspecific crosses in Cucumis (Betra 1953; Andrus and Fassuliotis 1965; Deakin et al. 1971; Chelliah and Sambandam 1972; Fassuliotis 1977; Dane et al. 1980; Kho et al. 1980; Visser and den Nijs 1983; Singh and Yadava 1984; den Nijs and Visser 1985; den Nijs and Custers 1990; Chatterjee and More 1991, etc.).
The first comprehensive crossability analysis of the genus was published by Deakin et al. (1971), who observed that crosses among wild species were frequently possible, but that all attempts to cross any of these with the two cultivated species, C. sativus and C. melo, failed. Raamsdonk et al. (1989) summarized the data in two crossing polygons. The species C. heptadactylus, C. humifructus, C. melo, and C. sativus have never been successfully crossed with any other species of Cucumis to produce a fertile F1 generation. The following species can be crossed to a limited extent among themselves: C. africanus, C. anguria, C. dipsaceus, C. ficifolius, C. metuliferus, C. myriocarpus, C. prophetarum, C. pustulatus, and C. zeyheri (Kirkbride 1993). Some wide-cross attempts between cultivated and wild Cucumis species are presented in Table 6.5.
A successful interspecific hybridization between C. hystrix and C. sativus was reported (Chen et al. 1997). It was the first reproducible cross between a cultivated Cucumis species and a wild relative, and it represented a breakthough in interspecific hybridization in Cucumis. The success of this cross was even more surprising because the parental species have different chromosome numbers. The original F1 hybrid (2n = 19), obtained by embryo rescue following pollination of C. sativus by C. hystrix (Fig. 6.6), has 7 chromosomes from C. sativus and 12 from C. hystrix and was both male- and female-sterile. To restore fertility, reciprocal crosses were made and the chromosome numbers of the progeny were successfully doubled (Chen et al. 1998). Pollen grains were produced by these progeny when C. hystrix was used as the seed parent; the plants produced fertile flowers and set fruit with viable seeds, indicating that fertility was restored. This restoration of fertility marked the creation of a new synthetic species, which has close phylogenetic relationships with its parental species, but is distinctively different from each parent. The new species (Cucumis × hytivus Chen and Kirkbride; Fig. 6.7) has genome HHCC, where H represents the genome of C. hystrix and C represents the genome of C. sativus and chromosome number 2n = 4x = 38 (Chen and Kirkbride 2000). This synthetic species might be useful as a new Cucumis crop. In addition, as a C. hystrix × C. sativus hybrid, it might be useful as a bridging species for transfer of useful traits to cucumber. Figure 6.8 shows the polygon of crossability in Cucumis species.
Polygon of crossability in Cucumis species according to Zhuang (2003). Arrows point to the female parent; moderately to strongly self-fertile and cross-fertile hybrids (thick solid line); sparingly self-fertile and moderately cross-fertile hybrids (thin solid line); self-fertile, usually not cross-fertile hybrids (dashed and dotted line); inviable seeds or seedlings (dashed line); self-sterile and cross-sterile hybrids (thick dashed line); self-sterile and cross-fertile hybrids (long dashed line); absence of a line indicates that seed fruits were not obtained
6.5.2 Major Problems in Interspecific Hybridization
6.5.2.1 Hybridization Barriers
Many experiments have indicated the presence of a strong barrier to interspecific hybridization in Cucumis. The nature of cross-incompatibility between cultivated Cucumis species and their wild relatives is not well understood. Incompatibility is characterized by delayed growth of pollen, or arrested pollen tube growth to reach the ovules (Kishi and Fujishita 1969), as well as lack of cell division of the zygote, and abortion of the endosperm (Kishi and Fujishita 1970).
Several traditional approaches in interspecific hybridization have been used to overcome the hybridization barriers in Cucumis. These include growth regulator application (Custers and Den Nijs 1986), pollen irradiation (Beharav and Cohen 1994), use of mentor pollen (Kho et al. 1980), and bud pollination (Chatterjee and More 1991). Biotechnological techniques, such as somatic hybridization, have also been applied as possible tools for overcoming these barriers in Cucumis (Tang and Punja 1989; Chatterjee and More 1991). Likewise, fusion of C. sativus and C. melo protoplasts has been attempted, but the results indicated that successful hybridization is still unpredictable (Fellner et al. 1996).
The interspecific hybrid between C. sativus and C. hystrix represents an important step in interspecific hybridization in Cucumis. If C. hystrix and C. melo are cross-compatible and if the F1 derived from either interspecific hybridization can be made fertile through crossing and/or chromosome doubling, then C. hystrix could act as bridge species between C. melo and C. sativus.
6.5.2.2 Post-fertilization Abortion and Embryo Rescue
In higher plants, post-zygotic failure of hybrid embryos is not often due to incompatibility between the parental chromosomes, but incompatibility problems in the endosperm. In such cases, embryos from interspecific hybridization have to be rescued; otherwise, they will fail due to embryo abortion and/or endosperm degeneration. Successful embryo rescue in tissue culture allows further advances in interspecific hybridization.
Embryos can sometimes be rescued, even if they are immature or lack endosperm (Laibach 1925). In Cucumis, fruits with non-viable seeds were obtained in the cross between C. prophetarum L. and C. melo (Singh and Yadava 1984). The barriers between these two species may be post-zygotic. If the embryo rescue technique had been employed, the experiment might have been successful.
6.5.2.3 Sterility in F1 Hybrids
A common problem on utilizing germplasm of wild species for crop improvement was sterility in F1 hybrids. In many cases, this sterility was associated with meiotic abnormalities and was a large obstacle that followed hybridization and hindered utilization.
