Abstract
MicroRNAs are a critical class of regulators for cells to deal with DNA damage. Abnormal miRNA function is associated with tumor initiation and progression, and altered miRNA expression found in tumor tissues are frequently associated with heterogeneity of tumor responses to therapeutic agents, including radiotherapy. In this chapter, we review recent advances of the functional role of microRNAs in the context of the DNA damage response, tissue specific tumor initiation and progression. We further discuss clinical implications of using miRNA signatures as biomarkers for radiosensitivity and targeting specific miRNAs as therapeutic approaches.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
8.1 Introduction
Radiotherapy is used to treat more than half of patients diagnosed with cancer, either as the primary mode of treatment or in combination with chemotherapy or surgical resection. The success of radiotherapy relies on the inability of cancerous cells to efficiently repair DNA damage relative to their normal tissue counterparts, pushing the cancerous cells into death pathways. Specifically, ionizing radiation (IR) is intended to induce DNA damage, engage the DNA double-stranded break repair machinery, and to push the cell into mitotic catastrophe, apoptosis, or stress-induced senescence. Some tumors, however, exhibit an insensitivity to an otherwise curative dose and are deemed radioresistant. Tumor radioresistance is a common problem linked to tumor heterogeneity and underlying biochemical factors such as abnormal DNA damage response pathways, microenvironment alterations, deregulated survival pathways, and altered expression of oncogenes and tumor suppressors. An increasing number of studies have shown that regulation of these pathways is modulated in part by microRNAs.
MicroRNAs (miR) are highly conserved, small, non-coding RNAs that are involved post-transcriptional regulation of target mRNAs, and also regulate roughly 30 % of human genes at the DNA level [1]. Biogenesis of these miRs involves a series of enzymatic cleavages that begins with primary microRNA transcripts (primiRNAs). These primiRNAs are converted to hairpin pre-miRs via activity of the Drosha/DGCR8 complex for export out of the nucleus and into the cytoplasm. Finally, the pre-miRs are cleaved by a Dicer, a RNase, and the cleaved mature miR is assembled into the RISC complex. The RISC then targets a specific mRNA for repression and degradation [2]. Regulation of protein expression subsequently affects the pathways in which the proteins function, resulting in a measurable change in intracellular processes. MiR biogenesis can be triggered by a number of external and internal cellular signals. Here the focus will be on the miR expression patterns, and the affected pathways in the context of ionizing radiation (IR), the DNA damage response (DDR), and tumor progression.
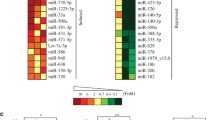
Exogenous genotoxic agents, in the form of IR and oxidative stress inducers, have been shown to influence the biogenesis of a certain subset of the identified ~3700 miRs [3], and within that subset there are tumor-type specific miR expression profiles [4]. While there are type-specific differences in miR expression, there also exists a significant overlap [Fig. 8.1]. So, it is within the differential expression, coupled with the common alterations, that diagnostic and therapeutic approaches can be taken to determine on a patient-by-patient basis whether radioresistance is likely. The use of miRs as prognosticators of tumor sensitivity to ionizing radiation has clinical significance in determining the best course of action for treatment, sparing the patient from therapies that are not viable options based on tissue micro-environment.
8.2 The Role of miRs in the DDR
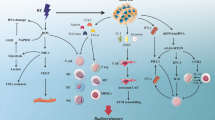
A properly functioning DDR is essential for the maintenance of genomic integrity. Double-stranded breaks (DSBs) trigger activation of the Ataxia-Telangiectasia Mutated (ATM) kinase which results in the phosphorylation of H2AX at lesion sites and recruitment of repair proteins. This process stalls cell cycle progression until the lesions are repaired, or, if the damage is too catastrophic, transitions the cell into senescence and apoptosis. The DDR pathway is extensively regulated by miRs and components of the miR biogenesis pathway. Specifically, the enzymatic functions of Dicer and Drosha regulate the expression of miRs that target pATM and its substrates that subsequently sense and form foci at DNA damage sites [5]. ATM is also directly modulated by miR-18a, or indirectly by miR-421 and miR-106a, underscoring the importance of regulation of the DDR pathway via ATM activity [6–8]. Additionally, DNA damage directly regulates biogenesis of a small subset of miRs linked to Drosha/DGCR8 by complexing with p68 and p72 to facilitate processing of pri-miRs to pre-miRs. ATM activates KH-type splicing regulatory protein (KSRP) which also complexes with Drosha/DCGR8 to allow for pri-miR processing [9] [Fig. 8.2.].
