Abstract
Multi-phase contrast-enhanced CT images can provide abundant and complementary tumor information, and thus radiologists often use multi-phase images to assist in segmenting and diagnosing liver tumors. However, the current multi-stage liver tumor segmentation methods are based on convolutional neural networks (CNNs), which make them ineffective in extracting global information during the multi-phase information fusion process. In this study, we propose a novel multi-phase liver tumor segmentation approach using delayed phase images to aid in portal vein phase tumor segmentation. The proposed method employs a Transformer structure to extract both global information and local information of tumors, which contributes to the precise segmentation of tumor boundaries. More importantly, we design a cross-phase aggregator (CFA), which facilitates the bidirectional interaction of cross-phase features to take full advantage of the complementary information from multi-phase images. A dataset of 164 multi-phase abdominal CT scans was collected with Institutional Review Board approval to evaluate the performance of the proposed approach. The experimental results showed that the proposed approach can better utilize multi-phase information and is superior to several state-of-the-art methods. Ablation study is performed to further validate the effectiveness of each module in the proposed model. The proposed method has the potential to assist radiologists to locate more accurate liver tumors and improve their diagnosis efficiency.
Xiangyuan Ma contributed equally to this work.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Liver cancer is a severe and fatal form of cancer that affects millions of individuals across the globe [1]. As per the World Cancer Research Fund International's data from 2020, liver cancer ranks as the sixth most common cancer and third leading cause of cancer-related deaths globally [2]. Early detection of liver lesions is very crucial for effective liver cancer treatment. Traditionally, radiologists have detected and diagnosed tumors by manually segmenting liver tumors, which is time-consuming, difficult to produce, and susceptible to subjective personal experience. Therefore, it is essential to develop an automatic and accurate method for liver tumor segmentation.
Single-phase computed tomography (CT) images have been used to segment liver tumors in most existing liver tumor segmentation (LiTS) methods [3,4,5,6,7,8]. However, these methods often fail to achieve satisfactory results due to inherent problems such as low contrast of medical images and blurred tumor boundaries. Therefore, some multi-phase-based segmentation methods for liver tumors have been proposed [9,10,11]. Compared to single images, multi-phase images can provide rich and complementary information of tumors, in which tumor areas are different from other tissues in morphology or grayscale. The multi-phase contrast-enhanced CT images are acquired in four phases: the noncontract-enhanced (NC) phase, arterial (ART) phase, portal vein (PV) phase and delay (DL) phase. Among the four phases, the PV phase is the preferred stage for liver tumor segmentation because the maximum contrast between the tumor and the surrounding liver tissue occurs in the PV phase. However, not all types of tumors are evident in the PV stage, hence we also need to employ other phases to assist in the segmentation of liver tumors [10].
The current multi-phase liver tumor segmentation methods can be categorized in accordance with multi-phase fusion strategies: input-level fusion (ILF) methods [12], decision-level fusion (DLF) methods [12,13,14] and feature-level fusion (FLF) methods [8, 10, 11]. Under these three fusion strategies, the FLF methods can achieve the best performance because they exploited the features from multiple phases. Wu et al. [9] proposed an MW-UNet, which integrated different phases by weighting their features from hidden layers of U-Net [6]. Xu et al. [10] developed a PA-ResSeg model based on residual networks [15] that re-weights features from different phases using the channel-attention mechanism. Zhang et al. [11] modified the PA-ResSeg with a spatial aggregation module (SAM) and a uncertain region inpainting module (URIM) to further improve the performance of liver tumor segmentation. Although known FLF methods focus on inter-phase relationships, it is still difficult to fully utilize the key global cues between different phases due to the inherent limitations of CNNs in extracting features in local sliding windows.
In recent years, Transformer has shown excellent performance not only in natural language processing (NLP) tasks [16], but also in semantic segmentation [17,18,19]. Because the Transformer based methods are better at leveraging global information and modeling long-term dependencies than the CNN approach, some studies have begun to apply the Transformer approach to medical image analyzing tasks [18, 20]. These methods utilize the self-attention mechanism of transformers to selectively focus on important regions of the image, allowing for better global feature extraction and improved segmentation performance. Therefore, Transformer based architecture has the potential to better utilizes information from different phases.
