Abstract
MicroRNAs (miRNAs) are a class of short non-coding RNAs (ncRNAs) with typical sequence lengths of 19–25 nucleotides and extraordinary abilities to regulate gene expression. Because miRNAs regulate multiple important biological functions of the cell (proliferation, migration, invasion, apoptosis, differentiation, and drug resistance), their expression is highly controlled. Genetic and epigenetic alterations frequently found in cancer cells can cause aberrant expression of miRNAs and, consequently, of their target genes. The tumor microenvironment can also affect miRNA expression through soluble factors (e.g., cytokines and growth factors) secreted by either tumor cells or non-tumor cells (such as immune and stromal cells). Furthermore, like hormones, miRNAs can be secreted and regulate gene expression in recipient cells. Altered expression levels of miRNAs in cancer cells determine the acquisition of fundamental biological capabilities (hallmarks of cancer) responsible for the development and progression of the disease.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- miRNAs
- ncRNAs
- Epigenetic
- Genetic
- Regulation
- Exosomes
- Cell-to-cell communication
- Tumor microenvironment
- Immunology
1 Introduction
The central dogma of molecular biology, which describes the flow of genetic information from DNA to proteins through the synthesis of RNA, can be summarized in a short sentence: “DNA transcribes RNA, which is then translated into proteins” [1]. However, protein-coding genes represent only 3% of the human genome. Our remaining genes code for RNAs that are not translated into proteins (ncRNAs). As their function was unknown for many years, these ncRNAs have been considered the dark matter of the human genome. Recently, there has been increasing evidence that ncRNAs actually play a critical role in the regulation of gene expression in both healthy and diseased cells [2, 3]. The family of ncRNAs can be divided into three major classes according to size: short, mid-length, and long ncRNAs. MiRNAs are short ncRNAs with an approximate length of 21 nucleotides, and due to their extraordinary abilities to regulate gene expression, they represent the most studied and characterized of all ncRNAs. Multiple cellular processes are controlled by miRNAs: proliferation, migration, invasion, apoptosis, differentiation, and drug resistance. Therefore, aberrant levels of miRNAs can alter these processes and lead to the development of cancer [4]. Genetic mutations (deletions, amplifications, mutation, translocations) and epigenetic alterations (methylation and histone modifications) are frequently observed in cancer cells and can be responsible for the dysregulated expression of miRNAs [5]. The tumor microenvironment can also regulate the expression levels of miRNAs of cancer cells. Soluble factors secreted in the tumor microenvironment, such as growth factors [e.g., epidermal growth factor (EGF)] and cytokines [e.g., transforming growth factor beta (TGF-β), IL-6], can affect miRNA expression [6]. Therefore, an altered tumor microenvironment, for example, during chronic inflammation, can have a great impact on tumor cell functions through the regulation of miRNA expression.
In the past few years, it has been discovered that miRNAs can also perform their regulatory functions outside the cells where they are expressed. Indeed, bioactive miRNA molecules can be secreted from their cells of origin into the extracellular space, be delivered to recipient cells (tumor and non-tumor cells), and regulate the recipient cells’ gene expression in a hormone-like fashion [7]. In this way, tumor cells can regulate their surrounding microenvironments and create favorable conditions allowing them to survive, proliferate, escape from attacks by immune cells, and eventually disseminate to distant organs and generate metastases.
2 MiRNA Biogenesis
The past decade has seen an increasing interest in the roles of miRNAs in tumor cells, due to miRNAs’ abilities to regulate the expression of genes controlling multiple cell processes frequently altered in cancer (e.g., cell cycle, proliferation, migration/invasion, differentiation, apoptosis) [4]. MiRNAs are some of the most abundant genes: there are 2588 miRNAs in humans, according to the latest miRNA database (miRBase). MiRNAs are short ncRNAs ~21 nucleotides (nts) in length, and they are encoded by sequences located within introns or exons of genes or in intergenic regions [8,9,10,11]. MiRNA genes are transcribed in the nucleus by RNA polymerase II (Pol II) as primary transcripts (pri-miRNAs), which are long several kilobases and contain characteristic hairpin structures. Following transcription, the pri-miRNA hairpin structure is recognized by a complex called Microprocessor—composed of the nuclear ribonuclease DROSHA (RNase III) and the essential cofactor DGCR8—which processes the stem-loop and generates a small hairpin-shaped RNA (pre-miRNA) of ~65 nucleotides in length. Then, the pre-miRNA is exported to the cytoplasm, where it matures. The export step is mediated by a transport complex composed of the pre-miRNA, Exportin 5, and the GTP-binding nuclear protein Ran-GTP. The complex drives the pre-miRNA through the nuclear pores and into the cytoplasm where it is released following the disassembly of the transport complex. Next, the pre-miRNA is cleaved by a complex composed of DICER (RNase III-type endonuclease) and the transactivation-responsive RNA-binding protein (TRBP), forming a double-stranded miRNA molecule. During the final maturation step, this RNA duplex is incorporated into the RNA-induced silencing complex (RISC), whose primary component is the Argonaut protein 2 (AGO2). In the RISC, the double-stranded miRNA is unwound, generating two single miRNA strands: the mature (guide) and passenger strands [12]. In general, the passenger strand is quickly degraded, whereas the mature miRNA strand, which remains in the RISC, is guided to the target-site sequence of a messenger RNA (mRNA) to either inhibit mRNA’s translation into protein or initiate its degradation. Particularly, miRNAs bind to specific binding sites in the 3′-untranslated regions (3′ UTRs) of mRNAs with different levels of complementarity, affecting the mechanism of translation inhibition. Perfect or near-perfect annealing between a miRNA and its binding site sequence induces the degradation of the target mRNA by RISC, whereas imperfect or partial annealing inhibits ribosomes’ access to the target mRNA, blocking translation. The power of miRNAs in gene expression regulation is vast, as a single miRNA can target hundreds of different mRNAs. Therefore, altered expression of miRNAs can have significant impacts on many biological functions and pathways regulated by target mRNAs, leading to transformation of normal cells into tumor cells and the progression of cancer.
3 Regulation of miRNA Expression in Human Cancers
3.1 Genetic Alteration
Human miRNAs are frequently located at chromosomal fragile sites and in genomic regions involved in cancer [13]. These regions are associated with increased probabilities of various genetic alterations (such as deletions, insertions, amplifications, single point mutations, transitions, and transversions) that occur in different cancer types with different frequencies [5]. These genetic alterations have significant impacts on the cellular levels of miRNAs, leading to their aberrant expression and, accordingly, altered regulatory functions in several pathways involved in tumorigenesis and tumor progression. The first evidence of the correlation between genetic alterations and aberrant miRNA expression was found for the tumor suppressors miR-15a and miR-16-1, which are located at chromosome 13q14. Deletion of this region, which occurs in more than 65% of chronic lymphocytic leukemia (CLL) cases, as well as in 50% of mantle cell lymphomas, 16–40% of multiple myelomas, and 60% of prostate cancers, causes a reduction in miR-15a and miR-16-1 levels [14, 15]. Other examples include the tumor suppressor miR-34a mapped on 11q23-q24, which is lost in breast and lung cancer; miR-123 located at 9q33, which is frequently deleted in non-small cell lung cancer (NSCLC); and miR-145 and miR-143 located at 5q32, which is frequently deleted in myelodysplastic syndrome [13]. On the other hand, genomic amplifications induce elevated levels of some miRNAs. The oncogenic miR-21 is located at 17q23 and is amplified in many tumors, including breast, colon, lung, pancreas, stomach, and prostate tumors [16]. The potent oncogenic miR-17-92 cluster is encoded by the 13q31-32 locus, which undergoes amplification in large B-cell lymphoma, follicular lymphoma, mantle cell lymphoma, and primary cutaneous B-cell lymphoma [13, 17] and is overexpressed in many cancer types, including lymphoma, colon, lung, breast, pancreas, and prostate cancer [16, 18, 19]. Elevated levels of miR-569 are associated with amplification of the 3q26.2 locus in ovarian and breast cancer [20]. Mutations at the DNA level can also affect the transcription of miRNAs or the maturation of pri- and pre-miRNAs, resulting in altered expression of mature miRNAs. Point mutations located in regions containing pri-miRNA recognition and processing motifs, which enhance pri-miRNA processing, determine reductions in mature miRNA levels. For instance, mutations in the basal UG and/or CNNC motifs affect the processing of pri-miR-16 and pri-miR-30a into their mature forms [21].