During the course of the DDR, nearly all major players involved in the process of clearing the damage are subject to direct regulation by miRs. From the damage sensor H2AX, to ATM as a signal transducer, to downstream effector pathways miRs target and directly regulate key components of the process. Indirectly, miRs modulate the expression of upstream regulators of the process in order to provide a fine-tuning of the pathway [10]. The extent to which the DDR, gatekeeper of genomic stability, is regulated by miRs underscores the importance of proper expression and function of these RNAs in this pathway [Fig. 8.3].
8.3 MiRs Expression in Carcinogenesis
MiRs, by their nature, regulate both tumor suppressor genes and oncogenes, and are divided into oncomircroRNAs which regulate tumor suppressors or anti-oncomicroRNAs which regulate oncogenes [11]. This regulation depends on the tissue in which the miR is expressed. The first indication that miRs could be implicated in cancer pathology was from the early descriptive studies in C. elegans and drosophila model systems where mutations in let-7 resulted in loss-of-function phenotypes as seen in loss of proliferation regulation [12, 13]. The association of miR expression with carcinogenesis was shown initially in chronic lymphocytic leukemia (CLL) which linked loss of expression of miRs 15 and 16 to disease progression [14]. This was the first in a series of studies that critically examined the miR expression patterns, mutation rates, and physiological consequences in neoplastic tissue arising from every tissue type [4]. Determination of the gene locus of these miRs found that a majority of these coding sites can be found in fragile sites, susceptible to alteration, and also in genomic regions frequently associated with carcinogenesis [15]. The distribution of these genes is not random as most of these coding regions are positioned to be flanking oncogenes and common translocation sites that result in altered expression or even deletion of these miRs [16]. As such, a stressed system is likely to expose these fragile sites to damage [17].
The effect of genomic changes to miR sequences or transcription rates is compounded because of the mechanisms by which miRs are processed into maturity. MiR clusters encompass precursors to mature miRNA products; as such alterations in any given cluster can have wide-ranging, deleterious effects. The localization of these miR coding genes at fragile sites, combined with the documented high mutation rate is further affected by alterations in protein-coding genes crucial in miR biogenesis, specifically Dicer and Argonaute coding genes [16]. Such alterations fall into the following catagories: (1) loss of miR expression due to deletion, transcription error, or mutation, (2) over-expression due to gene translocation, and (3) altered expression due to changes in the biogenesis pathway and machinery. The physiological consequences manifest themselves in the form of hyper-proliferation, evasion of apoptosis, and invasiveness due to the inability to properly regulate target mRNAs.
8.4 MiRs as Biomarkers for Radiation Sensitivity
An ideal biomarker meets the following characteristics: it must be specific to the pathology in question, is rapidly detectable upon onset of the pathology, is proportional, or inversely proportional, to the severity of the pathology, is a preclinical predictor of a clinical outcome, and is readily accessible [18]. The quest for biomarkers as prognosticators for positive therapeutic outcomes has focused attention on circulating miRs. Since miRs are critical regulatory elements of key pathways, changes in their expression levels could be reliably linked to the mRNA targets and subsequent pathways they regulate. The discovery of detectable miRs in serum and other bodily fluids points to use of these extracellular miRs as potential biomarkers. These miRs are packaged in such a way as to avoid RNase digestion and likely act as cell-to-cell communicators, and because of their stability and specificity, can indicate tissue specific pathology.
Normal, non-cancerous tissues exhibit a predictable change in miR expression in response to radiation. Notably, the biogenesis of miRs that regulate cell cycle progression and DDR are regulated in a dose-dependent manner to IR [19, 20]. Acting as anti-oncomiRs, the let-7 cluster of miRs is linked to cell cycle progression and apoptosis regulation through regulation of KRAS, while the miR-34 family is targeted by p53 to halt cell cycle progression and modulate apoptosis proteins [21]. Additionally, miR-21 is reliably up-regulated in both normal and cancerous tissue and is reported to target key components in the apoptotic pathway such as the transcript for programmed cell death 4 (hPDCD4) [22]. At the front end of the DDR, initiated by ionizing radiation-induced double-stranded DNA breaks, ATM expression is regulated by miR-421 [8] and miR-101 [23] while at the back end, H2AX, a histone variant phosphorylated by ATM, is targeted by miRs-24 and -138 [24, 25]. Over-expression of any of these miRs results in down-regulation of their protein targets and subsequently leads to accumulation of chromosomal damage and sensitivity to IR.
