Abstract
Wall shear stress induced remodelling of endothelial cell morphology is a focal cause of atherosclerosis. While the force distribution within endothelial cells has been quantified using computational modelling, no studies have included an image-informed cytoskeleton, nor have any studies examined the effect of population variation of the cytoskeleton. In this paper we quantified the spatial variation of the cytoskeleton and primary cilium in a population of endothelial cells to enable future mechanical analysis.
This was achieved using custom spatial descriptors for the microtubule and actin filament networks and the primary cilium. Using these descriptors our quantitative findings are in close agreement with previous findings of actin distribution (occupying a planar layer in the cell <20% of total cell height, at the periphery in the xy plane), and microtubule distribution (median of 38 microtubules per cell, in a tree-like network with branching tubules within 30° of the parent tubule).
Our set of descriptors improve on previous cell spatial descriptors by representing specific cells with sufficient accuracy to enable current modelling approaches. Additionally, the descriptors enable the generation of virtual cells with a cytoskeleton and primary cilium characteristic of the entire population. Thus reducing the computational cost needed to represent the entire population, without significant information loss.
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
Keywords
- Endothelial mechanotransduction
- Atherosclerosis
- Cell morphology
- Spatial descriptors
- Population morphology
- Cytoskeleton
1 Introduction
Blood-flow induced wall shear stress (WSS) is a focal risk factor for atherosclerosis [1]. Disturbed oscillatory WSS elicits an inflammatory endothelial cell (EC) response that contributes to the pathogenesis of atherosclerosis. Conversely, steady WSS results in a protective EC response [2]. A key EC response to WSS is the change in cell morphology: in particular, cytoskeletal morphology. Steady WSS results in cell elongation and alignment in the flow direction. In contrast, disturbed WSS causes greater shape variation in ECs (similar to cells grown in static conditions) [3,4,5,6,7,8,9].
Chien postulated that EC morphological dependency on WSS is caused by ECs maintaining an internal force homeostasis: cells adapt their morphology to keep internal forces within a “normal” range. In the case of disturbed WSS, the preferential flow direction cannot be sensed. This leads to internal force extremes causing endothelial dysfunction and an inflammatory response [10].
Computational models have already been used to quantify the stresses and strains within a single EC [11] or a population of ECs [12] in response to WSS (see [13] for a recent review). However, the effect of EC morphology on internal force transmission of WSS has only recently been examined. In particular, the effect of morphological variation in the nucleus and cell membrane on the internal force response to WSS has been quantified [14]. Further extension is required to include the cytoskeleton—a key component in cell mechanical stability [15]. One barrier to this is the lack of quantitative spatial descriptors (also known as shape/morphometric/summary descriptors) to define cytoskeletal morphology. Once this challenge has been overcome, it will be possible to test Chien’s theory by quantifying the internal force distributions in populations of ECs with differing morphologies that are exposed to different WSS environments.
In this study, we have extended the spatial descriptor analysis to include the cytoskeleton and primary cilia. These descriptors were then used to recreate specific cells. Furthermore, our set of spatial descriptors were used to generate virtual cells characteristic of the entire population. Thus, the morphological variation of the entire population can be represented by a smaller set of virtual cells without a significant loss in information.
Our spatial descriptor analysis aims to provide a spatial domain suitable for future mechanical analysis. In doing so, we hope to enable future studies on the role of EC mechanics in atherosclerosis.
2 Materials and Methods
Human microvascular endothelial cells (HMEC-1s) were cultured using previously described methods [14, 16]. The primary cilia, acetylated α-tubulin, F-actin and the nuclei from 15 HMEC-1s were then co-imaged as previously described [14, 16].
2.1 Spatial Descriptors
The framework for spatial descriptors is based on the series of studies published by Murphy et al. [17,18,19].
Reference system, nucleus and cell membrane shape: The reference coordinate system of our model has previously been described [14]. Briefly, the nucleus centroid is the origin of coordinate system, the long axis of the nucleus forms the first axis, the apical-basal direction forms the second axis and the 3rd axis is perpendicular to the other two. Nuclei are chosen as the reference point as they are easily identified, and there is only one in every EC. The spatial descriptors of the nucleus and cell membrane have also been previously described [14]. Image processing and analysis was carried out in MATLAB (version 2013b), ImageJ (version 1.48o) and AMIRA (version 5.6).
