Abstract
An arduous biomedical task involves condensing evidence derived from multiple interrelated studies, given a context as input, to generate reviews or provide answers autonomously. We named this task context-aware multi-document summarization (CA-MDS). Existing state-of-the-art (SOTA) solutions require truncation of the input due to the high memory demands, resulting in the loss of meaningful content. To address this issue effectively, we propose a novel approach called Ramses, which employs a retrieve-and-rank technique for end-to-end summarization. The model acquires the ability to (i) index each document by modeling its semantic features, (ii) retrieve the most relevant ones, and (iii) generate a summary via token probability marginalization. To facilitate the evaluation, we introduce a new dataset, FAQsumC19, which includes the synthesizing of multiple supporting papers to answer questions related to Covid-19. Our experimental findings demonstrate that Ramses achieves notably superior ROUGE scores compared to state-of-the-art methodologies, including the establishment of a new SOTA for the generation of systematic literature reviews using Ms2. Quality observation through human evaluation indicates that our model produces more informative responses than previous leading approaches.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Given the paramount societal role of biomedicine and related natural language processing (NLP) tasks [11,12,13,14,15, 38], aggregating information from multiple topic-related biomedical papers to help search, synthesize, and answer questions is of great interest [7]. Real-world applications require indexing, combining, and summarizing evidence from clinical trials on a research background to produce systematic literature reviews (SLRs) or answer medical inquiries. Consequently, we define such activities as context-aware multi-document summarization (CA-MDS) due to the presence of an input context (i.e., background or question) that conditions the downstream summarization task (Fig. 1). In real life, biomedical articles usually contain several thousands of words that compose lingo and complicated expressions, making understanding them a time- and labor-consuming process even for professionals. Thus, automation support for biomedical activities is practical and beneficial in facilitating knowledge acquisition.
The overview of biomedical CA-MDS. In our experiments, we use the input contexts (i.e., the background or question) to retrieve the salient studies and aggregate them to generate the target ( Input \(\rightarrow \) Output). For example, given a set of topic-related scientific papers, the goal is to select the ones more correlated to a user’s input question to produce a single answer. (Color figure online)
CA-MDS solutions for biomedical applications should process all inputs without ignoring any details, reducing the risk of model hallucination, namely generating unfaithful outputs due to training on targets having facts unfounded by the source. Therefore, state-of-the-art (SOTA) models rely on sparse transformers [2], Fusion-in-Decoder strategies [18], and marginalization-based decoding [34]. However, such methods either (i) need high memory requirements that force input truncation for organizations operating in low-resource regimes [29,30,31, 33, 35], or (ii) lack end-to-end learning, reducing the potential of cooperating neural modules.
In this paper, we introduce Ramses,Footnote 1 a retrieve-and-rank summarization approach trained via end-to-end learning to retrieve salient biomedical documents by their semantic meaning and synthesize them given an input context. Ramses comprises a biomedical bi-encoder and a generative aggregator. The bi-encoder reads all the documents, represents their semantics via embeddings, and retrieves and scores salient documents related to an input context. Then, the aggregator is conditioned by the context along with these latent documents to decode the summary by marginalizing the token probability distribution weighted by their relevance score.
We evaluated Ramses in two biomedical CA-MDS tasks: (i) producing SLRs on the Ms2 dataset [7] and (ii) answering frequently asked questions (FAQs) about Covid-19 in our proposed dataset FAQsumC19. In detail, we collected 514 Covid-19 FAQs with high-quality abstractive answers written by experts. Then we augmented each instance with 30 supporting scientific papers containing the information needed to answer the question, producing 15,420 articles. In particular, FAQsumC19 has two essential features: (i) includes abstractive answers authored by experts, unlike other related datasets that use extractive targets [41]; (ii) is the first CA-MDS dataset for Covid-19, becoming a crucial benchmark for producing multi-document summaries to answer questions on Covid-19 with the support of updated related biomedical papers.
We perform extensive experiments, showing that Ramses achieves new SOTA performance in the Ms2 dataset and outperforms previous solutions in FAQsumC19, whose inferred answers are also rated as of more quality by human experts.
