Abstract
Whilst the depletion of poly(ADP-ribose) polymerase (PARP) activity associated with PARP-1 and PARP-2 in cells causes radiosensitivity, its inhibition by small molecule inhibitors is showing great potential as a mechanism to potentiate the effects of radiotherapy. Indeed as over 50 % of all cancer patients will receive radiotherapy at some point in their treatment there is considerable interest in the development of radiosensitisers that can replace chemotherapeutic agents without the associated dose-limiting toxicities. Potent and specific inhibitors of PARP activity that compete with NAD+ at the enzyme’s activity site have been developed. Their use in preclinical in vitro and in vivo models is associated with enhanced radiosensitivity through mechanisms that not only involve the trapping of the PARP proteins at sites of strand breaks and the modulation of DNA repair in a proliferation dependent manner but also through the targeting of the endothelium and tumour vasculature and changes in tumour oxygenation and thus hypoxia-related radioresistance. However as both PARP-1 and PARP-2 are also involved in transcription regulation, chromatin modification and cellular homeostasis, the impact of PARP inhibition on these processes and long-term therapeutic responses needs to be investigated, as well as issues relating to scheduling, dose and radiation quality on the efficacy of this combined therapy.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Poly(ADP-ribose) polymerase 1
- Poly(ADP-ribose) polymerase 2
- Poly(ADP-ribose) polymerase inhibitors
- Radiosensitivity
- DNA repair
- DNA strand breaks
- Radiotherapy
1 Introduction
The inhibition of DNA repair and other damage response proteins by small molecule inhibitors can potentially be exploited to sensitise tumour cells when used in combination with chemo- and radiotherapy or in certain genetic backgrounds. Many therapies, including alkylating agents such as temozolomide (TMZ), campthotecin and radiation produce DNA single strand breaks (SSBs) and inhibitors of poly(ADP-ribose) polymerase (PARP) activity, an important post-translational modification in the repair of SSBs in mammalian cells, have shown great potential as sensitizers in a variety of cell and tumour types. Poly(ADP-ribosy)lation is a dynamic, energy consuming process carried out by members of the PARP family, also known as ADP-ribosyltransferases (ADPRTs) or poly(ADP-ribose) synthetases. PARP-1, the founding member of this family discovered 50 years ago [1], is responsible for the synthesis of the majority of poly(ADP)ribose (PAR) in eukaryotic cells and after the histones, is the most abundant nuclear protein [2]. Based on structural homology with its catalytic domain, 17 PARP family members have been identified using bioinformatics approaches [3], of these PARP-2 has the greatest structural similarity to PARP-1. PARP-1, PARP-2 and PARP-3 are activated by binding to DNA strand breaks and, using NAD+ as a substrate, catalyse the formation of long homopolymers of ADP-ribose that are involved in many cellular processes and in particular DNA repair. PARP-1 itself is the primary target for Poly(ADP-ribosy)lation in vivo, with more than 90 % of PAR being found on PARP-1 [4]. The histones are also poly(ADP-ribos)ylated in response to DNA damage, resulting in a loosening of the chromatin structure. This post-translational modification facilitates the repair of the DNA strand break and promotes the recruitment of the proteins that repair the break, in particular XRCC1 through its direct interaction with poly(ADP-ribos)ylated PARP-1 [5]. Once in place XRCC1 acts a scaffold for DNA polymerases and ligases that will then complete the repair of the SSB [6].
Because of the association between PARP activity and DNA damage responses, there has been a long-standing interest in the impact of the inhibition of this activity on enhancing cytotoxicity. One of the first reports came from the group of Sydney Shall over 30 years ago [7]. They demonstrated that 3-aminobenzamide (3-AB) prevented the rejoining of DNA strand breaks caused by the alkylating agent dimethyl sulphate and increased its toxicity in L1210 mouse leukemia lymphoblast cells. Sensitivity to γ-radiation was also reported using this prototype PARP inhibitor [8]. The development of second and third generation PARP inhibitors, more applicable to clinical use, combined with a better understanding of the impact of PARP inhibition on cellular processes in addition to DNA repair, has led to their assessment in preclinical and clinical trials either as single agent treatments or in combination with chemo- and/or radiotherapy to potentiate the cytotoxic effects of these therapeutic agents. This article will review the preclinical evidence that demonstrates the radiosensitisation obtained by PARP inhibition, in comparison to the radiosensitivity seen in the absence of PARP expression, in the light of the 5Rs of radiobiology namely repair, repopulation, redistribution, reoxygenation and radiosensitivity, that govern the probability of tumour control; readers are referred to other chapters in this book for detailed reviews of PARP family members, their activities and the development of PARP inhibitors.
2 Factors Affecting Cancer Cell Radiosensitivity: The Absence of PARPs
As reviewed by Powell et al. [9], when tumours are treated with radiotherapy, their ability to repair DNA damage, the number of clonogenic cells and their rate of repopulation, their redistribution in the cell cycle over time, their intrinisic radiosensitivity and the presence of tissue hypoxia will all influence the probability of tumour control. Ionising radiation used in the clinical treatment of cancer generates DNA SSBs and double strand breaks (DSBs) in an approximate ratio of 25:1 and PARP-1 can bind to both [10]. Although its role in SSB and DSB repair is well established, how the DNA-end binding versus the enzymatic activity of PARP-1 contributes to DNA repair regulation remains to be fully understood [11]. As mentioned above (and reviewed in detail in other chapters) binding of PARP-1 to SSBs that are induced either directly or as intermediate products of base excision repair (BER) appears to protect the damaged site from inappropriate recombination events, with one of the characteristics of PARP-1 depletion being an increase in the frequency of sister chromatid exchanges [12–14].
The absence of the PARP-1 protein through gene depletion in mouse models and derived cells (for example 3T3 PARP-1 −/− MEFs reported by De Murcia and colleagues in 1997) results in radiation sensitivity [12, 15]. PARP-1 −/− mice died in less than 14 days after total body γ-irradiation with a dose of 8 Gy. Autopsy at 3 days post-irradiation revealed a shortening of the villi in the duodenum and massive necrosis of epithelial cells. Human cells depleted in PARP-1, for example HeLa cells stably silenced for PARP-1 expression (PARP-1KD) using pEBVsiRNA vectors as reported by [16], also show increased radiosensitivity and particularly in the S-phase of the cell cycle (Fig. 11.1). Dissecting the role of PARP-1 in this radiation sensitivity is complicated because of its multiple roles in both the repair of SSB and DSBs. There is substantial evidence that SSB repair (SSBR) occurs in the absence of PARP-1 but with altered kinetics (for instance see [15, 17, 18]). In the HeLa cell model, stable PARP-1 silencing had no impact on SSB rejoining in the S phase of the cell cycle which goes to completion and was as fast as in control cells. However in the G1 phase, PARP-1 silencing delayed SSBR by 2.2-fold based on t1/2 values although repair reached completion in < 2 h [16]. This result suggests the existence of an alternative, PARP-1 independent SSBR pathway that can operate more rapidly in the S phase than in G1. Indeed, although the absence of PARP-1 prevents XRCC1-YFP recruitment to sites of laser micro-irradiation (405 nm), GFP-PCNA recruitment was not affected [16] (Fig. 11.2) suggesting that the long-patch sub-pathway is an alternative mechanism under these circumstances. This switching from the short-patch to the long-patch BER sub-pathway in PARP-1 depleted cells could be more favourable in S-phase because most of the long-patch effectors participate in the replication process. These results would suggest that the radiosensitivity seen in PARP-1 depleted cells is due to PARP-1’s involvement in processes other than SSBR.
Radiation survival in control, PARP-1KD and XRCC1KD HeLa cells synchronized in S-phase. Cells were synchronized by a double thymidine block, allowed to progress in S phase for 2 h and exposed to increasing doses of γ-rays. Following treatment cells were allowed to grow as colonies for 10–15 days, fixed, stained with Coomassie blue and counted. The experimental data for control and XRCC1KD lay within the same envelope of statistical deviation whereas the PARP-1KD cells showed enhanced radiation sensitivity. (Reproduced with permission from Godon et al. [16)])
Scheme summarizing the differential effect of PARP-1 inhibition versus silencing on SSB repair via the BER pathway. The experimental evidence would suggest that SSB rejoining proceeds via the LPR sub-pathway when the SPR is deficient because of a lack of PARP-1. In contrast, inhibition of poly(ADP-ribosyl)ation in PARP-1 proficient cells was found to result in the accumulation of PARP-1 and PCNA in the vicinity of DNA damaged sites, with a 10-fold reduction of the bulk rate of SSBR. This does not impact on radiosensitivity in the G1 phase of the cell cycle, probably because cells have enough time to perform SSB repair. However, under such conditions in S phase collision of unrepaired SSBs with replication forks results in the formation of DSBs that contributes to the S-phase radiosensitivity. (Reproduced with permission from Godon et al. [16])
As mentioned above PARP-1 has been found to bind to DNA DSBs and interacts with or modifies proteins involved in the DSB response [19–24]. The hyper-recombination phenotype associated with the loss of PARP-1 expression would also support a role for PARP-1 activity during recombination [12–14]. However the observation that the depletion of PARP-1 does not affect the repair efficiency of I-Sce1 endonuclease induced DSBs between direct repeats [25–27], suggested that PARP-1 does not directly influence the repair of DSBs by homologous recombination repair (HRR). It has to be noted too that PARP-1 depletion in 3T3 fibroblasts did not affect the survival response to neocarzinostatin, an inducer of DSBs [28]. In addition PARP-1 has been implicated in the regulation of non-homologous end joining (NHEJ) , although the full extent of the interplay of the PARP-1/DNA-PK/Ku complex remains to be understood [29, 30]. It is now well accepted that the PARP-1 protein is involved in the alternative-NHEJ repair pathway that promotes microhomology dependent end rejoining in the absence of the Ku proteins [31] and the fact that the PARP-1 and the Ku proteins can interact with the same DNA-ends [32] suggests that the binding of the PARP-1 or the Ku proteins prevents extensive DNA end resection and could also affect NHEJ sub-pathway usage. More recent studies have highlighted a role of PARP-1 in DNA repair during replication, despite the fact that it is not required for normal replication fork progression in untreated cells [23, 26, 33]. PARP-1 depletion results in an hypersensitivity to γ-radiation in S phase cells [16, 28] and to replication fork blocking agents including camptothecin [33], hydroxyurea [23, 26, 34] or thymidine [23]. PARP-1 also facilitates the recruitment of the MRN complex [23], suggesting that PARP-1 could participate in replication fork restart mediated recombination. Thus in the absence of PARP-1 changes in the efficiency of all these processes would be expected and more specifically its role in the processing of DNA damage during replication might contribute to the radiation hypersensitivity phenotype seen in S-phase cells.
