Abstract
Single nucleotide polymorphisms (SNPs) in putative microRNA (miRNA) target sites (miRSNPs) could affect the binding of miRNA with the target and contribute to the susceptibility of human cancers. However, the role of miRSNPs in gastric cancer susceptibility remains largely unknown. Since the over-expression of B7-H1 protein has been reported to be closely related to disease progression of gastric cancer, we investigated the possible role of miRSNPs at the 3′-untranslated region (3′-UTR) of B7-H1 in the risk of developing gastric cancer. In this association study on 205 gastric adenocarcinoma patients and 393 non-cancer controls, we found that the genotype distribution of a common C>G polymorphism (rs4143815) was significantly different between the cases and controls (P = 1.32 × 10−8). Compared with CC homozygotes, GG homozygotes and G allele carriers showed 3.73-fold (P = 2.98 × 10−8) and 1.85-fold (P = 0.002) increased risk of gastric adenocarcinoma, respectively. Stratified analyses indicated that variant genotypes had a strong association with the clinic-pathological features of gastric cancer including differentiation grade, depth of tumor infiltration, and tumor node metastasis (TNM) stage (P < 0.001). Luciferase reporter assay indicated that this SNP might be responsible for aberrant B7-H1 protein expression in gastric cancer by disrupting the interaction between miR-570 and B7-H1 mRNA. These results are consistent with our hypothesis and indicate that genetic polymorphisms influencing B7-H1 expression modify cancer susceptibility.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer remains the fourth most common malignancy and the second leading cause of death by cancer in the world, although a steady decline in the incidence and mortality rates of this disease has been observed in most countries in the last decades (Parkin et al. 2005). Almost two-thirds of the cases occur in developing countries and 42 % in China alone (Parkin et al. 2005). There is now compelling evidence to show that the initiation and progression of gastric cancer are closely related to the cancer microenvironment, in which the B7 superfamily co-stimulatory molecules play an important role. These molecules include activating and inhibitory co-stimulatory molecules that positively and negatively regulate immune responses, respectively. B7-H1 (also known as PD-L1 or CD274) (Dong et al. 1999), one of the most important members of the B7 superfamily co-stimulatory molecules, has been confirmed to negatively regulate immune response by inhibiting T cell activation and proliferation. B7-H1 was also determined to affect the immune escape of cancer cells such that many human cancer types have shown high levels of B7-H1 expression (Dong et al. 2002). Furthermore, B7-H1 has become an indicator of poor prognosis in patients with renal cell carcinoma (Thompson et al. 2004), esophageal cancer (Ohigashi et al. 2005), gastric carcinoma (Wu et al. 2006), breast cancer (Ghebeh et al. 2006), ovarian cancer (Hamanishi et al. 2007), bladder urothelial carcinoma (Inman et al. 2007), or pancreatic cancer (Nomi et al. 2007). We also previously confirmed the wide expression of the B7-H1 protein in gastric cancer, which was significantly related to the clinic-pathological features including tumor size, depth of invasion, lymph node metastasis, and prognosis of patients (Wang et al. 2012; Wu et al. 2006). The broad expression patterns of B7-H1 in cancer microenvironments and its prevalence in clinical and pathological parameters in patients with cancer provide strong evidence that B7-H1 is pathologically relevant.
It was found that the B7-H1 mRNA is abundant in many human non-lymphoid tissues, but low or rare B7-H1 protein is observed in these tissues, excepting for activated immune cells including T cell, B cell, and monocyte/macrophage cells, etc. (Dong et al. 2002). These findings indicate that post-transcriptional regulation could have a crucial role in the control of B7-H1 protein expression. This was confirmed by recent reports that the phosphatase and tensin homolog/phosphatidylinositol-3-kinase (PTEN/PI3K) pathway (Parsa et al. 2007), miR-513 (Gong et al. 2009), and miR-570 (Wang et al. 2012) played important roles in the post-transcriptional regulation of B7-H1 cell-surface expression.