The overview of Ramses. The input is a biomedical context and multiple studies that are encoded by two different BioBert models. Then, we compute the relevance score of each document conditioned by the context. Finally, the top-k most salient documents are concatenated with the context and given to Bart that marginalizes their token probability distribution, weighted by the relevance scores, at decoding time.
2 Related Work
Semantic Neural Retriever Applications. The semantic representation skill exhibited by neural networks has catalyzed the emergence of groundbreaking neural methodologies in information retrieval [10] First, the algorithm Bm25 has been exceeded by dense passage retrieval (Dpr) [21], a remarkable neural application that has since evolved into a fundamental element within numerous neural-driven retrieval solutions [42, 43]. These neural retrievers have been fused with a language model to enrich and improve input [23], generating superior models characterized by increased efficiency and improved performance [3, 12]. Despite their promising results, the end-to-end application of these solutions in MDS remains unexplored.
NLP for Biomedical Documents. Much recent work in NLP has concentrated on the biomedical domain [28], including CA-MDS [7], which can decrease the burden on medical workers by highlighting and aggregating key points while reducing the amount of information to read. Previous contributions focused on the automatic generation of SLRs. In detail, cutting-edge solutions rely on three different neural architectures: (i) transformer-based models with linear complexity in the input size thanks to sparse attention [2], which concatenate the input context along with all documents in the cluster producing a single source sequence; (ii) quadratic transformers with Fusion-in-Decoder [18], which join the hidden states of documents after encoding them individually; (iii) marginalization-based decoding augmented by frozen retrievers [34], which first pinpoints salient documents w.r.t. a query and produces a single summary by summing the probability distribution of the inferred token for each document.
MDS Solutions in Other Domains. Flat approaches with MDS-specific pre-training [49] concatenate the sources in a single text, treating MDS as a single-input task. Hierarchical approaches merge document relations to obtain semantically rich representations by leveraging graph-based methods [1] and multi-head pooling and interparagraph attention [19]. Marginalization-based approaches [17] apply marginalization to the token probability distribution at the decoding time to produce a single output from many inputs. The two-stage approaches [25] adopt different strategies to rank sources before producing the summary. Unlike previous work, Ramses is trained in end-to-end learning to retrieve relevant text from biomedical articles and marginalize the probability distribution of the latent extracted information at decoding time.
Covid-19 Datasets. With the appearance of Covid-19, thousands of articles have been published quickly. To aid experts in accessing this knowledge, large organizations collected corpora such as Cord-19 [47] and LitCovid [6], encouraging the proposal of task-specific datasets. Covid-QA [27] study question-answering using annotated pairs extracted from 147 papers. Covid-Q [48] collects 16,690 questions about Covid-19, classifying them into 15 categories. [40] scrapped over 40 trusted websites for Covid-19 FAQs, creating a collection of 2100 questions. [45, 52] proposed two datasets for the retrieval of FAQs, where user queries are semantically paired with existing FAQs. FAQsumC19 fills this gap, introducing the first CA-MDS dataset to answer Covid-19 FAQs by summarizing multiple related studies.
Fine-grained comparisons with previous work are in Sect. 6.1.
3 Preliminary
We provide details for context-aware multi-document summarization (CA-MDS).
Definition. CA-MDS aims to compile a summary from a cluster of related articles given an input context, analogous to the query in query-focused summarization [46]. Yet, unlike answering FAQs, SLR generation does not consider questions. Thus, we define the task we face as CA-MDS. The biomedical tasks we address in this work, such as SLR generation and FAQ answering, are CA-MDS tasks because they both have an input context (i.e., the research issue in SLRs and the human question in FAQs) and many topic-related documents from which produce the output.
Problem Formulation. In the CA-MDS setting, we have \((c, \textbf{D}, y)\), where c is the input context, \(\textbf{D}\) is the cluster of topic-related documents, and y is the target generated from \(\textbf{D}\) given c. Formally, we want to predict y from \(\{c, d_1,...,d_n | d \in \textbf{D}\}\).
