Abstract
Understanding disease-disease associations can not only help us gain deeper insights into complex diseases, but also lead to improvements in disease diagnosis, drug repositioning and new drug development. Due to the growing body of high-throughput biological data, a number of methods have been proposed for the computation of similarity among diseases during past decades. Recently, the disease module theory has been presented, which states that disease-related genes or proteins tend to interact with each other in the same neighborhood of protein-protein interaction network. In this study, we propose a new method called ModuleSim to measure associations between diseases by using disease-gene association data and protein-protein interaction network data based on disease module theory. By considering the interactions between disease modules and each module’s modularity, ModuleSim outperforms other four popular methods for predicting disease-disease similarity.
This work is supported by the National Science Fund for Excellent Young Scholars under Grant No. 61622213, the National Natural Science Foundation of China under grant No. 61370024 and No. 61472133, and the Program of Independent Exploration Innovation in Central South University (2016zzts354).
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Quantifying the associations among diseases is now playing an important role in modern biology and medicine, as discovering associations among diseases could be helpful for us to get a deeper knowledge of pathogenic mechanisms of complex diseases. Based on the hypothesis that similar diseases may be caused by the same or similar genes, the measurement of disease-disease associations is widely used in the study of disease gene prediction [1, 2, 33] and drug repositioning [3].
A number of approaches measuring disease-disease associations have been proposed during last decade [4,5,6,7,8]. Different approaches measures disease-disease associations from different perspectives by taking advantage of different biological data. These approaches can be broadly grouped into two classes: semantic-based methods and function-based methods [9]. Semantic-based methods take advantage of the structure of disease terminology such as Disease Ontology (DO) [10] and Medical Subject Headings (MeSH) [11] to measure the semantic similarity of diseases [12, 13]. Function-based methods are basically based on the hypothesis that similar diseases may have more same or similar causing genes/gene products [5, 14].
Mathur et al. proposed a method called BOG [15] which calculates disease similarity by comparing the overlapping of disease-related gene sets. Further, Mathur et al. proposed another method called PSB [16] which computes disease similarity based on biological process terms of Gene Ontology (GO) [17] associated with disease-related genes. By exploiting functional associations among disease-related genes based on GO, PSB outperforms BOG. To get a better performance, many other methods take advantage of disease-related genes’ interactions in protein-protein interaction networks (PPIN). FunSim [9] measures disease similarity by using a weighted human PPIN in which the weight of each interaction measures the functional association of a gene pair [32]. However, FunSim takes only the first neighbors of each gene into account, rather than making full use of the entire PPIN. Sun et al. [18] applied graphlet theory [19] to calculate gene similarity in PPIN. Then they inferred disease similarity by using disease-related genes’ graphlet similarity. Hamaneh et al. [20] proposed a method that first assigns weights to all proteins from a disease to the PPIN and back. Then the method calculates similarity between two diseases as cosine of the angel between their corresponding weight vectors. NetSim [21] uses random walk with restart (RWR) [22] to score the functional relevance between a gene and a disease. The functional relevance scores are then used to measure disease similarity.
Although there have been many methods (such as Sun’s method [18], Hamaneh’s method [20] and NetSim [21]) which take advantage of PPIN to discover disease-disease associations, these methods rarely consider the modularity of genes related to each disease in PPIN. According to the disease module theory, the disease-related genes or proteins are not scattered randomly in PPIN, but tend to interact with each other, forming one or several connected subgraphs which can be called the disease module [23, 40]. However, as the PPIN and our knowledge of disease-related genes remain incomplete, there also exist lots of disease modules that are not observable in PPIN. In this study, we propose a method to relate diseases based on disease module theory. In this method, we consider the related genes of two diseases as two modules in PPIN. We take advantage of shortest path of each gene pair between the two modules to measure the association of the two modules. Furthermore, for the purpose of overcoming the incompleteness of disease modules, we also take the modularity of each disease module into account. In the comparison with other proposed methods used PPIN, our method shows the best performance.
