Abstract
Cyclic nucleotide phosphodiesterase Type 4 specifically metabolizes Cyclic Adenosine Monophosphate and has widespread distribution across the human body. The aim of this study was to generate a well-validated ligand-based 3D Quantitative Structure Activity Relationship pharmacophore model to identify potential phosphodiesterase Type 4 inhibitors using a set of 18 known chemically diverse phosphodiesterase Type 4 inhibitors. The HypoGen module of Discovery Studio v4.1 was used to generate the aforementioned pharmacophore model which was then employed as 3D query for virtual screening of four chemical and two natural product databases. The top hits were evaluated for their drug-like properties. The binding orientations of the best fits were predicted by molecular docking. Orbital energies were predicted for top hits and the density functional theory based minimum energy gap (Highest Occupied Molecular Orbital–Lowest Unoccupied Molecular Orbital gap) was used to further cull the selection and identify the most potential phosphodiesterase Type 4 inhibitors. Chemical similarity search was performed and structural analogs of the best hits were designed to discover novel potential phosphodiesterase Type 4 inhibitors. Use of Hypo1 as 3D query for virtual screening yielded 1243 compounds and subsequent molecular docking studies narrowed it down to 19 potential phosphodiesterase Type 4 inhibitors while a density functional theory-based study further culled this selection down to six most potential inhibitors. Six structurally diverse chemical structures with novel scaffolds and six analogs of the best hits were identified using pharmacophore modeling to be potential phosphodiesterase Type 4 inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cyclic Adenosine Monophosphate (cAMP) is one of the most crucial secondary messengers in eukaryotic systems. It has pivotal roles in myriad essential cellular functions like cell proliferation and differentiation, gene expression, immune and inflammatory responses, smooth muscle relaxation, energy homeostasis, visual transduction, fluid homeostasis, cardiac rate and contractility, cognition, depression, oocyte maturation, and apoptosis (Maurice et al. 2014; Lugnier 2011; Azevedo et al. 2013; Jin et al. 2012). Phosphodiesterases (PDEs) are a super-family of enzymes that metabolize cyclic nucleotides-cAMP and/or cyclic guanosine monophosphate through hydroxylation and are thus involved in the intracellular regulation of these signal transducers. There are 11 distinct phosphodiesterase gene families (PDE1 to PDE11) based on sequence similarity, substrate specificity, and pharmacological properties and are encoded by a total of 21 genes (Lugnier 2006; Richter et al. 2013). Transcription level regulations of these genes give rise to about 100 distinct PDE isoforms with precise functions and with widespread tissue-specific distribution throughout the body. Of these, the PDE4 family is the most complex and considered to be having the widest distribution in the human body. This family is encoded by four genes, viz., PDE4A, PDE4B, PDE4C, and PDE4D and comprises of about 20 distinct is forms with unique non-redundant physiological functions (Parker et al. 1992; Chong et al. 2013). Elevated levels of PDE4 have been found to occur in cells involved with inflammation like leukocytes and activated lymphocytes, and hence inhibitors to these enzymes have been introduced for the treatment of chronic inflammatory diseases (Maurice et al. 2014). Further PAN-selective PDE4 inhibitors have also been reported to exhibit neuroprotective and neuroregenerative properties that exert memory and cognition enhancing effects in preclinical models.
However though selective PDE4 inhibitors like Rolipram, Ibudilast, Roflumilast and Apremilast are now prescribed, they have significant side-effects which include nausea, emesis, diarrhea, and weight loss. These dose-limiting side effects are often attributed to the ubiquitous distribution of PDE4 in human cells and tissues (Beghè et al. 2013; Castro et al. 2010). To combat this, inhibitors with more specific binding to the targeted PDE4 subtype are required. In the present study, pharmacophore modeling, database screening and molecular docking approaches have been employed for identification of lead candidates which would serve as a basis for potent PDE4 inhibitor drug design with high therapeutic index.
Herein, we have employed the pharmacophore modeling approach towards the identification of selective PDE4 inhibitors. Further, well validated hypothesis was used to screen database molecules with potential activity to PDE4. Virtual Filters such as Drug Likeness, ADMET Screening, Docking Studies and Density functional theory (DFT) Computation were used to identify the most selective PDE4 inhibitors. Structural analogs of the best hit were further generated in order to discover novel scaffold for PDE4 inhibition.
