Abstract
Heterogeneity and variability is ubiquitous in biology and physiology and one of the great modelling challenges is how we cope with and quantify this variability. There are a wide variety of approaches. We can attempt to ignore spatial effects and represent the heterogeneity through stochastic models that evolve only in time, or we can attempt to capture some key spatial components. Alternatively we can perform very detailed spatial simulations or we can attempt to use other approaches that mimic stochasticity in some way, such as by the use of delay models or by using populations of deterministic models. The skill is knowing when a particular model is appropriate to the questions that are being addressed.
In this review, we give a brief introduction to modelling and simulation in Computational Biology and discuss the various different sources of heterogeneity, pointing out useful modelling and analysis approaches. The starting point is how we deal with intrinsic noise; that is, the uncertainty of knowing when a chemical reaction takes place and what that reaction is. These discrete stochastic methods do not follow individual molecules over time; rather they track only total molecular numbers. This leads, in the first instance, to the Stochastic Simulation Algorithm that describes the time evolution of a discrete nonlinear Markov process. From there we consider approaches that are more efficient and effective but still preserve the discreteness of the simulation, the so-called tau-leaping algorithms. We then move to approximations that are continuous in time based around the Chemical Langevin stochastic differential equation. In these contexts we will focus, later in this chapter, on a particular application, namely the behaviour of ion channels dynamics.
In the second part of the review, we address the question of spatial heterogeneity. This involves consideration on the nature of diffusion in crowded spaces and, in particular, anomalous diffusion, a relevant topic (for example) in the analysis and simulation of cell membrane dynamics. We discuss different approaches for capturing this spatial heterogeneity through generalisations of the Stochastic Simulation Algorithm and that eventually leads us to the concept of fractional differential equations. Finally we consider the use of delays in capturing stochastic effects. For each case we attempt to give a discussion of applicable methods and an indication of their advantages and disadvantages.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Intrinsic noise
- Stochastic simulation
- Discrete stochastic methods
- Continuous stochastic methods
- Anomalous diffusion
- Delay
1 Introduction
Heterogeneity is a key property of biological systems at all scales: from the molecular level to the population level [1,2,3]. Many systems have evolved ways of either minimising or exploiting this noise [4, 5]. In particular, some critical cellular systems have evolved to minimise or take advantage of noise [6, 7], for example, persistence in bacteria [8, 9]. We can classify heterogeneity as arising from three main sources: genetic (nature), environmental (nurture) and stochastic (chance). In this chapter we focus on the latter. We make the distinction between Environmental heterogeneity often called extrinsic noise and intrinsic noise, which arises from random thermal fluctuations [1, 9].
Intrinsic noise was perhaps first observed in a cell biology setting by Spudich and Koshland [10]. They noticed that individual bacteria from an isogenic population maintained different swimming patterns throughout their entire lives. Although they called this ‘non-genetic individuality’, they believed it arose from random fluctuations of low copy-number molecule or intrinsic noise. This affects the microscopic DNA, RNA and protein molecules in many ways, most notably via the Brownian motion of molecules and the randomness associated with their reactions. It ensures that the process of gene expression, one of the most fundamental processes of a cell, proceeds differently each time, even in the absence of the previous two sources of heterogeneity [9, 11].
Some biological systems have evolved to make use of intrinsic noise: a good example is persister-type bacteria, which can withstand antibiotic treatments even though they do not have genetic mutations for resistance [12]. These effects can arise through the phenomenon of stochastic switching, where cells randomly transition from one state to another [13, 14]. Another well-known example of exploiting stochasticity is the bacteriophage Lambda decision circuit [15]. Stochasticity also plays a crucial role in causing genetic mutations [16, 17]; these are essential for creating the heritable heterogeneity upon which natural selection can act, thus allowing evolution to occur [18].
On the other hand, intrinsic noise can interfere with the precise regulation of molecular numbers [19], and cells have to compensate for this by adopting mechanisms that enable robustness of certain key properties [4, 5], such as feedback loops [9, 11]. Furthermore, mutations may also have negative effects, as certain mutations in, for example, stem and somatic cells are thought to be the cause of cancer [20,21,22].
2 Temporal Modelling in Computational Cell Biology
There has been a long and successful history in computational cell biology of using rate kinetic ordinary differential equations to model chemical kinetics within a living cell. For instance, these techniques have been applied on the plasma membrane, in the cytosol and in the nucleus of eukaryotic cells to understand cell processes ranging from gene regulation to transport between cellular compartments. Modifications via delay differential equations were first considered as early as [23], in order to represent the fact that the complex regulatory processes of transcription and translation were not immediate but were in fact examples of delayed processes.
Thus in a purely temporal homogeneous setting and when there are large numbers of molecules present, chemical reactions are modelled by ordinary differential equations that are based on the Law of Mass Action and that estimate reaction rates on the basis of average values of the reactant density. Any set of m chemical reactions can be characterised by two sets of quantities: the stoichiometric vectors (update rules for each reaction) ν 1 , … , ν m and the propensity functions a 1(X(t)) , … , a m (X(t)). The propensity functions represent the relative probabilities of each of the m reactions occurring. These are formed by multiplying the rate constant and the product of the reactants on the left-hand side of each reaction. Here X(t) is the vector of concentrations at time t of the N species involved in the reactions. The ODE that describes this chemical system is
This formulation may have many different timescales (stiffness) but there are a wide variety of numerical methods that can deal effectively with such systems.
It was the pioneering work of Gillespie [24] and Kurtz [25] who challenged this deterministic view of cellular kinetics. They argued that when the cellular environment contained small to moderate numbers of proteins, then the Law of Mass Action is not an adequate description of the underlying chemical kinetics as it only describes the average behaviour. In this regard, the fundamental underlying principle is that of intrinsic noise. Intrinsic noise is associated with the inherent uncertainty in knowing when a reaction occurs and what that reaction is. The variance associated with this uncertainty increases as the number of proteins in the cellular environment becomes small. [24, 25] showed how to model intrinsic noise through the concept of nonlinear discrete Markov processes and Poisson processes, respectively. These two approaches both model the same processes and are now lumped together under the title the Stochastic Simulation Algorithm (SSA), although their formulations are different. The essential observation underlying the SSA is that the waiting time between reactions is exponentially distributed and that the most likely reaction to occur in this time interval is based on the relative sizes of the propensity functions.
More formally, the SSA is an exact procedure that describes the evolution of a discrete nonlinear Markov process. It accounts for the inherent stochasticity of the m reactions within a system and only assigns integer numbers of molecules to the state vector. At each step, the SSA samples two random numbers from the uniform distribution U[0,1] to evaluate an exponential waiting time, τ, for the next reaction to occur and an integer j between 1 and m that indicates which reaction occurs. The state vector is updated at the new time point by the addition of the j th stoichiometric vector to the previous value of the state vector, that is,
Several more efficient, but more complex, variants of the SSA have been developed [26,27,27]. However, despite these increases in computational speed, the SSA has an inherent limitation: it must simulate every single reaction, and in cases where there are many reactions or molecular populations become too large, it is computationally intensive.
In a slightly different vein, the SSA describes the evolution of a nonlinear discrete Markov process and as such this stochastic process has a probability density function whose solution is described by the Chemical Master Equation (CME). The CME is a discrete parabolic partial differential equation in which there is one equation for each configuration of the ‘state space’. When the state space is enumerated, the CME becomes a linear ODE and the probability density function takes the form
Here A is the state-space matrix. Thus the solution of the CME can be reduced to the computation of the evolution of the exponential of a matrix times an initial probability vector. As there is one equation for each possible configuration of the state space this can be very computationally challenging, although recently developed methods can cope with some of these computational costs [28,30,31,32,32] to make this a very feasible technique.
