Abstract
During wound repair, embryonic development, and cancer invasion, cells migrate in cooperative groups and clusters. Collective migration has been studied for decades, for example with experiments to determine how quickly a sheet of cells migrates to close a wound. Until recently, those experiments have been limited to investigating molecular mechanisms due to a lack of experimental tools to quantify the kinematics and forces. Now that digital image correlation has gained wide acceptance in engineering, physics, and biology, new possibilities exist to reveal quantitative understanding of the connections between signaling, motions, and forces. Here we review applications of digital image correlation in measuring cell velocities, cell-to-substrate tractions, and cell-to-cell stresses. The data show that forces and motions follow no typical constitutive relationship such as that of an elastic solid or a viscous fluid; in many cases even the orientations of force and motion are misaligned. Ongoing research is seeking to connect force and motion, often by modeling the active processes associated with cell signaling. While it remains unclear how to reduce the complicated and numerous signaling pathways into a physical picture of collective cell migration, the experimental tools now available offer a useful place to start.
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
Keywords
- Digital image correlation
- Traction force microscopy
- Monolayer stress microscopy
- Cell mechanics
- Cell migration
12.1 Introduction
In pattern formation, would healing, and invasive cancer, cells move most often not as individual entities but as collective sheets, ducts, strands, or clusters. Groups of collectively moving cells also form within a cellular monolayer, resulting in fluctuations in velocity and local cellular density [1]. Together with these fluctuations in kinematics, fluctuations in the tensile stress between cells result in a rugged stress landscape [2]. The fluctuations have been linked to a change in phase from a fluid-like unjammed state to a solid-like jammed one [1–3]. In the unjammed state, cells can move freely past their neighbors whereas in the jammed state, each cell within the monolayer remains stuck in place. To quantify this collective motion, we use digital image correlation on phase contrast images of the cell monolayer. The force required to move is generated internally by each cell and applied to both the substrate beneath the cell and the cell’s immediate neighbors. To begin to relate the active cell forces to the motions, we describe experimental techniques for quantifying both force and motion here.
12.2 Experimental Methods
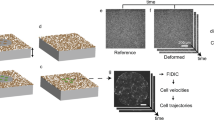
Here we give a summary of our experimental procedures, which include digital image correlation (DIC), traction force microscopy, and monolayer stress microscopy. Full details are in our recent publication [4]. Cells are typically micropatterned into circular monolayers of diameter 1 mm by using standard micropatterning techniques to adhere adhesive collagen to the substrate. The substrate is a compliant gel, typically polyacrylamide, embedded with fluorescent particles. The particles serve as a high contrast pattern to quantify the substrate’s displacements using DIC. We centrifuge the gel upside down during polymerization, which is sufficient to move the particles to a thin layer at the top of the gel. This thin layer allows for the microscope to focus exactly on the particles and prevents errors due to imaging particles that are out of the focal plane. The displacements in the substrate result from the tractions applied by the cells. To compute tractions from displacements we used the Boussinesq-Cerruti solution for the displacement field induced by a point force on a semi-infinite half space [5] as a Green’s function. Solving for tractions required an inverse solution, which we have implemented in the Fourier domain [6] with a correction for a substrate of finite thickness [7, 8]. We then apply monolayer stress microscopy [2, 9] to compute the two dimensional (2D) stress tensor σ ij within the monolayer. The basic idea of monolayer stress microscopy is illustrated in Fig. 12.1. Stresses within the monolayer are computed from a summation of the tractions, much like stress builds up in the rope of a tug-of-war game. In two dimensions, stresses are computed using the stress equilibrium equations, ∂σ ij /∂x j + b i = 0, i , j = 1 , 2, with body force b i . For a 2D monolayer, the traction t i , applied by the substrate to the monolayer, acts as the body force, which is given by b i = t i /h where h is the height of the monolayer [2, 9]. The 2D stress tensor has three unique components whereas there are only two equilibrium equations. To solve for all three unique components of the stress tensor, a third equation is required. The third equation we use is compatibility, which assumes the displacement or velocity field of the monolayer is continuous. Note that the earliest version of monolayer stress microscopy assumed the monolayer to be a linear elastic solid, but we have recently shown that the results are identical if the monolayer is a viscous fluid [4]. Thus the only assumptions are that the monolayer can be treated as a continuum and that there is a linear relationship between gradients in cell velocity and stress. It has been shown that dependence on these assumptions in minimal [9], which makes sense given that two of the three equations are simply force equilibrium which make no assumptions on the constitutive properties of the monolayer.
