Abstract
This chapter describes crystal structures of phosphodiesterases (PDEs) that are involved in CNS diseases and their interactions with family selective inhibitors. The structural comparison identifies a small hydrophobic pocket next to the active site, which may be valuable for improvement of selectivity of PDE inhibitors.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Phosphodiesterase (PDE) is a superfamily of enzymes hydrolyzing the second messengers cGMP and cAMP. Human genome encodes 21 genes that are categorized into 11 PDE families and express >100 isoforms of proteins (Conti and Beavo 2007; Maurice et al. 2014). PDE molecules contain a conserved catalytic domain at the C-terminus and a variable regulatory domain at the N-terminus. The catalytic domains of PDEs are well conserved but selectively hydrolyze their preferable substrates: PDE5, PDE6, and PDE9 specifically recognize cGMP as their substrate, while PDE4, PDE7, and PDE8 are cAMP-specific. The remaining PDE families are capable of degrading both substrates. For critical roles of the second messengers cAMP and cGMP in physiological processes, PDE inhibitors have been widely studied as therapeutics for treatment of human diseases (Conti and Beavo 2007; Maurice et al. 2014). A well-known example is the PDE5 inhibitor sildenafil that has been approved for the treatment of erectile dysfunction and pulmonary hypertension (Rotella 2002; Galie et al. 2005). This chapter will describe interactions between the inhibitors and PDEs in CNS to provide structural insight into design of PDE inhibitors.
2 Conservation of the PDE Active Sites for Binding of Substrates and Inhibitors
After the first crystal structure of the PDE4B catalytic domain (Xu et al. 2000), structures of catalytic domains of nine out of eleven PDE families (PDE1–5 and PDE7–10) are available and have greatly guided design of PDE family selective inhibitors (Ke and Wang 2007). The catalytic domains of PDEs contain 300–350 amino acids and their active site pockets are well conserved (Table 17.1).
The active site can be divided into two major sub-pockets respectively for metal and inhibitor binding (Fig. 17.1). Two metal ions occupy the metal binding sub-pocket. A zinc ion was identified by the anomalous scattering experiment and coordinates with four invariant residues in PDE families: His529, His563, Asp 564, and Asp 674 in PDE10A2 (Table 17.1). Another metal ion has been assigned but not confirmed as magnesium or manganese, dependent on PDE family, and forms only one coordination with Asp564. A hydroxide ion bridges the two metal ions and presumably serves as nucleophile to initiate the hydrolysis of the cyclic nucleotide (Xu et al. 2000; Zhan and Zheng 2001; Huai et al. 2003). The two ions chelate with several water molecules and form an octahedral configuration. The sub-pocket for inhibitor binding shows two characteristic features: (1) an invariant glutamine (Gln726 in PDE10A2) forms a hydrogen bond with substrate or inhibitors and (2) a conserved phenylalanine (Phe729 in PDE10A2) stacks against a hydrophobic group of substrates/inhibitors such as adenosine of cAMP (Fig. 17.1b). In addition, two hydrophobic residues (Ile693 and Phe696 in PDE10A2) together with the conserved phenylalanine (Phe729) sandwich the adenosine of cAMP to form a hydrophobic slot.
The active site of PDE10A2 catalytic domain. (a) Surface presentation of the active site of PDE10A2. Substrate cAMP is shown as yellow sticks and zinc and magnesium are presented with green and pink balls. (b) Ribbon diagram for cAMP binding to PDE10A2. Dotted lines represent hydrogen bonds or charge-charge interactions with zinc and magnesium
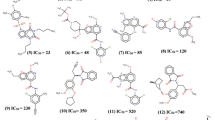
The above features are well conserved for binding of most PDE inhibitors. For example, the classic PDE4 inhibitor rolipram uses its catechin group to form two hydrogen bonds with the invariant glutamine (Gln369 in PDE4D2) and stacks against the conserved phenylalanine (Phe372 in PDE4D2, Huai et al. 2003). When the rolipram analogs such as roflumilast, a drug for the treatment of severe chronic obstructive pulmonary disease (Field 2008), are superimposed one another, they can be divided into five pharmacophores of CORE, R1-R4 (Fig. 17.2). The CORE pharmacophore or catechin forms two hydrogen bonds with Gln369 and stacks against Phe372 of PDE4D2 (Fig. 17.2), and may account for basic affinity of most inhibitors. The remaining pharmacophores are located in slightly different environments of PDE families and may thus contribute to selective binding of the inhibitors.
