Abstract
Otitis media (OM) is a complex disease of multifactorial etiology with a strong genetic component as evidenced by heritability studies and previously identified rare and common variants that confer susceptibility to OM. Recent scientific and technological advancements have helped propel our understanding of OM in the genetic context, with numerous genetic loci and genes brought into focus. Many of these genes relate the immune system and craniofacial development to OM susceptibility. Findings have also highlighted the complexity of OM pathogenesis with the implication of gene–environment interactions and changes to the microbiota of the middle ear and nasopharynx due to genetic variants. Using the latest genetic tools, the identification of OM-pathogenic variants in diverse populations will further increase our knowledge of OM pathophysiology, which can then be used to optimize prevention and treatment of OM.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Otitis media
- Candidate gene study
- Exome sequencing
- Genome-wide association studies
- Linkage analysis
- Mouse models
- Transcriptome
Introduction
Otitis media (OM) is a complex disease with various risk factors for its development, chronicity, and recurrence as well as a multifactorial etiology that is yet to be fully understood. An intricate interplay among environmental, genetic, microbial, anatomic, and immunological factors contributes to the pathogenesis of OM. Within this context, young age, lack of breastfeeding, use of pacifiers, day-care attendance or overcrowding at home, and exposure to tobacco smoke and particulate matter, to name a few, have been identified as environmental risk factors for the development of OM [1,2,3].
Within the otolaryngology practice, the presence of family history remains one of the most important inquiries made during an evaluation of a child with OM. Inherited or genetic factors were found to influence the early onset and development of OM, with a higher incidence observed in certain populations [4,5,6,7]. For example, the frequency of OM is increased in children with chromosomal abnormalities such as Down syndrome (MIM 190685) and Turner syndrome (MIM 300082), in which the affected chromosomes often contain numerous genes of various functions [8, 9]. Chromosome 21, which is affected in Down syndrome, contains many genes that are involved in the immune system, particularly those that play a role in interferon signaling, thereby causing a predisposition to viral and bacterial infections [10,11,12,13,14,15]. Compounded by distinct craniofacial structures of Down syndrome and its associated dysfunctions, such as small middle ear compartments, poor palatal tone, and maldeveloped Eustachian tubes, individuals with Down syndrome often experience OM that does not resolve easily, requiring multiple tympanostomy tube procedures, and subsequent persistent tympanic membrane perforation and hearing loss [16,17,18,19]. Turner syndrome is similar in its clinical manifestation with immune alterations found in T cells and immunoglobulins (Igs) along with craniofacial abnormalities of the palate, jaw, and ears that range from the external to inner ear system [20,21,22]. As a result, affected individuals are predisposed to a range of otologic problems from recurrent or chronic OM to conductive or sensorineural hearing loss and tympanic membrane pathology [9, 23].
The socioeconomic impact of health-care costs and delay in speech and language for OM-affected individuals underline the need for further research in OM. Although our current understanding of the main contributors of genetic susceptibility to OM remains scant compared to our overall genetic knowledge of other common diseases, such as cardiovascular disorders and cancer, continued efforts toward the identification of the determinants of human susceptibility and host genomic responses to OM will help elucidate its multifactorial etiology. Recent advances in genomic databases, sequencing, and analytic techniques will benefit the ongoing genetic investigation into OM and ultimately lead to a better understanding of its pathogenesis, which, in turn, can improve strategies for its prevention and treatment.
Human DNA Studies
Heritability
Heritability, in the narrow sense, is an estimate of the proportion of the variance in phenotypic or trait values under study (in this case, OM), which is due to genetic factors [24]. Because OM is a complex trait, heritability has been estimated using data from related individuals as was done in twin studies performed in three cohorts from Norway, the United States, and the United Kingdom (UK). The initial twin study from Norway in 1997 consisted of 2850 twins, which was then followed up in 2004 with an enlarged cohort of 4247 twins [25, 26]. In the initial study, gender differences were detected in female twins who had higher heritability for OM at 74% compared to males with 45% heritability [25]. In the follow-up study that included an additional 1397 twins, gender-based differences for heritability estimates were not observed, although the overall heritability for OM remained high at 61–72% [26].
In Pittsburgh, a prospective cohort study of 168 same-sex twins and 7 same-sex triplets was conducted to determine the heritability of time or duration of middle ear effusion (MEE) [27]. The same cohort was reassessed after 5 years, and it was determined that the heritability of the time with MEE as a phenotype remained consistent [28]. Heritability was 73% in the initial study and 72% in the follow-up study, and both had p < 0.001 [27, 28].
Rovers et al. (2002) assessed the heritability of both OM and chronic airway blockage in a prospective study of 1373 twins from the UK [29]. The heritability of acute OM was estimated to be 57%. For both OM and chronic airway blockade, an increasing genetic contribution to the phenotypic variance was observed at ages 2 years (49%), 3 years (66%), and 4 years (71%). However, no gender differences were observed [29].
Valuable information can also be gleaned from familial studies with multiple OM-affected siblings as in the study by Hafrén [30]. Heritability estimates of recurrent acute and chronic OM were determined in their survey of 2436 children, who underwent surgery for OM, and their relatives from 590 Finnish families. Chronic OM had lower heritability (22%) compared to recurrent acute OM (38%), and the overall heritability of all types of OM was estimated at 48% [30]. As expected, heritability estimates are higher in identical twins than in other familial relations due to the greater similarity in both genetic makeup and environmental background of twins. Nonetheless, these heritability studies, albeit with various heritability estimates, phenotypic definitions, ethnic background, and cohort composition, confirm the significant contribution of genetic factors to OM susceptibility.
Genetic Linkage Studies
Genetic linkage studies test whether a pattern of inheritance of a disease or trait coincides with the genotypes of families: genetic linkage is therefore a test of co-segregation of a variant or variants inherited together (e.g., a haplotype) with the known disease status of family members. The main metric for linkage is the logarithm of odds (LOD) score, which is usually considered statistically significant if a LOD score of 3.3 or greater is obtained for a variant or haplotype within pedigrees that co-segregate a disease or a trait [31]. To obtain a significant LOD score, either a large enough pedigree or a consanguineous family with multiple affected individuals is needed. Alternatively, smaller families that each on its own cannot obtain linkage but, when analyzed together, can lead to mapping of significant loci may be used. Before the advent of next-generation sequencing, genetic variants that are distributed across the genome were genotyped and then tested for linkage in order to identify a genomic region where the pathogenic variant is statistically the most likely to occur. Early linkage studies using different cohorts of families identified multiple loci in various regions of the human genome, most significantly on chromosome 10q (Table 10.1). It should be noted, however, that in many of the mapped loci, the pathogenic variants responsible for OM susceptibility remain unknown. This is partly due to the lack of genetic studies performed in families with OM and potentially due to genetic heterogeneity for OM where families carry variants within multiple genes that are mostly rare or private variants, such that current family cohorts lack the power to detect significant OM loci. Below is a review of loci that have been mapped using families with OM.
19q13.42–q13.43
Daly et al. (2004) first identified the 19q13.42–q13.43 region as a suggestive susceptibility locus (LOD 2.61, p = 5.3 × 10−4) using genotype data from Minnesotan families with OM [32]. Approximately 100 annotated genes are present in this region, but many of their functions remain unknown. The leukocyte receptor cluster (LRC) genes were suggested as potential candidate genes within the 10q locus. Within this region are genes encoding the leukocyte Ig-like receptor (LIR; also known as Ig-like transcripts) and the killer cell immunoglobulin-like receptor (KIR), which are transmembrane proteins expressed on cells of immune function for various protein activation or inhibition [33]. Such proteins include tyrosine phosphatases Src homology region 2 domain-containing phosphatase (SHP)-1 and/or SHP-2 [34]. This region was replicated by Chen et al. (2011), who fine-mapped the locus to chromosome 19q13.43 using additional single nucleotide variant genotypes (LOD 3.75, p = 1.6 × 10−5) [35]. The additional candidate genes identified within the 19q locus include genes related to zinc fingers, the tumor necrosis factor-alpha (TNFα), bone morphogenetic protein (BMP), and fibroblast growth factor-beta (FGFβ) pathways, lymphocyte activation, and the inflammasome protein complex, all of which regulate innate immunity in response to harmful stimuli [35,36,37].