Mutations can also occur in the key regulators of miRNA biogenesis, DICER and DROSHA, greatly contributing to aberrant expression of miRNAs and cancer development. Indeed, mutations in DICER and DROSHA have been identified in diverse types of cancers and enhance cellular transformation and tumorigenesis [22,23,24,25].
3.2 Epigenetic Regulation
Besides the genetic alteration reported above, epigenetic regulation can also contribute to the aberrant expression of miRNAs in cancer. Indeed, DNA methylation and histone modification play important roles in the regulation of miRNA expression. Many miRNAs are embedded in CpG islands, and their promoter regions can undergo heavy methylation. DNA methylation and histone acetylation induce chromatin remodeling that regulates the transcription machinery’s access to promoter regions, controlling miRNA expression [26]. The first evidence of alteration of miRNA expression by epigenetic changes was in breast and bladder cancer. Particularly, a rapid alteration of miRNA levels was measured in response to inhibitors of histone deacetylase (HDACi) [27] and DNA methylation [28]. Another important example of epigenetic regulation is represented by the miR-34 family, whose expression is repressed by hypermethylation in a variety of cancer types including breast, ovarian, esophageal, gastric, colon, renal, pancreatic, NSCLC, acute lymphocytic leukemia (ALL), and CLL. The downregulation of the tumor suppressor miR-34 is particularly relevant, as miR-34 cooperates with TP53 to suppress prostate cancer by regulating the stem cell compartment [29]. On the other hand, hypomethylation can induce the overexpression of some miRNAs, such as the oncogenic miRNAs (oncomiRs) miR-21 and miR-29b. Hypomethylation determines high levels of these miRNAs in breast cancer and is associated with aggressive characteristics of tumor cells [30].
3.3 MiRNA Regulation by Oncogenes and Tumor Microenvironment
Other mechanisms can regulate the expression levels of miRNAs. Oncogenes and tumor suppressor genes can activate or repress the expression of miRNAs working as transcription factors or repressors. Therefore, either overexpression of oncogenes or downregulation of tumor suppressor genes can impact miRNA levels in cancer cells. For instance, TP53 directly transactivates miR-34 transcription by binding to the miR-34 promoter [31]. The oncogene MYC positively regulates the transcription of the miR-17-92 cluster and can also repress the expression of miR-34 [32].
The tumor microenvironment can also play an important role in regulating miRNA expression in cancer cells. A variety of cytokines produced in the tumor microenvironment are released directly by tumor cells or by tumor associated cells (e.g., immune and stromal cells) [33, 34] and can regulate the expression of miRNAs involved in tumor pathogenesis and progression. For example, the pro-inflammatory cytokine IL-6 induces the expression of TWIST, SNAIL, and ZEB1, key transcription factors that regulate the epithelial-mesenchymal transition (EMT). During EMT, tumor cells lose epithelial characteristics and acquire motility and invasive abilities [35, 36]. TWIST can bind directly to the putative promoter of miR-10b and induce its transcription [37], whereas SNAIL and ZEB1 bind to E-boxes in the miR-34a promoter, repressing miR-34a expression [38]. ZEB1 can also repress the expression of members of the miR-200 family by binding directly to their promoter sequences [39,40,41]. IL-6 can also suppress the expression of miR-34 through the activation of its signaling pathway mediator STAT3, which can bind to a conserved STAT3-binding site in the first intron of the miR-34 gene [42]. The TGF-β/Smad pathway regulates the expression of miR-155, and increased levels of miR-155 induce the acquisition of migration and invasion ability in breast cancer by targeting RHOA [43]. Additionally, growth factors can regulate the expression of miRNAs. For instance, EGF can modulate the expression of miR-30b, miR-30c, miR221, and miR-222, which play important roles in gefitinib-induced apoptosis and EMT in NSCLC [44].
Therefore, in addition to genetic alterations in tumor cells, soluble factors released in the tumor microenvironment play significant roles in the regulation of the expression of miRNAs in tumor cells. This may be particularly important under conditions of chronic inflammation [45, 46], both in the early stages of cancer formation and later during the progression of the disease, as cytokines released in the microenvironment can provide signals that regulate the expression of miRNAs controlling tumor cell growth, differentiation, motility, angiogenesis, and resistance to treatment, as well as anti-tumor immune response [47,48,49,50,51].
4 miRNAs Regulate All Cancer Hallmarks
The hallmarks of cancer are the six fundamental biological capabilities acquired by tumor cells during the multistep process regulating the development of human tumors: self-sufficiency in growth signals, insensitivity to growth-inhibitory (antigrowth) signals, evasion of programmed cell death (apoptosis), limitless replicative potential, sustained angiogenesis, and tissue invasion and metastasis [52]. MiRNAs are involved in the regulation of all of these hallmarks. Accordingly, the aberrant expression of miRNAs significantly impacts the dysregulation of the stepwise cancer development. Some examples are reported below.
To move into an active proliferative state, normal cells require growth signaling provided by growth factors, extracellular matrix components, and cell-to-cell adhesion/interaction molecules. Whereas this dependence on growth signaling is a characteristic of normal cells, tumor cells show a greatly reduced dependence on exogenous growth stimulation. An example is the activation of the oncogene RAS that allows tumor cells to grow independently of external signals. It has been reported that all three RAS genes (K-RAS, N-RAS, and H-RAS) are directly regulated by the let-7 miRNA family [53].
Normal tissue is characterized by the presence of multiple antiproliferative signals that aim to maintain cellular quiescence and tissue homeostasis. The transcription factor FOXO1 is a tumor suppressor that regulates cell cycle progression, proliferation, and apoptosis. FOXO1 is the target of three miRNAs, miR-96, miR-182, and miR-183, and its repression by increased levels of these miRNAs leads to increased proliferation and reduced apoptosis [54].
Tumor cells’ ability to expand in number is determined by the balance between the rates of cell proliferation and cell attrition. Apoptosis, also known as programmed cell death, represents a major source of attrition. Tumor cells are characterized by resistance to apoptosis. The tumor suppressor miR-34 induces cell cycle arrest and subsequent caspase-dependent apoptosis by targeting BCL-2, an important anti-apoptotic regulator [55].
Cells have a finite replicative potential, and after a certain number of cell divisions, they stop growing and enter into senescence. It was reported that during senescence, miR-29 and miR-30 are upregulated and target the oncogene MYBL2, inhibiting DNA synthesis [56].
Angiogenesis is an important process that supplies oxygen and nutrients for cell function and survival. Significant amounts of proangiogenic factors are produced and secreted in tumor tissue to promote neovascularization, which supports the growth of tumor cells. The vascular endothelial growth factor (VEGF) is the most important proangiogenic factor regulating neoangiogenesis and is highly expressed in most cancers. It is induced by low oxygen concentration (hypoxia) in the tumor microenvironment. Hypoxia can modulate expression of several hypoxia-regulated microRNAs (HRMs), some of which, such as miR-210, miR-26, and miR-181, are directly controlled by hypoxia-inducible factor (HIF) [57]. It was found that miR-210 is involved in the regulation of endothelial cell chemotaxis and tubulogenesis [58]. Another HRM, miR-27a, targets the zinc finger gene ZBTB10, a negative regulator of the specific-protein (SP) transcription factors (such as Sp1, Sp3, and Sp4), resulting in the induction of Sp-dependent survival and angiogenic genes, including survivin (BIRC5), VEGF, and VEGF receptor 1 (VEGFR1) [59].
Metastasis is a complex process involving multiple steps that endow tumor cells with invasive properties (migration and invasion ability), intravasation (blood or lymphatic vasculature), blood circulation survival, extravasation, and growth at a new site [60]. It was reported that miR-10b can promote invasion and metastasis by targeting the transcription factor homeobox D10 (HOXD10) [37], which represses the expression of genes involved in cell migration and extracellular matrix remodeling: RHOC, α3 integrin (ITGA3), matrix metalloproteinase-14 (MMP-14), and urokinase-type plasminogen activator receptor (UPAR) [61].
