Abstract
Melanocytes are derived from the neural crest and enter the eye during embryogenesis. Uveal melanoma of the eye is a rare but deadly disease. About 50% of patients will eventually develop metastatic disease with an inevitable fatal end. Predisposing factors are race, skin and hair color, and familial tumor predisposition syndromes.
Although clinically uveal melanoma phenotype gives the impression of one disease, genetically uveal melanoma can be classified into at least two subgroups which can be distinguished by DNA-based and mRNA-based technologies. While patients with disomy 3 and gene expression profile class 1 have only a low risk of developing metastatic disease, patients with monosomy 3 and/or class 2 gene expression profile are likely to die from metastases. In addition to prognostic information, genetic testing also provides new insights into molecular pathobiology of uveal melanoma. Mutations in GNAQ, GNA11, and BAP1 have been found to be the crucial steps in tumor development.
Those insights raise the hope for targeted therapies and improved prognoses for uveal melanoma patients in the near future.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
9.1 Background
Melanocytes are found ubiquitously in the eye and periocular region. It is our current understanding that during embryogenesis pigment cells derived from the neural crest migrate along the nerve sheaths of the trigeminal branch V1 and reach the eye via the branches of the ciliary nerves. Still being melanoblasts, those cells enter the eye close to the optic disc to be distributed throughout the uvea—the choroid, the ciliary body, and the iris—where they mature and become melanocytes [1]. Disturbance of this process of maturation may lead to proliferation of melanocytic cells. They appear benign, round, or oval in shape if the proliferation started after the cells had reached their final destination. The lesion may affect a large sector of the iris and choroid or even the periocular skin if proliferation began early in posteroanterior migration, at a central branch of the trigeminal nerve (e.g., nevus of Ota) [2, 3]. Additional genetic alterations acquired later in life may cause further growth and increased proliferation rate and finally result in the development of a malignant uveal melanoma. Over the past decade, our understanding of the molecular and genetic mechanisms which deregulate the cell cycle of melanocytes and eventually turn nevi into melanomas has increased significantly. Many researchers and research groups contribute to elucidation of the pathogenesis of uveal melanoma, with the ultimate goal of finding a cure for or even prevent this disease.
In the following chapter, we present the current status of research in the molecular carcinogenesis of uveal melanoma, show how the clinical appearance results from molecular and genetic mechanisms, and demonstrate the clinical impact which might result from those. For readers who are not experts in ophthalmic oncology, we will give a brief general overview of the disease, as far as this is necessary for the understanding.
9.2 Epidemiology of Uveal Melanoma
The incidence of uveal melanoma varies among different races and ethnicities over the world. An analysis of the SEER (Surveillance, Epidemiology, and End Results) program database in the United States covering 36 years, from 1973 to 2008, revealed a mean age-adjusted incidence of 5.1 cases per million per year [4]. However, the majority of cases (97.8%) were observed in the white population. The ratio of black population-Asian population-Hispanic population-non-Hispanic white population is 1:1.2:5:19 (SEER program data, 1992–2000) [5]. Whereas the incidence rate was not significantly different between the registries in the United States, an analysis of the data from the European Cancer Registry-based (EUROCARE) study, which combined the data from 67 individual European cancer registries over the years 1983–1994, showed a dependency on latitude. The incidence decreased from north to south, with eight cases per million per year in Scandinavia and only two cases in the south of Europe (Spain and Southern Italy) [6, 7]. In comparison to cutaneous melanoma, with an incidence of 21.8 cases per 100,000 men and women per year, uveal melanoma appears rare (data from SEER Stat Fact Sheets 2016). Only 5% of all melanomas arise from the eye [8]. However, uveal melanoma is still the most common primary intraocular malignancy in adults [9].
9.3 Predisposing Factors
Predisposing factors can help elucidate the genetic mechanisms underlying carcinogenesis. Epidemiological, familial, clinical, and occupational factors contribute to tumor development. The incidence of uveal melanoma increases up to an age of 70 years, and the incidence peaks at 24.5 cases per million males and 17.8 cases per million females per year in the United States, resulting in a mean age at diagnosis of 60 years [6]. Overall, sex does not seem to be a predisposing factor. Data from the EUROCARE study and from the United States demonstrate that race (see the above), skin and eye color, and the ability to tan are the main predisposing factors [4, 7, 10]. This suggests a possible role of UV radiation in the carcinogenesis of uveal melanoma [11,12,13]. Though this hypothesis is still under discussion and there is no good evidence available, the preferred occurrence of iris melanoma in the inferior half of the iris and the higher incidence of choroidal melanomas at the posterior pole of the eye could be explained by a higher exposure to UV light in those locations. A cohort effect found in the data from the EUROCARE study, with a higher incidence of uveal melanoma in the birth cohorts from 1910 to 1935, might be interpreted that way, as changes in the profiles of work and occupational UV light exposure and the use of sunglasses could explain the decrease in incidence in the younger cohorts [7].
Familial uveal melanoma: Occurrence of uveal melanoma in more than one family member is a rare event, accounting for only 0.6% of patients [14]. However, several cases have been reported, dating back until 1905 [15]. In the published cases, no pattern of inheritance could be found, and only few individuals were affected in each family [14,15,16]. Other features of a genetic predisposition, such as early age at diagnosis, bilateral involvement, or phenotypic associations were not present [17]. Therefore, it might be possible that two individual family members might be affected by chance alone, though the likelihood of such an occurrence is small [18]. On the other hand, uveal melanoma has been reported to occur more frequently in patients showing oculodermal melanocytosis, familial atypical mole and melanoma syndrome, neurofibromatosis, Li-Fraumeni syndrome, or germline BAP1 mutations, which all have a hereditary and therefore familial background.
Oculodermal melanocytosis: Patients with oculodermal melanocytosis present with congenital increased pigmentation within the distribution of the first and/or second branch of the trigeminal nerve. The affected organs include not only the periocular skin/eyelid, episclera/sclera, uvea, and orbit but also the meninges and tympanic membrane. Patients with oculodermal melanocytosis have a significantly increased risk for uveal melanoma, with an estimated lifetime risk of 1:400 for developing uveal melanoma [19]. It remained unclear whether the increased risk for uveal melanoma results simply from the increased number of melanocytes in the uvea or from an underlying predisposing condition. The large affected sector of the trigeminal nerve points toward an early event in melanogenesis, altering melanoblasts before final maturation [3]. The frequent occurrence of mutations in GNAQ found in intradermal melanocytic proliferations/blue nevi and uveal melanomas appears to drive melanocytic proliferation and might explain the increased risk of malignant transformation [20].
Familial atypical mole and melanoma syndrome (FAM-M): According to the NIH consensus conference, the FAM-M syndrome is diagnosed in individuals with multiple atypical cutaneous nevi, showing distinct histological features and with cutaneous melanoma in one or more first- or second-degree relatives [21]. Dysplastic or atypical nevi of the skin may also occur isolated and are associated with an increased risk of cutaneous melanoma [22]. However, the FAM-M syndrome is a hereditary syndrome, caused by a mutation of the CDKN2A gene, coding for the INK4a and ARF proteins, which regulate the cell cycle and act as tumor suppressors. Germline mutations in CDKN2A are associated with an increased risk ofcutaneous melanoma, glioblastoma, and pancreatic cancer [23, 24]. Several case series suggested a role for CDKN2A and the FAM-M syndrome in the development of uveal nevi and melanoma. Uveal nevi and uveal melanomas seem to occur more frequently in patients with FAM-M syndrome. And the FAM-M syndrome has been observed more frequently in patients with uveal melanoma [25,26,27,28,29,30,31,32]. Despite this clinical evidence, the connection between FAM-M is still not fully understood and unproven, as CDKN2A mutations do not seem to play a role in uveal melanoma development and/or progression [33,34,35,36].
Neurofibromatosis-1 is a frequent autosomal dominant inherited genetic disorder affecting about 1 in 3000 individuals [37]. Because of the high prevalence, repeated cases of uveal melanoma in NF 1 patients may be coincidental. A connection has been suggested because half of uveal melanomas show reduced expression of NF1 tumor suppressor gene causing an increased activity of Ras and of MAPK activation [38].
Li-Fraumeni syndrome is an autosomal dominant inherited cancer predisposition syndrome caused by a germline p53 mutation [39].
BAP1 mutation: Germline mutations of BAP1 cause a hereditary tumor predisposition syndrome, and affected individuals frequently develop uveal melanoma, lung adenocarcinoma, mesothelioma, and meningioma [40,41,42]. The spectrum and number of tumors vary between individuals and the affected kindreds, and several cases of familial uveal melanoma have been associated with BAP1 mutations [43, 44]. The reduced penetrance of the germline mutation, however, can impede the identification of those patients as familial cases. Somatic mutations of BAP1 have been identified in metastasizing uveal melanoma and are thought to be a critical step for development of an aggressive tumor phenotype [45]. In case of a germline BAP1 mutation, loss of the wild-type allele of BAP1 on the remaining allele on chromosome 3 will promote melanoma development. The role of BAP1 mutations and the presumed genetic mechanisms are described later in this chapter.
9.4 Prognosis
Uveal melanoma is generally reported to have an overall chance of 50% for spreading to the liver and cause metastatic disease, which is usually fatal within a few months. However, for counselling patients, this information is not very helpful. It has been shown that for most patients, the prognosis is actually much better or much worse and that we need to obtain an individualized risk, by combining clinical, histological, and genetic risk factors [46].
Clinical factors: Older age and male gender have been associated with reduced survival. However, there remains the possibility that those results are biased by a delay in diagnosis in older patients, a higher general mortality rate in older patients (competing risks), and prolonged survival of younger patients with metastatic disease compared to older ones [47,48,49,50].