4 Method
The end-to-end learning of Ramses allows the cooperating modules to jointly retrieve and aggregate key information from multiple sources in one output (Fig. 2).
Given the context c and the documents \(\textbf{D}\), our method first generates relevance scores on \(\textbf{D}\) with a biomedical solution based on Dpr [20]:
where \(Enc_{\beta }\) and \(Enc_{\theta }\) are two different BioBert-base models trained to produce a dense representation of documents and the context [39], respectively, \(\oplus \) is the inner product between them, and p(d|c) is the relevance score associated to the document d given c. Thus, our solution finds the most top-k relevant texts according to c. Then, given c and each \(d \in \) top-k, a Bart-base model [22] draws a distribution for each next output token for each d, before marginalizing:
where \(d^{'} = [c, tok, d]\) is the concatenation of c and \(d \in \text {top-}k\) with a special text separator token (<doc-sep>) to make the model aware of the textual boundary, N is the target length, and \(p_\gamma (y_z | d^{'}, y_{1:z-1})\) is the probability of generating the target token \(y_z\) given \(d'\) and the previously generated tokens \(y_{1:z-1}\).
We train our Ramses model by minimizing the negative marginal log-likelihood of each target with the following loss function:
End-to-End Learning. The model (Eq. 2) allows the gradient to backpropagate to all modules. For clarity, we rewrite the formula as a continuous function, as follows:
where \((d_j, s_j) \in \text {top-}k\) and \(B_\gamma \) is Bart.
The presence of \(s_j\) in Eq. 4 allows the gradient, computed by minimizing the objective function, to reach \(Enc_{\beta }\) and \(Enc_{\theta }\). For this reason, the documents and context embeddings are adjusted during the training to improve the generated summary, making all modules of our solution learn jointly in an end-to-end fashion.
5 FAQsumC19 Dataset
We introduce a new dataset, FAQsumC19, containing 514 Covid-19-related FAQs with abstractive answers written by experts, each supported by 30 abstracts of scientific articles, for a total of 15,420 documents. We obtained from the Covid-19 FAQ section on WHOFootnote 2 all available question-answer pairs. We then augmented each instance with 30 Covid-19 scientific articles strictly related to the question from the updated version of the Cord-19 dataset [47]. Specifically, we experimented with the selection of supporting articles with different information retrieval methods, such as a random baseline, Bm25 [44], and Sublimer [38]. We used the concatenation between the question and the answer to retrieve the first 30 ranked documents regarding semantic similarity, creating a knowledge base to support the answer generation. We finally split the dataset into 464 instances for training (\(\approx 90\%\)) and 50 for the test (\(\approx 10\%\)).
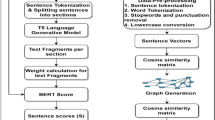
To assess the quality of question-cluster pairs in our dataset, we computed the content coverage with ROUGE-1 precision [24] and BERTScore [51] of the question-answer concatenation w.r.t. each document in the cluster, and calculate the average score. We evaluate the syntactic and semantic overlap between the question and answer and the texts. Table 1 reveals that Sublimer achieves the best scores, as expected.
6 Experiments
6.1 Experimental Setup
Datasets. Table 2 reports the statistics of the datasets used to test Ramses in different biomedical tasks: Ms2 [7] consists of 15,597 instances derived from the scientific literature. Each sample is composed of (i) the background statement, which describes the context research issue, (ii) the target statement, which is the summary to generate; and (iii) the studies, which are the abstracts of biomedical documents that contain the needed information for the research issue. FAQsumC19 is our proposed dataset that comprises 514 Covid-19 FAQs with abstractive answers written by experts, each supported by 30 abstracts of scientific papers.
