Abstract
The PURESystem™(for short: PS) is a defined set of about 80 different macromolecular species which can perform protein synthesis starting from a coding DNA. To understand the processes that take place inside a liposome with entrapped PS, several simulation approaches, of either a deterministic or stochastic nature, have been proposed in the literature. To correctly describe some peculiar phenomena that are observed only in very small liposomes (such as power-law distribution of solutes and supercrowding effect), a stochastic approach seems necessary, due to the very small average number of molecules contained in these liposomes. Here we recall the results reported in other works published by us and by other Authors, discussing the importance of a stochastic simulation approach and of a fine description of the system: both these aspects, in fact, were not properly acknowledged in such previous papers.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Among the different cell-free protein synthesis systems, the PUREsystem™ (Protein synthesis Using Recombinant Elements; PS for short hereafter), developed in 2001 by Ueda and coworkers [1] and commercially available, is one of the most known and employed, in particular in synthetic biology studies. Despite the relatively small number of chemical species contained in it (mainly DNA, ribosomes, a tRNA mixture, 36 purified E. coli enzymes, plus several small metabolites, for a total of about 80 different macromolecular species), it shows excellent performance in producing proteins starting from coding DNA [2]. Its description and preparation meet the standardization requirements of synthetic biology, it is listed in the Registry of Standard Biological Parts [3].
The studies of biochemical pathways in vitro proceeded in parallel to those performed in vivo, since in vitro approaches allow to test or measure quantities, kinetic parameters and other variables, that are not easily accessible in vivo. Together with the experiments, the necessity of a model able to describe the principal pathways lead several Authors to propose different strategies, focused on different pathways and using different detail level and different formal approaches. From early works where cell-free synthesis has been used to synthetize different isoforms of a protein [4], to more recent models where protein synthesis is studied in vivo or in vitro, by either “standard” differential-equation based models [5] or stochastic approaches to understand the role of the noise in the main biochemical patways [6, 7], trying to investigate emergent features due to the biochemical network complexity [8].
Apart from these general approaches, the PS has been successfully applied in lipo Footnote 1, allowing the possibility to monitor and study gene expression in small volumes.
For instance, an interesting study used the PS to study membrane proteins and pore-complex assembly using a novel approach called liposome display, where different DNA constructs are used as a template for in vitro protein production and assembly on the liposome membrane [9]. Other numerous applications of the compartmentalized PS were focused on understanding gene expression inside liposomes, such as in the presence of different membrane lipid composition [10], with different preparation methods [11], and in different-sized vesicles [12].
Understanding gene expression in small compartments is of crucial importance for defining the so-called minimal cell [10, 13, 14] a minimal entity able to display the fundamental properties of living system; the PS is the most used system for the construction of semi-synthetic minimal cells [13, 14].
Indeed, semi-synthetic minimal cells often consist of a liposome enclosing defined biochemical pathways, specifically pathways capable to synthesize proteins that could, in principle, close the circle towards a complete self-sustainability of the minimal cell: in other words, the different components of the minimal cells can cooperate to restore themselves. As of today, only a few papers have been published dealing with theoretical simulations, at different detail level, of PS behavior and time course [7, 15–17], and only three deal with PS entrapped in liposomes [18–20]. This last condition is very interesting since the smaller the liposome, the lower the probability that a large amount of molecules of each constituent chemical species be entrapped in the volume of the liposome. This implies that a standard simulation of PS based on deterministic ODE formalism is no longer suitable to describe a biochemical system beyond the deterministic limit. If even a single chemical species is present with only a few molecules, the behavior of all the PS is turned to stochastic.
To describe a stochastic chemical system, the most often used approach is to employ Gillespie algorithm [21]: this is an algorithm derived from collision theory that operates by selecting, by means of two suitably generated random numbers, which reaction r to execute at the current step, and the waiting time\(\tau \) for the next reaction. The probability distribution of r is uniform over the propensity of the reactions, while the \(\tau \) distribution is an exponential decay (i.e., the reaction system is described as a pure Markovian process). Gillespies algorithm has been proved to be correct: for large numbers of molecules and for long times, the time courses described by the algorithm are identical to those described by the Chemical Master Equation for the continuous case. Because of this characteristic, Gillespies algorithm (with several variants and improvements) has been applied in a very large number of stochastic simulations of chemical, and even biochemical, systems [22].