In this study, we propose a framework for accurate liver tumors segmentation from multiphase contrast-enhanced CT images. Specifically, our method takes advantage of the complementary information from delayed phase (DL) images to assist in segmenting liver tumors in portal venous (PV) phase images. We introduce the Transformer architecture into the traditional encoder-decoder structure to facilitate the model to fully learn the explicit global information. More importantly, we design a cross-phase aggregator to better exploit the features from PV and DL phases for comprehensively multi-phase information fusion. Specifically, our contributions can be summarized as follows: (1) Based on the tokens-to-tokens vision Transformer (T2T-ViT) architecture, we build a symmetric Transformer Encoder-Decoder architecture with skip-connection. In the encoder, the local and global information is extracted by self-attention, and in the decoder, the local and global features are up-sampled to the input resolution for corresponding pixel-level liver tumor prediction. (2) The designed cross-phase aggregator that can efficiently utilize the complementary information from DL phase images to help the tumor segmentation in PV phase images.
2 Method
In this paper, we propose a multi-phase liver tumor segmentation framework consisting of the encoder, cross-phase feature aggregator (CFA), decoder, and skip connection as Fig. 1. Tokens-to-tokens (T2T) strategy is used in the framework. Given input multi-phase (PV and DL phases) images and their multi-phase features are extracted utilizing the encoder. The multi-phase feature tokens generated in the encoder are interacted in the cross-phase feature aggregator, which also outputs feature tokens for the decoder. In the decoder, the Reverse T2T process gradually recovers the resolution of the features. In order to improve the capacity of global cue utilization and reduce tumor boundary ambiguity, we apply self-attention [16] in both the encoder and decoder. Furthermore, a task-related token and a skip connection mechanism assist in the recovery of finer segmentation results.
2.1 Encoder
Comparing the wide-applying convolution encoder module, we adopt the T2T-ViT model [17,18,19] as the backbone, and convolutional windowing similar to that in CNN is used to locally aggregate adjacent tokens, which helps model local features.
Giving the inputs of the PV phase and DL phase image, the encoder first unfolds these images into tokens by setting \(k=7, s=3\) and \(p=3\). Then, we perform the T2T Transformer and T2T Process twice to obtain the two phases’ feature tokens separately. The dimensions of the feature token in the three stages of the encoder can be expressed as \({{\text{T}}}_{i}^{{\text{PV}}}\in {\mathbb{R}}^{{l}_{i}\times c}\) and \({{\text{T}}}_{i}^{{\text{DL}}}\in {\mathbb{R}}^{{l}_{i}\times c}\), where PV and DL denote PV phase and DL phase, separately, \(i=1, 2, 3,\) and \(c=64.\)
T2T Transformer. For a particular input token from the preceding step, a linear projection is used to project the embedding dimension from c = 64 to d = 384. Then, it is processed by the self-attention block, as formulated:
where MSA represents the multi-head-attention mechanism with layer normalization and MLP denotes multi-layer perceptron with layer normalization in the standard transformer.
T2T Process. As shown in Fig. 1, after obtaining the feature tokens from the T2T Transformer, we reshaped the feature tokens as an image \(I\) in the spatial dimension. Then, \(I\) is split into a \(k\times k\) patch that uses the overlapping of \(s\) and the zero padding of \(p\) to fill the image boundary. finally, the patch is unfolded into tokens sequence and the length of the output token is defined as the below formula:
During the two times of the T2T Process, the patch size is set as \(k=\left[3, 3\right]\), the overlapping is set as \(s=\left[\mathrm{1,1}\right]\) and the padding size is set as \(P=\left[\mathrm{1,1}\right]\).
2.2 Cross-Phase Feature Aggregator
The Cross-phase feature aggregator (CFA) is introduced to better fuse the feature tokens which are output by the encoder session from PV and DL. The CFA module contains three parts, first is the linear projection, the second is the cross-phase attention, and the third part is the concatenation of the cross-phase features. The encoder output initially passes via linear projection to project the dimension to d = 384. A modified cross-phase self-attention is used to help multi-phase feature interactions and the features obtained after the interactions are aggregated, formulated as:
where Concat represents concatenation (superimposed on channel dimension) and MLP is the multi-layer Perceptron that is used to recover the dimensional size of the superimposed C-channel.