10q26.3
In an initial linkage study for OM using the Minnesota family cohort, the 10q26.3 locus was identified with a LOD score of 3.78 (p = 3.0 × 10−5) and was replicated in another cohort of Western Australia-based trios by Rye et al. in 2014 (Zlr 2.69, p = 3.6 × 10−3) [32, 33]. Herein, trios are composed of the OM-affected individual (proband) and both parents, without affected or unaffected siblings. Within the 10q26.3 locus, the Minnesotan and Australian studies identified candidate genes encoding a disintegrin and metalloproteinase (ADAM) domain. ADAM8 is found in leukocytes in response to allergen exposure in asthma and is also upregulated in epithelial cells of the airway during allergic inflammation [38, 39]. The function of ADAM12 is most strongly associated with cell adhesion and fusion, extracellular matrix restructuring, and cell signaling [40]. It has been found to be upregulated in the middle ear in response to tobacco smoke [41]. Other genes implicated in this region include DOCK1 (MIM 601403), which is involved in phagocytosis; TCERG1L encoding transcription elongation regulator-like protein, and PPP2R2D (MIM 613992), which modulates the modulator of the transforming growth factor (TGF)/Activin/Nodal pathway [33, 42, 43].
10q22.3
Chromosomal region 10q22.3 has also been identified (LOD >2, p = 1.81 × 10−3) and replicated with suggestive evidence (Zlr 1.64, p = 0.05), though not reaching statistical significance [2, 33]. SFTPA1 (MIM 178630) and SFTPA2 (MIM 178642) are genes within this region that together encode the human surfactant protein (SP-A) and have previously been implicated in OM [44]. SP-A haplotype and genotype variations have been observed in children with the first episode of OM before 6 months, and common variants in these genes were associated with protection against OM in infants at risk of asthma [45, 46]. SP-A is expressed in the middle ear mucosa and in the Eustachian tube and is known to contribute to innate immune responses by increasing the phagocytosis of otopathogens [45,46,47,48].
Candidate Gene Association Studies
Using a gene of interest, candidate gene association studies estimate the frequency with which the minor allele, or the less prevalent allele in the background population, is found in OM-affected patients in comparison to a control group. Healthy or unaffected individuals that are related to the OM-affected patients within the same families, or unrelated healthy individuals with no previous history of OM, may be used as controls (case–control study). Numerous studies have been conducted, particularly using candidate genes, relating to inflammation and innate immune responses, but many candidate gene studies will not lead to a significant association when a more stringent threshold for genome-wide significance is applied. Nonetheless, some genes that were initially identified in smaller case–control association studies, such as the HLA and ABO (MIM 110300) genes, were also deemed significant loci in genome-wide association studies (GWASs) for OM [49,50,51,52,53,54,55,56].
Genome-Wide Association Studies (GWASs)
A GWAS is currently the starting point in identifying the loci of interest. It assesses the association of variants spread throughout the genome with the trait or disease under investigation and, in contrast to candidate gene association studies, does not assume an association between a locus of interest and the trait. Instead, a GWAS employs an agnostic approach to identifying the loci of interest, which may be a regulatory variant often found in the non-coding region, a haplotype encompassing variants that are inherited together, or a gene harboring multiple rare variants associated with the trait. For OM, a series of GWASs have been conducted using common variant genotypes from microarrays, with each describing independent findings (Table 10.2). With advances in computational efficiency, continuous decline in the cost of sequencing, and increasing availability of large-scale biobank data with genotypes and OM phenotypes, the discovery of additional novel findings from GWASs, which include both common and rare variants, is possible in the near future.
In the first GWAS on OM, Rye et al. (2012) used data from Australian trios and identified three novel candidate genes/gene clusters, namely, (1) rs6755194 within chromosome 2p23.1, which is either upstream of or intronic to two isoforms of CAPN14 (MIM 610229); (2) rs1862981, also located on 2p23.1, intronic to GALNT14 (MIM 60822) and proximal to CAPN14; and (3) BPIFA clusters, including BPIFA1 (MIM 607412), BPIFA2, and BPIFA3 on the genomic region 20q11.21 [57]. However, these findings could not be replicated in an independent cohort [58]. Other genes identified in this study with trends toward association were linked to the TGF-β pathway [57].
In the Minnesota-based cohort, Allen et al. (2013) identified an intergenic locus rs10497394 between the genes CDCA7 (MIM 609937) and SP3 (MIM 601804) on 2q31.1, which was subsequently replicated in an independent Pittsburgh-based US cohort as being associated with both chronic OM with effusion and recurrent acute OM [59]. This locus was found to regulate the expression of LDLR (MIM 606945) on chromosome 19, a gene that is expressed in ciliated epithelial cells of the airway with its transcribed protein predicted to be a binding site for human rhinovirus C, a known pathogen for upper respiratory tract infections and OM [60]. Newer data from the Genotype-Tissue Expression (GTEx) database showed that the rs10497394 variant significantly regulates RNA levels of CDCA7 in thyroid and arterial tissues; however, GTEx does not include data from middle ear tissues [61]. CDCA7 mutations cause immunodeficiency, centromeric instability, and facial anomalies syndrome (MIM 616910), wherein hypo- or agammaglobulinemia leads to recurrent life-threatening infections [62]. Additional loci on chromosomes 5 and 15 (rs386057, rs1110060, and rs10775247) were also identified but did not reach genome-wide significance in the replication study [59]. The genes affected by these three variants are involved in microtubule function, regulation of mammalian sonic hedgehog and Indian hedgehog pathways, and DNA replication [63,64,65].
Einarsdottir et al. (2016) found three variants, all on chromosome 19, to be associated with childhood OM in the Finnish population [66]. None of the identified regions overlapped with those previously found in association with OM on chromosome 19. Genome-wide significance was established for rs16974263, a variant intronic to the PRX (MIM 605725) gene, which was found in association with chronic OM with effusion in a UK cohort, albeit with opposite directions of effect [66]. Out of the several genes identified within this region are three candidate genes, PLD3 (MIM 615698), SERTAD1 (MIM 617850), and BLVRB (MIM 600941), which are previously known to be associated with immune function with expression found in macrophages [67, 68]. In the GTEx database, the rs16974263 variant regulates either the RNA levels or splicing of isoforms for SERTAD3, HIPK4 (MIM 611712), PLD3, and PRX in various tissues [61]. In particular, SERTAD3 is expressed in the mucosal tissue and its protein inhibits the replication of influenza A virus upon induction by type I interferon responses during an infection [61, 69].
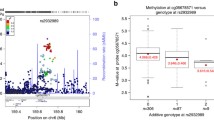
Additional studies in Americans of European-descent identified genome-wide significance for a variant on chromosome 6 (rs2932989) that alters the methylation status of the gene encoding fibronectin type III domain containing 1 (FNDC1, MIM 609991) [70]. Although its function is not clearly elucidated, the study found an upregulation of this gene in the middle ear tissue under pro-inflammatory conditions [70]. Other candidate genes from suggestive loci largely contained those related to immune responses.
Finally, 14 genome-wide significant loci were identified by Pickrell et al. (2016) and Tian et al. (2017) using GWAS data from 23&Me [49, 53]. Among these loci, the OM-associated genes included FUT2 (MIM 182100) and ABO, both involved in glycosphingolipid biosynthesis, as well as TBX1 (MIM 602054) and MKX (MIM 601332), which encode transcription factors. Genes important for embryogenesis and neurodevelopment (FGF3 (MIM 164950) and AUTS2 (MIM 607270)), as well as those implicated in asthma (CDHR3 (MIM 615610)), and spontaneous chronic OM in mice (PLG (MIM 173350)) were also found in association with OM [71,72,73]. To date, only 4 of these 14 loci, FUT2, TBX1, ABO, and CDHR3, have been replicated in independent human cohorts with additional evidence from functional or multi-omics studies performed for FUT2 and CDHR3 [49,50,51,52, 74,75,76,77,78].
To identify pathways that are potentially important for OM susceptibility, 21 genes with p < 5 × 10−5 available in the literature were selected from candidate gene association studies and GWAS results. Using NetworkAnalyst software [79,80,81], these genes were then used as an input for defining protein–protein interaction networks. Within these networks exist subnetworks that are involved in significant pathways, highlighting the underlying pathophysiology and serving as potential targets for novel therapeutics (Fig. 10.1). The major pathways identified included a rhythmic process, a viral process, negative regulation of an? apoptotic process, transcription, transcription by RNA polymerase II, chromatin remodeling, regulation of cell cycle, and circadian rhythm (Table 10.3). These identified pathways suggest that basic cellular processes are involved in OM susceptibility with viral infection as a primary target for OM prevention and treatment.