Cytoskeleton—deconvolution: The acetylated α-tubulin and F-actin components of cytoskeleton were imaged and the morphology quantified. As a preliminary step, deconvolution was carried out to increase contrast of the cytoskeletal structures, thus allowing automatic segmentation. Image stacks of actin and tubulin were deconvolved with an artificial point spread function (PSF) calculated using the Diffraction PSF 3D ImageJ plugin (http://www.optinav.com/Diffraction-PSF-3D.htm, [20]). Deconvolution was then carried out using the Iterative Deconvolve 3D ImageJ plugin (http://www.optinav.com/Iterative-Deconvolve-3D.htm, [20]).
F-actin: Each individual actin fibre was traced semi-automatically using the simple neurite tracer tool in ImageJ [21], hence the 3D coordinates of each fibre (at pixel resolution) were quantified. We found that nearly all stress fibres were within two slices (0.9 μm layer), in a layer one slice (0.45 μm) below the base of the nucleus. Hence the morphology of the actin fibre network can be captured by a 2D analysis. Five spatial descriptors were used to represent F-actin (see Fig. 1). The first is the number of actin fibres present in each cell, n f-actin. All other F-actin spatial descriptors are arrays of size n f-actin. The other descriptors are the fibre length, L f-actin; the fibre orientation with respect to the nuclear axis, θ f-actin; the length between the centroid of the fibre and the nucleus centroid, L actin-cent and angle of the vector between actin fibre centroid and the nucleus centroid, θ actin-cent.
Spatial descriptors used to quantify F-actin morphology: (a) n f-actin, the number of actin fibres present in each cell, in this simplified example there are only three fibres (blue); (b) L f-actin, fibre length; (c) θ f-actin, fibre orientation with respect to the nuclear axis; (d) L actin-cent, the length between the actin fibre centroid and the nucleus centroid (shown in green) and (e) θ actin-cent, angle of the vector between actin fibre centroid and the nucleus centroid
Acetylated α-tubulin: Acetylatedα-tubulin migrates outward from the centrosome, and therefore can be modelled as a branching network, with straight segments. The centrosome was not explicitly imaged, but instead was estimated to be the voxel with maximum intensity in the tubulin images [22]. Again, the simple neurite tracer tool [21] was used to segment the acetylated a-tubulin morphology. The six spatial descriptors used to quantify α-tubulin morphology were: (1) the position of the centrosome with respect to the nucleus centroid; (2) number of microtubules in the cell, n tubule (similar to F-actin, the remaining descriptors are arrays of size n tubule); (3) the length of each microtubule, L tubule; (4) the collinearity of a microtubule against its parent microtubule (the microtubule that it branched out from), θ cotubule; (5) the change in height (z value) of the centroid microtubule compared to the height of the parent microtubule and (6) the ancestry number of the microtubule, which identifies the number of its parent microtubule. These descriptors are illustrated in Fig. 2.
Spatial descriptors used to quantify acetylated α-tubulin morphology: (a) the position of the centrosome relative to nuclear centroid; (b) number of microtubules in the cell, n tubule; (c) the length of each microtubule, L tubule; (d) the collinearity of a microtubule with its parent microtubule, θ cotubule, note this consists of an in-plane angle and a z-direction change in slice and (e) the ancestry number of the microtubule, which identifies the number of its parent microtubule. In this example the ancestry numbers are 0, 0, 2, 2, indicating that the first two microtubules originate at the centrosome, and the third and fourth tubules originate from the second microtubule
Primary cilium: Due to spectral overlap, a separate population (n = 39), from the population that was used in cell shape, nucleus and cytoskeleton analysis, of cells was used to examine primary cilium morphology using methods described in Lim et al. 2015 [16]. To avoid bias, every cilium in each image was analysed. Two spatial descriptors were formulated for primary cilia, cilia length and cilia-centroid position—the vector between the nucleus centroid and the base of the cilium.
2.2 Generating Virtual Cells
Virtual cells were generated from the complete population dataset of previously described morphological descriptors. Statistical boot-strapping was used to obtain a distribution for each of the descriptors: first, the n-sized vector (where n is n f-actin or n tubule, respectively) of every spatial descriptor was resampled with replacement, to generate 1000 n-sized vectors. This large dataset is normal distributed. The F-actin and tubulin networks and primary cilia could then generated by sampling from these distributions. As the nucleus centroid is the reference point in the set of tubulin, actin and primary cilia descriptors, it is possible to generate a virtual cell with all of these components.