The ability to cross C. sativus and C. hystrix offered the promise of introgressing desirable characters from C. hystrix to C. sativus. However, self-pollination and backcrossing of the F1 plants to either parent was unsuccessful because the original hybrid was both male- and female-sterile, probably because of the non-functional gametes containing odd chromosome numbers. When chromosomes were doubled, each chromosome had a homologous partner for pairing during meiosis; if there were no cytoplasmic incompatibility, the chromosome-doubled F1 hybrid might have produced viable gametes, and fertility restoration was anticipated.
External application of chemical agents is the usual way to double chromosome number. Among various agents, colchicine was one of the antimitotic substances most frequently used for this purpose (Chen and Staub 1997). Colchicine at an aqueous solution of 0.05–0.5% (w/v) is believed to be most effective for many plant species. Since colchicine is poisonous to plants, germinating seeds or young seedlings are often preferred for treatment because they grow rapidly and recover more readily than more mature plants do.
When the experimental material does not respond well to chemical treatment, in vitro chromosome doubling (spontaneous polyploidy as a consequence of tissue culture) could be an alternative (D’Amato 1977). When and how the polyoloidization happened in tissue culture was not entirely clear, but it occurred at a low rate during plant formation from axillary buds (Adelberg et al. 1994), callus (Osifo et al. 1989) and culture of protoplasts (Tabei et al. 1992). Polyploidization can be generalized as a universal phenomenon in melon tissue culture (Ezura et al. 1992), although genotype is an important factor in determining the rate of chromosome doubling (Adelberg and Chen 1998).
6.6 Utilization of Wild Species
Although there are many wild Cucumis species, not all of them have been fully and properly characterized and documented. Still a few species have been actually used in breeding programs, despite the fact that several of them have been reported to have one or more excellent features to be used as donors (Fassuliotis 1967; Norton and Granberry 1980). Intergroup incompatibility has been assigned as the main reason for this (Deakin et al. 1971; Fassuliotis and Nelson 1988). However, the research work on transfer of useful genes from wild relatives was strengthened with the successful interspecific hybridization between C. hystrix and C. sativus.
6.6.1 Resistance in C. hystrix
C. hystrix is a wild species of Cucumis subgen. Cucumis, which originated in Asia (Kirkbride 1993). It has a sour taste that is not really like cucumber. As it has been described before that although it bears morphological similarity with cucumber, its diploid chromosome number is 24, the same number as in melon (Chen et al. 1997). Since the first interspecific hybridization successfully made in 1997 and development of the synthetic allotetraploid species (C. × hytivus Chen and Kirkbride, 2n = 4x = 38) in 2000, several disease screens were undertaken to characterize the response of C. hystrix and its progenies derived from the interspecific hybridization to common cucurbit diseases (Chen and Lewis 2000; Chen et al. 2004b).
6.6.1.1 Gummy Stem Blight
Resistance evaluations were made in a field at Cornell University using the highly virulent Didymella bryoniae isolate NY1. Resistance was found in C. hystrix plants in the field (Fig. 6.9a, b). Few symptoms were found on stem and slightly more on the leaves. There was no segregation observed among the plants.
6.6.1.2 Downy Mildew
Resistance tests were conducted in Nanjing Agricultural University, China. The disease index in C. hystrix was 5.3, indicating that it is highly resistant to downy mildew. This resistance was partially transmitted to the C. hytivus, and the progenies from backcross. Compared to the susceptible cucumber cultivar “Jinlu,” all the materials derived from this interspecific hybridization possess at least moderate resistance.
6.6.1.3 Viruses
C. hystrix was evaluated for resistance to four viruses: CMV-C, WMV-2, PRV, and ZYMV in the field at Cornell University. C. hystrix plants generally suffered from the WMV-2 inoculation (Fig. 6.9c), but showed stronger resistance to PRV and moderate resistance to CMV-C and ZYMV (Fig. 6.9d).
6.6.1.4 Nematodes
Nematode resistance tests were carried out. The three groups (C. hystrix, C. sativus, and reciprocal interspecific hybrids) varied greatly in their response to M. incognita. C. hystrix had a high level of resistance to M. incognita with mean gall index of 1.8 (Fig. 6.10). In contrast, cucumbers were confirmed as being highly susceptible possessing a mean gall index of 4.8–5.0. The interspecific F1 hybrid was intermediate in resistance to the two parents, with a mean gall index 3.4. The transmission of resistance was observed in backcross progeny of the chromosome-doubled F1 to cucumber.
6.6.2 Synthesis, Characterization, and Utilization of Novel Germplasm
6.6.2.1 Allotetraploid
The first repeatable interspecific hybridization between cucumber (C. sativus L., 2n = 14) and C. hystrix (2n = 24) was successfully made through embryo rescue (Chen et al. 1997). Hybrid plants (2n = 19; 12 from C. hystrix and 7 from cucumber) were sterile, but morphologically uniform. The multiple-branching habit, densely brown hairs (on corolla and pistil), orange–yellow corolla, and ovate fruit of F1 hybrid plants were similar to that of the C. hystrix parent, and the appearance of the first pistillate flower was more similar to that of C. sativus parent. The diameter and internode length of the stem and the shape and size of leaves and flowers were intermediate when compared to the parents.