2 Describing the PS in a Stochastic Simulator
There are several problems to address in order to have a detailed description of the PS in a Gillespies algorithm-based simulator. These problems arise from the not negligible differences between a common chemical system as hypothesized by Gillespie, and a cell-free system, which hosts several complex multi-step processes. For instance, two major problems are represented by the description of transcription and translation: as they imply the presence of a nucleic acid (either DNA or RNA) with an extensive reaction machinery (either polymerases or ribosomes) bound to it. These complexes are spaced from one another by a certain minimum distance, and slide on the nucleic acid molecules moving one step only when the complex ahead has itself moved.
Even this description is very simplified; at any rate, it is incorrect to represent this as a standard stoichiometric system. The concept of sequentially ordered movements is not contemplated in Gillespies approach, and there is no natural way to describe it. A possible solution to bypass these obstacles is to use some dummy chemical species, which do not exist in reality, but which can be used to the required effect (the term dummy is introduced by analogy with computer programming, where dummy variables indicate placeholders variables). Therefore, we simulated the attachment and movement of polymerases and ribosomes through a series of dummy chemical species simulating binding sites and movement from a nucleic acid segment to the next.
Adding to the complexity of this scenario, we have to note that translation and transcription are simultaneous events in cell-free PS, and translation starts immediately after the transcription of the first RNA segment, while polymerases are still working on the DNA molecule. This situation is close to that of prokaryotic cells, where translation and transcription take place both in the cytoplasm. As a consequence, the dummy species and the real reactions for both transcription and translation should be arranged to work together.
In the following, we would like to explain in detail the structure of our stochastic description of liposome-entrapped PS.
3 Characteristics and Properties of a Stochastically Simulated PS
3.1 Principles and General Remarks About the Simulator Used (QDC)
In our case study, the PS is used to synthesize Green Fluorescent Protein (GFP): the DNA input molecules code for the standard GFP sequence. Accordingly with Gillespies approach, all the described chemical species can interact independently of each other, and the solutes entrapped inside the liposome, even in the case of dummy species, are supposed to be well-stirred.
We simulated the system by using QDC, a lab-made simulator, whose core is based on Gillespies direct method, with extensions allowing the user to simulate a metabolic experiment, rather than an isolated metabolic system. A detailed survey of QDC is presented in the literature [23]. Here we briefly recall only the most important concepts. QDC can simulate a metabolic experiment since it implements control functions that allow the user to exert some control on the metabolic system during the simulation. In particular, QDC allows: to simulate the addition of molecules at a given time; to simulate continuous feeding or leaking of some species at a set time rate; to specify the so-called “immediate” reactions (reactions with theoretically infinite propensity, that take place immediately after their stoichiometric conditions are met). In our experiments, we have extensively used immediate reactions to describe the advancement of polymerases and ribosomes on their respective nucleic acids.
QDC outputs three main result files: the time courses of all the chemical species present in the system; the propensity time course for each reaction; the number of times that each reaction has been executed. By analyzing and comparing all these three outputs one can extract information about the status of the system and detect possible biases that might have perturbed the simulation results (e.g., a possible stiffness of the system).
3.2 Transcription and Translation Processes
The GFP DNA sequence was divided (according to its length) into 80-bp-long segments, each of which is treated as a distinct chemical species, and named, for instance, DNA1, DNA2, DNA3,.... As a consequence, the polymerization process is represented by several reactions: their core module includes a second-order reaction for nucleotide binding, and a first-order reaction for nucleotide incorporation, which returns the polymerase molecule (which, in turn, can bind to another nucleotide) and a dummy product designed to track the number of nucleotides incorporated in the RNA molecule (these will be named RNA1, RNA2, RNA3, ...