The post-transcriptional regulation through microRNA (miRNA) is one of the most important mechanisms governing the expression of gene. miRNA is an endogenous non-coding RNA of 22 nucleotides, regulating gene by pairing to the 3′-untranslated region (3′-UTR) of messenger RNA (mRNA) of target gene and specifying mRNA cleavage or translational repression (Bartel 2009). When miRNA binds to the 3′-UTR of its mRNA target with an imperfect complementarity, the translation can be inhibited. In this case, there is a reduced level of protein without reduction in the mRNA level. The maximal complementarity is usually restricted to the nucleotides 2–7 in the 5′-end of the miRNA and this region is called ‘seed region’ (Lewis et al. 2003). If a genetic variant occurs in the miRNA target site, the miRNA regulation function can be affected. Several studies have identified single nucleotide polymorphisms (SNPs), the most common genetic variants, in the miRNA target sites are associated with the risk of cancers (Ryan et al. 2010), including lung cancer (Chin et al. 2008), colorectal cancer (Landi et al. 2008), bladder cancer (Yang et al. 2008), oral cancer (Christensen et al. 2009), thyroid cancer (Jazdzewski et al. 2009), and breast cancer (Nicoloso et al. 2010; Saetrom et al. 2009). However, the role of miRSNPs in the susceptibility of gastric cancer is largely unknown.
Previous studies from our laboratory discovered a common guanine-to-cytosine somatic mutation at the SNP rs4143815 locus within the 3′-UTR of B7-H1 gene. Such mutation contributed to the B7-H1 over-expression in gastric cancer by disrupting post-transcriptional and translational controls mediated by miR-570 (Wang et al. 2012). Based on these observations, we hypothesized that the miRSNPs located at the 3′-UTR of B7-H1 may be associated with the risk of gastric cancer. The present report is a case–control study to test this hypothesis.
Materials and methods
Study subjects
A total of 598 genetically unrelated subjects including 393 healthy controls and 205 gastric cancer patients participated in this study after giving written informed consent. They were recruited from the First Affiliated Hospital of Soochow University between March 2007 and May 2009. All subjects were Han Chinese and were raised in Suzhou. They were histologically confirmed by two pathologists. Individuals that were previously cancerous or had metastasizing cancer from other origins were excluded. For gastric cancer patients, the clinic-pathological variables, including tumor markers, histological type, tumor size, tumor area, differentiation grade, depth of tumor infiltration, lymph node metastasis, distant metastasis, and TNM stage, were obtained from the medical records (Table 1). None of the patients had undergone radiotherapy or chemotherapy before surgery. The variables of depth of tumor infiltration, lymph node metastasis, distant metastasis, and TNM stage were examined and staged according to the American Joint Commission for Cancer Staging in 2002.
The controls have no gastrointestinal disorders or personal and familial history of cancers, which were traced back ≥3 generations and laterally to 2nd and 3rd degree relatives. They were obtained from the routine health examinations for early detection of cancer conducted during the same period as the cases were collected. They were randomly selected from a pool of 450 individuals based on a physical examination, and the response rate was 87 %. They were matched with the gastric adenocarcinoma patients by age (59.7 ± 10.9 years for controls and 60.2 ± 11.2 years for patients) and sex (female: 25.2 % for controls and 26.3 % for patients). The selection criteria included no individual history of cancer, and frequency-matched to gastric cancer cases on sex and age. At recruitment, informed consent was obtained from each subject, and each participant was then interviewed to collect detailed information on demographic characteristics. The research protocol was approved by the institutional review board of Soochow University.
Genomic DNA samples
The whole blood samples from patients and controls were collected and stored in Vacutainer® tubes (BD Franklin Lakes, NJ) containing anticoagulant of EDTA. Total genomic DNA was extracted from the whole blood according to the phenol/chloroform method. The purity and concentration of the extracted DNA were determined by UV–vis spectrophotometer. The extracted DNA was stored at 4 °C in TE buffer (10 mmol/L Tris–HCl, 1 mmol/L EDTA, pH 8.0).