To restore fertility, chromosome doubling of the F1 hybrid plants was carried out (Chen and Staub 1997). Sixty-two chromosome-doubled plants were obtained. The chromosome-doubled F1 plants were morphologically distinct from the parents and other progeny in traits such as a curve on leaf margins and shorter and stronger internodes. The fruits at two ploidy levels vary in morphology (Fig. 6.11). While diploid fruit (2n = 19, seedless) was longer and spindle-like in shape, the tetraploid fruit was shorter and column-shaped. After fertility selection, two primary allotetraploid produced fertile flowers and set fruit with viable seeds (Chen et al. 1998). The restoration of fertility in the chromosome-doubled F1 hybrid marks the creation of a new combination of genomes and a new synthetic species that did not exist previously.
Morphological comparison between the interspecific hybrid and synthetic allotetroploid C. × hytivus (a and b). The interspecific hybrid F1 diploid, sterile, hybrid plant from embryo rescue (left) and its chromosome-doubled tetraploid, fertile plant (right). (c) Female flowers of F1 hybrid (left) and allotetraploid (right). (d) Fruit of F1 hybrid (left) and allotetraploid (right). (e) Seeds harvested from the allotetraploid and its diploid progenitors
Several photosynthetic characters of the hybrid species Cucumis × hytivus Chen and Kirkbride under weak light condition were studied (Qian et al. 2002). The light compensation point of allotetraploid was 11.25 μE m−2 s−1. After treatment with low intensity light for 2 weeks, the leaf contents of chlorophyll a and b increased, while the value of chlorophyll a/b decreased, indicating that the allotetraploid has good tolerance to low irradiance.
Zhuang et al. (2002) investigated the responses of seedlings of the new species Cucumis × hytivus and its progenies from backcross with cucumber to chilling injury. The abnormal metabolism was observed in C. hytivus as it was subjected to low temperature treatment; however, the progenies from backcrossing the new species to cucumber showed high tolerance to chilling injury.
Chen et al. (2007) studied the genomic events in the early generations of the synthesized allotetraploid. Extensive genomic changes were detected by amplified fragment length polymorphism (AFLP) analysis. The changes mainly involved loss of parental restriction fragments and gain of novel fragments. The total detectable changes were from 11.1 to 32.1%, and the frequency of losing parental fragments was much higher than that of gaining novel fragments. Although no significant differences were detected in the reciprocal crosses, the data showed that the frequency of sequence loss in C. sativus was two times higher than that in C. hystrix. The results demonstrated that the sequence elimination was the major event of genomic changes, and it might provide the physical basis for the diploid-like meiotic behavior in the diploidization of the newly formed allopolyploids.
In order to explore the molecular involvement of epigenetic phenomena, cytosine methylation was investigated in C. × hytivus by using methylation-sensitive amplified polymorphism (MSAP) (Chen and Chen 2008). Twofold difference in the level of cytosine methylation in the reciprocal F1 hybrids and in the allotetraploid was observed. Pattern analysis found that 2.0–6.4% of total sites changed in both the F1 hybrids and the allotetraploid compared to their corresponding parents. 68.2–80.0% of the changed sites showed an increase in cytosine methylation, and most of the methylated sites were from the maternal parent. The extent of cytosine methylation pattern changes was greatly decreased during selfing process, suggesting stability in advanced generations.
Changes of gene expression played an important role in the evolution of plant allopolyploids. Characters of the changes of gene expression between the allotetraploid C. × hytivus and its diploid parental species were analyzed by using cDNA-AFLP technique. The results indicated that most genes from parents could be stably expressed in the allotetraploid, while some genes expressed differentially. A total of 36 (3.37%) differentially expressed transcripts were detected and classified into three types: no expression of genes from parents, expression of genes from one parent, and novel expression of new genes. The majority was the expression of genes from one parent. The data also showed that the genes from female parent were easier to be changed. Those results indicated a rapid change in gene expression in early generations of C. × hytivus.
In Cucurbitaceae, amphidiploidy was reported from C. maxima × C. moschata (Pearson et al. 1951) and C. anguria × C. dipsaceus (Yadava et al. 1986). There were no successful efforts on the two most commercially important Cucumis spp.: cucumber and melon. C. × hytivus as a synthesized allopolyploid can be a useful model system to study polyploidization and may also serve as a genetic bridge in Cucumis and thus is a source for broadening the genetic base of C. sativus.
6.6.2.2 Allotriploid
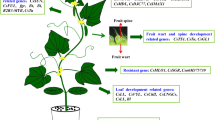
A primary allotriploid cucumber (2n = 3x = 26; HCC) was obtained through backcrossing the synthetic allotetraploid (HHCC, maternal parent) to cultivated cucumber (CC, paternal parent) (Chen et al. 2003). The allotriploid showed many novel characters, such as strong heterosis, sequential fruit set, tolerance to low temperature and low light, and high nutritional value of the fruit. In morphology, fruit weight, number of branches, ovary length, fruit length, and spine and mature fruit of progeny more closely resembled their maternal C. × hytivus parent (Fig. 6.12); developmental vigor (i.e., relative growth rate) and leaf color (i.e., dark green vs. reseda of C. × hytivus) were more similar to and characteristic of the paternal, C. sativus, parent. Progeny were more parthenocarpic than either parent bearing abundant seedless fruit (i.e., 84.8% developed fruit under greenhouse conditions). While their parents, C. × hytivus and C. sativus, were lower (26.7 and 45.2%, respectively) (Table 6.6).
Recently, 19 cucumber cultivars were used to make crosses with C. × hytivus to identify crosses producing allotriploid fruits with viable seeds. Only three crosses had fruits with viable seeds. However, this marked significant progress toward commercial use of this seedless novel cucumber. The putative allotriploid plants were confirmed by molecular and cytological analysis.