2 Materials and Methods
2.1 Materials
Disease-Gene Associations:
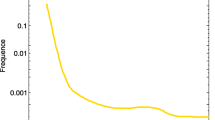
The disease-gene association data are downloaded from two databases: SIDD [25] and DisGeNET [24]. By integrating disease-gene associations from five databases (GeneRIF [34], Online Mendelian Inheritance in Man (OMIM) [35], Comparative Toxicogenomics Database (CTD) [36], Genetic Association Database (GAD) [37], and SpliceDisease [38]), SIDD contains 99658 associations between 2423 diseases and 10527 genes in total (Fig. 1). SIDD uses DOID [10] as the unique identifier for each disease.
Evaluation of ModuleSim against DO classification by using different datasets (the barplot shows similarity scores between disease pairs from the same DO categories, compared with those from different DO categories and all disease pairs). Note that two diseases are said to be in the same category if they have at least one common ancestor in the 3rd-level DO categories.
DisGeNET integrates human disease-gene associations from various expert curated databases and text-mining derived associations including Mendelian, complex and environmental diseases [24]. DisGeNET v4.0 contains 429036 associations between 17381 genes and 15,093 diseases. Because of the low reliability of disease-gene associations from literature in DisGeNET, a disease-gene association is adopted only if its DisGeNET score is not less than 0.06 [24]. DisGeNET uses Unified Medical Language System Identifier (UMLS ID) [39] as the unique identifier for each disease. After mapping disease ids from UMLS ID into DOID, in total, we got 1511 diseases, 6929 genes and 20787 associations between them from DisGeNET.
PPIN:
Two PPIN datasets were adopted. One is called hPPIN. As Li et al. [21] did, hPPIN was built by integrating four existing protein interaction databases (BioGrid [26], HPRD [27], IntAct [28], and HomoMINT [29]). In total, hPPIN contains 17506 proteins and 284476 interactions. The other is human interactome which was formed by experimentally documented molecular interactions as Menche et al. [23] did. The interactome integrates protein-protein and regulatory interactions, and metabolic pathway and kinase-substrate interactions. The union of all interactions in the interactome forms a network which contains 13460 proteins and 141296 physical interactions between them.
2.2 Methods
In disease module theory, a disease is considered as a subgraph consisting of genes related to the disease and the interactions between these genes in PPIN [23, 40]. In other words, any perturbation of the nodes in a disease module can be linked to the disease. If genes in two disease modules overlap or stay in the same neighborhood, the perturbations leading to one disease will likely disrupt the other disease modules as well, which results in shared clinical characteristics [23]. However, limited to the fact that our knowledge of disease-related genes and PPIN are still incomplete, lots of disease modules are not observable. Based on disease module theory and the fragmentation of disease modules, we proposed a method called ModuleSim to calculate disease-disease associations. Firstly, we use the length of the shortest path to calculate the strength of two genes’ relevance as follows:
where sp(g 1 ,g 2 ) represents the length of the shortest path between node g 1 and node g 2 in PPIN, A and b are two constants. To keep the value of sim(g 1 ,g 2 ) within the range [0, 1], we used A = 1 and b = 1, respectively. A higher sim(g 1 ,g 2 ) value represents a closer relationship between g 1 and g 2 . Suppose G is a disease module, which means G is a gene set associated with a disease, we then measure a gene’s relevance to a disease as follows:
As in Eq. (2), the relevance score of a gene g with the disease is calculated as the average transformed distance between g and genes in G.
Suppose G 1 = {g 11 , g 12 , …, g 1m } is a disease module which contains m genes, G 2 = {g 21 , g 22 , …, g 2n } is another disease module which contains n genes. The relatedness between the two disease modules is quantified by Eq. (3).
Our knowledge of disease-associated genes and PPIN remain incomplete [23]. This is to say, there also exist lots of diseases of whose modularity is not obvious. To overcome the incompleteness of disease modules, we normalize the relatedness score between G 1 and G 2 by dividing the average of relatedness scores of themselves as Eq. (4).
In Eq. (4), ModuleSim(G 1 , G 2 ) represents the ModuleSim of disease module G 1 and G 2 . A higher ModuleSim value represents a closer connection between G 1 and G 2 .
