Abstract
Autophagy is a cellular process that creates double-membraned vesicles, engulfs and degrades cytoplasmic material, and generates and recycles nutrients. A recognized participant in the innate immune response to microbial infection, a functional autophagic response can help to control the replication of many viruses. However, for several viruses, there is functional and mechanistic evidence that components of the autophagy pathway act as host factors in viral replicative cycles, viral dissemination, or both. Investigating the mechanisms by which viruses subvert or imitate autophagy, as well as the mechanisms by which they inhibit autophagy, will reveal cell biological tools and processes that will be useful for understanding the many functional ramifications of the double-membraned vesicle formation and cytosolic entrapment unique to the autophagy pathway.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Viruses, especially RNA viruses, have the capacity to evolve quickly in response to changes in host cells. As a consequence, it is difficult to think of any antiviral response for which there is not an example of a viral mechanism to avoid the response or, even more beneficially for the virus, to subvert the response to provide some advantage for the virus. A good example of the wide range and mechanisms of viral reactions to antiviral mechanisms is that of viral strategies to contend with cell death by apoptosis, an effective defense against many catastrophes, including infection by microorganisms. Most viruses, especially those whose success has not elevated them to fame as pathogens, are undoubtedly cleared by cellular apoptosis before they have a chance to spread. Almost every virus that is known to thrive in animals or in tissue culture cells has, of necessity, developed a mechanism to inhibit cellular apoptosis. Indeed, one of the first antiapoptotic proteins identified, p35, was discovered by Lois Miller and her laboratory through their studies of baculoviruses, a class of insect-infecting DNA viruses (Clem et al. 1991). Inhibitors of nearly every step in the extrinsic and intrinsic apoptotic pathways have been identified from viruses, and the actions of these range from derailing signal transduction to blocking mitochondrial channel formation to inhibiting the effector caspases themselves (reviewed in Best 2008; Galluzzi et al. 2008). We could conclude that the viral repertoire is exceedingly thorough, but this would be a bit unfair, because often (as was the case for p35) it was the identification of the viral inhibitor that defined the step in the first place.
It is perhaps not surprising that, given the sequence and cell biological space that viruses can explore, some viruses would have evolved to exploit the apoptotic pathway for their own benefit, rather than just inhibiting it. How could the induction of apoptosis facilitate viral replication? The eventual lysis of infected cells is the mechanism of spread of many viruses; it is likely that the ultimate failure of antiapoptotic mechanisms in many viral infections results in a delayed apoptosis that facilitates cell spread. Another, more surprising mechanism has also been demonstrated: for mink cell focus-forming virus (Best et al. 2003), human astrovirus (Mendez et al. 2004) and feline calicivirus (Al-Molawi et al. 2003), caspases induced during apoptosis are required for viral capsid processing (reviewed in Best 2008).
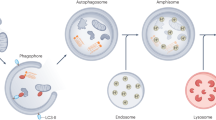
The cellular autophagy pathway, as discussed throughout this volume, was originally discovered as a response to starvation, and provides a route for cytoplasmic constituents to be targeted to the lysosomal machinery for degradation (reviewed in Mizushima et al. 2002, 2008). Cytosolic components become enwrapped by a double-membraned structure, termed the “immature autophagosome.” The resulting double-membraned structures contain the marker LC3, which becomes covalently linked to phosphatidylethanolamine and thereby membrane-associated upon the induction of autophagy. LC3 was originally identified as MAP-LC3, a microtubule-associated protein (Kuznetsov and Gelfand 1987), and may be part of the mechanism by which autophagosomes interact with the microtubule network, on which they traffic by an anterograde route toward the microtubule organizing center (Fass et al. 2006; Jahreiss et al. 2008; Kimura et al. 2008; Koechl et al. 2006). Immature autophagosomes mature by fusion with vesicles from the endosomal pathway, then with lysosomes, in experimentally separate steps. At the “mature autophagosome” stage, the inner membranes become degraded, accomplishing the topological transformation of cytosol into lumen, something that happens rarely within cells. Eventually, condensed “autolysosomes” release amino acids, lipids and nucleotides for further cellular use. A possibility considered here is that autophagosomes and autolysosomes have the capacity to release cytosolic contents to the extracellular milieu by a process similar to the known pathways of lysosomal exocytosis (reviewed in Stinchcombe et al. 2004).