6.6.2.3 Monosomic Alien Addition Lines
Two monosomic alien addition lines (MAALs) (14 CC + 1 H, 2n = 15) were recovered among 252 regenerated plants, when the allotriploid was treated with colchicine to induce polyploidy (Chen et al. 2004a). Both the putative MAALs, plant numbers 87 and 517, grew more slowly and were easily differentiated morphologically from the allotriploids, C. × hytivus, and C. sativus. The leaf shape of these plants was palmate and hastate, respectively, in contrast to the pentagon shape observed in allotriploid and the wavy leaf edges observed in the allotetraploid C. × hytivus. Fruit length of plant numbers 87 and 517 was 23.6 ± 0.9 and 25.4 ± 1.5 cm, respectively, much longer than the allotriploids (12.1 ± 2.1 cm) (Fig. 6.13). Both plants showed white spines, characteristic of C. sativus, in contrast to black spines observed in progenies of the interspecific hybrids from the female parent, C. hystrix, indicating that the gene for black spines is not located on the alien chromosome in either plant number 87 or 517; both MAALs also showed a multibranching character reminiscent of C. hystrix.
Alien chromosome addition lines harboring one single chromosome from the wild species C. hystrix might be used as a bridge to transfer genes of interest originating in C. hystrix to individual chromosomes of C. sativus via recombination or translocation events.
6.6.2.4 Introgression Lines
C. sativus–hystrix introgression lines were obtained among progenies of interspecific hybrids after the allotetraploid was backcrossed to cultivated cucumber. These introgression lines were genetically stable and differed from each other as well as their two parents in many morphological traits (Fig. 6.14). Substantial genomic changes were detected in introgression lines, including C. hystrix-specific fragments, deletion of fragments originally observed in C. sativus, and novel bands not from both parents (Zhou et al. 2009).
Zhou et al. (2009) used SSR markers to detect the introgression from C. hystrix to C. sativus, and one locus at 210 bp was revealed and assigned to introgressive fragment of C. hystrix genome in introgression line 56. This line was characterized to have small-sized leaf, short fruit, and multibranching habit, which were closer to the wild parent (C. hystrix), and had fertility as high as that of cultivated cucumber. Moreover, this line was found to have a desirable response to downy mildew and Fusarium wilt. Several molecular markers linked to these introgressed traits are now being developed. The introgression lines could be used as vehicle for transferring desirable characters from C. hystrix and valuable for the improvement of cucumber cultivars.
6.6.2.5 New Pickling Cucumber F1 Hybrid
A new pickling cucumber line 7012A was developed through subsequent backcrossing of the hybrid to cucumber and selection from the selfed progenies (Chen et al. 2005). This line was used to cross with an elite American pickling cucumber line (7011A) from the University of Wisconsin to produce F1 (Fig. 6.15). The results indicated that the F1 has significant heterosis over its parents in yield and growth vigor. The plants set uniform fruits with good quality. Fruits could be set on both the main and lateral branches. It has highest late yield and its total yield was higher than all the other cultigens tested and this F1 was subsequently designated as Ningjia #1.
6.7 Genomic Resources
The development of genomic tools in Cucumis species has been very limited in spite of the importance of these species. In recent years, there has been an effort toward the development of genomic resources mainly in melon and cucumber, the cultivated species of Cucumis.
The Cucurbit genomics database of the International Cucurbit Genomics Initiative offers comprehensive information on express sequence tags (ESTs) and maps developed in the cucurbits. The version 2 of melon EST collection contains 34,451 ESTs and published genes, representing 16,128 unigenes. In cucumber, 4,331 ESTs are also available from leaf, fruit, and flower tissues representing around 3,000 unigenes. Several genetic maps are available for melon and cucumber. They have been constructed using different types of molecular markers and populations; however, these maps are still far from saturation (Garcia-Mas 2008).
Several bacterial artificial chromosome (BAC) libraries have been constructed in melon (Luo et al. 2001; van Leeuwen et al. 2003) and cucumber (Nam et al. 2005). A fosmid library of cucumber was also constructed (Havey et al. 2008). These libraries can be accessed from the website of the Clemson University Genomics Institute (http://www.cugi.org). Construction of a BAC library of C. hystrix is now undergoing in China.
The Cucumber Genome Initiative has reported the progress of the sequencing of the cucumber genome. The sequencing of the cucumber genome has been completed and about 30,000 genes were annotated. A genetic map was constructed, consisting of 1,200 diversity array technology (Dart) markers and 1,000 SSR (microsatellite) markers, distributed over seven linkage groups. By fluorescent in situ hybridization (FISH) mapping of 60 SSR-anchored fosimids, an integrated genetic and cytogenetic map was constructed and chromosomal rearrangements between subspecies of cucumber were discovered (Huang et al., pers comm). The cucumber genome sequence will provide a rich resource for investigating the pathways of gene and genome evolution.
6.8 Recommendations for Future Actions and Future Prospects
Wild species are an important reservoir of useful genes and offer great potential to incorporate such genes into commercial cultivars for resistance to major diseases, insects, and tolerance to various abiotic stresses. Moreover, many of the useful alien genes are different from those of the cultivated species and are thus useful in broadening the sources of resistance/tolerance to various stresses. However, despite the importance of wild species of Cucumis, very little information is available on the studies of wild species. Morever, use of exotic germplasm for the development of lines and populations with unique traits has been limited. The application of genetic markers to germplasm management or for marker-assisted selection has not been clearly defined in Cucumis. Therefore, we hope that future attention should be paid to research on the wild species.