3 Experiments and Results
3.1 Correlation with Disease Classification of DO
The results obtained by ModuleSim were first evaluated against the disease classification of DO. DO is a standardized ontology for human disease concepts with stable identifiers organized by disease etiology [10]. DO (version: releases/2016-05-27) contains 6930 non-obsolete disease terms and 6921 disease terms under the 3rd-level categories. We say that two diseases are in the same class, if they have at least one common ancestor in the 3rd-level DO categories. To investigate the correlation between ModuleSim and the disease classification of DO, we tested whether disease pairs from the same DO classes tends to have higher similarity scores than disease pairs from different DO classes (Fig. 1). Our results show that for all four situations when using different disease-gene association datasets and PPIN datasets, similarity scores of disease pairs from the same classes are higher than those from different classes.
3.2 Evaluation of ModuleSim on the Benchmark Set
We adopted the benchmark set method [9] to evaluate ModuleSim with other methods. 70 disease pairs with high similarity derived from two manually checked datasets by Suthram et al. [30] and Pakhomov et al. [31] were taken as the benchmark set. Receiver operating characteristic (ROC) curves were then drawn with the benchmark set against 100 random sets. Each random set contains 700 randomly selected pairs.
We compared ModuleSim with other four popular methods which are all using disease-gene association data and PPIN data to measure disease-disease associations: Hamaneh [20], FunSim [9], Sun_topo [18], NetSim [21]. As shown in Fig. 2A, when using disease-gene associations from SIDD [25] and hPPIN as the PPIN, the Hamaneh method [20], with an average area under the ROC curve (AUC) of 93.7%, had the worst performance. By considering the functional weights between disease-related genes in PPIN, FunSim [9] got an AUC of 94.4%. NetSim [21] which took the entire interaction network into account by using RWR improved the AUC to 95.1%. By using graphlet theory [19], Sun_topo [18] got a higher AUC of 96.1%. The proposed method, ModuleSim, got the highest AUC of 96.9%. For a further comparison, we also checked how many answer disease pairs out of the top-ranking disease pairs can be found by ranking the benchmark pairs and the random pairs in descending order based on each method. From Fig. 2B we can see that, ModuleSim always find the most answer disease pairs in the top-ranking 150 disease pairs. Furthermore, ModuleSim find all 70 benchmark pairs by using the least top-ranking disease pairs, which showed a quite good performance. For example, “pneumonia” (DOID:552) and “meningitis” (DOID:9471) are two diseases which are validated to have high similarity with each other in the benchmark set. There are only six genes related to “meningitis” based on SIDD [25], which leads to the result that the disease module of “meningitis” is fragmentary. Thus, the average ranking of “pneumonia” and “meningitis” in the 770 disease pairs (70 benchmark pairs and 700 randomly selected pairs) is very low for all five methods, as shown in Table 1. However, by considering the modularity of each diseases, ModuleSim obtained an average ranking of 251 of “pneumonia” and “meningitis”, which raised about 100 places compared with Hamaneh and Sun_topo.
Only 55.3% of disease-gene associations in DisGeNET [24] and 11.5% of disease-gene associations in SIDD [25] are shared with each other, which shows that the two databases have a big difference in quantity with each other. Similarly, different PPIN datasets are also very different. The two PPIN datasets (interactome [23] and hPPIN [21]) used in this paper only have 12560 genes and 90938 interactions in common. To test the influence of different datasets, we further evaluated the five methods by using these two different disease-gene association databases and two different PPIN datasets. As shown in Fig. 3, ModuleSim got the best performance in all four situations, which indicated that ModuleSim have a stable and strong power for discovering disease-disease associations.
4 Conclusion and Discussion
It is a big challenge to get a deeper insight into the mechanisms between diseases in modern biology [41, 42]. Measuring disease-disease associations is helpful for us to gain more knowledge about diseases. A number of methods have been proposed for measuring disease-disease associations up to now. The methods which take advantage of disease-gene associations and PPIN have shown a great power to infer disease-disease associations. However, these methods rarely consider the modularity of genes related to each disease in PPIN.
According to the disease module theory, the disease-related genes or proteins are not scattered randomly in PPIN, but tend to interact with each other [23, 40]. In this study, we proposed a method ModuleSim to discovering disease-disease associations based on disease module theory. In the result of ModuleSim, similarity scores of disease pairs from the same DO classes are higher than those from different DO classes. Furthermore, ModuleSim outperformed other four methods (Hamaneh [20], FunSim [9], Sun_topo [18], NetSim [21]) in the evaluation of benchmark set.