From first principles, how might the autophagy pathway be related to the replicative cycles of viruses? A clear role in the degradation of viruses and viral factories can be readily envisaged, given the known ability of the autophagy pathway to degrade cellular components. Indeed—as discussed in the chapters by Tal and Iwasaki, Orvedahl and Levine, and Seay et al. in this volume—autophagy has been shown to participate in the control of viral infections as diverse as human herpes and plant RNA viruses, and has been justifiably hailed as a branch of the innate immune response. In such cases, reducing the function of the autophagy pathway removes this control and allows increased proliferation of the virus
2 Viruses for Which the Autophagy Pathway is Advantageous
In contrast, the opposite situation, in which reduction in the function of some component of the autophagy pathway reduces viral replication, has been demonstrated for a handful of viruses. As shown in Fig. 1, reduction of the intracellular concentration of either LC3 or Atg12 using small interfering RNAs was shown to reduce both the intracellular and extracellular yields of poliovirus, a small, nonenveloped positive-strand RNA virus, in single-cycle infections (Jackson et al. 2005). For coxsackievirus B3, a closely related picornavirus, reduction in the intracellular concentration of Atg7 substantially reduced the amount of viral capsid protein synthesized in a single-cycle infection (Fig. 1c) (Wong et al. 2008). For dengue virus, an enveloped positive-strand RNA virus, experiments to compare single-cycle infections of murine embryonic fibroblasts derived from autophagy-proficient and autophagy-deficient mice showed clear reductions in the yield of extracellular virus in the absence of a functional autophagy pathway (Lee et al. 2008). Recently, a genomic screen was performed to identify those protein-coding human genes whose targeting by specific small interfering RNAs reduced the yield of human immunodeficiency virus. This screen identified 36 known host factors, including coreceptors, and more than 100 others, which included autophagy genes ATG7, GABARAPL2 (a homolog of LC3), ATG12, and ATG16 (Brass et al. 2008). As is the case for the positive-strand RNA viruses mentioned, the mechanism of any advantage conferred to the replicative cycle, and whether the effects are direct or indirect, is not yet known.
Functional evidence that autophagy pathway function can have a positive effect on viral replicative cycles in tissue culture. The intracellular and extracellular yields of poliovirus were quantified in HeLa cells after treatment with (a) 12.5 pm each of eight different RNA duplexes targeted to the LC3A and LC3B mRNAs or with 100 pm of an RNA duplex targeted to firefly luciferase, or (b) with 25 pm each of four RNA duplexes targeted to ATG12 mRNA or with 100 pm of an RNA duplex targeted to firefly luciferase, for 24 h at 37°C. For both panels, insets show the relative abundance of the targeted protein upon treatment. Taken from Jackson et al. (2005) with permission. (c) Effect of reducing Atg7 protein abundance on intracellular accumulation of VP1 during infection with coxsackievirus B3 (Wong et al. 2008). (d) Differences in the yield of dengue virus after infection of murine embryonic fibroblasts from Atg5 +/+ and Atg5 −/− mice at two different times postinfection. Taken from Lee et al. (2008) with permission
What mechanistic inferences can be drawn from data such as those in Fig. 1? First, the effects are not absolute: in no case is the yield of intracellular virus or virus product reduced more than threefold, and, for extracellular virus, the largest observed reduction was 20-fold (Fig. 1a). For the RNAi experiments, it is likely that the reductions in autophagy protein concentration were not complete, so residual autophagy function remained. However, for genetic knockouts such as that shown for dengue virus in Atg5 −/− cells in Fig. 1d, the conclusion is inescapable that the autophagy pathway or its components are helpful to the formation or release of extracellular dengue virus, but not essential.