Histopathological factors: Uveal melanoma is commonly classified on cytomorphology according to a classification scheme proposed by Callender in 1931 and modified by McLean in 1983 [51]. Cells are divided into fusiform spindle cells and the larger polygonal and pleomorphic epithelioid cells. Both cell types may be present in the same tumor, which is then classified as “mixed cell type.” Epithelioid cell type has been associated with a higher metastasis rate [52]. Problems with the classification lie in the significant inter- and intraobserver variability and the missing consensus on how many epithelioid cells must be present for a melanoma to be classified as “mixed” or “epithelioid” [53]. Cytological tumor heterogeneity here indicates an underlying genetic heterogeneity. It has been shown by several authors that epithelioid cell type and cytogenetic risk factors (monosomy 3) are correlated [54, 55]. As the presence of even a low number of cells with monosomy 3 already causes deterioration of prognosis, agreement of a cutoff level for a histopathological classification appears critical. Recently, after the mutations in the gene encoding BRCA1-associated protein 1 (BAP1) on chromosome 3p21.1 have been identified as critical factor for the development of metastatic tumor phenotype, it was shown that depletion of BAP1 resulted in less differentiated spindle morphology, again drawing a connection between molecular changes and histopathological findings [56]. In addition to epithelioid cell type, the presence of specific extracellular matrix patterns (closed loops; networks) has been found to be associated with poor prognosis [57,58,59]. Those patterns are best depicted on PAS-stained sections and were initially thought to represent blood vessels but then identified as fluid-conducting channels lacking endothelial cells (vasculogenic mimicry). As cell type, they also correlate with other risk factors for metastasis, reflecting putative molecular changes in the melanoma cells [60]. In addition, increased microvascular density and the presence of tumor infiltrating macrophages and lymphocytes worsen prognosis [61,62,63]. Today, histopathological risk factors have lost much of their importance for counselling patients about their likely prognosis, as the identification of genetic risk factors has been proven to be superior predictors. However, they are still in use in multivariate mathematical models for prognostication as the inclusion of numerous predictors enables the model to correct for errors in even genetic results to some extent [46, 60].
Tumor parameters: Tumor dimensions, especially the largest tumor basal diameter (LBD), are strongly correlated with patient survival. While less than 5% of melanomas with a LBD less than 10 mm will be fatal within 5 years from diagnosis, 5-year mortality rises to over 50% in melanomas with a LBD over 15 mm [64, 65]. LBD remains an independent risk factor adding prognostic information even in addition to gene expression profiling (GEP) [66]. However, we do not know whether metastasis happens because the tumor has grown large and has had more opportunity to spread because it has been there for a longer time or whether large tumor size is simply an indicator of a greater growth rate [46]. Tumor height appears to be a less important parameter for patient survival. Tumor location, on the other hand, has long been recognized as an important predictor of metastasis. Ciliary body involvement is associated with reduced survival; however, it correlates with larger tumor size and it did not remain an independent risk factor for metastatic disease when information from GEP was available [53, 66, 67]. Melanomas of the iris, in contrast, have a favorable prognosis and a much lower mortality rate [68, 69]. At the time of diagnosis, iris melanomas are usually smaller than choroidal or ciliary body melanomas, and they show less aggressive histological features [53]. Cytogenetic changes with known unfavorable impact could be demonstrated in iris melanoma as well. Due to the small number of cases, however, the impact on survival has not been analyzed in detail [70].
9.5 Cytogenetics
Since the first report on cytogenetic changes in uveal melanoma more than 30 years ago, cytogenetic testing has gradually evolved from a research tool to a routine clinical test in the management of uveal melanoma [71]. Twenty years ago, Prescher et al. demonstrated the prognostic significance of cytogenetic abnormalities (i.e., monosomy 3), which was shown to be far superior to any clinical or histopathological marker [72]. Since then, the techniques to identify chromosomal anomalies have evolved and replaced chromosome G-banding. Today, fluorescence in situ hybridization (FISH), spectral karyotyping (SKY), (array) comparative genomic hybridization (a-CGH), and multiplex ligation-dependent probe amplification (MLPA) are in routine use for predictive testing and have been evaluated in large patient series [73, 74]. Microsatellite analysis (MSA) and single-nucleotide polymorphism (SNP) arrays offer the additional opportunity to identify loss of heterogeneity (LOH) and detect isodisomy of chromosomes which might be missed by the other techniques. Using SNP data, an attempt at creating an evolutionary tree for uveal melanoma has been published [75].
9.5.1 Chromosome 3
The loss of one copy of chromosome 3 (monosomy 3) is the most frequent and prognostically most important chromosomal aberration in uveal melanoma and can be found in 50–61% of tumors [72, 76,77,78,79,80,81]. In 1996, a dramatic reduction in the 3-year survival probability from approximately 100% to less than 50% was reported for the first time in patients with monosomy 3 melanoma [72]. Since then, this finding has been confirmed by other authors several times [46, 55, 60, 82]. Only about 5–20% of patients with disomy 3 melanoma are expected to eventually develop metastatic disease [83]. Several possible explanations why metastasis occurs in a disomy 3 tumor have been discussed. First, intratumoral heterogeneity may lead to sampling errors if only a single small biopsy is used for analysis. The risk for misclassification with fine needle aspiration biopsy (FNAB) has been estimated to be less than 1% and however has to be considered [78, 84]. Second, e.g., FISH, CGH, and MLPA cannot detect loss of heterozygosity (LOH) caused by isodisomy and give the impression of metastasis occurring without monosomy 3. Partial deletions of chromosome 3 might also be missed by FISH. However, the prognostic significance of partial deletions is still unclear. Identified deletions affected regions of the short arm (3p11–3p14, 3p25–3p26, 3p25.1–3p25.2), as well as smaller regions on 3q (3q13–3q21 and 3q24–3q26) [85,86,87,88]. A critical region of deletion causing metastatic disease has not been identified, until Harbour et al. identified mutations of BAP1 gene located on chromosome 3p21.1 in the majority of metastasizing uveal melanomas by exome sequencing [45]. Loss of chromosome 3 seems to be an early event in development of uveal melanoma and can be found in combination with other chromosomal aberrations [89, 90]. Unmasking of the second allele with mutated BAP1 induces tumor progression and also determines an aggressive tumor phenotype [56]. It is therefore not surprising that the loss of chromosome 3 is associated with other predictors of a poor prognosis, such as increased tumor diameter, ciliary body involvement, epithelioid cell type, and extravascular matrix patterns [55, 72, 91, 92].
9.5.2 Chromosome 8
Aberrations of chromosome 8 may affect both the short and the long arm. Depending on the technique used, gains of 8q can be found in 35–75% and loss of 8p in 15–30% [76, 93]. Gains of 8q and losses of 8p may occur together by formation of an isochromosome [90, 94, 95].
As for chromosome 3 changes of chromosome 8 are also associated with poor prognosis. Combining information on chromosome 3 and on chromosome 8 status improves the accuracy of prediction of metastatic disease compared to monosomy 3 or chromosome 8 status alone [46, 60]. Monosomy 3 and 8q gains occur together in about 45% of tumors and are associated with large tumor size, ciliary body involvement, and aggressive histology [80, 82]. In addition, chromosome 8 abnormalities are found in virtually all metastases from uveal melanoma, either to the liver or the brain [71, 94, 96]. Because of this and the variable copy number of 8q in one tumor, gains of chromosome 8 are thought to be a secondary event in uveal melanoma development. Improved analysis technique showed 8q amplification to be far more common than previously thought and high-resolution CGH identified frequent partial deletions on chromosome 8 [93]. Gains could be localized to 8q23–8q24 in many cases, and several oncogenes in that region have been evaluated for the potential significance. MYC, located on chromosome 8q24 and coding for a transcription factor, DDEF1 (development and differentiation factor 1), enhancing the motility of uveal melanoma cells, and NBS1 (Nijmegen breakage syndrome 1) were all analyzed for their possible role in promoting metastasis [97,98,99,100]. Though frequently overexpressed in uveal melanoma, an association to prognosis could not be established. ENPP2, also located on 8q24 and coding for autotaxin or ectonucleotide pyrophosphatase/phosphodiesterase family member 2 (ENPP2), has been identified by gene expression profiling as a predictor of survival [101]. Autotaxin is an enzyme producing lysophosphatidic acid (LPA), a small motility-enhancing and angiogenic lipid molecule [102]. Its role for tumor progression and prognosis in uveal melanoma still needs to be established.
Losses of chromosome 8p have also been analyzed in detail to identify possible tumor suppressor genes. Deletions could be localized on 8p12–8p22 by array CGH, and silencing of LZTS1 on the retained hemizygous allele was found [103]. This tumor suppressor genes code for the leucine zipper tumor suppressor-1 and inhibit motility and invasion of uveal melanoma in cell cultures. Interestingly, loss of 8p correlates with the prognostically unfavorable class 2B on gene expression profiling, further emphasizing the importance of 8p loss [89, 104].
9.5.3 Chromosome 6
Gains of chromosome 6p are observed in 28–54% of uveal melanomas and losses of 6q in 35–37% [76, 105]. Gains of 6p seem to be associated with a good prognosis. Possible explanations are that 6p gains are preferably found in spindle cell melanomas and rarely occur together with monosomy 3 [90, 106, 107]. Gains of 6p also correlate with gene expression profile subclass 1b, which is associated with a good prognosis. The protective effect of 6p gain might result from an alternative molecular pathway in tumor development dividing UM in two separate subgroups with good (6p gain) or worse (monosomy 3) prognosis. Loss of 6q, on the other hand, has been associated with a metastasizing phenotype. The effect of simultaneous 6p gain and 6q loss caused by chromosome rearrangements is unknown [77, 80].
Some authors have tried to identify tumor suppressor genes and oncogenes on chromosome 6; however, the genetic mechanism underlying chromosome 6 abnormalities remains unclear [108,109,110].
9.5.4 Chromosome 1
About a quarter of uveal melanomas show deletion on the short arm of chromosome 1, indicating a poorer prognosis. Because deletions of 1p are frequently found in larger tumors and in association with monosomy 3 and changes of chromosome 8, they are thought to develop later in tumor progression [78, 80, 84, 94, 111, 112]. However, 1p loss adds prognostic information independent of chromosome 3 status [112].
Several genetic mechanisms could be affected by 1p deletion, e.g., the NOTCH pathway and TP73 [113].
9.6 Molecular Genetics
9.6.1 MYC
The MYC gene is located on chromosome 8q24, a region frequently amplified in uveal melanoma, and therefore has been supposed to play a role in tumor progression [76, 90, 94, 95]. It codes for a transcription factor regulating the expression of numerous genes, controlling the cell cycle and apoptosis, and modifies the chromatin structure via histone acetyltransferases (HATs) [114, 115]. Many tumors show constitutive overexpression of MYC, which has also been identified in uveal melanoma and proposed as a potential prognostic marker [100, 116]. In cutaneous melanoma, overexpression of MYC seems to be associated with poor prognosis in cutaneous melanoma [117]. In uveal myeloma MYC is also frequently overexpressed but surprisingly associated with improved prognosis [97, 100]. Hence, the role of MYC overexpression in uveal melanoma remains unclear.
