Abstract
Recent developments in experimental technologies have transformed traditional microbial physiology into a data-rich or -omics discipline. As a result, it has caused a renaissance of the mathematical analysis of biological systems and stimulated the development of systems biology workflows which aim to provide a holistic vision of all cellular functions through genomics, transcriptomics, proteomics, metabolomics, and fluxomic data. In silico modeling of metabolic systems has become a powerful tool, providing insight into the complex processes in cellular metabolism and their underlying regulatory mechanisms, as well as potentially improving the biotechnological design of microbial strains with desired properties. In this chapter, we provide an overview of the systems biology of methane utilization, as an example of one unique microbial function that has been dissected using - omics technologies. We discuss the most recent advances in large-scale investigation and computational representation of related metabolic networks as well as highlight some challenges for further developments in the field.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
7.1 Introduction
Recent developments in experimental technologies have transformed traditional microbial physiology into a data-rich or -omics discipline (Kalyuzhnaya et al. 2015; Khadem et al. 2011; Wertz et al. 2012). As a result, it has caused a renaissance of the mathematical analysis of biological systems (Karr et al. 2012) and stimulated the development of systems biology workflows which aim to provide a holistic vision of all cellular functions through genomics, transcriptomics, proteomics, metabolomics, and fluxomic data (Cavill et al. 2015; Covert et al. 2001; Crowther et al. 2008; Haque et al. 2015; Leak and Dalton 1986b; Lee et al. 2006b; Machado and Herrgård 2014; Yizhak et al. 2010). In silico modeling of metabolic systems has become a powerful tool, providing insight into the complex processes in cellular metabolism and their underlying regulatory mechanisms, as well as potentially improving the biotechnological design of microbial strains with desired properties (Alon 2006; Lee et al. 2006b). In this chapter, we provide an overview of the systems biology of methane utilization, as an example of one unique microbial function that has been dissected using -omics technologies. We discuss the most recent advances in large-scale investigation and computational representation of related metabolic networks as well as highlight some challenges for further developments in the field.
7.2 Bacterial Methane Metabolism
The ability to use methane, i.e., methanotrophy, has been always considered as one of the most unique microbial functions. For years methanotrophy has been attributed to bacteria, known as methanotrophs, and described for Alphaproteobacteria, Gammaproteobacteria, and Verrucomicrobia (reviewed in Kalyuzhnaya et al. 2015; Trotsenko and Murrell 2008). While the ability of methane oxidation has also been demonstrated for some members of Archaea, in this chapter, we will only cover systems approaches applied to bacterial metabolic networks. The core elements of methane utilization were well established by early 1980s (Anthony 1982). Over the past decade, systems biology approaches helped to refine these established metabolic networks (summarized in Table 7.1). The massive amount of -omics information also highlighted a myriad of complexities and exceptions, which continue to challenge our knowledge of methane utilization.
Methanotrophs oxidize methane through the use of copper-dependent particulate methane monooxygenase (pMMO) or iron-linked soluble methane monooxygenase (sMMO) enzymes (Hakemian and Rosenzweig 2007; Culpepper and Rosenzweig 2012; Sirajuddin and Rosenzweig 2015; Chan and Yu 2008). Both enzymes require oxygen and convert methane into methanol, which is further oxidized into formaldehyde by a PQQ-dependent methanol dehydrogenase (MDH) (Chistoserdova 2011). While MDH and its corresponding genes mxaFI are the most well-known methanol oxidizers, there are homologues (xoxF and mdh2) that exist in other species (Kalyuzhnaya et al. 2008; Semrau et al. 2018; reviewed in Chap. 4). Many steps of methanotrophy are interconnected with carbon assimilation (reviewed in Chistoserdova and Lidstrom 2013; Kalyuzhnaya et al. 2015; Trotsenko and Murrell 2008). In some species, formaldehyde is incorporated into fructose 6-phosphate in a two-step reaction driven by one fused or two individual enzymes, hexulose phosphate synthase and isomerase (Orita et al. 2005, 2006; Rozova et al. 2017). These two steps of assimilation are the core of the ribulose monophosphate (RuMP) pathway, which additionally includes pentose phosphate pathway (PPP) reactions and at least one of the glycolytic pathways (Embden–Meyerhof–Parnas, Entner–Doudoroff, or Bifidobacterium shunt). Thus, methane-derived carbon enters the canonical sugar catabolic pathways for redox power regeneration or the production of the main precursors for biosynthesis (Fig. 7.1). So far, the RuMP pathway has been found only in gammaproteobacterial methanotrophs. In the majority of known methane-consuming Alphaproteobacteria, formaldehyde is first oxidized to formate, which is then incorporated into biomass via tetrahydrofolate-linked C 1 -transfer reactions and the serine cycle (Matsen et al. 2013; Yang et al. 2013). It has been postulated that to be an efficient and self-sustained pathway for C 1 -carbon utilization, the serine cycle must be coupled with a glyoxylate regeneration pathway, such as the glyoxylate shunt (GS) or ethylmalonyl-coA (EMC) pathway (Anthony 1982; Erb et al. 2007; Fig. 7.2). Both variants of the serine cycle, linked to the GS or EMC pathways, have been identified in methanotrophs. The genetic signatures of the key serine pathway enzymes have also been detected in the genomes of methanotrophic Gammaproteobacteria (reviewed in Kalyuzhnaya et al. 2015); however, none of them indicates the presence of a known glyoxylate regeneration pathway. The serine cycle seems to be a functional pathway in bacteria and contributes to carbon assimilation, most likely as a supplementary metabolic module interlinked with the pyruvate node (Kalyuzhnaya et al. 2015; Ward et al. 2004). All methanotrophic Verrucomicrobia and some Proteobacteria are autotrophs, which assimilate CO2 via the Calvin cycle (Khadem et al. 2011; Taylor et al. 1981; Vorobev et al. 2011).