3 What Can Autophagy Do for a Virus?
A few possibilities for positive roles of the autophagy pathway or autophagosome formation in viral infections have been suggested by data in the literature thus far. In 1965, electron microscopic observations of poliovirus-infected cells revealed the presence of large numbers of double-membraned vesicles (Dales et al. 1965), which were recognized by these authors as “autophagolysomes.” Then, led by the work of Kurt Bienz and Denise Eggers (Bienz et al. 1983), it became increasingly appreciated that all positive-strand RNA viruses replicate their RNA genomes on the cytosolic faces of cytoplasmic membranes. In the case of poliovirus, these cytosolic-facing surfaces were shown to be those of the hundreds of membranous vesicles that accumulated in the cytoplasm of infected cells. Might these membranous vesicles be related to the double-membraned, autophagosome-like structures originally observed by the laboratory of George Palade? Over the last several years, my laboratory has shown that such double-membraned structures are readily visualized in both poliovirus-infected cells (Fig. 2a, b) and human rhinovirus-infected cells by electron microscopy following fixation by high-pressure freezing and freeze substitution (Schlegel et al. 1996; Dodd et al. 2001; Suhy et al. 2000; Jackson et al. 2005). Immunoelectron microscopy has shown that virally encoded proteins that are known to be found in the RNA replication complex localize to the cytoplasmic surface of these double-membraned vesicles, as do LC3 and LAMP-1 (Schlegel et al. 1996; Suhy et al. 2000; Jackson et al. 2005). Mechanistic insight into the assembly of double-membraned vesicles in cells may be provided by the finding that two viral proteins, termed 2BC and 3A, are sufficient to induce double-membraned vesicles that contain LC3 and LAMP-1 (Jackson et al. 2005; Suhy et al. 2000), with 2BC alone being sufficient to cause the accumulation of lipidated LC3 (Taylor and Kirkegaard, 2007). The observation that RNA replication complexes can assemble on the cytoplasmic surfaces of double-membraned vesicles that resemble autophagosomes in many respects has led to the hypothesis that the autophagy pathway is subverted by poliovirus, and possibly other positive-strand viruses, to generate such membranous surfaces. As can be seen in Fig. 2, double-membraned vesicles have recently been observed in cells infected with coxsackievirus B3 and Dengue virus as well, both of which show some functional dependence on the autophagy pathway (Fig. 1) (Lee et al. 2008; Wong et al. 2008).
Ultrastructural changes in cells infected with viruses that are suspected of subverting the cellular autophagy pathway. Transmission electron microscopy of uninfected (a) and poliovirus-infected (b) Hela cells preserved by high-pressure freezing and freeze substitution in the presence of osmium tetroxide. Size bar: 1000 nm. Taken from Schlegel et al. (1996) with permission. Uninfected (c) and coxsackievirus 3-infected (d) HEK293 cells visualized following glutaraldehyde fixation and osmium tetroxide staining. Size bar in inset: 200 nm. Taken from Wong et al. (2008) with permission. Dengue virus-infected murine embryonic fibrobasts from either Atg5 +/+ (e) or Atg5 −/− mice (f) after glutaraldehyde fixation and osmium tetroxide staining. Taken from Lee et al. (2008) with permission
But why would viruses use double-membraned vesicles and autophagosome-related proteins? While the hypothesis that the membranes and membrane constituents of the autophagy pathway contribute to the platforms on which viral RNA replication complexes can be built is attractive, it is clear that even poliovirus, coxsackievirus B3 and dengue viruses must be able to utilize other membrane surfaces as well, given that the reduced availability of autophagic proteins resulted in only small reductions in virus production (Fig. 1). For example, in Atg5 −/− cells infected with dengue virus (Fig. 1f), no double-membraned vesicles were observed (Lee et al. 2008), yet the virus must have assembled its RNA replication complex somewhere. And what is to be made of related viruses, such as human rhinovirus and murine hepatitis virus, which induce the accumulation of double-membraned vesicles in infected cells, but for which single replicative cycles show no apparent dependence on the presence or absence of a functional autophagic pathway substitution (Brabec-Zaruba et al. 2007; Zhao et al. 2007)? It is interesting, in this context, to consider the case of flock house virus, a positive-strand RNA virus that normally infects insects, and whose RNA replicative cycle can be observed in a variety of cell types including S. cerevisiae In all cell types examined, the normal site of assembly of the RNA replication complexes of flock house virus is the outer mitochondria membrane. However, clever genetic engineering experiments allowed the assembly of these RNA replication complexes on the ER of yeast cells; at this retargeted location, the rate of viral RNA replication increased sixfold (Miller et al. 2003). Thus, either nature has made a mistake by targeting flock house virus RNA replication to the outer mitochondrial membrane, or it is targeted there for some reason other than that which can be observed as an increased rate of a mechanistic process in a single RNA replicative cycle.