3 Results
3.1 F-Actin
The median of n f-actin was 20 fibres, with a maximum of 32 and a minimum of ten. There were fewer actin stress fibres than microtubules on average. L f-actin is right skewed, with a median value of 4.6 μm (see Fig. 3a). From the angle histogram of θ f-actin (see Fig. 3b) there does not appear to be a preferential angle of orientation of actin fibres. Median L actin-cent was 13 μm (see Fig. 3c), suggesting that the majority of the actin stress fibres were beyond the xy area bounded by the nucleus. From the angle histogram of θ actin-cent (see Fig. 3d), it appears that the centroid of actin fibres are preferentially to the left and right (270° and 90°) of the nucleus centroid, rather than above–below (0° and 180°) the nucleus centroid (nucleus axis is parallel to the above–below direction).
Quantification of spatial descriptors of F-actin in the population. (a) The histogram of L f-actin shows L f-actin is right skewed and not normal (Shapiro–Wilk test, p < 0.0001). (b) The angle histogram of θ f-actin with units in degrees: no preferential direction is evident. (c) The histogram of L actin-cent. L actin-cent is not normal, (Shapiro–Wilk test, p < 0.0001). Most of the values of L actin-cent are above 5 μm, suggesting that actin fibres are far from the nucleus. (d) The angle histogram of θ actin-cent. Fibres are preferentially to the left and right (270° and 90°) of the nucleus centroid, rather than above–below (0° and 180°) the nucleus centroid
3.2 Acetylated α-Tubulin
The distance in the xy plane between the centrosome and the nucleus centroid for all cells in the population are illustrated in Fig. 4a, with the nucleus length and width of that cell for comparison. The median of n tubule was 38, with a maximum of 71 and minimum of 15. L tubule was right skewed, with a median value of 5.9 μm (see Fig. 4b). The median of θ cotubule was 3°, with the majority of microtubules within 30° of the parent microtubule (see Fig. 4c). The ancestry number and change in height are not shown here, but are stored to allow specific cells to be recreated. The actin filament and microtubule networks of a specific cell can also be recreated from its spatial description (see Fig. 5). The methods presented in this study result in straight actin filaments and straight microtubules. Nearly all actin filaments are captured, whereas the finer microtubule network cannot be resolved with our imaging technique. Note, these recreated images are in 2D. While actin filaments are mostly in plane, the microtubule network is not, hence further development is needed to extend these recreated models to 3D.
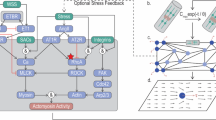
Quantification of spatial descriptors of acetylated α-tubulin in the population. (a) Column graph of distance between centrosome and nucleus (blue), half of nucleus length (orange) and half of nucleus width (grey), inset: cartoon cell illustrating these lengths with nucleus in green. The centrosome distance is shown by the dashed blue circle. (b) The histogram of L tubule shows microtubule length is right skewed and not normal (Shapiro–Wilk test, p < 0.0001). (c) Angle histogram of θ cotubule, with units in degrees. The majority of microtubules are within 30° of their parent microtubule
Actin filament (a–c) and microtubule (d–f) morphology in a specific cell can be regenerated directly from its spatial descriptors. (a) F-actin (pink) and nucleus (red) in a HMEC-1. (b) Actin filament network recreated from spatial descriptors described in Fig. 1. Consisting of actin filaments (blue), nucleus axis (red) and filament centroids (blue crosses). (c) Overlap of the original image with the recreated geometry. While the recreated filaments are straighter than they appear in the original image, there is a close resemblance between the two. (d) Image slice of acetylated α-tubulin in a HMEC-1. The nucleus position is evident, as microtubules form a cage structure around it. (e) The geometry of the microtubule network (in 2D), recreated from spatial descriptors described in Fig. 2. The nucleus axis is shown in red, with the microtubules in blue. (f) Overlap of the original image with the recreated geometry. Similarly, recreated microtubules are straighter than the original image, and only the larger, thicker microtubules are detected
3.3 Primary Cilium
A histogram of primary cilium length is shown in Fig. 6a. The median ± interquartile range of primary cilium length was 3.0 μm ± 1.0 μm. The position of the base of the cilia relative to the nucleus centroid is shown in Fig. 6b. There does not appear to be a directional pattern in cilia location.