Materials and methods
Selection and preparation of data set
A total of 78 chemical compounds with diverse structural features and having a wide range of inhibitory activity against PDE4- identified using the same type of biological assay were collected from literature (Buckley et al. 2002; Buckley et al. 2002; Provins et al. 2006; Ochiai et al. 2004; Gilbert et al. 2000; Muller et al. 1998) and the ChEMBL database (Davies et al. 2015) for data set preparation. Of these, 18 structurally diverse compounds with activity (IC50) value between 1.2–9700 nmol/L- spanning four orders of magnitude were used for training set preparation to warrant statistical relevance (Fig. 1). The remaining 60 compounds were used to construct the Test set for pharmacophore validation. 2D chemical structures of all the compounds that constituted the data set were sketched using ChemSketch, v6.2.0 (ACD Inc., Toronto, Canada) and subsequently converted to 3D structures in BIOVIA Discovery Studio v4.1 (DS). The compounds of the training set and test set were categorized as Most active IC50 < 75 nmol/L (++++); Active 75 nmol/L ≤ IC50 < 750 nmol/L (+++); Moderately Active 750 nmol/L ≤ IC50 < 1750 nmol/L (++); Inactive IC50 ≥ 1750 nmol/L (+). Dataset compounds were further checked for added hydrogens and minimized using CHARMM based smart minimizer that performs 1000 steps of steepest descent with ensuing conjugate gradient algorithms with a convergence gradient of 0.001 kcal mol−1 (John et al. 2011). After these steps of minimization, 230 acceptable conformers of each of the compounds were derived using the Poling algorithm at an energy threshold of 15 kcal mol−1. The Diverse Conformer Generation protocol running with Best/Flexible conformer generation option (Discovery Studio v4.1 (DS) was applied to generate these multiple conformers. The principle of rigorous energy minimization in both torsional and cartesian space that is employed in this option ensures the best coverage of conformational space by application of the poling algorithm (Arooj et al. 2011; Niu et al. 2013).
3D pharmacophore modeling
A quantitative hypothesis that established a correlation between the experimental and the predicted activity values of the inhibitory compounds using the HypoGen module of DS v4.1 was generated. The HypoGen algorithm estimates the activity of each training set compound by computing regression analysis-employing parameters such as the relationship of geometric fit value at variance with the negative logarithm of the activity. Identification and selection of the best hypothesis is based on its ability to predict most accurately the biological activity of unknown compounds in the chemical databases. While performing the hypothesis, the uncertainty value (default of 3.0) was changed to 2.0 for the training set compounds. The ratio of the reported activity value in relation to the minimum value is represented by the uncertainty, and the maximum values greater than 1.0. The uncertainty value classifies ligands in the data set as either active or inactive compounds and is used during the constructive and subtractive phases of pharmacophore model derivation. An uncertainty value of 2.0 was deemed suitable for this data set because the compound activities encompassed the requisite 4 orders of magnitude, which is supported by previous literature (Brogi et al. 2009; Kansal et al. 2010; Kumar et al. 2015) Prior to pharmacophore modeling, 1000 possible pharmacophores for all the selected ligands were predicted by first identifying the common features of the active inhibitors against Type 4 cAMP phosphodiesterase using Feature mapping/DS protocol of DS v4.1. These features served as the basis for potential inputs to complete the final pharmacophore hypotheses generation (Sakkiah and Lee 2012). The numbers of maximum and minimum features were set at 5 and 0 respectively for development of the series of hypotheses. Using specified parameters and the aforementioned algorithms, we have developed ten pharmacophore models. These models were further used in database screening on the basis of distinct computed parameters like Cost Value, Root Mean Square, and Fit Values (Sakkiah and Lee 2012; Debnath 2002).
Hypothesis validation
The quintessential attributes of pharmacophore models are their statistical significance, accurate prediction of the desired activity of molecules, and correct retrieval of active compounds from databases. The validation of the best pharmacophore model in this study has been performed using robust approaches such as Fischer’s Randomization and Test set (Debnath 2002). The former was performed simultaneously during the pharmacophore model development and yielded a number of spreadsheets by the random shuffling of the activity value of the training set compounds which is dependent upon the selected significance level (90, 95, 98, and 99%). The selected level of significance in this study was 95% and 19 spread sheets were thus computed. Test set was employed for estimation of the predictive power of the best hypothesis to identify molecules with orders of magnitude of activity similar to that of the active training set. It could give us distinct values of the PDE4 inhibitor activity estimated in relation to experimental activity of the same.