Baselines. We compare Ramses with SOTA solutions: Bart-FiD [7], which is Bart with the Fusion-in-Decoder strategy [18], encodes all sources individually and combines their hidden states before decoding. Led-Gaq [7], which is Led [2] with global attention on the input query, concatenates all texts in a single input of up to 16,384 tokens. Damen [34], a retrieval-enhanced solution with marginalization-based decoding, discriminates important fragments of the cluster with a frozen Bert-base model and marginalizes their probability distribution during decoding. Primera [49], which is Led pre-trained with a multi-document summarization-specific objective, concatenates the texts with a special separator token up to 4096 tokens in size.
Evaluation Metrics. We use ROUGE-1/2/L [24] to assess fluency and informativeness. We also adopt \(\mathcal {R}\) [32] as an aggregated judgment that considers the variance of the ROUGE scores. Finally, we perform qualitative analysis to bridge the superficiality of automatic evaluation measures.
Implementation. We fine-tune the models using PyTorch and the HuggingFace library, setting the seed to 42 for reproducibility. Ramses is trained on an NVIDIA RTX 3090 GPU of 24 GB memory from an internal cluster for 1 epoch with a learning rate of 3e-5 on Ms2 and for 3 epochs with a learning rate of 1e-5 on FAQsumC19. For decoding, we use the beam search with 4 beams and the following min-max target size: 32–256 for Ms2 and 100-256 for FAQsumC19.
6.2 Results
Table 3 reports the performance of the models in the two evaluation datasets. Ramses yields better scores, suggesting that the retrieve-and-rank end-to-end learning is more effective than prior SOTA approaches in both biomedical CA-MDS tasks.
The Impact of k. As our method relies on learning to select the best top-k relevant documents from the cluster, the value of k is crucial for model performance and GPU memory occupation. Therefore, we analyze the impact of k on model performance by experimenting with a different number of documents to retrieve: 3, 6, 9, 12, 15, 18. Table 4 reports a slight performance improvement as k increases until a threshold is reached (e.g., \(k=9\) for Bart-base), indicating that the marginalization approach with more documents helps produce better ROUGE scores. However, a high k (i.e., \(k\ge 12\)) can also increase information redundancy and contradiction, lowering the final performance. Table 4 also lists the results of different models on single text summarization as the aggregator’s checkpoint, such as Bart and Pegasus [50]. We notice that Bart-large achieves better ROUGE scores, although Pegasus is the largest model. However, as Bart-base achieved a slightly lower result despite the noticeably fewer trainable parameters, we chose to use it for all experiments. Therefore, we tested the best checkpoint of Bart-base trained with \(k=9\) with a different k at the inference time in Ms2. Table 5 reports that the best performance has been achieved with \(k=12\). Furthermore, Table 5 also shows the results on FAQsumC19 with a different k at training time, revealing a trend similar to Ms2.
Memory Requirements. Figure 3 shows the memory complexity at the training time of Ramses for each k. We notice that the memory occupation is linear w.r.t. k, indicating that our solution is not computationally expensive, even for large clusters.
6.3 Ablation Studies
Table 6 reports the ablation studies on Ms2 using Ramses with Bart-base and \(k=9\) with the same hyperparameter settings for all experiments.
Excluding the input context from the input concatenation to give to the generative aggregator (w/o context) leads to the most significant decrease in performance. Indeed, the context is the research question shared by all documents in the cluster, so it contains important information for the final summary.
Training a single model to encode both the context and documents (w/o bi-encoder), namely using a shared BioBert model, decreases performance. Indeed, since the context and the documents have two different purposes (i.e., we need the context to select context-related documents), two models are needed to specialize and differentiate the text representation. Despite the similarity of the two tasks, they differ for two main reasons: (i) the context is relatively shorter than the documents in the cluster, and (ii) the concept expressiveness is denser in the context than in the more verbose documents.
Removing the token separator <doc-sep> between the context and document (w/o token-sep) decreases performance. Indeed, this token is needed to make the model aware of the textual boundary between the context and the documents.
Using cosine similarity instead of the inner product (w/o inner-product) to score the documents against the input context achieves the worst results.
Freezing \(Enc_\beta \) (w/o trained-retrieval) decreases performance, highlighting the usefulness of end-to-end learning to allow the model to select more informative documents.