The pathway-specificity and ubiquitous expression profiles of these few miRs provide promising biomarkers for predicting radiosensitivity. Nearly all have direct targets at key non-redundant junctures in the DDR, cell-cycle, proliferation and apoptotic pathways. The differential expression in normal tissues upon treatment with ionizing radiation gives a baseline with which to measure efficient pathway execution when compared to cancerous tissue. The ability to detect and quantify these miRs prior to and in the course of treatment has the potential to increase therapeutic efficacy and improve patient outcome. Use of patient databases such as The Cancer Genome Atlas (http://tcga-data.nci.nih.gov) and cBioPortal (http://www.cbioportal.org) provide standardized sets of data gathered from patient biopsies. Meta-analysis of these datasets links expression patterns in tumor types to survivability and treatment efficacy, identifying clinically relevant miRs that may be key regulators in the response to radiation. These studies are currently being done for most tissue specific cancers from head and neck cancers to glioblastomas [26–28].
Can circulating miRs be used to predict radiosensitivity? Recent studies have indicated that, yes, tumor specific circulating miRs can be detected in blood serum. Examples of this can be found in breast cancer [29–31], and in a recent study that showed xenograft-specific miR detection in the blood serum of mice correlated to clinical samples from patients with pancreatic, lung and colorectal cancers [32]. The real key to the feasibility of miRs as biomarkers in a clinical setting is that the detectable, circulating miRs at least partially overlap with the tumor-specific ones. To date, radiosensitivity miR biomarkers have been identified in every major tissue specific cancer [Table 8.1]. The development of clinical assays to rapidly and accurately evaluate the radiosensitivity of cancers depends in part on the ability to isolate and detect miRs in a minimally invasive manner.
8.5 MiRs as Therapeutic Targets to Enhance Radiation Efficacy
Given the baseline expression of key miRs involved in the radiation response, coupled with changes in expression when exposed to radiation, a list of therapeutic targets begins to emerge in disregulated systems. The therapeutic avenues for rescuing normal phenotypes are to either inhibit the activity of over-expressed miRs or exogenously express deficient miRs. The major obstacle to this is the ability to package and deliver the therapeutic agent in a way that will ensure both stability and tissue specificity. Current strategies in miR-based cancer therapy have used modified, or locked, nucleic acids to competitively bind and inhibit mature miRs [33]. To combat under-expression or deletion of key miRs, synthetic naked miRs can be loaded into tagged lipid vesicles or nanoparticles for delivery [34]. Viral delivery of deficient miRs has shown promise in laboratory settings and even in vivo, but the concern of chromosomal integration and other off target effects has limited success in pre-clinical and clinical trials [35, 36].
A less direct method to affect miRNA expression is through the use of drug compounds to induce miR biogenesis pathways subsequently resulting in a synthetic lethality in cancerous cells when coupled with radiation treatment. A prime example of this is the identification of MiRNA-145 as an indicator of tumor sensitivity to radiation in prostate cancer [37]. This miR also plays a role in ovarian cancer where its expression can be induced by the flavonoid quecetin subsequently inducing apoptosis [38]. Also within the class of flavonoids, Rhamnetin and cirsiliol have been shown to induce miR-34a expression. MiR-34a inhibits Notch-1 expression and renders tumors more susceptible to IR treatment [39]. Another example is cantharidin, a terpenoid, affecting the expression of miR-214 regulating p53-mediated apoptotic pathway to swing the Bax/Bcl-2 balance towards cell death [40]. The use of these compounds and others like them depends heavily on identification and targeting of the tissue specific miR responsible for pathway dysregulation.