Some immediate reactions determine the transition to the next step, ensuring the following conditions are met: (a) an adjacent DNA site is available, (b) the correct number of nucleotides has been added to the RNA sequence, (c) the corresponding RNA sequence is produced, (d) the previously occupied DNA site is released. Here is an example for transcription at the fourth step (GTP and ATP are considered):

This reaction routine was repeated different times ensuring the correct succession of molecular states; when only one DNA molecule is available, the polymerases advance one by one, separated by at least one DNA site between each other (elongating RNA polymerases are separated from each other by at least 80 bp [24]). The same strategy was used to describe the translation reactions.
An elongating ribosome (eR2) binds the complex carrying the aminoacid (EFaRGTP), after which it moves to the next codon, aided by the elongation factor EFg charged with GTP (EFgGTP); this translocation reaction yields an additional product used to regulate the progression to the next state.

After a fixed number of translocation steps (the minimal space between two elongating ribosomes is, as with polymerases, \(80\ \text {nt} \sim 20\ \text {codons}\) [25]) an immediate reaction occurs in a similar fashion as seen for transcription:
-
1.
the correct amount of aminoacids are incorporated, and thus consumed;
-
2.
the next free RNA site is occupied, and
-
3.
the previous one is therefore freed;
-
4.
an entity named PEPT is also produced, tracking the length of the peptide sequence produced so far:
For example, if 4 species named PEPT3 are present in a certain time of the simulation, this means that there are 4 peptides, still bound to the ribosomes, with a length spanning from \(27\times 3= 81\) to \((27 \times 4) -1 = 107\) aminoacids. This “segmental” partition of DNA and RNA, with the discrete and stepwise occupation by the transcription and translation molecular machineries respectively, is schematically represented in Fig. 1.
A schematic representation of the stepwise transcription and co-translation processes (redrawn from [20]).
3.3 The Overall PS
As this system focuses on the synthesis of GFP starting from its DNA and other basic molecules (nucleotides, aminoacids, etc.), it hosts only a few reactions other than translation and transcription, namely the initialization of both processes and the eventual ribosome inactivation (which takes place 3 h after the start of protein synthesis, see [17]).
4 Results: Simulating in-lipo PS at Nanoscale Range
4.1 Poisson vs. Power-Law: Which Distribution for Solute Entrapment in Nano-Vesicles?
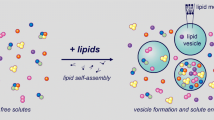
The process of solute entrapment during the formation of liposomes is of special interest, as it affects the distribution of molecules inside them - a relevant issue for studies on the origin of life. Theoretically, when no interactions are supposed to exist between the chemical species to be entrapped, or between these and the nascent lipid bilayer, a standard Poisson process well describes the entrapment mechanism. Recent experimental findings, however, show that, for small liposomes (100 nm diameter), the distribution of entrapped molecules is best described by a power-law function [26], where supercrowded liposomes (i.e., liposomes showing a very high inner solute concentration) are present in a very small but not negligible number. This is a matter of great consequence, as the two random processes generate two completely different scenarios.
We used QDC to simulate a GFP-synthesizing PS encapsulated into liposomes, with the solute partition inside the vesicles obtained by the two entrapment models: a pure Poisson process or a power law. The protein synthesis in lipo was studied in both cases to highlight experimental observables that could be measured to test which model best fits the real entrapment process.
The time course of protein synthesis within each kind of vesicle was then simulated, as a function of vesicle size. Our study can predict translation yield in a population of small liposomes down to the attoliter (\(10^{-18}\) L) range. Our results show that the efficiency of protein synthesis peaks at approximately \(3\cdot 10^{-16}\) L (840 nm diam.) with a Poisson distribution of solutes, while a relative optimum is found at around \(10^{-17}\) L (275 nm diam.) for the power-law statistics. Figure 2 shows the time course of GFP synthesis in a vesicle of \(10^{-17}\) L volume: the protein copies accumulate over time, until the depletion of inner reagents stops the generation of new copies.
Time course of the GFP synthesis in a \(10^{-17}\) L vesicle, populated accordingly to a power-law distribution. In the abscissa the time (in s), in the ordinate the number of GFP molecules synthesized (redrawn from [19]).