Protein–protein interactions in the network analysis of genes implicated in otitis media from exome sequencing and genome-wide association studies. The green nodes represent proteins encoded by input or seed genes with a p < 5 × 10−5 from exome sequencing and genome-wide association studies. The orange nodes represent proteins that are found in association with the seed proteins within a significant cellular process
Exome Sequencing
The coding regions or exome, i.e., all the exons comprising the parts of all genes that are transcribed and translated into proteins, only make up approximately 1% of the human genome. With growing knowledge and advancements in technology, exome sequencing allows for the assessment of variants within all coding regions of the genome and has become a cheaper alternative for identifying pathogenic variants for various diseases, particularly for rare variants with strong phenotypic effects. Using exome sequence data, rare pathogenic variants in three genes, A2ML1 (MIM 610627), FUT2, and SPINK5 (MIM 605010), were found in association with OM in an Indigenous Filipino pedigree and has since been replicated in various OM cohorts of different ethnicities [74, 82,83,84].
The A2ML1 gene encodes a protease inhibitor that is specifically expressed in the middle ear [83]. It is similar to alpha-2-macroglobulin (A2M), also a protease found in the middle ear that is associated with recurrent acute OM [85]. Two loss-of-function A2ML1 variants, i.e., a frameshift variant and a splice variant, were identified in a cohort of Indigenous Filipino population with an estimated 50% prevalence of OM [83]. The same frameshift variant in A2ML1 was identified in US-based otitis-prone children with early-onset severe OM (p = 3.34 × 10−14) [83]. An additional 25 variants of the same gene were found in probands with OM from different cohorts [82, 83]. Moreover, differential expression analysis of RNA sequence data from Coloradan children with OM revealed co-upregulation of A2ML1 with genes (e.g., SPINK5) that are involved in several pathways, including keratinocyte differentiation [82]. Follow-up microbiota studies showed a significantly higher relative abundance of the Leptotrichia spp. in the middle ears of individuals with A2ML1 variants [84, 86]. In contrast, for a rare pathogenic SPINK5 variant that was also identified in the Indigenous Filipino population through exome sequencing and linkage analysis, a greater biodiversity in the oral cavity and increased relative abundance of Microbacteriaceae were observed [84]. Notably, A2ML1 is expressed in the middle ear mucosa, whereas SPINK5 is faintly localized to the mucosal tissues but strongly localized to the outer ear and tympanic membrane in mouse middle ears [83, 84].
FUT2, located on chromosome 19, encodes 2-fucosyltransferase and is involved in the synthesis of blood group H antigens and the regulation of its expression on various mucosal surfaces [87]. Several common and rare variants within FUT2 were associated with an increased risk for OM in US trios and various ethnic cohorts [74]. These include two stop variants, p.Trp154* and p.Arg202*, and two missense variants, p.Ala104Val and p.Arg138Cys [74]. For these four FUT2 variants, levels of A antigen, a common binding spot for potential otopathogens and commensal bacteria, were reduced on the cell surface of mutant epithelial cells [74]. FUT2 variants also affect gene transcription and alter the mucosal microbiome in the setting of OM. Differential expression analysis revealed downregulation of FN1, KMT2D, MUC16, and NBPF20 and upregulation of MTAP in individuals with the FUT2 p.Trp154* variant [75]. Changes in regulation of these four genes were also seen in the middle ears of mice inoculated with the human otopathogen non-typeable Haemophilus influenzae (NTHi), except upregulation of Fn1 found in the inoculated mice [75]. Additionally, the FUT2 variant was associated with an increased load of otopathogens within the middle ear with concordant decrease in relative taxa abundance of commensal bacteria in the nasopharynx [75].
Also utilizing exome sequencing, Jamieson et al. reported two genes, NR3C1 (MIM 138040) and NREP (MIM 607332), as candidates for severe OM in a cohort from the Australian Aboriginal population [88]. Although these genes failed to attain genome-wide significance, NR3C1 and NREP implicate a role of gene–environment interaction in the expression of inflammatory modulators due to environmental stress and shifts in the microbiome, respectively [89,90,91,92,93].
Mouse DNA Studies
A critical limitation to human studies is the inability to adequately control for genetic or environmental factors due to variability and heterogeneity across human subjects. Animal models present an alternative method in the exploration of the genetics of OM through the control of genetic and environmental factors and have led to a growing number of genes identified in association with OM [94]. Despite the availability of numerous animal models in the study of OM, the murine model has been largely favored in genetic studies. In all, 99% of mouse genes are homologous to human genes and, consequently, mice and humans share similar physiology and development of many basic functions [95]. Owing to the early mapping of the mouse genome, tools to manipulate mouse genes for phenotyping and susceptibility studies are widely available, along with a growing number of knockout and transgenic mouse models [96].
Gene-driven models and phenotype-driven N-ethyl-N-nitrosourea (ENU) mutagenesis methods have been used to understand the genes involved in OM in mice. In the gene-driven method, a growing availability of transgenic and knockout mouse strains through efforts such as the Knockout Mouse Phenotyping Program (KOMP2) is harnessed to assess the effect of mutations on OM susceptibility [97]. In the phenotype-driven model, ENU is used as a mutagen to induce random nucleotide changes [95]. This is then followed by screening of the phenotype of interest (e.g., increased OM prevalence), allowing for the discovery of novel genes. Genetically altered mice from either method may express the desired phenotypes, for example, by developing spontaneous OM, or in other studies, changes in OM expression after inoculation of otopathogens through transbullar injection or pressure-induced translocation of microbes to the middle ear from the nasopharynx [98, 99]. The genes identified thus far mainly involve craniofacial development or innate immune responses. Here, we will discuss a number of genes with a distinctive expression of OM phenotypes, illustrating the key points.
Mice with Craniofacial Abnormalities
Mice with impaired craniofacial development from DNA modifications often develop chronic OM with effusion and are frequently associated with congenital syndromes. Gene-driven models have led to the implication of numerous genes in knockout mice with similar phenotypic observations. E2f4 is a key transcription factor that interacts with pRB during cell cycle progression [100]. Many E2f4−/− mice die early on, but the surviving mice develop OM due to craniofacial defects that increase their susceptibility to opportunistic infections [101]. In humans, mutations in the EYA gene family, comprised of nuclear phosphatases that act as transcriptional coactivators, are known to cause branchiootorenal syndrome 1 (MIM 113650) from variants in EYA1 (MIM 601653) and sensorineural hearing loss (MIM 601316) from variants in EYA4 (MIM 603550) [102,103,104,105,106]. In mice, deficiency of Eya4 caused middle ear and Eustachian tube abnormalities, ultimately leading to OM with effusion [107]. Similar findings were observed in haploinsufficient Ets1 and Fli1 mice that developed OM and had craniofacial abnormalities consistent with those of the small middle ear cavity, short nasal bone, and malformation of the nasal bony-cartilaginous junction [108]. Specific to the middle ear, anatomical abnormalities, including ossicular fusion to the middle ear wall and stapedial malformations, were observed. The genes ETS1 (MIM 164720) and FLI1 (MIM 193067) lie within a genomic region of the human genome on the long arm of chromosome 11, which is deleted in Jacobsen syndrome (MIM 147791); Jacobsen syndrome includes craniofacial dysmorphisms and isoimmune thrombocytopenia among its clinical features [109]. These genes are E26 transformation-specific (ETS) transcription factors that are expressed in neural crest cells during embryogenesis [110,111,112].
ENU-derived edison and Jeff variants both lead to spontaneous chronic OM and mild craniofacial defects in mice [113, 114]. In both edison and Jeff variants, their respective mutations in the mouse genes Nisch and Fbxo11 are also associated with faulty innate immune responses (further described below) [115]. Eya4 is another gene with a possible regulatory role in the immune system [116].