Switching the context with the documents in the input concatenation (w/o context-first) decreases performance, indicating that the leading context in the input helps the generative aggregator to focus better on how to join context-related information.
6.4 Human Evaluation
Considering the drawbacks of automatic metrics such as ROUGE [9], which is still the standard for evaluating text generation, we qualitatively evaluated the answers inferred from the FAQs of the entire FAQsumC19 test set with three domain experts with master’s degrees in medical and biological areas.
Instructions. We gave evaluators a table, with each row containing the question and three possible answers in random order: (i) the “gold” from WHO, (ii) the prediction of Ramses, and (iii) the prediction of Led-Gaq (the second-best model in FAQsumC19 according to \(\mathcal {R}\)). Each expert was asked to order the answers according to how thoroughly they answered the question, focusing primarily on the factuality. For fairness, we did not inform the evaluators about the answers’ origins and the test’s goal. Overall, experts completed the task in two days, reporting no difficulty ordering the 50 answers.
Results. Evaluation results, reported in Table 7, show that our method produces better informative abstractive answers to a given open question than a linear transformer with sparse attention. To be precise, the experts rated 76% of the answers of our solution as better than those of Led, with 46% agreement between the annotators (which means that 46% of the time, the three evaluators agree). Furthermore, the evaluators also found that 7.33% of the answers inferred by Ramses are more informative than “gold” from WHO. Nevertheless, model generations are still far from being as informative as gold answers, indicating the limitations of current neural language models in FAQsumC19.
7 Conclusion
In this paper, we introduced Ramses, a retrieve-and-rank end-to-end learning solution for CA-MDS of biomedical studies. Ramses is designed to simultaneously acquire indexing capabilities and retrieve pertinent documents to generate comprehensive summaries. Through multiple experiments on two biomedical datasets (including our proposed FAQsumC19 to answer Covid-19 FAQs), we found that Ramses outperforms SOTA models. This finding suggests that the integrated retrieval mechanism significantly benefits the CA-MDS task. Yet, human assessments indicate that there is still notable room for improvement, motivating further research in pursuit of novel retrieval applications within the realm of biomedical multi-document summarization.
Future works can investigate and include multimodal [36, 37], cross-domain [8], and knowledge propagation [4, 5, 26] approaches.Footnote 3
References
Amplayo, R.K., Lapata, M.: Informative and controllable opinion summarization. In: EACL, Online, April 19–23 2021, pp. 2662–2672. ACL (2021)
Beltagy, I., Peters, M.E., Cohan, A.: Longformer: The long-document transformer. CoRR abs/2004.05150 (2020)
Borgeaud, S., Mensch, A., Hoffmann, J., Cai, T., et al.: Improving language models by retrieving from trillions of tokens. In: ICML. PMLR, vol. 162, pp. 2206–2240. PMLR (2022)
Cerroni, W., Moro, G., Pasolini, R., Ramilli, M.: Decentralized detection of network attacks through P2P data clustering of SNMP data. Comput. Secur. 52, 1–16 (2015). https://doi.org/10.1016/j.cose.2015.03.006
Cerroni, W., Moro, G., Pirini, T., Ramilli, M.: Peer-to-peer data mining classifiers for decentralized detection of network attacks. In: ADC. CRPIT, vol. 137, pp. 101–108. ACS (2013)