As a first step, germplasm acquisition of all the Cucumis species available from every country should be initiated. Our objective should not be limited to the maintenance of germplasm. Instead, what is required is a well-organized research program on germplasm characterization, utilization, and enhancement.
An important long-term objective for Cucumis breeders is the introduction of genes from wild relatives. Some wild relatives, such as C. metuliferus E. Meyer ex Naudin (nematode resistance) and C. figarei Naudin (virus resistance), have long been attractive to scientists. However, the limits to utilization of wild relatives depend on the breeders’ ability to produce interspecific hybrids, but hybrids may be sterile. Future research should focus on producing fertile interspecific hybrids through biotechnological or other approaches.
The rapid advancement in the molecular genetics has now opened a new era of technologies and the application of MAS in crop breeding has become more and more important. There is thus a need to develop molecular markers tightly linked to the useful traits of wild species. Generation and utilization of novel genetic stocks such as introgression lines, substitution lines, and deletion lines will further facilitate the genetic analysis of complex agronomic traits and the introgression of desirable genes into commercial cultivars. Construction of the bacterial artificial chromosome (BAC) library of wild species of Cucumis, along with the tightly linked markers of desirable traits, may greatly promote the identification and isolation of useful genes for cucumber and melon improvement.
Wild relatives are useful sources of characters desired to improve cultivated Cucumis species. Although many of these wild species are resistant to pests and diseases or adapted to adverse environments, the utilization of wild relatives is limited due to the cross-incompatiability problems. However, the successful development of the synthetic allotetraploid species C. × hytivus Chen and Kirkbride via chromosome doubling of an interspecific hybrid between cucumber and C. hystrix Chakr. provides a step toward transferring desirable genes from the wild species C. hystrix. Future works emphasized on map-based cloning of the downy mildew and root-knot nematode resistance genes from C. hystrix will be of great value for the improvement of cucumber and melon.
References
Adelberg JW, Chen JF (1998) Genetic control of regeneration was altered during one-week ripening of immature melon cotyledons on liquid/membrane system. Presented at IAPTC World Congress on Cell Culture, Jerusalem, Israel
Adelberg JW, Rhodes BB, Skorupska HT, Bridges WC (1994) Explant origin affects the frequency of tetraploid plants from tissue cultures of melon. HortScience 29:689–692
Andrus CF, Fassuliotis G (1965) Crosses among Cucumis species. Veg Improv Newsl 7:3
Ayyangar KR (1967) Taxomomy of Cucurbitaceae. Bull Natl Inst Sci India 34:380–396
Beharav A, Cohen Y (1994) Effect of gamma radiation on vitality and fertilization ability of Cucumis melon and C. metuliferus pollen. Cucurbit Genet Coop Rep 17:94–96
Betra S (1953) Interspecific hybridization in the genus Cucumis. Sci Cult 18(9):445–446
Bhaduri PN, Bose PC (1947) Cyto-genetical investigations in some common cucurbits, with special reference to fragmentation of chromosomes as a physical basis of speciation. J Genet 48:237–256
Bowley SR, Taylor NL (1987) Introgressive hybridization. In: Christie BR (ed) CRC handbook of plant science in agriculture, vol 1. CRC, Boca Raton, FL, USA, pp 23–59
Chatterjee M, More TA (1991) Interspecific hybridization in Cucumis spp. Cucurbit Genet Coop 14:69–70
Chelliah S, Sambandam CN (1972) Inheritance of resistance to the fruit fly, Dacus cucurbitae C., in the interspecific cross between Cucumis callosus (Rottl.) Cogn. and Cucumis melo L. Auara 4(5):169–171
Chen LZ, Chen JF (2008) Changes of cytosine methylation induced by wide hybridization and allopolyploidy in Cucumis. Genome 51:789–799
Chen JF, Kirkbride J (2000) A new synthetic species Cucumis (Cucurbitaceae) from interspecific hybridization and chromosome doubling. Brittonia 52:315–319
Chen JF, Lewis S (2000) New source of nematode resistance was identified in Cucumis. Cucurbit Genet Coop Rep 23:32–35
Chen JF, Staub JE (1997) Attempts at colchicine doubling of an interspecific hybrid of Cucumis sativus L. × C. hystrix Chakr. Cucurbit Genet Coop Rep 20:24–26
Chen JF, Zhang S, Zhang X (1994) The xishuangbanna gourd, a traditional cultivated plant of the Hanai People, Xishuangbanna, Yunnan, China. Cucurbit Genet Coop Rep 17:18–20
Chen JF, Isskiki S, Tashiro Y, Miyazaki S (1995) Studies on a wild cucumber from China (Cucumis hystrix Chakr.) I. Genetic distances between C. hystrix and two cultivated Cucumis species (C. sativus L., and C. melo L.) based on isozyme analysis. J Jpn Soc Hortic Sci 64:264–265