2.3 Decoder
The decoder in the proposed framework is designed to decode the patch token into a significant graph. Considering that directly adopting the low-resolution feature to perform segmentation prediction cannot achieve a satisfactory result, we introduce the decoder that consists of three main components, the first is RT2T Process that gradually up-sampled the main feature tokens, and the second is Task-related Transformer Module which is aimed to perform patch-task-attention between patch token and task-related token for better improve the segmentation performance. The last one is the Prediction that utilizes the high-resolution main feature tokens to perform liver tumor prediction.
RT2T Process. Given the input tokens, we adopt a linear projection to extend the embedding dimension from c to \({ck}^{2}\). Then, each token is treated as a k × k image patch with \(s\) overlapping and the tokens are folded as an image with P zero-padding. finally, the generated image is reshaped back to the unsampled tokens.
Task-Related Transformer Module. Inspired by T2T-ViT [17,18,19], which introduces a class token to improve the classification accuracy, we also introduce a task-related token in our proposed model to help us improve the accuracy of liver tumor segmentation.
We first concatenate a low-level PV feature from the encoder with the multi-phase feature token from the cross-phase feature aggregator to generate the main feature tokens. Then, we introduce a task-related token \({T}_{ts}\), and concatenate it with the main feature tokens \({T}_{i}^{M}\). The concatenated tokens will be processed by Transformer module. The processed tokens are divided into task-related tokens and main-phase features.
For the overall processes of the decoder, we adopt the RT2T Process and Task-related Transformer Module twice to obtain the main feature tokens at the \(\frac{1}{4}\) scale, and set \(k\) = [1, 1], \(s=[\mathrm{1,1}]\), and \(p =[\mathrm{1,1}]\) in the two RT2T Processes. To recover the major feature tokens to full resolution, we project the embedding dimension of main feature tokens from \(c\) to \(c{k}^{2}\) (\(k\) = 7). Then we fold the tokens into an image, setting \(k=7, s=3,\) and \(p = 3.\) Finally, we use a linear projection to predict the final segmentation result.
3 Experiment and Discussion
3.1 Dataset
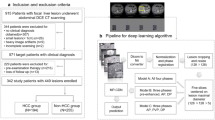
With Institutional Review Board approval (IRB) approval, a clinical multi-phase contrast-enhanced CT liver tumor dataset was collected in this study. The dataset includes 164 multi-phase abdominal CT scans. For the source hospital of the dataset, Sun Yat-Sen University’s Third Affiliated Hospital contributes to 108 cases of multiphase abdominal CT scans, and Sun Yat-Sen University’s Fifth Affiliated Hospital contributes to 56 cases of multiphase abdominal CT scans.
The typology of CT images we captured included the non-contrast phase, arterial phase, portal venous phase, and delay phase. Below Table 1 shows the details of the characteristic of CT scans.
3.2 Preprocessing
The dataset is separated into 80 and 20% for training and evaluation.
For data pre-processing, first, all the input images are set as 224 × 224. Second, we set the limit of the intensity value between −80 and 220 HU. Third, normalization is performed on these scans to eliminate interference and solve abnormal data scale distribution.
3.3 Experiment Configuration
For the hyperparameters and training strategies for learning, the setting of the initial learning rate is 0.0001, using the Adam gradient descent with momentum for the optimizer in the model and the binary cross entropy loss is used. For the strategy in the learning rate, occurring a reduction in the learning rate up to 10 times when the loss of training stops falling and continues for 10 times.
3.4 Evaluation Metrics
We introduce three metrics for evaluation, which are Dice similarity coefficient (DSC), absolute relative volume difference (ARVD), and average symmetric surface distance (ASSD). First, the accuracy of segmentation is evaluated by the DSC and ARVD. Second, ASSD is capable to evaluate the segmentation distance error.
A great performance method is proven by high values of DCS and sensitivity with low values of ARVD and ASSD.