Chen, Q., Allot, A., Lu, Z.: Litcovid: an open database of COVID-19 literature. Nucleic Acids Res. 49(Database-Issue), D1534–D1540 (2021)
DeYoung, J., Beltagy, I., van Zuylen, M., Kuehl, B., et al.: Ms \(\hat{}\) 2: Multi-document summarization of medical studies. In: EMNLP, Punta Cana, 7–11 November, 2021, pp. 7494–7513. ACL (2021). https://doi.org/10.18653/v1/2021.emnlp-main.594
Domeniconi, G., Moro, G., Pagliarani, A., Pasolini, R.: On deep learning in cross-domain sentiment classification. In: IC3K (Volume 1), Funchal, Madeira, Portugal, November 1–3, 2017, pp. 50–60. SciTePress (2017). https://doi.org/10.5220/0006488100500060
Fabbri, A.R., Kryscinski, W., McCann, B., Xiong, C., et al.: Summeval: re-evaluating summarization evaluation. TACL 9, 391–409 (2021). https://doi.org/10.1162/tacl_a_00373
Formal, T., Piwowarski, B., Clinchant, S.: Match your words! a study of lexical matching in neural information retrieval. In: Hagen, M., et al. (eds.) Advances in Information Retrieval: 44th European Conference on IR Research, ECIR 2022, Stavanger, Norway, April 10–14, 2022, Proceedings, Part II, pp. 120–127. Springer, Cham (2022). https://doi.org/10.1007/978-3-030-99739-7_14
Frisoni, G., Italiani, P., Salvatori, S., Moro, G.: Cogito Ergo \(Summ\): Abstractive Summarization of Biomedical Papers via Semantic Parsing Graphs and Consistency Rewards. In: AAAI 2023, Washington, DC, USA, February 7–14, 2023. AAAI Press, Washington, DC, USA (2023)
Frisoni, G., Mizutani, M., Moro, G., Valgimigli, L.: Bioreader: a retrieval-enhanced text-to-text transformer for biomedical literature. In: EMNLP 2022, pp. 5770–5793. ACL, Abu Dhabi, United Arab Emirates (2022)
Hammoudi, Slimane, Quix, Christoph, Bernardino, Jorge (eds.): Data Management Technologies and Applications: 9th International Conference, DATA 2020, Virtual Event, July 7–9, 2020, Revised Selected Papers. Springer, Cham (2021)
Frisoni, G., Moro, G., Carbonaro, A.: Learning interpretable and statistically significant knowledge from unlabeled corpora of social text messages: a novel methodology of descriptive text mining. In: DATA, pp. 121–134. SciTePress (2020)
Frisoni, G., Moro, G., Carbonaro, A.: A survey on event extraction for natural language understanding: Riding the biomedical literature wave. IEEE Access 9, 160721–160757 (2021). https://doi.org/10.1109/ACCESS.2021.3130956
Grusky, M., Naaman, M., Artzi, Y.: Newsroom: A dataset of 1.3 million summaries with diverse extractive strategies. In: NAACL (Long Papers), pp. 708–719. ACL, New Orleans, Louisiana (2018). https://doi.org/10.18653/v1/N18-1065
Hokamp, C., Ghalandari, D.G., Pham, N.T., Glover, J.: Dyne: Dynamic ensemble decoding for multi-document summarization. CoRR abs/2006.08748 (2020)
Izacard, G., Grave, E.: Leveraging passage retrieval with generative models for open domain question answering. In: EACL: Main Volume, pp. 874–880. ACL, Online (2021). https://doi.org/10.18653/v1/2021.eacl-main.74
Jin, H., Wang, T., Wan, X.: Multi-granularity interaction network for extractive and abstractive multi-document summarization. In: ACL, Online, July 5–10 2020, pp. 6244–6254. ACL (2020). https://doi.org/10.18653/v1/2020.acl-main.556
Karpukhin, V., Oğuz, B., Min, S., Lewis, P., et al.: Dense passage retrieval for open-domain question answering. arXiv preprint arXiv:2004.04906 (2020)
Karpukhin, V., Oguz, B., Min, S., Lewis, P.S.H., et al.: Dense passage retrieval for open-domain question answering. In: EMNLP 2020, Online, November 16–20, 2020, pp. 6769–6781. ACL (2020). https://doi.org/10.18653/v1/2020.emnlp-main.550
