Abstract
The necessity of genetic diversity for evolution and the relationship between heterozygosity and population fitness are important arguments for conserving genetic diversity. The loss of genetic diversity can be detrimental to the short-term viability of individuals and populations, and to the evolutionary potential of populations and species. Genetic erosion can be defined as the permanent reduction in richness or evenness of common local alleles or as the loss of combinations of alleles over time in a defined area. Various international and intergovernmental organizations and networks have therefore recognized the need to assess and monitor plant genetic erosion in order to prevent such effects. The rare tree species Chihuahua spruce (Picea chihuahuana Martínez), which is endemic to Mexico, is an excellent model for estimating genetic erosion. The species occurs in about 40 isolated relict populations in the Sierra Madre Occidental, in the northwest of the country. Here, we will review a study assessing the degree of genetic erosion that was evaluated in five populations of P. chihuahuana M. in the State of Durango (Mexico), by comparing genetic diversity across diameter classes (which were assumed to be a surrogate for age classes). In the two largest populations, there was a moderate loss of genetic diversity at AFLP loci from older trees to saplings, and to young seedlings. Significant genetic erosion was only detected in the very small population of San José de las Causas (SJ). Hence, if genetic diversity at AFLP loci reflects diversity in the whole genome, genetic erosion per se does not explain the relict status of Chihuahua spruce, except for very small populations, such as SJ. However, further researches with candidate genes are necessary to assess the putative loss of evolutionary potential in these stands. Activities that increase population size should be helpful to preserve genetic diversity.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
7.1 Introduction
7.1.1 Genetic Erosion
The necessity of genetic diversity for evolution and the relationship between heterozygosity and population fitness are important arguments for conserving genetic diversity (Reed and Frankham 2003). In the short term, genetic diversity is related to increased inbreeding, which at its turns affects individual fitness. In the longer term, standing genetic variation can be associated with a species’ ability to respond to changing selection pressures (Young et al. 1996; Reed and Frankham 2003; De Carvalho et al. 2010; Kremer et al. 2012). On the other hand, genetic erosion can be detrimental to the viability of individuals and the evolutionary potential of populations and species, thus affecting the direct use of genetic resources (Brown et al. 1997).
Various international and intergovernmental organizations and networks [e.g., the World Conservation Union (IUCN), Species Survival Commission, Convention on Biological Diversity (CBD), UNEP World Conservation Monitoring Centre (UNEP/WCMC), Organisation for Economic Co-operation and Development (OECD), European Union (EU), Bioversity International (formerly IPGRI) and FAO] have recognized the need to assess and monitor genetic erosion, in order to prevent such effects (Diulgheroff 2006). A literature review has shown that surprisingly few studies have measured and assessed this important process in forest tree communities (e.g., Lee et al. 2002), particularly for subtropical threatened taxa.
Genetic erosion can be viewed as the “loss of genetic diversity, in a particular location and over a particular period of time, including the loss of individual genes, and the loss of particular combinations of genes such as those manifested in landraces or varieties. It is thus a function of change of genetic diversity over time” (FAO/IPGRI 2002). Maxted and Guarino (2006) suggested that genetic erosion may also be defined as a “permanent reduction in richness or evenness of common local alleles or the loss of combination of alleles over time in a defined area”. However, it must be noted that these definitions do not specify whether genetic erosion is caused by adaptation (selection), genetic drift, or inbreeding. For instance, the latter includes the total number of variants and their relative frequencies, as two important components of diversity that are well-balanced, and considered in the Simpson index (Simpson 1949) and in the “effective number” of variants (υ 2 ) (υ 2 = 1/Σp 2 i ) (Gregorius 1978). As a complement, Brown et al. (1997) provided a useful list of features or indicators for estimating the potential risk of genetic erosion, namely: (i) the number of subspecific entities, (ii) population sizes, numbers, and isolation, (iii) environmental amplitude, (iv) genetic diversity at marker loci, (v) quantitative genetic variation, (vi) interpopulation genetic structure, and (vii) amount and patterns of mating.