Virtual screening
Pharmacophore-based virtual screening was performed to identify potential PDE4 inhibitors from chemical databases through the well-validated hypothesis that served as 3D structural search query. The 460695 compounds that were used for screening and identification of potential PDE4 inhibitors belonged to the following databases- ChemBridge (www.chembridge.com), MayBridge (www.maybridge.com), NCI (www.cactus.nci.nih.gov), ChemDiv (www.chemdiv.com), and two specific natural product databases, viz., TCM Database of Taiwan (www.tcm.cmu.edu.tw) and IB Screen NP (www.ibscreen.com). The Best/Flexible Search Protocol from Ligand Pharmacophore Mapping of DS v4.1 was applied to identify and retrieve novel hits. The 300 best hits (Best S = 300) of each database, based on the Fit value with minimum predicted IC50 value were considered for ADMET study. Prior to virtual screening, the hit compounds were screened for their drug-likeness. The identification of compounds which tested positive for Lipinski’s Rule of Five and Veber’s drug-like filtration was conducted using the Ligand Filtration module of DS v4.1. Lipinski’s rule of five imposes the following criteria to be met for selection of a compound as Lipinski-positive: the octanol/water partition coefficient < 5, molecular weight < 500, number of H–bond donors < 5 and the number of H-bond acceptors < 10. Veber further decreed that for a compound to be drug-like it should have no more than 10 rotatable bonds and a polar surface area of 140 Å, or the sum of H-bond donors and acceptors of the compound should be no more than 12, while the number of rotatable bonds should be no more than 10 (Lipinski et al. 1997; Veber et al. 2002). Compounds not fulfilling more than one criterion of these specifications were phased out prior to ADMET prediction.
ADMET prediction
Optimum pharmacokinetics properties are quintessential requirements for consideration of any compound as a candidate drug molecule. Hits obtained through the 3D pharmacophore-based virtual screening were hence further filtered by ADMET prediction by deploying the ADMET properties prediction protocol of DS 4.1. The relevant properties that were examined are blood–brain barrier permeability, solubility, human intestinal absorption, Oral bioavailability and hepatotoxicity. The AMES test is an inexpensive, easy and fast screening procedure to identify the mutagenic potential of a chemical compound or a mixture to induce mutations in DNA. This is of primary significance as mutagens are often identified as potential carcinogens (Mortelmans and Zeiger 2000). The compounds which qualified for accepted levels of ADME were hence further subjected to AMES mutagenicity screening. The non-mutagenic compounds were finally selected for further molecular docking and interaction studies.
Molecular docking
The receptor model of PDE4 with a resolution of 1.72Ả (PDB ID- 1XON) was acquired from Protein Data Bank. Prior to docking, the protein model was prepared using the Protein Preparation Module of DS v4.1, and subsequent energy optimization was performed using CHARMM force field, selecting the steepest descent method followed by conjugate gradient at default mode. The active site of the protein was predicted and identified using two distinct docking algorithms, viz., MolDock and LibDock, and validation of the same was performed by docking the co-crystal ligand (Piclamilast) to the active site of the protein. The hits depicting the identified PDE4 inhibitors obtained through database screening and ADMET screening were then subjected to molecular docking studies using the aforementioned docking softwares. MolDock- a component of Molegro Virtual Docker v6.0 (MVD) is a molecular docking software which is based on a differential evolution algorithm the solution of which takes into account the sum of the intermolecular interaction energy between the ligand and the protein, and the intramolecular interaction energy of the ligand (Thomsen and Christensen 2006) The docking energy scoring function is based on the modified piecewise linear potential and is inclusive of new hydrogen bonding and electrostatic terms (De Azevedo and Walter 2010). MolDock was set at a maximum iteration of 1500 with a simplex evolution size of 50 and a minimum of 20 runs. The flexibility of the bonds of the ligands and the side chain flexibility of the amino acids in the binding cavity were set with a tolerance of 1.10 and strength of 0.90 for docking simulations. RMSD threshold for multiple cluster poses was computed at < 2.00 Å. LibDock is a component of the DS v4.1 software. Consensus docking scores were used to compare the binding orientation of top hits with the active site of the PDE4 receptor.
DFT-based analysis
Calculation of the orbital energies has been performed with an objective to reveal significant information chiefly on the electrostatic properties of the known PDE4 inhibitors. DFT analysis has found increasing application in quantum chemistry through quantum mechanical simulation of a periodic system (Sakkiah and Lee 2012).