Increasing evidence suggests that two additional hallmarks may be involved in the regulation of cancer development: reprogramming of energy metabolism and evasion of immune destruction. Tumor cells are able to adjust their metabolic pathways according to their energy requirements. The metabolism of glucose and glutamine represents the major source of energy for cells, and pathways using these two nutrients are often altered in cancer cells [62]. Oncogenes and miRNAs are involved in the regulation of these metabolic pathways. For instance, MYC modulates the metabolism of glutamine by repressing miR-23a/b, resulting in increased expression of their target protein, mitochondrial glutaminase [63]. Furthermore, glycolytic pathways are regulated by several miRNAs, including miR-378 [64] and miR-143 [65].
According to the immune surveillance hypothesis, the immune system plays an important role in recognizing and eliminating incipient and advanced stage tumors (metastasis). However, many tumors are able to evade immune cell attack using tumor immune escape mechanisms that include alterations in: tumor antigen processing and presentation by human leukocyte antigen (HLA) class I and II; signal transduction pathways; expression of co-stimulatory and co-inhibitory molecules; and secretion of immune-suppressive mediators [66]. Recently, the roles of miRNAs in regulating tumor immunogenicity and antitumor immune responses have been unveiled [67, 68]. For instance, miR-9 and miR-346 regulate the expression of MHC class I antigen processing machinery (APM) components and interferon (IFN)-induced genes [69, 70], and miR-148a and miR-181a target the expression of HLA-C and HLA-A, respectively [71, 72]. The major histocompatibility complex class I-related molecules (MICs) A and B are the ligands of the activating NK cell receptor NKG2D, which mediates NK cell-mediated cytotoxicity. The expression of MICA and MICB is controlled by miR-20a in breast cancer stem cells, resulting in reduced sensitivity to NK cell-mediated lysis and enhanced metastatic potential [73]. The B7 family includes both co-stimulatory and co-inhibitory molecules (CD80, CD86, CD28, CTLA-4, PD-1, PD-L1, PDL2, ICOSL) that play important roles in immune responses [74]. It has been found that miRNAs can regulate the expression of B7 family members. For instance, the expression of PD-L1 is controlled by miR-570 in gastric cancer [75], miR-34a in acute myeloid leukemia (AML) [76], miR-200 in NSCLC [77], and miR-138-5p in colorectal cancer (CRC) [78]. Recently, it was found that the expression of high levels of miR-124 could reverse the immunosuppressive phenotype of glioma cancer stem cells by targeting STAT3 signaling and reducing the generation of FoxP3+ regulatory T cells (Treg) [79].
5 Circulating miRNAs as Tumor Biomarkers
Recently, it was observed that miRNAs can be released into the extracellular space [80] and can be detected in many biological fluids, such as serum, plasma, urine, saliva, and breast milk [81]. These circulating miRNAs can be actively secreted outside the cell either encapsulated within exosomes [82] or in an extracellular vesicle-free manner associated with the Ago2 protein [83, 84]. They can also be passively secreted into the blood circulation as a result of apoptotic [85] or necrotic cell death [86]. Importantly, aberrant levels of miRNAs can be detected not only in tumor cells but also in the biological fluids of cancer patients, possibly reflecting the expression patterns of the tumor tissues from which circulating miRNAs originate [87]. Due to their extraordinary stability in body fluids, resistance to storage handling, and the ease of assessment by quantitative PCR and miRNA microarrays [80, 88], circulating miRNAs are considered suitable biomarker molecules to differentiate normal from diseased states and monitor both progression of cancer and response to therapy (Table 6.1). Indeed, tumor-specific miRNAs were identified for the first time in the serum of patients with diffuse large B-cell lymphoma; in these patients, high levels of miR-21 were correlated with improved relapse-free survival [89]. Since then, many other studies have been published reporting the potential use of circulating miRNAs as tumor biomarkers in different types of cancer [7], such as miR-141 in prostate cancer [80], miR-486, miR-30d, miR-1, and miR-499 in NSCLC [95], miR-17-3p and miR-92 in CRC [96], miR-195 and let7-a in breast cancer [87], and miR-500 in liver cancer [97].
The potential use of tumor-specific miRNAs as diagnostic markers for cancer has been confirmed not only in serum and plasma but also in other body fluids, such as saliva and urine. For instance, levels of miR-125a and miR-200a were significantly lower in the saliva of cancer patients with oral squamous-cell carcinoma compared to healthy controls [100]. In another study, it was reported that miR-31 levels were higher in the saliva of patients with oral squamous-cell carcinoma compared to healthy controls, and a decrease in miR-31 levels was measured after tumor resection. The latter result was also demonstrated in plasma [101]. Finally, increased levels of miR-126, miR-152, and miR-182 were found in the urine of patients with bladder cancer, and the ratios of miR-126 to miR-152 and miR-182 to miR-152 could indicate the presence of bladder cancer with a specificity of 82% and a sensitivity of 72% [102].
Circulating miRNAs were also studied for their ability to predict prognosis and response to therapy. For instance, miR-125b and miR-532-3p predict the efficacy of rituximab-mediated lymphodepletion in chronic lymphocytic leukemia patients [90]. Six serum microRNAs can predict lymph node metastasis in cervical squamous cell carcinoma patients: miR-1246, miR-20a, miR-2392, miR-3147, miR-3162-5p, and miR-4484 [98]. In hormone-refractory prostate cancer, high serum miR-21 levels could identify patients who were resistant to docetaxel-based chemotherapy [94]. Circulating let-7a and miR-16 levels can predict progression-free survival and overall survival in patients with myelodysplastic syndrome [92]. Circulating miR-200c levels significantly predict prognosis and response to therapy in patients undergoing neoadjuvant chemotherapy for esophageal cancer [99]. Plasma levels of miR-155 can predict response to therapy in patients with chronic lymphocytic leukemia [91]. Finally, high levels of miR-141 in the serum of breast cancer patients were associated with shorter brain metastasis-free survival and were an independent predictor of both progression-free survival and overall survival [93].
6 Extracellular Vesicles Mediate Intercellular Communication in the Tumor Microenvironment
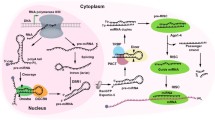
During the past few years, there has been increasing evidence to support the concept that miRNAs are able to mediate intercellular communication. This exchange of genetic information is mediated by extracellular vesicles (EVs), carrying miRNAs and other molecules, that are secreted by donor cells and taken up by recipient cells through several mechanisms [103] (Fig. 6.1). The first evidence of miRNA transfer was provided by Valadi et al. [82], who showed that functional RNA molecules (mRNAs and miRNAs) are transferred between mast cells through exosomes. Exosomes are extracellular vesicles of endosomal origin with diameters ranging from 30 to 100 nm [104]. The generation of exosomes is a highly controlled multi-step process [105] (Fig. 6.1). In the first step, the cell membrane buds inward, forming early endosomes in the endocytic pathway. Then, the early endosome membrane invaginates to generate multivesicular bodies (MVBs), each consisting of a large endosome containing exosomes of different sizes (called intraluminal vesicles or ILVs). During this second inward budding, the exosomes are loaded with different cellular components, including coding RNAs (mRNAs), short and long non-coding RNAs (miRNAs and lncRNAs), proteins, and DNA [82, 106,107,108,109]. In the final step, the fusion of MVBs with the plasma membrane allows the release of ILVs into the extracellular space as exosomes. The regulation of exosome formation, cargo loading, and secretion involves several mechanisms [110]. Ceramide, synthesized by the neutral sphingomyelinase 2 (nSmase2), is involved in the budding of ILVs from MVBs and in exosome secretion [111, 112]. RAB proteins, such as RAB11, RAB27 and RAB35, participate in vesicle trafficking and exosome secretion [113]. The endosomal sorting complex required for transport (ESCRT) mainly regulates protein sorting into MVBs in a ubiquitin-dependent manner [114]; whereas miRNA loading into exosomes is regulated by ceramide and heterogeneous nuclear ribonucleoprotein A2B1 (hnRNPA2B1) [112, 115].