Chen JF, Staub JE, Tashiro Y, Isshiki S, Miyazaki S (1997) Successful interspecific hybridization between Cucumis sativas L. and C. hystrix Chakr. Euphytica 96:413–419
Chen JF, Adelberg JW, Staub JE, Skorupska HT, Rhodes BB (1998) A new synthetic amphidiploid Cucumis from a C. sativus × C. hystrix F1 interspecific hybrid. In: McCreight J (ed) Cucurbitaceae’98 – Evaluation and enhancement of cucurbit germplasm. ASHS, Alexandria, VA, USA, pp 336–339
Chen JF, Luo XD, Staub JE, Jahn MM, Qian ChT, Zhuang FY, Ren G (2003) An allotriploid derived from a amphidiploid × diploid mating in Cucumis I: production, micropropagation and verification. Euphytica 131:235–241
Chen JF, Luo XD, Qian ChT, Molly MJ, Staub JE, Zhuang FY, Lou QF, Ren G (2004a) Cucumis monosomic alien addition lines: morphological, cytological, and genotypic analyses. Theor Appl Genet 108:1343–1348
Chen JF, Moriarty G, Jahn M (2004b) Some disease resistance tests in Cucumis hystrix and its progenies from interspecific hybridization with cucumber. In: Lebeda A, Paris HS (eds) Proceedings of 8th EUCARPIA meet on cucurbit genetics and breeding, Palacký University, Olomouc, Czech Republic, pp 189–196
Chen LZ, Chen JF, Staub J, Qian CT (2005) A new pickling cucumer F1 hybrid bred from interspecific hybridization. China Veg 3:4–6
Chen LZ, Lou QF, Zhuang Y, Chen JF, Zhang XQ, Wolukau JN (2007) Cytological diploidization and rapid genome changes of the newly synthesized allotetraploids Cucumis hytivus. Planta 225:603–614
Chung SF, Staub JE, Chen JF (2006) Molecular phylogeny of Cucumis species as revealed by consensus chloroplast SSR marker length and sequence variation. Genome 49:219–229
Custers JBM, Den Nijs APM (1986) Effects of aminoethoxyvinylglycine (AVG), environment, and genotype in overcoming hybridization barriers between Cucumis species. Euphytica 35:639–647
D’Amato F (1977) Cytogenetics of differentiation in tissue and cell cultures. In: Reinert J, Bajaj YPS (eds) Plant cell, tissue and organ culture. Springer, Berlin, Germany, pp 34–393
Dane F, Tsuchiya T (1976) Chromosome studies in the genus Cucumis. Euphytica 25:367–374
Dane F, Tsuchiya T (1979) Meiotic chromosome and pollen studies of polyploidy Cucumis species. Euphytica 28:563–567
Dane F, Denna DW, Tsuchiya T (1980) Evolutionary studies of wild species in the genus Cucumis. Z Pflanzenzucht 85:89–109
Danin-Poleg Y, Reis N, Tzuri G, Katzir N (2001) Development and characterization of microsatellite markers in Cucumis. Theor Appl Genet 102:61–72
Deakin JR, Bohn GW, Whitaker TW (1971) Interspecific hybridization in Cucumis. Econ Bot 25:195–211
den Nijs APM, Custers JBM (1990) Introducing resistance into cucumbers by interspecific hybridization. In: Bates DM, Robinson RW, Jeffery C (eds) Biology and utilization of the Cucurbitaceae. Cornell University Press, Ithaca, NY, USA, pp 382–396
den Nijs APM, Visser DL (1985) Relationships between African species of the genus Cucumis L. estimated by the production, vigour and fertility of F1 hybrids. Euphytica 34:279–290
Esquinas-Alcazar JT (1977) Alloenzyme variation and relationships in the genus Cucumis. PhD Dissertation, University of California, Davis, CA, USA
Ezura H, Amagai H, Yoshioka K, Oosawa K (1992) Highly frequent appearance of tetraploidy in regenerated plants, a universal phenomenon in tissue culture of melon (Cucumis melo L.). Plant Sci 85:209–213
Fassuliotis G (1967) Species of Cucumis resistant to the root-knot nematode, Meloidogyne incognita acrita. Plant Dis Rep 51:720–723
Fassuliotis G (1977) Self-fertilization of Cucumis metuliferus Naud. and its cross-compatilibity with C. melo L. J Am Hortic Soc 102(3):336–339
Fassuliotis G, Nelson BV (1988) Interspecific hybrids of Cucumis metuliferus × C. anguria obtained through embryo culture and somatic embryogenesis. Euphytica 37:53–60
Fellner M, Binarova P, Lebeda A (1996) Isolation and fusion of Cucumis sativas and Cucumis melo protoplasts. In: Gomez-Guillamon ML, Soria C, Cuartero J, Tores JA, Fernandez-Munoz R (eds) Cucurbits towards 2000, Proceedings of 6th EUCARPIA meeting on cucurbit genetics and breeding, Malaga, Spain, pp 202–209
Franken J, Custers JBM, Bino RJ (1988) Effects of temperature on pollen tube growth and fruit set in reciprocal crosses between Cucumis sativus and C. metuliferus. Plant Breeding 100:150–153
Garcia-Mas J (2008) Current genomic resources in melon and other cucubits. In: Pitrat M (ed) Proceedings of 9th EUCARPIA Cucurbitaceae Symposium, Avignon, France, pp 201–205
Garcia-Mas J, Monforte AJ, Arus P (2004) Phylogentic relationships among Cucumis species based on the ribosomal internal transcribed spacer sequence and microsatellite markers. Plant Syst Evol 248:191–203
Ghebretinsae AG, Thulin M, Barber JC (2007) Relationships of cucumbers and melons unraveled: molecular phylogenetics of Cucumis and related genera (Benincaseae, Cucurbitaceae). Am J Bot 94:1256–1266