Studies of Innate Immune Responses in Mice
An innate immune response is broad and encompasses cellular functions of macrophages and neutrophils to mucosal integrity and ciliary function. As such, the genetic findings related to innate immunity are accordingly diverse. Toll-like receptors (Tlrs) are a class of pattern recognition receptors involved in innate immunity at the cellular level. Numerous Tlr mutations have been identified through knockout mouse models (gene-driven) as causing or prolonging otitis media in mice. Tlr4 deletion caused a delay in immune responses, leading to an early development of OM, which was often chronic [117, 118]. Knockout of Tlr2 and Tlr9 also resulted in similar findings with prolonged or severe infections [119, 120]. A similar phenotype was observed with the deletion of Myd88, which encodes an adaptor protein recruited in response to Tlr and interleukin (IL)-1 for activation of downstream signaling pathways [121, 122]. After Myd88 knockout, a delay in macrophage and neutrophil recruitment resulted in chronic OM and prolonged mucosal thickening, which were more severe than those found with Tlr mutations [123].
In response to TLR activation, the NF-κB pathway is triggered downstream to initiate an inflammatory response [124]. Through phenotype-driven models, a number of genes have been associated with the NF-κB pathway and OM. As described above, the edison mouse variant displayed disruption in immune response in addition to the observed craniofacial abnormalities. The deletion of the Nisch gene further impacted the downstream signaling of the NF-κB pathway as well as the LIMK1 pathway, which is associated with vascular permeability and perturbation that lead to middle ear effusion and mucoperiosteal inflammation [113, 125, 126]. Another ENU-derived mouse strain is Junbo, which has a deletion of the Evi1 gene and demonstrated spontaneous acute OM and chronic OM [127]. Evi1 encodes a transcription factor that is known to be involved in multiple pathways [128, 129]. In the NF-κB pathway, Evi1 binds to a subunit of NF-κB, preventing its interaction with DNA, thereby downregulating inflammation [130].
In the TGF-β pathway, the activation of TGF-β causes a cell signaling cascade with SMAD proteins, leading to transcription of target genes, RNA processing, translation of messenger RNA (mRNA), and protein regulation for numerous cellular processes, including craniofacial development and inflammation [131,132,133,134]. The genes implicated in association with the TGF-β pathway include Tgif1, Fbxo11, and Evi1. As previously mentioned, the phenotypic changes observed in Jeff mice were attributed to a mutation in the Fbxo11 gene, which encodes for a ubiquitination protein involved in tumor suppression [135, 136]. Jeff mice displayed nuclear accumulation of the Smad2 protein, which mediates transcription of target genes within the TGF-β pathway [137]. Evi1, defects in which are responsible for the OM observed in Junbo mice, interacts with Smad3 proteins, resulting in the suppression of the growth-inhibiting mechanism of the TGF-β pathway [127, 129, 138]. Based on the mutagenesis studies that implicated the TGF-β pathway in OM in these gene-based mouse models, TGIF1, a homeodomain protein that acts as a negative regulator of the TGF-β pathway, binds to Smad2 to recruit a co-repressor and subsequently recruits histone deacetylases, leading to inhibition of transcription [132, 139, 140]. In Tgif1-knockout mice, conductive hearing loss was associated with chronic effusion, middle ear mucosal thickening, and an increase in goblet cells [141].
Other pathways and functions presumed to be associated with OM include the c-Jun N-terminal kinase (JNK) pathway, mucociliary clearance, mitogen-activated protein kinase (MAPK) pathway, and bony development [76, 129, 142,143,144,145]. Despite the vast expansion of genetic knowledge gained from these mouse studies, there are a number of limitations that prevent rapid application of knowledge from mouse models to application in the management of OM in humans. Because the otopathogens in humans differ from those in mice, these mouse models require inoculation of human otopathogens into the middle ear, which often cause a shorter course of OM in mice. There are also immunological and anatomical differences. The lymphocyte-rich immune system of mice versus the neutrophil-rich immune system in humans as well as the lack of mastoid air cells in mice require additional consideration [146]. At the genetic level, alternative splicing and innumerable factors contributing to complex disease processes like OM require additional complementary studies in humans.
RNA Studies
There are various types of RNAs, each serving numerous functions beyond protein translation to the regulation of gene expression. Examination of RNA profiles can thus provide a more accurate and detailed picture of the cellular mechanisms at play for various disease processes. A messenger RNA (mRNA) is a single-stranded RNA, complementary to the DNA, used as codes for protein synthesis. In contrast, a microRNA (miR) provides insights into gene expression. A microRNA is a 23 nucleotide-long RNA that functions to control gene expression posttranscriptionally by binding to the 3′ untranslated region (UTR) of a target mRNA [147]. Accordingly, investigators have studied mRNAs and miRNAs in order to understand OM pathophysiology.
mRNAs
mRNA expression in OM has been studied through the assessment of single gene transcription or whole transcriptome of various cells within the middle ear. Single-gene studies performed on human samples showed differential expression of aquaporins, C-type lectin receptors, mucins, beta-defensins, and cytokines in different types of OM, implicating the roles of cellular homeostatic and inflammatory responses in OM pathogenesis [148,149,150,151,152]. Differential expression of the protooncogene C/EBP-homologous protein (DDIT3, MIM 126337) was observed in otitis-prone children, associating the frequency of OM with the endoplasmic reticulum stress response through the PERK (protein kinase RNA-like endoplasmic reticulum kinase) signaling pathway [153].
Various methods of genetic expression analysis, including reverse transcription PCR (RT-PCR), microarray, and RNA sequencing (RNA-seq), have been used to profile the genetic expression of middle ear epithelial cells (MEECs) in association with OM. The types of information that can be gathered from these studies are wide-ranging. Inoculation of murine models with a common human otopathogen, NTHi, helped identify differential expression of 3657 genes, most of which are involved in innate immune response modulation, cell marker variation, and recruitment of neutrophils and macrophages [154]. The substantial role that innate immunity plays in OM was reinforced in the study by Ryan et al. (2020) with single-cell transcriptome analysis in mice [155]. Considerable cellular diversity in the mouse middle ear mucosa was observed with identification of 17 distinct cell types, all with expression of innate immune genes [155]. The cellular diversity of the middle ear mucosal tissue is further demonstrated in the variable types of MEECs identified [156, 157]. In the process of MEEC differentiation, approximately 500 genes that are associated with secretory proteins and ciliogenesis were upregulated [158]. Such findings provide a pathway for improving the characterization of the unrestrained responses of MEECs with the cellular remodeling that occurs in OM. Stabenau et al. (2021) studied human MEECs in OM with effusion in comparison with normal mucosal cells and identified 1282 differentially expressed genes [159]. The functions of identified genes encompassed inflammation, bacterial immunity, mucociliary clearance, cellular proliferation and transformation, and auditory cell differentiation. Of these functions, the most upregulated genes involved mucin production, immunity, and cell cycle regulation.
MicroRNAs (miRs)
miRs have diverse actions and targets that are context-specific. Most commonly, miRs bind to the 3′UTR of an mRNA in perfect pairs or with imperfect complementation but with additional base pairings at the 5′ of the mRNA [160]. miRs have also been observed to bind to 5′UTRs, the coding region of RNA, or directly to gene promoters [161,162,163]. The complexity in understanding the mechanism of miRs is also due to the ability of one miR to bind to numerous distinct mRNAs. Furthermore, one mRNA can be bound by different miRs, with each pairing leading to a distinct outcome [161]. As a result of miR binding to mRNA and 3′UTR, translational suppression can occur as deadenylases and decapping factors are recruited [164]. With its protective ends exposed, the resulting mRNA strand will be prone to degradation. Binding of miR to 5′UTR and the coding region causes silencing of expression, but its interaction with a promotor region has been observed to lead to transcription [165, 166]. The presence of miRs in various cellular compartments, the abundance of miRs, mRNAs, and their various combinations, and the miRNA-induced silencing complex mediating translational inhibition make the function of miRs dynamic [162].
The role of miRs in OM is increasingly being examined. In vitro analysis of human MEECs revealed differential expression of 15 genes as a result of both up- and downregulating functions of miRs. These changes ultimately led to an increase in cell differentiation, endocytosis, cellular communication, IκBK/NF-κB cascade, developmental process, complement activation, innate immune response, and cell adhesion [167]. miR-146 has been implicated in numerous inflammatory diseases owing to its negative regulatory role in activating TLRs and its ability to fine-tune signaling cascades. In OM, miR-146 expression increased in response to in vitro exposure to pro-inflammatory cytokines, which, in turn, was correlated with middle ear mucosal thickness and observed decline in the expression of tumor necrosis factor receptor (TNFR)-associated factor 6 (TRAF6), a modulator in the TLR pathway [168].