The main limiting feature of SSA is that the time step can become very small, especially if there are large numbers of molecules or widely varying rate constants. Thus tau-leap methods have been proposed in which the sampling of likely reactions is taken from either Poisson [33] or Binomial [34] distributions. In these approaches a much larger time step can be used at the loss of a relatively small amount of accuracy. The tau-leap method [33] allows steps that are much larger than the SSA by estimating the total number of occurrences of each type of reaction over that step. Thus if there are m reactions, we take m Poisson random number samples based on the sizes of the propensity functions evaluated at the beginning of the step. The algorithm can thus be written as
Note that now we have complete flexibility in choosing τ and it can be done adaptively in such a way as to control the local error at each step. This approach has the key advantage that individual reactions need not be simulated. However, the main drawback is a loss of accuracy compared to the SSA as a function of the step size. Several schemes have been devised to optimise the time step [35, 36], while some implementations combine the SSA with tau-leaping via a threshold on the number of molecules in the system at any given time. However, it is possible that molecular species that are depleted in any reaction can go negative if the time step is too large and schemes have been developed that allow the choice of larger time steps whilst avoiding negative populations [34, 36,37,38,39,40]. One important way in which this can be done is to sample from the Binomial distribution rather than the Poisson distribution [34]. Another approach is to consider the order of accuracy as a function of the step size. In this regard the (weak) order of accuracy of the tau-leap method can be shown to be one [41,43,43], meaning that error decreases proportionally to the time step. Higher-order methods allow larger time steps to achieve the same error, thus decreasing computational time [43,44,45,46,47].
Although it is not uncommon for chemical systems to be rather complicated, a difficult situation arises when a system has reactions that operate at very different timescales, for instance, slow and fast. Although standard tau-leap and higher-order tau-leap methods are able to simulate these systems, their time steps must be reduced, which can dramatically slow down simulation time when the separation of the scales is significant and the fast reactions occur frequently.
In such cases, there are two options: either to use special methods for these ‘stiff’ systems or to use multiscale methods. The former are often based on deterministic methods for stiff ordinary differential equation systems and allow the use of normal time steps by expanding the range of time steps where the method is stable, thus opening the door for stiff systems to be simulated in similar time periods as non-stiff ones [48,49,50]. Multiscale methods, on the other hand, partition the reactions into fast and slow types, and simulate the fast reactions using an approximate method and the slow reactions using an accurate stochastic method. The partitioning implies that the slow reactions are constant over the timescale of the fast reactions and that the fast reactions have relaxed to asymptotic values between each slow reaction. Thus, the two sets of reactions are simulated iteratively to take into account the coupling. This can also be generalised to three regimes: fast, medium and slow [51]. This approach allows for very significant reductions in computational time as the fast reactions, which would take up the most computational effort, can be simulated very quickly with continuous methods. However, it also introduces errors associated with coupling between two scales, as well as the possibility of errors from the simulation of the fast reactions. An interesting approach runs short bursts of a single SSA for the fast reactions, which is used to infer parameters for a differential equation approximation of the slow reactions [52, 53].
There is in fact an intermediate regime that can still capture the inherent stochastic effects but reduce the computational complexity associated with the SSA. This intermediate framework is called the Chemical Langevin Equation (CLE). It is described by an Itô stochastic differential equation (SDE) driven by a set of Wiener processes that describes the fluctuations in concentrations of the molecular species. Various numerical methods can then be applied to this equation—the simplest method being the Euler–Maruyama method [54].
The CLE attempts to preserve the correct dynamics for the first two moments of the SSA and takes the form
Here W(t) = (W 1(t), … , W N (t))T is a vector of N independent Wiener processes whose increments ΔW j = W j (t + h) − W j (t) are N(0, h) and where
Here h is the time discretisation step. Effective methods designed for the numerical solution of SDEs [54,55,56] can be used to simulate the chemical kinetics in this intermediate regime. [57] have shown how to construct the CLE so that it minimises the number of Wiener processes. Furthermore, adaptive multiscale methods have been developed that attempt to move back and forth between the deterministic and stochastic regimes as the numbers of molecules change [51].
These temporal approaches are applied under the hypothesis of homogeneous and well-mixed systems. It is well known, for example that diffusion on the cell membrane is not only highly anomalous but the diffusion rate of proteins on live cell membranes is also between one and two orders of magnitude slower than in reconstituted artificial membranes with the same composition [58]. Furthermore, diffusion is dependent on the dimensions of the medium so that diffusion on the highly disordered cell membrane is not a perfectly mixed process and therefore the assumptions underlying the classical theory of chemical kinetics fail, requiring either new approaches to modelling chemistry on a spatially crowded membrane [59] or methods based on detailed spatial simulations.
However, rather than abandoning temporal models entirely, it is possible to capture important spatial aspects and incorporate them into temporal models. This can be done in a number of ways. For example, compartmental models have been developed that couple together the plasma membrane, cytosol and nucleus—see, for example [60], in which an SSA implementation of Ras nanoclusters on the plasma membrane is coupled with an ODE model for the MAPK pathway in the cytosol. Diffusion and translocation can be captured through the use of distributed delays that can then be incorporated into mathematical frameworks through the use of delay differential equations or delay variants of the Stochastic Simulation Algorithm (see [61], for example). [62] have explored a number of spatial scenarios, run detailed spatial simulations to capture diffusion and translocation processes and then incorporated this information into purely temporal models through distributed delays—see Sect. 5 for more details. Another way in which spatial information can be captured and then incorporated into purely temporal models is the area of anomalous diffusion, where spatial crowding and molecular binding can affect chemical kinetics. In this setting the mean square deviation of a diffusing molecule is no longer linear but sublinear in time t and of the form
Here, α is called the anomalous diffusion parameter. If the value of α can be estimated, either experimentally or from detailed Monte Carlo simulations, then the SSA can be modified so that the waiting time between reactions is no longer exponentially distributed but has a heavy tail [59].
3 Spatial Models in Computational Cell Biology
One of the fundamental goals of integrative Computational Biology is to understand complex spatio-temporal processes within cells. However, such a task may become exceedingly difficult, due to the intrinsic multiscale nature of these processes. For example, in order to fully understand cell signal transduction, a careful description of membrane-bound receptor activation processes must be accounted for. However, the plasma membrane is a highly complex structure, compartmentalised on multiple length and timescales stemming from lipid–lipid, lipid–protein interactions with the cytoskeleton. As a result, detailed simulations accounting for all processes may be computationally prohibitively expensive, or simply infeasible.
Of the class of spatial methods, the approach with the lowest computational cost consists of solving reaction-diffusion partial differential equations, representing the concentration of a given molecular species in the system. However, this approach is only valid if and when: (1) all molecular species in the system have large molecular concentrations, and (2) noise is not amplified throughout the system. If at least one of these conditions fails to hold, we must rely on spatial stochastic simulation that can be discrete or continuous in form. In the discrete spatial setting either lattice or off-lattice particle based methods are appropriate.