ModuleSim considers modularity of each disease module when measuring disease-disease associations. However, our knowledge of disease-related genes and PPIN remains incomplete. Therefore, lots of disease modules remain incomplete. In the future, more disease-gene associations and gene-gene interactions with high quality need to be discovered. In addition, the application of ModuleSim on disease-gene prediction and drug repositioning is worthy of further investigation.
References
Vanunu, O., Magger, O., Ruppin, E., et al.: Associating genes and protein complexes with disease via network propagation. PLoS Comput. Biol. 6(1), e1000641 (2010)
Li, M., Zheng, R., Li, Q., et al.: Prioritizing disease genes by using search engine algorithm. Curr. Bioinform. 11(2), 195–202 (2016)
Luo, H., Wang, J., Li, M., et al.: Drug repositioning based on comprehensive similarity measures and Bi-Random walk algorithm. Bioinformatics 32(17), 2664–2671 (2016)
van Driel, M.A., Bruggeman, J., Vriend, G., et al.: A text-mining analysis of the huamn phenome. Eur. J. Hum. Genet. 14(5), 535–542 (2006)
Goh, K.I., Cusick, M.E., Valle, D., et al.: The human disease network. Proc. Natl. Acad. Sci. 104(21), 8685–8690 (2007)
Jung, J., Lee, D.: Inferring disease association using clinical factors in a combinatorial manner and their use in drug repositioning. Bioinformatics 29(16), 2017–2023 (2013)
Sun, K., Buchan, N., Larminie, C., et al.: The integrated disease network. Integr. Biol. 6(11), 1069–1079 (2014)
Frick, J.M., Guha, R., Peryea, T., et al.: Evaluating disease similarity using latent Dirichlet allocation. bioRxiv: 030593 (2015)
Cheng, L., Li, J., Ju, P., et al.: SemFunSim: a new method for measuring disease similarity by integrating semantic and gene functional association. PLoS One 9(6), e99415 (2014)
Schriml, L.M., et al.: Disease ontology: a backbone for disease semantic integration. Nucleic Acids Res. 40(D1), D940–D946 (2012)
Lipscomb, C.E.: Medical subject headings (MeSH). Bull. Med. Libr. Assoc. 88(3), 265 (2000)
Yu, G., Wang, L.G., Yan, G.R., et al.: DOSE: an R/Bioconductor package for disease ontology semantic and enrichment analysis. Bioinformatics 31(4), 608–609 (2015)
Wang, D., Wang, J., Lu, M., et al.: Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 26(13), 1644–1650 (2010)
Zhang, X., Zhang, R., Jiang, Y., et al.: The expanded human disease network combining protein–protein interaction information. Eur. J. Hum. Genet. 19(7), 783–788 (2011)
Mathur, S., Dinakarpandian, D.: Automated ontological gene annotation for computing disease similarity. AMIA Summits Transl. Sci. Proc. 2010, 12–16 (2010)
Mathur, S., Dinakarpandian, D.: Finding disease similarity based on implicit semantic similarity. J. Biomed. Inform. 45(2), 363–371 (2012)
Ashburner, M., Ball, C.A., Blake, J.A., et al.: Gene ontology: tool for the unification of biology. Nat. Genet. 25(1), 25–29 (2000)
Sun, K., Gonçalves, J.P., Larminie, C., et al.: Predicting disease associations via biological network analysis. BMC Bioinform. 15(1), 1 (2014)
Milenkoviæ, T., Pržulj, N.: Uncovering biological network function via graphlet degree signatures. Cancer Inform. 6, 257 (2008)
Hamaneh, M.B., Yu, Y.K.: Relating diseases by integrating gene associations and information flow through protein interaction network. PLoS ONE 9(10), e110936 (2014)
Li, P., Nie, Y., Yu, J.: Fusing literature and full network data improves disease similarity computation. BMC Bioinform. 17(1), 326 (2016)
Köhler, S., Bauer, S., Horn, D., et al.: Walking the interactome for prioritization of candidate disease genes. Am. J. Hum. Genet. 82(4), 949–958 (2008)