In this chapter, we will discuss one of the few documented examples of genetic erosion found the endangered, rare, relictic, and fragmented endemic Mexican spruce, P. chihuahuana M., in Durango State, northwestern Mexico. It studied five populations, by comparing genetic diversity among diameter at breast height (DBH) classes (as a surrogate variable for age classes), estimated using dominant gene markers (AFLP) and Gregorius’ total population differentiation (δ T) (Gregorius 1987). These populations bear less adult trees and should be more affected by the ongoing climate change than the northern ones, and thus represent ideal models to estimate genetic erosion. Results were previously reported in Wehenkel and Sáenz-Romero (2012).
7.1.2 Genetic Diversity and Structure of Picea chihuahuana Martínez
The rare tree species Chihuahua spruce (P. chihuahuana Martinéz), an endemic of Mexico, is an excellent model for estimating potential genetic erosion (Ledig et al. 1997). This species occurs in about 40 isolated relict populations at elevetions between 2,155 and 2,990 m above sea level in the Sierra Madre Occidental in the states of Durango and Chihuahua, in Northwestern Mexico. The size of the populations varies from 21 to 5546 individuals, including trees, saplings, and seedlings (Ledig et al. 2000; Farjon 2001).
As this species is a relict stranded by a warming climate during the current interglacial period, Mahlman (1997) and Ledig et al. (2000) proposed that Chihuahua spruce can serve as a signal species for the projected climate change in the twenty-first century. It has thus became emblematic of the challenges that Mexico will face in implementing management actions, such as assisted colonization, to prevent extinctions due to global warming (Ledig et al. 2010). Some studies have been carried out to establish the genetic diversity and structure of this species (Ledig et al. 1997, 2004; Jaramillo-Correa et al. 2006; Wehenkel and Sáenz-Romero 2012; Wehenkel et al. 2012; Quiñones-Pérez et al. 2014a, b; www.mapforgen.org).
Ledig et al. (1997) analyzed 24 loci in 16 enzyme systems to estimate genetic diversity (H e) and the number of alleles per locus (A) in 10 populations comprising 15 to 2441 mature trees, based on seeds and sample sizes of 7.7–22.9 trees per locus. They concluded that “if genetic diversity at isozyme loci reflects diversity in the genome as a whole, lack of diversity per se is not the reason for the relictual status of Chihuahua spruce.” These authors also found that H e and A were closely related to the logarithm of the number (N) of mature trees in the population (r He,N = 0.93, P = 0.004; r A,N = 0.78, P = 0.047), which confirms the theory relating population size and genetic diversity (Frankham 1996).
Jaramillo-Correa et al. (2006) determined and observed numbers of mitotypes and chlorotypes, and mitochondrial and chloroplast diversity estimates (H; equivalent to the expected heterozygosity; H e, for diploid data) for 16 Chihuahua spruce populations, based on seeds and sample sizes of 8–10 mature trees per population. These authors found that none of the 16 P. chihuahuana populations surveyed was polymorphic for the mtDNA markers, while for the cpDNA markers, three of the 16 stands surveyed were fixed for a particular chlorotype. In addition, they noted that the diversity in cpDNA decreased from northern to southern latitudes (Lat) (r H,Lat = 0.626; P < 0.01) and that genetic and geographic distances were not related.
However, a marginal correlation was observed when comparing the diversity in cpDNA with the population census (r H,ln(N) = 0.064; P = 0.863) and with all other ecological or demographic factors previously considered (Ledig et al. 2000) in the stands surveyed therein. Jaramillo-Correa et al. (2006) therefore assumed that P. chihuahuana has been subjected to strong bottlenecks and has suffered from genetic drift in the recent past (i.e., during the Holocene). Thus, according to the previous definition, genetic erosion has proceeded in the species as a whole, at least since the end of the last glacial period. However, the question remains as to whether genetic erosion took place recently (or is still taking place) in single isolated populations, because not all fragmentation events lead to reduced genetic variation in plants (Young et al. 1996).