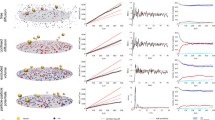
As a particular example of lattice based methods, we consider the plasma membrane. The plasma membrane is a complex and crowded environment that has many roles including signalling, cell–cell communication, cell feeding and excretion and protection of the interior of a cell. It is heterogeneous—the cytoskeletal structure just inside the plasma membrane can corral and compartmentalise membrane proteins. Chemically inert objects can form barriers to protein diffusion on the plasma membrane. Trying to capture such complexity using higher-level mathematical frameworks such as partial differential equations is extremely challenging, so instead a stochastic spatial model using Monte-Carlo simulation is appropriate. In such an approach the plasma membrane can be mapped to a two dimensional lattice, usually regular but not necessarily so. The size of each computational cell ‘voxel’ depends on the biological questions that are being addressed, but taking into account volume-exclusion effects, usually the voxel is such that at most one protein per voxel is allowed. A protein undergoes a random walk, so that at each time step a protein is selected at random, and a movement direction (north, south, east or west, in the case of a rectangular lattice arrangement) is randomly determined. The distance moved depends on the diffusion rates for each species. Chemical reactions can be simulated by checking the chemical reaction rules and then replacing that protein and/or creating a new protein at that location whenever a collision (volume exclusion event) occurs [63, 64]. Although only relatively small sections of the membrane on short timescales can be simulated with this approach due to the very slow computational performance, we note that diffusion can be considered as a unimolecular reaction. Thus if we order the voxel elements within the lattice into a vector, we can consider this method in the SSA framework and apply the same tools that have been developed in the purely temporal setting. This can allow us to make use of vectorisation to speed up the performance. Furthermore, there is a spatial CME associated with this approach [65] and again techniques used in the purely temporal case can be used, although at the cost of very significantly increased computational complexity.
In off-lattice methods, all particles in the system have explicit spatial coordinates at all times. At each time step, molecules are able to move, in a random walk fashion, to new positions. In many cases, reaction zones whose size depends on the particular diffusion rates are drawn around each particle. If one or more molecules happen to be inside such a zone, appropriate chemical reactions can take place with a certain probability; if a reaction is readily performed, the reactant particles are flagged, to avoid repetition of chemical events. Noticeably, in off-lattice methods, the domains and/or compartments can still be discretised, to aid the localisation of particles within the simulation domain. In this vein, [66] have considered how to combine tau-leaping and compartmentalisation in a spatial setting.
A less computationally intensive alternative, albeit still costly in many scenarios, is to consider molecular interactions in the mesoscopic realm. Here, the discretisation of the Reaction-Diffusion Master Equation (RDME) results in reactive neighbouring subvolumes within which several particles can coexist, with well-mixedness assumed in each subvolume. There are a number of algorithms extending discrete stochastic simulators to approximate solutions of the RDME by introducing diffusion steps as first-order reactions, with a reaction rate constant proportional to the diffusion coefficient. In [67, 68] the authors provide the specific outline for extending discrete stochastic simulators to the RDME regime, while the algorithms in [69, 70] provide clever extensions of the ‘next reaction method’ [26], commonly known as the ‘next subvolume method’. A review on the construction of such methods is given in [71].
The Next Subvolume Method [69, 70, 72, 73] is a generalisation of the SSA [24], where the simulation domain is divided into uniform separate subvolumes that are small enough to be considered homogeneous by diffusion over the timescale of the reaction. At each step, the state of the system is updated by performing an appropriate reaction within a single subvolume or by allowing a molecule to jump to a randomly selected neighbouring subvolume. Diffusion is then modelled as a unary reaction. In a two dimensional setting the rate is proportional to the diffusion coefficient divided by the length of a side of the subvolume. In this way, diffusion inside the algorithm becomes another possible event with a regular propensity function, and follows the same update procedure as any chemical reaction. The expected time for the next event in a subvolume is calculated in a similar way to the SSA algorithm, including the reaction and diffusion propensities of all molecules contained in that subvolume, at that particular time. However, times for subsequent events will only be recalculated for those subvolumes that were involved in the current time step, and they are subsequently re-ordered in an event queue. Similar to accelerated approaches to simulate exact trajectories from the CME, there exist methods to coarse-grain, and therefore accelerate, computations for the RDME [66]. Separately, [74] split the time integration of the RDME into a macroscopic diffusion (for species with large numbers of molecules) and a stochastic mesoscopic reaction/diffusion part (for species with small numbers) obtaining the mesoscopic diffusion coefficients from appropriate Finite Element discretisations.
In addressing these spatial issues, we are led to consider the role of anomalous diffusion. Anomalous diffusion refers to processes where the mean squared displacement (MSD) of a particle is no longer linear in time [75]. Anomalous diffusion can be viewed in two ways: as a mechanism to localise molecules and control signalling [76], or a macroscopic result of underlying microscopic events. From the experimental perspective, various techniques have been used to study such processes, including Single Particle Tracking [77], Fluorescence Recovery after Photo bleaching [78] and Fluorescence Correlation Spectroscopy [79]. However, the quantification of the degree and nature of the anomalous diffusion has shown to be more difficult than anticipated, due to experimental limitations [76, 80]. In spite of these discussions and recent technical developments, the nature of anomalous subdiffusion is still not well understood, either from experimental or simulation perspectives.
There are many reasons for these discrepancies including the fact that often only short tracks are recorded and the MSD relationship is not necessarily a robust metric for inferring complex spatial information. A more robust metric would be to construct a probability density function evolving in space and time, but this would require a very time-consuming experimental study. From the simulation perspective, a number of simulation studies have reported crowding-dependent values of α [64, 81]. But a prediction of percolation theory is that for immobile obstacles α depends only on the embedding space dimension and not on the obstacle density [82]. From theoretical perspectives, in the case of immobile objects, we can expect to observe Brownian diffusion for initial small time periods (when no obstacles have yet been encountered), an anomalous regime at intermediate times and Brownian diffusion in the asymptotic regime [83, 84]. It is these crossovers that can create confusion when trying to interpret diffusive characteristics.
It remains to be seen whether anomalous diffusion scenarios can only be captured by explicit spatial models, or whether equivalent dynamics could be obtained using off-lattice simulations with single molecules devoid of explicit obstacles, thus capturing essential dynamics while significantly reducing computational time. Other possibilities would include deriving an SSA and its associated CME that would work in an anomalous diffusion temporal setting only. We could then attempt to replicate the deterministic and stochastic regimes in the time anomalous setting. This would lead to time fractional or space fractional representations. For example, molecular concentration dynamics C(x, t) in a typical subdiffusive setting could be represented by the time-fractional differential equation
where x and t are the space and time variables, respectively, and the anomalous exponent α is the fractional order of the time derivative operator. Here \( {D}_t^{1-\alpha }\ g(t) \) is the fractional Riemann–Liouville derivative operator that reduces to the identity operator when α = 1 while K α is the fractional equivalent to the classical diffusion coefficient and has dimension [K α ] = [l]2[t]−α.
4 Modelling and Simulating Stochastic ion Channel Dynamics
Ion channels are multiconformational proteins that form a pore in the membrane of excitable cells. They open and close due to conformational changes in the proteins as a result of variations in membrane potential, and thus regulate the movement of ions across the lipid bilayer [85]. The dynamics of these proteins are fundamental to the generation of an action potential (AP) in excitable cells [86]. Single-channel recordings have demonstrated that the conformational changes the protein undergoes as it opens and closes occur at random [87]. This internal stochasticity causes fluctuations in individual ionic conductances, [86], and has important effects on the electrical dynamics of the cell [88,89,90,91].
In neuronal cells, stochastic ion channel behaviour can modify a number of electrical properties of the cell including the firing threshold [92] and spike timings [93]. In cardiac myocytes this intrinsic randomness leads to variability in the duration of successive APs [89, 90], termed beat-to-beat variability, which is thought to be an indicator of arrhythmias [94]. It can even cause alterations to the AP morphology under pathological conditions, resulting early-after-depolarisations (EADs) [89].