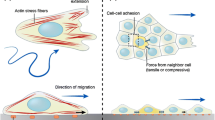
Stresses within the monolayer are computed with monolayer stress microscopy. (a) Each cell within the monolayer applies traction to the substrate beneath it (red arrows) and stress to its neighbors (blue arrows). (b) The intercellular stresses are computed by summing the tractions, similar to how tension builds up in the rope of a tug-of-war game. Image adapted from Ref. [10]
12.3 Results
Typical results are shown in Fig. 12.2. Cell velocities and tractions are both vector quantities; the components in the radial direction are plotted here. The stress is a tensor representing the in-plane normal and shear stresses within the monolayer. The principal stresses are shown in the figure. The two principal stresses are nearly always positive, indicating that there is only tension, no pressure, within the monolayer. This finding is consistent with the notion that cells are contractile and therefore generate tensile stresses.
Typical results showing cell image, radial component of velocity, radial component of traction, first principal stress (σ1) and second principal stress (σ2). For velocity and traction, positive is defined as outward from the center of the cell island. Notice that both principal stresses are positive, indicating there are no compressive stresses within the monolayer
References
Angelini, T.E., Hannezo, E., Trepat, X., Marquez, M., Fredberg, J.J., Weitz, D.A.: Proc. Natl. Acad. Sci. U. S. A. 108, 4714–4719 (2011)
Tambe, D.T., Hardin, C.C., Angelini, T.E., Rajendran, K., Park, C.Y., Serra-Picamal, X., Zhou, E.H., Zaman, M.H., Butler, J.P., Weitz, D.A., Fredberg, J.J., Trepat, X.: Nat. Mater. 10, 469–475 (2011)
Sadati, M., Taheri Qazvini, N., Krishnan, R., Park, C.Y., Fredberg, J.J.: Differentiation. 86, 121–125 (2013)
Notbohm, J., Banerjee, S., Utuje, K.J., Gweon, B., Jang, H., Park, Y., Shin, J., Butler, J.P., Fredberg, J.J., Marchetti, M.C.: Biophys. J. 110, 2729–2738 (2016)
Landau, L.D., Lifshitz, E.M.: Theory of Elasticity, 3rd edn. Pergamon Press (1986)
Butler, J.P., Tolic-Norrelykke, I.M., Fabry, B., Fredberg, J.J.: Am. J. Physiol.-Cell Phys. 282, C595–C605 (2002)
del Alamo, J.C., Meili, R., Alonso-Latorre, B., Rodriguez-Rodriguez, J., Aliseda, A., Firtel, R.A., Lasheras, J.C.: Proc. Natl. Acad. Sci. U. S. A. 104, 13343–13348 (2007)
Trepat, X., Wasserman, M.R., Angelini, T.E., Millet, E., Weitz, D.A., Butler, J.P., Fredberg, J.J.: Nat. Phys. 5, 426–430 (2009)
Tambe, D.T., Croutelle, U., Trepat, X., Park, C.Y., Kim, J.H., Millet, E., Butler, J.P., Fredberg, J.J.: PLoS One. 8, e55172 (2013)
Trepat, X., Fredberg, J.J.: Trends Cell Biol. 21, 638–646 (2011)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 The Society for Experimental Mechanics, Inc.
About this paper
Cite this paper
Saraswathibhatla, A., Notbohm, J. (2017). Applications of DIC in the Mechanics of Collective Cell Migration. In: Sutton, M., Reu, P. (eds) International Digital Imaging Correlation Society. Conference Proceedings of the Society for Experimental Mechanics Series. Springer, Cham. https://doi.org/10.1007/978-3-319-51439-0_12
Download citation
DOI: https://doi.org/10.1007/978-3-319-51439-0_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-51438-3
Online ISBN: 978-3-319-51439-0
eBook Packages: EngineeringEngineering (R0)