Biogenesis of exosomes and cell-to-cell communication. Exosome biogenesis starts with the inward budding of the plasma membrane to form a clathrin-coated vesicle (CCV) and then an early endosome. Next, a second inward budding of the endosome membrane will generate a multivesicular body (MVB) containing exosomes. During the second inward budding, exosomes are loaded with their cargo (mRNAs, ncRNAs, proteins, and DNA fragments). The MVB can be directed either to the lysosome for degradation and recycling of MVB components or to the plasma membrane for secretion. Finally, the MVB fuses with the plasma membrane, and exosomes are released into the extracellular space. Secreted exosomes can be taken up by recipient cells through several mechanisms: (1) receptor-mediated endocytosis; (2) phago- and micropino-cytosis; (3) direct fusion with the recipient plasma membrane; (4) clathrin-, caveolin-, and lipid raft-mediated endocytosis
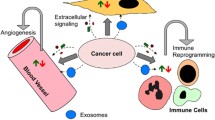
Exosomes are secreted from all types of cells and represent potent vehicles for cell-to-cell communication [116], as they can naturally deliver genetic and protein cargo to recipient cells and regulate these cells’ biological functions. This intercellular communication mechanism is particularly important in cancer, as tumor cells produce significant amounts of EVs. Accordingly, the altered composition of cancer cell-derived exosomes’ cargo can mediate dysregulated signaling. Furthermore, other components of the tumor microenvironment, such as mesenchymal stromal cells (MCS), cancer associated fibroblasts (CAFs), and immune cells (macrophages, dendritic cells, T cells, and NK cells), can participate in EV-mediated crosstalk with tumor cells and regulate their biological functions (Fig. 6.2). This generates a niche that facilitates tumor progression by regulating proliferation, differentiation, angiogenesis, metastasis, anti-tumor immune responses, and drug resistance.
Exosome-mediated cell-to-cell communication in the tumor microenvironment. Exosomes mediate intercellular communication between tumor cells and other cellular components of the tumor microenvironment. EC endothelial cells, MM BM-MSCs multiple myeloma bone marrow mesenchymal stromal cells, DC dendritic cells, Th cells T helper cells, CD8+ CTL CD8+ cytotoxic T lymphocytes, Treg T regulatory lymphocytes, NK natural killer cells, TAM tumor-associated macrophages, CAFs cancer-associated fibroblasts, NF normal fibroblasts
Bone marrow mesenchymal stromal cells from the tumor microenvironment of multiple myeloma patients (MM BM-MSCs) support the growth of multiple myeloma (MM) cells, whereas the bone marrow mesenchymal stromal cells from healthy donors (BM-MSCs) inhibit the growth of MM cells. Exosomes from MM BM-MSCs contain lower levels of the tumor suppressors miR-15 and miR-16 and higher levels of IL-6 and CCL2 compared with exosomes from BM-MSCs. The exosomes released from MM BM-MSCs can deliver their cargo to MM cells, playing a crucial role in MM pathogenesis, tumor growth, and disease progression [117]. The transfer of exosomal miRNAs to endothelial cells can promote angiogenesis and metastasis. It was found that exosomal miR-9 secreted by tumor cells induced endothelial cell migration and an in vivo increase in endothelial density, which promoted tumor growth [118]. Exosomes secreted by the leukemia cell line K562 carry miR-210, which increases tube formation by human umbilical vein endothelial cells [119]. Exosomal miR-210 can also be secreted by breast cancer cells and taken up by endothelial cells, promoting angiogenesis [112]. Exosomal miR-105 secreted from breast cancer cells can target cellular tight junctions and disrupt vascular endothelial barriers during early premetastatic niche formation [120].
Exosomes released by tumor cells can also contribute to the dissemination of malignant cells by remotely regulating a metastatic site. Indeed, exosomes from melanoma cells conditioned lymph node tissue and induced microanatomic niches that allowed metastasis of melanoma cells to lymph nodes [121]. Another study showed that exosomes released by renal cancer stem cells stimulated angiogenesis and the formation of a premetastatic niche in lung tissue [122]. Exosomes can also mediate communication between tumor and immune cells. Exosomes secreted from tumor associated macrophages (TAMs) can deliver miR-223 to breast cancer tumor cells, increasing their invasive abilities [123]. Cancer cell-derived exosomes containing miRNAs can also regulate the functions of immune cells. For instance, exosomal miR-21 and miR-29 released by mouse lung cancer cells can bind to Toll-like receptors (TLRs) 8 and 7 of mouse macrophages and activate the NF-κB pathway. This induces an inflammatory response mediated by TNF-α and IL-6, which promote tumor growth and metastasis [124]. An interesting miRNA-mediated bidirectional crosstalk between neuroblastoma (NBL) cells and monocytes was recently described. Particularly, exosomal miR-21 released by NBL cells can induce the expression of miR-155 in human monocytes. In turn, miR-155 is delivered from monocytes to NBL cells through exosomes and regulates resistance to cisplatin treatment [125].
Tumor cell-derived exosomes can also regulate the functions of immune cells present within the tumor microenvironment [126, 127]. Dendritic cells (DCs) play an important role in activation of anti-tumor immune responses, and their functions can be altered by tumor-derived-exosomes. For instance, exosomal miR-212-3p released from pancreatic tumor cells (PANC-1) can be transferred to DCs and affect their immune functions by inducing immune tolerance [128]. MiR-203 is expressed in PANC-1 cells, and its exosome-mediated delivery induced the downregulation of TLR4, TNF-α, and IL-12, resulting in impairment of immune response activation [129]. Exosomes from nasopharyngeal cancer cells containing miR-24-3p impair T cell proliferation and differentiation into Th1 and Th17 cells and promote the induction of T regulatory CD4+ CD25high Foxp3+ lymphocytes (Tregs) [130]. Tumor-derived microvesicles can deliver miR-214 to CD4+ T cells and promote the expansion of Tregs by targeting phosphatase and tensin homolog (PTEN), resulting in enhanced immune suppression [131]. High levels of miR-210 and miR-23a are present in hypoxic tumor-derived microvesicles and can be transferred to natural killer (NK) cells, leading to impairment of cytotoxicity against different tumor cells in vitro and in vivo [132]. Tumor-derived microvesicles inhibit proliferation and induce apoptosis of activated CD8+ T cells [133].
Cancer-associated fibroblasts (CAFs) are the major components of tumor stroma and can participate in exosome-mediated crosstalk with tumor cells. Indeed, it was found that exosomes released by breast cancer CAFs had increased levels of miRs-21, -378e, and -143 compared to normal fibroblasts. Transfer of these exosomes to breast cancer cells induced a significantly increased capacity to form mammospheres, induced stem cell and EMT markers, and promoted anchorage-independent cell growth [134]. High levels of the pro-metastatic miR-9 are found in various breast cancer cell lines. Exosome-mediated delivery of miR-9 can modify the properties of human breast fibroblasts and promote in vivo tumor growth by enhancing the switch from the normal fibroblast (NF) state to the CAF state [135].
Exosomes can also play important roles in regulating drug resistance [136,137,138]. It was reported that exosomes from tamoxifen-resistant breast cancer cells (MCF-7TamR) could transfer miR-221/222 and induce drug resistance in recipient breast cancer cells by targeting p27 (CDKN1B) and ERα (ESR1) [139]. In advanced renal cell carcinoma (RCC), the bioactive lncRNA named lncRNA Activated in RCC with Sunitinib Resistance (lncARSR) can be incorporated into exosomes. LncARSR can then transmit sunitinib resistance to sensitive cells by competitively binding miR-34/miR-449 to facilitate AXL and MET expression [140]. Exosomes can also perform their regulatory functions by interacting with drugs in the extracellular space. Indeed, it was found that exosomes released by the HER2-overexpressing tumor cell lines SKBR3 and BT474 express a full-length HER2 molecule, can bind to trastuzumab (anti-HER2 humanized monoclonal antibody), and accordingly reduce free molecules of trastuzumab. Exosomes with bound trastuzumab have been found in both HER2-positive tumor cell-conditioned supernatants and serum from breast cancer patients, which resulted in modulation of sensitivity to trastuzumab [141]. Drug-efflux pumps inserted in the exosome membrane can mediate drug sequestration from the cytoplasm. The presence of drug efflux transporters P-glycoprotein (P-gp) and ATP-Binding Cassette G2 (ABCG2) on EVs of breast cancer cells enabled the influx of drugs into the microvesicular compartment, resulting in an active sequestration of chemotherapeutic drugs from the cytoplasm [142, 143]. Then, chemotherapeutic drugs encapsulated inside EVs/exosomes can be expelled by active secretion [144, 145]. Interestingly, the transfer of EVs containing drug efflux pumps from drug-resistant to drug-sensitive tumor cells can contribute to the acquisition of multidrug resistance phenotypes by recipient cells [138].