9.6.2 TGF-b
TGF-b1 is located on chromosome 19, a region without frequent structural abnormalities in uveal melanoma. It has become of interest, because intraocular melanomas are growing in an immune privileged location, and the immunosuppressive properties inside the eye are mediated by cytokines, such as TGF-b [118]. TGF-b has antiproliferative and anti-apoptotic effects on various cell types (epithelial, endothelial, neuronal, leukocytes) [119]. TGF-b is secreted into the extracellular matrix and stored until its activation by numerous often unspecific stimuli, like as irradiation, reactive oxygen, and proteases such as plasmin and metalloproteinases [120, 121]. Metalloproteinases (MMP) have been of interest as a marker for tumor progression as they are involved in tumor invasion, angiogenesis, and metastasis, and TGF-b enhance and modify these effects [122, 123]. Whereas MMP-9 is predominantly expressed in epithelioid melanomas and associated with poorer prognosis, immunohistochemical staining showed TGF-b positivity in uveal melanoma regardless of cell type, tumor size, or location [118]. TGF might also play a direct role in hematogenous metastasis to the liver, by increasing adhesion of uveal melanoma to the hepatic endothelium [124]. For metastases to grow to detectable size, induction of angiogenesis is a crucial step in tumor progression [125]. By its angiogenic effects, TGF-b could further promote growth of uveal melanoma metastases, and expression of TGF-b receptor endoglin has been found to correlate with metastatic death [126, 127]. A high number of tumor infiltrating leukocytes and macrophages as well as high HLA class I and II expression have also been linked with bad prognosis, as those are preferably found in monosomy 3 melanomas [128]. By downregulation of MHC class I antigen, on the other hand, TGF-b renders uveal melanoma cells more susceptible to cytolysis by natural killer cells [129, 130]. It has been suggested that uveal melanoma might prepare its own microenvironment for growth by secretion of local factors into the vitreous and aqueous humor [131].
9.6.3 Bcl-2
The Bcl family of proto-oncogenes comprises pro-apoptotic proteins, like Bax, Bad, and Bak, and anti-apoptotic proteins, like Bcl-2, Bcl-xL, and Bcl-w [132]. Blc-2 seems to be of specific importance in regulation of tumor cell survival and apoptosis. As the pro-apoptotic members, Bcl-2 resides on the outer membrane of the mitochondria, inhibiting the initiation of the apoptotic cascade by the intrinsic pathway [133, 134]. Bcl-2 is strongly expressed in uveal melanoma, as shown by immunohistochemistry [97, 135,136,137]. Uveal melanoma is known to be resistant against radiation, and high radiation dose has to be used for the treatment of primary tumor. It is also resistant to standard chemotherapy. This is explained by the anti-apoptotic effects of Bcl-2. Therefore Bcl-2 has become a possible starting point in the search for a targeted therapy [138, 139]. Bcl-2 inhibitors could revert the inhibition of apoptosis and allow for the initiation of apoptosis by pro-apoptotic members of the Bcl family. Nemati et al. showed increased response of uveal melanoma to chemotherapy with fotemustine after the administration of a Bcl-2 inhibitor in animal xenografts [140]. Until recently no Bcl inhibitor with acceptable toxicity was available for use in humans [141, 142]. However, novel Bcl-2 inhibitors with low toxicity have been approved for use in chronic lymphocytic leukemia (CLL) with 17p deletion (affecting p53), and clinical testing in uveal melanoma is expected.
9.6.4 P53
P53 plays an important role as a tumor suppressor, inactivated in more than 50% of all tumors. The gene for p53 is located on chromosome 17p13.1. It is a transcription factor controlling the expression of other downstream genes to regulate cell proliferation and growth. Upregulation of p53 has been described in uveal melanoma after irradiation [143, 144]. This appears plausible as p53 is a main factor in the cellular response to stress and DNA damage. Until DNA repair is completed, the cell is stopped in the G1 or G2 phase of the cell cycle [145]. If the damage is too severe to be repaired, the cell will undergo apoptosis. In contrast to other tumors, which frequently show altered expression or mutations of p53, the signaling pathway upstream of p53 seems to be intact in uveal melanoma [146]. However, there seems to be a functional inhibition of p53 in uveal melanoma. P53 interacts with several downstream molecules in a negative-feedback loop [147]. First, the cyclin-dependent kinase inhibitor CDKN2A stabilizes p53 by degrading Mdm2 through p14(ARF) [148, 149]. Second, it induces the expression of Hdm2/Mdm2 (human/mouse double minute 2). Overexpressed Hdm2/Mdm2 in reverse represses p53 transcriptional activity and also enhances its degradation [150,151,152]. This mechanism seems to be important in uveal melanoma, and overexpression of Mdm2 hasbeen shown to be of prognostic value [143, 153]. Some authors evaluated the possible therapeutic effect of inhibition of Mdm2 by a small synthetic peptide in animal models; however, this has never been evaluated in clinical trials [141, 154, 155].
9.6.5 Rb
The Rb pathway has been shown to be frequently altered and functionally inhibited in uveal melanoma [143, 156,157,158]. To arrest cells in the G1 or G1/S phase, the Rb protein has to be kept in a hypophosphorylated state. This is accomplished by the cyclin-dependent kinase inhibitor CDKN2A coding for p16(INK)4a, keeping Rb active. In this hypophosphorylated state, Rb binds E2Fs, thereby repressing their transcriptional activity. Phosphorylation of Rb by cyclin-dependent kinases (CDK) can occur at multiple phosphorylation sites in Rb throughout the cell cycle, which gradually inhibits Rb function and releases E2F [159, 160]. Loss or inactivation of Rb will result in deregulated cell cycle progression and cell proliferation. In uveal melanoma Rb protein is frequently hyperphosphorylated and inactivated because of cyclin D1 overexpression and CDKN2A promotor methylation [143]. Still, most tumors, including retinoblastoma, need additional mutations of other tumor suppressor genes to develop [161].
9.6.6 Ras-Raf-MAPK Pathway (GNAQ, GNA11, CYSLTR2, PLCB4)
Mutations of proteins in the mitogen-activated protein kinase pathway (MAPK) have long been implicated in the development of conjunctival melanomas [162,163,164]. BRAF mutations are found in more than 65% of cutaneous melanoma, resulting in a constitutively active protein leading to MAPK activation [165]. Such BRAF mutations are rarely observed, on the other hand, in uveal melanoma [166,167,168,169]. Activation of the MAPK pathway has been demonstrated to be a frequent event in UM; however, this does not seem to occur through mutations of BRAF [170]. The mechanisms behind the activation remained obscure until the discovery of mutations in GNAQ and GNA11 in uveal melanocytic lesions. A connection between mutations in GNAQ/GNA11 and proliferation of melanocytes was first established in 2004 during the analysis of determinants of skin color in mice [171]. Thereupon those mutations were found in almost 85% of uveal melanomas [20, 172, 173]. GNAQ and GNA11 encode the alpha subunit of heterotrimeric g-proteins (g-alpha-q and g-alpha-11). G-protein-coupled receptors (GPCR) are a large group of transmembrane receptors exerting multiple physiologic functions, such as in the sensory system, vision, olfaction, and taste [174]. Their function is transmitted by g-proteins from the extracellular to intracellular space. In their basal and inactive state, the alpha, beta, and gamma subunits are bound together with GDP. Upon activation through ligand binding to the g-protein-coupled receptor, the heterotrimeric g-protein dissociates and exchanges GDP to GTP [175]. The GTPase activity of the alpha subunit hydrolysis GTP to GDP. Mutations in the alpha subunit of GNAQ and GNA11 keep them in the activated state, leading to constitutive MAP-kinase pathway activation and cell proliferation without further extracellular stimuli. This raised the hope for targeted therapy with MEK inhibitors. In fact, a randomized open-label phase II clinical trial comparing selumetinib versus chemotherapy showed for the first time prolonged progression-free survival in patients with uveal melanoma metastasis [176]. However, this has not been confirmed in later studies [177].
In patients without detectable mutations in GNAQ/GNA11 of the g-protein-coupled receptor CYSLTR2 and PLCB4, the gene encoding for the 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta-4, a downstream target of GNAQ/GNA11, could be identified, further emphasizing the importance of MAP-kinase pathway activation through this mechanism [178, 179].
9.6.7 BAP1
Enormous efforts have been made to identify the critical mutation on chromosome 3 that promotes metastasis. Harbour et al. identified mutations of BAP1, located onchromosome 3p21.1, in 47% of metastasizing monosomy 3 uveal melanomas [45]. Interestingly, one of their patients had a germline mutation which was uncovered by loss of the second allele (monosomy 3). Soon after germline BAP1 mutations were found to cause a hereditary tumor predisposition syndrome, the spectrum of this is still evolving [40, 41, 180]. The BRCA1-associated protein-1 (BAP1) was firstdescribed by Jensen et al. as a potential tumor suppressor gene [181, 182]. BAP1 encodes a deubiquitinating enzyme and acts on BRCA1, histone H2A, host cell factor-1, and O-linked N-acetylglucosamine transferase (OGT) [181]. RNAi-mediated depletion of BAP1 in uveal melanoma cells resulted in loss of differentiation and gain of stemlike properties, similar to GEP class 2 tumors [56]. Though the molecular mechanisms of BAP1 are still not fully understood, a targeted therapy seems possible. Histone deacetylase (HDAC) inhibitors might restore histone H2A function which has been shown to be accompanied by increased melanocytic differentiation [183, 184].
9.6.8 SF3B1 and EIF1AX
Recently, additional somatic mutations have been identified in UM by exon sequencing, specifically occurring in patients with disomy 3 and partial monosomy 3 [185, 186]. SF3B1 is found in about 15–20% of uveal melanomas and associated with a favorable prognosis and prolonged metastasis-free interval compared to uveal melanomas with BAP1 mutations [186,187,188,189]. Mutations were mutually exclusive with BAP1 mutations and seem to support the bifurcated model of tumor progression in uveal melanoma [108, 190]. However, within the disomy 3 and partial monosomy 3 tumors, uveal melanomas with SF3B1 mutations had a worse prognosis than those without this mutation [189]. Interestingly, the mutational spectrum in SF3B1 of tumors with and without metastasis was different, and a further subgrouping of tumors according to mutation type might help to identify patients at risk of metastatic disease [186]. Tumors harboring EIF1AX mutations also are associated with a longer disease-free survival and rarely demonstrate metastasis [189]. Both mutations EIF1AX and SF3B1 are associated with good prognostic features, such as disomy 3; spindle cell type, positive BAP1 immunohistochemistry staining; and the absence of closed vascular loops [191].