Geometry optimization and Hessian calculations of ligands and corresponding calculations were performed using Becke’s three parameter exchange functionals (B3LYP), in combination with the Lee-Yang-Parr correlation energy functional (Lee et al. 1988). DFT has been identified as one of the best methods to study medium-sized and larger molecular systems (Devereux et al. 2009).The best DFT methods have also been shown to achieve substantially greater precision than the Hartree—Fock theory by incurring only a modest augment in cost (Panchmatia et al. 2010). This is efficiently accomplished by incorporating a few effects of electron correlation with less affluence than traditional correlated methods. On the basis of the treatment in exchange and correlation components, a range of functionals has been identified- and the best-known of the hybrid functionals is Becke’s three-parameter formulation- B3LYP (Lee et al. 1988). The complete geometry optimization for final hits along with the known PDE4 inhibitors was performed using DFT with B3LYP, by employing a basis set of 6-31G* level at Gaussian suite 3. Vibrational frequencies were computed at the same B3LYP/6-31G* level to identify the stationary points on the corresponding potential energy surfaces (Arooj et al. 2013).
The orbital energies of the frontier orbitals, viz., Highest Occupied Molecular Orbital (HOMO) and Lowest Unoccupied Molecular Orbital (LUMO) were computed for all the final hits. These orbital energy values play a significant role in terms of electron donor and acceptor properties of a molecule and hence can be utilized to suitably understand the reactivity of a molecule to its receptor. In the present study, the energy gap, ∆E (HOMO–LUMO) was computed for all the potential hits and compared with that of known PDE4 inhibitors.
Chemical similarity search and design of structural analogs
Chemical Similarity Search was performed using selected lead molecules to identify novel scaffolds towards the inhibition of PDE4. The best hit with the minimum energy gap (LUMO–HOMO) was submitted to the PubChem database and compounds with > 90% similarity was retrieved. These compounds were then submitted to the SciFinder Scholar and PubMed databases to ascertain their novelty as PDE4 inhibitors. The best hit was selected to develop bioisosteres which would serve as more efficient PDE4 inhibitors. Thus a series of analogs were designed by replacing the reactive side chain of the best hit using molecular replacements acquired from the Swiss Bioisostere database (2012) (www.swissbioisostere.ch).This database is a compilation of information on 4.5 million molecular sub-structural replacements and their relevant data in biochemical assays. This has been sourced through detection of matching molecular pairs and through the process of mining bioactivity data in the ChEMBL database (Wirth et al. 2012). The entire library of designed analogs was then subjected to molecular docking analysis using MVD v6.0 software, and the selected analogs with the best interactions with the PDE4 binding site were proposed as PDE4 candidate inhibitors.
Results and discussion
The HypoGen algorithm was used to construct a quantitative hypothesis that correlated the experimental and the predicted activity values of the inhibitors. Ten hypotheses were developed based on the structurally diverse PDE4 inhibitors that were acquired from literature. The fixed cost, null cost, and cost difference values were predicted for all of these 10 hypotheses. Statistical parameters such as correlation coefficient (r) and RMSD for each hypothesis were also computed as shown in Table 1. The top scoring hypothesis i.e., Hypo 1 was finally considered for further processing, based on the cost analysis and correlation (r) values. Hypo 1 comprised of two HBA (Hydrogen bond acceptor) and three HY (Hydrophobic) features as shown in Fig. 2. Experimental and estimated activity values of the training set compounds as predicted by the best hypothesis Hypo1 have been listed in Table 2.
Hypo1 was further tested using a set of 60 compounds as compiled in Table 3. As shown in Fig. 3, correlation (r) values between the actual and predicted activities of training and test set compounds are 0.97 and 0.96, respectively. Fisher’s randomization test was further performed to evaluate the significance of Hypo1 based on the statistical validation. Nineteen spreadsheets as shown in Fig. 4 were generated to establish Hypo1 as a novel model. Significance of the hypothesis was calculated using the formula S = [1 − (1 + X)/Y] × 100, where X is the total number of hypotheses having a total cost lower than the original hypothesis, and Y is the total number of HypoGen runs (initial + random runs). Here, X = 0 and Y = (1 + 19), hence 95% = {1 − [(1 + 0)/(19 + 1)]} × 100
Hypo 1 was then employed to screen a large library of 4,60,695 compounds from six chemoinformatics databases. Drug-likeness filtration was performed prior to the pharmacophore screening. The best 50 hits from each of the six databases amounting to a total of 300 hits were further subjected to ADMET and carcinogenicity tests. These hits have been derived on the basis of maximum fit value to Hypo 1. Finally, 1243 compounds that were obtained through this screening were considered for molecular docking study against PDE4 receptors. MolDock (MVD) and LibDock (DS v4.1) were two algorithms used in molecular docking, and they further yielded a total of 19 compounds.