7.2 Genetic Erosion in Populations of Picea chihuahuana M.
7.2.1 Populations Studied and Methodological Approach of Detecting Genetic Erosion
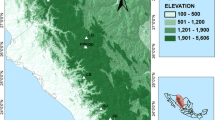
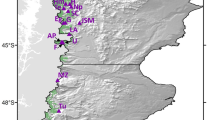
The study was conducted in the State of Durango which occupies about 23 % of the Sierra Madre Occidental ecosystem. Branches were collected from each of 254 randomly chosen specimens of P. chihuahuana M., distributed in five populations as follows: (a) Paraje Piedra Rayada (PPR), (b) Quebrada de los Durán (Arroyo del Indio Ignacio) (QD), (c) La Pista (LP), (d) Santa Barbara (Arroyo del Infierno) (SB), and (e) San José de Causas (SJ), which covers most of the latitudinal range of the species in Durango State, Mexico (Figs. 7.1 and 7.2). In addition, the diameter at breast height (DBH) of trees and saplings, and the diameter at ground level of seedlings were assessed for every individual studied.
Map of the 40 already detected populations of Picea chihuahuana M. and the five locations of the populations under study: Paraje Piedra Rayada (PPR), Quebrada de los Durán (Arroyo del Indio Ignacio) (QD), La Pista (LP), Santa Barbara (Arroyo del infierno) (SB), and San José de Causas (SJ) (red circles), in the State of Durango, Mexico (Elevation in m)
Because P. chihuahuana is an endangered species (IUCN Red List of Threatened Species 2011; www.iucnredlist.org), the sample trees were not cored for growth ring counting to obtain the actual ages (permission to core the stems was not obtained from federal authorities). However, Gordon (1968) and Narvaéz-Flores (1984) - in Ledig et al. (2000) - reported a positive relationship between tree diameter, height, and tree age in Chihuahua spruce. According to Ledig et al. (2000), “P. chihuahuana probably grows slowly, about 0.25 to 0.75 m per year, on average, over its first 100 years”; this was estimated from a regression of height and age of 29 trees. DBH was therefore used as a surrogate for age (see below for more details). Ledig et al. (2000) also provided detailed descriptions of the first four populations shown in Table 7.1 along with some demographic and ecological parameters.
Given that DBH can represent the approximate tree age (Seymour and Kenefic 1998; Nord-Larsen and Cao 2006) in Chihuahua spruce (Ledig 2000), it was assumed that each diameter class reflects natural regeneration during a particular period within the past 160 years. For instance, the dc of 5 cm was assumed to represent the natural regeneration in the past 20 years, the dc of 75 cm the natural regeneration that occurred about 130–160 years ago, etc. This enabled Wehenkel and Sáenz-Romero (2012) comparing the genetic diversity of Chihuahua spruce across different time periods for a defined area.
According to Wehenkel and Sáenz-Romero (2012), DNA data from frozen needles were obtained by the amplified fragment length polymorphism (AFLP) technique. AFLP fingerprints were generated using a modified protocol described by Vos et al. (1995). As for other dominant markers, each band detected (presence) at each given position (locus) corresponds to a dominant genotype (plus phenotype).
Further methodological details for measuring Gregorius’ total differentiation (δ T) can be found in Simpson (1949) and Gregorius (1987); for determining the proportion of polymorphic fragments (pr poly) and down-weighted marker values (DW) in Schönswetter and Tribsch (2005); for computing arithmetic the mean genetic distances calculated with all pairs of individuals in each cohort (d 0,ind,m) and with four randomly chosen pairs of individuals in each cohort to compensate for the different sample sizes (d 0,ind,m(4)) in Weir et al. (2006) and Gregorius et al. (2007); and for calculating covariation (C) as well as for statistical tests in Gregorius et al. (2007). Those methods were used by Wehenkel and Sáenz-Romero (2012).
7.2.2 Significant Genetic Erosion Detected in the Very Small Population
Wehenkel and Sáenz-Romero (2012) reported that a significant loss of genotype diversity was only detected among the 319 AFLP loci studied in the very small San José de Causas (SJ) stand (Fig. 7.3b and Table 7.2); although the demographical stem-number distributions were almost balanced when using 10-cm DBH classes (Fig. 7.5). One of the main reasons for the loss of diversity in SJ may be the smaller proportion of mature and reproductively competent individuals (Reed and Frankham 2003), which can be explained by the detection of significantly less genetic differentiation (and thus higher relatedness) between individuals in the younger generations (Fig. 7.4 and Table 7.2).