As mentioned above as a consequence of poly(ADP-ribosyl)ation, chromatin adopts a more relaxed structure which is thought to facilitate DNA repair. It is known that PARP-1 binds nucleosomes and plays a major role in the condensation of chromatin and transcriptional repression [35–37]. It would appear that the DNA binding domain and catalytic domain of PARP-1 cooperate to regulate chromatin structure and chromatin-dependent transcription and that the enzymatic activation of PARP-1 protein depends on the dynamics of nucleosomal core histone mediation [35, 37]. It was proposed by Ding et al. based on the susceptibility of chromatin to digestion by deoxyribonuclease I or micrococcal nuclease, that the structure of chromatin is altered in cells lacking PARP-1 [17, 38]. The contribution of these chromatin structure changes in the absence of PARP-1 to radiation sensitivity remains to be fully evaluated but clearly will impact on the dynamics of chromatin changes after exposure to ionising radiation. For instance recent data has shown that the chromatin remodelling enzyme ALC1 is recruited to nucleosomes and activated in a poly(ADP-ribose)-dependent manner, most likely via interactions with chromatin associated PAR [39, 40], and in the absence of PARP-1 or in PARP inhibitor treated cells is not recruited to sites of laser induced DNA damage (Fig. 11.3). Other proteins involved in the chromatin remodelling complexes have been found to be recruited at DNA damage site in a PAR-dependent manner [41, 42]. Thus in the absence of PARP-1 there will be some considerable perturbation of the PAR levels in cells and, in particular, the levels that can be generated in response to DNA damage that would be expected to have consequences on the dynamics of changes in chromatin structure in response to DNA damage.
Recruitment of ALC1 to UVA laser damage sites is dependent on the PARP-1 protein and PARP-1 activity a Quantitative analysis of the relative spot intensity with time of YFP-ALC1 recruitment profiles at laser induced DNA damage sites in Control cells treated with or without the PARP inhibitor ABT-888 and in PARP-1KD cells (squares, circles and rhombi). Data represents mean relative spot intensity ± SEM, n = 6–30 individual cells from several independent experiments. Inset: western blot analysis confirming stable depletion of PARP-1 in HeLa PARP-1KD cells. b Representative images of the recruitment of fluorescent tagged ALC1 to DNA damage sites in Control cells treated with or without the PARP inhibitor ABT-888 and in PARP-1KD cells. Scale bar: 5 μm
In several model systems it has been noted that in the absence of PARP-1 there is a concomitant reduction in the level of the poly(ADP-ribose) glycohydrolase (PARG) protein [41–43]. Furthermore, it was recently reported that PARP-1 mRNA and protein expression is down-regulated by the knockdown of the Parg gene [43–45]. Whether these changes are an adaptive means of maintaining the substantially lower PAR levels that can be generated by the other PARPs to allow some PAR dependent processes to occur remains to be established but they highlight the complex inter-relationship of these proteins and are another potential contributing factor to the radiosensitivity phenotype seen in the absence of PARP-1.
The absence of PARP-2 is also associated with radiosensitivity although PARP-2 knockout mice are generally less sensitive to DNA damage produced by ionising radiation than PARP-1 knockout mice [46]. This could be due to the lower abundance of PARP-2 and/or different roles of the two proteins and the availability of back-up mechanisms that are operational in its absence. PARP-2 −/− mice present a similar radiosensitivity phenotype to PARP-1 −/− mice with dilated duodenum, reduction of villi’s length and degeneration of epithelial crypt cells. In both knockout models, cause of death is attributed to radio-induced intestinal toxicities. In addition to the intestinal phenotype, both thymocytes and hematopoietic cells isolated from PARP2 −/− animals are hypersensitive to radiation exposure [47, 48] revealing a crucial role of PARP-2 in cell survival. Hematopoietic cells isolated from PARP2 −/− after 2 Gy γ-irradiation presented an increase of chromatid breaks suggesting a deficiency in DNA damage response [48]. After sub-lethal γ-irradiation, PARP2 −/− mice showed an impaired DNA damage response characterized by increased γ-H2AX foci in hematopoietic stem cells associated with increased apoptosis and reduced regenerative capacity of the hematopoietic stem cell pool. PARP-2 knockout cells were reported to be more sensitive to doses below 2 Gy, raising the possibility that PARP-2 plays a role in the G2/M phase cell cycle checkpoint activated in response to low doses of ionising radiation [49, 50]. Radiation sensitivity was also seen in a HeLa cell model depleted for PARP-2 but in contrast to PARP-1 depleted cells this radiation sensitivity was not increased specifically in S-phase cells [43]. Although several studies have shown that the depletion of the PARP-2 protein is associated with a reduction in the total PAR synthesis [51, 52], others have reported increased PARP activity in the absence of PARP-2 [47, 53]. Increased PARP activity in the absence of PARP-2 has been observed in thymocytes and stimulated B lymphocytes from PARP2 −/− mice that show at least a 20 % increase in PAR synthesis compared to Control cells [47, 53]. A similar increase was also seen in PARP-2 depleted HeLa cells [43]. The mechanistic basis of the increase in PARP-1 activity in the absence of PARP-2 is unknown. The observation that the PAR synthesis rate catalysed in the presence of both PARP-1 and PARP-2 was lower than in the presence of PARP-1 alone [51, 54] suggests that the PARP-1/PARP-2 heterodimers might be less active than monomers or homo-dimers of PARP-1. Alternatively following activation through binding to DNA structures the poly(ADP-ribosyl)ation of PARP-1 by PARP-2 could impact on its enzymatic activity. In the HeLa cell model used in our laboratory, the lower PARG protein levels seen in the PARP-2 stably depleted cells was a consequence of decreased PARG mRNA level (our unpublished data) suggesting a direct role of PARP-2 in the regulation of the expression of PARG. Indeed, recent reports have revealed that the depletion of PARP-2 modifies the activity of multiple transcription factors and in particular it is involved in the transcriptional regulation of metabolism and oxidative stress responses [55].
In addition to an increased sensitivity to ionising radiation, PARP-2 depleted cells were sensitive to neocarzinostatin in comparison to PARP-1 [43] and PARP-2 suppressed translocations between c-myc and IgH during induced class switch recombination [53]. Taken together these data suggest that PARP-2 may play a role in the processing of DSBs. Clearly the perturbation of SSB and DSB repair in the absence of PARP-2 will contribute to the radiosensitivity seen in PARP-2 depleted cells; however a contribution from an alteration in the dynamics of chromatin structure changes could also be a contributing factor. As mentioned above poly(ADP-ribosy)lation is a major regulatory factor in the recruitment of chromatin remodelling factors. The absence of PARP-2 impacts on the dissociation of ALC1 from sites of DNA damage suggesting that chromatin structure may also be modified in response to DNA damage in the absence of PARP-2 (our unpublished data). The loss of PARP-2 expression has also been reported to increase the frequency of spontaneous chromosome and chromatid breaks and the number of DNA ends lacking detectable telomere repeats [56]. This might result from faulty chromosome segregation during the late steps of cytokinesis. Thus, as for the radiation sensitivity phenotype seen in the absence of PARP-1 there may be many contributory factors to the radiosensitivity phenotype seen in PARP-2’s absence.