While a discrete-state modelling and simulation approach is often seen as the ‘gold-standard’, it becomes increasingly computationally costly as the number of channels increases beyond a few hundred [95]. This has led to the increasing popularity of using stochastic differential equations to describe ion channel dynamics [89, 96, 97]. Fox and Lu [98] were the first to take such an approach, which they applied to the Hodgkin–Huxley model, and their method has since been extensively used to model neuronal cells [96, 97], cardiac myocytes [89, 90] and pancreatic beta cells [91]. Their approach reduces the dynamics of the whole channel to the collective dynamics of a series of gating variables that can be either open or closed. The proportion of each gating variable in the open state is described by a Stochastic Differential Equation (SDE) and the proportion of open channels is given by the product of the number of open gates.
However, a number of studies have demonstrated discrepancies between the SDE and discrete-state Markov chain model [95, 99, 100] [101]. Goldwyn et al. [101] demonstrate that constructing the SDE model in terms of the kinetic dynamics of the channel, rather than the individual gating variables, preserves the stochastic behaviour of the discrete-state Markov chain model, see the recent review [104] for a further discussion.
In order to illustrate these ideas we give more details on the form of the Chemical Langevin equation for ion channel dynamics. We first consider an ion channel transitioning just between the closed and open states. Let the proportion of channels in the open state be y, let N be the total number of ion channels and W be a Wiener process. Then the form of the CLE is
However, the complicating factor is that the parameters a and b are themselves functions of the trans-membrane voltage. In the more general setting when there are a number of transitions between the various ion channel states then the formulation is given by
For the moment take k = 0. Here the columns of v are state changes resulting from a transition. D is a diagonal matrix whose entries are chance of transition occurring (namely the propensity functions), b p(y(t)) are columns of matrix B, where BBT = vD(y(t))v T. In the case of, say p, unimolecular transitions, which is the setting for ion channels, then this simplifies to B = ES, where E is a p × d/2 matrix and S is a d/2 × d/2 diagonal matrix. In particular, E is a matrix with 1s on the diagonal, −1s in certain lower triangular positions and 0s elsewhere. Here S is a diagonal matrix with square roots of linear combinations of certain pairs of components of y [57].
For example, in the case of Sodium and Potassium ion channels, that form key elements of the Hodgkin-Huxley ion channel model [103], the transitions are given~by
In the case of Sodium, for example, then E and S are given by
In passing, we note that due to the structure of the ion channels there is an explicit formulation to the underlying probability density function in terms of multinomials [104] but as the transition rates are nonlinear functions of the voltage this does not really help in simulating the ion channel dynamics.
Although this approach gives the correct dynamics, the solution to the SDE must remain non-negative for the path to have any biological relevance. Yet it has been shown that the solution can become negative [105]. Furthermore, since the noise term in the SDE model involves the square root of some function of the state variable, this can result in numerical solutions becoming imaginary [105]. Alterations to the numerical scheme can be made to force the solution to remain positive, for example, the Wiener increment can be continually resampled [89]. However, such alterations can bias the results [105,106,107]. Another approach is to replace the variable in the noise term with its equilibrium value, [96, 101] so that the square root term is independent of the state of the system. However, such an approach can still result in the proportion of channels becoming negative [101]. In [105] a hybrid simulation method for the Hodgkin–Huxley model is developed that attempts to improve the computational efficiency of the discrete-state Markov chain model whilst ensuring individual simulations remain non-negative by switching between regimes.

Example of the reflection process on a three dimensional bounded domain
We argue that the boundary conditions are not naturally incorporated into the standard CLE and that they can do so by the use of reflected SDEs [108] in which k t in the general formulation is a reflected process that only comes into play at the boundaries of the domain. The basic idea is to evaluate the non-reflected process at the next time step using the Euler–Maruyama method. If the simulation is in the desired region, set Y at the next time step to be this value. Otherwise orthogonally project onto the boundary of the domain D. We assume that the process will reflect y(t) into the interior of the domain in the direction of the inward pointing unit normal. It can be shown that this approach will converge with strong order ½-h for all h > 0. This can be visualised for the three state models—see figure. Thus in our general formulation we can interpret k t as a reflecting process and this determines the behaviour at the boundary.
Finally, very recently Schnoerr et al. [109] show in a very nice paper that, by extending the domain of the CLE to the Complex space, the CLE’s accuracy for unimolecular systems is restored. This is at the cost of having to perform simulations in complex arithmetic and taking care with the use of pre-defined functions.
5 The Role of Delays in Modelling Biochemical Reaction Systems and Model Reduction
The origins of delay differential equations (DDEs) date, most likely, back to the second half of the eighteenth century. According to Schmidt [110], some of the early work on DDEs (and, more generally, the so-called functional differential equations) originates from famous mathematicians such as Laplace, Poisson and Lacroix (see references in [110]). Two of the earliest examples of DDEs from the early twentieth century found in the mathematical medicine and mathematical biology literature are by Lotka, who studied a model of malaria epidemics with incubation delays (in particular in [111]), and by Volterra, who investigated delayed predator–prey models [112]. Ever since, DDEs have become an integral part of mathematical modelling of biological, biomedical and physiological processes. Examples of delay models can be found in areas such as population dynamics and epidemiology [113, 114], gene regulation [115], cell signalling [116], viral dynamics [117], tumour growth [118], drug therapy [118, 119], immune response [120] and respiratory systems [121].
The use of delays and systems’ histories was driven by the desire for more realistic and, consequently, more accurate mathematical models. Indeed, introducing delays is many times essential for reconciling models with observations and experimental data. Moreover, delays provide a way for a more intuitive modelling approach, i.e. phenomenological rather than mechanistic. In this case, complex processes are lumped while underlying mechanisms and inherent intermediate steps are not explicitly accounted for. Yet, the time that such processes require is included in the form of a constant delay or delay distribution.
With growing interest in the stochastic dynamics of chemical reaction networks, delays have also been introduced into stochastic simulation algorithms (SSAs). Several so-called delay SSAs (DSSAs) have been proposed [61, 122,123,124,125], and the Delay Chemical Master Equation (DCME) was introduced as a generalisation to the Chemical Master Equation for reactions with associated constant delays [61, 126] or delay distributions [126]. However, contrary to CMEs, for which either closed solutions have been presented or a large number of numerical and computational strategies have been suggested [104, 127,128,129,130,131,132], DCMEs have been largely disregarded. This is because DCMEs do no longer represent Markovian processes, since transitions between states depend on the current state and the process history. This leads to terms in a DCME that, for a delay reaction with stoichiometric vector ν and constant delay τ, look like
where the sum is taken over all previous states x i prior to the current state x. The joint probabilities are usually unknown and these terms can only be simplified if (a) the coupling of the system states at times t and t − τ is weak, in which case we obtain a reasonably good approximation; or (b) the triggering of the delayed reaction is fully independent of the occurrences of other reactions and of the state x i at the time of triggering. The latter implies in particular that none of the reactions in the system, including the delayed reaction, change the number of reactants of the delayed reaction, nor its kinetic function [126]. In this case, the propensity function of the delayed reaction is constant, i.e. a(x i ) = c for all x i , and the above sum simplifies to
If the delay is given in the form of a distribution instead of a constant delay, then its cumulative density function appears as just another factor. The DCME remains a homogeneous system of linear first-order ODEs, except that it now includes a time-dependent coefficient, and can then be solved numerically using the available tools for CMEs.
Moreover, the analytic solution for CMEs of monomolecular reaction systems was shown to be the convolution of multinomial and product Poisson distributions [104]. For a simple delayed, unidirectional reaction scheme, a multinomial distribution was recently derived as its solution using purely probabilistic arguments [126]. This suggests that such a distribution may also be the solution to a more general class of monomolecular, delayed reaction systems.