(a) The ribulose monophosphate (RuMP) pathway variants. (b) Energy and carbon balance of each RuMP. Key steps for aldol condensation of formaldehyde with ribulose monophosphate and isomerization of hexulose 6-phosphate to fructose 6-phosphate. (2–5) Various routes that the fructose 6-phosphate utilizes: (2) the Bifidobacterium shunt contributes to fermentation and recycling of acetyl-CoA back to pentose phosphate pathway (PPP), (3) the Embden–Meyerhof–Parnas (EMP) pathway, (4) the Entner–Doudoroff (ED) pathway, (5) the dissimilatory pentose phosphate pathway (dPPP). Green labels indicate intermediates which enter the pentose phosphate pathway (PPP) for regeneration of ribulose 5-phosphate. Key enzymes: (A) hexulose phosphate synthase, (B) hexulose phosphate isomerase, (C) phosphoketolase, (D) PPi-dependent phosphofructokinase, (E) 6-phosphogluconate dehydratase, (F) 2-keto-3-deoxy-6-phosphogluconate (KDPG) aldolase, (G) 6-phosphogluconate dehydrogenase (decarboxylating)
Serine cycle and its variants. (1) Three various pathways have been proposed for methylenetetrahydrofolate formation, including spontaneous condensation with tetrahydrofolate, condensation facilitated by a formaldehyde-activating enzyme (Fae), and tetrahydrofolate-linked C1-transfer; (2) the core serine pathway involves the assimilation of methylenetetrahydrofolate into phosphoglycerate, which is then used for assimilation and regeneration of C1-accepting molecule, glycine (formed from glyoxylate); (3) the glyoxylate shunt is a truncated TCA cycle allowing replenishment of glyoxylate (Anthony 1982); (4) the ethylmalonyl-CoA cycle is an alternative pathway for glyoxylate regeneration for bacteria lacking isocitrate lyase (Peyraud et al. 2009, 2011; Yang et al. 2013). Key enzymes: (A) serine hydroxymethyltransferase, (B) serine glyoxylate aminotransferase, (C) hydroxypyruvate reductase, (D) glyceratekinase, (E) enolase, (F) phospho(enol)pyruvate (PEP) carboxylase or alternative pathway to converting C3 to C4 compounds, (G) malate dehydrogenase, (H) malate thiokinase, (I) malyl-CoA lyase, (J) ß-methylmalyl-CoA lyase, (K) isocitrate lyase
All three groups of methanotrophs have functional TCA cycles and quite complex and often redundant sets of electron transfer systems (ETS) (Fig. 7.3). It is predicted that electrons from methane oxidation are transferred to oxygen, regenerating energy for biosynthesis. However, anaerobic respiratory pathways and fermentation pathways have also been described (Kalyuzhnaya et al. 2013; Kits et al. 2015).
Electron transfer reactions in Methylomicrobium alcaliphilum 20Z. 1–3 represent different pathways for redox supply for pMMO reaction: (1) direct coupling mode, (2) uphill electron transfer mode, and (3) redox mode. M. alcaliphilum 20Z has various terminal cytochrome oxidases, whose expressions depend on availability of oxygen and copper
It has been predicted that the methane oxidation machinery has relatively high basal energy requirements and about 25% of consumed methane is directed toward functions required to sustain a metabolically active state (Akberdin et al. 2018). That might explain the conservation of numerous PPi-dependent reactions in all main groups of methanotrophs (reviewed in Chap. 6). The estimated growth-dependent energy consumptions fall into a range that is typical for many microbial species (Akberdin et al. 2018).
7.2.1 Genomics
Whole-genome sequencing is a powerful tool that allows one to gain an initial basis for and a holistic insight into the metabolic potential of a studied microorganism. A number of complete or gapped genome sequences for a variety of methanotrophs are now available, which not only provide a fundamental platform for implementation of systems biology strategies for metabolic network reconstruction (Sipkema et al. 2000; Vuilleumier et al. 2009, 2012) but also facilitate comparative and evolutionary analysis of microbial diversity across all genera of methanotrophs (Chistoserdova 2011; Tamas et al. 2014; Tavormina et al. 2017). Genome mining led to the discovery of redundant methanol-utilization systems (Chistoserdova 2011; Chu and Lidstrom 2016; Ettwig et al. 2010), genetic elements for a copper acquisition system (such as methanobactin and its biosynthesis) (Semrau et al. 2013), carbon assimilation pathways in methanotrophic Verrucomicrobia (Op den Camp et al. 2009; Anvar et al. 2014; Khadem et al. 2011, 2012), and key elements of nitrogen metabolism, from N2-fixation to denitrification (Wertz et al. 2012).
The availability of genomes from a variety of pure methanotrophic cultures empowered metagenomics analyses, leading to the identification of molecular and metabolic mechanisms of interactions between methanotrophic strains in complex microbial communities (Beck et al. 2013). Finally, genomic information revealed a number of functions for transfer to and implementation in non-methanotrophic hosts and enabled genome-directed reconstruction and engineering of methanotrophy for biotechnological applications (Fei et al. 2014; Haque et al. 2016; Kalyuzhnaya et al. 2015; Lee and Kim 2015; Henard et al. 2017, also reviewed in Chap. 8).