Lewis, M., Liu, Y., Goyal, N., Ghazvininejad, M., et al.: BART: denoising sequence-to-sequence pre-training for natural language generation, translation, and comprehension. In: ACL, July 5–10 2020, pp. 7871–7880 (2020). https://doi.org/10.18653/v1/2020.acl-main.703
Lewis, P.S.H., Perez, E., Piktus, A., Petroni, F., et al.: Retrieval-augmented generation for knowledge-intensive NLP tasks. In: NeurIPS 2020, December 6–12, 2020, virtual (2020)
Lin, C.Y.: ROUGE: A package for automatic evaluation of summaries. In: Text Summarization Branches Out, pp. 74–81. ACL, Barcelona, Spain (2004)
Liu, Y., Lapata, M.: Hierarchical transformers for multi-document summarization. In: ACL, Florence, Italy, July 28- August 2 2019, pp. 5070–5081. ACL (2019). https://doi.org/10.18653/v1/p19-1500
Lodi, S., Moro, G., Sartori, C.: Distributed data clustering in multi-dimensional peer-to-peer networks. In: (ADC), Brisbane, 18–22 January, 2010. CRPIT, vol. 104, pp. 171–178. ACS (2010)
Möller, T., Reina, A., Jayakumar, R., Pietsch, M.: Covid-qa: a question answering dataset for Covid-19 (2020)
Moro, G., Masseroli, M.: Gene function finding through cross-organism ensemble learning. BioData Min. 14(1), 14 (2021)
Moro, G., Piscaglia, N., Ragazzi, L., Italiani, P.: Multi-language transfer learning for low-resource legal case summarization. Artif. Intell. Law 31 (2023)
Moro, G., Ragazzi, L.: Semantic self-segmentation for abstractive summarization of long documents in low-resource regimes. In: AAAI 2022, Virtual Event, February 22 - March 1, 2022, pp. 11085–11093. AAAI Press (2022). www.ojs.aaai.org/index.php/AAAI/article/view/21357
Moro, G., Ragazzi, L.: Align-then-abstract representation learning for low-resource summarization. Neurocomputing 548, 126356 (2023). https://doi.org/10.1016/j.neucom.2023.126356
Moro, G., Ragazzi, L., Valgimigli, L.: Carburacy: summarization models tuning and comparison in eco-sustainable regimes with a novel carbon-aware accuracy. AAAI 37(12), 14417–14425 (2023). https://doi.org/10.1609/aaai.v37i12.26686
Moro, G., Ragazzi, L., Valgimigli, L.: Graph-based abstractive summarization of extracted essential knowledge for low-resource scenario. In: ECAI 2023, Kraków, Poland, September 30 - October 4, 2023, pp. 1–9 (2023)
Moro, G., Ragazzi, L., Valgimigli, L., Freddi, D.: Discriminative marginalized probabilistic neural method for multi-document summarization of medical literature. In: ACL, pp. 180–189. ACL, Dublin, Ireland (May 2022). https://doi.org/10.18653/v1/2022.acl-long.15
Moro, G., Ragazzi, L., Valgimigli, L., Frisoni, G., Sartori, C., Marfia, G.: Efficient memory-enhanced transformer for long-document summarization in low-resource regimes. Sensors 23(7) (2023). https://doi.org/10.3390/s23073542, www.mdpi.com/1424-8220/23/7/3542
Moro, G., Salvatori, S.: Deep vision-language model for efficient multi-modal similarity search in fashion retrieval, pp. 40–53 (09 2022). https://doi.org/10.1007/978-3-031-17849-8_4
Moro, G., Salvatori, S., Frisoni, G.: Efficient text-image semantic search: A multi-modal vision-language approach for fashion retrieval. Neurocomputing 538, 126196 (2023). https://doi.org/10.1016/j.neucom.2023.03.057
Moro, G., Valgimigli, L.: Efficient self-supervised metric information retrieval: A bibliography based method applied to COVID literature. Sensors 21(19) (2021). https://doi.org/10.3390/s21196430
Papanikolaou, Y., Bennett, F.: Slot filling for biomedical information extraction. CoRR abs/2109.08564 (2021)