Overlay of PDE4 inhibitors with catechin scaffold suggests five pharmacophores. The CORE group is well superimposed and stacks against a conserved phenylalanine (Phe372 in PDE4D2), in addition to hydrophobic interactions with conserved Phe341 and Ile336, thus contributing basic affinity for non-selective binding of the inhibitors. Other pharmacophores R1-R4 appear to contribute to the selective binding of the PDE4 inhibitors
3 A Unique Subpocket for Improvement of Selective Binding of PDE Inhibitors
An early study on the crystal structure of the Leishmania major phosphodiesterase B1 (LmjPDEB1) revealed a small hydrophobic pocket neighboring the active site (Wang et al. 2007). Approach to this subpocket is gated by two residues: Met874 and Gly886 of LmjPDE1 (Fig. 17.3). Since the size of these two residues is apparently large in human PDE families, the subpocket was thought to specially belong to Leishmania major phosphodiesterases and was named as the L-pocket (Wang et al. 2007). Later, a careful examination on the structure of Trypanosoma brucei phosphodiesterase B1 and the sequences of other parasite PDEs show possible accession to the pocket in the parasite PDEs and thus the pocket was renamed as the P-pocket (Jansen et al. 2013). However, this claim was not completely justified by the crystal structure of human PDE10 catalytic domain in complex with inhibitor PF-2545902, in which a fragment of the PDE10 inhibitors interacts with the P-pocket and thus the pocket was named as the PDE10 selective pocket (Verhoest et al. 2009).
Recently, the crystal structure of PDE9 in complex with inhibitor C33 revealed that the tyrosyl group of (S)-enantiomer of C33 oriented to and interact with a small subpocket that is composed of a portion of helices H14 and H15 and the M-loop (Fig. 17.4), while its (R)-enantiomer adopted different conformation. Since the M-loop is a major contributor, the pocket was named as the M-pocket. Helices H14 (Leu420, Leu421 and Phe425) and H15 contribute two walls of the pocket, while Val447 and backbone of the M-loop form the bottom of the pocket. In fact, the M-pocket is identical to the P-pocket or PDE10-selective pocket, and may thus serve as a general pocket for improvement of selectivity of PDE inhibitors. Sequence alignment reveals that the gating residues vary significantly across PDE families (Fig. 17.3) so as to make the pocket inaccessible for many human PDEs (Wang et al. 2007; Wang et al. 2012).
In addition to the accessibility gated by two residues, the M-pocket shows significant different size, conformation, and sequence components (Fig. 17.3), and thus may serve as an important element for improvement of selective binding of PDE inhibitors. For example, the M-pocket in PDE5, which is composed of Val782, Ala783, Phe786, Phe787 and Ile813 (Fig. 17.5a), is small and slightly deeper than that of PDE9. Two large gating residues of Leu804 and Met816 would allow small group to penetrate into the pocket, such as an ethoxyl fragment in the PDE5-sildenafil structure (Wang et al. 2008). The affinity of the PDE5 inhibitors was significantly improved by the replacement of ethoxyl with propoxy group (Salem et al. 2006). The M-pocket in the PDE8A1 structure, which is made up of residues Ser745, Tyr748, Phe749, Phe767, and Cys772, is much shallower and smaller than that of PDE9 and might not well accommodate their inhibitors (Fig. 17.5b). In the structure of PDE1, the M-loop has a good conservation of amino acids to those of PDE9, as shown by correspondence of Leu388, Met389, and Phe392 of PDE1B to Leu420, Leu421, and Y424 of PDE9, respectively. Since part of the M-pocket is disordered in PDE1, it might reasonably predict a poor selectivity of PDE9 inhibitors against PDE1, as observed in the PDE9 structure (Wunder et al. 2005).
The M-pockets in some PDE families. (a) Surface presentation of the M-pocket of PDE5. (b) Surface presentation of the M-pocket of PDE8A. IBMX is a non-selective inhibitor of PDEs. (c) Ribbon presentation of the superposition of PDE9 (green) over PDE1B (cyan). The M-loop of PDE1B is partially disordered. The corresponding residues between PDE9A2 and PDE1B are: M365/M336, N405/H373, L420/L388, L421/M389, Y424/F392, Q453/Q421, and F456/F424
4 Conclusive Remarks
We believe that the M-pocket may serve as a selectivity determinant for inhibitors binding and is useful for improvement of inhibitor affinity and selectivity.
Abbreviations
- cAMP:
-
cyclic adenosine monophosphate
- cGMP:
-
cyclic guanosine monophosphate
- IBMX:
-
3-isobutyl-1-methylxanthine
- PDE:
-
phosphodiesterase
References
Conti M, Beavo J. Biochemistry and physiology of cyclic nucleotide phosphodiesterases: essential components in cyclic nucleotide signaling. Annu Rev Biochem. 2007;76:481–511.