Cell lines and cell culture
The human gastric cancer cell line SGC-7901 was purchased from the Type Culture Collection of Chinese Academy of Sciences (Shanghai, China). It was cultured in RPMI 1640 medium supplemented with 10 % fetal calf serum (GBICO BRL, Rockville, MD, USA) at 37 °C in a humidified atmosphere containing 5 % CO2. Cells in the logarithmic growth phase were used for experiments.
miRSNPs selection and genotyping assays
The SNPs in the 3′-UTR of the B7-H1 gene were obtained from the published databases of NCBI dbSNP BUILD 129 and ENSEMBL v58. The online softwares of miRanda (http://www.microrna.org/) and TargetScan v5.1 (http://targetscan.org/) were used to predict the possible miRNA binding sites in B7-H1 3′-UTR (with the GenBank accession no. NM_014143). The SNPs located in putative miRNA binding sites were regarded as miRSNPs.
Then the genotypes of the miRSNPs were determined using sequencing technology (Invitrogen, Shanghai, China). The primers of 5′-GAT ACA CAT TTG GAG GAG ACG-3′ (forward) and 5′-CAA ATA CTC CAT GTT TTA CTA G-3′ (reverse) were used for amplification of B7-H1 3′-UTR. Briefly, each 50-μL reaction contained 10 mM Tris–HCl (pH 8.0), 50 mM KCl, 1.5 mM MgCl2, 0.2 mM of each dNTP, 0.4 μM of each primer, 0.625 U of Taq DNA polymerase, and 1 μL genomic DNA (~100 ng). Thermal cycling was performed in PTC-200 Thermal Cycler (Bio-Rad) with an initial denaturation of 5 min at 94 °C, followed by 35 cycles of 94 °C for 20 s, 55 °C for 30 s, and 72 °C for 1 min. Final extension was completed at 72 °C for 7 min. PCR products were used as templates for sequencing reactions using BigDye terminator kits (PE Biosystems, USA).
3′-UTR luciferase reporter assays
The 3′-UTRs of B7-H1 mRNA (G allele or C allele for SNP rs4143815) were amplified using the primers of 5′-GGC TAG TCT AGA TCC AGC ATT GGA ACT TCT GAT C-3′ (forward) and 5′-GGC TAG TCT AGA CAA ATA CTC CAT GTT TTA CTA G-3′ (reverse). They were then cloned downstream of the firefly luciferase gene in pGL3-Control Vector (Promega), according to the manufacturer’s protocol. Positive clones were selected by restriction digestion using XbaI (MBI) and confirmed by DNA sequencing method. For luciferase reporter assays, human gastric cancer cell line SGC-7901 was plated at 0.2 × 106 cells per well in 24-well dishes and 24 h later co-transfected with Lipofectamine 2000 (Invitrogen), according to the manufacturer’s recommendations. Each co-transfection reaction contained 200 ng of pGL3-G or pGL3-C constructs, 50 pmol/μL of chemically synthesized miRNAs miR-570 (Ambion) or negative control miRNA, 20 ng of pRL-TK plasmid (Promega) that served as a normalizing control, and pGL3-Control Vector or Lipofectamine 2000 alone as blank controls. The transfected SGC-7901 cells were cultured in optimum medium (Invitrogen) with 10 % FBS and 1 % penicillin streptomycin. Each transfection was carried out in triplicate. After 24 h of incubation, cells were collected and analyzed for luciferase activity with the Dual-Luciferase Reporter Assay System (Promega).
Statistical analysis
Chi-square test was used to evaluate differences in frequency distributions of each allele and genotype of the miRSNPs between the cases and controls. Chi-square test was also used to assess the difference between the miRSNPs and the clinic-pathological features. The crude and adjusted odds ratios (ORs) and 95 % confidence intervals (CIs) were obtained to assess the association of the miRSNPs with gastric cancer risk using Unconditional Univariate and Multivariate Logistic Regression models, respectively. Hardy–Weinberg equilibrium of the genotype distribution among control groups was tested by a goodness-of-fit Chi-square test. The genotypes were further stratified by subgroups of the clinic-pathological variables. All of the statistical analyses were carried out using STATA software (version 10.0; StataCorp LP, TX, USA), and were done by two persons independently in a blind fashion. For all tests, P < 0.05 was considered statistically significant.