Our simulation clearly shows that the wet-lab measurement of an effective protein synthesis at smaller volumes than \(10^{-17}\) L would rule out, according to our models, a Poisson distribution of solutes, and thus would indirectly support that a supercrowding effect took place. This suggestion fits well with a discussion found in [27].
4.2 How Is Intra-vesicle Solute Composition Driving Gene Expression from DNA to Protein?
The next step was to determine whether all chemical constituents of the PS are entrapped in liposomes according to a power law (then, giving rise to a supercrowding effect) independently of each other. Independence (or lack thereof) in solute partition is a foundational feature which we hypothesized would have a significant effect on the final state of the system (e.g. efficiency of gene expression). Finding key end-state parameters dependent on the original entrapment dynamics would also offer an obvious pathway to empirical testability. In this study, the simulations were firstly performed for large liposomes (2.67 \(\mu \)m diameter) entrapping the PS to synthesize GFP. By varying the initial concentrations of the three main classes of molecules involved in the PS (DNA, enzymes, consumables), we were able to stochastically simulate the time-course of GFP production. A mathematical fitting of the simulated GFP production curves allowed us to extract quantitative parameters describing the protein production kinetics; as expected, these parameters resulted significantly dependent on the initial inner solute compositions. Then we extended this study to small-volume liposomes (575 nm diameter), where intra-vesicle composition is harder to infer due to the expected anomalous entrapment phenomena.
In our in silico perturbation of the PS composition, we observed how the competition between DNA transcription and translation for energy resources (here represented by the sum of ATP and GTP available molecules) shapes the protein production dynamics, in terms of total protein yield and its production kinetics. Figure 3 offers a global survey of these results: it shows, for each class of initial DNA inner concentration, the resulting global production in GFP and the corresponding use of energy molecules by the different biochemical processes.
The graph shows the final GFP production obtained (ordinate) according to the initial DNA inner concentration (abscissa). The pie charts placed in correspondence of the abscissa summarize the global energy molecules use for all the case studied (redrawn from [20]).
A mathematical formalization of the PS reaction network allowed us to focus on different aspects of in lipo protein production, and a discussion about our computational strategy together with its scientific relevance is presented in the following sections.
5 On the Relevance of a Detailed Stochastic PS in Silico Model
Different in silico approaches have been tackled for the description of the complex system of reactions encompassing the PS. Despite the large number of biochemical species, the interactions between the single molecules are well-known. Moreover, higher-order interactions between different molecular classes seem to have a modest impact on the overall protein production [29]. Therefore, it is reasonable to approach a comprehensive description of the PS by modeling every reactant as a single entity.
A deterministic formulation of transcription and translation processes can be a useful representation of the PS [17] under the assumption of a continuous, homogeneous distribution of solutes. When dealing with compartmentalized biochemical networks, the intrinsic stochasticity of molecular reactions plays an important role in the observed protein production. A very recent study [30] highlighted the importance of stochasticity in cell-sized lipid compartments, showing how fluctuations in the number of DNA molecules is one of the main responsible of protein production noise using the PS. As volume decreases, well-described anomalous entrapment phenomena [26] create a drastic vesicle-to-vesicle variability in terms of protein production. Moreover, the extremely low number of molecules that eventually can govern a cellular process poses a great challenge in modeling the PS biochemical network. A stochastic representation of the PS deals with those limitations, allowing researchers to focus on single components of the network, with the possibility to focus on nanoscale-sized compartments. This creates the ability to further test different assumptions of the encapsulation efficiencies for different groups of reactant, which, by differentially interacting with the lipidic bilayer, can differentially drive protein production in a size-dependent manner.
Moreover, quantifying the extent to which the intrinsic vesicle-to-vesicle variability influences the protein production in small compartments is of great relevance to discriminate between different components of variability in nano-scaled compartmentalized gene expression: one coming from the random collision between molecules, and the other coming from different internal vesicles composition.
6 Conclusions and Future Directions
Cell-free technologies like the in lipo-PS allow us to monitor and study gene expression kinetics under controlled, known conditions, whereas this is not possible in vivo as well as for whole cell-extracts (e.g. E. coli extracts) also used in similar synthetic biology approaches. Such information is of great value for the most diverse applications, in both applied and basic research [30, 31].