Exosomal release of endocytic vesicles leads to extracellular release of mRNAs, miRs, and proteins to be then transported between cells [169, 170]. Particularly, exosomal miRs have been found to influence expressions of distant cells and promote positive or negative effects on pro-inflammatory signaling of the receiving cell [171,172,173,174]. Human MEECs produce a baseline level of extracellular miR, the microRNAome, which is composed of 110 different miRs during in vitro stimulation [175]. For example, stimulation by NTHi induces elevation in the levels of five distinct miRs, namely, miR-378a-3p + miR-378i, miR-200a-3p, miR-378g, miR30d-5p, and miR-222-3p, all known to target genes related to innate immunity [175,176,177,178,179,180]. The targeted mRNAs are involved in apoptosis, cancer, cell growth, and the IL-8 pathway mediated by CXCR1/2 that activates NF-κB, oxidative stress, inflammation, chemotaxis, angiogenesis, and neutrophil functions [175]. Of the miRs, miR-320e has been further associated with the presence of allergies in children with OM [181]. In an exosomal miR analysis in patients with middle ear effusion, 17 miRs unique to middle ear effusions compared to serum controls were identified, with the most abundant being miR-223-3p [170]. These exosomal miRs regulate a total of 442 target genes, most of which were involved in the IL-8 signaling process and were also mediated by CXCR1/2. miR-223 regulates innate immune response, protease activity, and a few key functions of neutrophils [182, 183].
Another form of RNA that plays a role similar to miR is long non-coding RNA (lncRNA). As the name suggests, it is longer than miR with its length greater than 200 nucleotides and does not code any proteins [184, 185]. It has the ability to interact with DNA, RNA, and proteins and modulate transcription, epigenetic changes, RNA and protein stabilization, translation, and posttranslational modifications [186,187,188,189]. Interestingly, it can also interact with miR to modulate cellular function. In OM, the lncRNA nuclear-enriched abundant transcript 1 (NEAT1) targets miR-495 to activate p38 MAPK, allowing the release of inflammatory cytokines and promoting the expression of genes involved in acute inflammatory responses [190,191,192].
The extent of miR interactions and their functions are still in the process of discovery with much to uncover. Understanding the role of RNA in the expression of final protein products helps delineate complex and dynamic disease processes such as OM.
Future Directions
Although prevention and diagnostic and treatment strategies are available for OM patients, we still lack the understanding of the progression of OM from acute to chronic forms. This lack of knowledge undermines efforts to develop novel therapies and fine-tune the current management protocols, particularly for patients with undiscovered genetic susceptibility to OM. To reduce the massive burden on global healthcare due to OM, ongoing investigations furthering our understanding of the genetic component of OM susceptibility are important to ultimately deliver effective therapeutic solutions to patients. We must continue to identify OM-pathogenic variants within different populations, especially to include underrepresented groups across the world. Advancements in technology bring about efficiency and speed in our ability to sequence genetic materials, which must be harnessed in studying complex diseases with multifactorial contributors at play such as OM. Currently, many human studies involve patients of European descent. A few available studies on minority groups focus on populations with an especially high incidence of OM; however, it is equally as important to study underrepresented groups without such context. Using bioinformatics, we can glean the genetic networks at play in various types of OM. The influence of pathogens on gene expression requires an in-depth understanding of the microbiome and gene expression responses to different pathogens. Genetic variations from splicing, epigenetic modifications, and different factors that influence production of proteins are especially important. It is essential to continue harnessing the availability of animal models, especially in studying therapeutic solutions, but even more important is the application of the findings from animal models to treating OM in humans.
References
Swanson JA, Hoecker JL. Concise review for primary-care physicians. Mayo Clin Proc. 1996;71:179–83.
Neto JFL, Hemb L, Silva DB, et al. Systematic literature review of modifiable risk factors for recurrent acute otitis media in childhood. J Pediatr. 2006;82:87–96. http://www.jped.com.br/conteudo/Ing_resumo.asp?varArtigo=1453&cod=&idSecao=3.
Uhari M, Mäntysaari K, Niemelä M. A meta-analytic review of the risk factors for acute otitis media. Clin Infect Dis. 1996;22:1079–83.
Hudson HM, Rockett IR. An environmental and demographic analysis of otitis media in rural Australian aborigines. Int J Epidemiol. 1984;13:73–82.
Daly KA, Hoffman HJ, Kvaerner KJ, Kvestad E, Casselbrant ML, Homoe P, et al. Epidemiology, natural history, and risk factors: panel report from the Ninth International Research Conference on Otitis Media. Int J Pediatr Otorhinolaryngol. 2010;74:231–40.
Gunasekera H, Haysom L, Morris P, Craig J. The global burden of childhood otitis media and hearing impairment: a Systematic review. Pediatrics. 2008;121:S107. https://doi.org/10.1542/peds.2007-2022QQ.
Santos-Cortez RLP, Reyes-Quintos MRT, Tantoco MLC, Abbe I, Llanes EGV, Ajami NJ, et al. Genetic and environmental determinants of otitis media in an indigenous Filipino population. Otolaryngol Head Neck Surg. 2016;155:856–62. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5093071/.
Austeng ME, Akre H, Øverland B, Abdelnoor M, Falkenberg E-S, Kværner KJ. Otitis media with effusion in children with in Down syndrome. Int J Pediatr Otorhinolaryngol. 2013;77:1329–32. https://linkinghub.elsevier.com/retrieve/pii/S0165587613002644.
Bois E, Nassar M, Zenaty D, Léger J, Van Den Abbeele T, Teissier N. Otologic disorders in Turner syndrome. Eur Ann Otorhinolaryngol Head Neck Dis. 2018;135:21–4. https://linkinghub.elsevier.com/retrieve/pii/S1879729617301291.
Ram G, Chinen J. Infections and immunodeficiency in Down syndrome: immunodeficiency in Down syndrome. Clin Exp Immunol. 2011;164:9–16. https://doi.org/10.1111/j.1365-2249.2011.04335.x.
Chung H, Green PHR, Wang TC, Kong X-F. Interferon-driven immune dysregulation in Down syndrome: a review of the evidence. J Inflamm Res. 2021;14:5187–200. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8504936/.
Kong AM, Hurley D, Evans KA, Brixner D, Csoboth C, Visootsak J. A retrospective, longitudinal, claims-based comparison of concomitant diagnoses between individuals with and without Down syndrome. J Manag Care Spec Pharm. 2017;23:761–70. https://doi.org/10.18553/jmcp.2017.23.7.761.
Sullivan KD, Evans D, Pandey A, Hraha TH, Smith KP, Markham N, et al. Trisomy 21 causes changes in the circulating proteome indicative of chronic autoinflammation. Sci Rep. 2017;7:14818.
Kong X-F, Worley L, Rinchai D, Bondet V, Jithesh PV, Goulet M, et al. Three copies of four interferon receptor genes underlie a mild type I interferonopathy in Down syndrome. J Clin Immunol. 2020;40:807–19. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7418179/.
Hüls A, Costa ACS, Dierssen M, Baksh RA, Bargagna S, Baumer NT, et al. Medical vulnerability of individuals with Down syndrome to severe COVID-19–data from the Trisomy 21 Research Society and the UK ISARIC4C survey. EClinicalMedicine. 2021;33:100769. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7897934/.
Nogaki T, Paparella MM, Cureoglu S. A structural analysis of tympanic compartments of the middle ear in patients with Down’s syndrome: a temporal bone study. Otol Neurotol. 2020;41:1149–57.
Omar M, McCoy JL, McCormick AA, Vellody K, Chi DH. Repeat tympanostomy tubes in children with Down syndrome. Int J Pediatr Otorhinolaryngol. 2021;148:110811.
Ghadersohi S, Ida JB, Bhushan B, Billings KR. Outcomes of tympanoplasty in children with Down syndrome. Int J Pediatr Otorhinolaryngol. 2017;103:36–40.
Kreicher KL, Weir FW, Nguyen SA, Meyer TA. Characteristics and progression of hearing loss in children with Down syndrome. J Pediatr. 2018;193:27–33.e2.