7.2.2 Transcriptomics
Transcriptomics, or gene-expression profiling, became essential for interpreting genome functionality in a given environmental condition. A number of transcriptomic-based studies have demonstrated how the approach can rapidly advance and facilitate our understanding of the metabolic pathways of C1-metabolism and its underlying regulatory mechanisms. In particular, Luesken et al. (2012) used the approach to understand oxygen production and consumption in Candidatus Methylomirabilis oxyfera. Transcriptomic data has complemented known enzymatic and genomic information to provide a global overview of the metabolic map for methane assimilation in Methylosinus trichosporium OB3b, highlighting the importance of the ethylmalonyl pathways for carbon assimilation (Matsen et al. 2013; Yang et al. 2013). Gene-expression profiles helped to reevaluate C1-assimilation pathways in Methylomicrobium alcaliphilum 20Z (Kalyuzhnaya et al. 2013) and were essential for the discovery of fermentation pathways and the reconstruction of the complete network for sucrose metabolism (But et al. 2015). Detailed transcriptomic analysis of the facultative methanotroph Methylocystis sp. strain SB2 (Vorobev et al. 2014) provided insights into pathways for C2-carbon utilization and metabolic switches between lanthanum (La)- and Ca-dependent growth. The gene expression study conducted by Larsen and Karlsen (2016) highlighted additional copper-linked regulatory switches in M. capsulatus. A total of 137 genes related to energy and transport metabolism were found to be differentially expressed between cells producing sMMO and pMMO. The study led to the detection of novel c-type cytochromes linked to copper-limited growth.
Tavormina and coauthors employed global gene-expression profiling to characterize cellular responses to methane starvation and recovery in the deep-sea aerobic methanotroph Methyloprofundus sedimenti (Tavormina et al. 2017). High transcript levels of methane monooxygenase genes and genes related to methanol utilization and lower transcript levels for other metabolic and housekeeping genes were demonstrated under active growth, while significant reduction of their expression including transcripts encoding methanol dehydrogenases (mxa and xox) was observed during starvation with one notable exception—transcript abundances for genes coding for methane monooxygenases increased considerably during starvation, but more notably, the pmo transcript abundance decreased during the early stage of recovery after methane starvation. Very similar responses have been found in numerous metatranscriptomic datasets, indicating significant metabolic bottlenecks for in situ methane utilization.
7.2.3 Proteomics
In order to discover microbial proteome profiles, high-resolution two-dimensional gel electrophoresis techniques and mass spectrometry approaches have been developed (Bensimon et al. 2012; Otto et al. 2014; Van Oudenhove and Devreese 2013). The data provide key insight into enzyme representation at the whole cell level and highlight posttranslational regulation of metabolic fluxes under different environmental growth conditions. Despite the advances in quantitative proteomics, however, there are not many examples of ubiquitous application of the approach to methanotrophy (Berven et al. 2006; Crombie and Murrell 2014; Gourion et al. 2006; Kao et al. 2004; Laukel et al. 2004). The most obvious cause is due to the structural complexity of the intracytoplasmic membrane that occupies a large portion of the cell volume and contains most of the essential proteins for the initial steps of assimilation pathways (Best and Higgins 1981; Semrau et al. 2010). However, Berven et al. (2006) were able to analyze the outer membrane subproteome of M. capsulatus (Bath). Twenty-eight unique polypeptides were identified from proteins enriched in the outer membrane using two-dimensional gel electrophoresis coupled with electrospray ionization mass spectrometry. Of these, only the location and function of six of the polypeptides were previously known. Bioinformatics allowed predictions to be made for the functions of many of the previously unidentified proteins (β-barrel outer membrane proteins, lipoproteins, or cell surface proteins) that were in very good agreement with experimental data. In addition to this study for M. capsulatus (Bath), Kao et al. (2004) conducted a comprehensive quantitative analysis of the methanotroph’s proteome for cells grown in the presence of different copper ion concentrations. Combining growth-limiting experiments with copper further led to interesting new discoveries such as the presence of all the genes for the serine cycle as well as key differences in expression between copper-starved and control bacteria at key metabolic enzymes such as formyl-methanofuran hydrolase and many of the first or second enzymes in the C1 assimilatory pathways (serine pathway, TCA, and RuMP) (Kao et al. 2004). Similar effects on transcriptional regulation of oxidative enzymes have been shown with MDH and the ratio of lanthanides to calcium in other methanotrophs (Chu and Lidstrom 2016; Haque et al. 2015). The most recent example of the application of proteomics is a study in which the ability of a single bacterial strain, Methylocella silvestris, to grow on methane or short-chain alkane was evaluated. The underlying mechanisms by which the methanotrophic strain used methane or propane as a carbon and energy source was determined (Crombie and Murrell 2014).