Menche, J., Sharma, A., Kitsak, M., et al.: Uncovering disease-disease relationships through the incomplete interactome. Science 347(6224), 1257601 (2015)
Piñero, J., Queralt-Rosinach, N., Bravo, À., et al.: DisGeNET: a discovery platform for the dynamical exploration of human diseases and their genes. Database 2015, bav028 (2015)
Cheng, L., Wang, G., Li, J., et al.: SIDD: a semantically integrated database towards a global view of human disease. PLoS ONE 8(10), e75504 (2013)
Chatr-Aryamontri, A., Breitkreutz, B.J., Heinicke, S., et al.: The BioGRID interaction database: 2013 update. Nucleic Acids Res. 41(D1), D816–D823 (2013)
Prasad, T.S.K., Goel, R., Kandasamy, K., et al.: Human protein reference database—2009 update. Nucleic Acids Res. 37(suppl 1), D767–D772 (2009)
Orchard, S., Ammari, M., Aranda, B., et al.: The MIntAct project—IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 42(D1), D358–D363 (2013)
Persico, M., Ceol, A., Gavrila, C., et al.: HomoMINT: an inferred human network based on orthology mapping of protein interactions discovered in model organisms. BMC Bioinform. 6(4), 1 (2005)
Suthram, S., Dudley, J.T., Chiang, A.P., et al.: Network-based elucidation of human disease similarities reveals common functional modules enriched for pluripotent drug targets. PLoS Comput. Biol. 6(2), e1000662 (2010)
Pakhomov, S., McInnes, B., Adam, T., et al.: Semantic similarity and relatedness between clinical terms: an experimental study. In: AMIA annual symposium proceedings. American Medical Informatics Association, p. 572 (2010)
Lee, I., Blom, U.M., Wang, P.I., et al.: Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res. 21(7), 1109–1121 (2011)
Ni, J., Koyuturk, M., Tong, H., et al.: Disease gene prioritization by integrating tissue-specific molecular networks using a robust multi-network model. BMC Bioinform. 17(1), 453 (2016)
Mitchell, J.A., Aronson, A.R., Mork, J.G., et al.: Gene indexing: characterization and analysis of NLM’s GeneRIFs. In: AMIA (2003)
Amberger, J.S., Bocchini, C.A., Schiettecatte, F., et al.: OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 43(D1), D789–D798 (2015)
Davis, A.P., Murphy, C.G., Johnson, R., et al.: The comparative toxicogenomics database: update 2013. Nucleic Acids Res. 41(D1), D1104–D1114 (2012)
Becker, K.G., Barnes, K.C., Bright, T.J., et al.: The genetic association database. Nat. Genet. 36(5), 431–432 (2004)
Wang, J., Zhang, J., Li, K., et al.: SpliceDisease database: linking RNA splicing and disease. Nucleic Acids Res. 40(D1), D1055–D1059 (2012)
Bodenreider, O.: The unified medical language system (UMLS): integrating biomedical terminology. Nucleic Acids Res. 32(suppl 1), D267–D270 (2004)
Barabási, A.L., Gulbahce, N., Loscalzo, J.: Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12(1), 56–68 (2011)
Guo, X., Zhang, J., Cai, Z., et al.: Searching genome-wide multi-locus associations for multiple diseases based on Bayesian Inference. In: IEEE/ACM transactions on computational biology and bioinformatics (2016)
Teng, B., Yang, C., Liu, J., et al.: Exploring the genetic patterns of complex diseases via the integrative genome-wide approach. IEEE/ACM Trans. Comput. Biol. Bioinform. 13(3), 557–564 (2016)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this paper
Cite this paper
Ni, P. et al. (2017). Relating Diseases Based on Disease Module Theory. In: Cai, Z., Daescu, O., Li, M. (eds) Bioinformatics Research and Applications. ISBRA 2017. Lecture Notes in Computer Science(), vol 10330. Springer, Cham. https://doi.org/10.1007/978-3-319-59575-7_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-59575-7_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-59574-0
Online ISBN: 978-3-319-59575-7
eBook Packages: Computer ScienceComputer Science (R0)