9.6.9 Gene Expression Profiling (GEP)
While over the past decades methods for detection of cytogenetic changes and mutations have evolved and allowed for more detailed analyses, some other groups chose another approach offering a different view of the tumor and its microenvironment. Gene expression profiling (GEP), based on mRNA signature, has been described as a functional snapshot of the tumor microenvironment, which is perhaps less variable across the tumor and therefore less affected by intratumoral heterogeneity [192]. GEP was studied first in uveal melanoma cell lines, comparing those to normal melanocytes [193]. Then, Tschentscher et al. developed oligonucleotide microarrays to describe and compare the expression profile of monosomy and disomy 3 melanomas [194]. They also classified uveal melanomas according to their gene expression profile.
Soon afterwards, Onken et al. improved this technique and, leaving chromosomal status behind, classified uveal melanomas based on results from GEP, using unsupervised clustering methods to divide uveal melanomas in class 1 melanomas with low risk and class 2 tumors with a high risk of metastasis. Since then, numerous other investigators have compared the accuracy of GEP versus cytogenetic prognostic indicators. Unfortunately, as DNA-based techniques continuously evolved (MLPA, a-CGH, and SNP array), no direct comparison of GEP classification versus latest DNA-based techniques has been made. Onken et al. reduced the number of genes necessary for classification and developed a PCR-based microfluidics platform for routine clinical testing [192, 195].
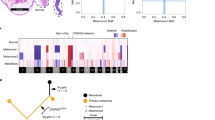
Gene expression profiling closely correlates with cytogenetic findings in most cases [196, 197]. Class 1 tumors usually show disomy 3 and spindle cell type melanoma, whereas class 2 profile is associated with epithelioid cell type, monosomy 3. A further subclassification is possible into four groups (1A, 1B, and 2A, 2B). Class 1A predicts the best prognosis, and class 1B is slightly worse, corresponding to disomy 3 melanomas with 6p gain. For class 2 melanomas, additional loss of chromosome 8p corresponds to the subclass 2B with the worst prognosis [89]. Based on this, a modified bifurcated tumor progression model has been proposed which is shown in Fig. 9.1. Some problems, however, remain. First, the significance and predictive value of GEP after radiation has not been evaluated so far and, as in contrast, has been done for DNA-based tests [198,199,200,201]. Also biopsy of non-melanoma tissue will reveal class 1 profile. For DNA-based tests, identification of GNAQ/GNA11 mutations can easily be done which will be positive in 85% of cases and ensure that melanocytic tissue has been biopsied.
Proposed bifurcated tumor progression model. GNAQ/GNA11 mutations induce growth of uveal nevi, which subsequently evolve into melanoma by acquiring (a) mutations in SF3B1 and/or EIF1AX, associated with a rather good prognosis, or (b) mutations in BAP1 and loss of the second allele by monosomy 3, which is associated with a high risk for metastasis. A more advanced model of an evolutionary tree for uveal melanoma was published by Nakul Singh et al. [75]
9.7 Conclusion and Future Prospective
New genetic techniques have provided new and sometimes unsuspected insights into the pathobiology and the molecular carcinogenesis of uveal melanoma. Extended use of next-generation sequencing and proteomics will provide further insights, and targeted therapies will hopefully lead to an improved prognosis for patients with metastatic disease. Still, one should not forget to consider and reconsider again basic facts of melanogenesis in the light of latest findings as explained in the introduction of this chapter. It is still unknown whether choroidal nevi are congenital and slowly grow to clinically detectable size during life or whether they arise from normal melanocytes, which acquire their first transforming event (mutation of GNAQ/GNA11) later in life. Findings from clinical syndromes like the oculodermal melanocytosis still pose many questions to be answered. Unifying concepts of pigment cell distribution and ocular melanogenesis may answer some questions but even more arise.
References
Schwab C, Wackernagel W, Grinninger P, Mayer C, Schwab K, Langmann G, Richtig E, Wedrich A, Hofmann-Wellenhof R, Zalaudek I. A unifying concept of uveal pigment cell distribution and dissemination based on an animal model: insights into ocular melanogenesis. Cells Tissues Organs. 2016;201:232–8.
Saida T. Histogenesis of congenital and acquired melanocytic nevi: a unifying concept. Am J Dermatopathol. 2006;28:377–9.
Schwab C, Zalaudek I, Mayer C, Riedl R, Wackernagel W, Juch H, Aigner B, Brunasso AM, Langmann G, Richtig E. New insights into oculodermal nevogenesis and proposal for a new iris nevus classification. Br J Ophthalmol. 2015;99:644–9.
Singh AD, Turell ME, Topham AK. Uveal melanoma: trends in incidence, treatment, and survival. Ophthalmology. 2011;118:1881–5.
Hu DN, Yu GP, Mccormick SA, Schneider S, Finger PT. Population-based incidence of uveal melanoma in various races and ethnic groups. Am J Ophthalmol. 2005;140:612–7.
Singh AD, Topham A. Incidence of uveal melanoma in the United States: 1973-1997. Ophthalmology. 2003;110:956–61.
Virgili G, Gatta G, Ciccolallo L, Capocaccia R, Biggeri A, Crocetti E, Lutz JM, Paci E, Group EW. Incidence of uveal melanoma in Europe. Ophthalmology. 2007;114:2309–15.
Chang AE, Karnell LH, Menck HR. The National Cancer Data Base report on cutaneous and noncutaneous melanoma: a summary of 84,836 cases from the past decade. Cancer. 1998;83:1664–78.
Egan KM, Seddon JM, Glynn RJ, Gragoudas ES, Albert DM. Epidemiologic aspects of uveal melanoma. Surv Ophthalmol. 1988;32:239–51.
Margo CE, Mulla Z, Billiris K. Incidence of surgically treated uveal melanoma by race and ethnicity. Ophthalmology. 1998;105:1087–90.
Marshall JC, Gordon KD, Mccauley CS, De Souza Filho JP, Burnier MN. The effect of blue light exposure and use of intraocular lenses on human uveal melanoma cell lines. Melanoma Res. 2006;16:537–41.
Singh AD, Rennie IG, Seregard S, Giblin M, Mckenzie J. Sunlight exposure and pathogenesis of uveal melanoma. Surv Ophthalmol. 2004;49:419–28.
Vajdic CM, Kricker A, Giblin M, Mckenzie J, Aitken J, Giles GG, Armstrong BK. Sun exposure predicts risk of ocular melanoma in Australia. Int J Cancer. 2002;101:175–82.
Singh AD, Shields CL, De Potter P, Shields JA, Trock B, Cater J, Pastore D. Familial uveal melanoma. Clinical observations on 56 patients. Arch Ophthalmol. 1996a;114:392–9.
Jay M, Mccartney AC. Familial malignant melanoma of the uvea and p53: a Victorian detective story. Surv Ophthalmol. 1993;37:457–62.
Canning CR, Hungerford J. Familial uveal melanoma. Br J Ophthalmol. 1988;72:241–3.
Singh AD, Wang MX, Donoso LA, Shields CL, Potter PD, Shields JA, Elston RC, Fijal B. Familial uveal melanoma, III. Is the occurrence of familial uveal melanoma coincidental? Arch Ophthalmol. 1996b;114:1101–4.
Singh AD, Demirci H, Shields CL, Shields JA, Smith AF. Concurrent choroidal melanoma in son and father. Am J Ophthalmol. 2000;130:679–80.
Singh AD, De Potter P, Fijal BA, Shields CL, Shields JA, Elston RC. Lifetime prevalence of uveal melanoma in white patients with oculo(dermal) melanocytosis. Ophthalmology. 1998;105:195–8.
Van Raamsdonk CD, Bezrookove V, Green G, Bauer J, Gaugler L, O’brien JM, Simpson EM, Barsh GS, Bastian BC. Frequent somatic mutations of GNAQ in uveal melanoma and blue naevi. Nature. 2009;457:599–602.
NIH Consensus conference. Diagnosis and treatment of early melanoma. JAMA. 1992;268:1314–9.
Naeyaert JM, Brochez L. Clinical practice. Dysplastic nevi. N Engl J Med. 2003;349:2233–40.
Feng J, Kim ST, Liu W, Kim JW, Zhang Z, Zhu Y, Berens M, Sun J, Xu J. An integrated analysis of germline and somatic, genetic and epigenetic alterations at 9p21.3 in glioblastoma. Cancer. 2012;118:232–40.
Soura E, Eliades PJ, Shannon K, Stratigos AJ, Tsao H. Hereditary melanoma: update on syndromes and management: genetics of familial atypical multiple mole melanoma syndrome. J Am Acad Dermatol, 2016; 74: 395–407; quiz 408–10.
Abramson DH, Rodriguez-Sains RS, Rubman R. B-K mole syndrome. Cutaneous and ocular malignant melanoma. Arch Ophthalmol. 1980;98:1397–9.
Bellet RE, Shields JA, Soll DB, Bernardino EA. Primary choroidal and cutaneous melanomas occurring in a patient with the B-K mole syndrome phenotype. Am J Ophthalmol. 1980;89:567–70.
Richtig E, Langmann G, Mullner K, Smolle J. Ocular melanoma: epidemiology, clinical presentation and relationship with dysplastic nevi. Ophthalmologica. 2004;218:111–4.
Rodriguez-Sains RS. Ocular findings in patients with dysplastic nevus syndrome. Ophthalmology. 1986;93:661–5.
Rodriguez-Sains RS. Ocular findings in patients with dysplastic nevus syndrome. An update. Dermatol Clin. 1991;9:723–8.
Singh AD, Shields CL, Shields JA, Eagle RC, De Potter P. Uveal melanoma and familial atypical mole and melanoma (FAM-M) syndrome. Ophthalmic Genet. 1995;16:53–61.
Smith JH, Padnick-Silver L, Newlin A, Rhodes K, Rubinstein WS. Genetic study of familial uveal melanoma: association of uveal and cutaneous melanoma with cutaneous and ocular nevi. Ophthalmology. 2007;114:774–9.