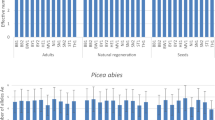
Relationships between DBH classes and mean values of Gregorius’ total differentiation (δ T), ( a ) across all populations and for observed relative diameter distribution (f), and ( b ) total differentiation (δ T) for each population [Paraje Piedra Rayada (PPR), Quebrada de los Durán (Arroyo del Indio Ignacio) (QD), La Pista (LP), Santa Barbara (Arroyo del infierno) (SB) and San José de Causas (SJ)] for all 254 individuals of Picea chihuahuana M. studied in Durango, Mexico
Relationships between DBH classes and mean values of mean genetic distance between two individuals with four randomly chosen pairs of individuals in each cohort (d 0,m(4)) for each population [Paraje Piedra Rayada (PPR), Quebrada de los Durán (Arroyo del Indio Ignacio) (QD), La Pista (LP), Santa Barbara (Arroyo del infierno) (SB), and San José de Causas (SJ)] of Picea chihuahuana M. studied in Durango, Mexico
The observed relative diameter distribution in the diameter classes of 2.5–82.5 cm in the populations Paraje Piedra Rayada (PPR), Quebrada de los Durán (Arroyo del Indio Ignacio) (QD), La Pista (LP), Santa Barbara (Arroyo del infierno) (SB), and San José de Causas (SJ) populations of Picea chihuahuana M., Durango, Mexico
Hence, the results of Wehenkel and Sáenz-Romero (2012) are consistent with the findings of Ledig et al. (1997), i.e., if genetic diversity at a set of loci reflects the diversity in the whole genome, then genetic erosion per se does not explain the relict status of Chihuahua spruce, except in the very small populations, such as SJ. This population, consisting of about 120 trees, and has probably fallen below the level of a minimum viable population size when demographic and environmental stochasticity, as well as natural catastrophes, are ignored. The standing genetic diversity of this population would therefore not be sufficient to prevent a dangerous accumulation of inbreeding depression, while future mutations should not compensate for the loss of alleles due to genetic drift (Wright 1938; Millar and Libby 1991; Frankham et al. 2002; Bücking 2003).
It is important to note that the trend of genetic erosion in the population SJ could be reversed if pollen and/or seedlings originating from older trees in SJ (genetically more variable than younger individuals) or from neighboring populations are (re)introduced (Wehenkel and Sáenz-Romero 2012). It may be argued that the introduction of foreign alleles may cause outbreeding depression. However, there is probably a greater risk of allowing this population to continue its genetic decline in what could become a vortex of extinction (Frankham et al. 2002). Furthermore, given that all populations studied therein have shown identical mtDNA signatures and similar cpDNA patterns, they should be derived from the same ancestral stand and bear similar adaptive alleles. Therefore, the risk of outbreeding depression should be low. It must also be noted that some models predict that gene flow fosters adaptation in forest trees (see Kremer et al. 2012 for a review), and although these effects might be different for the rear edge populations (such as those surveyed by Wehenkel and Sáenz-Romero 2012), this could be an ideal opportunity to empirically test the putative responses of these threatened stands to future environmental changes.
7.2.3 Implications for Management and Conservation
Traill et al. (2007) reported in a meta-analysis based on 141 sources and 212 species that the minimum viable population size is context-specific and is on average 4,824 individuals for plant species (95 % CI = 2,512 − 15,992). Such an estimate suggests that all populations of Chihuahua spruce have low chances of survival without assistance, given that the maximum population size currently observed is only 3564 trees (>2 m tall; Ledig et al. 2000). Therefore, all activities that increase population size would be helpful regarding the genetic structure and diversity of the species as a whole. Such activities might include the following: (i) protect natural regeneration against livestock, wild animals and forest fires, (ii) establish artificial regeneration with autochthonous reproductive material in well-selected locations in the vicinity of (but not inside) the particular population, (iii) remove competing vegetation (including other tree species) in the vicinity of the particular population, (iv) support putative biotic dispersal vectors, and (v) promote the establishment of local mycorrhiza and other microorganisms that might help increase nutrient intakes in seedlings and saplings in and around populations.