The third PARP family member implicated in the responses to DNA damage is PARP-3. PARP-3 depleted MRC5 or A549 cells displayed a significant increase in γ-H2AX foci in untreated cells, suggesting the presence of spontaneous strand breaks, and after X-irradiation a significant delay in the repair of radio-induced DSBs compared with control cells [57, 58]. PARP-3 has been shown to be stimulated by DNA DSBs in vitro and facilitates DNA ligation through a mechanism involving PARP-3 in the accumulation of APLF at DSBs that in turn promotes the retention of XRCC4/DNA ligase IV [58]. Although, PARP-3 is known to localize with the daughter centriole and may play a specific role in cytokinesis, possibly at the mitotic fidelity checkpoint [59], the depletion of PARP-3 was not found to significantly impact on the long-term sensitivity to X-irradiation using colony-forming ability as an endpoint suggesting a compensating repair activity over time by another DNA damage-induced PARP family member (i.e., PARP-1 and/or PARP-2). This hypothesis was confirmed by comparing the radio-induced sensitivity of PARP-3 depleted and control cells to 1 Gy X-ray irradiation in the absence or presence of the PARP inhibitor KU-0058948 (100 nM). The additional inhibition of the remaining PARP activity in PARP-3 depleted cells significantly reduced their survival (by two-fold) after X-irradiation compared with mock-treated PARP-3 depleted cells. These observations were confirmed in mouse models, where the loss of PARP-3 alone did not decrease survival after X-irradiation, whilst the combined loss of PARP-1 and PARP-3 had a significant impact [60]. The variation in PARP-3 mRNA levels has been assessed in human breast tumours where an under-expression was observed in 10.4 % of tumours, more frequently in the hormone receptor-negative tumours (25.4 %) than the hormone receptor-positive tumours (5.9 %) [61]. Intriguingly this PARP-3 under-expression was mutually exclusive with a PARP-1 over-expression [61]. Clearly the balance of the expression of the different PARPs is fine-tuned and it’s tempting to speculate that such changes may reflect compensating activities between the different members of the PARP families and the PARG protein.
3 Radiosensitisation and PARP Inhibition
The prototype PARP inhibitor 3-AB was first shown to enhance the response of mammalian cells to ionising radiation in the mid-1980s [8, 62]. These earlier studies recognised that the inhibitor exposure time impacted on the enhanced radiation response and that a good correlation existed between the potency of the inhibitor and enhanced cell killing. They also showed that repair deficient cells responded to lower concentrations of inhibitor compared to radiation resistant cell lines.
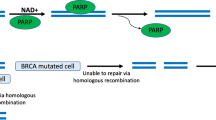
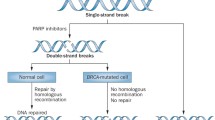
3-AB, like many of the second and third generation inhibitors subsequently developed, acts through binding to the enzyme’s catalytic domain. Thus whilst the inhibitors are specific for PARP activity they show varying degrees of selectivity for the different members of the PARP superfamily [63]. A direct consequence of this inhibition of multiple PARPs is that the effects are likely to be greater and/or different than the depletion of a single PARP family member. Indeed these competitive inhibitors do not inhibit PARP-1 binding to DNA but do inhibit PAR formation that is essential for the dissociation of PARPs from the strand break. Thus their use would be expected to result in an accumulation of PARP-1 at DNA damage sites. This potential consequence of PARP inhibition was first proposed by Satoh and Lindahl based on results obtained investigating the role of PAR formation in DNA repair in a human cell-free system over 20 years ago [64]. The trapping of the PARP-1 protein and probably other PARPs too, will generate a situation where the recruitment of other repair factors could be obstructed. The recruitment of PARP-1 to sites of micro-laser induced DNA damage can be visualised using GFP-tagged PARP-1 by live cell video-microscopy. Under such experimental conditions Mortusewicz et al. found that the recruitment of GFP-PARP-1 in MEFs lacking PARP-1 or in HeLa cells was efficient but transient and that treatment with the inhibitor NU1025 lead to a delayed and prolonged accumulation of GFP-PARP-1 to damage sites [65]. Using a mutant form of GFP-PARP-1, that converts PARP-1 into a mono-ADP-ribosyl-transferase, expressed in MEFs lacking PARP-1 a delayed accumulation and longer persistence at DNA damage sites in comparison to the wildtype protein was observed [65]. Under similar experimental conditions and using a HeLa cell model we observed an immediate recruitment of GFP-PARP (< 10 s) at damage sites that formed discrete foci that had dissipated in about 20 min. In contrast, in the presence of the PARP inhibitor 4-amino-1,8-naphthalimide (ANI), GFP-PARP-1 formed larger intense spots of fluorescence at 5 min post-irradiation compared to untreated cells that still persisted at 20 min. A similar recruitment pattern was observed for GFP-PCNA in the presence of ANI [16]. In addition XRCC1 recruitment is hampered when PARP activity is inhibited [5, 16, 65–67]. This altered recruitment caused by PARP inhibition impairs SSBR in PARP-1 proficient cells with different outcomes in S and G1 phase cells. As discussed above in our HeLa cell model SSBR, measured by alkaline elution analysis, in the absence of ANI proceeded to completion in < 20 min, irrespective of whether the cells where in the G1 or S phases of the cell cycle. Although ANI slowed down the SSBR in both phases, in the G1 cells SSBR was completed after 5 h whilst in cells treated in the S phase as much as 26 % of the initial load of SSBs remained unjoined from 2 to 6 h post-irradiation, demonstrating a clear block in SSBR [16]. This block correlated with the induction of a large number of de novo DSBs that are suppressed by aphidicolin, indicating that they originate from the progression of a replication fork towards unrepaired SSBs [68]. Chalmers and colleagues also showed that the PARP-1 and PARP-2 inhibitor KU-0059436/AZD-2281/Olaparib increased the radiosensitivity of human glioma cell lines by a replication-dependent mechanism that generates persistent DSBs [69]. The cell cycle dependency of PARP inhibition had been reported as early as 1984 when Ben-Hur et al. showed that the inhibitor nicotinamide enhanced the killing mainly of late S-phase cells and did not affect cells at the G1/S border [62]. These de novo DSBs are repaired by HRR and PARP inhibition results in the accumulation of recombinogenic substrates marked by Rad51 and γ-H2AX nuclear foci [68, 70]. This explains why cancer cells deficient in BRCA1 and BRCA2 are selectively hypersensitive to PARP inhibitors [71, 72]. More recent studies have also suggested that disruption of the Fanconi anemia, PTEN, ATM and other HRR-related genes also sensitises to PARP inhibitors but the response of tumoral cells is complex and enhanced cytoxicity is not always observed.
Murai et al. showed that both PARP-1 and PARP-2 can be trapped at damaged DNA by PARP inhibitors and that the trapped complexes are more cytotoxic than unrepaired SSBs in the absence of PARP [73, 74]. They also showed that the potency of trapping PARP differed between inhibitors with niriparib (MK-4827) > olaparib (AZD-2281) >> veliparib (ABT-888), a pattern not correlated with the catalytic inhibitory properties of each drug [74]. More recently they have shown that the novel inhibitor BMN 673 is approximately. 100 fold more potent at trapping PARP-DNA complexes, shows stereospecific binding to PARP-1 and is more cytotoxic as a single agent and in combination with the alkylating agent methyl methanesulphonate (MMS) and TMZ than olaparib [73]. They also reported that PARP-1/2 knockout cells have a high level of resistance to BMN 673, as did half of the NCI-60 cell lines even at doses of BMN 673 as high as 100 µmol/L [73]. The PARP-1 trapping model can explain why wildtype cells are more sensitive to a PARP inhibitor combined with the alkylating agent MMS than PARP-1 −/− mouse cells [75–78]. These results highlight important considerations for the clinical use of PARP inhibitors as the expression of genes that are critical for the repair of PARP-DNA complexes would be expected to modulate the sensitivity to PARP inhibitors.
4 In Vivo Radiation Sensitivity After PARP Inhibition: Additional Benefits and Use in Combination with New Radiation Techniques
A number of PARP inhibitors (ABT-888/Veliparib, AZD-2281/Olaparib, AG014699/Rucaparib, MK-4827/Niraparib, AG14361, GPI-15427, E7016) have been shown to enhance tumour sensitivity to radiation in syngeneic and xenograft models of breast [79, 80], colon [81–83], lung [79, 84, 85], nasopharyngeal [86], head and neck [87], and brain tumours [88–91] (Table 11.1). PARP inhibition combined with ionising radiation resulted in delayed tumour growth and extended median survival time. Immunohistochemical analysis of tumour sections after combined treatment revealed an increased number of apoptotic cells and a decrease in proliferating cells [84, 87]. As previously described for nicotinamide and other benzamide analogs [92], PARP inhibitors can induce temporary vasodilation increasing perfusion of tumour blood vessels [82, 85]. This vascular side effect of PARP inhibitors treatment enhances drug delivery (e.g. TMZ in glioblastoma) and can modulate the oxygen concentration in the tumour microenvironment. Hypoxic cells are 2.5-3-fold more radioresistant than oxic cells and intra-tumoural hypoxia can be a significant cause of treatment failure after radical radiotherapy. Work from the Bristow lab has demonstrated that the PARP inhibitor ABT-888 was a radiosensitizer in hypoxic conditions in prostate and lung cancer cell lines [93], highlighting their role as attractive adjuncts for radiotherapy via mechanisms other than the inhibition of DNA repair.