7 Conclusions
MiRNAs play crucial roles in the regulation of physiological functions in normal cells; therefore, alterations in miRNA expression levels have significant impacts on cells’ biology. Indeed, aberrant miRNA levels are associated with carcinogenesis and cancer progression. Dysregulation of the expression of miRNAs results from regulatory events at both the intracellular level (genetic and epigenetic modifications) and the extracellular level (signaling from the tumor microenvironment). Because a single miRNA has the potential to regulate the expression of up to 100–200 target genes, it is easy to understand how one or a few genetic alterations inside a cell or a disruption in homeostasis in the microenvironment can significantly impact many biological functions, as shown by the roles of miRNAs in regulating the hallmarks of cancer. Altered levels of circulating miRNAs can reflect a pathological status; therefore, these miRNAs can serve as predictive and prognostic biomarkers of cancer. Via secretion into the extracellular space, miRNAs can also perform their regulatory functions outside their cells of origin. Indeed, circulating miRNAs are important mediators of cell-to-cell communication, regulating crosstalk both locally, among different cellular components of the tumor microenvironment, and remotely, by regulating premetastatic niches.
A comprehensive understanding of miRNA functions at multiple levels will allow for the development of more precise and less toxic targeted treatments for cancer.
References
Crick F (1970) Central dogma of molecular biology. Nature 227:561–563
Morris KV, Mattick JS (2014) The rise of regulatory RNA. Nat Rev Genet 15:423–437
Esteller M (2011) Non-coding RNAs in human disease. Nat Rev Genet 12:861–874
Berindan-Neagoe I, Monroig Pdel C, Pasculli B, Calin GA (2014) MicroRNAome genome: a treasure for cancer diagnosis and therapy. CA Cancer J Clin 64:311–336
Tuna M, Machado AS, Calin GA (2016) Genetic and epigenetic alterations of microRNAs and implications for human cancers and other diseases. Genes Chromosomes Cancer 55:193–214
Kedmi M, Sas-Chen A, Yarden Y (2015) MicroRNAs and growth factors: an alliance propelling tumor progression. J Clin Med 4:1578–1599
Cortez MA, Bueso-Ramos C, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA (2011) MicroRNAs in body fluids—the mix of hormones and biomarkers. Nat Rev Clin Oncol 8:467–477
Monteys AM, Spengler RM, Wan J, Tecedor L, Lennox KA, Xing Y, Davidson BL (2010) Structure and activity of putative intronic miRNA promoters. RNA 16:495–505
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A (2004) Identification of mammalian microRNA host genes and transcription units. Genome Res 14:1902–1910
Kim VN, Han J, Siomi MC (2009) Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10:126–139
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M, Croce CM (2004) Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci U S A 101:2999–3004
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, Aldler H, Rattan S, Keating M, Rai K et al (2002) Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A 99:15524–15529
Dong JT, Boyd JC, Frierson HF Jr (2001) Loss of heterozygosity at 13q14 and 13q21 in high grade, high stage prostate cancer. Prostate 49:166–171
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F, Visone R, Iorio M, Roldo C, Ferracin M et al (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A 103:2257–2261
Ota A, Tagawa H, Karnan S, Tsuzuki S, Karpas A, Kira S, Yoshida Y, Seto M (2004) Identification and characterization of a novel gene, C13orf25, as a target for 13q31-q32 amplification in malignant lymphoma. Cancer Res 64:3087–3095
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, Powers S, Cordon-Cardo C, Lowe SW, Hannon GJ, Hammond SM (2005) A microRNA polycistron as a potential human oncogene. Nature 435:828–833
Hayashita Y, Osada H, Tatematsu Y, Yamada H, Yanagisawa K, Tomida S, Yatabe Y, Kawahara K, Sekido Y, Takahashi T (2005) A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 65:9628–9632
Chaluvally-Raghavan P, Zhang F, Pradeep S, Hamilton MP, Zhao X, Rupaimoole R, Moss T, Lu Y, Yu S, Pecot CV et al (2014) Copy number gain of hsa-miR-569 at 3q26.2 leads to loss of TP53INP1 and aggressiveness of epithelial cancers. Cancer Cell 26:863–879
Auyeung VC, Ulitsky I, McGeary SE, Bartel DP (2013) Beyond secondary structure: primary-sequence determinants license pri-miRNA hairpins for processing. Cell 152:844–858
Foulkes WD, Priest JR, Duchaine TF (2014) DICER1: mutations, microRNAs and mechanisms. Nat Rev Cancer 14:662–672
Kumar MS, Lu J, Mercer KL, Golub TR, Jacks T (2007) Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet 39:673–677
Wegert J, Ishaque N, Vardapour R, Georg C, Gu Z, Bieg M, Ziegler B, Bausenwein S, Nourkami N, Ludwig N et al (2015) Mutations in the SIX1/2 pathway and the DROSHA/DGCR8 miRNA microprocessor complex underlie high-risk blastemal type Wilms tumors. Cancer Cell 27:298–311
Merritt WM, Lin YG, Han LY, Kamat AA, Spannuth WA, Schmandt R, Urbauer D, Pennacchio LA, Cheng JF, Nick AM et al (2008) Dicer, Drosha, and outcomes in patients with ovarian cancer. N Engl J Med 359:2641–2650
Cedar H, Bergman Y (2009) Linking DNA methylation and histone modification: patterns and paradigms. Nat Rev Genet 10:295–304
Scott GK, Mattie MD, Berger CE, Benz SC, Benz CC (2006) Rapid alteration of microRNA levels by histone deacetylase inhibition. Cancer Res 66:1277–1281
Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA, Jones PA (2006) Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9:435–443
Cheng CY, Hwang CI, Corney DC, Flesken-Nikitin A, Jiang L, Oner GM, Munroe RJ, Schimenti JC, Hermeking H, Nikitin AY (2014) miR-34 cooperates with p53 in suppression of prostate cancer by joint regulation of stem cell compartment. Cell Rep 6:1000–1007
Li Y, Zhang Y, Li S, Lu J, Chen J, Wang Y, Li Y, Xu J, Li X, Genome-wide DNA (2015) methylome analysis reveals epigenetically dysregulated non-coding RNAs in human breast cancer. Sci Rep 5:8790
Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH, Feldmann G, Yamakuchi M, Ferlito M, Lowenstein CJ et al (2007) Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 26:745–752
Chang TC, Yu D, Lee YS, Wentzel EA, Arking DE, West KM, Dang CV, Thomas-Tikhonenko A, Mendell JT (2008) Widespread microRNA repression by Myc contributes to tumorigenesis. Nat Genet 40:43–50
Dranoff G (2004) Cytokines in cancer pathogenesis and cancer therapy. Nat Rev Cancer 4:11–22
Liotta LA, Kohn EC (2001) The microenvironment of the tumour-host interface. Nature 411:375–379
Sullivan NJ, Sasser AK, Axel AE, Vesuna F, Raman V, Ramirez N, Oberyszyn TM, Hall BM (2009) Interleukin-6 induces an epithelial-mesenchymal transition phenotype in human breast cancer cells. Oncogene 28:2940–2947
Ortiz-Montero P, Londono-Vallejo A, Vernot JP (2017) Senescence-associated IL-6 and IL-8 cytokines induce a self- and cross-reinforced senescence/inflammatory milieu strengthening tumorigenic capabilities in the MCF-7 breast cancer cell line. Cell Commun Signal 15:17
Ma L, Teruya-Feldstein J, Weinberg RA (2007) Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449:682–688