Two docking algorithms-each with a different approach to identify the binding orientations of selected hits with the PDE4 receptor model, were employed in this study. Molecular docking was performed for 1243 compounds which qualified optimum ADMET parameters from all the compounds chosen for the study. Thirty one compounds were identified based on their docking scores in comparison to both the co-crystal ligand (Piclamilast) and the most active compound in the training set. On the basis of the minimum H–Bond energy derived from the molecular docking analysis and the possible interactions with the amino acid residues at the binding site of PDE4, nineteen compounds were subjected to DFT-analysis.
DFT was employed to generate the HOMO and LUMO values which were calculated for the 31 compounds screened from the molecular docking study. According to Fukui’s frontier orbital approximation, the frontier orbitals HOMO and LUMO of a chemical species are crucial in defining its reactivity. The significance of frontier orbitals as principal constituents which influence the ease of chemical reactions and the stereoselective path was established by Fukui while Parr and Yang demonstrated that most frontier theories can be rationalized from DFT (Li and Evans 1995) In the present study, we have used the B3LYP/6-31G* basis set in order to estimate the orbital energy values of the predicted hits. The B3LYP, run with a 6-31G* or better basis set, is on average the best choice of a model chemistry for most systems. B3LYP (Becke 3-term correlation functional; Lee, Yang, and Parr exchange functional) /6-31G* is particular good for organic molecules, but less so for metal containing compounds.
In the current study, we have also compared the HOMO and LUMO energy values with known PDE4 inhibitors as presented in Table 4. The top hit was selected based on the minimum energy gap (ΔE, eV = HOMO–LUMO). Finally, six compounds (Fig. 5) were identified as potential PDE4 inhibitors based on the comparative inspection of the overall docking score, hydrogen bond interactions and the HOMO–LUMO energy gaps (Please also refer to Fig. S1 in supplementary data). The compound ZINC15273803, with a minimum band energy gap of −5.0425 eV, was accordingly identified to be the best hit and was proposed as a potential PDE4 inhibitor, along with the other identified compounds.
The compound ZINC15273803 was identified as the most potential PDE4 inhibitor on the basis of least band energy gap (HOMO–LUMO) (Fig. 6) along-with consideration of important criteria like mapping to Hypo1 and obtaining suitable docking scores and acceptable ADMET values. It was hence further submitted to the PubChem Database for chemical similarity search. Compounds with more than 90% chemical similarity to ZINC15273803 were retrieved and their drug-likeness parameters were computed the chemical structures of these are displayed in Fig. S3 and Supplementary File Table S1. A total of 200 analogs of ZINC15273803 were subsequently developed by the use of molecular fragment replacements provided by the Swiss Bioisostere database (Fig. S4). These were then subjected to molecular docking analysis using the Piclamilast binding site as reference to finally cull the selection down to six structural analogs of ZINC15273803 as potential PDE4 candidate inhibitors (Supplementary files Table S2, Fig. S4).
Conclusions
A ligand-based pharmacophore model was developed to elucidate the spatial arrangement of pharmacophore features necessary for the potential inhibition of type 4 phosphodiesterase (PDE4). A well-validated hypothesis (Hypo1) was generated using a training set of 18 known PDE4 inhibitors and its total cost, cost difference, RMSD and correlation values were derived to be 78.49, 93.94, 0.81, and 0.97, respectively. These pharmacophore parameters were compared with nine other hypotheses and Hypo1 was found to be the most suitable for 3D query in virtual screening of large chemical databases to identify novel hits and was thus employed for the same.
