Abstract
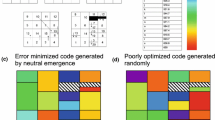
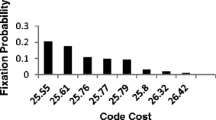
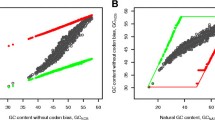
One of hypotheses explaining the origin of the genetic code assumes that its evolution has minimised the deleterious effects of mutations in coded proteins. To estimate the level of such optimization, we calculated optimal codes for genes located on differently replicating DNA strands separately assuming the rate of amino acid substitutions in proteins as a measure of code’s susceptibility to errors. The optimal code for genes located on one DNA strand was simultaneously worse than the universal code for the genes located on the other strand. Furthermore, we generated 20 million random codes of which only 23 were better than the universal one for genes located on both strands simultaneously while about two orders of magnitude more codes were better for each of the two strands separately. The result indicates that the existing universal code, the mutational pressure, the codon and amino acid compositions are highly optimised for the both differently replicating DNA strands.
Chapter PDF
Similar content being viewed by others
Keywords
References
Di Giulio, M.: On the origin of the genetic code. J. Theor. Biol. 187, 573–581 (1997)
Knight, R.D., Freeland, S.J., Landweber, L.F.: Selection, history and chemistry: the three faces of the genetic code. Trends Biochem. Sci. 24, 241–247 (1999)
Knight, R.D.: The origin and evolution of the genetic code: statistical and experimental investigations. Ph.D. Thesis, Department of Ecology and Evolutionary Biology, Princeton University (2001)
Dunnill, P.: Triplet nucleotide-amino-acid pairing a stereochemical basis for the division between protein and non-protein amino-acids. Nature 210, 1265–1267 (1966)
Pelc, S.R., Welton, M.G.: Stereochemical relationship between coding triplets and amino-acids. Nature 209, 868–872 (1966)
Woese, C.R., Dugre, D.H., Saxinger, W.C., Dugre, S.A.: The molecular basis for the genetic code. Proc. Natl. Acad. Sci. USA 55, 966–974 (1966)
Yarus, M., Caporaso, J.G., Knight, R.: Origins of the genetic code: the escaped triplet theory. Annu. Rev. Biochem. 74, 179–198 (2005)
Wong, J.T.-F.: A Co-Evolution Theory of the Genetic Code. Proc. Natl. Acad. Sci. USA 72, 1909–1912 (1975)
Taylor, F.J.R., Coates, D.: The code within the codons. BioSystems 22, 177–187 (1989)
Di Giulio, M.: On the relationships between the genetic code coevolution hypothesis and the physicochemical hypothesis. Z. Naturforsch. [C] 46, 305–312 (1991)
Amirnovin, R.: An analysis of the metabolic theory of the origin of the genetic code. J. Mol. Evol. 44, 473–476 (1997)
Di Giulio, M., Medugno, M.: The Historical Factor: The Biosynthetic Relationships Between Amino Acids and Their Physicochemical Properties in the Origin of the Genetic Code. J. Mol. Evol. 46, 615–621 (1998)
Alff-Steinberger, C.: The genetic code and error transmission. Proc. Natl. Acad. Sci. USA 64, 584–591 (1969)
Haig, D., Hurst, L.D.: A quantitative measure of error minimization in the genetic code. J. Mol. Evol (Erratum in J. Mol. Evol. 49, 708 (1999)) 33, 412–417 (1991)
Ardell, D.H.: On error minimization in a sequential origin of the standard genetic code. J. Mol. Evol. 47, 1–13 (1998)
Freeland, S.J., Hurst, L.D.: The genetic code is one in a million. J. Mol. Evol. 47, 238–248 (1998)
Freeland, S.J., Knight, R.D., Landweber, L.F., Hurst, L.D.: Early fixation of an optimal genetic code. Mol. Biol. Evol. 17, 511–518 (2000)
Sella, G., Ardell, D.H.: The impact of message mutation on the fitness of a genetic code. J. Mol. Evol. 54, 638–651 (2002)
Freeland, S.J., Wu, T., Keulmann, N.: The case for an error minimizing standard genetic code. Orig. Life Evol. Biosph. 33, 457–477 (2003)
Crick, F.H.: The origin of the genetic code. J. Mol. Evol. 38, 367–379 (1968)
Nowicka, A., Mackiewicz, P., Dudkiewicz, M., Mackiewicz, D., Kowalczuk, M., Cebrat, S., Dudek, M.R.: Correlation between mutation pressure, selection pressure, and occurrence of amino acids. In: Sloot, P.M.A., Abramson, D., Bogdanov, A.V., Gorbachev, Y.E., Dongarra, J., Zomaya, A.Y. (eds.) ICCS 2003. LNCS, vol. 2658, pp. 650–657. Springer, Heidelberg (2003)