As already mentioned above, delays can also be used for model reduction. A number of reduction techniques have been proposed in the past, including the classical equilibrium approximation by Michaelis and Menten, the quasi-steady state approximation (QSSA) by Briggs and Haldane, several variations of the QSSA [133, 134], methods based on the linear noise approximation [135, 136], and the finite-state projection method [130]. However, they all approximate the true solution and/or make certain assumptions, for instance, a timescale separation, which, if not met, can lead to inaccurate results. The method presented in [137, 138] uses delayed reactions between species of interest that replace a number of intermediate species and reactions between these. The individual delays are obtained as first-passage time distributions in analytic form. These can then be placed into DSSA implementations for generating sample trajectories of the abridged system’s dynamics. This method is fully accurate for all bidirectional, unimolecular reaction chains, including degradations, synthesis and bypass reactions, and allows for large computational savings. This holds true in particular if, over the same simulation time span, the number of reactions in the unabridged system is considerably larger than the number of DSSA steps in the abridged system.
Lastly, it has become evident that spatial aspects play a crucial role in biological and biomedical processes. Even in relatively simple biochemical systems, the observed behaviour can vary considerably from the often assumed, well-mixed scenario, where spatial dependencies, geometries and structures are not taken into account. Thus, it is important to consider these spatial aspects in modelling approaches, both for our understanding and for accurate predictions of such processes. While detailed spatial models are more realistic they are also much more computationally demanding—if not prohibitively expensive. Alternatively, we can try to incorporate the effects of such spatial features into temporal models, without any explicit spatial representation. As has been shown recently, delay distributions may provide an appropriate tool [62]. The proposed methodology is similar to the model reduction with delays described above: the first step consists of obtaining proper delay distributions. These can stem from diffusion profiles and can be directly obtained from particle simulations, in vitro experiments or corresponding PDE solutions. Such tailored distributions are then used along with their associated reactions in a modified version of the DSSA. When applied to a variety of simple scenarios of molecular translocation processes large computational savings were achieved.
6 Conclusions
There is still considerable ongoing research to refine both the SSA and approximate methods. However, there are four areas where significant improvements can be made easily. Firstly, graphical processing units on graphics cards are now very powerful for some applications. Parallelised stochastic methods allow us to harness this power in order to run many thousands of simulations simultaneously. Secondly, multiscale problems are common in many real applications and they are often large in nature. The use of adaptive multiscale approaches where processes on many different scales are integrated into one model could, for example, play an important role in personalised medicine. Thirdly, we can attempt to reduce the number of simulations needed in order to gain a certain accuracy using ideas from multi-level theory developed by Mike Giles—this is a form of variance reduction. Fourthly, the limitation of non-spatial methods is that they can only be accurately applied to macroscopically homogeneous systems, but this assumption does not hold in many (or even most) cases of interest. Therefore it is important to develop methods that can take appropriate account of such heterogeneous environments. Finally, we note that one area that we have not covered in this chapter is parameter inference and model selection. This area involves using statistical methods to find model parameters from experimental data, and to discriminate between those models best fit this data [139,140,141]. This is typically not an easy task, as the data may be noisy, missing or sparse; Bayesian approaches offer a way of addressing these issues [142].
References
H.H. McAdams, A. Arkin, It's a noisy business! Genetic regulation at the nanomolar scale. Trends Genet. 15, 65–69 (1999)
S. Huang, Non-genetic heterogeneity of cells in development: More than just noise. Development 136, 3853–3862 (2009)
S.V. Avery, Microbial cell individuality and the underlying sources of heterogeneity. Nat. Rev. Microbiol. 4, 577–587 (2006)
N. Barkai, B. Shilo, Variability and robustness in biomolecular systems. Mol. Cell 28, 755–760 (2007)
C.V. Rao, D.M. Wolf, A.P. Arkin, Control, exploitation and tolerance of intracellular noise. Nature 420, 231–237 (2002)
H.B. Fraser, A.E. Hirsh, G. Giaever, J. Kumm, M.B. Eisen, Noise minimization in eukaryotic gene expression. PLoS Biol. 2, 834–838 (2004)
B. Lehner, Selection to minimise noise in living systems and its implications for the evolution of gene expression. Mol. Syst. Biol. 4, 170 (2008)
D. Fraser, M. Kaern, A chance at survival: Gene expression noise and phenotypic diversification strategies. Mol. Microbiol. 71, 1333–1340 (2009)
M. Kaern, T. Elston, W. Blake, J. Collins, Stochasticity in gene expression: From theories to phenotypes. Nat. Rev. Genet. 6, 451–464 (2005)
J.L. Spudich, D.E. Koshland Jr., Non-genetic individuality: Chance in the single cell. Nature 262, 467–471 (1976)
N. Maheshri, E.K. O'Shea, Living with noisy genes: How cells function reliably with inherent variability in gene expression. Annu Rev Bioph Biom 36, 413–434 (2007)
N.Q. Balaban, J. Merrin, R. Chait, L. Kowalik, S. Leibler, Bacterial persistence as a phenotypic switch. Science 305, 1622–1625 (2004)
M. Acar, J.T. Mettetal, A. van Oudenaarden, Stochastic switching as a survival strategy in fluctuating environments. Nat. Genet. 40, 471–475 (2008)
P.J. Choi, L. Cai, K. Frieda, X.S. Xie, A stochastic single-molecule event triggers phenotype switching of a bacterial cell. Science 322, 442–446 (2008)
A. Arkin, J. Ross, H.H. McAdams, Stochastic kinetic analysis of developmental pathway bifurcation in phage λ-infected Escherichia coli cells. Genetics 149, 1633–1648 (1998)
E.C. Friedberg, G.C. Walker, W. Siede, R.D. Wood, DNA Repair and Mutagenesis, 2nd edn. (ASM Press, Washington, DC, 2006)
J.M. Pennington, S.M. Rosenberg, Spontaneous DNA breakage in single living Escherichia coli cells. Nat. Genet. 39, 797–802 (2007)
T.R. Gregory, Understanding natural selection: Essential concepts and common misconceptions. Evo Edu Outreach 2, 156–175 (2009)
N. Rosenfeld, J.W. Young, U. Alon, P.S. Swain, M.B. Elowitz, Gene regulation at the single-cell level. Science 307, 1962–1965 (2005)