The Hypo1 model consisted of five features with two HBA and three HY bonds. The features mapping to the most and least active compounds with this model have been presented in Fig. 7a, b). Hypo1 also showed the highest fit value of 8.95 and was hence employed to estimate the inhibitory activities of 18 structurally diverse compounds to elucidate its predictive accuracy. Since it was able to predict the inhibitory activity values of training set compounds in the same order of magnitude, Hypo1 could be established as a novel hypothesis in the present study. It was also used for the prediction of a series of test set compounds consisting of 60 known PDE4 inhibitors. The correlation between actual and predicted activity derived by Hypo1 for all 60 compounds was found to be 0.96 and this value served as validation for the same. Fisher’s randomization was also applied to Hypo1 and nineteen random sheets were generated at the confidence level of 95%. The result of this test further corroborated that Hypo1 incorporated all necessary features to identify any potential PDE4 inhibitors.
Subsequently, Hypo1 was used to screen a large virtual chemical library which included two natural product databases, viz. Traditional Chinese Medicine and InterBioScreen. The preference of inclusion of natural product databases was based on the reported literature that drug candidates having a biological origin are generally seen to exhibit lesser side effects.
The 50 best compounds from each database, amounting to a total of 300 compounds, based on their maximum fit value to Hypo1 were further selected for molecular docking. Drug-likeness parameters such as fulfillment of Lipinski’s and Veber’s rules were used to filter the database compounds to a manageable library for subsequent studies. Docking was performed at the binding site of Piclamilast, a selective PDE4 inhibitor. This was done for a total of 1243 compounds which included the most active compounds of the training set. Protein molecules were prepared and optimized and Piclamilast was re-docked to validate its binding site. Thirty one compounds were selected based on their docking scores in comparison to both the co-crystal ligand (Piclamilast) and the most active compound in the training set. Finally, 19 compounds were chosen based on the minimum H–Bond energy and the possible interactions with the amino acid residues at the binding site of PDE4. Predicted compounds were found to interact with the potential neighboring amino acids of the Piclamilast binding active site viz., Met273, Leu319, Asn321, Phe372, Ile336, Phe340 etc (Fig. 8)
The orbital energies such as HOMO and LUMO were calculated for all of the 19 hits screened through molecular docking studies, Taking the energies of the known PDE4 inhibitors as reference, the calculated energy values of the potential PDE4 inhibitors were found to be higher than the reference compounds especially the HOMO energy. Comparison with PDE4 inhibitors like Apremilast and Roflumilast (Table 4) was indicative of the final hits as stable molecules at the PDE4 active sites. Further, the energy gap values between the HOMO and LUMO orbitals were calculated for all of the hits and ZINC15273803 was found to possess the lowest energy gap of −5.239 eV. This compound was thus proposed as the best PDE4 inhibitor in the current investigation. Moreover, the compound ZINC15273803 was also found to have optimal binding interactions in the PDE4 binding pocket (Fig. 8 and Supplementary Fig S2). ZINC15273803 was found to interact with Gln369 with a conventional Hydrogen bond while a van der Waal interaction was observed with Met357. Important Pi–Pi stacking, Pi–Pi T-shaped and Pi-Alkyl interactions with Phe340, Phe372, and Leu319 of PED4 protein were other significant molecular interactions of this compound with PDE4. These predicted interactions of ZINC15273803 with PDE4 further corroborates potential PDE4 the importance of the proposed hit as inhibitor. The summative virtual screening process to identify the best hit compound has been presented in Fig. 9.
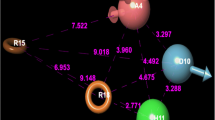
The six compounds forwarded by DFT analysis also mapped well with Hypo1 as presented in Fig. 10. Interestingly, out of these six hits, 4 compounds, viz., ZINC85852509, ZINC00662319, ZINC15273803, and ZINC15121039 are found to be natural products screened from the TCM database of Taiwan. The other two compounds, viz., CB7725955 and 93263702 belong to the ChemBridge database.
The DFT-based band energy gap of the compound ZINC15273803 was found to be minimum, which qualified it to be proposed as the best hit compound. Further, compounds with structural similarity to ZINC15273803 (>90%) too satisfied all drug-likeness parameters, hence establishing ZINC15273803 as a novel inhibitor of PDE4. Structural analogs of ZINC15273803 were further screened in order to identify its potency of interaction with PDE4 and six analogs (Ana11, Ana22, Ana125, A52, A125, and A161) were finally selected to be potential PDE4 inhibitors based on the LibDock Score (Supplementary file Fig S4 and Table 5). On the basis of the summative results above, we propose six known chemical compounds from the databases screened, and six structural analogs of the best hit ZINC15273803 to be novel and potential PDE4 inhibitors which can be further researched as potential drug candidates.