Toth-Molnar E, Olah J, Dobozy A, Hammer H. Ocular pigmented findings in patients with dysplastic naevus syndrome. Melanoma Res. 2004;14:43–7.
Abdel-Rahman MH, Pilarski R, Massengill JB, Christopher BN, Noss R, Davidorf FH. Melanoma candidate genes CDKN2A/p16/INK4A, p14ARF, and CDK4 sequencing in patients with uveal melanoma with relative high-risk for hereditary cancer predisposition. Melanoma Res. 2011b;21:175–9.
Buecher B, Gauthier-Villars M, Desjardins L, Lumbroso-Le Rouic L, Levy C, De Pauw A, Bombled J, Tirapo C, Houdayer C, Bressac-De Paillerets B, Stoppa-Lyonnet D. Contribution of CDKN2A/P16 (INK4A), P14 (ARF), CDK4 and BRCA1/2 germline mutations in individuals with suspected genetic predisposition to uveal melanoma. Fam Cancer. 2010;9:663–7.
Mouriaux F, Maurage CA, Labalette P, Sablonniere B, Malecaze F, Darbon JM. Cyclin-dependent kinase inhibitory protein expression in human choroidal melanoma tumors. Invest Ophthalmol Vis Sci. 2000;41:2837–43.
Zeschnigk M, Tschentscher F, Lich C, Brandt B, Horsthemke B, Lohmann DR. Methylation analysis of several tumour suppressor genes shows a low frequency of methylation of CDKN2A and RARB in uveal melanomas. Comp Funct Genomics. 2003;4:329–36.
Friedman JM. Epidemiology of neurofibromatosis type 1. Am J Med Genet. 1999;89:1–6.
Foster WJ, Fuller CE, Perry A, Harbour JW. Status of the NF1 tumor suppressor locus in uveal melanoma. Arch Ophthalmol. 2003;121:1311–5.
Varley JM. Germline TP53 mutations and Li-Fraumeni syndrome. Hum Mutat. 2003;21:313–20.
Abdel-Rahman MH, Pilarski R, Cebulla CM, Massengill JB, Christopher BN, Boru G, Hovland P, Davidorf FH. Germline BAP1 mutation predisposes to uveal melanoma, lung adenocarcinoma, meningioma, and other cancers. J Med Genet. 2011a;48:856–9.
Wiesner T, Fried I, Ulz P, Stacher E, Popper H, Murali R, Kutzner H, Lax S, Smolle-Juttner F, Geigl JB, Speicher MR. Toward an improved definition of the tumor spectrum associated with BAP1 germline mutations. J Clin Oncol. 2012;30:e337–40.
Wiesner T, Obenauf AC, Murali R, Fried I, Griewank KG, Ulz P, Windpassinger C, Wackernagel W, Loy S, Wolf I, Viale A, Lash AE, Pirun M, Socci ND, Rutten A, Palmedo G, Abramson D, Offit K, Ott A, Becker JC, Cerroni L, Kutzner H, Bastian BC, Speicher MR. Germline mutations in BAP1 predispose to melanocytic tumors. Nat Genet. 2011;43:1018–21.
Hoiom V, Edsgard D, Helgadottir H, Eriksson H, All-Ericsson C, Tuominen R, Ivanova I, Lundeberg J, Emanuelsson O, Hansson J. Hereditary uveal melanoma: a report of a germline mutation in BAP1. Genes Chromosomes Cancer. 2013;52:378–84.
Maerker DA, Zeschnigk M, Nelles J, Lohmann DR, Worm K, Bosserhoff AK, Krupar R, Jagle H. BAP1 germline mutation in two first grade family members with uveal melanoma. Br J Ophthalmol. 2014;98:224–7.
Harbour JW, Onken MD, Roberson ED, Duan S, Cao L, Worley LA, Council ML, Matatall KA, Helms C, Bowcock AM. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science. 2010;330:1410–3.
Damato B, Eleuteri A, Taktak AF, Coupland SE. Estimating prognosis for survival after treatment of choroidal melanoma. Prog Retin Eye Res. 2011;30:285–95.
Damato BE, Heimann H, Kalirai H, Coupland SE. Age, survival predictors, and metastatic death in patients with choroidal melanoma: tentative evidence of a therapeutic effect on survival. JAMA Ophthalmol. 2014;132:605–13.
Kujala E, Makitie T, Kivela T. Very long-term prognosis of patients with malignant uveal melanoma. Invest Ophthalmol Vis Sci. 2003;44:4651–9.
Rietschel P, Panageas KS, Hanlon C, Patel A, Abramson DH, Chapman PB. Variates of survival in metastatic uveal melanoma. J Clin Oncol. 2005;23:8076–80.
Virgili G, Gatta G, Ciccolallo L, Capocaccia R, Biggeri A, Crocetti E, Lutz JM, Paci E, Group EW. Survival in patients with uveal melanoma in Europe. Arch Ophthalmol. 2008;126:1413–8.
Mclean IW, Foster WD, Zimmerman LE, Gamel JW. Modifications of Callender’s classification of uveal melanoma at the Armed Forces Institute of Pathology. Am J Ophthalmol. 1983;96:502–9.
Mclean IW, Foster WD, Zimmerman LE. Uveal melanoma: location, size, cell type, and enucleation as risk factors in metastasis. Hum Pathol. 1982;13:123–32.
Mudhar HS, Parsons MA, Sisley K, Rundle P, Singh A, Rennie IG. A critical appraisal of the prognostic and predictive factors for uveal malignant melanoma. Histopathology. 2004;45:1–12.
Bronkhorst IH, Maat W, Jordanova ES, Kroes WG, Schalij-Delfos NE, Luyten GP, Jager MJ. Effect of heterogeneous distribution of monosomy 3 on prognosis in uveal melanoma. Arch Pathol Lab Med. 2011b;135:1042–7.
Scholes AG, Damato BE, Nunn J, Hiscott P, Grierson I, Field JK. Monosomy 3 in uveal melanoma: correlation with clinical and histologic predictors of survival. Invest Ophthalmol Vis Sci. 2003;44:1008–11.
Matatall KA, Agapova OA, Onken MD, Worley LA, Bowcock AM, Harbour JW. BAP1 deficiency causes loss of melanocytic cell identity in uveal melanoma. BMC Cancer. 2013;13:371.
Folberg R, Rummelt V, Parys-Van Ginderdeuren R, Hwang T, Woolson RF, Pe’er J, Gruman LM. The prognostic value of tumor blood vessel morphology in primary uveal melanoma. Ophthalmology. 1993;100:1389–98.
Makitie T, Summanen P, Tarkkanen A, Kivela T. Microvascular loops and networks as prognostic indicators in choroidal and ciliary body melanomas. J Natl Cancer Inst. 1999b;91:359–67.
Maniotis AJ, Folberg R, Hess A, Seftor EA, Gardner LM, Pe’er J, Trent JM, Meltzer PS, Hendrix MJ. Vascular channel formation by human melanoma cells in vivo and in vitro: vasculogenic mimicry. Am J Pathol. 1999;155:739–52.
Damato B, Duke C, Coupland SE, Hiscott P, Smith PA, Campbell I, Douglas A, Howard P. Cytogenetics of uveal melanoma: a 7-year clinical experience. Ophthalmology. 2007;114:1925–31.
Bronkhorst IH, Ly LV, Jordanova ES, Vrolijk J, Versluis M, Luyten GP, Jager MJ. Detection of M2-macrophages in uveal melanoma and relation with survival. Invest Ophthalmol Vis Sci. 2011a;52:643–50.
Makitie T, Summanen P, Tarkkanen A, Kivela T. Microvascular density in predicting survival of patients with choroidal and ciliary body melanoma. Invest Ophthalmol Vis Sci. 1999a;40:2471–80.
Whelchel JC, Farah SE, Mclean IW, Burnier MN. Immunohistochemistry of infiltrating lymphocytes in uveal malignant melanoma. Invest Ophthalmol Vis Sci. 1993;34:2603–6.
Damato B, Eleuteri A, Fisher AC, Coupland SE, Taktak AF. Artificial neural networks estimating survival probability after treatment of choroidal melanoma. Ophthalmology. 2008;115:1598–607.
Gamel JW, Mclean IW, Mccurdy JB. Biologic distinctions between cure and time to death in 2892 patients with intraocular melanoma. Cancer. 1993;71:2299–305.
Walter SD, Chao DL, Feuer W, Schiffman J, Char DH, Harbour JW. Prognostic implications of tumor diameter in association with gene expression profile for uveal melanoma. JAMA Ophthalmol. 2016;134:734–40.
Schmittel A, Bechrakis NE, Martus P, Mutlu D, Scheibenbogen C, Bornfeld N, Foerster MH, Thiel E, Keilholz U. Independent prognostic factors for distant metastases and survival in patients with primary uveal melanoma. Eur J Cancer. 2004;40:2389–95.
Shields CL, Kaliki S, Shah SU, Luo W, Furuta M, Shields JA. Iris melanoma: features and prognosis in 317 children and adults. J AAPOS. 2012;16:10–6.
Shields CL, Shields JA, Materin M, Gershenbaum E, Singh AD, Smith A. Iris melanoma: risk factors for metastasis in 169 consecutive patients. Ophthalmology. 2001;108:172–8.
Shields CL, Ramasubramanian A, Ganguly A, Mohan D, Shields JA. Cytogenetic testing of iris melanoma using fine needle aspiration biopsy in 17 patients. Retina. 2011;31:574–80.
Rey JA, Bello MJ, De Campos JM, Ramos MC, Benitez J. Cytogenetic findings in a human malignant melanoma metastatic to the brain. Cancer Genet Cytogenet. 1985;16:179–83.
Prescher G, Bornfeld N, Hirche H, Horsthemke B, Jockel KH, Becher R. Prognostic implications of monosomy 3 in uveal melanoma. Lancet. 1996;347:1222–5.
Coupland SE, Lake SL, Zeschnigk M, Damato BE. Molecular pathology of uveal melanoma. Eye (Lond). 2013;27:230–42.
Naus NC, Van Drunen E, De Klein A, Luyten GP, Paridaens DA, Alers JC, Ksander BR, Beverloo HB, Slater RM. Characterization of complex chromosomal abnormalities in uveal melanoma by fluorescence in situ hybridization, spectral karyotyping, and comparative genomic hybridization. Genes Chromosomes Cancer. 2001;30:267–73.