Nowicka, A., Mackiewicz, P., Dudkiewicz, M., Mackiewicz, D., Kowalczuk, M., Banaszak, J., Cebrat, S., Dudek, M.R.: Representation of mutation pressure and selection pressure by PAM matrices. Applied Bioinformatics 3, 31–39 (2004)
Dudkiewicz, M., Mackiewicz, P., Nowicka, A., Kowalczuk, M., Mackiewicz, D., Polak, N., Smolarczyk, K., Banaszak, J., Dudek, M.R., Cebrat, S.: Correspondence between mutation and selection pressure and the genetic code degeneracy in the gene evolution. FGCS 21, 1033–1039 (2005)
Lobry, J.R.: Asymmetric substitution patterns in the two DNA strands of bacteria. Mol. Biol. Evol. 13, 660–665 (1996)
Mrazek, J., Karlin, S.: Strand compositional asymmetry in bacterial and large viral genomes. Proc. Natl. Acad. Sci. USA 95, 3720–3725 (1998)
Frank, A.C., Lobry, J.R.: Asymmetric substitution patterns: a review of possible underlying mutational or selective mechanisms. Gene 238, 65–77 (1999)
Tillier, E.R., Collins, R.A.: The contributions of replication orientation, gene direction, and signal sequences to base-composition asymmetries in bacterial genomes. J. Mol. Evol. 50, 249–257 (2000)
Kowalczuk, M., Mackiewicz, P., Mackiewicz, D., Nowicka, A., Dudkiewicz, M., Dudek, M.R., Cebrat, S.: DNA asymmetry and the replicational mutational pressure. J. Appl. Genet. 42, 553–577 (2001a)
Rocha, E.P.: The replication-related organization of bacterial genomes. Microbiology 150, 1609–1627 (2004)
Gierlik, A., Kowalczuk, M., Mackiewicz, P., Dudek, M.R., Cebrat, S.: Is there replication-associated mutational pressure in the Saccharomyces cerevisiae genome? J. Theor. Biol. 202, 305–314 (2000)
Niu, D.K., Lin, K., Zhang, D.Y.: Strand compositional asymmetries of nuclear DNA in eukaryotes. J. Mol. Evol. 57, 325–334 (2003)
Touchon, M., Nicolay, S., Audit, B., Brodie of Brodie, E.B., d’Aubenton-Carafa, Y., Arneodo, A., Thermes, C.: Replication-associated strand asymmetries in mammalian genomes: Toward detection of replication origins. Proc. Natl. Acad. Sci. USA 102, 9836–9841 (2005)
McInerney, J.O.: Replicational and transcriptional selection on codon usage in Borrelia burgdorferi. Proc. Natl. Acad. Sci. USA 95, 10698–10703 (1998)
Lafay, B., Lloyd, A.T., McLean, M.J., Devine, K.M., Sharp, P.M., Wolfe, K.H.: Proteome composition and codon usage in spirochaetes: species-specific and DNA strand-specific mutational biases. Nucleic Acids Res. 27, 1642–1649 (1999)
Mackiewicz, P., Gierlik, A., Kowalczuk, M., Dudek, M.R., Cebrat, S.: How does replication-associated mutational pressure influence amino acid composition of proteins? Genome Res. 9, 409–416 (1999a)
Rocha, E.P., Danchin, A., Viari, A.: Universal replication biases in bacteria. Mol. Microbiol. 32, 11–16 (1999)
Zhu, C.T., Zeng, X.B., Huang, W.D.: Codon usage decreases the error minimization within the genetic code. J. Mol. Evol. 57, 533–537 (2003)
Mackiewicz, P., Gierlik, A., Kowalczuk, M., Szczepanik, D., Dudek, M.R., Cebrat, S.: Mechanisms generating long-range correlation in nucleotide composition of the Borrelia burgdorferi genome. Physica A 273, 103–115 (1999b)
Fraser, C.M., Casjens, S., Huang, W.M., Sutton, G.G., Clayton, R., Lathigra, R., White, O., Ketchum, K.A., Dodson, R., Hickey, E.K., et al. (38 co-authors): Genomic sequence of a Lyme disease spirochaete Borrelia burgdorferi. Nature 390, 580–586 (1997)
GenBank, ftp://www.ncbi.nlm.nih.gov
Kowalczuk, M., Mackiewicz, P., Mackiewicz, D., Nowicka, A., Dudkiewicz, M., Dudek, M.R., Cebrat, S.: High correlation between the turnover of nucleotides under mutational pressure and the DNA composition. BMC Evol. Biol. 1, 13 (2001b)
Drake, J.W., Charlesworth, B., Charlesworth, D., Crow, J.F.: Rates of spontaneous mutation. Genetics 148, 1667–1686 (1998)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2008 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Mackiewicz, P. et al. (2008). Optimisation of Asymmetric Mutational Pressure and Selection Pressure Around the Universal Genetic Code. In: Bubak, M., van Albada, G.D., Dongarra, J., Sloot, P.M.A. (eds) Computational Science – ICCS 2008. ICCS 2008. Lecture Notes in Computer Science, vol 5103. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-69389-5_13
Download citation
DOI: https://doi.org/10.1007/978-3-540-69389-5_13
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-69388-8
Online ISBN: 978-3-540-69389-5
eBook Packages: Computer ScienceComputer Science (R0)