8.6 Clinical Significance
The extent to which miRs regulate critical survival and repair pathways in a cell makes these small oligonucleotides ideal as predictors and therapeutic targets in the treatment of a wide array of cancers. MiR biogenesis pathways are incredibly sensitive to internal and external changes, allowing for the characterization of expression patterns in the face of DNA damage resulting from ionizing radiation. It is important to establish and validate cancer-specific biomarkers using meta-data analysis and bench-top verification of radiosensitive miR signatures for relevant carryover into the clinical setting. Properly vetted biomarkers can be used both as prognosticators as to whether the cancerous tissue will be responsive to therapeutic doses of radiation, and if readily accessible, as with circulating miRs, can be assessed during the course of treatment to assure that the cells are responding to treatment. This is beneficial from the perspective of patient wellbeing because radiotherapy can be discounted out right, fine-tuned in dosage, or coupled with other therapies with greater confidence in the sensitivity of the cells to treatment.
As clinically relevant therapeutic targets, miRs can either be inhibited or induced. The balance in this strategy is in the delivery mechanism and ensuring that drug delivery is tumor specific with limited off target effects. Because of the nature of miR regulation, in that one mature miR has multiple mRNA targets, it is critical to target miRs with direct effects on pathways involved in DNA damage. Over-expressed miRs are an easier target to address because the drug or oligomimetic cargo can be packaged in such a way to direct the therapy to specific cells with specific receptors. Drug-induced specific miR biogenesis with the intent of weakening a survival or repair pathway could prove more challenging. A more focused approach to manipulating miR expression is needed to effectively produce the synthetically lethal radiosensitivity. As a class of nucleotides though, miRs are emerging as critical components.
References
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20.
Garzon R, Calin GA, Croce CM. MicroRNAs in cancer. Annu Rev Med. 2009;60:167–79.
Londin E, Loher P, Telonis AG, Quann K, Clark P, Jing Y, Hatzimichael E, Kirino Y, Honda S, Lally M, Ramratnam B, Comstock CE, Knudsen KE, Gomella L, Spaeth GL, Hark L, Katz LJ, Witkiewicz A, Rostami A, Jimenez SA, Hollingsworth MA, Yeh JJ, Shaw CA, Mckenzie SE, Bray P, Nelson PT, Zupo S, van Roosbroeck K, Keating MJ, Calin GA, Yeo C, Jimbo M, Cozzitorto J, Brody JR, Delgrosso K, Mattick JS, Fortina P, Rigoutsos I. Analysis of 13 cell types reveals evidence for the expression of numerous novel primate- and tissue-specific microRNAs. Proc Natl Acad Sci U S A. 2015;112:E1106–15.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Francia S, Michelini F, Saxena A, Tang D, de Hoon M, Anelli V, Mione M, Carninci P, d’adda di Fagagna F. Site-specific DICER and DROSHA RNA products control the DNA-damage response. Nature. 2012;488:231–5.
Boohaker RJ, Xu B. The versatile functions of ATM kinase. Biomed J. 2014;37:3–9.
Guo X, Yang C, Qian X, Lei T, Li Y, Shen H, Fu L, Xu B. Estrogen receptor alpha regulates ATM Expression through miRNAs in breast cancer. Clin Cancer Res. 2013;19:4994–5002.
Hu H, Du L, Nagabayashi G, Seeger RC, Gatti RA. ATM is down-regulated by N-Myc-regulated microRNA-421. Proc Natl Acad Sci U S A. 2010;107:1506–11.
Liu Y, Liu Q. ATM signals miRNA biogenesis through KSRP. Mol Cell. 2011;41:367–8.
Liu Y, Lu X. Non-coding RNAs in DNA damage response. Am J Cancer Res. 2012;2:658–75.
Fabbri M, Ivan M, Cimmino A, Negrini M, Calin GA. Regulatory mechanisms of microRNAs involvement in cancer. Expert Opin Biol Ther. 2007;7:1009–19.
Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell. 2003;113:25–36.
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE, Horvitz HR, Ruvkun G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature. 2000;403:901–6.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, Aldler H, Rattan S, Keating M, Rai K, Rassenti L, Kipps T, Negrini M, Bullrich F, Croce CM. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A. 2002;99:15524–9.
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M, CROCE CM. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci U S A. 2004;101:2999–3004.
Zhang L, Huang J, Yang N, Greshock J, Megraw MS, Giannakakis A, Liang S, Naylor TL, Barchetti A, Ward MR, Yao G, Medina A, O’brien-Jenkins A, Katsaros D, Hatzigeorgiou A, Gimotty PA, Weber BL, Coukos G. microRNAs exhibit high frequency genomic alterations in human cancer. Proc Natl Acad Sci U S A. 2006;103:9136–41.