References
Arooj M, Sakkiah S, Kim S, Arulalapperumal V, Lee KW (2013) A combination of receptor-based pharmacophore modeling & QM techniques for identification of human chymase inhibitors. PLoS One 8(4):e63030
Arooj M, Thangapandian S, John S, Hwang S, Park JK, Lee KW (2011) 3D QSAR pharmacophore modeling, in silico screening, and density functional theory (DFT) approaches for identification of human chymase inhibitors. Int J Mol Sci 12(12):9236–9264
Azevedo MF, Faucz FR, Bimpaki E, Horvath A, Levy I, de Alexandre RB, Ahmad F, Manganiello V, Stratakis CA (2013) Clinical and molecular genetics of the phosphodiesterases (PDEs). Endocr Rev 35(2):195–233
Beghè B, Rabe KF, Fabbri LM (2013) Phosphodiesterase-4 inhibitor therapy for lung diseases. Am J Respir Crit Care Med 188(3):271–278
Brogi S, Kladi M, Vagias C, Papazafiri P, Roussis V, Tafi A (2009) Pharmacophore modeling for qualitative prediction of antiestrogenic activity. J Chem Inf Model 49(11):2489–2497
Buckley GM, Cooper N, Davenport RJ, Dyke HJ, Galleway FP, Gowers L, Haughan AF, Kendall HJ, Lowe C, Montana JG, Oxford J (2002a) 7-Methoxyfuro [2, 3-c] pyridine-4-carboxamides as PDE4 inhibitors: A potential treatment for asthma. Bioorg Med Chem Lett 12(3):509–512
Buckley GM, Cooper N, Dyke HJ, Galleway FP, Gowers L, Haughan AF, Kendall HJ, Lowe C, Maxey R, Montana JG, Naylor R (2002b) 8-Methoxyquinoline-5-carboxamides as PDE4 inhibitors: a potential treatment for asthma. Bioorg Med Chem Lett 12(12):1613–1615
Castro LR, Gervasi N, Guiot E, Cavellini L, Nikolaev VO, Paupardin-Tritsch D, Vincent P (2010) Type 4 phosphodiesterase plays different integrating roles in different cellular domains in pyramidal cortical neurons. J Neurosci Methods 30(17):6143–6151
Chong J, Leung B, Poole P (2013) Phosphodiesterase 4 inhibitors for chronic obstructive pulmonary disease. Cochrane Database Syst Rev 11:CD002309
Davies M, Nowotka M, Papadatos G, Dedman N, Gaulton A, Atkinson F, Bellis L, Overington JP (2015) ChEMBL web services: streamlining access to drug discovery data and utilities. Nucleic Acids Res 43:W612–W620
De Azevedo J, Walter F (2010) MolDock applied to structure-based virtual screening. Curr Drug Targets 11(3):327–334
Debnath AK (2002) Pharmacophore mapping of a series of 2, 4-diamino-5-deazapteridine inhibitors of Mycobacterium avium complex dihydrofolate reductase. J Med Chem 45(1):41–53
Devereux M, Popelier PL, McLay IM (2009) Quantum isostere database: a web-based tool using quantum chemical topology to predict bioisosteric replacements for drug design. J Chem Inf Model 49(6):1497–1513
Gilbert AM, Caltabiano S, Roberts D, Sum SF, Francisco GD, Lim K, Asselin M, Ellingboe JW, Kharode Y, Cannistraci A, Francis R (2000) Novel and selective calcitonin-inducing agents. J Med Chem 43(6):1223–1233
Jin SL, Ding SL, Lin SC (2012) Phosphodiesterase 4 and its inhibitors in inflammatory diseases. Chang Gung Med J 35(3):197–210
John S, Thangapandian S, Arooj M, Hong JC, Kim KD, Lee KW (2011) Development, evaluation and application of 3D QSAR pharmacophore model in the discovery of potential human renin inhibitors. BMC Bioinform 12(14):S4
Kansal N, Silakari O, Ravikumar M (2010) Three dimensional pharmacophore modelling for c-Kit receptor tyrosine kinase inhibitors. Eur J Med Chem 45(1):393–404
Kumar R, Son M, Bavi R, Lee Y, Park C, Arulalapperumal V, Cao GP, Kim HH, Suh JK, Kim YS, Kwon YJ (2015) Novel chemical scaffolds of the tumor marker AKR1B10 inhibitors discovered by 3D QSAR pharmacophore modeling. Acta Pharmacol Sin 36(8):998–1012
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B 37(2):785
Li Y, Evans JN (1995) The Fukui function: a key concept linking frontier molecular orbital theory and the hard-soft-acid-base principle. J Am Chem Soc 117(29):7756–7759