3.5 Comparison Study
We provide qualitative comparison results on the dataset, as shown in Fig. 2. We compare the ground truth with segmentation results of three methods—The original U-Net applies the PV phase images as input; PA-ResSeg which fuses the DL and PV phases as input; and our proposed method which fuses the DL and PV phases as input. In this example, our network achieves the best performance compared with the multi-phase segmentation method and the single-phase segmentation method, which demonstrates the validity of utilizing delay and portal vein images for segmentation.
U-Net and PA-ResSeg are both based on the traditional CNN architecture. The performance of PA-ResSeg is great but still exists some puzzle. The loss of high-frequency information in Up and Down-sampling operations, and low contrast between the tumor and surroundings result in blurred boundaries. In the proposed method, the structure of the Transformer has a wider perceptual area than the convolutional neural network, allowing for a more complete segmentation of the entire tumor region.
Table 2 shows the results of three evaluation metrics for different methods. Our proposed method achieved the best performance under the three metrics compared to the state-of-the-art CNN-based models, hence demonstrating the great effectiveness of our network. Both the U-Net and the PA-ResSeg prediction exist some blurred boundary issues. Instead, as the transformer has strong global properties and modal fusion capabilities, it allows the proposed method to better understand the information about the tumor, leading to the demonstration of better predictive capabilities.
3.6 Ablation Study
In the baseline testing phase, there occur two issues that should be improved. First, the baseline divides the image into multiple tokens and then uses multiple transformers to model the global picture, but this loses localization, and in fact, local information such as edges, lines, and textures are essential for the network to comprehend the visual information. Second, the attention backbone of the baseline contains redundancy and limited feature richness. Based on this, the RT2T module was added to iteratively reduce the length of Tokens, adding locality to simulate the local structural information of surrounding Tokens. CFA is added to help multi-phase feature interactions and multi-level token fusion is added to support low-level fine-grained information.
We conduct ablation studies to verify all our proposed model components. The experimental results of all the validation and results are shown in Table 3. The advantage of our method is shown in both DSC, ARVD, and ASSD. The step-by-step addition module shows that using boundary detection brings further performance gain in the perception of global information and edge characteristics.
3.7 Discussion
The results from the comparison study and ablation study demonstrate the proposed method achieved the best performance on all three-evaluation metrics for segmenting tumors of different sizes. The results indicate that our new method has good stability and thus possesses the ability for clinical application.
There are several reasons why our method outperformed other approaches. First, our backbone of T2T-ViT enables the network to extract local tumor details, which facilitates accurate segmentation of small regions. Second, our token module addresses the issue in ViT [21] that modeling local regions loses the line as well as edge information of the image and uses the CFA module to fuse multimodal tumor features and leads to more precise segmented tumor boundaries, particularly for small tumors.
Despite these strengths, our method still has some drawbacks. For instance, in the multiphase fusion stage, our multiphase data need to be strictly aligned, but the tumor location of different phases in the same location may be shifted due to the possible movement of the patient during the shooting. Future work will focus on finding algorithms that can achieve liver position alignment to obtain a finer tumor profile.
4 Conclusions
The transformer architecture is well-known for its robust self-attention mechanisms and has been widely adopted in various fields of deep learning in recent years. In our study, we design a symmetric Transformer Encoder-Decoder architecture with skip-connection to extract the multi-phase features and predict the liver tumor. More importantly, we introduce a cross-phase aggregator, which can assist the interaction of the PV phase feature and the DL phase feature. Extensive experiments on a clinical multi-phase contrast-enhanced CT liver tumor dataset have demonstrated that the proposed method outperforms state-of-the-art methods and highlights the promising potential of Transformers for liver tumor segmentation tasks.