L.B. Alexandrov, S. Nik-Zainal, D.C. Wedge, S.A.J.R. Aparicio, S. Behjati, et al., Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
M.R. Stratton, P.J. Campbell, P.A. Futreal, The cancer genome. Nature 458, 719–724 (2009)
I.A. Rodriguez-Brenes, N.L. Komarova, D. Wodarz, Evolutionary dynamics of feedback escape and the development of stem-cell driven cancers. Proc. Natl. Acad. Sci. U. S. A. 108, 18983–18988 (2011)
B.C. Goodwin, Oscillatory behavior in enzymatic control processes. Adv. Enzym. Regul. 3, 425–438 (1965)
D.T. Gillespie, Exact stochastic simulation of coupled chemical reactions, J. Phys. Chemistry 81(25), 2340–2361 (1977)
T.G. Kurtz, The relationship between stochastic and deterministic models for chemical reactions. J. Chem. Phys. 57(7), 2976–2978 (1972)
M.A. Gibson, J. Bruck, Efficient exact stochastic simulation of chemical systems with many species and many channels. J. Phys. Chem. A 104(9), 1876–1889 (2000)
Y. Cao, H. Li, L.R. Petzold, Efficient formulation of the stochastic simulation algorithm for chemically reacting systems. J. Chem. Phys. 121, 4059–4067 (2004)
S. MacNamara, A.M. Bersani, K. Burrage, R.B. Sidje, Stochastic chemical kinetics and the total quasi-steady-state assumption: Application to the stochastic simulation algorithm and chemical master equation. J. Chem. Phys. 129(9), 095105 (2008)
S. MacNamara, K. Burrage, R.B. Sidje, Multiscale modeling of chemical kinetics via the master equation. SIAM J Multiscale Modelling and Simulation Multiscale 6(4), 1146–1168 (2008)
S. Peleš, B. Munsky, M. Khammash, Reduction and solution of the chemical master equation using time scale separation and finite state projection. J. Chem. Phys. 125, 204104-1–20410413 (2006)
T. Jahnke, S. Galan,Solving chemical master equations by an adaptive wavelet method, in Numerical Analysis and Applied Mathematics: International Conference on Numerical Analysis and Applied Mathematics, 16–20 Sept, ed. by T. E. Simos, G. Psihoyios, C. Tsitouras, vol. 1048 (AIP Conference Proceedings, Psalidi, Kos, Greece, 2008), pp. 290–293
S. Engblom, Galerkin spectral method applied to the chemical master equation. Commun Comput Phys v5(i5), 871–896 (2009)
D.T. Gillespie, Approximate accelerated stochastic simulation of chemically reacting systems. J. Chem. Phys. 115(4), 2001 (1716—1733)
T. Tian, K. Burrage, Binomial leap methods for simulation stochastic chemical kinetics. J. Chem. Phys. 121, 10356–10364 (2004)
D.T. Gillespie, L.R. Petzold, Improved leap-size selection for accelerated stochastic simulation. J. Chem. Phys. 119, 8229–8234 (2003)
Y. Cao, D.T. Gillespie, L.R. Petzold, Efficient step size selection for the tau-leaping simulation method. J. Chem. Phys. 124, 044109 (2006)
A. Chatterjee, D.G. Vlachos, M.A. Katsoulakis, Binomial distribution based tau-leap accelerated stochastic simulation. J. Chem. Phys. 122, 024112 (2005)
X. Peng, W. Zhou, Y. Wang, Efficient binomial leap method for simulating chemical kinetics. J. Chem. Phys. 126, 224109 (2007)
M.F. Pettigrew, H. Resat, Multinomial tau-leaping method for stochastic kinetic simulations. J. Chem. Phys. 126, 084101 (2007)
C.A. Yates, K. Burrage, Look before you leap: A confidence-based method for selecting species criticality while avoiding negative populations in tau-leaping. J. Chem. Phys. 134, 084109 (2011)
M. Rathinam, L.R. Petzold, Y. Cao, D.T. Gillespie, Consistency and stability of tau-leaping schemes for chemical reaction systems. Multiscale Model Sim 4, 867–895 (2005)
T. Li, Analysis of explicit tau-leaping schemes for simulating chemically reacting systems. Multiscale Model Sim 6, 417–436 (2007)
Y. Hu, T. Li, B. Min, A weak second order tau-leaping method for chemical kinetic systems. J. Chem. Phys. 135, 024113 (2011)
Y. Hu, T. Li, Highly accurate tau-leaping methods with random corrections. J. Chem. Phys. 130, 124109 (2009)
D.F. Anderson, M. Koyama, Weak error analysis of numerical methods for stochastic models of population processes. Multiscale Model Sim 10, 1493–1524 (2012)
T. Székely Jr., K. Burrage, R. Erban, K.C. Zygalakis, A higher-order numerical framework for stochastic simulation of chemical reaction systems. BMC Syst. Biol. 6, 85 (2012)
Z. Xu, X. Cai, Unbiased tau-leap methods for stochastic simulation of chemically reacting systems. J. Chem. Phys. 128, 154112 (2008)
M. Rathinam, L.R. Petzold, Y. Cao, D.T. Gillespie, Stiffness in stochastic chemically reacting systems: The implicit tau-leaping method. J. Chem. Phys. 119, 12784–12794 (2003)
Y. Cao, D.T. Gillespie, L.R. Petzold, The adaptive explicit-implicit tau-leaping method with automatic tau selection. J. Chem. Phys. 126, 224101 (2007)
P. Rué, J. Villa-Freixà, K. Burrage, Simulation methods with extended stability for stiff biochemical kinetics. BMC Syst. Biol. 4, 110–123 (2010)
K. Burrage, T. Tian, P. Burrage, A multi-scaled approach for simulating chemical reaction systems. Prog Biophys Mol Bio 85, 217–234 (2004)
R. Erban, I.G. Kevrekidis, D. Adalsteinsson, T.C. Elston, Gene regulatory networks: A coarse-grained, equation-free approach to multiscale computation. J. Chem. Phys. 124, 084106 (2006)
S.L. Cotter, K.C. Zygalakis, I.G. Kevrekidis, R. Erban, A constrained approach to multiscale stochastic simulation of chemically reacting systems. J. Chem. Phys. 135, 094102 (2011)
P.E. Kloeden, E. Platen, Numerical Solution of Stochastic Differential Equations (Springer-Verlag, Berlin, 1992)
P.M. Burrage, K. Burrage, A variable stepsize implementation for stochastic differential equations. SIAM J. Sci. Comput. 24(3), 848–864 (2002)
P.M. Burrage, R. Herdiana, K. Burrage, Adaptive stepsize based on control theory for SDEs. J Comp and App Math 170, 317–336 (2004)
B. Mélykúti, K. Burrage, K.C. Zygalakis, Fast stochastic simulation of biochemical reaction systems by alternative formulations of the chemical Langevin equation, J. Chem. Phys. 132, 1 (2010)
K. Murase, T. Fujiwara, T.Y. Umemura, Ultrafine membrane compartments for molecular diffusion as revealed by single molecule techniques. Biophys. J. 86, 4075–4093 (2004)
D.V. Nicolau, K. Burrage, Stochastic simulation of chemical reactions in spatially complex media. Computers & Mathematics with Applications 55(5), 1007–1018 (2008)
T. Tian, A. Harding, E. Westbury, J. Hancock, Plasma membrane nano-switches generate robust high-fidelity Ras signal transduction. Nat. Cell Biol. 9, 905–914 (2007)
M. Barrio, K. Burrage, A. Leier, T. Tian, Oscillatory regulation of Hes1: Discrete stochastic delay modelling and simulation. PLoS Comp Bio 2(9), e117 (2006). doi:10.1371/journal.pcbi.0020117
T.T. Marquez-Lago, A. Leier, K. Burrage, Probability distributed time delays: Integrating spatial effects into temporal models. BMC Syst. Biol. 4, 19 (2010)
D.V. Nicolau Jr., K. Burrage, R.G. Parton, et al., Identifying optimal lipid raft characteristics required to promote nanoscale protein-protein interactions on the plasma membrane. Mol. Cell. Biol. 26(1), 313–323 (2006)