Mendell JT, Olson EN. MicroRNAs in stress signaling and human disease. Cell. 2012;148:1172–87.
Etheridge A, Lee I, Hood L, Galas D, Wang K. Extracellular microRNA: a new source of biomarkers. Mutat Res. 2011;717:85–90.
Lee KF, Chen YC, Hsu PW, Liu IY, Wu LS. MicroRNA expression profiling altered by variant dosage of radiation exposure. Biomed Res Int. 2014;2014:456323.
Metheetrairut C, Slack FJ. MicroRNAs in the ionizing radiation response and in radiotherapy. Curr Opin Genet Dev. 2013;23:12–9.
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A, Labourier E, Reinert KL, Brown D, Slack FJ. RAS is regulated by the let-7 microRNA family. Cell. 2005;120:635–47.
Chaudhry MA, Omaruddin RA, Kreger B, DE Toledo SM, Azzam EI. Micro RNA responses to chronic or acute exposures to low dose ionizing radiation. Mol Biol Rep. 2012;39:7549–58.
Yan D, Ng WL, Zhang X, Wang P, Zhang Z, Mo YY, Mao H, Hao C, Olson JJ, Curran WJ, Wang Y. Targeting DNA-PKcs and ATM with miR-101 sensitizes tumors to radiation. PLoS One. 2010;5:e11397.
Lal A, Pan Y, Navarro F, Dykxhoorn DM, Moreau L, Meire E, Bentwich Z, Lieberman J, Chowdhury D. miR-24-mediated downregulation of H2AX suppresses DNA repair in terminally differentiated blood cells. Nat Struct Mol Biol. 2009;16:492–8.
Wang Y, Scheiber MN, Neumann C, Calin GA, Zhou D. MicroRNA regulation of ionizing radiation-induced premature senescence. Int J Radiat Oncol Biol Phys. 2011;81:839–48.
Liu N, Boohaker RJ, Jiang C, Boohaker JR, Xu B. A radiosensitivity MiRNA signature validated by the TCGA database for head and neck squamous cell carcinomas. Oncotarget. 2015;6:34649–57.
Moskwa P, Zinn PO, Choi YE, Shukla SA, Fendler W, Chen CC, Lu J, Golub TR, Hjelmeland A, Chowdhury D. A functional screen identifies miRs that induce radioresistance in glioblastomas. Mol Cancer Res. 2014;12:1767–78.
Zhan C, Yan L, Wang L, Jiang W, Zhang Y, Xi J, Chen L, Jin Y, Qiao Y, Shi Y, Wang Q. Identification of reference miRNAs in human tumors by TCGA miRNA-seq data. Biochem Biophys Res Commun. 2014;453:375–8.
Kodahl AR, Lyng MB, Binder H, Cold S, Gravgaard K, Knoop AS, Ditzel HJ. Novel circulating microRNA signature as a potential non-invasive multi-marker test in ER-positive early-stage breast cancer: a case control study. Mol Oncol. 2014;8:874–83.
Mar-Aguilar F, Mendoza-Ramirez JA, Malagon-Santiago I, Espino-Silva PK, Santuario-Facio SK, Ruiz-Flores P, Rodriguez-Padilla C, Resendez-Perez D. Serum circulating microRNA profiling for identification of potential breast cancer biomarkers. Dis Markers. 2013;34:163–9.
Schrauder MG, Strick R, Schulz-Wendtland R, Strissel PL, Kahmann L, Loehberg CR, Lux MP, Jud SM, Hartmann A, Hein A, Bayer CM, Bani MR, Richter S, Adamietz BR, Wenkel E, Rauh C, Beckmann MW, Fasching PA. Circulating micro-RNAs as potential blood-based markers for early stage breast cancer detection. PLoS One. 2012;7:e29770.
Greystoke A, Ayub M, Rothwell DG, Morris D, Burt D, Hodgkinson CL, Morrow CJ, Smith N, Aung K, Valle J, Carter L, Blackhall F, Dive C, Brady G. Development of a circulating miRNA assay to monitor tumor burden: from mouse to man. Mol Oncol. 2015;10(2):282–91.