Havey MJ, Meyer JDF, Deleu W, Garcia-Mas J (2008) Construction of a fosmid library of cucumber (Cucumis sativus) and comparative analyses of the eIF4E and eIF(iso)4E regions from cucumber and melon (Cucumis melo). Mol Genet Genom 279(5):473–480
Jeffrey C (1962) Notes on Cucurbitaceae including a proposed new classification for the family. Kew Bull 15:337–371
Jeffrey C (1967) On the classification of Cucurbitaceae. Kew Bull 20:417–426
Jeffrey C (1980) A review of the Cucurbitaceae. Bot J Linn Soc 81:233–247
Jeffrey C (1990) Systematics of Cucurbitaceae: an overview. In: Bates DM, Robinson RW, Jeffrey C (eds) Biology and utilization of the Cucurbitaceae. Cornell University Press, Ithaca, NY, USA, pp 3–9
Jeffrey C (2005) A new system of Cucurbitaceae. Bot Zhurn 90:332–335
Jobst J, King K, Hemleben V (1998) Molecular evolution of the internal transcribed spacers (ITS1 and ITS2) and phylogenetic relationships among species of the family Cucurbitaceae. Mol Phylogenet Evol 9:204–219
Kho YO, den Nijs APM, Franken J (1980) Interspecific hybridization in Cucumis L. II. The crossability of species, an investigation of in vivo pollen tube growth and seed set. Euphytica 29:661–672
Kirkbride JH (1993) Biosystematic monograph of the genus Cucumis (Cucurbitaceae). Parkway, Boone, NC, USA
Kishi Y, Fujishita N (1969) Studies on interspecific hybridization in the genus Cucumis. I. Pollen germination and pollen tube growth in selfings and incompatible crossings. J Jpn Soc Hortic Sci 38:329–334
Kishi Y, Fujishita N (1970) Studies on interspecific hybridization in the genus Cucumis II Pollen tube growth, fertilization and embryogenesis of post-fertilization stage in incompatible crossings. J Jpn Soc Hortic Sci 39:51–57
Kocyan A, Zhang LB, Schaefer H, Renner SS (2007) A multi-locus chloroplast phylogeny for the Cucurbitaceae and its implications for character evolution and classification. Mol Phylogenet Evol 44:553–577
Kristkova E, Lebeda A, Vinter V, Blahousek O (2003) Genetic resources of the genus Cucumis and their morphological description (English-Czech version). HortScience (Prague) 30(1):14–42
Laibach F (1925) Das Taubwerden von Bastardsamen und die kunstliche Aufzucht fruh absterbender Bastardembryonen. Z Bot 17:417–459
Leppick EE (1966) Searching gene centers of the genus Cucumis. Euphytica 15:323–328
Liu Q (2007) Distribution and molecular phylogenentics of Cucumis species. MA Dissertation, Nanjing Agricultural University, PR China
Lower RL, Edwards MD (1986) Cucumber breeding. In: Basset MJ (ed) Breeding vegetable crops. AVI, Westport, CN, USA, pp 173–207
Luo M, Wang YH, Frisch D, Joobeur T, Wing RA, Dean RA (2001) Melon bacterial chromosome (BAC) library construction using improved methods and identification of clones linked to the locus conferring resistance to melon Fusarium wilt (Fom-2). Genome 44:154–162
Ma SW, Wang JM, Qiu JT (2003) Present situation of collection and presevation of watermelon and melon germplasm in China. China Watermelon Muskmelon 5:17–19
McCreight JD, Nerson H, Grumet R (1993) Melon, Cucumis melo L. In: Kalloo G, Bergh BO (eds) Genetic improvement of vegetable crops. Pergamon, New York, USA, pp 267–294
Nam YW, Lee JR, Song KH, Lee MK, Robbins MD, Chung SM, Staub JE, Zhang HB (2005) Construction of two BAC libraries from cucumber (Cucumis sativus L.) and identification of clones linked to yield component quantitative trait loci. Theor Appl Genet 111(1):150–161
Naudin C (1859) Revue des Cucurbitaceae cultiveesau museum en 1859. Ann Sci Nat Ser 4 Bot 12:79–164
Norton JD, Granberry DM (1980) Characteristics of progeny from an interspecific cross of Cucumis melo with C. metuliferus. J Am Soc Hortic Sci 105:174–180
Niemirowicz-Szczytt K, Kubiki B (1979) Cross fertilization between cultivated species of genera Cucumis L. and Cucurbit L. Genetica Polonica 20:117–125
Osifo E, Webb JK, Henshaw GG (1989) Variation amongst callus derived potato plants Solanum bredidens. J Plant Physiol 134:1–4
Pangalo KJ (1950) Melons as the independent genus Melo Adans. Bot Z (Moscow and Leningrad) 35:571–580
Pearson OH, Hopp R, Bohn GW (1951) Notes on species crosses in Cucurbita. Proc Am Soc Hortic Sci 57:310
Perl-Treves R, Galun E (1985) The Cucumis plastome: physical map, intrageneric variation and phylogenetic relationships. Theor Appl Genet 71:417–429
Pitrat M, Chauvet M, Foury C (1999) Diversity, history, and production of cultivated cucurbits. Acta Hortic 492:21–28
Puchalski T, Robinson RW (1990) Electrophoretic analysis of isozymes in Cucurbita and Cucumis and its application for phylogenetic studies. In: Bates DM, Robinson RW, Jeffrey C (eds) Biology and utilization of the Cucurbitaceae. Cornell University Press, Ithaca, NY, USA, pp 60–76
Qian CT, Chen JF, Zhuang FY, Xu YB, Li SJ (2002) Several photosynthetic characters of the synthetic species Cucumis hytivus Chen and Kirkbride under weak light condition. Plant Physiol Commun 38:336–338