Our main focus is the quantitative understanding of compartmentalized gene expression with respect to the internal solute distribution. By testing different entrapment mechanisms and their effect on protein production in the different PS processes, it is possible to create a link between an observable feature (protein production) and the internal liposomal solute distribution. The ability to infer the precise internal vesicle content is of course of great interest for understanding the different kinetics in different size compartments. This means that the ideal experimental approaches are those able to perform single vesicle measurements, as population measurements could be unable to characterize the high complexity of vesicles behavior.
The results coming from our stochastic modeling are geared to aid experimental design, and experimental validation of our predictions will be needed to further explore and improve our understanding of the compartmentalized PS. A better description of liposome formation kinetics will also help the community to test different encapsulation efficiencies for different molecules, which will in turn affect the internal gene expression kinetics.
As more data about PS use in compartmentalized systems become available, our interpretation of in silico experiments will also improve, allowing us to further tackle the problem of understanding nano-scaled protein-producing liposomes.
References
Shimizu, Y., Inoue, A., Tomari, Y., Suzuki, T., Yokogawa, T., Nishikawa, K., Ueda, T.: Cell-free translation reconstituted with purified components. Nat. Biotechnol. 19, 751–755 (2001)
Shimizu, Y., Kanamori, T., Ueda, T.: Protein synthesis by pure translation systems. Methods 36, 299–304 (2005)
http://parts.igem.org/Chassis/Cell-Free_Systems. Accessed 14 December 2015
Giudice, L.C., Chaiken, I.M.: Cell-free biosynthesis of different high molecular weight forms of bovine neurophysins I and II coded by hypothalamic mRNA. J. Biol. Chem. 254, 11767–11670 (1979)
Drew, D.D.: A mathematical model for prokaryotic protein synthesis. Bull. Math. Biol. 63, 329–351 (2001)
Swain, P.S., Elowitz, M.B., Siggia, E.D.: Intrinsic and extrinsic contributions to stochasticity in gene expression. PNAS 99, 12795–12800 (2002)
Frazier, J.M., Chushak, Y., Foy, B.: Stochastic simulation and analysis of biomolecular reaction networks. BMC Syst. Biol. 3, 64 (2009)
Mier-y-Terán-Romero, L., Silber, M., Hatzimanikatis, V.: The origins of time-delay in template biopolymerization processes. PLoS Comp. Biol. 6, e1000726 (2010)
Fujii, S., Matsuura, T., Sunami, T., Nishikawa, T., Kazuta, Y., Yomo, T.: Liposome display for in vitro selection and evolution of membrane proteins. Nature Protocol 9, 1578–1591 (2014)
Nishimura, K., Matsuura, T., Nishimura, K., Sunami, T., Suzuki, H., Yomo, T.: Cell-free protein synthesis inside giant unilamellar vesicles analyzed by flow cytometry. Langmuir 28, 8426–8432 (2012)
Torre, P., Keating, C.D., Mansy, S.S.: Multiphase water-in-oil emulsion droplets for cell-free transcription? Translation. Langmuir 30, 5695–5699 (2014)
Matsuura, T., Hosoda, K., Kazuta, Y., Ichihashi, N., Suzuki, H., Yomo, T.: Effects of compartment size on the kinetics of intracompartmental multimeric protein synthesis. ACS Synth. Biol. 1, 431–437 (2012)
Murtas, G., Kuruma, Y., Bianchini, P., Diaspro, A., Luisi, P.L.: Protein synthesis in liposomes with a minimal set of enzymes. Biochem. Biophys. Res. Commun. 363, 12–17 (2007)
Kuruma, Y., Stano, P., Ueda, T., Luisi, P.L.: A synthetic biology approach to the construction of membrane proteins in semi-synthetic minimal cells. Biochim. Biophys. Acta 1788, 567–574 (2009)