One such potentially alternative function for autophagosome-like membranes in virally infected cells was suggested to us by the data in Fig. 1. At 6 h after infection with poliovirus, no apparent cell lysis is usually observed. Yet, 1,000 plaque-forming units of extracellular virus accumulated in control infected cells, and 20-fold less when the abundance of LC3 was reduced. This effect was much more pronounced than the threefold reduction in intracellular virus titer, and it led us to hypothesize that, since it is known that the double-membraned vesicles that form late in poliovirus infection can contain cytoplasmic contents that include infectious virions and other cytoplasmic components, these membranes can provide a route of nonlytic viral escape from infected cells. We term this proposed pathway of cell exit “autophagic exit without lysis” or AWOL (Taylor et al., 2009), and suspect that it may be a component of viral pathogenesis.
There is ample precedent in the literature for the nonlytic spread of nonenveloped cytoplasmic viruses. Elegant experiments with Theiler’s virus, a murine virus related to poliovirus, have argued that the transfer of Theiler’s virus from optic nerves to the oligodendrocytes that myelinate them is an obligate step in the establishment of persistent infections, and can occur without neuronal lysis (Roussarie et al. 2007). In another potential example from picornaviruses, hepatitis A virus is known to spread through liver tissue in the absence of any apparent cell lysis; no mechanism for this observation has been demonstrated (reviewed in Hollinger and Ticehurst 1990).
Even at early times postinfection, it is difficult to distinguish whether a few cells lyse or many cells leak when so few submicroscopic viruses are present in the extracellular milieu (Fig. 1). Similarly, although the sporadic, apparently nonlytic spread of toxic aggregated proteins between cells has been observed, it is difficult to exclude stochastic events like occasional cell lysis (Ren et al. 2009). However, the demonstrably nonlytic release of Cryptococcus neoformans, a pathogenic fungus, from infected macrophages, which has been shown to correlate with association with complex membranes that contain LAMP-1 (Alvarez and Casadevall 2006), may be a spectacular example of AWOL by an intracellular pathogen.
The integrity of contents delivered by a pathway such as AWOL should depend critically on the extent of maturation of the autophagosome-like compartment. As illustrated in Fig. 3, the fusion of an immature autophagosome with the plasma membrane would secrete a lipid-rich, single-membraned vesicle, similar to an exosome, into the extracellular milieu. On the other hand, a mature autolysosome would, upon fusion with the plasma membrane, directly release degraded material and any cytosolic constituents that could withstand the environment of the lysosome. Clearly, intermediates between these stages are likely to exist as well.
Model for AWOL: autophagosome-mediated exit without lysis. Cytosolic constitutents can be surrounded by double-membraned vesicles derived from or resembling membranes of the autophagy pathway. Partial or complete degradation of the inner membrane can effectively mix cytosol and luminal components. Then, a process akin to the known lysosomal secretion pathway could facilitate the extracellular release of the autophagosomal contents. Modified from Jackson et al. (2005) with permission
Recent experiments have shown directly that autophagosomal-like membranes within virally infected cells can be less degradative than bona fide autophagosomes. As shown in Fig. 4a, decreasing the concentration of LAMP-2 was shown to increase the intracellular accumulation of coxsackievirus B3 VP1 protein, presumably by changing the functionality of the virus-induced membranes. During hepatitis C infection, autophagy is known to be induced, although any benefit of this induction for viral replication is not yet known (Ait-Goughoulte et al. 2008; Sir et al. 2008). In any case, as shown in Fig. 4b, The GFP-LC3-containing structures induced during HCV infection do not colocalize with a marker of acidified lysosomes, unlike the GFP-LC3 in starved cells. It remains to be explored whether these viruses affect the maturation of autophagosomes to inhibit an antiviral response in the host cell, or for their own direct benefit.