Results
The SNPs in the B7-H1 3′-UTR in gastric adenocarcinoma patients and controls
From the databases of NCBI dbSNP BUILD 129 and ENSEMBL v58, we found two SNPs rs2297136 and rs4143815 were located in the 3′-UTR of the B7-H1 gene. Using the online softwares of miRanda and TargetScan v5.1, we found 195 miRNAs and 73 miRNAs had possible binding sites in the 3′-UTR of the B7-H1 gene, respectively. From these results, we found that the SNPs rs2297136 and rs4143815 were located in the binding sites of miR-324-5p and miR-570, respectively (Fig. 1a). Moreover, the SNP rs4143815 was located in the ‘seed region’ of the binding site of miR-570. To test whether these particular SNPs are associated with the occurrence of gastric cancer, we performed a case–control study on 205 gastric adenocarcinoma patients and 393 non-cancer controls. We typed these two SNPs using sequencing method. The typical genotyping results are presented in Fig. 1b.
The SNPs in the B7-H1 3′-UTR. a The schematic of B7-H1 mRNA harbors putative miR-324-5p and miR-570 binding sites and two SNPs in the 3′-UTR. The SNP rs4143815 G>C is located within the ‘seed region’ of the miR-570 binding site. The sequences of miR-324-5p and miR-570 as well as their binding sites are shown. b The typical genotyping results of SNPs rs2297136 and rs4143815
The association of SNP rs4143815 with gastric adenocarcinoma risk
The genotype frequencies of rs2297136 and rs4143815 and their associations with gastric adenocarcinoma risk are presented in Table 2. The goodness-of-fit Chi-square test results showed that the genotype distributions of these two SNPs were in Hardy–Weinberg equilibrium (HWE) for the controls. Two-sided Chi-square test revealed that the genotype frequencies of the SNP rs4143815 were significantly different between the cases and controls (P = 1.32 × 10−8), but not for the SNP rs2297136 (P = 0.366). Multivariate logistic regression analysis indicated that the individuals with GG genotype of SNP rs4143815 were associated with a significantly higher risk of gastric adenocarcinoma compared to those with CC genotype (adjusted OR = 3.73, 95 % CI = 2.34–5.94). Moreover, the G allele carriers (individuals with CG or GG genotype) were also related to a significantly higher risk of gastric adenocarcinoma compared to the C allele homozygotes (adjusted OR = 1.85, 95 % CI = 1.25–2.74). Chi-square test results showed that the G allele frequency of rs4143815 was significantly higher (P = 9.70 × 10−10) in the case group (60.5 %) than in the control group (41.7 %). No significant difference in the allele frequencies of rs2297136 was found between the cases and controls (18.8 vs. 16.5 % for the G allele; P = 0.335).
We also analyzed the correlations between the genotypes of SNP rs4143815 and clinicopathological features of gastric adenocarcinoma in this study (Table 3). Strong associations between the variant genotypes and differentiation grade, depth of tumor infiltration, and TNM stage were observed. However, we did not note the statistically significant association of the polymorphism with tumor markers and tumor area.
The effect of SNP rs4143815 on the interaction between miR-570 and the B7-H1 3′-UTR
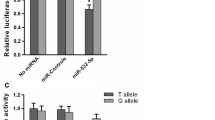
We then determined the effect of this SNP on the gene regulation mediated by miR-570. We devised the B7-H1 3′-UTR luciferase reporter plasmids (rs4143815 G or C allele) and co-transfected them with miR-570 mimics in SGC-7901 cells. pRL-TK plasmids were used to normalize the transfections. For both constructs, in the presence of chemically synthesized miR-570 mimics, the expression of luciferase was significantly reduced (Fig. 2), confirming the functional potential of the mRNA–miRNA duplex. In contrast, the negative control miRNA with no predicting binding site in the B7-H1 3′-UTR has no effect on the luciferase expression (Fig. 2). These findings indicate that miR-570-mediated repression is specific to B7-H1. Furthermore, we found that the C allele resulted in a modest but statistically significant increase of luciferase expression compared to that of the G allele (Fig. 2).