Sunami, T., Hosoda, K., Suzuki, H., Matsuura, T., Yomo, T.: Cellular compartment model for exploring the effect of the lipidic membrane on the kinetics of encapsulated biochemical reactions. Langmuir 26, 8544–8547 (2010)
Karzbrun, E., Shin, J., Bar-Ziv, R.H., Noireaux, V.: Coarse-grained dynamics of protein synthesis in a cell-free system. Phys. Rev. Lett. 106, 048104 (2011)
Stögbauer, T., Windhager, L., Zimmer, R., Rädlerab, J.O.: Experiment and mathematical modeling of gene expression dynamics in a cell-free system. Integr. Biol. 4, 494–501 (2012)
Mavelli, F., Marangoni, R., Stano, P.: A simple protein synthesis model for the PURE system operation. Bull. Math. Biol. 77, 1185–1212 (2015)
Lazzerini-Ospri, L., Stano, P., Luisi, P.L., Marangoni, R.: Characterization of the emergent properties of a synthetic quasi-cellular system. BMC Bioinfo. 13, S9 (2012)
Calviello, L., Stano, P., Mavelli, F., Luisi, P.L., Marangoni, R.: Quasi-cellular systems: stochastic simulation analysis at nanoscale range. BMC Bioinfo. 14, S7 (2013)
Gillespie, D.T.: Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81, 2340–2361 (1977)
Li, H., Cao, Y., Petzold, L.R., Gillespie, D.T.: Algorithms and software for stochastic simulation of biochemical reacting systems. Biotechnol. Prog. 24, 56–61 (2008)
Cangelosi, D., Fabbiano, S., Felicioli, C., Freschi, L., Marangoni, R.: Quick direct-method controlled (QDC): a simulator of metabolic experiments. EMBnet.journal 19, 39 (2013)
Kubori, T., Shimamoto, N.: Physical interference between Escherichia coli RNA polymerase molecules transcribing in tandem enhances abortive synthesis and misincorporation. N.A.R. 25, 2640–2647 (1997)
Brandt, F., Etchells, S.A., Ortiz, J.O., Elcock, A.H., Hartl, F.U., Baumeister, W.: The native 3D organization of bacterial polysomes. Cell 136, 261–271 (2009)
Luisi, P.L., Allegretti, M., de Souza, T.P., Steiniger, F., Fahr, A., Stano, P.: Spontaneous protein crowding in liposomes: a new vista for the origin of cellular metabolism. Chembiochem 11, 1989–1992 (2010)
De Sousa, P., Stano, P., Luisi, P.L.: The minimal size of liposome-based model cells brings about a remarkably enhanced entrapment and protein synthesis. ChemBioChem 10, 1056–1063 (2009)
Matsuura, T., Kazuta, Y., Alta, T., Adachi, J., Yomo, T.: Quantifying epistatic interactions among the components constituting the protein translation system. Molec. Syst. Biol. 5, 297–300 (2011)
Nishimura, K., Tsuru, S., Suzuki, H., Yomo, T.: Stochasticity in gene expression in a cell-sized compartment. ACS Synth. Biol. 4, 566–576 (2015)
Murray, C.J., Baliga, R.: Cell-free translation of peptides and proteins: from high throughput screening to clinical production. Curr. Op. Chem. Biol. 17, 420–426 (2013)
Chizzolini, F., Forlin, M., Cecchi, D., Mansy, S.S.: Gene position more strongly influences cell-free protein expression from operons than T7 transcriptional promoter strength. ACS Synth. Biol. 3, 363–371 (2014)
Acknowledgments
The Authors will thank Pasquale Stano and Fabio Mavelli for the valuable discussions during the preparation of this review.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this paper
Cite this paper
Calviello, L., Lazzerini-Ospri, L., Marangoni, R. (2016). On Fine Stochastic Simulations of Liposome-Encapsulated PUREsystem™. In: Rossi, F., Mavelli, F., Stano, P., Caivano, D. (eds) Advances in Artificial Life, Evolutionary Computation and Systems Chemistry. WIVACE 2015. Communications in Computer and Information Science, vol 587. Springer, Cham. https://doi.org/10.1007/978-3-319-32695-5_14
Download citation
DOI: https://doi.org/10.1007/978-3-319-32695-5_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-32694-8
Online ISBN: 978-3-319-32695-5
eBook Packages: Computer ScienceComputer Science (R0)