Poliak, A., Fleming, M., Costello, C., Murray, K.W., et al.: Collecting verified COVID-19 question answer pairs. In: NLP4COVIDEMNLP. ACL (2020)
Rajpurkar, P., Jia, R., Liang, P.: Know what you don’t know: Unanswerable questions for squad. In: ACL 2018, Melbourne, Australia, July 15–20, 2018, pp. 784–789. ACL (2018). https://doi.org/10.18653/v1/P18-2124
Ren, R., Lv, S., Qu, Y., Liu, J., et al.: PAIR: leveraging passage-centric similarity relation for improving dense passage retrieval. In: ACL/IJCNLP (Findings). Findings of ACL, vol. ACL/IJCNLP 2021, pp. 2173–2183. Association for Computational Linguistics (2021)
Ren, R., Qu, Y., Liu, J., Zhao, W.X., et al.: Rocketqav2: A joint training method for dense passage retrieval and passage re-ranking. In: EMNLP (1), pp. 2825–2835. ACL (2021)
Croft, Bruce W.., van Rijsbergen, C.. J.. (eds.): SIGIR ’94. Springer London, London (1994). https://doi.org/10.1007/978-1-4471-2099-5
Sun, S., Sedoc, J.: An analysis of bert faq retrieval models for Covid-19 infobot (2020)
Vig, J., Fabbri, A.R., Kryscinski, W., Wu, C., et al.: Exploring neural models for query-focused summarization. In: NAACL 2022, Seattle, WA, United States, July 10–15, 2022, pp. 1455–1468. ACL (2022). https://doi.org/10.18653/v1/2022.findings-naacl.109
Wang, L.L., Lo, K., Chandrasekhar, Y., Reas, R., et al.: CORD-19: the Covid-19 open research dataset. CoRR abs/2004.10706 (2020)
Wei, J.W., Huang, C., Vosoughi, S., Wei, J.: What are people asking about Covid-19? A question classification dataset. CoRR abs/2005.12522 (2020)
Xiao, W., Beltagy, I., Carenini, G., Cohan, A.: PRIMERA: Pyramid-based masked sentence pre-training for multi-document summarization. In: ACL, pp. 5245–5263. ACL, Dublin (2022). https://doi.org/10.18653/v1/2022.acl-long.360
Zhang, J., Zhao, Y., Saleh, M., Liu, P.J.: PEGASUS: pre-training with extracted gap-sentences for abstractive summarization. In: ICML, 13–18 July 2020. vol. 119, pp. 11328–11339. PMLR (2020)
Zhang, T., Kishore, V., Wu, F., Weinberger, K.Q., et al.: Bertscore: Evaluating text generation with BERT. In: ICLR, Addis Ababa, Ethiopia, April 26–30, 2020. OpenReview.net (2020)
Zhang, X.F., Sun, H., Yue, X., Lin, S.M., et al.: COUGH: A challenge dataset and models for COVID-19 FAQ retrieval. In: EMNLP 2021, Virtual Event, 7–11 November, 2021, pp. 3759–3769. ACL (2021)
Acknowledgements
This research is partially supported by (i) the Complementary National Plan PNC-I.1, “Research initiatives for innovative technologies and pathways in the health and welfare sector” D.D. 931 of 06/06/2022, DARE—DigitAl lifelong pRevEntion initiative, code PNC0000002, CUP B53C22006450001, (ii) the PNRR, M4C2, FAIR—Future Artificial Intelligence Research, Spoke 8 “Pervasive AI,” funded by the European Commission under the NextGeneration EU program. The authors thank the Maggioli Group for granting the Ph.D. scholarship to Luca Ragazzi and Lorenzo Valgimigli.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Moro, G., Ragazzi, L., Valgimigli, L., Molfetta, L. (2023). Retrieve-and-Rank End-to-End Summarization of Biomedical Studies. In: Pedreira, O., Estivill-Castro, V. (eds) Similarity Search and Applications. SISAP 2023. Lecture Notes in Computer Science, vol 14289. Springer, Cham. https://doi.org/10.1007/978-3-031-46994-7_6
Download citation
DOI: https://doi.org/10.1007/978-3-031-46994-7_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-46993-0
Online ISBN: 978-3-031-46994-7
eBook Packages: Computer ScienceComputer Science (R0)