In addition to the effects of PARP inhibitors on the ability to repair DNA damage and the presence of tissue hypoxia and vascularisation, factors that will all influence the probability of tumour control, additional therapeutic benefits of PARP inhibition have been noted when combined with radiotherapy. Radiotherapy is frequently used in the treatment of mediastinal malignant diseases during which the lungs, aorta and coronary arteries are often in the irradiation fields. The side effects of such irradiation include lung and intimal fibrosis and the obliteration of irradiated vessels which can lead in extreme cases to cardiac failure. Damage to the vasculature also occurs to other tissues and is a hallmark of the deleterious complications of radiotherapy [94]. It has long been hypothesized (reviewed in [95]) that long-lasting oxidative stress may be a mechanism of radiation-induced vascular injury. In particular peroxynitrite that is formed by the rapid reaction of superoxide anion and nitric oxide has been established as a powerful, endogenous trigger of DNA damage whose repair requires PARP-1. Indeed the high levels of base damage and peroxynitrite adducts in DNA generated under such situations can trigger in PARP-1 over-activation resulting in the rapid depletion of intracellular ATP pools and impaired mitochondrial respiration and eventually leading to cellular dysfunction and the death of endothelial cells [96]. Reactive oxygen species also elicit specific signalling in vascular smooth muscle cells [97]. For these reasons one of the first PARP inhibitors were originally designed to protect the endothelium from PARP-1 over-activation induced in response to ischaemia-reperfusion injury, circulatory shock, diabetic complications and atherosclerosis [98, 99].
PARP inhibitors were subsequently found to have anti-angiogenic effects in ex vivo models [100, 101]. For instance the PARP inhibitor INO-1001 was found to decrease nitrosative stress and protect rat aorta in an ex vivo model from loss of vasomotor function following 20 Gy radiation in a single fraction [102]. INO-1001 also reduced neointimal hyperplasia and activation of the peroxynitrite-PARP pathway subsequent to irradiation of endarterectomized rats [103] and protected rat aortic rings ex vivo from loss of the endothelial function induced by H2O2 [104]. Taken together these data suggest that, in addition to acting as potentiators of DNA-damaging drugs and radiation in strictly concomitant administration, PARP inhibitors might be used long after treatment to protect normal tissues from the deleterious effects of radiotherapy, notably chronic inflammatory processes and fibrosis.
Radiation induced fibrosis is characterized by a progressive and excessive accumulation of extracellular matrix (ECM) which may severely compromise normal tissue function. Among the factors that play a role in remodelling of the ECM two extracellular proteases play a crucial role, namely, the plasminogen activator inhibitor-1 (PAI-1) and the matrix metalloproteinase system. The transforming growth factor-β1 (TGF-β1) is the most potent transcriptional activator of these systems and the late, deleterious effects of radiotherapy are likely to be related to dysfunction of tissue healing under control of the TGF-β pathway [105]. The links between PARP-1 and the TGF-β pathway remain to be fully dissected with data published to date suggesting that cell and tissue type specific differences exist. Upon activation, TGF-β1 dimerises and binds to a cell surface receptor complex that can activate either the activin-like kinase 5 (ALK5, also known as nexin) or activin-like kinase 1 (ALK1, also known as endoglin) receptors, ending ultimately in the activation of different members of the Smad family of transcription factors and their target genes [106]. ALK1 opposes ALK5 signalling and the expression of ECM components [107]. The activated Smads translocate to the nucleus and initiate gene-specific transcription in cooperation with other transcription factors. For instance, p53 cooperates transcriptionally with certain of the Smad complexes leading to the radio-induced transcriptional activation of the profibrotic plasminogen activator inhibitor-1 (PAI-1) gene [108–110]. There is growing evidence that ionising radiation shifts the balance between the PAI-1 vs. ID-1 (inhibitor of DNA binding-1) pathways [111] and that PARP-1 is involved in the regulation of the Smads downstream of TGF-β1 in endothelial and vascular smooth muscle cells. In animal models PARP inhibitors have been shown to alleviate fibrotic remodelling of blood arteries by inhibiting the TGF-β/Smad3 pathway [112], high glucose-induced peritoneal epithelial mesenchymal transition (EMT) and fibrosis [113], diabetic or hypertensive cardiomyopathy [114–118] and a variety of syndromes linked to chronic oxidative stress or exaggerated inflammation [119, 120]. Taken together, these data suggest that targeting PARP-1 might be a promising therapeutic approach against vascular or pro-fibrotic diseases induced by dysregulation of the TGF-β/Smad3 pathway following irradiation.
Over the past decade, a number of innovative radiotherapy techniques have been developed that are being introduced into clinical practice. These include volumetric-modulated arc therapy (RapidArc®, TomoTherapy®) and multi-beam stereotactic irradiation (Cyberknife®) that involve the rapid alternation of radiation beams and/or split-dose irradiation of tissues over a time scale spanning from a few seconds to minutes and pencil beam irradiation that uses high dose-rates, up to 200-fold higher than those used in conventional external-beam therapy. The effects of such high dose-rates and split-dose irradiation, or how PARP inhibitors can be used in combination with such techniques, have not yet been fully investigated in detail in vivo although recent data has shown that ultra-high dose-rate flash irradiation maintains antitumor efficiency and increases lung radio-tolerance in C57BL/6J mice [121]. No substantial difference in mammalian cell survival to pulsed, ultra-high dose-rate relativistic electrons compared to X-rays at conventional dose-rates was reported [122]. Whilst this observation is consistent with the fact that the density of energy deposit and ionisation at the microscopic scale is the same for relativistic electrons and X- or γ-rays with energies > 600 keV, the situation may be different when the dose is fractionated with delays in the same time range as DNA damage recognition and repair. Indeed using a linear electron accelerator operated in a chopped mode for time-resolved investigation of split-dose radiation recovery in mammalian cells in vitro, we demonstrated that the first (priming) irradiation induces fast, synchronous oscillations of the cellular radiosensitivity, yielding a tetraphasic survival curve as a function of the delay separating the two pulses [123, 124]. This phenomenon was shown to require the presence and catalytic activity of PARP [124] whilst NHEJ repair of DNA double-strand breaks was not involved [123]. It has also been shown that the incidence of delayed cell death after a single pulse of radiation was reduced by a factor of 2.5 compared to that occurring after protracted X-ray irradiation at a conventional dose-rate [123]. Thus it may be expected that the impact of PARP inhibitors will depend on the relative dose-rate of the radiation used and the extension of the use of PARP inhibitors in combination with these technologies will need to be carefully evaluated.
5 Conclusions
The inhibition of PARP activity has potential clinical benefits when combined with radiotherapy in strictly concomitant administration protocols through mechanisms that involve DNA repair perturbation, changes in intrinsic radiosensitivity and tissue hypoxia that all impact on tumour control. Preclinical evidence would suggest that there may also be a role for these drugs in adjuvant therapy to reduce the long-term side effects of radiotherapy . There are, however, many aspects of their use that still need to be established. It is known that the complex interaction of irradiation dose and schedule will influence the risk of developing adverse secondary reactions such as radiation-induced fibrosis and secondary cancers. Thus there is a clear need to optimize the PARP inhibitor administration protocols to maximise efficacy whilst minimising secondary side-effects and fully evaluate the impact of the inhibition of multiple PARPs. Finally, whether the introduction of these innovative techniques, and thus the increased possibility of modulating parameters such as the dose-rate or using radiation sources of different quality (protons vs. X-rays), can be exploited to generate a panel of DNA end lesions in treated cells that will maximise PARP protein binding and subsequent trapping by the inhibition of PARP activity also warrants further investigation to extend the clinical use of PARP inhibitors.