Singh N, Singh AD, Hide W. Inferring an evolutionary tree of uveal melanoma from genomic copy number aberrations. Invest Ophthalmol Vis Sci. 2015;56:6801–9.
Damato B, Dopierala JA, Coupland SE. Genotypic profiling of 452 choroidal melanomas with multiplex ligation-dependent probe amplification. Clin Cancer Res. 2010;16:6083–92.
Horsman DE, White VA. Cytogenetic analysis of uveal melanoma. Consistent occurrence of monosomy 3 and trisomy 8q. Cancer. 1993;71:811–9.
Naus NC, Verhoeven AC, Van Drunen E, Slater R, Mooy CM, Paridaens DA, Luyten GP, De Klein A. Detection of genetic prognostic markers in uveal melanoma biopsies using fluorescence in situ hybridization. Clin Cancer Res. 2002;8:534–9.
Singh AD, Boghosian-Sell L, Wary KK, Shields CL, De Potter P, Donoso LA, Shields JA, Cannizzaro LA. Cytogenetic findings in primary uveal melanoma. Cancer Genet Cytogenet. 1994;72:109–15.
Sisley K, Parsons MA, Garnham J, Potter AM, Curtis D, Rees RC, Rennie IG. Association of specific chromosome alterations with tumour phenotype in posterior uveal melanoma. Br J Cancer. 2000;82:330–8.
Wiltshire RN, Elner VM, Dennis T, Vine AK, Trent JM. Cytogenetic analysis of posterior uveal melanoma. Cancer Genet Cytogenet. 1993;66:47–53.
Sisley K, Rennie IG, Parsons MA, Jacques R, Hammond DW, Bell SM, Potter AM, Rees RC. Abnormalities of chromosomes 3 and 8 in posterior uveal melanoma correlate with prognosis. Genes Chromosomes Cancer. 1997;19:22–8.
Trolet J, Hupe P, Huon I, Lebigot I, Decraene C, Delattre O, Sastre-Garau X, Saule S, Thiery JP, Plancher C, Asselain B, Desjardins L, Mariani P, Piperno-Neumann S, Barillot E, Couturier J. Genomic profiling and identification of high-risk uveal melanoma by array CGH analysis of primary tumors and liver metastases. Invest Ophthalmol Vis Sci. 2009;50:2572–80.
Damato B, Dopierala J, Klaasen A, Van Dijk M, Sibbring J, Coupland SE. Multiplex ligation-dependent probe amplification of uveal melanoma: correlation with metastatic death. Invest Ophthalmol Vis Sci. 2009;50:3048–55.
Blasi MA, Roccella F, Balestrazzi E, Del Porto G, De Felice N, Roccella M, Rota R, Grammatico P. 3p13 region: a possible location of a tumor suppressor gene involved in uveal melanoma. Cancer Genet Cytogenet. 1999;108:81–3.
Cross NA, Ganesh A, Parpia M, Murray AK, Rennie IG, Sisley K. Multiple locations on chromosome 3 are the targets of specific deletions in uveal melanoma. Eye (Lond). 2006;20:476–81.
Parrella P, Fazio VM, Gallo AP, Sidransky D, Merbs SL. Fine mapping of chromosome 3 in uveal melanoma: identification of a minimal region of deletion on chromosomal arm 3p25.1-p25.2. Cancer Res. 2003;63:8507–10.
Tschentscher F, Prescher G, Horsman DE, White VA, Rieder H, Anastassiou G, Schilling H, Bornfeld N, Bartz-Schmidt KU, Horsthemke B, Lohmann DR, Zeschnigk M. Partial deletions of the long and short arm of chromosome 3 point to two tumor suppressor genes in uveal melanoma. Cancer Res. 2001;61:3439–42.
Landreville S, Agapova OA, Harbour JW. Emerging insights into the molecular pathogenesis of uveal melanoma. Future Oncol. 2008;4:629–36.
Prescher G, Bornfeld N, Friedrichs W, Seeber S, Becher R. Cytogenetics of twelve cases of uveal melanoma and patterns of nonrandom anomalies and isochromosome formation. Cancer Genet Cytogenet. 1995;80:40–6.
Mooy CM, De Jong PT. Prognostic parameters in uveal melanoma: a review. Surv Ophthalmol. 1996;41:215–28.
White VA, Chambers JD, Courtright PD, Chang WY, Horsman DE. Correlation of cytogenetic abnormalities with the outcome of patients with uveal melanoma. Cancer. 1998;83:354–9.
Hammond DW, Al-Shammari NS, Danson S, Jacques R, Rennie IG, Sisley K. High-resolution array CGH analysis identifies regional deletions and amplifications of chromosome 8 in uveal melanoma. Invest Ophthalmol Vis Sci. 2015;56:3460–6.
Aalto Y, Eriksson L, Seregard S, Larsson O, Knuutila S. Concomitant loss of chromosome 3 and whole arm losses and gains of chromosome 1, 6, or 8 in metastasizing primary uveal melanoma. Invest Ophthalmol Vis Sci. 2001;42:313–7.
Horsman DE, Sroka H, Rootman J, White VA. Monosomy 3 and isochromosome 8q in a uveal melanoma. Cancer Genet Cytogenet. 1990;45:249–53.
Parada LA, Maranon A, Hallen M, Tranberg KG, Stenram U, Bardi G, Johansson B. Cytogenetic analyses of secondary liver tumors reveal significant differences in genomic imbalances between primary and metastatic colon carcinomas. Clin Exp Metastasis. 1999;17:471–9.
Chana JS, Wilson GD, Cree IA, Alexander RA, Myatt N, Neale M, Foss AJ, Hungerford JL. c-myc, p53, and Bcl-2 expression and clinical outcome in uveal melanoma. Br J Ophthalmol. 1999;83:110–4.
Ehlers JP, Harbour JW. NBS1 expression as a prognostic marker in uveal melanoma. Clin Cancer Res. 2005;11:1849–53.
Ehlers JP, Worley L, Onken MD, Harbour JW. DDEF1 is located in an amplified region of chromosome 8q and is overexpressed in uveal melanoma. Clin Cancer Res. 2005;11:3609–13.
Royds JA, Sharrard RM, Parsons MA, Lawry J, Rees R, Cottam D, Wagner B, Rennie IG. C-myc oncogene expression in ocular melanomas. Graefes Arch Clin Exp Ophthalmol. 1992;230:366–71.
Singh AD, Sisley K, Xu Y, Li J, Faber P, Plummer SJ, Mudhar HS, Rennie IG, Kessler PM, Casey G, Williams BG. Reduced expression of autotaxin predicts survival in uveal melanoma. Br J Ophthalmol. 2007;91:1385–92.
Boutin JA, Ferry G. Autotaxin. Cell Mol Life Sci. 2009;66:3009–21.
Onken MD, Worley LA, Harbour JW. A metastasis modifier locus on human chromosome 8p in uveal melanoma identified by integrative genomic analysis. Clin Cancer Res. 2008a;14:3737–45.
Harbour JW. A prognostic test to predict the risk of metastasis in uveal melanoma based on a 15-gene expression profile. Methods Mol Biol. 2014;1102:427–40.
Hoglund M, Gisselsson D, Hansen GB, White VA, Sall T, Mitelman F, Horsman D. Dissecting karyotypic patterns in malignant melanomas: temporal clustering of losses and gains in melanoma karyotypic evolution. Int J Cancer. 2004;108:57–65.
Singh AD, Tubbs R, Biscotti C, Schoenfield L, Trizzoi P. Chromosomal 3 and 8 status within hepatic metastasis of uveal melanoma. Arch Pathol Lab Med. 2009;133:1223–7.
Sisley K, Tattersall N, Dyson M, Smith K, Mudhar HS, Rennie IG. Multiplex fluorescence in situ hybridization identifies novel rearrangements of chromosomes 6, 15, and 18 in primary uveal melanoma. Exp Eye Res. 2006;83:554–9.
Parrella P, Sidransky D, Merbs SL. Allelotype of posterior uveal melanoma: implications for a bifurcated tumor progression pathway. Cancer Res. 1999;59:3032–7.
Speicher MR, Prescher G, Du Manoir S, Jauch A, Horsthemke B, Bornfeld N, Becher R, Cremer T. Chromosomal gains and losses in uveal melanomas detected by comparative genomic hybridization. Cancer Res. 1994;54:3817–23.
Van Gils W, Kilic E, Bruggenwirth HT, Vaarwater J, Verbiest MM, Beverloo B, Van Til-Berg ME, Paridaens D, Luyten GP, De Klein A. Regional deletion and amplification on chromosome 6 in a uveal melanoma case without abnormalities on chromosomes 1p, 3 and 8. Melanoma Res. 2008;18:10–5.
Hausler T, Stang A, Anastassiou G, Jockel KH, Mrzyk S, Horsthemke B, Lohmann DR, Zeschnigk M. Loss of heterozygosity of 1p in uveal melanomas with monosomy 3. Int J Cancer. 2005;116:909–13.
Kilic E, Naus NC, Van Gils W, Klaver CC, Van Til ME, Verbiest MM, Stijnen T, Mooy CM, Paridaens D, Beverloo HB, Luyten GP, De Klein A. Concurrent loss of chromosome arm 1p and chromosome 3 predicts a decreased disease-free survival in uveal melanoma patients. Invest Ophthalmol Vis Sci. 2005;46:2253–7.
Kilic E, Bruggenwirth HT, Meier M, Naus NC, Beverloo HB, Meijerink JP, Luyten GP, De Klein A. Increased expression of p73Deltaex2 transcript in uveal melanoma with loss of chromosome 1p. Melanoma Res. 2008;18:208–13.
Adhikary S, Eilers M. Transcriptional regulation and transformation by Myc proteins. Nat Rev Mol Cell Biol. 2005;6:635–45.
Cotterman R, Jin VX, Krig SR, Lemen JM, Wey A, Farnham PJ, Knoepfler PS. N-Myc regulates a widespread euchromatic program in the human genome partially independent of its role as a classical transcription factor. Cancer Res. 2008;68:9654–62.
Zeller KI, Jegga AG, Aronow BJ, O’donnell KA, Dang CV. An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol. 2003;4:R69.
Ross DA, Wilson GD. Expression of c-myc oncoprotein represents a new prognostic marker in cutaneous melanoma. Br J Surg. 1998;85:46–51.