Lennox KA, Behlke MA. Chemical modification and design of anti-miRNA oligonucleotides. Gene Ther. 2011;18:1111–20.
Huang X, Schwind S, Yu B, Santhanam R, Wang H, Hoellerbauer P, Mims A, Klisovic R, Walker AR, Chan KK, Blum W, Perrotti D, Byrd JC, Bloomfield CD, Caligiuri MA, Lee RJ, Garzon R, Muthusamy N, Lee LJ, Marcucci G. Targeted delivery of microRNA-29b by transferrin-conjugated anionic lipopolyplex nanoparticles: a novel therapeutic strategy in acute myeloid leukemia. Clin Cancer Res. 2013;19:2355–67.
Broderick JA, Zamore PD. MicroRNA therapeutics. Gene Ther. 2011;18:1104–10.
Thomas CE, Ehrhardt A, Kay MA. Progress and problems with the use of viral vectors for gene therapy. Nat Rev Genet. 2003;4:346–58.
Gong P, Zhang T, He D, Hsieh JT. MicroRNA-145 modulates tumor sensitivity to radiation in prostate cancer. Radiat Res. 2015;184:630–8.
Zhou J, Gong J, Ding C, Chen G. Quercetin induces the apoptosis of human ovarian carcinoma cells by upregulating the expression of microRNA-145. Mol Med Rep. 2015;12:3127–31.
Kang J, Kim E, Kim W, Seong KM, Youn H, Kim JW, Kim J, Youn B. Rhamnetin and cirsiliol induce radiosensitization and inhibition of epithelial-mesenchymal transition (EMT) by miR-34a-mediated suppression of Notch-1 expression in non-small cell lung cancer cell lines. J Biol Chem. 2013;288:27343–57.
Tian X, Zeng G, Li X, Wu Z, Wang L. Cantharidin inhibits cell proliferation and promotes apoptosis in tongue squamous cell carcinoma through suppression of miR-214 and regulation of p53 and Bcl-2/Bax. Oncol Rep. 2015;33:3061–8.
Czochor JR, Glazer PM. microRNAs in cancer cell response to ionizing radiation. Antioxid Redox Signal. 2014;21:293–312.
Wang Y, Huang JW, Li M, Cavenee WK, Mitchell PS, Zhou X, Tewari M, Furnari FB, Taniguchi T. MicroRNA-138 modulates DNA damage response by repressing histone H2AX expression. Mol Cancer Res. 2011;9:1100–11.
Simone NL, Soule BP, Ly D, Saleh AD, Savage JE, Degraff W, Cook J, Harris CC, Gius D, Mitchell JB. Ionizing radiation-induced oxidative stress alters miRNA expression. PLoS One. 2009;4:e6377.
Brozovic A, Duran GE, Wang YC, Francisco EB, Sikic BI. The miR-200 family differentially regulates sensitivity to paclitaxel and carboplatin in human ovarian carcinoma OVCAR-3 and MES-OV cells. Mol Oncol. 2015;9:1678–93.
Muratsu-Ikeda S, Nangaku M, Ikeda Y, Tanaka T, Wada T, Inagi R. Downregulation of miR-205 modulates cell susceptibility to oxidative and endoplasmic reticulum stresses in renal tubular cells. PLoS One. 2012;7:e41462.
Lafferty-Whyte K, Cairney CJ, Jamieson NB, Oien KA, Keith WN. Pathway analysis of senescence-associated miRNA targets reveals common processes to different senescence induction mechanisms. Biochim Biophys Acta. 2009;1792:341–52.
Marta GN, Garicochea B, Carvalho AL, Real JM, Kowalski LP. MicroRNAs, cancer and ionizing radiation: where are we? Rev Assoc Med Bras. 2015;61:275–81.
Acknowledgements
This work was supported in part by NIH grants R01CA133093 and R01ES016354, the Alabama Innovation Fund, and Alabama Drug Discovery Alliance.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Boohaker, R.J., Xu, B. (2016). The Role of MicroRNAs in Modulating Tissue Response to Radiation. In: Anscher, M., Valerie, K. (eds) Strategies to Enhance the Therapeutic Ratio of Radiation as a Cancer Treatment. Springer, Cham. https://doi.org/10.1007/978-3-319-45594-5_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-45594-5_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-45592-1
Online ISBN: 978-3-319-45594-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)