Siemens H, Jackstadt R, Hunten S, Kaller M, Menssen A, Gotz U, Hermeking H (2011) miR-34 and SNAIL form a double-negative feedback loop to regulate epithelial-mesenchymal transitions. Cell Cycle 10:4256–4271
Brabletz S, Brabletz T (2010) The ZEB/miR-200 feedback loop—a motor of cellular plasticity in development and cancer? EMBO Rep 11:670–677
Rokavec M, Wu W, Luo JL (2012) IL6-mediated suppression of miR-200c directs constitutive activation of inflammatory signaling circuit driving transformation and tumorigenesis. Mol Cell 45:777–789
Ahn YH, Gibbons DL, Chakravarti D, Creighton CJ, Rizvi ZH, Adams HP, Pertsemlidis A, Gregory PA, Wright JA, Goodall GJ et al (2012) ZEB1 drives prometastatic actin cytoskeletal remodeling by downregulating miR-34a expression. J Clin Invest 122:3170–3183
Rokavec M, Oner MG, Li H, Jackstadt R, Jiang L, Lodygin D, Kaller M, Horst D, Ziegler PK, Schwitalla S et al (2014) IL-6R/STAT3/miR-34a feedback loop promotes EMT-mediated colorectal cancer invasion and metastasis. J Clin Invest 124:1853–1867
Kong W, Yang H, He L, Zhao JJ, Coppola D, Dalton WS, Cheng JQ (2008) MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol Cell Biol 28:6773–6784
Garofalo M, Romano G, Di Leva G, Nuovo G, Jeon YJ, Ngankeu A, Sun J, Lovat F, Alder H, Condorelli G et al (2011) EGFR and MET receptor tyrosine kinase-altered microRNA expression induces tumorigenesis and gefitinib resistance in lung cancers. Nat Med 18:74–82
Coussens LM, Werb Z (2002) Inflammation and cancer. Nature 420:860–867
Quail DF, Joyce JA (2013) Microenvironmental regulation of tumor progression and metastasis. Nat Med 19:1423–1437
Rupaimoole R, Calin GA, Lopez-Berestein G, Sood AK (2016) miRNA deregulation in cancer cells and the tumor microenvironment. Cancer Discov 6:235–246
Dews M, Homayouni A, Yu D, Murphy D, Sevignani C, Wentzel E, Furth EE, Lee WM, Enders GH, Mendell JT, Thomas-Tikhonenko A (2006) Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nat Genet 38:1060–1065
Ma J, Dong C, Ji C (2010) MicroRNA and drug resistance. Cancer Gene Ther 17:523–531
Mehta A, Baltimore D (2016) MicroRNAs as regulatory elements in immune system logic. Nat Rev Immunol 16:279–294
O’Connell RM, Rao DS, Chaudhuri AA, Baltimore D (2010) Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol 10:111–122
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A, Labourier E, Reinert KL, Brown D, Slack FJ (2005) RAS is regulated by the let-7 microRNA family. Cell 120:635–647
Xie L, Ushmorov A, Leithauser F, Guan H, Steidl C, Farbinger J, Pelzer C, Vogel MJ, Maier HJ, Gascoyne RD et al (2012) FOXO1 is a tumor suppressor in classical Hodgkin lymphoma. Blood 119:3503–3511
Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE, Zhai Y, Giordano TJ, Qin ZS, Moore BB et al (2007) p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol 17:1298–1307
Martinez I, Cazalla D, Almstead LL, Steitz JA, DiMaio D (2011) miR-29 and miR-30 regulate B-Myb expression during cellular senescence. Proc Natl Acad Sci U S A 108:522–527
Kulshreshtha R, Davuluri RV, Calin GA, Ivan M (2008) A microRNA component of the hypoxic response. Cell Death Differ 15:667–671
Fasanaro P, D’Alessandra Y, Di Stefano V, Melchionna R, Romani S, Pompilio G, Capogrossi MC, Martelli F (2008) MicroRNA-210 modulates endothelial cell response to hypoxia and inhibits the receptor tyrosine kinase ligand Ephrin-A3. J Biol Chem 283:15878–15883
Mertens-Talcott SU, Chintharlapalli S, Li X, Safe S (2007) The oncogenic microRNA-27a targets genes that regulate specificity protein transcription factors and the G2-M checkpoint in MDA-MB-231 breast cancer cells. Cancer Res 67:11001–11011
Fidler IJ (2003) The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer 3:453–458
Myers C, Charboneau A, Cheung I, Hanks D, Boudreau N (2002) Sustained expression of homeobox D10 inhibits angiogenesis. Am J Pathol 161:2099–2109
Ward PS, Thompson CB (2012) Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell 21:297–308
Gao P, Tchernyshyov I, Chang TC, Lee YS, Kita K, Ochi T, Zeller KI, De Marzo AM, Van Eyk JE, Mendell JT, Dang CV (2009) c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458:762–765
Eichner LJ, Perry MC, Dufour CR, Bertos N, Park M, St-Pierre J, Giguere V (2010) miR-378 (*) mediates metabolic shift in breast cancer cells via the PGC-1beta/ERRgamma transcriptional pathway. Cell Metab 12:352–361
Fang R, Xiao T, Fang Z, Sun Y, Li F, Gao Y, Feng Y, Li L, Wang Y, Liu X et al (2012) MicroRNA-143 (miR-143) regulates cancer glycolysis via targeting hexokinase 2 gene. J Biol Chem 287:23227–23235
Khong HT, Restifo NP (2002) Natural selection of tumor variants in the generation of “tumor escape” phenotypes. Nat Immunol 3:999–1005
Eichmuller SB, Osen W, Mandelboim O, Seliger B (2017) Immune modulatory microRNAs involved in tumor attack and tumor immune escape. J Natl Cancer Inst 109(10). https://doi.org/10.1093/jnci/djx034
Paladini L, Fabris L, Bottai G, Raschioni C, Calin GA, Santarpia L (2016) Targeting microRNAs as key modulators of tumor immune response. J Exp Clin Cancer Res 35:103
Gao F, Zhao ZL, Zhao WT, Fan QR, Wang SC, Li J, Zhang YQ, Shi JW, Lin XL, Yang S et al (2013) miR-9 modulates the expression of interferon-regulated genes and MHC class I molecules in human nasopharyngeal carcinoma cells. Biochem Biophys Res Commun 431:610–616
Bartoszewski R, Brewer JW, Rab A, Crossman DK, Bartoszewska S, Kapoor N, Fuller C, Collawn JF, Bebok Z (2011) The unfolded protein response (UPR)-activated transcription factor X-box-binding protein 1 (XBP1) induces microRNA-346 expression that targets the human antigen peptide transporter 1 (TAP1) mRNA and governs immune regulatory genes. J Biol Chem 286:41862–41870
Kulkarni S, Qi Y, O’HUigin C, Pereyra F, Ramsuran V, McLaren P, Fellay J, Nelson G, Chen H, Liao W et al (2013) Genetic interplay between HLA-C and MIR148A in HIV control and Crohn disease. Proc Natl Acad Sci U S A 110:20705–20710
Liu Y, Zhao JJ, Wang CM, Li MY, Han P, Wang L, Cheng YQ, Zoulim F, Ma X, DP X (2009) Altered expression profiles of microRNAs in a stable hepatitis B virus-expressing cell line. Chin Med J 122:10–14
Wang B, Wang Q, Wang Z, Jiang J, SC Y, Ping YF, Yang J, SL X, Ye XZ, Xu C et al (2014) Metastatic consequences of immune escape from NK cell cytotoxicity by human breast cancer stem cells. Cancer Res 74:5746–5757
Guo Y, Wang AY (2015) Novel immune check-point regulators in tolerance maintenance. Front Immunol 6:421
Wang W, Li F, Mao Y, Zhou H, Sun J, Li R, Liu C, Chen W, Hua D, Zhang X (2013) A miR-570 binding site polymorphism in the B7-H1 gene is associated with the risk of gastric adenocarcinoma. Hum Genet 132:641–648
Wang X, Li J, Dong K, Lin F, Long M, Ouyang Y, Wei J, Chen X, Weng Y, He T, Zhang H (2015) Tumor suppressor miR-34a targets PD-L1 and functions as a potential immunotherapeutic target in acute myeloid leukemia. Cell Signal 27:443–452
Chen L, Gibbons DL, Goswami S, Cortez MA, Ahn YH, Byers LA, Zhang X, Yi X, Dwyer D, Lin W et al (2014) Metastasis is regulated via microRNA-200/ZEB1 axis control of tumour cell PD-L1 expression and intratumoral immunosuppression. Nat Commun 5:5241
Zhao L, Yu H, Yi S, Peng X, Su P, Xiao Z, Liu R, Tang A, Li X, Liu F, Shen S (2016) The tumor suppressor miR-138-5p targets PD-L1 in colorectal cancer. Oncotarget 7:45370–45384