Mock, Markert UR, Vogelsang H, Jäger L. Selective T-cell deficiency in Turner’s syndrome. J Investig Allergol Clin Immunol. 2000;10:312–3.
Cacciari E, Masi M, Fantini MP, Licastro F, Cicognani A, Pirazzoli P, et al. Serum immunoglobulins and lymphocyte subpopulations derangement in Turner’s syndrome. J Immunogenet. 1981;8:337–44.
Makishima T, King K, Brewer CC, Zalewski CK, Butman J, Bakalov VK, et al. Otolaryngologic markers for the early diagnosis of Turner syndrome. Int J Pediatr Otorhinolaryngol. 2009;73:1564–7. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2757481/.
Stenberg AE, Nylén O, Windh M, Hultcrantz M. Otological problems in children with Turner’s syndrome. Hear Res. 1998;124:85–90.
Balding DJ, et al., editors. Handbook of statistical genetics, vol. 1 and 2. Somerset, NJ: John Wiley and Sons Ltd; 2003. https://www.biblio.com/book/handbook-statistical-genetics-volumes-1-2/d/1465841857.
Kvaerner KJ, Tambs K, Harris JR, Magnus P. Distribution and heritability of recurrent ear infections. Ann Otol Rhinol Laryngol. 1997;106:624–32.
Kvestad E, Kvaerner KJ, Røysamb E, Tambs K, Harris JR, Magnus P. Otitis media: genetic factors and sex differences. Twin Res. 2004;7:239–44.
Casselbrant ML, Mandel EM, Fall PA, Rockette HE, Kurs-Lasky M, Bluestone CD, et al. The heritability of otitis media: a twin and triplet study. JAMA. 1999;282:2125–30.
Casselbrant ML, Mandel EM, Rockette HE, Kurs-Lasky M, Fall PA, Bluestone CD, et al. The genetic component of middle ear disease in the first 5 years of life. Arch Otolaryngol Head Neck Surg. 2004;130:273–8.
Rovers M, Haggard M, Gannon M, Koeppen-Schomerus G, Plomin R. Heritability of symptom domains in otitis media: a longitudinal study of 1,373 twin pairs. Am J Epidemiol. 2002;155:958–64.
Hafrén L. Genetic background and the risk of otitis media. Int J Pediatr Otorhinolaryngol. 2012;76(1):41–4.
Risch N, Merikangas K. The future of genetic studies of complex human diseases. Science. 1996;273:1516–7.
Daly KA, Brown WM, Segade F, Bowden DW, Keats BJ, Lindgren BR, et al. Chronic and recurrent otitis media: a genome scan for susceptibility loci. Am J Hum Genet. 2004;75:988–97.
Rye MS, Scaman ESH, Thornton RB, Vijayasekaran S, Coates HL, Francis RW, et al. Genetic and functional evidence for a locus controlling otitis media at chromosome 10q26.3. BMC Med Genet. 2014;15:18.
Lanier LL. NK cell receptors. Annu Rev Immunol. 1998;16:359–93.
Chen W-M, Allen EK, Mychaleckyj JC, Chen F, Hou X, Rich SS, et al. Significant linkage at chromosome 19q for otitis media with effusion and/or recurrent otitis media (COME/ROM). BMC Med Genet. 2011;12:124.
Mataki C, Murakami T, Umetani M, Wada Y, Ishii M, Tsutsumi S, et al. A novel zinc finger protein mRNA in human umbilical vein endothelial cells is profoundly induced by tumor necrosis factor alpha. J Atheroscler Thromb. 2000;7:97–103.
Sabater L, Ashhab Y, Caro P, Kolkowski EC, Pujol-Borrell R, Domínguez O. Identification of a KRAB-containing zinc finger protein, ZNF304, by AU-motif-directed display method and initial characterization in lymphocyte activation. Biochem Biophys Res Commun. 2002;293:1066–72.
André P, Biassoni R, Colonna M, Cosman D, Lanier LL, Long EO, et al. New nomenclature for MHC receptors. Nat Immunol. 2001;2:661.
Schroder K, Tschopp J. The inflammasomes. Cell. 2010;140:821–32.
Nyren-Erickson EK, Jones JM, Srivastava DK, Mallik S. A disintegrin and metalloproteinase-12 (ADAM12): function, roles in disease progression, and clinical implications. Biochim Biophys Acta. 2013;1830:4445–55.
Cho J-G, Woo J-S, Lee H-M, Jung HH, Hwang S-J, Chae S. Effects of cigarette smoking on mucin production in human middle ear epithelial cells. Int J Pediatr Otorhinolaryngol. 2009;73:1447–51.
Hasegawa H, Kiyokawa E, Tanaka S, Nagashima K, Gotoh N, Shibuya M, et al. DOCK180, a major CRK-binding protein, alters cell morphology upon translocation to the cell membrane. Mol Cell Biol. 1996;16:1770–6.
Batut J, Schmierer B, Cao J, Raftery LA, Hill CS, Howell M. Two highly related regulatory subunits of PP2A exert opposite effects on TGF-beta/Activin/Nodal signalling. Dev Camb Engl. 2008;135:2927–37.
Floros J, DiAngelo S, Koptides M, Karinch AM, Rogan PK, Nielsen H, et al. Human SP-A locus: allele frequencies and linkage disequilibrium between the two surfactant protein A genes. Am J Respir Cell Mol Biol. 1996;15:489–98.
Rämet M, Löfgren J, Alho O-P, Hallman M. Surfactant protein-A gene locus associated with recurrent otitis media. J Pediatr. 2001;138:266–8. https://linkinghub.elsevier.com/retrieve/pii/S0022347601576544.
Pettigrew MM, Gent JF, Zhu Y, Triche EW, Belanger KD, Holford TR, et al. Association of surfactant protein A polymorphisms with otitis media in infants at risk for asthma. BMC Med Genet. 2006;7:68.
Abdel-Razek O, Ni L, Yang F, Wang G. Innate immunity of surfactant protein A in experimental otitis media. Innate Immun. 2019;25:391–400.
McNeely TB, Coonrod JD. Aggregation and opsonization of type A but not type B Hemophilus influenzae by surfactant protein A. Am J Respir Cell Mol Biol. 1994;11:114–22.
Pickrell JK, Berisa T, Liu JZ, Segurel L, Tung JY, Hinds D. Detection and interpretation of shared genetic influences on 42 human traits. Nat Genet. 2016;48:709–17. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5207801/.
Mortensen EH, Lildholdt T, Gammelgård NP, Christensen PH. Distribution of ABO blood groups in secretory otitis media and cholesteatoma. Clin Otolaryngol Allied Sci. 1983;8:263–5.
Apostolopoulos K, Labropoulou E, Konstantinos B, Rhageed S, Ferekidis E. Blood group in otitis media with effusion. ORL J Otorhinolaryngol Relat Spec. 2002;64:433–5.
Wiesen BM, Hafrén L, Einarsdottir E, Kere J, Mattila PS, Santos-Cortez RLP. ABO genotype and blood type are associated with otitis media. Genet Test Mol Biomark. 2019;23:823–7. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6857544/.
Tian C, Hromatka BS, Kiefer AK, Eriksson N, Noble SM, Tung JY, et al. Genome-wide association and HLA region fine-mapping studies identify susceptibility loci for multiple common infections. Nat Commun. 2017;8:599.
Kalm O, Johnson U, Prellner K. HLA frequency in patients with chronic secretory otitis media. Int J Pediatr Otorhinolaryngol. 1994;30:151–7.
Kalm O, Johnson U, Prellner K, Ninn K. HLA frequency in patients with recurrent acute otitis media. Arch Otolaryngol Head Neck Surg. 1991;117:1296–9.
Kalm O, Johnson U, Prellner K, Ninn K. HLA antigens and recurrent acute otitis media. Acta Otolaryngol Suppl. 1992;492:107–9.
Rye MS, Warrington NM, Scaman ESH, Vijayasekaran S, Coates HL, Anderson D, et al. Genome-wide association study to identify the genetic determinants of otitis media susceptibility in childhood. PLoS One. 2012;7:e48215.
Allen EK, Manichaikul A, Chen W-M, Rich SS, Daly KA, Sale MM. Evaluation of replication of variants associated with genetic risk of otitis media. PLoS One. 2014;9:e104212.