The effect of SNP rs4143815 on the interaction between the B7-H1 3′-UTR and miR-570. Relative luciferase activity in the presence of miR-570 or negative control miRNA is shown for the B7-H1 3′-UTR with G allele and C allele; error bars show maximum values; BC and NC represent blank control and negative control, respectively
Discussion
In this study, we speculated that there are SNPs at the miRNA target site in the 3′-UTR of the B7-H1 gene, which modulate the B7-H1 protein expression, at least in part, through alteration of miRNA target binding capability, ultimately leading to differences in the susceptibility to gastric cancer. This is based on the observations from several reports. Firstly, B7-H1 has been confirmed to promote apoptotic death in activated tumor antigen-specific human T cells and antigen-specific T cells in a mouse P815 tumor model, and also been identified and confirmed promoting the evasion of the cancer cells from apoptosis (Dong et al. 2002). Secondly, the B7-H1 protein has been shown to be over-expressed in many epithelial cancers, including cancers of the stomach (Wu et al. 2006). Increased expression of the B7-H1 protein has been associated with the clinicopathological features including tumor size, depth of tumor infiltration, lymph node metastasis, and prognosis of patients (Wu et al. 2006). Recently, this molecule has been demonstrated to directly participate in skin carcinogenesis (Cao et al. 2011). Thirdly, the expression of the B7-H1 protein has been showed to be post-transcriptionally regulated (Dong et al. 2002; Gong et al. 2009; Parsa et al. 2007; Wang et al. 2012). Fourthly, miRNAs play a critical role in the post-transcriptional regulation of the gene expression (Bartel 2009), and have been increasingly implicated in the control of many pathological processes such as cancers (Croce 2009). Importantly, increasing evidences have demonstrated that miRSNPs can have detrimental effects on the modulation of gene expression and, thus, contribute to the differences between individuals in susceptibility to and severity of cancers (Chin et al. 2008; Christensen et al. 2009; Jazdzewski et al. 2009; Landi et al. 2008; Nicoloso et al. 2010; Ryan et al. 2010; Saetrom et al. 2009; Yang et al. 2008).
We provided evidence in this report that a miRSNP rs4143815 in the miRNA miR-570 binding site in B7-H1 3′-UTR, which can affect B7-H1 protein expression by interfering with miR-570 suppressive function, was significantly related to the risk of gastric adenocarcinoma. On the basis of analysis with 205 gastric adenocarcinoma patients and 393 controls, we found that subjects carrying the GG genotype had a greater than threefold increased risk for developing gastric adenocarcinoma. Moreover, this SNP was also found to be significantly related to the clinicopathological features of gastric cancer including tumor size, differentiation grade, depth of tumor infiltration, lymph node metastasis, and TNM stage (Table 3). This was consistent with our previous observations that the expression of the B7-H1 protein in gastric adenocarcinoma was significantly associated with the clinicopathological features including tumor size, depth of tumor infiltration, lymph node metastasis, and prognosis of patients (Wu et al. 2006). Therefore, this SNP might be a potential hallmark of cancer predisposition. To the best of our knowledge, our findings provide the first evidence that whether B7-H1 gene polymorphisms contribute to the occurrence of gastric adenocarcinoma within a Chinese population. These results further support the hypothesis that the miRSNP may constitute a common susceptibility factor for certain cancers.
Several previous reports imply that multiple mechanisms may contribute to the expression of B7-H1 protein on cancer cells (Liu et al. 2007; Marzec et al. 2008; Parsa et al. 2007; Wang et al. 2012). The findings from our study support the idea that miR-570 mediates B7-H1 translational regulation. A significant decrease of luciferase activity was detected in cells transfected with a pGL-3 control luciferase construct that contains the putative miR-570 binding site in the 3′-UTR of B7-H1 compared with cells transfected with the control empty vector, suggesting that a 3′-UTR-associated B7-H1 translational inhibition exists in non-stimulated cells. Furthermore, the variant allele of the investigated SNP rs4143815 in the B7-H1 gene may cause a loss of the binding site for miR-570. This finding indicated that the SNP rs4143815 may enhance the ability of B7-H1 to promote cancer cell growth, survival and invasion and, thus, partly explain the observed association between the variant genotypes and the clinicopathological features.