Potential blockage to maturation of autophagosome-like compartments in virally infected cells. (a) Immunoblot analysis of extracts from mock-infected and coxsackievirus B3-infected HeLa cells in the absence and presence of reduced intracellular amounts of LAMP-1. Taken from Wong et al. (2008) with permission. (b) Fluorescence microscopy to determine colocalization of Lysotracker and GFP-LC3 in Huh-7 cells that were either starved to induce autophagy or infected with the JFH-1 strain of hepatitis C virus. Taken from Sir et al. with permission of John Wiley & Sons, Inc
4 Conclusion
In summary, for most viruses and other intracellular pathogens, cellular defenses such as apoptosis and autophagy are induced upon infection and participate actively in clearance. Any viruses famous enough to be studied, however, are likely to encode one or multiple inhibitors of both these processes. Several viruses have been shown to induce autophagosome-like structures and to benefit from their formation. These benefits, so far, are likely to be the proliferation of membranes for RNA replication complexes and the generation of unique topologies that allow the escape of cytoplasmic constituents without cell lysis, which may be critical for viral spread within infected tissues.
References
Ait-Goughoulte M, Kanda T, Meyer K, Ryerse JS, Ray RB, Ray R (2008) Hepatitis C virus genotype 1a growth and induction of autophagy. J Virol 82:2241–2249
Al-Molawi N, Beardmore VA, Carter MJ, Kass GE, Roberts LO (2003) Caspase-mediated cleavage of the feline calicivirus capsid protein. J Gen Virol 84:1237–1244
Alvarez M, Casadevall A (2006) Phagosome extrusion and host-cell survival after Cryptococcus neoformans phagocytosis by macrophages. Curr Biol 16:2161–2165
Best SM (2008) Viral subversion of apoptotic enzymes: escape from death row. Annu Rev Microbiol 62:171–192
Best SM, Shelton JF, Pompey JM, Wolfinbarger JB, Bloom ME (2003) Caspase cleavage of the nonstructural protein NS1 mediates replication of Aleutian mink disease parvovirus. J Virol 77:5305–5312
Bienz K, Egger D, Rasser Y, Bossart W (1983) Intracellular distribution of poliovirus proteins and the induction of virus-specific cytoplasmic structures. Virology 131:39–48
Brabec-Zaruba M, Berka U, Blaas D, Fuchs R (2007) Induction of autophagy does not affect human rhinovirus type 2 production. J Virol 81:10815–10817
Brass AL, Dykxhoorn DM, Benita Y, Yan N, Engelman A, Xavier RJ, Lieberman J, Elledge SJ (2008) Identification of host proteins requried for HIV infection through a functional genomic screen. Science 319:921–926
Clem RJ, Fechheimer M, Miller LK (1991) Prevention of apoptosis by a baculovirus gene during infection of insect cells. Science 254:1388–1390
Dales S, Eggers HJ, Tamm I, Palade GE. (1965) Electron microscopic study of the formation of poliovirus. Virology 26:379–389
Dodd DA, Giddings TH, Jr., Kirkegaard K. (2001) Poliovirus 3A protein limits interleukin-6 (IL-6), IL-8, and beta interferon secretion during viral infection. J Virol 75:8158–8165
Fass E, Shvets E, Degani I, Hirschberg K, Elazar Z (2006) Microtubules support production of starvation-induced autophagosomes but not their targeting and fusion with lysosomes. J Biol Chem 281:36303–36316
Galluzzi L, Brenner C, Morselli E, Touat Z, Kroemer G (2008) Viral control of mitochondrial apoptosis. PLoS Pathog 4:e1000018
Hollinger FB, Ticehurst J (1990) Hepatitis A virus. In: Fields BN et al. (eds) Fields virology, 2nd edn. Raven, New York, pp 631–667
Jackson WT, Giddings TH Jr, Taylor MP, Mulinyawe S, Rabinovitch M, Kopito RR, Kirkegaard K (2005) Subversion of cellular autophagosomal machinery by RNA viruses. PLoS Biol 3:e156
Jahreiss L, Menzies FM, Rubinsztein DC (2008) The itinerary of autophagosomes: from peripheral formation to kiss-and-run fusion with lysosomes. Traffic 9:574–587
Kimura S, Noda T, Yoshimori T (2008) Dynein-dependent movement of autophagosomes mediates efficient encounters with lysosomes. Cell Struct Funct 33:109–122