D.V. Nicolau Jr., J.F. Hancock, K. Burrage, Sources of anomalous diffusion on cell membranes: A Monte Carlo study. Biophys. J. 92, 1975–1987 (2007)
B. Drawert, M.J. Lawson, L. Petzold, M. Khammash, The diffusive finite state projection algorithm for efficient simulation of the stochastic reaction-diffusion master equation. J. Chem. Phys. 132(074101), 2010 (2010). doi:10.1063/1.3310809
T.T. Marquez-Lago, K. Burrage, Binomial tau-leap spatial stochastic simulation algorithm for applications in chemical kinetics. J. Chem. Phys. 127, 104101 (2007)
A.B. Stundzia, C.J. Lumsden, Stochastic simulation of coupled reaction-diffusion processes. J Comp Phys 127, 196–207 (1996)
F. Baras, M. Malek Mansour, Reaction-diffusion master equation: A comparison with microscopic simulations. Phys. Rev. E 54(6), 6139–6148 (1996)
J. Elf, A. Doncic, M. Ehrenberg, Mesoscopic reaction-diffusion in intracellular signaling. Proc. SPIE 5110, 114–125 (2003)
M. Ander, P. Beltrao, B. Di Ventura, et al., SmartCell, a framework to simulate cellular processes that combines stochastic approximation with diffusion and localisation: Analysis of simple networks. Syst. Biol. 1, 129–138 (2004)
R. Erban, S.J. Chapman, P.K. Maini, A practical guide to stochastic simulations of reaction–diffusion processes, arXiv:0704.1908 (2007
J. Elf, M. Ehrenberg, Spontaneous separation of bi-stable biochemical systems into spatial domains of opposite phases. Syst. Biol. 1, 230–236 (2004)
J. Hattne, D. Fange, J. Elf, Stochastic reaction-diffusion simulation with MesoRD. Bioinformatics 21, 2923–2924 (2005)
S. Engblom, L. Ferm, A. Hellander, P. Loetstedt, Simulation of stochastic reaction-diffusion processes on unstructured meshes. SIAM J. Sci. Comput. 31, 1774–1797 (2009)
R. Metzler, J. Klafter, The random walk’s guide to anomalous diffusion: A fractional dynamics approach. Phys Reports 339, 1–77 (2000)
D.S. Martin, M.B. Forstner, J.A. Kas, Apparent subdiffusion inherent to single particle tracking. Biophys. J. 83(4), 2109–2117 (2002)
P.R. Smith et al., Anomalous diffusion of major histocompatibility complex class I molecules on HeLa cells determined by single particle tracking. Biophys. J. 76(6), 3331–3344 (1999)
T.M. Jovin, W.L. Vaz, Rotational and translational diffusion in membranes measured by fluorescence and phosphorescence methods. Methods Enzymol. 172, 471–513 (1989)
M. Weiss, H. Hashimoto, T. Nilsson, Anomalous protein diffusion in living cells as seen by fluorescence correlation spectroscopy. Biophys. J. 84(6), 4043–4052 (2003)
B. Leitinger, N. Hogg, The involvement of lipid rafts in the regulation of integrin function. J. Cell Sci. 115(Pt 5), 963–972 (2002)
H. Berry, Monte Carlo simulations of enzyme reactions in two dimensions: Fractal kinetics and spatial segregation. Biophys. J. 83(4), 1891–1901 (2002)
R.A. Kerr et al., Fast Monte Carlo simulation methods for biological reaction-diffusion Systems in Solution and on surfaces. SIAM J. Sci. Comput. 30(6), 3126 (2008)
R. Hilfer, L. Anton, Fractional master equations and fractal time random walks. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 51(2), R848–R851 (1995)
D. Fulger, E. Scalas, G. Germano, Monte Carlo simulation of uncoupled continuous-time random walks yielding a stochastic solution of the space-time fractional diffusion equation. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 77(2 Pt 1), 021122 (2008)
F. Bezanilla, The voltage sensor in voltage-dependent ion channels. Physiol. Rev. 80, 555 (2000)
B. Hille, Ionic Channels of Excitable Membranes, 3rd edn. (Sinauer Associates, Sunderland, MA, 1991). isbn:0878933239
B. Sakmann, E. Neher, Single-Channel Recording (Plenum, New York, 1995)
J.A. White, J.T. Rubinstein, A.R. Kay, Channel noise in neurons. Trends Neurosci. 23, 131 (2000.) ISSN 0166-2236
E. Pueyo, A. Corrias, L. Virag, N. Jost, T. Szel, A. Varro, N. Szentandrassy, P.P. Nanasi, K. Burrage, B. Rodrıguez, A multiscale investigation of repolarization variability and its role in cardiac arrhythmogenesis. Biophys. J. 12, 2892 (2011)
M. Lemay, E. de Lange, J.P. Kucera, Effects of stochastic channel gating and distribution on the cardiac action potential. J. Theor. Biol. 281, 84 (2011.) ISSN 1095-8541
G. De Vries, A. Sherman, Channel sharing in pancreatic beta-cells revisited: Enhancement of emergent bursting by noise. J. Theor. Biol. 207, 513 (2000)
J.R. Clay, L.J. DeFelice, Relationship between membrane excitability and single channel open-close kinetics. Biophys. J. 42, 151 (1983). doi:10.1016/S0006-3495(83)84381-1
E. Schneidman, B. Freedman, I. Segev, Ion channel stochasticity may be critical in determining the reliability and precision of spike timing. Neural Comput. 10, 1679 (1998.) ISSN 0899-7667, URL http://neco.mitpress.org/cgi/content/abstract/10/7/1679
P. Oosterhoff, A. Oros, M.A. Vos, Anadolu kardiyoloji dergisi. Anatolian journal of cardiology 7(Suppl 1), 73 (2007.) ISSN 1302-8723, URL http://view.ncbi.nlm.nih.gov/pubmed/17584687
H. Mino, J.T. Rubinstein, J.A. White, Comparison of algorithms for the simulation of action potentials with stochastic sodium channels. Ann. Biomed. Eng. 30, 578 (2002.) ISSN 0090-6964
R. Fox, Stochastic versions of the Hodgkin-Huxley equations. Biophys. J. 72, 2068 (1997.) ISSN 00063495
X.J. Sun, J.Z. Lei, M. Perc, Q.S. Lu, S.J. Lv, Effects of channel noise on firing coherence of small-world Hodgkin-Huxley neuronal networks. The European Physical Journal B - Condensed Matter and Complex Systems 79, 61 (2011). doi:10.1140/epjb/e2010-10031-3
R.F. Fox, Y.N. Lu, Emergent collective behavior in large numbers of globally coupled independently stochastic ion channels. Phys. Rev. E 49, 3421 (1994)
I.C. Bruce, Evaluation of stochastic differential equation approximation of ion channel gating models. Ann. Biomed. Eng. 37, 824 (2009.) ISSN 1521-6047
B. Sengupta, S.B. Laughlin, J.E. Niven, Comparison of Langevin and Markov channel noise models for neuronal signal generation. Phys. Rev. E 81, 011918 (2010.), ISSN 1550-2376)
J.H. Goldwyn, N.S. Imennov, M. Famulare, E. Shea-Brown, Stochastic differential equation models for ion channel noise in Hodgkin-Huxley neurons. Phys. Rev. E 83, 041908 (2011)
A.L. Hodgkin, A.F. Huxley, A quantitative description of membrane current and its application to conduction and excitation in nerve. J. Physiol. 117, 500 (1952.) ISSN 0092-8240
T. Jahnke, W. Huisinga, Solving the chemical master equation for monomolecular reaction systems analytically. J. Math. Biol. 54, 1–26 (2007.) ISSN 0303-6812
J.H. Goldwyn, E. Shea-Brown, The what and where of adding channel noise to the Hodgkin-Huxley equations. PLoS Comput. Biol. 7, e1002247+ (2011). doi:10.1371/journal.pcbi.1002247
C.E. Dangerfield, D. Kay, K. Burrage, Stochastic models and simulation of Ion Channel dynamics. Procedia Computer Science 1, 1581 (2010.) ISSN 18770509