Raamsdonk LWD, den Nijs APM, Jongerius MC (1989) Meiotic analyses of Cucumis hybrids and an evolutionary evaluation of the genus Cucumis (Cucurbitaceae). Plant Syst Evol 163:133–146
Ramachandran C (1984) Nuclear DNA variation in Cucumis species. Cucurbit Genet Coop 7:97–98
Ramachandran C, Narayan RKJ (1985) Chromosomal DNA variation in Cucumis. Theor Appl Genet 69:497–502
Ramachandran C, Seshadri VS (1986) Cytological analysis of the genome of cucumber (Cucumis sativus L.) and muskmelon (Cucumis melo L.). Z Pflanzenzücht 96:25–38
Renner SS, Schaefer H (2008) Phylogenetics of Cucumis (Cucurbitaceae) as understood in 2008. Cucurbitaceae 2008. In: Pitrat M (ed) Proceedings of 9th EUCARPIA meeting on genetics and breeding of Cucurbitaceae. INRA, Avignon, France, 21–24 May 2008, pp 53–58
Renner SS, Schaefer H, Kocyan A (2007) Phylogenetics of Cucumis (Cucurbitaceae): Cucumber (C. sativus) belongs in an Asian/Australian clade far from melon (C. melo). BMC Evol Biol 7:58
Robinson RW, Decker-Walters DS (1997) Interspecific hybridization. In: Robinson R, Decker-Walters DS (eds) Cucurbits. CABI, Wallingford, Oxon, UK, pp 51–55
Rubatzky VE, Yamaguchi M (1997) World vegetables: principles, production, and nutritive values, 2nd edn. Chapman and Hall, New York, NY, USA
Schaefer H (2007) Cucumis (Cucurbitaceae) must include Cucumella, Dicoelospermum, Mukia, Myrmecosicyos, and Oreosyce: a recircumscription based on nuclear and plastid DNA data. Blumea 52:165–177
Sheng D, Li XX, Wang HP, Song JP (2006) Research progress and prospects on cucumber germplasm. China Veg Suppl 150:77–81
Singh BP (1991) Interspecific hybridization in between new and old-world species of Luffa and phylogenetic implication. Cytologia 56:35–365
Singh AK, Yadava KS (1984) An analysis of interspecific hybrids and phylogenetic implications in Cucumis (Cucurbitaceae). Plant Syst Evol 147:237–252
Soria C, Gomez-Guillamon ML, Esteva J, Nuez F (1990) Ten interspecific crosses in the genus Cucumis: a preparatory study to seek crosses resistant to melon yellowing diseases. Cucurbit Genet Coop Rpt 13:31–33
Staub JE, Serquen FC, Horejsi T, Chen JF (1999) Genetic diversity in cucumber (Cucumis sativus L.): IV. An evaluation of Chinese germplasm. Genet Resour Crop Evol 46:297–310
Tabei Y, Nishio T, Kanno T (1992) Shoo regeneration from cotyledonary protoplasts on melon (Cucumis melo L.cv. Charentais). J Jpn Soc Hortic Sci 61:317–322
Tang FA, Punja ZK (1989) Isolation and culture of protoplasts of Cucumis sativas and Cucumis metuliferus and methods for their fusion. Cucurbit Genet Coop Rep 12:29–32
Trivedi RN, Roy RP (1970) Cytological studies in Cucumis and Citrullus. Cytologia 35:561–569
Valvilov N (1925) Inter-genetic hybrids of melons, watermelons and squashes. Plant Breed 14:3–35
van Leeuwen H, Monfort A, Zhang HB, Puigdomenech P (2003) Identification and characterisation of a melon genomic region containing a resistance gene cluster from a constructed BAC library. Microcolinearity between Cucumis melo and Arabidopsis thaliana. Plant Mol Biol 51:703–718
Varekamp HQ, Visser DL, den Nijs APM (1982) Rectification of the names of certain accessions of the IVT- Cucumis collection. Cucurbit Genet Coop 5:59–60
Visser DL, den Nijs APM (1983) Variation for interspecific crossability of Cucumis anguria L. and C. zeyer Sond. Cucurbit Genet Coop 6:100–101
Weeden NF, Robinson RW (1986) Allozyme segregation ratios in the interspecific cross Cucurbita maxima × C. ecuadorensis suggest that hybrid breakdown is not causes by minor alterations in chromosome structure. Genetics 114:593–609
Whitaker TW, Davis GN (1962) Cucurbits: botany, cultivation and utilization. Interscience, New York, USA
Yadava KS, Singh AK, Roy RP, Jha UC (1986) Cytogenetics in Cucumis L. VI Synthetic amphidiploids. Nucleus 29:58–62
Zhang YB, Chen JF, YI HP, Feng JX, Wu MZ (2005) Staining and slide-preparing technique of mitotic chromosomes and its use in karyotype determination of Cucumis melo L. Acta Bot Boreal-Occident Sin 25(9):1735–1739
Zhou XH, Qian CT, Lou QF, Chen JF (2009) Molecular analysis of introgression lines from Cucumis hystrix Chakr. to C. sativus L. Sci Hortic 119:232–235
Zhuang FY, Chen JF, Qian ChT, Li ShJ, Ren G, Wang ZhJ (2002) Responses of seedlings of Cucumis × hytivus and progenies to low temperature. J Nanjing Agric Univ 25:27–30
Zhuang FY, Chen JF, Staub JE, Qian CT (2006) Taxonomic relationships of a rare Cucumis species (C. hystrix Chakr.) and its interspecific hybrid with cucumber. HortScience 41(3):571–574
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Chen, JF., Zhou, XH. (2011). Cucumis. In: Kole, C. (eds) Wild Crop Relatives: Genomic and Breeding Resources. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-20450-0_6
Download citation
DOI: https://doi.org/10.1007/978-3-642-20450-0_6
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-20449-4
Online ISBN: 978-3-642-20450-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)