7.2.4 Metabolomics
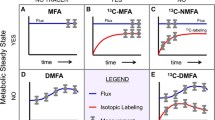
In the context of systems biology, metabolomic approaches for the comprehensive identification and the accurate quantification of metabolites are now regarded as a valuable asset for protein or transcript profiling (Yang et al. 2012). Both targeted and global nontargeted approaches have been applied to investigate metabolic pathways in methanotrophic bacteria (Akberdin et al. 2018; Yang et al. 2013). A dynamic flux analysis based on 13C metabolomics technology was the basis for the quantitative determination of a novel ethylmalonyl-CoA (EMC) pathway as an essential component for glyoxylate regeneration in M. trichosporium OB3b (Yang et al. 2013). A similar dynamic approach allowed the demonstration of the propensity for Methylomicrobium alcaliphilum 20Z to employ a pyrophosphate-mediated EMP variant of the RuMP pathway as the main route for C1-assimilation under oxygen-limiting conditions (Kalyuzhnaya et al. 2013). 13C-carbon tracings have determined the complete oxidative TCA cycle in M. buryantense (Fu et al. 2017). It should be mentioned that despite considerable advances in metabolomics technologies, many critical limitations must be considered, such as the leakage of intracellular metabolites into the solution, the overlap of many compartmentalized metabolic processes in the cell, and the complications linked to the interpretation of 13C single carbon-labeling patterns.
7.3 Metabolic Modeling of Methane Metabolism
Currently, mathematical modeling approaches have become a basic framework for the integration and analysis of experimental data and the iterative investigation of dynamic biological systems (Akberdin et al. 2013; Hübner et al. 2011; Mast et al 2014; Sanchez-Osorio et al. 2014). The general type of model is determined on the basis of the available information, the use of qualitative or quantitative data, and the problem to solve. In a broad sense, the modeling approach as a key component of systems biology is becoming a standard tool for theoretical interpretation of biological systems and prediction of novel genes and their functions.
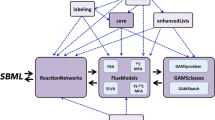
To build a metabolic model, it is necessary to have a reconstruction of the metabolism for the organism of interest. A number of genome-scale biochemical network reconstructions of biotechnology-relevant methanotrophic bacteria are available in BioCyc (http://www.biocyc.org; Caspi et al. 2016) or in the more commonly referred KEGG databases (http://www.genome.jp/kegg/; Kanehisa and Goto 2000). However, they are based on automatic reconstructions, which should be carefully evaluated in accordance with published data for the microbe of interest (substrate consumption and biomass accumulation rates, biomass composition analysis, metabolic pathway validation via enzymatic activity, gene/protein expression, etc.) and converted into a mathematical model that can be analyzed through constraint-based linear programming approaches, such as COBRA (http://opencobra.sourceforge.net/openCOBRA/Welcome.html; Schellenberger et al. 2011) or Pathway Tools (http://bioinformatics.ai.sri.com/ptools/; Karp et al. 2015), at a global systems level and through nonlinear kinetic modeling at a more local mechanistic level. In ideal situations, the reconstruction should be further validated through comparison of model predictions to phenotypic data. Eventually, the metabolic modeling approach provides a scaffold for the integration and analysis of high throughput data such as transcriptomics, proteomics, and metabolomics.
A few metabolic models focused on CH4 metabolism have been constructed (Table 7.2). The computational interpretation of this C1-network has been initiated with a steady-state model of the central metabolism of the facultative methylotrophs Methylobacterium extorquens AM1 and Methylobacillus flagellatum KT (Van Dien and Lidstrom 2002). The computational model has been further improved by implementation of a 13C-fluxomics technique that was also applied to measure the distribution of metabolic fluxes under methanol growth conditions (Peyraud et al. 2011). The network-level analysis of the model indicated that the C1-metabolic core in the methanotroph has a mosaic structure of embedded biochemical cycles. At the same time, it was demonstrated that multiple genes, which encode essential enzymes for methanol assimilation, are not functionally redundant, thereby explaining the structural fragility of the system. It has been concluded that the entire metabolism of the C1-utilization is redox limited (Leak and Dalton 1983; Sipkema et al. 2000; Yoon and Semra 2008). However, contrary to methylotrophic models, the theoretical calculation of methanotrophy showed very poor correlation with measured parameters (Leak and Dalton 1986a). Critical factors that continue to hinder the development of computational models of methane utilization include the lack of fundamental knowledge of the initial steps of methane metabolism, from the catalytic mechanism of methane activation to the structural organization of the methane oxidation apparatus in biological systems.
To address these challenges, the first stoichiometric flux balance model of Methylomicrobium buryatense strain 5G(B1) has been constructed and used for evaluating different metabolic arrangements of methane oxidation and assimilation (Torre et al. 2015). Three arrangements were considered for methane oxidation: redox mode, the currently accepted model in which electrons driving methane oxidation come from NADH produced by formate or formaldehyde oxidation, while electrons produced from methanol oxidation are linked to the redox and used for ATP production; the direct coupling mode, in which methanol oxidation supplies electrons for methane oxidation without any additional inputs; and, finally, the uphill electron transfer mode, in which electrons driving methane oxidation come from cytochrome c to ubiquinone. The model simulations suggested the direct coupling mode is the most compelling mode of methane oxidation, and only this arrangement can support measured growth parameters, while the scenario employing NADH as a possible source of electrons for particulate methane monooxygenase cannot. Recently a developed genome-scale model for a closely related species, Methylomicrobium alcaliphilum 20ZR, has highlighted the dynamic behavior of methane oxidation machinery (Akberdin et al. 2018) and indicated the necessity of an additional constraint on the O2 consumption rate to correctly reproduce experimentally observed parameters (growth rate and corresponding yields). The flux balance analysis of the model combined with global, nontargeted, metabolomic profiling and enzymatic assays highlighted the importance of the substitution of ATP-linked steps with PPi-dependent reactions and supported the presence of a carbon shunt from acetyl-CoA to the pentose-phosphate pathway and highly branched TCA cycle (Akberdin et al. 2018).