Field SK. Roflumilast: an oral, once-daily selective PDE-4 inhibitor for the management of COPD and asthma. Expert Opin Investig Drugs. 2008;17:811–8.
Galie N, Ghofrani HA, Torbicki A, Barst RJ, Rubin LJ, Badesch D, Fleming T, Parpia T, Burgess G, Branzi A, Grimminger F, Kurzyna M, Simonneau G. Sildenafil citrate therapy for pulmonary arterial hypertension. N Engl J Med. 2005;353:2148–57.
Huai Q, Colicelli J, Ke H. The crystal structure of AMP-bound PDE4 suggests a mechanism for phosphodiesterase catalysis. Biochemistry. 2003;42:13220–6.
Jansen C, Wang H, Kooistra AJ, de Graaf C, Orrling KM, Tenor H, Seebeck T, Bailey D, de Esch IJ, Ke H, Leurs R. Discovery of novel Trypanosoma brucei phosphodiesterase B1 inhibitors by virtual screening against the unliganded TbrPDEB1 crystal structure. J Med Chem. 2013;56:2087–96.
Ke H, Wang H. Crystal structures of phosphodiesterases and implications on substrate specificity and inhibitor selectivity. Curr Top Med Chem. 2007;7:391–403.
Maurice DH, Ke H, Ahmad F, Wang Y, Chung J, Manganiello VC. Advances in targeting cyclic nucleotide phosphodiesterases. Nat Rev Drug Discov. 2014;13:290–314.
Rotella DP. Phosphodiesterase 5 inhibitors: current status and potential applications. Nat Rev Drug Discov. 2002;1:674–82.
Salem EA, Kendirci M, Hellstrom WJ. Udenafil, a long-acting PDE5 inhibitor for erectile dysfunction. Curr Opin Investig Drugs. 2006;7:661–9.
Verhoest PR, Chapin DS, Corman M, Fonseca K, Harms JF, Hou X, Marr ES, Menniti FS, Nelson F, O’Connor R, Pandit J, Proulx-Lafrance C, Schmidt AW, Schmidt CJ, Suiciak JA, Liras S. Discovery of a novel class of phosphodiesterase 10A inhibitors and identification of clinical candidate 2-[4-(1-Methyl-4-pyridin-4-yl-1H-pyrazol-3-yl)-phenoxymethyl]-quinoline (PF-2545920). J Med Chem. 2009;52:5188–96.
Wang H, Kunz S, Chen G, Seebeck T, Wan Y, Robinson H, et al. Biological and structural characterization of Trypanosoma cruzi phosphodiesterase C and implications for the design of parasite selective inhibitors. J Biol Chem. 2012;287:11788–97.
Wang H, Yan Z, Geng J, Kunz S, Seebeck T, Ke H. Crystal structure of the Leishmania major phosphodiesterase LmjPDEB1 and insight into the design of the parasite-selective inhibitors. Mol Microbiol. 2007;66:1029–38.
Wang H, Ye M, Robinson H, Francis SH, Ke H. Conformational variations of both phosphodiesterase-5 and inhibitors provide the structural basis for the physiological effects of vardenafil and sildenafil. Mol Pharmacol. 2008;73:104–10.
Wunder F, Tersteegen A, Rebmann A, Erb C, Fahrig T, Hendrix M. Characterization of the first potent and selective PDE9 inhibitor using a cGMP reporter cell line. Mol Pharmacol. 2005;68:1775–81.
Xu RX, Hassell AM, Vanderwall D, Lambert MH, Holmes WD, Luther MA, et al. Atomic structure of PDE4: insight into phosphodiesterase mechanism and specificity. Science. 2000;288:1822–5.
Zhan CG, Zheng F. First computational evidence for a catalytic bridging hydroxide ion in a phosphodiesterase active site. J Am Chem Soc. 2001;123:2835–8.
Acknowledgment
We thank for supports of NIH GM59791 to HK and the National Natural Science Foundation of China (31271944, 31471626).
Conflict of Interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Wang, Y., Ke, H. (2017). A Unique Sub-Pocket for Improvement of Selectivity of Phosphodiesterase Inhibitors in CNS. In: Zhang, HT., Xu, Y., O'Donnell, J. (eds) Phosphodiesterases: CNS Functions and Diseases. Advances in Neurobiology, vol 17. Springer, Cham. https://doi.org/10.1007/978-3-319-58811-7_17
Download citation
DOI: https://doi.org/10.1007/978-3-319-58811-7_17
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-58809-4
Online ISBN: 978-3-319-58811-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)