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23(1-3):3–25
Lugnier C (2006) Cyclic nucleotide phosphodiesterase (PDE) superfamily: a new target for the development of specific therapeutic agents. Pharmacol Ther 109(3):366–398
Lugnier C (2011) PDE inhibitors: a new approach to treat metabolic syndrome ? Curr Opin Pharmacol 11(6):698–706
Maurice DH, Ke H, Ahmad F, Wang Y, Chung J, Manganiello VC (2014) Advances in targeting cyclic nucleotide phosphodiesterases. Nat Rev Drug Discov 13(4):290–314
Mortelmans K, Zeiger E (2000) The ames salmonella/microsome mutagenicity assay. Mutat Res Fund Mol Mech Mut 455(1):29–60
Muller GW, Shire MG, Wong LM, Corral LG, Patterson RT, Chen Y, Stirling DI (1998) Thalidomide analogs and PDE4 inhibition. Bioorg Med Chem Lett 8:2669–2674
Niu M, Dong F, Tang S, Fida G, Qin J, Qiu J, Liu K, Gao W, Gu Y (2013) Pharmacophore modeling and virtual screening for the discovery of new type 4 cAMP phosphodiesterase (PDE4) inhibitors. PLoS One 8(12):e82360
Ochiai H, Ohtani T, Ishida A, Kishikawa K, Obata T, Nakai H, Toda M (2004) Orally active PDE4 inhibitors with therapeutic potential. Bioorg Med Chem Lett 14(5):1323–1327
Panchmatia PM, Ali ME, Sanyal B, Oppeneer PM (2010) Halide ligated iron porphines: a DFT + U and UB3LYP study. J Phys Chem A 114(51):13381–13387
Parker KA, Ressa M, Skelley S, Smith DK (1992) Community resources in obese care. J Fla Med Assoc 79(6):389–391
Provins L, Christophe B, Danhaive P, Dulieu J, Durieu V, Gillard M, Lebon F, Lengelé S, Quéré L, van Keulen B (2006) First dual M 3 antagonists-PDE4 inhibitors: synthesis and SAR of 4, 6-diaminopyrimidine derivatives. Bioorg Med Chem Lett 16(7):1834–1839
Richter W, Menniti FS, Zhang HT, Conti M (2013) PDE4 as a target for cognition enhancement. Exp Opin Ther Targets 17(9):1011–1027
Sakkiah S, Lee KW (2012) Pharmacophore-based virtual screening and density functional theory approach to identifying novel butyrylcholinesterase inhibitors. Acta Pharmacol Sin 33(7):964–978
Thomsen R, Christensen MH (2006) MolDock: a new technique for high-accuracy molecular docking. J Med Chem 49(11):3315–3321
Veber DF, Johnson SR, Cheng HY, Smith BR, Ward KW, Kopple KD (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45(12):2615–2623
Wirth M, Zoete V, Michielin O, Sauer WH (2012) SwissBioisostere: a database of molecular replacements for ligand design. Nucleic Acids Res 41(D1):D1137–D1143
Acknowledgements
Authors thankfully acknowledge Department of Biotechnology, Government of India for providing financial support for the Bioinformatics Infrastructure Facility (BIF) and the DBT e-library Consortium (DelCON) facility at the Centre for Biotechnology and Bioinformatics, Dibrugarh University in which the present work has been performed. The authors are also grateful to Dr. R. L. Bezbaruah, Chief Scientist (Retired) and Coordinator, BIF, CSIR-NEIST, Jorhat for his advice in the study. The Authors also thankful to Dr. Bulumoni Kalita, Assistant Professor, Department of Physics and Mr. Vishwa Jyoti Baruah, Assistant Professor, Centre for Biotechnology and Bioinformatics, Dibrugarh University for their valuable support in the present work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interests
The authors declares that they have no competing interests.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Gogoi, D., Chaliha, A.K., Sarma, D. et al. Identification of potential type 4 cAMP phosphodiesterase inhibitors via 3D pharmacophore modeling, virtual screening, DFT and structural bioisostere design. Med Chem Res 26, 3000–3014 (2017). https://doi.org/10.1007/s00044-017-1998-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-017-1998-3