A genome-scale metabolic model of Methylococcus capsulatus, tentatively termed iCL656, has been constructed by extending and curating an automatically generated draft reconstruction published in 2012 as part of the Path2Models project (Büchel et al. 2013). As a genome-scale metabolic model, iCL656 includes all major biosynthetic pathways for amino acids, cell-wall components, fatty acids, membrane lipids, and cofactors. In addition, the presence of a detailed representation of the respiratory chain, the RuMP pathway, and the nitrogen metabolism (assimilation and interconversions) provide a comprehensive insight into the metabolism of M. capsulatus. The model’s predictions of growth yields and O2/CH4 ratios agree well with an experimental dataset published by Leak and Dalton (1986a) and indicate that like Methylomicrobium species, M. capsulatus may also use electrons from a methanol oxidation step.
In the most recent development, a kinetic modeling approach that accounts for systems dynamics at the metabolite level as well as regulatory effects has been applied (Akberdin et al. n.d.). Kinetic models are particularly suitable to the study of metabolic systems (Karr et al. 2012; Kitano 2001; Klipp et al. 2008) because they are capable of representing the complex biochemistry of cells in a more complete way compared to other types of models and provide quantitative predictions of the system in response to different inputs. To decipher the puzzle of electron transfer system in methanotrophs, the first kinetic model was recently constructed for Methylomicrobium alcaliphilum 20ZR (Akberdin et al. 2018). Model analysis combined with a mutagenesis study on components of the electron transport chain demonstrates that direct coupling is the most compelling mode of the methane oxidation in the steady state, while NADH is essential for the initial activation of pMMO upon substrate limitation.
7.4 Final Remarks
Overall, the metabolic reconstruction of the methane metabolic network coupled with systems-biology approaches has greatly advanced our understanding of methane utilization and highlighted the importance of further investigation of the initial steps of methane utilization. The redundancy of methane and methanol oxidation machineries and the importance of iron, copper, and lanthanum in governing the switch between the key enzymes also await a thorough investigation. We should also expect advances in metabolic modeling of Alphaproteobacterial and Verrucomicrobial systems, as well as descriptions of metabolic interplays between methanotrophic and non-methanotrophic bacteria in complex microbial communities.
References
Akberdin IR, Kazantsev FV, Ermak TV, Timonov VS, Khlebodarova TM, Likhoshvai VA (2013) In silico cell: challenges and perspectives. Math Biol Bioinform 8(1):295–315
Akberdin IR, Thompson M, Hamilton R, Desai N, Alexander D, Henard CA, Guarnieri MT, Kalyuzhnaya MG (2018) Methane utilization in Methylomicrobium alcaliphilum 20Z R: a systems approach. Sci Rep 8:2512
Akberdin IR, But S, Collins D, Kalyuzhnaya MG (n.d.) Methane utilization in Methylomicrobium alcaliphilum 20ZR: mutagenesis-based investigation and dynamic modeling of the initial steps of methane oxidation. BMC Syst Biol (unpublished data)
Alon U (2006) An introduction to systems biology: design principles of biological circuits. CRC Press, Boca Raton, FL
Anthony C (1982) Biochemistry of methylotrophs. Academic Press, London
Anvar SY, Frank J, Pol A, Schmitz A, Kraaijeveld K, den Dunnen JT, den Camp HJO (2014) The genomic landscape of the verrucomicrobial methanotroph Methylacidiphilum fumariolicum SolV. BMC Genom 15(1):914
Beck DA, Kalyuzhnaya MG, Malfatti S, Tringe SG, del Rio TG, Ivanova N et al (2013) A metagenomic insight into freshwater methane-utilizing communities and evidence for cooperation between the Methylococcaceae and the Methylophilaceae. PeerJ 1:e23
Bensimon A, Heck AJ, Aebersold R (2012) Mass spectrometry-based proteomics and network biology. Ann Rev Biochem 81:379–405
Berven FS, Karlsen OA, Straume AH, Flikka K, Murrell JC, Fjellbirkeland A et al (2006) Analysing the outer membrane subproteome of Methylococcus capsulatus (Bath) using proteomics and novel biocomputing tools. Arch Microbiol 184(6):362–377
Best DJ, Higgins IJ (1981) Methane-oxidizing activity and membrane morphology in a methanolgrown obligate methanotroph, Methylosinus trichosporium OB3b. Microbiology 125(1):73–84
Büchel F, Rodriguez N, Swainston N, Wrzodek C, Czauderna T, Keller R et al (2013) Path2Models: large-scale generation of computational models from biochemical pathway maps. BMC Syst Biol 7(1):116