Allen EK, Chen W-M, Weeks DE, Chen F, Hou X, Mattos JL, et al. A genome-wide association study of chronic otitis media with effusion and recurrent otitis media identifies a novel susceptibility locus on chromosome 2. J Assoc Res Otolaryngol. 2013;14:791–800. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3825021/.
Bochkov YA, Palmenberg AC, Lee W-M, Rathe JA, Amineva SP, Sun X, et al. Molecular modeling, organ culture and reverse genetics for a newly identified human rhinovirus C. Nat Med. 2011;17:627–32.
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013;45:580–5.
Thijssen PE, Ito Y, Grillo G, Wang J, Velasco G, Nitta H, et al. Mutations in CDCA7 and HELLS cause immunodeficiency–centromeric instability–facial anomalies syndrome. Nat Commun. 2015;6:7870. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4519989/.
Ovádi J, Orosz F. An unstructured protein with destructive potential: TPPP/p25 in neurodegeneration. BioEssays News Rev Mol Cell Dev Biol. 2009;31:676–86.
Hsu S-HC, Zhang X, Yu C, Li ZJ, Wunder JS, Hui C-C, et al. Kif7 promotes hedgehog signaling in growth plate chondrocytes by restricting the inhibitory function of Sufu. Dev Camb Engl. 2011;138:3791–801.
Kumagai A, Shevchenko A, Shevchenko A, Dunphy WG. Treslin collaborates with TopBP1 in triggering the initiation of DNA replication. Cell. 2010;140:349–59.
Einarsdottir E, Hafrén L, Leinonen E, Bhutta MF, Kentala E, Kere J, et al. Genome-wide association analysis reveals variants on chromosome 19 that contribute to childhood risk of chronic otitis media with effusion. Sci Rep. 2016;6:33240.
Hammarén-Malmi S, Tarkkanen J, Mattila PS. Analysis of risk factors for childhood persistent middle ear effusion. Acta Otolaryngol (Stockh). 2005;125:1051–4.
Jasenosky LD, Scriba TJ, Hanekom WA, Goldfeld AE. T cells and adaptive immunity to Mycobacterium tuberculosis in humans. Immunol Rev. 2015;264:74–87.
Sun N, Li C, Li X-F, Deng Y-Q, Jiang T, Zhang N-N, et al. Type-I interferon-inducible SERTAD3 inhibits influenza A virus replication by blocking the assembly of viral RNA polymerase complex. Cell Rep. 2020;33:108342.
van Ingen G, Li J, Goedegebure A, Pandey R, Li YR, March ME, et al. Genome-wide association study for acute otitis media in children identifies FNDC1 as disease contributing gene. Nat Commun. 2016;7:12792.
Bønnelykke K, Sleiman P, Nielsen K, Kreiner-Møller E, Mercader JM, Belgrave D, et al. A genome-wide association study identifies CDHR3 as a susceptibility locus for early childhood asthma with severe exacerbations. Nat Genet. 2014;46:51–5.
Eriksson P-O, Li J, Ny T, Hellström S. Spontaneous development of otitis media in plasminogen-deficient mice. Int J Med Microbiol. 2006;296:501–9.
Jolley A, Corbett M, McGregor L, Waters W, Brown S, Nicholl J, et al. De novo intragenic deletion of the autism susceptibility candidate 2 (AUTS2) gene in a patient with developmental delay: a case report and literature review. Am J Med Genet A. 2013;161A:1508–12.
Santos-Cortez RLP, Chiong CM, Frank DN, Ryan AF, Giese APJ, Bootpetch Roberts T, et al. FUT2 variants confer susceptibility to familial otitis media. Am J Hum Genet. 2018;103:679–90. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6217759/.
Elling CL, Scholes MA, Streubel S-O, Larson ED, Wine TM, Bootpetch TC, et al. The FUT2 variant c.461G>A (p.Trp154*) is associated with differentially expressed genes and nasopharyngeal microbiota shifts in patients with otitis media. Front Cell Infect Microbiol. 2021;11:798246.
Tian C, Harris BS, Johnson KR. Ectopic mineralization and conductive hearing loss in Enpp1asj mutant mice, a new model for otitis media and tympanosclerosis. PLoS One. 2016;11:e0168159.
Hirsch SD, Elling CL, Bootpetch TC, Scholes MA, Hafrén L, Streubel S-O, et al. The role of CDHR3 in susceptibility to otitis media. J Mol Med. 2021;99:1571–83. https://doi.org/10.1007/s00109-021-02118-7.
Everman JL, Sajuthi S, Saef B, Rios C, Stoner AM, Numata M, et al. Functional genomics of CDHR3 confirms its role in HRV-C infection and childhood asthma exacerbations. J Allergy Clin Immunol. 2019;144:962–71.
Xia J, Benner MJ, Hancock REW. NetworkAnalyst—integrative approaches for protein–protein interaction network analysis and visual exploration. Nucleic Acids Res. 2014;42:W167–74. https://doi.org/10.1093/nar/gku443.
Xia J, Gill EE, Hancock REW. NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat Protoc. 2015;10:823–44.
Zhou G, Soufan O, Ewald J, Hancock REW, Basu N, Xia J. NetworkAnalyst 3.0: a visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res. 2019;47:W234–41.
Larson ED, Magno JPM, Steritz MJ, Llanes EGV, Cardwell J, Pedro M, et al. A2ML1 and otitis media: novel variants, differential expression and relevant pathways. Hum Mutat. 2019;40:1156–71. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6711784/.
Santos-Cortez RLP, Chiong CM, Reyes-Quintos MRT, Tantoco MLC, Wang X, Acharya A, et al. Rare A2ML1 variants confer susceptibility to otitis media. Nat Genet. 2015;47:917–20. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4528370/.
Frank DN, Giese APJ, Hafren L, Bootpetch TC, Yarza TKL, Steritz MJ, et al. Otitis media susceptibility and shifts in the head and neck microbiome due to SPINK5 variants. J Med Genet. 2021;58:442–52.
Bondestam M, Foucard T, Gebre-Medhin M. Serum albumin, retinol-binding protein, thyroxin-binding prealbumin and acute phase reactants as indicators of undernutrition in children with undue susceptibility to acute infections. Acta Paediatr Scand. 1988;77:94–8.
Santos-Cortez RLP, Hutchinson DS, Ajami NJ, Reyes-Quintos MRT, Tantoco MLC, Labra PJ, et al. Middle ear microbiome differences in indigenous Filipinos with chronic otitis media due to a duplication in the A2ML1 gene. Infect Dis Poverty. 2016;5:97. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5088646/.
Ravn V, Dabelsteen E. Tissue distribution of histo-blood group antigens. Acta Pathol Microbiol Immunol Scand. 2000;108:1–28.
Jamieson SE, Fakiola M, Tang D, Scaman E, Syn G, Francis RW, et al. Common and rare genetic variants that could contribute to severe otitis media in an Australian Aboriginal Population. Clin Infect Dis. 2021;73:1860–70. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8599203/.
Barnes PJ. Corticosteroid effects on cell signalling. Eur Respir J. 2006;27:413–26.
Yue MM, Lv K, Meredith SC, Martindale JL, Gorospe M, Schuger L. Novel RNA-binding protein P311 binds eukaryotic translation initiation factor 3 subunit b (eIF3b) to promote translation of transforming growth factor β1-3 (TGF-β1-3). J Biol Chem. 2014;289:33971–83.
Mulligan CJ, D’Errico NC, Stees J, Hughes DA. Methylation changes at NR3C1 in newborns associate with maternal prenatal stress exposure and newborn birth weight. Epigenetics. 2012;7:853–7.
Lewis CR, Breitenstein RS, Henderson A, Sowards HA, Piras IS, Huentelman MJ, et al. Harsh parenting predicts novel HPA receptor gene methylation and NR3C1 methylation predicts cortisol daily slope in middle childhood. Cell Mol Neurobiol. 2021;41:783–93.
Deshpande NP, Riordan SM, Castaño-Rodríguez N, Wilkins MR, Kaakoush NO. Signatures within the esophageal microbiome are associated with host genetics, age, and disease. Microbiome. 2018;6:227.
Santos-Cortez RLP, Bhutta MF, Earl JP, Hafrén L, Jennings M, Mell JC, et al. Panel 3: Genomics, precision medicine and targeted therapies. Int J Pediatr Otorhinolaryngol. 2020;130:109835. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7155947/.