Moreover, it is necessary to continue monitoring the population sizes and genetic diversity of the existing stands in situ. Then, if genetic erosion, or any other significant stochastic force, is detected in a population, the most obvious tactic should be to restore gene flow to this population (Ledig et al. 1997). However, because of differences in the genetic population structure, seed transfer from the northern to the central or southern populations and vice versa should be avoided (see Jaramillo-Correa et al. 2006).
Interestingly, for the small La Pista (LP) population and the very small Santa Bárbara (SB) stand, the mean genetic diversity did not decrease throughout the DBH classes (and thus age classes) (Wehenkel and Sáenz-Romero 2012). Perhaps the purging of lethal alleles and facultative selfing suspected in many threatened spruces (Ledig et al. 2002, 2005; Aleksiċ and Geburek 2013) occurred earlier during a bottleneck event for these particular locations, thus maintaining higher mean levels of genetic diversity. It is also possible that an array of well-adapted individuals that retained a well-represented genetic diversity survived and sustained these populations since the last bottleneck. Because of the limitations imposed by the dominant marker used, Wehenkel and Sáenz-Romero (2012) could not determine whether the fitness and constant genetic diversity of these individuals were caused by high degrees of heterozygosity (Ledig 1986), which should be thus explored further. However, the results reported by Ledig et al. (1997) argue against this possibility given the excess of homozygosity found for eight of the 13 isozyme loci surveyed. In which case, it could be argued that any possible loss of genetic diversity in LP and SB due to genetic drift and inbreeding was compensated by incoming gene flow from the neighboring populations, such as proposed for isolated stands in the endemic and endangered Serbian spruce (Aleksiċ and Geburek 2013).
7.2.4 Adaptation to Climatic Change
The predicted reduction and eventual disappearance of a suitable habitat for P. chihahuana due to climatic change (Ledig et al. 2010) imposes an additional risk of extinction. Management actions such as migration (also called assisted colonization), e.g., establishing ex situ conservation plantations outside the present distribution, at localities where it is predicted that suitable habitat will occur, should be carried out, giving priority to seedlings grown from seed collected from older trees. According to Wehenkel and Sáenz-Romero (2012), these mature trees should be the most genetically diverse.
A priority area to establish ex situ conservation plantations would be the northwest corner of the State of Durango, near its border with the State of Chihuahua, where it was predicted that suitable climatic habitat will arise in the near future (i.e., between 2030 and 2060; Ledig et al. 2010). Also, in this area will collide projected suitable climatic habitats for what was identified as the northern mitotype and the central–southern mtDNA variant (Jaramillo-Correa et al. 2006). Since there are no available provenance test results so far, and then, it is not known if both mitotypes reflect genetic differentiation for quantitative traits of adaptive relevance, a conservative approach would be to keep separated both mitochondrial lineages at the ex situ conservation planting sites to prevent potential outbreeding depression.
Such ex situ conservation plantations should be big enough to equate a genetically viable effective population size (see Traill et al. 2007), where the population size might be large enough for at least to theoretically compensate the loss of genetic diversity due to genetic drift with new genetic variants provided by mutation (Millar and Libby 1991). As a general suggestion, it could be proposed to establish Forest Genetic Resource Conservation Units (FGRCUs), with a minimum of 4,660 trees at reproductive age, in order to maintain an average heterozygosity similar to that found across Mexican conifers (Sáenz-Romero et al. 2003). For the P. chihuahuana case, however, the FGRCU would be an ex situ plantation, instead of a designed natural stand. Nevertheless, further research might be needed to determine how many mature trees should be sampled to collect the seed to establish such a plantation. In any case, as a starting point, and considering the lack of information and the urgency to preserve P. chihuahuana, it seems reasonable to aim for ex situ conservation plantations of a minimum of 5,000 individuals at the above-mentioned border of Durango and Chihuahua States as soon as possible.