Esser P, Grisanti S, Bartz-Schmidt K. TGF-beta in uveal melanoma. Microsc Res Tech. 2001;52:396–400.
Siegel PM, Massague J. Cytostatic and apoptotic actions of TGF-beta in homeostasis and cancer. Nat Rev Cancer. 2003;3:807–21.
Derynck R, Akhurst RJ, Balmain A. TGF-beta signaling in tumor suppression and cancer progression. Nat Genet. 2001;29:117–29.
Koli K, Saharinen J, Hyytiainen M, Penttinen C, Keski-Oja J. Latency, activation, and binding proteins of TGF-beta. Microsc Res Tech. 2001;52:354–62.
El-Shabrawi Y, Ardjomand N, Radner H, Ardjomand N. MMP-9 is predominantly expressed in epithelioid and not spindle cell uveal melanoma. J Pathol. 2001;194:201–6.
Lai K, Conway RM, Crouch R, Jager MJ, Madigan MC. Expression and distribution of MMPs and TIMPs in human uveal melanoma. Exp Eye Res. 2008;86:936–41.
Woodward JK, Rennie IG, Burn JL, Sisley K. A potential role for TGFbeta in the regulation of uveal melanoma adhesive interactions with the hepatic endothelium. Invest Ophthalmol Vis Sci. 2005;46:3473–7.
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57–70.
Pardali E, Ten Dijke P. Transforming growth factor-beta signaling and tumor angiogenesis. Front Biosci (Landmark Ed). 2009;14:4848–61.
Ziemssen F, Wegner R, Wegner J, Tatar O, Susskind D, Gelisken F, Rohrbach M, Bartz-Schmidt KU, Grisanti S. Analysis of neovasculature in uveal melanoma by targeting the TGFbeta-binding receptor endoglin: is there prognostic relevance of proliferating endothelium? Graefes Arch Clin Exp Ophthalmol. 2006;244:1124–31.
Maat W, Ly LV, Jordanova ES, De Wolff-Rouendaal D, Schalij-Delfos NE, Jager MJ. Monosomy of chromosome 3 and an inflammatory phenotype occur together in uveal melanoma. Invest Ophthalmol Vis Sci. 2008;49:505–10.
Ijland SA, Jager MJ, Heijdra BM, Westphal JR, Peek R. Expression of angiogenic and immunosuppressive factors by uveal melanoma cell lines. Melanoma Res. 1999;9:445–50.
Ma D, Niederkorn JY. Transforming growth factor-beta down-regulates major histocompatibility complex class I antigen expression and increases the susceptibility of uveal melanoma cells to natural killer cell-mediated cytolysis. Immunology. 1995;86:263–9.
Canovas D, Rennie IG, Nichols CE, Sisley K. Local environmental influences on uveal melanoma: vitreous humor promotes uveal melanoma invasion, whereas the aqueous can be inhibitory. Cancer. 2008;112:1787–94.
Cory S, Huang DC, Adams JM. The Bcl-2 family: roles in cell survival and oncogenesis. Oncogene. 2003;22:8590–607.
Adams JM, Cory S. Bcl-2-regulated apoptosis: mechanism and therapeutic potential. Curr Opin Immunol. 2007;19:488–96.
Thornberry NA. Caspases: key mediators of apoptosis. Chem Biol. 1998;5:R97–103.
Hussein MR. Analysis of Bcl-2 protein expression in choroidal melanomas. J Clin Pathol. 2005;58:486–9.
Jay V, Yi Q, Hunter WS, Zielenska M. Expression of bcl-2 in uveal malignant melanoma. Arch Pathol Lab Med. 1996;120:497–8.
Mooy CM, Luyten GP, De Jong PT, Luider TM, Stijnen T, Van De Ham F, Van Vroonhoven CC, Bosman FT. Immunohistochemical and prognostic analysis of apoptosis and proliferation in uveal melanoma. Am J Pathol. 1995;147:1097–104.
Druker BJ. Perspectives on the development of a molecularly targeted agent. Cancer Cell. 2002;1:31–6.
Triozzi PL, Eng C, Singh AD. Targeted therapy for uveal melanoma. Cancer Treat Rev. 2008;34:247–58.
Nemati F, De Montrion C, Lang G, Kraus-Berthier L, Carita G, Sastre-Garau X, Berniard A, Vallerand D, Geneste O, De Plater L, Pierre A, Lockhart B, Desjardins L, Piperno-Neumann S, Depil S, Decaudin D. Targeting Bcl-2/Bcl-XL induces antitumor activity in uveal melanoma patient-derived xenografts. PLoS One. 2014;9:e80836.
Harbour JW, Worley L, Ma D, Cohen M. Transducible peptide therapy for uveal melanoma and retinoblastoma. Arch Ophthalmol. 2002;120:1341–6.
Vogler M, Dinsdale D, Dyer MJ, Cohen GM. Bcl-2 inhibitors: small molecules with a big impact on cancer therapy. Cell Death Differ. 2009;16:360–7.
Brantley Jr MA, Harbour JW. Deregulation of the Rb and p53 pathways in uveal melanoma. Am J Pathol. 2000a;157:1795–801.
Janssen K, Kuntze J, Busse H, Schmid KW. p53 oncoprotein overexpression in choroidal melanoma. Mod Pathol. 1996;9:267–72.
Wang JY. DNA damage and apoptosis. Cell Death Differ. 2001;8:1047–8.
Brantley Jr MA, Worley L, Harbour JW. Altered expression of Rb and p53 in uveal melanomas following plaque radiotherapy. Am J Ophthalmol. 2002;133:242–8.
Prives C, Hall PA. The p53 pathway. J Pathol. 1999;187:112–26.
Chin L, Pomerantz J, Depinho RA. The INK4a/ARF tumor suppressor: one gene--two products--two pathways. Trends Biochem Sci. 1998;23:291–6.
Pomerantz J, Schreiber-Agus N, Liegeois NJ, Silverman A, Alland L, Chin L, Potes J, Chen K, Orlow I, Lee HW, Cordon-Cardo C, Depinho RA. The Ink4a tumor suppressor gene product, p19Arf, interacts with MDM2 and neutralizes MDM2’s inhibition of p53. Cell. 1998;92:713–23.
Haupt Y, Maya R, Kazaz A, Oren M. Mdm2 promotes the rapid degradation of p53. Nature. 1997;387:296–9.
Honda R, Tanaka H, Yasuda H. Oncoprotein MDM2 is a ubiquitin ligase E3 for tumor suppressor p53. FEBS Lett. 1997;420:25–7.
Landers JE, Cassel SL, George DL. Translational enhancement of mdm2 oncogene expression in human tumor cells containing a stabilized wild-type p53 protein. Cancer Res. 1997;57:3562–8.
Coupland SE, Anastassiou G, Stang A, Schilling H, Anagnostopoulos I, Bornfeld N, Stein H. The prognostic value of cyclin D1, p53, and MDM2 protein expression in uveal melanoma. J Pathol. 2000;191:120–6.
Chene P, Fuchs J, Bohn J, Garcia-Echeverria C, Furet P, Fabbro D. A small synthetic peptide, which inhibits the p53-hdm2 interaction, stimulates the p53 pathway in tumour cell lines. J Mol Biol. 2000;299:245–53.
Vassilev LT, Vu BT, Graves B, Carvajal D, Podlaski F, Filipovic Z, Kong N, Kammlott U, Lukacs C, Klein C, Fotouhi N, Liu EA. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science. 2004;303:844–8.
Brantley Jr MA, Harbour JW. Inactivation of retinoblastoma protein in uveal melanoma by phosphorylation of sites in the COOH-terminal region. Cancer Res. 2000b;60:4320–3.
Pardo M, Pineiro A, De La Fuente M, Garcia A, Prabhakar S, Zitzmann N, Dwek RA, Sanchez-Salorio M, Dominguez F, Capeans C. Abnormal cell cycle regulation in primary human uveal melanoma cultures. J Cell Biochem. 2004;93:708–20.
Scholes AG, Liloglou T, Maloney P, Hagan S, Nunn J, Hiscott P, Damato BE, Grierson I, Field JK. Loss of heterozygosity on chromosomes 3, 9, 13, and 17, including the retinoblastoma locus, in uveal melanoma. Invest Ophthalmol Vis Sci. 2001;42:2472–7.
Halaban R. Rb/E2F: a two-edged sword in the melanocytic system. Cancer Metastasis Rev. 2005;24:339–56.
Harbour JW. Eye cancer: unique insights into oncogenesis: the Cogan Lecture. Invest Ophthalmol Vis Sci. 2006;47:1736–45.
Laurie NA, Donovan SL, Shih CS, Zhang J, Mills N, Fuller C, Teunisse A, Lam S, Ramos Y, Mohan A, Johnson D, Wilson M, Rodriguez-Galindo C, Quarto M, Francoz S, Mendrysa SM, Guy RK, Marine JC, Jochemsen AG, Dyer MA. Inactivation of the p53 pathway in retinoblastoma. Nature. 2006;444:61–6.
El-Shabrawi Y, Radner H, Muellner K, Langmann G, Hoefler G. The role of UV-radiation in the development of conjunctival malignant melanoma. Acta Ophthalmol Scand. 1999;77:31–2.
Gear H, Williams H, Kemp EG, Roberts F. BRAF mutations in conjunctival melanoma. Invest Ophthalmol Vis Sci. 2004;45:2484–8.
Spendlove HE, Damato BE, Humphreys J, Barker KT, Hiscott PS, Houlston RS. BRAF mutations are detectable in conjunctival but not uveal melanomas. Melanoma Res. 2004;14:449–52.
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JW, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR, Futreal PA. Mutations of the BRAF gene in human cancer. Nature. 2002;417:949–54.
Cohen Y, Goldenberg-Cohen N, Parrella P, Chowers I, Merbs SL, Pe’er J, Sidransky D. Lack of BRAF mutation in primary uveal melanoma. Invest Ophthalmol Vis Sci. 2003;44:2876–8.
Cruz 3rd F, Rubin BP, Wilson D, Town A, Schroeder A, Haley A, Bainbridge T, Heinrich MC, Corless CL. Absence of BRAF and NRAS mutations in uveal melanoma. Cancer Res. 2003;63:5761–6.