As previously reported, somatic mutations from G allele to C allele were commonly occurred at the SNP rs4143815 locus in the G allele carriers (72 of 88 GG homozygotes and 56 of 72 GC heterozygotes), which disrupted the suppressive role of miR-570, thereby contributing to the development of gastric cancer by elevating B7-H1 protein expression (Wang et al. 2012). For CC homozygotes, the situation is different as we did not find any mutations and B7-H1 expression in nearly all cases (41/45) (Wang et al. 2012). Low gastric cancer occurrence rate in this cohort may be due to B7-H1 regulatory factor(s) (e.g. miR-513 or PTEN/PI3K pathway) other than miR-570. As a result, the higher rate of gastric cancer in G allele carriers than CC homozygotes (OR = 1.85, 95 % CI 1.25–2.74; Table 2) is caused by the elevated B7-H1 due to mutation that disrupting the miR-570 binding.
Multiple risk factors have been proposed to play a role in gastric carcinogenesis, such as Helicobacter pylori infection, tobacco smoking, diet, and genetic factors (Correa 1992; Parsonnet et al. 1991). In addition, other factors, such as gastroesophageal reflux disease, obesity, and certain dietary components, are also known to be associated with risk of gastric cancer (Mayne and Navarro 2002). These factors might interact with B7-H1 genotypes or act as potential confounders in the analysis. Unfortunately, information on these factors in our case–control study is not available. It would be interesting to investigate the interaction between B7-H1 genotype and these risk factors on risk of gastric adenocarcinoma in additional studies. Moreover, without detailed information on the survival of gastric adenocarcinoma for our study population, our further analysis of the role of B7-H1 polymorphisms in cancer prognosis was prevented. Another limitation of this study was the small sample sizes. However, our study provides the first evidence that the SNP rs4143815 is a genetic susceptibility factor for the development of gastric adenocarcinoma, with the GG genotype being associated with increased risk of the cancer in a Chinese population. These results further supporting the hypothesis that B7-H1 may play an important role in gastric cancer carcinogenesis. Continued studies will expand on the impact of B7-H1 polymorphisms in patient susceptibility to gastric cancer from other ethnic populations and/or other cancer types, and such study will be of considerable value.
References
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Cao Y, Zhang L, Ritprajak P, Tsushima F, Youngnak-Piboonratanakit P, Kamimura Y, Hashiguchi M, Azuma M (2011) Immunoregulatory molecule B7-H1 (CD274) contributes to skin carcinogenesis. Cancer Res 71: 4737–4741
Chin LJ, Ratner E, Leng S, Zhai R, Nallur S, Babar I, Muller RU, Straka E, Su L, Burki EA, Crowell RE, Patel R, Kulkarni T, Homer R, Zelterman D, Kidd KK, Zhu Y, Christiani DC, Belinsky SA, Slack FJ, Weidhaas JB (2008) A SNP in a let-7 microRNA complementary site in the KRAS 3′ untranslated region increases non-small cell lung cancer risk. Cancer Res 68:8535–8540
Christensen BC, Moyer BJ, Avissar M, Ouellet LG, Plaza SL, McClean MD, Marsit CJ, Kelsey KT (2009) A let-7 microRNA-binding site polymorphism in the KRAS 3′ UTR is associated with reduced survival in oral cancers. Carcinogenesis 30:1003–1007