Kirkegaard K, Jackson WT (2005) Topology of double-membraned vesicles and the opportunity for non-lytic release of cytoplasm. Autophagy 1:182–184
Koechl R, Hu XW, Chan EYW, Tooze S (2006) Microtubules faciliate autophagosome formation and fusion of autophagosomes with endosomes. Traffic 7:129–145
Kuznetsov SA, Gelfand VI (1987) 18 kDa microtubule-associated protein: identification as a new light chain (LC-3) of microtubule-associated protein 1 (MAP-1). FEBS Lett 212:145–148
Lee YR, Lei HY, Liu MT, Wang JR, Chen SH, Jiang-Shieh YF, Lin YS, Yeh TM, Liu CC, Liu HS (2008) Autophagic machinery activated by dengue virus enhances virus replication. Virology 374:240–248
Mendez E, Salas-Ocampo E, Arias CF (2004) Caspases mediate processing of the capsid precursor and cell release of human astroviruses. J Virol 78:8601–8608
Miller DJ, Schwartz MD, Dye BT, Ahlquist P (2003) Engineered retargeting of viral RNA replication complexes to an alternative intracellular membrane. J Virol 77:12193–12202
Mizushima N, Ohsumi Y, Yoshimori T (2002) Autophagosome formation in mammalian cells. Cell Struct Funct 27:421–429
Mizushima N, Levine B, Cuervo AM, Klionsky DJ (2008) Autophagy fights disease through cellular self-digestion. Nature 451:1069–1075
Ren PH, Lauckner JE, Kachirskaia I, Heusor JE, Melki R, Kopita RR (2009) Cytoplasmic penetration and persistent infection of mammalian cells by polyglutamine. Nat Cell Biol 11(2):219–225 (epub 2009)
Roussarie JP, Ruffie C, Edgar JM, Griffiths I, Brahic M (2007) Axon myelin transfer of a non-enveloped virus. PLoS ONE 2:e1331
Schlegel A, Giddings TH, Jr., Ladinsky MS, Kirkegaard K. (1996) Cellular origin and ultrastructure of membranes induced during poliovirus infection. J Virol 70:6576–6588
Sir D, Chen WL, Choi J, Wakita T, Yen TS, Ou JH. (2008) Induction of incomplete autophagic response by hepatitis C virus via the unfolded protein response. Hepatology 48:1054–1061
Stinchcombe J, Bossi G, Griffiths GM. (2004) Linking albinism and immunity: the secrets of secretory lysosomes. Science 305:55–59
Suhy DA, Giddings TH, Jr., Kirkegaard K (2000) Remodeling the endoplasmic reticulum by poliovirus infection and by individual viral proteins: an autophagy-like origin for virus-induced vesicles. J Virol 74:8953–8965
Taylor MP, Kirkegaard K (2007) Modification of cellular autophagy protein LC3 by poliovirus. J Virol 81:12543–12553
Taylor MP, Burgon TB, Kirkegaard K, Jackson WT (2009) Role of microtubules in extracellular release of poliovirus. J Viral 83(13):6599–6609 (epub 2009)
Wong J, Zhang J, Si X, Gao G, Mao I, McManus BM, Luo H (2008) Autophagosome supports coxsackievirus B3 replication in host cells. J Virol 82:9143–9153
Zhao Z, Thackray L, Miller B, Lynn T, Becker M, Ward E, Mizushima N, Denison M, Virgin HT (2007) Coronavirus replication does not require the autophagy gene ATG5. Autophagy 3:581–585
Acknowledgments
I would especially like to thank Thomas H. Giddings, Jr., whose electron micrographs allowed us to rediscover the double-membraned character of poliovirus-induced vesicles, and Andrew Staehelin, who first taught me about the autophagy pathway. I am grateful to the members of the laboratory who have worked on this topic over several years: Andreas Schlegel, David Suhy, William T. Jackson and Matthew Taylor. I thank Michel Brahic for stimulating discussions of non-lytic viral release and Arturo Casadevall for communicating results prior to publication. This work has been funded by the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Kirkegaard, K. (2009). Subversion of the Cellular Autophagy Pathway by Viruses. In: Levine, B., Yoshimori, T., Deretic, V. (eds) Autophagy in Infection and Immunity. Current Topics in Microbiology and Immunology, vol 335. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-00302-8_16
Download citation
DOI: https://doi.org/10.1007/978-3-642-00302-8_16
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-00301-1
Online ISBN: 978-3-642-00302-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)