Wei J, Wang F, Kong LY, Xu S, Doucette T, Ferguson SD, Yang Y, McEnery K, Jethwa K, Gjyshi O et al (2013) miR-124 inhibits STAT3 signaling to enhance T cell-mediated immune clearance of glioma. Cancer Res 73:3913–3926
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O’Briant KC, Allen A et al (2008) Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A 105:10513–10518
Weber JA, Baxter DH, Zhang S, Huang DY, Huang KH, Lee MJ, Galas DJ, Wang K (2010) The microRNA spectrum in 12 body fluids. Clin Chem 56:1733–1741
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9:654–659
Arroyo JD, Chevillet JR, Kroh EM, Ruf IK, Pritchard CC, Gibson DF, Mitchell PS, Bennett CF, Pogosova-Agadjanyan EL, Stirewalt DL et al (2011) Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc Natl Acad Sci U S A 108:5003–5008
Turchinovich A, Weiz L, Langheinz A, Burwinkel B (2011) Characterization of extracellular circulating microRNA. Nucleic Acids Res 39:7223–7233
Zernecke A, Bidzhekov K, Noels H, Shagdarsuren E, Gan L, Denecke B, Hristov M, Koppel T, Jahantigh MN, Lutgens E et al (2009) Delivery of microRNA-126 by apoptotic bodies induces CXCL12-dependent vascular protection. Sci Signal 2:ra81
Laterza OF, Lim L, Garrett-Engele PW, Vlasakova K, Muniappa N, Tanaka WK, Johnson JM, Sina JF, Fare TL, Sistare FD, Glaab WE (2009) Plasma MicroRNAs as sensitive and specific biomarkers of tissue injury. Clin Chem 55:1977–1983
Heneghan HM, Miller N, Lowery AJ, Sweeney KJ, Newell J, Kerin MJ (2010) Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Ann Surg 251:499–505
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X et al (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18:997–1006
Lawrie CH, Gal S, Dunlop HM, Pushkaran B, Liggins AP, Pulford K, Banham AH, Pezzella F, Boultwood J, Wainscoat JS et al (2008) Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br J Haematol 141:672–675
Gagez AL, Duroux-Richard I, Lepretre S, Orsini-Piocelle F, Letestu R, De Guibert S, Tuaillon E, Leblond V, Khalifa O, Gouilleux-Gruart V et al (2017) miR-125b and miR-532-3p predict the efficiency of rituximab-mediated lymphodepletion in chronic lymphocytic leukemia patients. A French Innovative Leukemia Organization study. Haematologica 102:746–754
Ferrajoli A, Shanafelt TD, Ivan C, Shimizu M, Rabe KG, Nouraee N, Ikuo M, Ghosh AK, Lerner S, Rassenti LZ et al (2013) Prognostic value of miR-155 in individuals with monoclonal B-cell lymphocytosis and patients with B chronic lymphocytic leukemia. Blood 122:1891–1899
Zuo Z, Calin GA, de Paula HM, Medeiros LJ, Fernandez MH, Shimizu M, Garcia-Manero G, Bueso-Ramos CE (2011) Circulating microRNAs let-7a and miR-16 predict progression-free survival and overall survival in patients with myelodysplastic syndrome. Blood 118:413–415
Debeb BG, Lacerda L, Anfossi S, Diagaradjane P, Chu K, Bambhroliya A, Huo L, Wei C, Larson RA, Wolfe AR et al (2016) miR-141-mediated regulation of brain metastasis from breast cancer. J Natl Cancer Inst 108:djw026
Zhang HL, Yang LF, Zhu Y, Yao XD, Zhang SL, Dai B, Zhu YP, Shen YJ, Shi GH, Ye DW (2011) Serum miRNA-21: elevated levels in patients with metastatic hormone-refractory prostate cancer and potential predictive factor for the efficacy of docetaxel-based chemotherapy. Prostate 71:326–331
Hu Z, Chen X, Zhao Y, Tian T, Jin G, Shu Y, Chen Y, Xu L, Zen K, Zhang C, Shen H (2010) Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J Clin Oncol 28:1721–1726
Ng EK, Chong WW, Jin H, Lam EK, Shin VY, Yu J, Poon TC, Ng SS, Sung JJ (2009) Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut 58:1375–1381
Yamamoto Y, Kosaka N, Tanaka M, Koizumi F, Kanai Y, Mizutani T, Murakami Y, Kuroda M, Miyajima A, Kato T, Ochiya T (2009) MicroRNA-500 as a potential diagnostic marker for hepatocellular carcinoma. Biomarkers 14:529–538
Chen J, Yao D, Li Y, Chen H, He C, Ding N, Lu Y, Ou T, Zhao S, Li L, Long F (2013) Serum microRNA expression levels can predict lymph node metastasis in patients with early-stage cervical squamous cell carcinoma. Int J Mol Med 32:557–567
Tanaka K, Miyata H, Yamasaki M, Sugimura K, Takahashi T, Kurokawa Y, Nakajima K, Takiguchi S, Mori M, Doki Y (2013) Circulating miR-200c levels significantly predict response to chemotherapy and prognosis of patients undergoing neoadjuvant chemotherapy for esophageal cancer. Ann Surg Oncol 20(Suppl 3):S607–S615
Park NJ, Zhou H, Elashoff D, Henson BS, Kastratovic DA, Abemayor E, Wong DT (2009) Salivary microRNA: discovery, characterization, and clinical utility for oral cancer detection. Clin Cancer Res 15:5473–5477
Liu CJ, Kao SY, HF T, Tsai MM, Chang KW, Lin SC (2010) Increase of microRNA miR-31 level in plasma could be a potential marker of oral cancer. Oral Dis 16:360–364
Hanke M, Hoefig K, Merz H, Feller AC, Kausch I, Jocham D, Warnecke JM, Sczakiel G (2010) A robust methodology to study urine microRNA as tumor marker: microRNA-126 and microRNA-182 are related to urinary bladder cancer. Urol Oncol 28:655–661
Mulcahy LA, Pink RC, Carter DR (2014) Routes and mechanisms of extracellular vesicle uptake. J Extracell Vesicles 3:24641
Raposo G, Stoorvogel W (2013) Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol 200:373–383
Simons M, Raposo G (2009) Exosomes—vesicular carriers for intercellular communication. Curr Opin Cell Biol 21:575–581
Mittelbrunn M, Gutierrez-Vazquez C, Villarroya-Beltri C, Gonzalez S, Sanchez-Cabo F, Gonzalez MA, Bernad A, Sanchez-Madrid F (2011) Unidirectional transfer of microRNA-loaded exosomes from T cells to antigen-presenting cells. Nat Commun 2:282
Montecalvo A, Larregina AT, Shufesky WJ, Stolz DB, Sullivan ML, Karlsson JM, Baty CJ, Gibson GA, Erdos G, Wang Z et al (2012) Mechanism of transfer of functional microRNAs between mouse dendritic cells via exosomes. Blood 119:756–766
Ehnfors J, Kost-Alimova M, Persson NL, Bergsmedh A, Castro J, Levchenko-Tegnebratt T, Yang L, Panaretakis T, Holmgren L (2009) Horizontal transfer of tumor DNA to endothelial cells in vivo. Cell Death Differ 16:749–757
Ahadi A, Brennan S, Kennedy PJ, Hutvagner G, Tran N (2016) Long non-coding RNAs harboring miRNA seed regions are enriched in prostate cancer exosomes. Sci Rep 6:24922
Janas T, Janas MM, Sapon K, Janas T (2015) Mechanisms of RNA loading into exosomes. FEBS Lett 589:1391–1398
Trajkovic K, Hsu C, Chiantia S, Rajendran L, Wenzel D, Wieland F, Schwille P, Brugger B, Simons M (2008) Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 319:1244–1247
Kosaka N, Iguchi H, Hagiwara K, Yoshioka Y, Takeshita F, Ochiya T (2013) Neutral sphingomyelinase 2 (nSMase2)-dependent exosomal transfer of angiogenic microRNAs regulate cancer cell metastasis. J Biol Chem 288:10849–10859
Bobrie A, Colombo M, Raposo G, Thery C (2011) Exosome secretion: molecular mechanisms and roles in immune responses. Traffic 12:1659–1668
Gibbings DJ, Ciaudo C, Erhardt M, Voinnet O (2009) Multivesicular bodies associate with components of miRNA effector complexes and modulate miRNA activity. Nat Cell Biol 11:1143–1149
Villarroya-Beltri C, Gutierrez-Vazquez C, Sanchez-Cabo F, Perez-Hernandez D, Vazquez J, Martin-Cofreces N, Martinez-Herrera DJ, Pascual-Montano A, Mittelbrunn M, Sanchez-Madrid F (2013) Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat Commun 4:2980