Correa P (1992) Human gastric carcinogenesis: a multistep and multifactorial process–First American Cancer Society Award Lecture on Cancer Epidemiology and Prevention. Cancer Res 52:6735–6740
Croce CM (2009) Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet 10:704–714
Dong H, Zhu G, Tamada K, Chen L (1999) B7-H1, a third member of the B7 family, co-stimulates T-cell proliferation and interleukin-10 secretion. Nat Med 5:1365–1369
Dong H, Strome SE, Salomao DR, Tamura H, Hirano F, Flies DB, Roche PC, Lu J, Zhu G, Tamada K, Lennon VA, Celis E, Chen L (2002) Tumor-associated B7-H1 promotes T-cell apoptosis: a potential mechanism of immune evasion. Nat Med 8:793–800
Ghebeh H, Mohammed S, Al-Omair A, Qattan A, Lehe C, Al-Qudaihi G, Elkum N, Alshabanah M, Bin Amer S, Tulbah A, Ajarim D, Al-Tweigeri T, Dermime S (2006) The B7-H1 (PD-L1) T lymphocyte-inhibitory molecule is expressed in breast cancer patients with infiltrating ductal carcinoma: correlation with important high-risk prognostic factors. Neoplasia 8:190–198
Gong AY, Zhou R, Hu G, Li X, Splinter PL, O’Hara SP, LaRusso NF, Soukup GA, Dong H, Chen XM (2009) MicroRNA-513 regulates B7-H1 translation and is involved in IFN-gamma-induced B7-H1 expression in cholangiocytes. J Immunol 182:1325–1333
Hamanishi J, Mandai M, Iwasaki M, Okazaki T, Tanaka Y, Yamaguchi K, Higuchi T, Yagi H, Takakura K, Minato N, Honjo T, Fujii S (2007) Programmed cell death 1 ligand 1 and tumor-infiltrating CD8+T lymphocytes are prognostic factors of human ovarian cancer. Proc Natl Acad Sci USA 104:3360–3365
Inman BA, Sebo TJ, Frigola X, Dong H, Bergstralh EJ, Frank I, Fradet Y, Lacombe L, Kwon ED (2007) PD-L1 (B7-H1) expression by urothelial carcinoma of the bladder and BCG-induced granulomata: associations with localized stage progression. Cancer 109:1499–1505
Jazdzewski K, Liyanarachchi S, Swierniak M, Pachucki J, Ringel MD, Jarzab B, de la Chapelle A (2009) Polymorphic mature microRNAs from passenger strand of pre-miR-146a contribute to thyroid cancer. Proc Natl Acad Sci USA 106:1502–1505
Landi D, Gemignani F, Naccarati A, Pardini B, Vodicka P, Vodickova L, Novotny J, Forsti A, Hemminki K, Canzian F, Landi S (2008) Polymorphisms within micro-RNA-binding sites and risk of sporadic colorectal cancer. Carcinogenesis 29:579–584
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB (2003) Prediction of mammalian microRNA targets. Cell 115:787–798
Liu J, Hamrouni A, Wolowiec D, Coiteux V, Kuliczkowski K, Hetuin D, Saudemont A, Quesnel B (2007) Plasma cells from multiple myeloma patients express B7-H1 (PD-L1) and increase expression after stimulation with IFN-{gamma} and TLR ligands via a MyD88-, TRAF6-, and MEK-dependent pathway. Blood 110:296–304
Marzec M, Zhang Q, Goradia A, Raghunath PN, Liu X, Paessler M, Wang HY, Wysocka M, Cheng M, Ruggeri BA, Wasik MA (2008) Oncogenic kinase NPM/ALK induces through STAT3 expression of immunosuppressive protein CD274 (PD-L1, B7-H1). Proc Natl Acad Sci USA 105:20852–20857
Mayne ST, Navarro SA (2002) Diet, obesity and reflux in the etiology of adenocarcinomas of the esophagus and gastric cardia in humans. J Nutr 132:3467S–3470S
Nicoloso MS, Sun H, Spizzo R, Kim H, Wickramasinghe P, Shimizu M, Wojcik SE, Ferdin J, Kunej T, Xiao L, Manoukian S, Secreto G, Ravagnani F, Wang X, Radice P, Croce CM, Davuluri RV, Calin GA (2010) Single-nucleotide polymorphisms inside microRNA target sites influence tumor susceptibility. Cancer Res 70:2789–2798