Edmunds SC, Cree IA, Di Nicolantonio F, Hungerford JL, Hurren JS, Kelsell DP. Absence of BRAF gene mutations in uveal melanomas in contrast to cutaneous melanomas. Br J Cancer. 2003;88:1403–5.
Rimoldi D, Salvi S, Lienard D, Lejeune FJ, Speiser D, Zografos L, Cerottini JC. Lack of BRAF mutations in uveal melanoma. Cancer Res. 2003;63:5712–5.
Zuidervaart W, Van Nieuwpoort F, Stark M, Dijkman R, Packer L, Borgstein AM, Pavey S, Van Der Velden P, Out C, Jager MJ, Hayward NK, Gruis NA. Activation of the MAPK pathway is a common event in uveal melanomas although it rarely occurs through mutation of BRAF or RAS. Br J Cancer. 2005;92:2032–8.
Van Raamsdonk CD, Fitch KR, Fuchs H, De Angelis MH, Barsh GS. Effects of G-protein mutations on skin color. Nat Genet. 2004;36:961–8.
Onken MD, Worley LA, Long MD, Duan S, Council ML, Bowcock AM, Harbour JW. Oncogenic mutations in GNAQ occur early in uveal melanoma. Invest Ophthalmol Vis Sci. 2008b;49:5230–4.
Van Raamsdonk CD, Griewank KG, Crosby MB, Garrido MC, Vemula S, Wiesner T, Obenauf AC, Wackernagel W, Green G, Bouvier N, Sozen MM, Baimukanova G, Roy R, Heguy A, Dolgalev I, Khanin R, Busam K, Speicher MR, O’brien J, Bastian BC. Mutations in GNA11 in uveal melanoma. N Engl J Med. 2010;363:2191–9.
O’hayre M, Vazquez-Prado J, Kufareva I, Stawiski EW, Handel TM, Seshagiri S, Gutkind JS. The emerging mutational landscape of G proteins and G-protein-coupled receptors in cancer. Nat Rev Cancer. 2013;13:412–24.
Oldham WM, Hamm HE. Heterotrimeric G protein activation by G-protein-coupled receptors. Nat Rev Mol Cell Biol. 2008;9:60–71.
Carvajal RD, Sosman JA, Quevedo JF, Milhem MM, Joshua AM, Kudchadkar RR, Linette GP, Gajewski TF, Lutzky J, Lawson DH, Lao CD, Flynn PJ, Albertini MR, Sato T, Lewis K, Doyle A, Ancell K, Panageas KS, Bluth M, Hedvat C, Erinjeri J, Ambrosini G, Marr B, Abramson DH, Dickson MA, Wolchok JD, Chapman PB, Schwartz GK. Effect of selumetinib vs chemotherapy on progression-free survival in uveal melanoma: a randomized clinical trial. JAMA. 2014;311:2397–405.
Carvajal RD, Schwartz GK, Tezel T, Marr B, Francis JH, Nathan PD. Metastatic disease from uveal melanoma: treatment options and future prospects. Br J Ophthalmol. 2017;101(1):38–44.
Johansson P, Aoude LG, Wadt K, Glasson WJ, Warrier SK, Hewitt AW, Kiilgaard JF, Heegaard S, Isaacs T, Franchina M, Ingvar C, Vermeulen T, Whitehead KJ, Schmidt CW, Palmer JM, Symmons J, Gerdes AM, Jonsson G, Hayward NK. Deep sequencing of uveal melanoma identifies a recurrent mutation in PLCB4. Oncotarget. 2016;7:4624–31.
Moore AR, Ceraudo E, Sher JJ, Guan Y, Shoushtari AN, Chang MT, Zhang JQ, Walczak EG, Kazmi MA, Taylor BS, Huber T, Chi P, Sakmar TP, Chen Y. Recurrent activating mutations of G-protein-coupled receptor CYSLTR2 in uveal melanoma. Nat Genet. 2016;48:675–80.
Testa JR, Cheung M, Pei J, Below JE, Tan Y, Sementino E, Cox NJ, Dogan AU, Pass HI, Trusa S, Hesdorffer M, Nasu M, Powers A, Rivera Z, Comertpay S, Tanji M, Gaudino G, Yang H, Carbone M. Germline BAP1 mutations predispose to malignant mesothelioma. Nat Genet. 2011;43:1022–5.
Jensen DE, Proctor M, Marquis ST, Gardner HP, Ha SI, Chodosh LA, Ishov AM, Tommerup N, Vissing H, Sekido Y, Minna J, Borodovsky A, Schultz DC, Wilkinson KD, Maul GG, Barlev N, Berger SL, Prendergast GC, Rauscher 3rd FJ. BAP1: a novel ubiquitin hydrolase which binds to the BRCA1 RING finger and enhances BRCA1-mediated cell growth suppression. Oncogene. 1998;16:1097–112.
Jensen DE, Rauscher 3rd FJ. BAP1, a candidate tumor suppressor protein that interacts with BRCA1. Ann N Y Acad Sci. 1999;886:191–4.
Bommi PV, Dimri M, Sahasrabuddhe AA, Khandekar J, Dimri GP. The polycomb group protein BMI1 is a transcriptional target of HDAC inhibitors. Cell Cycle. 2010;9:2663–73.
Landreville S, Agapova OA, Matatall KA, Kneass ZT, Onken MD, Lee RS, Bowcock AM, Harbour JW. Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma. Clin Cancer Res. 2012;18:408–16.
Harbour JW, Roberson ED, Anbunathan H, Onken MD, Worley LA, Bowcock AM. Recurrent mutations at codon 625 of the splicing factor SF3B1 in uveal melanoma. Nat Genet. 2013;45:133–5.
Martin M, Masshofer L, Temming P, Rahmann S, Metz C, Bornfeld N, Van De Nes J, Klein-Hitpass L, Hinnebusch AG, Horsthemke B, Lohmann DR, Zeschnigk M. Exome sequencing identifies recurrent somatic mutations in EIF1AX and SF3B1 in uveal melanoma with disomy 3. Nat Genet. 2013;45:933–6.
Furney SJ, Pedersen M, Gentien D, Dumont AG, Rapinat A, Desjardins L, Turajlic S, Piperno-Neumann S, De La Grange P, Roman-Roman S, Stern MH, Marais R. SF3B1 mutations are associated with alternative splicing in uveal melanoma. Cancer Discov. 2013;3:1122–9.
Harbour JW. Genomic, prognostic, and cell-signaling advances in uveal melanoma. Am Soc Clin Oncol Educ Book. 2013;2013:388–91.
Yavuzyigitoglu S, Koopmans AE, Verdijk RM, Vaarwater J, Eussen B, Van Bodegom A, Paridaens D, Kilic E, De Klein A, Rotterdam Ocular Melanoma Study, G. Uveal melanomas with SF3B1 mutations: a distinct subclass associated with late-onset metastases. Ophthalmology. 2016;123:1118–28.
Harbour JW. The genetics of uveal melanoma: an emerging framework for targeted therapy. Pigment Cell Melanoma Res. 2012;25:171–81.
Kilic E, Koopmans AE, Yavuzyigitoglu S, Vaarwater J, Van Ijcken WFJ, Paridaens D, De Klein J. SF3B1 and EIF1AX mutations in uveal melanoma: a protective factor, or not? Acta Ophthalmol, 2014;92.
Onken MD, Worley LA, Tuscan MD, Harbour JW. An accurate, clinically feasible multi-gene expression assay for predicting metastasis in uveal melanoma. J Mol Diagn. 2010;12:461–8.
Zuidervaart W, Van Der Velden PA, Hurks MH, Van Nieuwpoort FA, Out-Luiting CJ, Singh AD, Frants RR, Jager MJ, Gruis NA. Gene expression profiling identifies tumour markers potentially playing a role in uveal melanoma development. Br J Cancer. 2003;89:1914–9.
Tschentscher F, Husing J, Holter T, Kruse E, Dresen IG, Jockel KH, Anastassiou G, Schilling H, Bornfeld N, Horsthemke B, Lohmann DR, Zeschnigk M. Tumor classification based on gene expression profiling shows that uveal melanomas with and without monosomy 3 represent two distinct entities. Cancer Res. 2003;63:2578–84.
Onken MD, Worley LA, Davila RM, Char DH, Harbour JW. Prognostic testing in uveal melanoma by transcriptomic profiling of fine needle biopsy specimens. J Mol Diagn. 2006;8:567–73.
Onken MD, Worley LA, Ehlers JP, Harbour JW. Gene expression profiling in uveal melanoma reveals two molecular classes and predicts metastatic death. Cancer Res. 2004;64:7205–9.
Petrausch U, Martus P, Tonnies H, Bechrakis NE, Lenze D, Wansel S, Hummel M, Bornfeld N, Thiel E, Foerster MH, Keilholz U. Significance of gene expression analysis in uveal melanoma in comparison to standard risk factors for risk assessment of subsequent metastases. Eye (Lond). 2008;22:997–1007.
Coupland SE, Kalirai H, Ho V, Thornton S, Damato BE, Heimann H. Concordant chromosome 3 results in paired choroidal melanoma biopsies and subsequent tumour resection specimens. Br J Ophthalmol. 2015;99:1444–50.
Dogrusöz M, Kroes WGM, Van Duinen SG, Creutzberg CL, Versluis M, Bleeker JC, Marinkovic M, Luyten GPM, Jager MJ. Radiation treatment affects chromosome testing in uveal melanoma. Invest Ophthalmol Vis Sci. 2015;56:5956–64.
Hussain RN, Kalirai H, Groenewald C, Kacperek A, Errington RD, Coupland SE, Heimann H, Damato B. Prognostic biopsy of choroidal melanoma after proton beam radiation therapy. Ophthalmology. 2016;123:2264–5.
Wackernagel W, Tarmann L, Mayer C, Langmann G, Wedrich A. Genetic analysis of uveal melanoma by array comparative genomic hybridization before and after radiotherapy. Spektrum Augenheilkd. 2013;27:286–91.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Wackernagel, W., Tarmann, L., Langmann, G., Singh, A.D. (2017). Molecular Carcinogenesis of Uveal Melanoma. In: Haybaeck, J. (eds) Mechanisms of Molecular Carcinogenesis – Volume 1. Springer, Cham. https://doi.org/10.1007/978-3-319-53659-0_9
Download citation
DOI: https://doi.org/10.1007/978-3-319-53659-0_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-53657-6
Online ISBN: 978-3-319-53659-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)