References
Chambon P, Weill J, Mandel P (1963) Nicotinamide mononucleotide activation of new DNA-dependent polyadenylic acid synthesizing nuclear enzyme. Biochem Biophys Res Commun 11:39–43
Virág L, Szabó C (2002) The therapeutic potential of poly(ADP-ribose) polymerase inhibitors. Pharmacol Rev 54:375–429
Schreiber V, Dantzer F, Ame J-C, de Murcia G (2006) Poly(ADP-ribose): novel functions for an old molecule. Nat Rev Mol Cell Biol 7:517–528. doi:10.1038/nrm1963
D’Amours D, Desnoyers S, D’Silva I, Poirier GG (1999) Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem J 342:249–268
El-Khamisy SF, Masutani M, Suzuki H, Caldecott KW (2003) A requirement for PARP-1 for the assembly or stability of XRCC1 nuclear foci at sites of oxidative DNA damage. Nucleic Acids Res 31:5526–5533. doi:10.1093/nar/gkg761
Caldecott KW (2003) XRCC1 and DNA strand break repair. DNA Repair (Amst) 2:955–969. doi:10.1016/S1568-7864(03)00118-6
Durkacz B, Omidiji O, Gray D, Shall S (1980) (ADP-ribose) n participates in DNA excision repair. Nature 283:593–596
Ben-Hur E, Chen C, Elkind M (1985) Inhibitors of poly (adenosine diphosphoribose) synthetase, examination of metabolic perturbations, and enhancement of radiation response in Chinese hamster cells. Cancer Res 45:2123–2127
Powell C, Mikropoulos C, Kaye SB, Nutting CM, Bhide SA, Newbold K, Harrington KJ (2010) Pre-clinical and clinical evaluation of PARP inhibitors as tumour-specific radiosensitisers. Cancer Treat Rev 36:566–575. doi:10.1016/j.ctrv.2010.03.003
Chalmers AJ, Lakshman M, Chan N, Bristow RG (2010) Poly(ADP-ribose) polymerase inhibition as a model for synthetic lethality in developing radiation oncology targets. Semin Radiat Oncol 20:274–281. doi:10.1016/j.semradonc.2010.06.001
De Vos M, Schreiber V, Dantzer F (2012) The diverse roles and clinical relevance of PARPs in DNA damage repair: current state of the art. Biochem Pharmacol 84:137–146. doi:10.1016/j.bcp.2012.03.018
De Murcia JM, Niedergang C, Trucco C, Ricoul M, Dutrillaux B, Mark M, Oliver FJ, Masson M, Dierich A, Lemeur M, Walztinger C, Chambon P, de Murcia G (1997) Requirement of poly (ADP-ribose) polymerase in recovery from DNA damage in mice and in cells Requirement of poly (ADP-ribose) polymerase in recovery from DNA damage in mice and in cells. Proc Natl Acad Sci U S A 94:7303–7307
Wang Z, Stingl L, Morrison C, Jantsch M, Los M, Schulze-Osthoff K, Wagner EF (1997) PARP is important for genomic stability but dispensable in apoptosis. Genes Dev 11:2347–2358. doi:10.1101/gad.11.18.2347
Meyer R, Müller M, Beneke S, Küpper JH, Bürkle A (2000) Negative regulation of alkylation-induced sister-chromatid exchange by poly(ADP-ribose) polymerase-1 activity. Int J Cancer 88:351–355
Trucco C, Oliver FJ, de Murcia G, Ménissier-de Murcia J (1998) DNA repair defect in poly(ADP-ribose) polymerase-deficient cell lines. Nucleic Acids Res 26:2644–2649
Godon C, Cordelières FP, Biard D, Giocanti N, Mégnin-Chanet F, Hall J, Favaudon V (2008) PARP inhibition versus PARP-1 silencing: different outcomes in terms of single-strand break repair and radiation susceptibility. Nucleic Acids Res 36:4454–4464. doi:10.1093/nar/gkn403
Ding R, Pommier Y, Kang VH, Smulson M (1992) Depletion of poly(ADP-ribose) polymerase by antisense RNA expression results in a delay in DNA strand break rejoining. J Biol Chem 267:12804–12812
Fisher AEO, Hochegger H, Takeda S, Caldecott KW (2007) Poly(ADP-ribose) polymerase 1 accelerates single-strand break repair in concert with poly(ADP-ribose) glycohydrolase. Mol Cell Biol 27:5597–5605. doi:10.1128/MCB.02248-06
Ariumi Y, Masutani M, Copeland TD, Mimori T, Sugimura T, Shimotohno K, Ueda K, Hatanaka M, Noda M (1999) Suppression of the poly(ADP-ribose) polymerase activity by DNA-dependent protein kinase in vitro. Oncogene 18:4616–4625. doi:10.1038/sj.onc.1202823
Galande S, Kohwi-Shigematsu T (1999) Poly(ADP-ribose) polymerase and Ku autoantigen form a complex and synergistically bind to matrix attachment sequences. J Biol Chem 274:20521–20528
Haince J-F, Kozlov S, Dawson VL, Dawson TM, Hendzel MJ, Lavin MF, Poirier GG (2007) Ataxia telangiectasia mutated (ATM) signaling network is modulated by a novel poly(ADP-ribose)-dependent pathway in the early response to DNA-damaging agents. J Biol Chem 282:16441–16453. doi:10.1074/jbc.M608406200
Haince J-F, McDonald D, Rodrigue A, Déry U, Masson J-Y, Hendzel MJ, Poirier GG (2008) PARP1-dependent kinetics of recruitment of MRE11 and NBS1 proteins to multiple DNA damage sites. J Biol Chem 283:1197–1208. doi:10.1074/jbc.M706734200
Bryant HE, Petermann E, Schultz N, Jemth A-S, Loseva O, Issaeva N, Johansson F, Fernandez S, McGlynn P, Helleday T (2009) PARP is activated at stalled forks to mediate Mre11-dependent replication restart and recombination. EMBO J 28:2601–2615. doi:10.1038/emboj.2009.206
Hochegger H, Dejsuphong D, Fukushima T, Morrison C, Sonoda E, Schreiber V, Zhao GY, Saberi A, Masutani M, Adachi N, Koyama H, de Murcia G, Takeda S (2006) PARP-1 protects homologous recombination from interference by Ku and Ligase IV in vertebrate cells. EMBO J 25:1305–1314. doi:10.1038/sj.emboj.7601015
Schultz N, Lopez E, Saleh-Gohari N, Helleday T (2003) Poly(ADP-ribose) polymerase (PARP-1) has a controlling role in homologous recombination. Nucleic Acids Res 31:4959–4964. doi:10.1093/nar/gkg703
Yang Y-G, Cortes U, Patnaik S, Jasin M, Wang Z-Q (2004) Ablation of PARP-1 does not interfere with the repair of DNA double-strand breaks, but compromises the reactivation of stalled replication forks. Oncogene 23:3872–3882. doi:10.1038/sj.onc.1207491
Adamson B, Smogorzewska A, Sigoillot F, King R, Elledge SJ (2012) A genome-wide homologous recombination screen identifies the RNA-binding protein RBMX as a component of the DNA-damage response. Nat Cell Biol 14:318–328. doi:10.1038/ncb2426.A
Noël G, Giocanti N, Fernet M, Mégnin-Chanet F, Favaudon V (2003) Poly(ADP-ribose) polymerase (PARP-1) is not involved in DNA double-strand break recovery. BMC Cell Biol 4:7. doi:10.1186/1471-2121-4-7
Ruscetti T, Lehnert BE, Halbrook J, Le Trong H, Hoekstra MF, Chen DJ, Peterson SR (1998) Stimulation of the DNA-dependent protein kinase by poly(ADP-ribose) polymerase. J Biol Chem 273:14461–14467
Spagnolo L, Barbeau J, Curtin NJ, Morris EP, Pearl LH (2012) Visualization of a DNA-PK/PARP1 complex. Nucleic Acids Res 40:4168–4177. doi:10.1093/nar/gkr1231
Audebert M, Salles B, Calsou P (2004) Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining. J Biol Chem 279:55117–55126. doi:10.1074/jbc.M404524200
Wang M, Wu W, Wu W, Rosidi B, Zhang L, Wang H, Iliakis G (2006) PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res 34:6170–6182. doi:10.1093/nar/gkl840
Sugimura K, Takebayashi S-I, Taguchi H, Takeda S, Okumura K (2008) PARP-1 ensures regulation of replication fork progression by homologous recombination on damaged DNA. J Cell Biol 183:1203–1212. doi:10.1083/jcb.200806068
Gottipati P, Vischioni B, Schultz N, Solomons J, Bryant HE, Djureinovic T, Issaeva N, Sleeth K, Sharma RA, Helleday T (2010) Poly(ADP-ribose) polymerase is hyperactivated in homologous recombination-defective cells. Cancer Res 70:5389–5398. doi:10.1158/0008-5472.CAN-09-4716
Wacker DA, Ruhl DD, Balagamwala EH, Hope KM, Zhang T, Kraus WL (2007) The DNA binding and catalytic domains of poly(ADP-ribose) polymerase 1 cooperate in the regulation of chromatin structure and transcription. Mol Cell Biol 27:7475–7485. doi:10.1128/MCB.01314-07
Huletsky A, de Murcia G, Muller S, Hengartner M, Ménard L, Lamarre D, Poirier GG (1989) The effect of poly(ADP-ribosyl)ation on native and H1-depleted chromatin. A role of poly(ADP-ribosyl)ation on core nucleosome structure. J Biol Chem 264:8878–8886