Spatial descriptors of the primary cilium. (a) The histogram of primary cilium length (n = 39). The median ± interquartile range of primary cilium length was 3.0 μm ± 1.0 μm. (b) A scatter plot of the xy position of the base of the cilia with respect to the nucleus centroid, (units are normalised by nucleus length of each individual cell). Orange circle indicates a half nucleus length away from nucleus centroid. Inset: cartoon of cell giving context to the orange circle. The base of the cilium is generally very close to the nucleus centroid, with the majority within half a nucleus length (of that specific cell). There does not appear to be a predominant direction, with respect to the nucleus axis
3.4 Generating Virtual Cell Components
To illustrate the ability to generate virtual cell components using our method, three 2D models of a single cell composed of a nucleus, cilium and cytoskeleton were created (see Fig. 7). These have a cytoskeletal network and primary cilium typical of the overall population of ECs.
Virtual cells generated using morphological descriptors, containing: actin stress fibres (green lines, with crosses at the centroid position); the microtubule network (in red), emerging from the centrosome (black circle) and the primary cilium (length, indicated, emerging from the centrosome). An idealised nucleus was also added (blue dashed oval), with length and width of the average nucleus calculated in [14]. Furthermore, the cells are enclosed by a cell membrane based on the smallest shape that can enclose the subcellular components
These virtual models are at a preliminary stage. Further extension is needed to examine the co-dependency of spatial descriptors. For instance, the nucleus position and tubulin network are likely to be co-dependent, as the tubulin forms a cage around the nucleus. As this dependency is not measured in our current analysis, the virtual cells shown in Fig. 7 have a cage like tubulin network that is independent of nucleus position. Other less obvious co-dependencies may exist, and are best tested for using a regression analysis. Further extension is also required to generate 3D models.
4 Discussion and Conclusions
In this paper, we have quantified the morphological variation of the EC cytoskeleton and primary cilium using newly developed spatial descriptors. Our findings are in close agreement with a number of earlier studies: We found that actin occupies a planar layer in the cell (15–18% of the total cell height), which is in close agreement with Galbraith et al. finding of 20% (Fig. 7, from [5]). We also agree with the findings of Chiu et al. [3] that actin stress fibres tend to localise at the periphery of the cells. Our estimates for number of microtubules (median of 38) and collinearity (median of 3°, with the majority of values within 30°) is within the range of the findings of Li et al. [23] for a number of different cancer cell lines.
We found that centrosomes do not have to appear to have a directional bias in static cells (they are equally likely to occur at any angle around the nucleus centroid with respect to the nucleus centroid). Furthermore, we found that centrosomes occur near the nucleus centroid. Both these findings are in agreement with the qualitative findings of Galbraith et al. (Fig. 1a, t = 0; [5]). Furthermore, the base of the primary cilium is closely associated with centrosomes [24]: we also found a lack of preferential direction of the base of the cilia with respect to the nucleus centroid and nucleus axis. This is in agreement with Galbraith et al.’s finding that the centrosomes in static cells are distributed evenly upstream and downstream of the cell centroid (Fig. 1b, t = 0; [24]). Our estimated HMEC-1 primary cilium length of 3.0 μm is within the reported range of primary cilia lengths in other EC types [25,26,27].
Our spatial descriptor analysis is an improvement over existing descriptors because it can represent the geometry of a particular cell (Fig. 5) with sufficient accuracy to allow mechanical modelling: the most sophisticated cytoskeletonised EC models to date only represent the larger elements of the cytoskeleton as straight elements (n < 50) [12, 28], which our approach captures. Furthermore, our approach allows the creation of virtual cytoskeletonised cells that are characteristic of the entire population. Although at a preliminary stage, it is hoped that this virtual cell approach will reduce the computational cost of analysing the mechanics of an entire cell population.