7.2.5 Future Research Needed
Estimations of current and historical gene flow among populations seem essential to enrich the current conservation programs for P. chihuahuana. Ideally, assisted migration should only include populations that have been or are still exchanging genetic material, and their identification can easily be done by scanning the genome with anonymous markers (such as AFLPs) and/or by transferring some of the genetic tools developed for other spruces. Recently, two different SNP arrays comprising more than 15,000 markers originally designed for P. glauca (a boreal conifer whose genome was recently published; Birol et al. 2013) were tested in seven spruce taxa, including P. chihuahuana (Pavy et al. 2013). Although the number of segregating markers was very low in this last species (i.e., 322), they are still more than enough to estimate basic population-genetics and demographic figures, including gene flow and inbreeding. Further re-sequencing of genes directly derived from these genomic resources might allow the estimation of other demographic parameters, such as the time and intensity of the past bottlenecks, and the times of divergence of the modern populations (e.g., Aleksiċ and Geburek 2013). Altogether, these parameters could be integrated into simulation frameworks to predict possible outcomes under different climate change scenarios, and to propose precautionary measures when needed.
Genomic tools can also be used to monitor effective population sizes, genetic diversity and inbreeding across time. Indeed, under a population decline, or an extinction vortex scenario, these parameters are expected to change rapidly from one generation to next (Frankham et al. 2002). Therefore, developing DNA arrays that are easy to apply would allow the direct estimation of these parameters once a new cohort is established and/or prior to the introduction of individuals (or pollen) from other populations, which could maximize the amount of preserved genetic diversity. Furthermore, the combination of these analyzes with test plantations in the field could help identify divergence patterns of adaptive relevance.
Nevertheless, because a significant loss of diversity had occurred in some of the studied Chihuahua spruce populations at different gene loci, a globally directed force, such as climate, which should operate as a directional selective force, should be less relevant than most stochastic pressures. Indeed, genetic drift and inbreeding due to reduction of the effective population size would be expected to have a greater influence (Franklin 1980). However, the search for putative adaptive gene variants that still remain in the populations, and which are related to other variables than climate (i.e., soil type), should be helpful to reinforce any future conservation efforts in order in correctly account for the evolutionary potential of each particular stand.
7.2.6 Conclusions
Genetic erosion was documented at one of the smallest population of the southern distribution of P. chihuahuana. Considering the progressive loss of genetic diversity along the DBH classes (a surrogate of age classes), the insufficient recruitment of young plants and the multiple threatening factors (illegal logging, grazing, forest fires), and climatic change, there is no reason to believe that such population would not go extinct, unless there is an active management for conservation. Among other conservation actions, it should be considered enrichment to perform some planting to increase genetic diversity and population size, and to develop ex situ conservation, by planting in sites where it is predicted that will occur the climate for which populations are adapted (as a measure for adaptation to climatic change).
References
Aleksić JM, Geburek T (2013) Quaternary population dynamics of an endemic conifer, Picea omorika, and their conservation implications. Conserv Genetics
Birol I, Raymond A, Jackman SD, Pleasance S, Coope R et al (2013) Assembling the 20 Gb white spruce (Picea glauca) genome from whole-genome shotgun sequencing. Bioinformatics 29(12):1492–1497
Brown A, Young A, Burdon J, Christides L, Clarke G, Coates D, Sherwin W (1997) Genetic indicators for state of the environment reporting. State of the Environment Technical Paper Series (Environmental Indicators), Department of Environment, Sport and Territories, Canberra ACT, Australia