Pinnola A, Naumova N, Shah M, Tulin AV (2007) Nucleosomal core histones mediate dynamic regulation of poly(ADP-ribose) polymerase 1 protein binding to chromatin and induction of its enzymatic activity. J Biol Chem 282:32511–32519. doi:10.1074/jbc.M705989200
Ding R, Smulson M (1994) Depletion of nuclear poly (ADP-ribose) polymerase by antisense RNA expression: influences on genomic stability, chromatin organization, and carcinogen cytotoxicity. Cancer Res 54(17):4627–4634
Gottschalk AJ, Timinszky G, Kong SE, Jin J, Cai Y, Swanson SK, Washburn MP, Florens L, Ladurner AG, Conaway JW, Conaway RC (2009) Poly(ADP-ribosyl)ation directs recruitment and activation of an ATP-dependent chromatin remodeler. Proc Natl Acad Sci U S A 106:13770–13774. doi:10.1073/pnas.0906920106
Ahel D, Horejsí Z, Wiechens N, Polo SE, Garcia-Wilson E, Ahel I, Flynn H, Skehel M, West SC, Jackson SP, Owen-Hughes T, Boulton SJ (2009) Poly(ADP-ribose)-dependent regulation of DNA repair by the chromatin remodeling enzyme ALC1. Science 325:1240–1243. doi:10.1126/science.1177321
Erdélyi K, Bai P, Kovács I, Szabó E, Mocsár G, Kakuk A, Szabó C, Gergely P, Virág L (2009) Dual role of poly(ADP-ribose) glycohydrolase in the regulation of cell death in oxidatively stressed A549 cells. FASEB J 23:3553–3563. doi:10.1096/fj.09-133264
Frizzell KM, Gamble MJ, Berrocal JG, Zhang T, Krishnakumar R, Cen Y, Sauve AA, Kraus WL (2009) Global analysis of transcriptional regulation by poly(ADP-ribose) polymerase-1 and poly(ADP-ribose) glycohydrolase in MCF-7 human breast cancer cells. J Biol Chem 284:33926–33938. doi:10.1074/jbc.M109.023879
Boudra MT, Bolin C, Chiker S, Fouquin A, Zaremba T, Vaslin L, Biard D, Cordelières FP, Mégnin-Chanet F, Favaudon V, Fernet M, Pennaneach V, Hall J (2014) PARP-2 depletion results in lower radiation cell survival but cell line-specific differences in poly(ADP-ribose) levels. Cell Mol Life Sci. Epub ahead of print
Uchiumi F, Watanabe T, Ohta R, Abe H, Tanuma S-I (2013) PARP1 gene expression is downregulated by knockdown of PARG gene. Oncol Rep 29:1683–1688. doi:10.3892/or.2013.2321
Ambrose HE, Willimott S, Beswick RW, Dantzer F, de Murcia JM, Yelamos J, Wagner SD (2009) Poly(ADP-ribose) polymerase-1 ( PARP-1)-deficient mice demonstrate abnormal antibody responses. Immunology 127:178–186. doi:10.1111/j.1365-2567.2008.02921.x
Ménissier de Murcia J, Ricoul M, Tartier L, Niedergang C, Huber A, Dantzer F, Schreiber V, Amé J-C, Dierich A, LeMeur M, Sabatier L, Chambon P, de Murcia G (2003) Functional interaction between PARP-1 and PARP-2 in chromosome stability and embryonic development in mouse. EMBO J 22:2255–2263. doi:10.1093/emboj/cdg206
Yélamos J, Monreal Y, Saenz L, Aguado E, Schreiber V, Mota R, Fuente T, Minguela A, Parrilla P, de Murcia G, Almarza E, Aparicio P, Ménissier-de Murcia J (2006) PARP-2 deficiency affects the survival of CD4 + CD8 + double-positive thymocytes. EMBO J 25:4350–4360. doi:10.1038/sj.emboj.7601301
Farrés J, Martín-Caballero J, Martínez C, Lozano JJ, Llacuna L, Ampurdanés C, Ruiz-Herguido C, Dantzer F, Schreiber V, Villunger A, Bigas A, Yélamos J (2013) PARP-2 is required to maintain hematopoiesis following sublethal γ-irradiation in mice. Blood 122:44–54. doi:10.1182/blood-2012-12-472845
Yélamos J, Schreiber V, Dantzer F (2008) Toward specific functions of poly(ADP-ribose) polymerase-2. Trends Mol Med 14:169–178. doi:10.1016/j.molmed.2008.02.003
Schreiber V, Ricoul M, Amé J, Dantzer F, Meder V, Spenlehauer C, Stiegler P, Niedergang C, Sabatier L, Favaudon V, Menissier-de Murcia J, de Murcia G (2006) PARP-2: structure-function relationship. In: Poly(ADP-Ribosy)ation edited by Alexander Bϋrkle ©2006 Landes biosecience and Springer Science+ Business media pp 13–31
Schreiber V, Amé J-C, Dollé P, Schultz I, Rinaldi B, Fraulob V, Ménissier-de Murcia J, de Murcia G (2002) Poly(ADP-ribose) polymerase-2 (PARP-2) is required for efficient base excision DNA repair in association with PARP-1 and XRCC1. J Biol Chem 277:23028–23036. doi:10.1074/jbc.M202390200
Szántó M, Rutkai I, Hegedus C, Czikora Á, Rózsahegyi M, Kiss B, Virág L, Gergely P, Tóth A, Bai P (2011) Poly(ADP-ribose) polymerase-2 depletion reduces doxorubicin-induced damage through SIRT1 induction. Cardiovasc Res 92:430–438. doi:10.1093/cvr/cvr246
Robert I, Dantzer F, Reina-San-Martin B (2009) PARP1 facilitates alternative NHEJ, whereas PARP2 suppresses IgH/c-myc translocations during immunoglobulin class switch recombination. J Exp Med 206:1047–1056. doi:10.1084/jem.20082468
Kutuzov MM, Khodyreva SN, Amé J-C, Ilina ES, Sukhanova MV, Schreiber V, Lavrik OI (2013) Interaction of PARP-2 with DNA structures mimicking DNA repair intermediates and consequences on activity of base excision repair proteins. Biochimie 95:1208–1215. doi:10.1016/j.biochi.2013.01.007
Szántó M, Brunyánszki A, Kiss B, Nagy L, Gergely P, Virág L, Bai P (2012) Poly(ADP-ribose) polymerase-2: emerging transcriptional roles of a DNA-repair protein. Cell Mol Life Sci 69:4079–4092. doi:10.1007/s00018-012-1003-8
Dantzer F, Giraud-Panis M, Jaco I, Ame J-C, Schultz I, Blasco M, Koering C-E, Gilson E, Menissier-de Murcia J, de Murcia G, Schreiber V (2004) Functional Interaction between Poly(ADP-Ribose) Polymerase 2 (PARP-2) and TRF2: PARP Activity Negatively Regulates TRF2. Mol Cell Biol 24:1595–1607. doi:10.1128/MCB.24.4.1595-1607.2004
Boehler C, Gauthier L, Yelamos J, Noll A, Schreiber V, Dantzer F (2011) Phenotypic characterization of PARP-1 and PARP-2 deficient mice and cells. Methods Mol Biol 780:313–336. doi:10.1007/978-1-61779-270-0_19
Rulten SL, Fisher AEO, Robert I, Zuma MC, Rouleau M, Ju L, Poirier G, Reina-San-Martin B, Caldecott KW (2011) PARP-3 and APLF function together to accelerate nonhomologous end-joining. Mol Cell 41:33–45. doi:10.1016/j.molcel.2010.12.006
Augustin A, Spenlehauer C, Dumond H, Ménissier-De Murcia J, Piel M, Schmit A-C, Apiou F, Vonesch J-L, Kock M, Bornens M, De Murcia G (2003) PARP-3 localizes preferentially to the daughter centriole and interferes with the G1/S cell cycle progression. J Cell Sci 116:1551–1562. doi:10.1242/jcs.00341
Boehler C, Gauthier LR, Mortusewicz O, Biard DS, Saliou J-M, Bresson A, Sanglier-Cianferani S, Smith S, Schreiber V, Boussin F, Dantzer F (2011) Poly(ADP-ribose) polymerase 3 (PARP3), a newcomer in cellular response to DNA damage and mitotic progression. Proc Natl Acad Sci U S A 108:2783–2788. doi:10.1073/pnas.1016574108
Bieche I, Pennaneach V, Driouch K, Vacher S, Zaremba T, Susini A, Lidereau R, Hall J (2013) Variations in the mRNA expression of poly(ADP-ribose) polymerases, poly(ADP-ribose) glycohydrolase and ADP-ribosylhydrolase 3 in breast tumors and impact on clinical outcome. Int J Cancer 133:2791–2800. doi:10.1002/ijc.28304
Ben-Hur E, Utsumi H, Elkind M (1984) Inhibitors of poly (ADP-ribose) synthesis enhance X-ray killing of log-phase Chinese hamster cells. Radiat Res 555:546–555
Wahlberg E, Karlberg T, Kouznetsova E, Markova N, Macchiarulo A, Thorsell A-G, Pol E, Frostell Å, Ekblad T, Öncü D, Kull B, Robertson GM, Pellicciari R, Schüler H, Weigelt J (2012) Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors. Nat Biotechnol 30:283–288. doi:10.1038/nbt.2121
Satoh MS, Lindahl T (1992) Role of poly(ADP-ribose) formation in DNA repair. Nature 356:356–358. doi:10.1038/356356a0
Mortusewicz O, Amé J-C, Schreiber V, Leonhardt H (2007) Feedback-regulated poly(ADP-ribosyl)ation by PARP-1 is required for rapid response to DNA damage in living cells. Nucleic Acids Res 35:7665–7675. doi:10.1093/nar/gkm933
Okano S, Lan L, Caldecott KW, Mori T, Yasui A (2003) Spatial and temporal cellular responses to single-strand breaks in human cells. Mol Cell Biol 23:3974–3981. doi:10.1128/MCB.23.11.3974-3981.2003
Lan L, Nakajima S, Oohata Y, Takao M, Okano S, Masutani M, Wilson SH, Yasui A (2004) In situ analysis of repair processes for oxidative DNA damage in mammalian cells. Proc Natl Acad Sci U S A 101:13738–13743. doi:10.1073/pnas.0406048101