But SY, Khmelenina VN, Reshetnikov AS, Mustakhimov II, Kalyuzhnaya MG, Trotsenko YA (2015) Sucrose metabolism in halotolerant methanotroph Methylomicrobium alcaliphilum 20Z. Arch Microbiol 197(3):471–480
Caspi R, Billington R, Ferrer L, Foerster H, Fulcher CA, Keseler IM et al (2016) The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res 44(D1):D471–D480
Cavill R, Jennen D, Kleinjans J, Briedé JJ (2015) Transcriptomic and metabolomic data integration. Brief Bioinform 17:bbv090
Chan SI, Yu SSF (2008) Controlled oxidation of hydrocarbons by the membrane-bound methane monooxygenase: the case for a tricopper cluster. Acc Chem Res 41(8):969–979
Chistoserdova L (2011) Modularity of methylotrophy, revisited. Environ Microbiol 13(10):2603–2622
Chistoserdova L, Lidstrom ME (2013) Aerobic methylotrophic prokaryotes. In: DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes. Springer, Berlin, pp 267–285
Chu F, Lidstrom ME (2016) XoxF acts as the predominant methanol dehydrogenase in the type I methanotroph Methylomicrobium buryatense. J Bacteriol 198(8):1317–1325
Covert MW, Schilling CH, Famili I, Edwards JS, Goryanin II, Selkov E, Palsson BO (2001) Metabolic modeling of microbial strains in silico. Trends Biochem Sci 26(3):179–186
Crombie AT, Murrell JC (2014) Trace-gas metabolic versatility of the facultative methanotroph Methylocella silvestris. Nature 510:148–151
Crowther GJ, Kosály G, Lidstrom ME (2008) Formate as the main branch point for methylotrophic metabolism in Methylobacterium extorquens AM1. J Bacteriol 190(14):5057–5062
Culpepper MA, Rosenzweig AC (2012) Architecture and active site of particulate methane monooxygenase. Crit Rev Biochem Mol Biol 47(6):483–492
Erb TJ, Berg IA, Brecht V, Müller M, Fuchs G, Alber BE (2007) Synthesis of C5-dicarboxylic acids from C2-units involving crotonyl-CoA carboxylase/reductase: the ethylmalonyl-CoA pathway. Proc Natl Acad Sci 104(25):10631–10636
Ettwig KF, Butler MK, Le Paslier D, Pelletier E, Mangenot S, Kuypers MM et al (2010) Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464(7288):543
Fei Q, Guarnieri MT, Tao L, Laurens LM, Dowe N, Pienkos PT (2014) Bioconversion of natural gas to liquid fuel: opportunities and challenges. Biotechnol Adv 32(3):596–614
Fu Y, Li Y, Lidstrom M (2017) The oxidative TCA cycle operates during methanotrophic growth of the Type I methanotroph Methylomicrobium buryatense 5GB1. Metab Eng 42:43–51
Gourion B, Rossignol M, Vorholt JA (2006) A proteomic study of Methylobacterium extorquens reveals a response regulator essential for epiphytic growth. Proc Natl Acad Sci 103(35):13186–13191
Hakemian AS, Rosenzweig AC (2007) The biochemistry of methane oxidation. Annu Rev Biochem 76:223–241
Haque MFU, Kalidass B, Bandow N, Turpin EA, DiSpirito AA, Semrau JD (2015) Cerium regulates expression of alternative methanol dehydrogenases in Methylosinus trichosporium OB3b. Appl Environ Microbiol 81(21):7546–7552
Haque MFU, Gu W, DiSpirito AA, Semrau JD (2016) Marker exchange mutagenesis of mxaF, encoding the large subunit of the Mxa methanol dehydrogenase, in Methylosinus trichosporium OB3b. Appl Environ Microbiol 82(5):1549–1555
Henard CA, Smith HK, Guarnieri MT (2017) Phosphoketolase overexpression increases biomass and lipid yield from methane in an obligate methanotrophic biocatalyst. Metab Eng 41:152–158
Hübner K, Sahle S, Kummer U (2011) Applications and trends in systems biology in biochemistry. FEBS J 278(16):2767–2857
Kalyuzhnaya MG, Hristova KR, Lidstrom ME, Chistoserdova L (2008) Characterization of a novel methanol dehydrogenase in representatives of Burkholderiales: implications for environmental detection of methylotrophy and evidence for convergent evolution. J Bacteriol 190(11):3817–3823
Kalyuzhnaya MG, Yang S, Rozova ON, Smalley NE, Clubb J, Lamb A et al (2013) Highly efficient methane biocatalysis revealed in a methanotrophic bacterium. Nat Commun 4:2785
Kalyuzhnaya MG, Puri AW, Lidstrom ME (2015) Metabolic engineering in methanotrophic bacteria. Metab Eng 29:142–152
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30
Kao WC, Chen YR, Eugene CY, Lee H, Tian Q, Wu KM, Tsai SF, Yu SS, Chen YJ, Aebersold R, Chan SI (2004) Quantitative proteomic analysis of metabolic regulation by copper ions in Methylococcus capsulatus (Bath). J Biol Chem 279(49):51554–51560
Karp PD, Latendresse M, Paley SM, Krummenacker M, Ong QD, Billington R et al (2015) Pathway tools version 19.0 update: software for pathway/genome informatics and systems biology. Brief Bioinform 17:bbv079
Karr JR, Sanghvi JC, Macklin DN, Gutschow MV, Jacobs JM, Bolival B, Assad-Garcia N, Glass JI, Covert MW (2012) A whole-cell computational model predicts phenotype from genotype. Cell 150(2):389–401