Mittelbrunn M, Sanchez-Madrid F (2012) Intercellular communication: diverse structures for exchange of genetic information. Nat Rev Mol Cell Biol 13:328–335
Roccaro AM, Sacco A, Maiso P, Azab AK, Tai YT, Reagan M, Azab F, Flores LM, Campigotto F, Weller E et al (2013) BM mesenchymal stromal cell-derived exosomes facilitate multiple myeloma progression. J Clin Invest 123:1542–1555
Zhuang G, Wu X, Jiang Z, Kasman I, Yao J, Guan Y, Oeh J, Modrusan Z, Bais C, Sampath D, Ferrara N (2012) Tumour-secreted miR-9 promotes endothelial cell migration and angiogenesis by activating the JAK-STAT pathway. EMBO J 31:3513–3523
Tadokoro H, Umezu T, Ohyashiki K, Hirano T, Ohyashiki JH (2013) Exosomes derived from hypoxic leukemia cells enhance tube formation in endothelial cells. J Biol Chem 288:34343–34351
Zhou W, Fong MY, Min Y, Somlo G, Liu L, Palomares MR, Yu Y, Chow A, O’Connor ST, Chin AR et al (2014) Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell 25:501–515
Hood JL, San RS, Wickline SA (2011) Exosomes released by melanoma cells prepare sentinel lymph nodes for tumor metastasis. Cancer Res 71:3792–3801
Grange C, Tapparo M, Collino F, Vitillo L, Damasco C, Deregibus MC, Tetta C, Bussolati B, Camussi G (2011) Microvesicles released from human renal cancer stem cells stimulate angiogenesis and formation of lung premetastatic niche. Cancer Res 71:5346–5356
Yang M, Chen J, Su F, Yu B, Su F, Lin L, Liu Y, Huang JD, Song E (2011) Microvesicles secreted by macrophages shuttle invasion-potentiating microRNAs into breast cancer cells. Mol Cancer 10:117
Fabbri M, Paone A, Calore F, Galli R, Gaudio E, Santhanam R, Lovat F, Fadda P, Mao C, Nuovo GJ et al (2012) MicroRNAs bind to Toll-like receptors to induce prometastatic inflammatory response. Proc Natl Acad Sci U S A 109:E2110–E2116
Challagundla KB, Wise PM, Neviani P, Chava H, Murtadha M, Xu T, Kennedy R, Ivan C, Zhang X, Vannini I et al (2015) Exosome-mediated transfer of microRNAs within the tumor microenvironment and neuroblastoma resistance to chemotherapy. J Natl Cancer Inst 107:djv135
Whiteside TL (2016) Exosomes and tumor-mediated immune suppression. J Clin Invest 126:1216–1223
Liu Y, Gu Y, Cao X (2015) The exosomes in tumor immunity. Oncoimmunology 4:e1027472
Ding G, Zhou L, Qian Y, Fu M, Chen J, Chen J, Xiang J, Wu Z, Jiang G, Cao L (2015) Pancreatic cancer-derived exosomes transfer miRNAs to dendritic cells and inhibit RFXAP expression via miR-212-3p. Oncotarget 6:29877–29888
Zhou M, Chen J, Zhou L, Chen W, Ding G, Cao L (2014) Pancreatic cancer derived exosomes regulate the expression of TLR4 in dendritic cells via miR-203. Cell Immunol 292:65–69
Ye SB, Zhang H, Cai TT, Liu YN, Ni JJ, He J, Peng JY, Chen QY, Mo HY, Jun C et al (2016) Exosomal miR-24-3p impedes T-cell function by targeting FGF11 and serves as a potential prognostic biomarker for nasopharyngeal carcinoma. J Pathol 240:329–340
Yin Y, Cai X, Chen X, Liang H, Zhang Y, Li J, Wang Z, Chen X, Zhang W, Yokoyama S et al (2014) Tumor-secreted miR-214 induces regulatory T cells: a major link between immune evasion and tumor growth. Cell Res 24:1164–1180
Berchem G, Noman MZ, Bosseler M, Paggetti J, Baconnais S, Le Cam E, Nanbakhsh A, Moussay E, Mami-Chouaib F, Janji B, Chouaib S (2016) Hypoxic tumor-derived microvesicles negatively regulate NK cell function by a mechanism involving TGF-beta and miR23a transfer. Oncoimmunology 5:e1062968
Wieckowski EU, Visus C, Szajnik M, Szczepanski MJ, Storkus WJ, Whiteside TL (2009) Tumor-derived microvesicles promote regulatory T cell expansion and induce apoptosis in tumor-reactive activated CD8+ T lymphocytes. J Immunol 183:3720–3730
Donnarumma E, Fiore D, Nappa M, Roscigno G, Adamo A, Iaboni M, Russo V, Affinito A, Puoti I, Quintavalle C et al (2017) Cancer-associated fibroblasts release exosomal microRNAs that dictate an aggressive phenotype in breast cancer. Oncotarget 8:19592–19608
Baroni S, Romero-Cordoba S, Plantamura I, Dugo M, D’Ippolito E, Cataldo A, Cosentino G, Angeloni V, Rossini A, Daidone MG, Iorio MV (2016) Exosome-mediated delivery of miR-9 induces cancer-associated fibroblast-like properties in human breast fibroblasts. Cell Death Dis 7:e2312
Azmi AS, Bao B, Sarkar FH (2013) Exosomes in cancer development, metastasis, and drug resistance: a comprehensive review. Cancer Metastasis Rev 32:623–642
Bach DH, Hong JY, Park HJ, Lee SK (2017) The role of exosomes and miRNAs in drug-resistance of cancer cells. Int J Cancer 141:220–230
Sousa D, Lima RT, Vasconcelos MH (2015) Intercellular transfer of cancer drug resistance traits by extracellular vesicles. Trends Mol Med 21:595–608
Wei Y, Lai X, Yu S, Chen S, Ma Y, Zhang Y, Li H, Zhu X, Yao L, Zhang J (2014) Exosomal miR-221/222 enhances tamoxifen resistance in recipient ER-positive breast cancer cells. Breast Cancer Res Treat 147:423–431
Qu L, Ding J, Chen C, ZJ W, Liu B, Gao Y, Chen W, Liu F, Sun W, Li XF et al (2016) Exosome-transmitted lncARSR promotes sunitinib resistance in renal cancer by acting as a competing endogenous RNA. Cancer Cell 29:653–668
Ciravolo V, Huber V, Ghedini GC, Venturelli E, Bianchi F, Campiglio M, Morelli D, Villa A, Della Mina P, Menard S et al (2012) Potential role of HER2-overexpressing exosomes in countering trastuzumab-based therapy. J Cell Physiol 227:658–667
Gong J, Luk F, Jaiswal R, George AM, Grau GE, Bebawy M (2013) Microparticle drug sequestration provides a parallel pathway in the acquisition of cancer drug resistance. Eur J Pharmacol 721:116–125
Ifergan I, Goler-Baron V, Assaraf YG (2009) Riboflavin concentration within ABCG2-rich extracellular vesicles is a novel marker for multidrug resistance in malignant cells. Biochem Biophys Res Commun 380:5–10
Shedden K, Xie XT, Chandaroy P, Chang YT, Rosania GR (2003) Expulsion of small molecules in vesicles shed by cancer cells: association with gene expression and chemosensitivity profiles. Cancer Res 63:4331–4337
Safaei R, Larson BJ, Cheng TC, Gibson MA, Otani S, Naerdemann W, Howell SB (2005) Abnormal lysosomal trafficking and enhanced exosomal export of cisplatin in drug-resistant human ovarian carcinoma cells. Mol Cancer Ther 4:1595–1604
Acknowledgements
Work in Dr. Calin’s laboratory is supported by National Institutes of Health (NIH/NCATS) grant UH3TR00943-01 through the NIH Common Fund, Office of Strategic Coordination (OSC), the NIH/NCI grant 1 R01 CA182905-01, a U54 grant—UPR/MDACC Partnership for Excellence in Cancer Research 2016 Pilot Project, a CLL Moonshot Flagship project and the Estate of C. G. Johnson, Jr.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Anfossi, S., Fu, X., Nagvekar, R., Calin, G.A. (2018). MicroRNAs, Regulatory Messengers Inside and Outside Cancer Cells. In: Mettinger, K., Rameshwar, P., Kumar, V. (eds) Exosomes, Stem Cells and MicroRNA. Advances in Experimental Medicine and Biology, vol 1056. Springer, Cham. https://doi.org/10.1007/978-3-319-74470-4_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-74470-4_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-74469-8
Online ISBN: 978-3-319-74470-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)