Nomi T, Sho M, Akahori T, Hamada K, Kubo A, Kanehiro H, Nakamura S, Enomoto K, Yagita H, Azuma M, Nakajima Y (2007) Clinical significance and therapeutic potential of the programmed death-1 ligand/programmed death-1 pathway in human pancreatic cancer. Clin Cancer Res 13:2151–2157
Ohigashi Y, Sho M, Yamada Y, Tsurui Y, Hamada K, Ikeda N, Mizuno T, Yoriki R, Kashizuka H, Yane K, Tsushima F, Otsuki N, Yagita H, Azuma M, Nakajima Y (2005) Clinical significance of programmed death-1 ligand-1 and programmed death-1 ligand-2 expression in human esophageal cancer. Clin Cancer Res 11:2947–2953
Parkin DM, Bray F, Ferlay J, Pisani P (2005) Global cancer statistics, 2002. CA Cancer J Clin 55:74–108
Parsa AT, Waldron JS, Panner A, Crane CA, Parney IF, Barry JJ, Cachola KE, Murray JC, Tihan T, Jensen MC, Mischel PS, Stokoe D, Pieper RO (2007) Loss of tumor suppressor PTEN function increases B7-H1 expression and immunoresistance in glioma. Nat Med 13:84–88
Parsonnet J, Friedman GD, Vandersteen DP, Chang Y, Vogelman JH, Orentreich N, Sibley RK (1991) Helicobacter pylori infection and the risk of gastric carcinoma. N Engl J Med 325:1127–1131
Ryan BM, Robles AI, Harris CC (2010) Genetic variation in microRNA networks: the implications for cancer research. Nat Rev Cancer 10:389–402
Saetrom P, Biesinger J, Li SM, Smith D, Thomas LF, Majzoub K, Rivas GE, Alluin J, Rossi JJ, Krontiris TG, Weitzel J, Daly MB, Benson AB, Kirkwood JM, O’Dwyer PJ, Sutphen R, Stewart JA, Johnson D, Larson GP (2009) A risk variant in an miR-125b binding site in BMPR1B is associated with breast cancer pathogenesis. Cancer Res 69:7459–7465
Thompson RH, Gillett MD, Cheville JC, Lohse CM, Dong H, Webster WS, Krejci KG, Lobo JR, Sengupta S, Chen L, Zincke H, Blute ML, Strome SE, Leibovich BC, Kwon ED (2004) Costimulatory B7-H1 in renal cell carcinoma patients: indicator of tumor aggressiveness and potential therapeutic target. Proc Natl Acad Sci USA 101:17174–17179
Wang W, Sun J, Li F, Li R, Gu Y, Liu C, Yang P, Zhu M, Chen L, Tian W, Zhou H, Mao Y, Zhang L, Jiang J, Wu C, Hua D, Chen W, Lu B, Ju J, Zhang X (2012) A frequent somatic mutation in CD274 3′-UTR leads to protein over-expression in gastric cancer by disrupting miR-570 binding. Hum Mutat 33:480–484
Wu C, Zhu Y, Jiang J, Zhao J, Zhang XG, Xu N (2006) Immunohistochemical localization of programmed death-1 ligand-1 (PD-L1) in gastric carcinoma and its clinical significance. Acta Histochem 108:19–24
Yang H, Dinney CP, Ye Y, Zhu Y, Grossman HB, Wu X (2008) Evaluation of genetic variants in microRNA-related genes and risk of bladder cancer. Cancer Res 68:2530–2537
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 30901360, No. 81270031, and No. 30930085), the 973 project (No. 2013CB530501), the Natural Science Foundation of the Higher Education Institutions of Jiangsu Province (No. 12KJB320012), and a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Conflict of interest
The authors have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Wang, W., Li, F., Mao, Y. et al. A miR-570 binding site polymorphism in the B7-H1 gene is associated with the risk of gastric adenocarcinoma. Hum Genet 132, 641–648 (2013). https://doi.org/10.1007/s00439-013-1275-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-013-1275-6