Noël G, Godon C, Fernet M, Giocanti N, Mégnin-Chanet F, Favaudon V (2006) Radiosensitization by the poly(ADP-ribose) polymerase inhibitor 4-amino-1,8-naphthalimide is specific of the S phase of the cell cycle and involves arrest of DNA synthesis. Mol Cancer Ther 5:564–574. doi:10.1158/1535-7163.MCT-05-0418
Dungey FA, Löser DA, Chalmers AJ (2008) Replication-dependent radiosensitization of human glioma cells by inhibition of poly(ADP-Ribose) polymerase: mechanisms and therapeutic potential. Int J Radiat Oncol Biol Phys 72:1188–1197. doi:10.1016/j.ijrobp.2008.07.031
Saleh-Gohari N, Bryant HE, Schultz N, Parker KM, Cassel TN, Helleday T (2005) Spontaneous homologous recombination is induced by collapsed replication forks that are caused by endogenous DNA single-strand breaks. Mol Cell Biol 25:7158–7169. doi:10.1128/MCB.25.16.7158-7169.2005
Bryant HE, Schultz N, Thomas HD, Parker KM, Flower D, Lopez E, Kyle S, Meuth M, Curtin NJ, Helleday T (2005) Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434:913–917. doi:10.1038/nature03443
Farmer H, McCabe N, Lord CJ, Tutt ANJ, Johnson DA, Richardson TB, Santarosa M, Dillon KJ, Hickson I, Knights C, Martin NMB, Jackson SP, Smith GCM, Ashworth A (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434:917–921. doi:10.1038/nature03445
Murai J, Huang SN, Renaud A, Zhang Y, Ji J, Takeda S, Morris J, Teicher B, Doroshow JH, Pommier Y (2014) Stereospecific PARP trapping by BMN 673 and comparison with olaparib and rucaparib. Mol Cancer Ther 13:433–443. doi:10.1158/1535-7163.MCT-13-0803
Murai J, Huang SN, Das BB, Renaud A, Zhang Y, Doroshow JH, Ji J, Takeda S, Pommier Y (2012) Trapping of PARP1 and PARP2 by Clinical PARP Inhibitors. Cancer Res 72:5588–5599. doi:10.1158/0008-5472.CAN-12-2753
Heacock ML, Stefanick DF, Horton JK, Wilson SH (2010) Alkylation DNA damage in combination with PARP inhibition results in formation of S-phase-dependent double-strand breaks. DNA Repair (Amst) 9:929–936. doi:10.1016/j.dnarep.2010.05.007
Horton JK, Stefanick DF, Naron JM, Kedar PS, Wilson SH (2005) Poly(ADP-ribose) polymerase activity prevents signaling pathways for cell cycle arrest after DNA methylating agent exposure. J Biol Chem 280:15773–15785. doi:10.1074/jbc.M413841200
Kedar PS, Stefanick DF, Horton JK, Wilson SH (2012) Increased PARP-1 association with DNA in alkylation damaged, PARP-inhibited mouse fibroblasts. Mol Cancer Res 10:360–368. doi:10.1158/1541-7786.MCR-11-0477
Helleday T (2011) The underlying mechanism for the PARP and BRCA synthetic lethality: clearing up the misunderstandings. Mol Oncol 5:387–393. doi:10.1016/j.molonc.2011.07.001
Wang L, Mason KA, Ang KK, Buchholz T, Valdecanas D, Mathur A, Buser-Doepner C, Toniatti C, Milas L (2012) MK-4827, a PARP-1/-2 inhibitor, strongly enhances response of human lung and breast cancer xenografts to radiation. Invest New Drugs 30:2113–2120. doi:10.1007/s10637-011-9770-x
Ali M, Kamjoo M, Thomas HD, Kyle S, Pavlovska I, Babur M, Telfer BA, Curtin NJ, Williams KJ (2011) The clinically active PARP inhibitor AG014699 ameliorates cardiotoxicity but does not enhance the efficacy of doxorubicin, despite improving tumor perfusion and radiation response in mice. Mol Cancer Ther 10:2320–2329. doi:10.1158/1535-7163.MCT-11-0356
Donawho CK, Luo Y, Penning TD, Bauch JL, Bouska JJ, Bontcheva-Diaz VD, Cox BF, DeWeese TL, Dillehay LE, Ferguson DC, Ghoreishi-Haack NS, Grimm DR, Guan R, Han EK, Holley-Shanks RR, Hristov B, Idler KB, Jarvis K, Johnson EF, Kleinberg LR, Klinghofer V, Lasko LM, Liu X, Marsh KC, McGonigal TP, Meulbroek JA, Olson AM, Palma JP, Rodriguez LE, Shi Y, Stavropoulos JA, Tsurutani AC, Zhu G, Rosenberg SH, Giranda VL, Frost DJ (2007) ABT-888, an orally active poly(ADP-ribose) polymerase inhibitor that potentiates DNA-damaging agents in preclinical tumor models. Clin Cancer Res 13:2728–2737. doi:10.1158/1078-0432.CCR-06-3039
Calabrese CR, Almassy R, Barton S, Batey MA, Calvert AH, Canan-Koch S, Durkacz BW, Hostomsky Z, Kumpf RA, Kyle S, Li J, Maegley K, Newell DR, Notarianni E, Stratford IJ, Skalitzky D, Thomas HD, Wang L-Z, Webber SE, Williams KJ, Curtin NJ (2004) Anticancer Chemosensitization and Radiosensitization by the Novel Poly(ADP-ribose) Polymerase-1 Inhibitor AG14361. JNCI J Natl Cancer Inst 96:56–67. doi:10.1093/jnci/djh005
Shelton JW, Waxweiler TV, Landry J, Gao H, Xu Y, Wang L, El-Rayes B, Shu H-KG (2013) In vitro and in vivo enhancement of chemoradiation using the oral PARP inhibitor ABT-888 in colorectal cancer cells. Int J Radiat Oncol Biol Phys 86:469–476. doi:10.1016/j.ijrobp.2013.02.015
Albert JM, Cao C, Kim KW, Willey CD, Geng L, Xiao D, Wang H, Sandler A, Johnson DH, Colevas AD, Low J, Rothenberg ML, Lu B (2007) Inhibition of poly(ADP-ribose) polymerase enhances cell death and improves tumor growth delay in irradiated lung cancer models. Clin Cancer Res 13:3033–3042. doi:10.1158/1078-0432.CCR-06-2872
Senra JM, Telfer BA, Cherry KE, McCrudden CM, Hirst DG, O’Connor MJ, Wedge SR, Stratford IJ (2011) Inhibition of PARP-1 by olaparib (AZD2281) increases the radiosensitivity of a lung tumor xenograft. Mol Cancer Ther 10:1949–1958. doi:10.1158/1535-7163.MCT-11-0278
Chow JPH, Man WY, Mao M, Chen H, Cheung F, Nicholls J, Tsao SW, Li Lung M, Poon RYC (2013) PARP1 is overexpressed in nasopharyngeal carcinoma and its inhibition enhances radiotherapy. Mol Cancer Ther 12:251725–251728. doi:10.1158/1535-7163.MCT-13-0010
Khan K, Araki K, Wang D, Li G, Li X, Zhang J, Xu W, Hoover RK, Lauter S, O’Malley B, Lapidus RG, Li D (2010) Head and neck cancer radiosensitization by the novel poly(ADP-ribose) polymerase inhibitor GPI-15427. Head Neck 32:381–391. doi:10.1002/hed.21195
Clarke MJ, Mulligan EA, Grogan PT, Mladek AC, Carlson BL, Schroeder MA, Curtin NJ, Lou Z, Decker PA, Wu W, Plummer ER, Sarkaria JN (2009) Effective sensitization of temozolomide by ABT-888 is lost with development of temozolomide resistance in glioblastoma xenograft lines. Mol Cancer Ther 8:407–414. doi:10.1158/1535-7163.MCT-08-0854
Russo AL, Kwon H-C, Burgan WE, Carter D, Beam K, Weizheng X, Zhang J, Slusher BS, Chakravarti A, Tofilon PJ, Camphausen K (2009) In vitro and in vivo radiosensitization of glioblastoma cells by the poly (ADP-ribose) polymerase inhibitor E7016. Clin Cancer Res 15:607–612. doi:10.1158/1078-0432.CCR-08-2079
Venere M, Hamerlik P, Wu Q, Rasmussen RD, Song LA, Vasanji A, Tenley N, Flavahan WA, Hjelmeland AB, Bartek J, Rich JN (2014) Therapeutic targeting of constitutive PARP activation compromises stem cell phenotype and survival of glioblastoma-initiating cells. Cell Death Differ 21:258–269. doi:10.1038/cdd.2013.136
Mueller S, Bhargava S, Molinaro AM, Yang X, Kolkowitz I, Olow A, Wehmeijer N, Orbach S, Chen J, Matthay KK, Haas-Kogan DA (2013) Poly (ADP-Ribose) polymerase inhibitor MK-4827 together with radiation as a novel therapy for metastatic neuroblastoma. Anticancer Res 33:755–762
Horsman MR (1995) Nicotinamide and other benzamide analogs as agents for overcoming hypoxic cell radiation resistance in tumours. A review. Acta Oncol 34:571–587
Liu SK, Coackley C, Krause M, Jalali F, Chan N, Bristow RG (2008) A novel poly(ADP-ribose) polymerase inhibitor, ABT-888, radiosensitizes malignant human cell lines under hypoxia. Radiother Oncol 88:258–268. doi:10.1016/j.radonc.2008.04.005
Baker DG, Krochak RJ (1989) The response of the microvascular system to radiation: a review. Cancer Invest 7:287–294
Dubner D, Gisone P, Jaitovich I, Perez M (1995) Free radicals production and estimation of oxidative stress related to gamma irradiation. Biol Trace Elem Res 47:265–270. doi:10.1007/BF02790126
Jagtap P, Szabó C (2005) Poly(ADP-ribose) polymerase and the therapeutic effects of its inhibitors. Nat Rev Drug Discov 4:421–440. doi:10.1038/nrd1718