Khadem AF, Pol A, Wieczorek A, Mohammadi SS, Francoijs KJ, Stunnenberg HG et al (2011) Autotrophic methanotrophy in Verrucomicrobia: Methylacidiphilum fumariolicum SolV uses the Calvin-Benson-Bassham cycle for carbon dioxide fixation. J Bacteriol 193(17):4438–4446
Khadem AF, Wieczorek AS, Pol A, Vuilleumier S, Harhangi HR, Dunfield PF et al (2012) Draft genome sequence of the volcano-inhabiting thermoacidophilic methanotroph Methylacidiphilum fumariolicum strain SolV. J Bacteriol 194(14):3729–3730
Kitano H (ed) (2001) Foundations of systems biology. MIT Press, Cambridge, pp 1–36
Kits KD, Campbell DJ, Rosana AR, Stein LY (2015) Diverse electron sources support denitrification under hypoxia in the obligate methanotroph Methylomicrobium album strain BG8. Front Microbiol 6:1072
Klipp E, Herwig R, Kowald A, Wierling C, Lehrach H (2008) Systems biology in practice: concepts, implementation and application. Wiley, Weinheim
Larsen Ø, Karlsen OA (2016) Transcriptomic profiling of Methylococcus capsulatus (Bath) during growth with two different methane monooxygenases. Microbiologyopen 5(2):254–267
Laukel M, Rossignol M, Borderies G, Völker U, Vorholt JA (2004) Comparison of the proteome of Methylobacterium extorquens AM1 grown under methylotrophic and nonmethylotrophic conditions. Proteomics 4(5):1247–1264
Leak DJ, Dalton H (1983) In vivo studies of primary alcohols, aldehydes and carboxylic acids as electron donors for the methane mono-oxygenase in a variety of methanotrophs. Microbiology 129(11):3487–3497
Leak DJ, Dalton H (1986a) Growth yields of methanotrophs. Appl Microbiol Biotechnol 23(6):470–476
Leak DJ, Dalton H (1986b) Growth yields of methanotrophs 2. A theoretical analysis. Appl Microbiol Biotechnol 23(6):477–481
Lee SW, Keeney DR, Lim DH, Dispirito AA, Semrau JD (2006) Mixed pollutant degradation by Methylosinus trichosporium OB3b expressing either soluble or particulate methane monooxygenase: can the tortoise beat the hare? Appl Environ Microbiol 72(12):7503–7509
Lee SY, Kim HU (2015) Systems strategies for developing industrial microbial strains. Nat Biotechnol 33(10):1061–1072
Luesken FA, Wu ML, Op den Camp HJ, Keltjens JT, Stunnenberg H, Francoijs KJ, Strous M, Jetten MS (2012) Effect of oxygen on the anaerobic methanotroph ‘Candidatus Methylomirabilis oxyfera’: kinetic and transcriptional analysis. Environ Microbiol 14(4):1024–1034
Machado D, Herrgård M (2014) Systematic evaluation of methods for integration of transcriptomic data into constraint-based models of metabolism. PLoS Comput Biol 10(4):e1003580
Mast FD, Ratushny AV, Aitchison JD (2014) Systems cell biology. J Cell Biol 206(6):695–706
Matsen JB, Yang S, Stein LY, Beck DA, Kalyuzhanaya MG (2013) Global molecular analyses of methane metabolism in methanotrophic alphaproteobacterium, Methylosinus trichosporium OB3b. Part I: transcriptomic study. Front Microbiol 4:40
Op den Camp HJ, Islam T, Stott MB, Harhangi HR, Hynes A, Schouten S et al (2009) Environmental, genomic and taxonomic perspectives on methanotrophic Verrucomicrobia. Environ Microbiol Rep 1(5):293–306
Orita I, Yurimoto H, Hirai R, Kawarabayasi Y, Sakai Y, Kato N (2005) The archaeon Pyrococcus horikoshii possesses a bifunctional enzyme for formaldehyde fixation via the ribulose monophosphate pathway. J Bacteriol 187(11):3636–3642
Orita I, Sato T, Yurimoto H, Kato N, Atomi H, Imanaka T, Sakai Y (2006) The ribulose monophosphate pathway substitutes for the missing pentose phosphate pathway in the archaeon Thermococcus kodakaraensis. J Bacteriol 188(13):4698–4704
Otto A, Becher D, Schmidt F (2014) Quantitative proteomics in the field of microbiology. Proteomics 14(4-5):547–565
Peyraud R, Kiefer P, Christen P, Massou S, Portais JC, Vorholt JA (2009) Demonstration of the ethylmalonyl-CoA pathway by using 13C metabolomics. Proc Natl Acad Sci 106(12):4846–4851
Peyraud R, Schneider K, Kiefer P, Massou S, Vorholt JA, Portais JC (2011) Genome-scale reconstruction and system level investigation of the metabolic network of Methylobacterium extorquens AM1. BMC Syst Biol 5(1):189
Rozova ON, But SY, Khmelenina VN, Reshetnikov AS, Mustakhimov II, Trotsenko YA (2017) Characterization of two recombinant 3-hexulose-6-phosphate synthases from the halotolerant obligate methanotroph Methylomicrobium alcaliphilum 20Z. Biochemistry (Mosc) 82(2):176–185
Sanchez-Osorio I, Ramos F, Mayorga P, Dantan E (2014) Foundations for modeling the dynamics of gene regulatory networks: a multilevel-perspective review. J Bioinform Comput Biol 12(01):1330003
Schellenberger J, Que R, Fleming RM, Thiele I, Orth JD, Feist AM et al (2011) Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox v2. 0. Nat Protoc 6(9):1290–1307
Semrau JD, DiSpirito AA, Yoon S (2010) Methanotrophs and copper. FEMS Microbiol Rev 34(4):496–531