Bücking W (2003) Are there threshold numbers for protected forests? J Environ Manag 67:37–45
De Carvalho D, Ingvarsson PK, Joseph J, Suter L, Sedivy C, Macaya-Sanz D, Cottrell J, Heize B, Schanzer I, Lexer C (2010) Admixture facilitates adaptation from standing variation in the European aspen (Populus tremula L.), a widespread forest tree. Mol Ecol 19(8):1638–1650
Diulgheroff S (2006) A global overview of assessing and monitoring genetic erosion of crop wildrelatives and local varieties using WIEWS and other elements of the FAO global system on PGR. In Ford-Lloyd BV, Dias SR, Bettencourt E (eds) Genetic erosion and pollution assessment methodologies. Proceedings of PGR Forum Workshop 5, Terceira Island, Autonomous Region of the Azores, Portugal, 8–11 September 2004, pp. 35–45. Published on behalf of the European Crop Wild Relative Diversity Assessment and Conservation Forum, by Bioversity International, Rome, Italy, pp 100, Available at http://www.bioversityinternational.org/fileadmin/bioversity/publications/pdfs/1171.pdf (accessed 11 January 2010)
FAO/IPGRI (2002) Review and development of indicators for genetic diversity, genetic erosion and genetic vulnerability (GDEV): summary report of a joint FAO/IPGRI workshop (Rome, 11–14 Sept 2002)
Farjon A (2001) World checklist and bibliography of conifers, 2nd edn. Royal Bot Gardens, Kew, UK
Frankham R (1996) Relationship of genetic variation to population size in wildlife. Cons Biol 10(6):1500–1508
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Franklin JR (1980) Evolutionary change in small populations. in: M.E. Soule and B.A. Wilcox (editors). Conservation biology: an evolutionary-ecological perspective, Sinauer, Sunderland MA, USA, pp 135–150
Gordon AG (1968) Ecology of Picea chihuahuana Martínez. Ecology 49:880–896
Gregorius HR (1978) The concept of genetic diversity and its formal relationship to heterozygosity and genetic distance. Math Biosci 41:253–271
Gregorius HR (1987) The relationship between the concepts of genetic diversity and differentiation. TAG 74:397–401
Gregorius HR, Degen B, König A (2007) Problems in the analysis of genetic differentiation among populations—a case study in Quercus robur. Silvae Genet 56:190–199
Jaramillo-Correa JP, Beaulieu J, Ledig FT, Bousquet J (2006) Decoupled mitochondrial and chloroplast DNA population structure reveals Holocene collapse and population isolation in a threatened Mexican-endemic conifer. Mol Ecol 15:2787–2800
Kremer A, Ronce O, Robledo-Arnuncio JJ, Guillaume F, Bohrer G, Nathan R, Bridle JR, Gomulkiewicz R, Klein EK, Ritland K, Kuparinen A, Gerber S, Schueler S (2012) Long-distance gene flow and adaptation of forest trees to rapid climate change. Ecol Lett 15(4):378–392
Ledig FT (1986) Heterozygosity, heterosis and fitness in outbreeding plants. In: Soulé ME (ed) Conservation Biology. The Science of Scarcity and Diversità, Sinauer Associates, Sunderland, pp 77–104
Ledig FT, Jacob-Cervantes V, Hodgskiss PD, Eguiluz-Piedra T (1997) Recent evolution and divergence among populations of a rare Mexican endemic, Chihuahua spruce, following Holocene climatic warming. Evolution 51(6):1815–1827
Ledig FT, Mápula-Larreta M, Bermejo-Velázquez B et al (2000) Locations of endangered spruce populations in México and the demography of Picea chihuahuana. Madroño 47:71–88
Ledig FT, Hodgskiss PD, Jacob-Cervantes V (2002) Genetic diversity, mating system, and conservation of a Mexican subalpine relict, Picea mexicana Martínez. Cons Genet 3:113–122
Ledig FT, Hodgskiss PD, Krutovskii KV, Neale DB, Eguiluz- Piedra T (2004) Relationships among the spruces (Picea, Pinaceae) of southwestern North America. Syst Bot 39:275–295
Ledig FT, Hodgskiss PD, Johnson DR (2005) Genetic diversity, genetic structure, and mating system of Brewer spruce (Pinaceae), a relict of the Arcto-Tertiary forest. Am J Bot 92(12):1975–1986
Ledig FT, Rehfeldt GE, Sáenz-Romero C, Flores-López C (2010) Projections of suitable habitat for rare species under global warming scenarios. Am J Bot 97(6):970–987
Lee CT, Wickneswari R, Mahani MC, Zakri AH (2002) Effect of selective logging on the genetic diversity of Scaphium macropodum. Biol Cons 104(1):107–118
Mahlman JD (1997) Uncertainties in projections of human-caused climate warming. Science 278:1416–1417