We are in the process of developing a tensegrity-based mechanical cell model using the morphological data of the cytoskeleton obtained in this study. Our model consists of custom geometrically non-linear truss elements based on those developed by Crisfield [29, 30], modified to incorporate a prestress. A load controlled Newton–Raphson method is used to solve the static analysis of this model [31]. This modelling framework is suitable for modelling a cytoskeleton. Furthermore, we will use this model together with force-relaxation/force-minimisation methods [32,33,34] to determine force equilibrium. Forces acting on the membrane can be calculated using existing continuum methods [14] and mapped on to the nearest nodes in the pseudo-tensegrity model. It is anticipated that preliminary results of our model will be presented at the conference.
References
Davies PF, Polacek DC, Shi C, Helmke BP (2002) The convergence of haemodynamics, genomics, and endothelial structure in studies of the focal origin of atherosclerosis. Biorheology 39(3):299–306
Davies PF (2009) Hemodynamic shear stress and the endothelium in cardiovascular pathophysiology. Nat Clin Pract Cardiovasc Med 6(1):16–26
Chiu J, Wang DL, Chien S, Skalak R, Usami S (1998) Effects of disturbed flow on endothelial cells. J Biomech Eng 120(1):2–8
Davies PF, Remuzzi A, Gordon EJ, Dewey CF, Gimbrone MA (1986) Turbulent fluid shear stress induces vascular endothelial cell turnover in vitro. Proc Natl Acad Sci U S A 83(7):2114–2117. doi:10.1073/pnas.83.7.2114
Galbraith CG, Skalak R, Chien S (1998) Shear stress induces spatial reorganization of the endothelial cell cytoskeleton. Cell Motil Cytoskeleton 40(4):317–330
Helmlinger G, Geiger RV, Schreck S, Nerem RM (1991) Effects of pulsatile flow on cultured vascular endothelial cell morphology. J Biomech Eng 113(2):123–131
Levesque MJ, Sprague EA, Schwartz CJ, Nerem RM (1989) The influence of shear stress on cultured vascular endothelial cells: the stress response of an anchorage-dependent mammalian cell. Biotechnol Prog 5(1):1–8
Nerem RM, Levesque MJ, Cornhill JF (1981) Vascular endothelial morphology as an indicator of the pattern of blood flow. J Biomech Eng 103(3):172–176
Rouleau L, Farcas M, Tardif J, Mongrain R, Leask RL (2010) Endothelial cell morphologic response to asymmetric stenosis hemodynamics: effects of spatial wall shear stress gradients. J Biomech Eng 132(8):081013–081013
Chien S (2007) Mechanotransduction and endothelial cell homeostasis: the wisdom of the cell. Am J Physiol Heart Circ Physiol 292(3):H1209–H1224
Ferko M, Bhatnagar A, Garcia M, Butler PJ (2007) Finite-element stress analysis of a multicomponent model of sheared and focally-adhered endothelial cells. Ann Biomed Eng 35(2):208–223
Dabagh M, Jalali P, Butler PJ, Tarbell JM (2014) Shear-induced force transmission in a multicomponent, multicell model of the endothelium. J R Soc Interface 11(98):20140431
Lim YC, Cooling MT, Long DS (2014) Computational models of the primary cilium and endothelial mechanotransmission. Biomech Model Mechanobiol 14:665–678
Lim YC, McGlashan S, Cooling MT, Long DS (2016) Mechanical models of endothelial mechanotransmission based on a population of cells. In: Joldes GR, Doyle B, Wittek A, PMF N, Miller K (eds) Computational biomechanics for medicine. Springer, New York
Chen TJ, Wu CC, Su FC (2012) Mechanical models of the cellular cytoskeletal network for the analysis of intracellular mechanical properties and force distributions: a review. Med Eng Phys 34(10):1375–1386
Lim YC, McGlashan S, Cooling MT, Long DS (2015) Culture and detection of primary cilia in endothelial cell models. Cilia 4(1):11
Buck TE, Li J, Rohde GK, Murphy RF (2012) Toward the virtual cell: automated approaches to building models of subcellular organization “learned” from microscopy images. BioEssays 34(9):791–799
Peng T, Murphy RF (2011) Image-derived, three-dimensional generative models of cellular organization. Cytometry A 79(5):383–391