References
McGlynn, K.A., Petrick, J.L., El-Serag, H.B.: Epidemiology of hepatocellular carcinoma. Hepatology 73, 4–13 (2021)
Sung, H., Ferlay, J., Siegel, R.L., et al.: Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a Cancer J. Clin. 71, 209–249 (2021)
Han, X.: Automatic liver lesion segmentation using a deep convolutional neural network method (2017). arXiv:170407239
Bi, L., Kim, J., Kumar, A., et al.: Automatic liver lesion detection using cascaded deep residual networks (2017). arXiv:170402703
Bellver, M., Maninis, K.-K., Pont-Tuset, J., et al.: Detection-aided liver lesion segmentation using deep learning (2017). arXiv:171111069
Ronneberger, O., Fischer, P., Brox, T.: U-net: convolutional networks for biomedical image segmentation. Springer; pp 234–24 (2015)
Seo, H., Huang, C., Bassenne, M., et al.: Modified U-Net (mU-Net) with incorporation of object-dependent high level features for improved liver and liver-tumor segmentation in CT images. IEEE Trans. Med. Imaging 39, 1316–1325 (2019)
Huang H, Lin L, Tong R et al (2020) Unet 3+: A full-scale connected unet for medical image segmentation, pp. 1055–1059. IEEE
Wu Y, Zhou Q, Hu H et al (2019) Hepatic lesion segmentation by combining plain and contrast-enhanced CT images with modality weighted U-Net, pp. 255–259. IEEE
Xu, Y., Cai, M., Lin, L., et al.: PA-ResSeg: a phase attention residual network for liver tumor segmentation from multiphase CT images. Med. Phys. 48, 3752–3766 (2021). https://doi.org/10.1002/mp.14922
Zhang, Y., Peng, C., Peng, L., et al.: Multi-phase liver tumor segmentation with spatial aggregation and uncertain region inpainting, pp 68–77. Springer (2021)
Ouhmich, F., Agnus, V., Noblet, V., et al.: Liver tissue segmentation in multiphase CT scans using cascaded convolutional neural networks. Int. J. Comput. Assist. Radiol. Surg. 14, 1275–1284 (2019)
Sun, C., Guo, S., Zhang, H., et al.: Automatic segmentation of liver tumors from multiphase contrast-enhanced CT images based on FCNs. Artif. Intell. Med. 83, 58–66 (2017)
Raju, A., Cheng, C.-T., Huo, Y., et al.: Co-heterogeneous and adaptive segmentation from multi-source and multi-phase CT imaging data: a study on pathological liver and lesion segmentation, pp. 448–465. Springer (2020)
He, K., Zhang, X., Ren, S., et al.: Deep residual learning for image recognition, pp. 770–778 (2016)
Vaswani, A., Shazeer, N., Parmar, N., et al.: Attention is all you need. In: Advances in Neural Information Processing Systems, vol. 30 (2017)
Yuan, L., Chen, Y., Wang, T., et al.: Tokens-to-token vit: training vision transformers from scratch on imagenet, pp. 558–567 (2021)
Cao, H., Wang, Y., Chen, J., et al.: Swin-unet: unet-like pure transformer for medical image segmentation (2021). arXiv:210505537
Zheng, S., Lu, J., Zhao, H., et al.: Rethinking semantic segmentation from a sequence-to-sequence perspective with transformers, pp. 6881–6890 (2021)
Chen, J., Lu, Y., Yu, Q., et al.: Transunet: transformers make strong encoders for medical image segmentation (2021). arXiv:210204306
Dosovitskiy, A., Beyer, L., Kolesnikov, A., et al.: An image is worth 16 × 16 words: transformers for image recognition at scale (2020). arXiv:201011929
Acknowledgment
This work is supported by Shantou University (STU Scientific Research Foundation for Talents: NTF21004) and Zhuhai Basic and Applied Basic Research Project Foundation, under Grant ZH22017003200001PWC.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Zhang, W. et al. (2024). Automatic Segmentation of Liver Tumor from Multi-phase Contrast-Enhanced CT Images Using Cross-Phase Fusion Transformer. In: Wang, G., Yao, D., Gu, Z., Peng, Y., Tong, S., Liu, C. (eds) 12th Asian-Pacific Conference on Medical and Biological Engineering. APCMBE 2023. IFMBE Proceedings, vol 103. Springer, Cham. https://doi.org/10.1007/978-3-031-51455-5_15
Download citation
DOI: https://doi.org/10.1007/978-3-031-51455-5_15
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-51454-8
Online ISBN: 978-3-031-51455-5
eBook Packages: EngineeringEngineering (R0)