Maxted N, Guarino L (2006) Genetic erosion and genetic pollution of crop wild relatives. In Ford-Lloyd BV, Dias SR, Bettencourt E (eds) Genetic erosion and pollution assessment methodologies. Proceedings of PGR Forum Workshop 5, Terceira Island, Autonomous Region of the Azores, Portugal, 8–11 Sept 2004, pp 35–45. Published on behalf of the European Crop Wild Relative Diversity Assessment and Conservation Forum, by Bioversity International, Rome, Italy, pp 100 Available at http://www.bioversityinternational.org/fileadmin/bioversity/publications/pdfs/1171.pdf (accessed 11 Jan 2010)
Millar CI, Libby WJ (1991) Strategies for conserving clinal, ecotipic, and disjunct population diversity in widespread species. In: Falk DA, Holsinger KE (eds) Genetics and Conservation of Rare Plants. Oxford University Press, New York, pp 149–170
Narvaéz-Flores R (1984) Contribucion al conocimiento de la ecología de Picea chihuahuana Martínez. Unpublished thesis. Universidad Autónoma de Nuevo León, Monterrey, Nuevo León
Nord-Larsen T, Cao QV (2006) A diameter distribution model for even-aged beech in Denmark Forest. Forest Ecol Manag 231(1–3):218–225
Pavy N, Gagnon F, Rigault P, Blais S, Deschênes A et al (2013) Development of high-density SNP genotyping arrays for white spruce (Picea glauca) and transferability to subtropical and nordic congeners. Mol Ecol Res 13:324–336
Quiñones-Pérez CZ, Sáenz-Romero C, Wehenkel C (2014a) Influence of neighbouring tree species on AFLP variants of endangered Picea chihuahuana populations on the Sierra Madre Occidental, Northeastern México. Polish J Ecol 62(1):69–79
Quiñones-Perez CZ, Simental-Rodriguez SL, Saenz-Romero C, Jaramillo-Correa JP, Wehenkel C (2014b) Spatial genetic structure in the very rare and species-rich Picea chihuahuana tree community (Mexico). Silvae Genetica, submitted
Reed DH, Frankham R (2003) Correlation between fitness and genetic diversity. Conserv Biol 17:230–237
Sáenz-Romero C, Snively A, Lindig-Cisneros R (2003) Conservation and restoration of pine forest genetic resources in México. Silvae Genet 52(5–6):233–237
Schönswetter P, Tribsch A (2005) Vicariance and Dispersal in the Alpine Perennial Bupleurum stellatum L. (Apiaceae). Taxon 54:725–732
Seymour RS, Kenefic LS (1998) Balance and sustainability in multi-aged stands: a northern conifer case study. J Forest 96:12–17
Simpson EH (1949) Measurement of diversity. Nature 163:688
Traill LW, Bradshaw JA, Brook BW (2007) Minimum viable population size: a meta-analysis of 30 years of published estimates. Biol Cons 139(1–2):159–166
Vos P, Hogers R, Bleeker M et al (1995) AFLP: a new concept for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Wehenkel C, Saenz-Romero C (2012) Estimating genetic erosion using the example of Picea chihuahuana Martínez. Tree Genet Genomes 8(5):1085–1094
Wehenkel C, Martínez-Guerrero JH, Pinedo-Alvarez A, Carrillo A (2012) Adaptive genetic differentiation in Picea chihuahuana M. caused by different copper concentrations in the top soil. Forstarchiv 83:48–51
Weir BS, Anderson AD, Hepler AB (2006) Genetic relatedness analysis: modern data and new challenges. Nat Rev Genet 7:771–780
Wright S (1938) Size of population and breeding structure in relation to evolution. Science 87:430–431
Young A, Boyle T, Brown A (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418
Acknowledgments
This study was supported by joint funding from the Mexican Council of Science and Technology (CONACyT) and the Ministry of Education (SEP; Project CB-2010-01 158054), the Durango State Ministry of Natural Resources and Environment, the Universidad Juárez of the State of Durango, México and ISOGEN, Göttingen, Germany. With kind permission from Springer Science+Business Media, we used here four figures and two tables from: Wehenkel and Saenz-Romero (2012).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Wehenkel, C., Sáenz-Romero, C., Jaramillo-Correa, J.P. (2016). Estimating Genetic Erosion in Threatened Conifers: The Example of Picea chihuahuana Martínez. In: Ahuja, M., Jain, S. (eds) Genetic Diversity and Erosion in Plants. Sustainable Development and Biodiversity, vol 8. Springer, Cham. https://doi.org/10.1007/978-3-319-25954-3_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-25954-3_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-25953-6
Online ISBN: 978-3-319-25954-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)