Clempus RE, Griendling KK (2006) Reactive oxygen species signaling in vascular smooth muscle cells. Cardiovasc Res 71:216–225. doi:10.1016/j.cardiores.2006.02.033
Affar EB, Germain M, Poirier GG (2000) Role of poly(ADP-ribose) polymerase in cell death. In: de Murcia G, Shall S (eds) From DNA damage stress signal. To cell death. Oxford University Press, New York, pp 125–150
Szabo C (2000) Activation of poly(ADP-ribose) polymerase in the pathogenesis of ischaemia-reperfusion injury. In: de Murcia G, Shall S (eds) From DNA damage stress signal. To cell death. Poly ADP-ribosylation react. Oxford University Press, New York, pp 151–176
Rajesh M, Mukhopadhyay P, Bátkai S, Godlewski G, Haskó G, Liaudet L, Pacher P (2006) Pharmacological inhibition of poly(ADP-ribose) polymerase inhibits angiogenesis. Biochem Biophys Res Commun 350:352–357. doi:10.1016/j.bbrc.2006.09.049
Rajesh M, Mukhopadhyay P, Godlewski G, Bátkai S, Haskó G, Liaudet L, Pacher P (2006) Poly(ADP-ribose)polymerase inhibition decreases angiogenesis. Biochem Biophys Res Commun 350:1056–1062. doi:10.1016/j.bbrc.2006.09.160
Beller CJ, Radovits T, Seres L, Kosse J, Krempien R, Gross M-L, Penzel R, Berger I, Huber PE, Hagl S, Szabó C, Szabó G (2006) Poly(ADP-ribose) polymerase inhibition reverses vascular dysfunction after gamma-irradiation. Int J Radiat Oncol Biol Phys 65:1528–1535. doi:10.1016/j.ijrobp.2006.03.058
Beller CJ, Kosse J, Radovits T, Gerö D, Krempien R, Gross M-L, Berger I, Hagl S, Szabó C, Szabó G (2006) Poly(ADP-ribose) polymerase inhibition combined with irradiation: a dual treatment concept to prevent neointimal hyperplasia after endarterectomy. Int J Radiat Oncol Biol Phys 66:867–875. doi:10.1016/j.ijrobp.2006.06.055
Radovits T, Lin L, Zotkina J, Gero D, Szabó C, Karck M, Szabó G (2007) Poly(ADP-ribose) polymerase inhibition improves endothelial dysfunction induced by reactive oxidant hydrogen peroxide in vitro. Eur J Pharmacol 564:158–166. doi:10.1016/j.ejphar.2007.02.060
Yarnold J, Brotons M-CV (2010) Pathogenetic mechanisms in radiation fibrosis. Radiother Oncol 97:149–161. doi:10.1016/j.radonc.2010.09.002
Milliat F, François A, Isoir M, Deutsch E, Tamarat R, Tarlet G, Atfi A, Validire P, Bourhis J, Sabourin J-C, Benderitter M (2006) Influence of endothelial cells on vascular smooth muscle cells phenotype after irradiation: implication in radiation-induced vascular damages. Am J Pathol 169:1484–1495. doi:10.2353/ajpath.2006.060116
Finnson KW, Parker WL, ten Dijke P, Thorikay M, Philip A (2008) ALK1 opposes ALK5/Smad3 signaling and expression of extracellular matrix components in human chondrocytes. J Bone Miner Res 23:896–906. doi:10.1359/jbmr.080209
Cordenonsi M, Dupont S, Maretto S, Insinga A, Imbriano C, Piccolo S (2003) Links between tumor suppressors: p53 is required for TGF-beta gene responses by cooperating with Smads. Cell 113:301–314
Hageman J, Eggen BJ, Rozema T, Damman K, Kampinga HH, Coppes RP (2005) Radiation and transforming growth factor-beta cooperate in transcriptional activation of the profibrotic plasminogen activator inhibitor-1 gene. Clin Cancer Res 11:5956–5964. doi:10.1158/1078-0432.CCR-05-0427
Niemantsverdriet M, de Jong E, Langendijk JA, Kampinga HH, Coppes RP (2010) Synergistic induction of profibrotic PAI-1 by TGF-β and radiation depends on p53. Radiother Oncol 97:33–35. doi:10.1016/j.radonc.2010.04.002
Scharpfenecker M, Kruse JJCM, Sprong D, Russell NS, Ten Dijke P, Stewart FA (2009) Ionizing radiation shifts the PAI-1/ID-1 balance and activates notch signaling in endothelial cells. Int J Radiat Oncol Biol Phys 73:506–513. doi:10.1016/j.ijrobp.2008.09.052
Wang Y, Wang L, Zhang F, Zhang C, Deng S, Wang R, Zhang Y, Huang D, Huang K (2013) Inhibition of PARP prevents angiotensin II-induced aortic fibrosis in rats. Int J Cardiol 167:2285–2293. doi:10.1016/j.ijcard.2012.06.050
Lei P, Jiang Z, Zhu H, Li X, Su N, Yu X (2012) Poly(ADP-ribose) polymerase-1 in high glucose-induced epithelial-mesenchymal transition during peritoneal fibrosis. Int J Mol Med 29:472–478. doi:10.3892/ijmm.2011.859
Palfi A, Toth A, Hanto K, Deres P, Szabados E, Szereday Z, Kulcsar G, Kalai T, Hideg K, Gallyas F, Sumegi B, Toth K, Halmosi R (2006) PARP inhibition prevents postinfarction myocardial remodeling and heart failure via the protein kinase C/glycogen synthase kinase-3beta pathway. J Mol Cell Cardiol 41:149–159. doi:10.1016/j.yjmcc.2006.03.427
Pacher P, Liaudet L, Mabley JG, Cziráki A, Haskó G, Szabó C (2006) Beneficial effects of a novel ultrapotent poly(ADP-ribose) polymerase inhibitor in murine models of heart failure. Int J Mol Med 17:369–375
Chiu J, Farhangkhoee H, Xu BY, Chen S, George B, Chakrabarti S (2008) PARP mediates structural alterations in diabetic cardiomyopathy. J Mol Cell Cardiol 45:385–393. doi:10.1016/j.yjmcc.2008.06.009
Bartha E, Kiss GN, Kalman E, Kulcsár G, Kálai T, Hideg K, Habon T, Sumegi B, Toth K, Halmosi R (2008) Effect of L-2286, a poly(ADP-ribose)polymerase inhibitor and enalapril on myocardial remodeling and heart failure. J Cardiovasc Pharmacol 52:253–261. doi:10.1097/FJC.0b013e3181855cef
Bartha E, Solti I, Kereskai L, Lantos J, Plozer E, Magyar K, Szabados E, Kálai T, Hideg K, Halmosi R, Sumegi B, Toth K (2009) PARP inhibition delays transition of hypertensive cardiopathy to heart failure in spontaneously hypertensive rats. Cardiovasc Res 83:501–510. doi:10.1093/cvr/cvp144
Pagano A, Métrailler-Ruchonnet I, Aurrand-Lions M, Lucattelli M, Donati Y, Argiroffo CB (2007) Poly(ADP-ribose) polymerase-1 (PARP-1) controls lung cell proliferation and repair after hyperoxia-induced lung damage. Am J Physiol Lung Cell Mol Physiol 293:L619–L629. doi:10.1152/ajplung.00037.2007
Anjos SM, Robert R, Waller D, Zhang DL, Balghi H, Sampson HM, Ciciriello F, Lesimple P, Carlile GW, Goepp J, Liao J, Ferraro P, Phillipe R, Dantzer F, Hanrahan JW, Thomas DY (2012) Decreasing poly(ADP-Ribose) polymerase activity restores ΔF508 CFTR trafficking. Front Pharmacol 3:165. doi:10.3389/fphar.2012.00165
Favaudon V, Caplier L, Monceau V, Pouzoulet F, Sayarath M, Fouillade C, Poupon MF, Brito I, Hupé P, Bourhis J, Hall J, Fontaine JJ, Vozenin MC (2014) Ultrahigh dose-rate FLASH irradiation increases the differential response between normal and tumor tissue in mice. Sci Transl Med 6:245ra93. doi:10.1126/scitranslmed.3008973
Zackrisson BU, Nyström UH, Ostbergh P (1991) Biological response in vitro to pulsed high dose rate electrons from a clinical accelerator. Acta Oncol 30:747–751
Ponette V, Le Péchoux C, Deniaud-Alexandre E, Fernet M, Giocanti N, Tourbez H, Favaudon V (2000) Hyperfast, early cell response to ionizing radiation. Int J Radiat Biol 76:1233–1243
Fernet M, Ponette V, Deniaud-Alexandre E, Ménissier-De Murcia J, De Murcia G, Giocanti N, Megnin-Chanet F, Favaudon V (2000) Poly(ADP-ribose) polymerase, a major determinant of early cell response to ionizing radiation. Int J Radiat Biol 76:1621–1629
Acknowledgements
Research in Inserm U612 is supported by Inserm and Institut Curie and is part of the Comprehensive Cancer Centre “SIRIC” program (INCa 2011-189). CF is supported by INCa (PL-BIO 2013-11) and AF by a PhD fellowship from the French Ministry of Research. MTB was supported by Institute Curie’s International Postdoctoral fellow program.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Fouillade, C., Fouquin, A., Boudra, MT., Favaudon, V., Pennaneach, V., Hall, J. (2015). Radiosensitisation by Poly(ADP-ribose) Polymerase Inhibition. In: Curtin, N., Sharma, R. (eds) PARP Inhibitors for Cancer Therapy. Cancer Drug Discovery and Development, vol 83. Humana Press, Cham. https://doi.org/10.1007/978-3-319-14151-0_11
Download citation
DOI: https://doi.org/10.1007/978-3-319-14151-0_11
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-14150-3
Online ISBN: 978-3-319-14151-0
eBook Packages: MedicineMedicine (R0)