C.E. Dangerfield, D. Kay, S. MacNamara, K. Burrage, A boundary preserving numerical algorithm for the Wright-fisher model with mutation. BIT Numer. Math. 52, 283 (2011)
R. Lord, R. Koekkoek, D.V. Dijk, A comparison of biased simulation schemes for stochastic volatility models. Quantitative Finance 10, 177 (2010)
C.E. Dangerfield, D. Kay, K. Burrage, Modelling ion channel dynamics through reflected stochastic differential, equations. Phys. Rev. E 85, 051907 (2012)
D. Schnoerr, G. Sanguinetti, R. Grima, The complex chemical Langevin equation. J.Chem.Phys. 141(2), 024103 (2014)
E. Schmidt, Über eine Klasse linearer funktionaler Differentialgleichungen. Math. Ann. 70(4), 499–524 (1911)
F.R. Sharpe, A.J. Lotka, Contribution to the analysis of malaria epidemiology. IV. Incubation lag. Am. J. Epidemiology. 3(supp1), 96–112 (1923)
V. Volterra, Variazioni e fluttuazioni del numero d’individui in specie animali conviventi. Memorie del R. Comitato Talassografico Italiano 43, 1–142 (1927)
Y. Kuang, Delay Differential Equations: With Applications in Population Dynamics (Academic Press, Boston, MA, 1993)
J. Arino, P. van den Driessche, Time delays in epidemic models: Modeling and numerical considerations, in Delay Differential Equations and Applications, ed. by O. Arino et al. (Springer, New York, 2006), pp. 539–578
J. Lewis, Autoinhibition with transcriptional delay: A simple mechanism for the zebrafish somitogenesis oscillator. Curr. Biol. 13(16), 1398–1408 (2003)
J. Srividhya, M.S. Gopinathan, A simple time delay model for eukaryotic cell cycle. J. Theor. Biol. 241(3), 617–627 (2006)
S. Yi, P.W. Nelson, A.G. Ulsoy, Time-Delay Systems: Analysis and Control Using the Lambert W Function (New Jersey World Scientific, 2010)
M. Villasana, A. Radunskaya, A delay differential equation model for tumor growth. J. Math. Biol. 47(3), 270–294 (2003)
M.V. Barbarossa, C. Kuttler, J. Zinsl, Delay equations modeling the effects of phase-specific drugs and immunotherapy on proliferating tumor cells. Math. Biosc. Eng 9(2), 241 (2012)
K. Cooke, Y. Kuang, B. Li, Analyses of an antiviral immune response model with time delays. Canad. Appl. Math. Quart. 6(4), 321–354 (1998)
C.-S. Kim, J.M. Ansermino, J.-O. Hahn, A comparative data-based Modeling study on respiratory CO2 gas exchange during mechanical ventilation. Front. Bioeng. Biotechnol. 4, 8 (2016). doi:10.3389/fbioe.2016.00008
D. Bratsun, D. Volfson, J. Hasty, L. Tsimring, Delay-induced stochastic oscillations in gene regulation. PNAS 102(41), 14593–14598 (2005). doi:10.1073/pnas.0503858102
M.R. Roussel, R. Zhu, Validation of an algorithm for delay stochastic simulation of transcription and translation in prokaryotic gene expression. Phys. Biol. 3(4), 274–284 (2006)
X. Cai, Exact stochastic simulation of coupled chemical reactions with delays. J. Chem. Phys. 126(12), 124108 (2007). doi:10.1063/1.2710253
D.F. Anderson, A modified next reaction method for simulating chemical systems with time dependent propensities and delays. J. Chem. Phys. 127, 214107 (2007)
A. Leier, T.T. Marquez-Lago, Delay chemical master equation: Direct and closed-form solutions. Proc. R. Soc. A 471, 20150049 (2015)
V. Sunkara, M. Hegland, An optimal finite state projection method. Procedia Comput. Sci. 1, 1579–1586 (2010)
V. Wolf, R. Goel, M. Mateescu, T. Henzinger, Solving the chemical master equation using sliding windows. BMC Syst. Biol. 4, 42 (2010)
K. Burrage, M. Hegland, S. MacNamara, R. Sidje, in A Krylov-based finite state projection algorithm for solving the chemical master equation arising in the discrete modelling of biological systems. Markov Anniversary Meeting: An International Conference to Celebrate the 150th Anniversary of the Birth of A.A. Markov (2006), pp. 21–38
B. Munsky, M. Khammash, The finite state projection algorithm for the solution of the chemical master equation. J. Chem. Phys. 124(4), 044104 (2006)
R.B. Sidje, H.D. Vo, Solving the chemical master equation by a fast adaptive finite state projection based on the stochastic simulation algorithm. Math. Biosci. 269, 10–16 (2015)
K.N. Dinh, R.B. Sidje, Understanding the finite state projection and related methods for solving the chemical master equation. Phys. Biol. 13(3), 035003 (2016)
A. Borghans, R.J. de Boer, L.A. Segel, Extending the quasi-steady state approximation by changing variables. Bull. Math. Biol. 58(1), 43 (1996)
E.A. Mastny, E.L. Haseltine, J.B. Rawlings, Two classes of quasi-steady-state model reductions for stochastic kinetics. J. Chem. Phys. 127(9), 094106 (2007)
P. Thomas, A.V. Straube, R. Grima, The slow-scale linear noise approximation: An accurate, reduced stochastic description of biochemical networks under timescale separation conditions. BMC Syst. Biol. 6(1), 39 (2012)
P. Thomas, R. Grima, A.V. Straube, Rigorous elimination of fast stochastic variables from the linear noise approximation using projection operators. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 86(4 Pt 1), 041110 (2012)
M. Barrio, A. Leier, T.T. Marquez-Lago, Reduction of chemical reaction networks through delay distributions. J. Chem. Phys. 138, 104114 (2013). doi:10.1063/1.4793982
A. Leier, M. Barrio, T.T. Marquez-Lago, Exact model reduction with delays: Closed-form distributions and extensions to fully bi-directional monomolecular reactions. J R Soc Interface (2014). https://doi.org/10.1098/rsif.2014.0108
A. Robson, K. Burrage, M.C. Leake, Inferring diffusion in single live cells at the single-molecule level. Phil Trans Roy Soc B 368, 20120029 (2013)
G. Lillacci, M. Khammash, Parameter estimation and model selection in computational biology. PLoS Comput. Biol. 6, e1000696 (2010)
T. Toni, D. Welch, N. Strelkowa, A. Ipsen, M.P.H. Stumpf, Approximate Bayesian computation scheme for parameter inference and model selection in dynamical systems. J Roy Soc Interface 6, 187–202 (2009)
D.J. Wilkinson, Bayesian methods in bioinformatics and computational systems biology. Brief. Bioinform. 8, 109–116 (2007)
Acknowledgements
The authors would like to thank the Isaac Newton Institute for Mathematical Sciences at the University of Cambridge, UK under the scientific program entitled Stochastic Dynamical Systems in Biology: Numerical Methods and Applications for supporting our visit and participation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Burrage, K., Burrage, P., Leier, A., Marquez-Lago, T. (2017). A Review of Stochastic and Delay Simulation Approaches in Both Time and Space in Computational Cell Biology. In: Holcman, D. (eds) Stochastic Processes, Multiscale Modeling, and Numerical Methods for Computational Cellular Biology. Springer, Cham. https://doi.org/10.1007/978-3-319-62627-7_11
Download citation
DOI: https://doi.org/10.1007/978-3-319-62627-7_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-62626-0
Online ISBN: 978-3-319-62627-7
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)