Semrau JD, Jagadevan S, DiSpirito AA, Khalifa A, Scanlan J, Bergman BH et al (2013) Methanobactin and MmoD work in concert to act as the ‘copper-switch’ in methanotrophs. Environ Microbiol 15(11):3077–3086
Semrau JD, DiSpirito AA, Wenyu G, Yoon S, Cann I (2018) Metals and Methanotrophy. Appl Environ Microbiol 84(6):e02289-17
Sipkema EM, de Koning W, Ganzeveld KJ, Janssen DB, Beenackers AA (2000) NADH-regulated metabolic model for growth of Methylosinus trichosporium OB3b. Model presentation, parameter estimation, and model validation. Biotechnol Prog 16(2):176–188
Sirajuddin S, Rosenzweig AC (2015) Enzymatic oxidation of methane. Biochemistry 54(14):2283–2294
Tamas I, Smirnova AV, He Z, Dunfield PF (2014) The (d) evolution of methanotrophy in the Beijerinckiaceae—a comparative genomics analysis. ISME J 8(2):369
Tavormina PL, Kellermann MY, Antony CP, Tocheva EI, Dalleska NF, Jensen AJ, Dubilier N, Orphan VJ (2017) Starvation and recovery in the deep-sea methanotroph Methyloprofundus sedimenti. Mol Microbiol 103(2):242–252
Taylor SC, Dalton H, Dow CS (1981) Ribulose-1, 5-bisphosphate carboxylase/oxygenase and carbon assimilation in Methylococcus capsulatus (Bath). Microbiology 122(1):89–94
Torre A, Metivier A, Chu F, Laurens LM, Beck DA, Pienkos PT et al (2015) Genome-scale metabolic reconstructions and theoretical investigation of methane conversion in Methylomicrobium buryatense strain 5G (B1). Microb Cell Fact 14(1):1
Trotsenko YA, Murrell JC (2008) Metabolic aspects of aerobic obligate methanotrophy. Adv Appl Microbiol 63:183–229
Van Dien SJ, Lidstrom ME (2002) Stoichiometric model for evaluating the metabolic capabilities of the facultative methylotroph Methylobacterium extorquens AM1, with application to reconstruction of C3 and C4 metabolism. Biotechnol Bioeng 78(3):296–312
Van Oudenhove L, Devreese B (2013) A review on recent developments in mass spectrometry instrumentation and quantitative tools advancing bacterial proteomics. Appl Microbiol Biotechnol 97(11):4749–4762
Vorobev A, Jagadevan S, Jain S, Anantharaman K, Dick GJ, Vuilleumier S, Semrau JD (2014) Genomic and transcriptomic analyses of the facultative methanotroph Methylocystis sp. strain SB2 grown on methane or ethanol. Appl Environ Microbiol 80(10):3044–3052
Vorobev AV, Baani M, Doronina NV, Brady AL, Liesack W, Dunfield PF, Dedysh SN (2011) Methyloferula stellata gen. nov., sp. nov., an acidophilic, obligately methanotrophic bacterium that possesses only a soluble methane monooxygenase. Int J Syst Evol Microbiol 61(10):2456–2463
Vuilleumier S, Chistoserdova L, Lee MC, Bringel F, Lajus A, Zhou Y et al (2009) Methylobacterium genome sequences: a reference blueprint to investigate microbial metabolism of C1 compounds from natural and industrial sources. PLoS One 4(5):e5584
Vuilleumier S, Khmelenina VN, Bringel F, Reshetnikov AS, Lajus A, Mangenot S et al (2012) Genome sequence of the haloalkaliphilic methanotrophic bacterium Methylomicrobium alcaliphilum 20Z. J Bacteriol 194(2):551–552
Ward N, Larsen Ø, Sakwa J, Bruseth L, Khouri H, Durkin AS et al (2004) Genomic insights into methanotrophy: the complete genome sequence of Methylococcus capsulatus (Bath). PLoS Biol 2(10):e303
Wertz JT, Kim E, Breznak JA, Schmidt TM, Rodrigues JL (2012) Genomic and physiological characterization of the Verrucomicrobia isolate Diplosphaera colitermitum gen. nov., sp. nov., reveals microaerophily and nitrogen fixation genes. Appl Environ Microbiol 78(5):1544–1555
Yang S, Sadilek M, Lidstrom M (2012) Metabolite profiling and dynamic 13C metabolomics of one-carbon assimilation pathways in methylotrophic and methanotrophic bacteria. J Metabolomics Metab 1:2
Yang S, Matsen JB, Konopka M, Green-Saxena A, Clubb J et al (2013) Global molecular analyses of methane metabolism in methanotrophic Alphaproteobacterium, Methylosinus trichosporium OB3b. Part II. Metabolomics and 13C-labeling study. Front Microbiol 4:70
Yizhak K, Benyamini T, Liebermeister W, Ruppin E, Shlomi T (2010) Integrating quantitative proteomics and metabolomics with a genome-scale metabolic network model. Bioinformatics 26(12):i255–i260
Yoon S, Semrau JD (2008) Measurement and modeling of multiple substrate oxidation by methanotrophs at 20 C. FEMS Microbiol Lett 287(2):156–162
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Akberdin, I.R., Thompson, M., Kalyuzhnaya, M.G. (2018). Systems Biology and Metabolic Modeling of C1-Metabolism. In: Kalyuzhnaya, M., Xing, XH. (eds) Methane Biocatalysis: Paving the Way to Sustainability. Springer, Cham. https://doi.org/10.1007/978-3-319-74866-5_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-74866-5_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-74865-8
Online ISBN: 978-3-319-74866-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)