Zhao T, Murphy RF (2007) Automated learning of generative models for subcellular location: building blocks for systems biology. Cytometry A 71(12):978–990
Dougherty R (2005) Extensions of DAMAS and benefits and limitations of deconvolution in beamforming. Am Inst Aeronaut Astronaut. doi:10.2514/6.2005-2961
Longair MH, Baker DA, Armstrong JD (2011) Simple neurite tracer: open source software for reconstruction, visualization and analysis of neuronal processes. Bioinformatics 27(17): 2453–2454
Shariff A, Murphy RF, Rohde GK (2011) Automated estimation of microtubule model parameters from 3-D live cell microscopy images. In: Anonymous biomedical imaging: from nano to macro, 2011 IEEE international symposium on, p. 1330
Li J, Shariff A, Wiking M, Lundberg E, Rohde GK, Murphy RF (2012) Estimating microtubule distributions from 2D immunofluorescence microscopy images reveals differences among human cultured cell lines. PLoS One 7(11):e50292
Hagiwara H, Ohwada N, Aoki T et al (2008) The primary cilia of secretory cells in the human oviduct mucosa. Med Mol Morphol 41(4):193–198
Iomini C, Tejada K, Mo W et al (2004) Primary cilia of human endothelial cells disassemble under laminar shear stress. J Cell Biol 164(6):811–817
Van der Heiden K, Groenendijk BCW, Hierck BP, Krams R, de Com R, Cheng C, Baiker M, Pourquie MJBM, Alkemade FE, deRuiter MC, Gittenberger-de Groot AC, Poelmann RE (2006) Monocilia on chicken embryonic endocardium in low shear stress areas. Dev Dyn 235(1):19–28
Geerts WJC, Vocking K, Schoonen N, Haarbosch L, van Donselaar EG, Regan-Klapisz E, Post JA (2011) Cobblestone HUVECs: a human model system for studying primary ciliogenesis. J Struct Biol 176(3):350–359
Khayyeri H, Barreto S, Lacroix D (2015) Primary cilia mechanics affects cell mechanosensation: a computational study. J Theor Biol 379:38–46
Crisfield MA (1991) Non-linear finite element analysis of solids and structures volume 1: essentials. Wiley, New York
Crisfield MA (1997) Non-linear finite element analysis of solids and structures volume 2: advanced topics. Wiley, New York
Zienkiewicz OC, Taylor RL (2000) The finite element method: the basis. Butterworth-Heinemann, Oxford
Bel Hadj Ali N, Rhode-Barbarigos L, Smith IFC (2011) Analysis of clustered tensegrity structures using a modified dynamic relaxation algorithm. Int J Solids Struct 48(5):637–647
Estrada GG, Bungartz H-J, Mohrdieck C (2006) Numerical form-finding of tensegrity structures. Int J Solids Struct 43(22–23):6855–6868
Paul C, Lipson H, Cuevas FJV (2005) Evolutionary form-finding of tensegrity structures. In: GECCO ‘05 Proceedings of the 7th annual conference on Genetic and evolutionary computation on, pp. 3–10
Ades EW, Candal FJ, Swerlick RA, George VG, Summers S, Bosse DC, Lawley TJ (1992) HMEC-1: establishment of an immortalized human microvascular endothelial cell line. J Investig Dermatol 99(6):683–690
Acknowledgments
HMEC-1s were kindly provided by Dr. Edwin Ades, Mr. Francisco J. Candal (CDC, Atlanta GA, USA) and Dr. Thomas Lawley (Emory University, Atlanta, GA, USA) NCEZID-R147589-00 [35]. The authors would also like to acknowledge Dr. Sue McGlashan, Ms. Hilary Holloway and Ms. Jacqui Ross from the Biomedical Imaging Research Unit, University of Auckland for assistance in microscope training and image acquisition. This work was supported by a University of Auckland Faculty Research Development Fund grant (3702516, D.S.L.). The first author is grateful for financial support from the University of Kassel.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this paper
Cite this paper
Lim, Y.C., Kuhl, D., Cooling, M.T., Long, D.S. (2017). Quantifying Cytoskeletal Morphology in Endothelial Cells to Enable Mechanical Analysis. In: Wittek, A., Joldes, G., Nielsen, P., Doyle, B., Miller, K. (eds) Computational Biomechanics for Medicine. Springer, Cham. https://doi.org/10.1007/978-3-319-54481-6_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-54481-6_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-54480-9
Online ISBN: 978-3-319-54481-6
eBook Packages: EngineeringEngineering (R0)