Justice MJ, Noveroske JK, Weber JS, Zheng B, Bradley A. Mouse ENU mutagenesis. Hum Mol Genet. 1999;8:1955–63.
Ringwald M, Iyer V, Mason JC, Stone KR, Tadepally HD, Kadin JA, et al. The IKMC web portal: a central point of entry to data and resources from the International Knockout Mouse Consortium. Nucleic Acids Res. 2011;39:D849–55. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3013768/.
Knockout Mouse Phenotyping. 2013. https://commonfund.nih.gov/komp2.
Ryan AF, Ebmeyer J, Furukawa M, Pak K, Melhus A, Wasserman SI, et al. Mouse models of induced otitis media. Brain Res. 2006;1091:3–8. https://linkinghub.elsevier.com/retrieve/pii/S0006899306003799.
Stol K, van Selm S, van den Berg S, Bootsma HJ, Blokx WAM, Graamans K, et al. Development of a non-invasive murine infection model for acute otitis media. Microbiology. 2009;155:4135–44.
Brehm A, Kouzarides T. Retinoblastoma protein meets chromatin. Trends Biochem Sci. 1999;24:142–5.
Humbert PO, Rogers C, Ganiatsas S, Landsberg RL, Trimarchi JM, Dandapani S, et al. E2F4 is essential for normal erythrocyte maturation and neonatal viability. Mol Cell. 2000;6:281–91.
Ohto H, Kamada S, Tago K, Tominaga SI, Ozaki H, Sato S, et al. Cooperation of six and eya in activation of their target genes through nuclear translocation of Eya. Mol Cell Biol. 1999;19:6815–24.
Rayapureddi JP, Kattamuri C, Steinmetz BD, Frankfort BJ, Ostrin EJ, Mardon G, et al. Eyes absent represents a class of protein tyrosine phosphatases. Nature. 2003;426:295–8.
Tootle TL, Silver SJ, Davies EL, Newman V, Latek RR, Mills IA, et al. The transcription factor Eyes absent is a protein tyrosine phosphatase. Nature. 2003;426:299–302.
Krug P, Morinière V, Marlin S, Koubi V, Gabriel HD, Colin E, et al. Mutation screening of the EYA1, SIX1, and SIX5 genes in a large cohort of patients harboring branchio-oto-renal syndrome calls into question the pathogenic role of SIX5 mutations. Hum Mutat. 2011;32:183–90. https://doi.org/10.1002/humu.21402.
Pfister M, Tóth T, Thiele H, Haack B, Blin N, Zenner H-P, et al. A 4-bp insertion in the eya-homologous region (eyaHR) of EYA4 causes hearing impairment in a Hungarian family linked to DFNA10. Mol Med Camb Mass. 2002;8:607–11.
Depreux FFS, Darrow K, Conner DA, Eavey RD, Liberman MC, Seidman CE, et al. Eya4-deficient mice are a model for heritable otitis media. J Clin Invest. 2008;118:651–8. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2213371/.
Carpinelli MR, Kruse EA, Arhatari BD, Debrincat MA, Ogier JM, Bories J-C, et al. Mice haploinsufficient for Ets1 and Fli1 display middle ear abnormalities and model aspects of Jacobsen syndrome. Am J Pathol. 2015;185:1867–76.
Jones C, Slijepcevic P, Marsh S, Baker E, Langdon WY, Richards RI, et al. Physical linkage of the fragile site FRA11B and a Jacobsen syndrome chromosome deletion breakpoint in 11q23.3. Hum Mol Genet. 1994;3:2123–30.
Hollenhorst PC, McIntosh LP, Graves BJ. Genomic and biochemical insights into the specificity of ETS transcription factors. Annu Rev Biochem. 2011;80:437–71.
Kola I, Brookes S, Green AR, Garber R, Tymms M, Papas TS, et al. The Ets1 transcription factor is widely expressed during murine embryo development and is associated with mesodermal cells involved in morphogenetic processes such as organ formation. Proc Natl Acad Sci U S A. 1993;90:7588–92.
Mélet F, Motro B, Rossi DJ, Zhang L, Bernstein A. Generation of a novel Fli-1 protein by gene targeting leads to a defect in thymus development and a delay in Friend virus-induced erythroleukemia. Mol Cell Biol. 1996;16:2708–18.
Crompton M, Purnell T, Tyrer HE, Parker A, Ball G, Hardisty-Hughes RE, et al. A mutation in Nischarin causes otitis media via LIMK1 and NF-κB pathways. PLoS Genet. 2017;13:e1006969.
Hardisty RE, Erven A, Logan K, Morse S, Guionaud S, Sancho-Oliver S, et al. The deaf mouse mutant Jeff (Jf) is a single gene model of otitis media. J Assoc Res Otolaryngol. 2003;4:130–8. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3202714/.
Hardisty-Hughes RE, Tateossian H, Morse SA, Romero MR, Middleton A, Tymowska-Lalanne Z, et al. A mutation in the F-box gene, Fbxo11, causes otitis media in the Jeff mouse. Hum Mol Genet. 2006;15:3273–9. http://academic.oup.com/hmg/article/15/22/3273/713947/A-mutation-in-the-Fbox-gene-Fbxo11-causes-otitis.
Okabe Y, Sano T, Nagata S. Regulation of the innate immune response by threonine-phosphatase of Eyes absent. Nature. 2009;460:520–4.
MacArthur CJ, Hefeneider SH, Kempton JB, Trune DR. C3H/HeJ mouse model for spontaneous chronic otitis media. Laryngoscope. 2006;116:1071–9.
Hirano T, Kodama S, Fujita K, Maeda K, Suzuki M. Role of Toll-like receptor 4 in innate immune responses in a mouse model of acute otitis media. FEMS Immunol Med Microbiol. 2007;49:75–83.
Leichtle A, Hernandez M, Pak K, Yamasaki K, Cheng C-F, Webster NJ, et al. TLR4-mediated induction of TLR2 signaling is critical in the pathogenesis and resolution of otitis media. Innate Immun. 2009;15:205–15.
Leichtle A, Hernandez M, Lee J, Pak K, Webster NJ, Wollenberg B, et al. The role of DNA sensing and innate immune receptor TLR9 in otitis media. Innate Immun. 2012;18:3–13.
Akira S, Takeda K. Toll-like receptor signalling. Nat Rev Immunol. 2004;4:499–511.
Fritz JH, Girardin SE. How Toll-like receptors and Nod-like receptors contribute to innate immunity in mammals. J Endotoxin Res. 2005;11:390–4. https://doi.org/10.1177/09680519050110060301.
Hernandez M, Leichtle A, Pak K, Ebmeyer J, Euteneuer S, Obonyo M, et al. Myeloid differentiation primary response gene 88 is required for the resolution of otitis media. J Infect Dis. 2008;198:1862–9.
Kawai T, Akira S. TLR signaling. Semin Immunol. 2007;19:24–32.
Gorovoy M, Han J, Pan H, Welch E, Neamu R, Jia Z, et al. LIM kinase 1 promotes endothelial barrier disruption and neutrophil infiltration in mouse lungs. Circ Res. 2009;105:549–56.
Rothwarf DM, Karin M. The NF-kappa B activation pathway: a paradigm in information transfer from membrane to nucleus. Sci STKE Signal Transduct Knowl Environ. 1999;1999:RE1.
Parkinson N, Hardisty-Hughes RE, Tateossian H, Tsai H-T, Brooker D, Morse S, et al. Mutation at the Evi1 locus in Junbo mice causes susceptibility to otitis media. PLoS Genet. 2006;2:e149.
Alliston T, Ko TC, Cao Y, Liang Y-Y, Feng X-H, Chang C, et al. Repression of bone morphogenetic protein and activin-inducible transcription by Evi-1. J Biol Chem. 2005;280:24227–37.
Kurokawa M, Mitani K, Yamagata T, Takahashi T, Izutsu K, Ogawa S, et al. The evi-1 oncoprotein inhibits c-Jun N-terminal kinase and prevents stress-induced cell death. EMBO J. 2000;19:2958–68.
Xu X, Woo C-H, Steere RR, Lee BC, Huang Y, Wu J, et al. EVI1 acts as an inducible negative feedback regulator of NF-κB by inhibiting p65 acetylation. J Immunol. 1950;2012(188):6371–80. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3370108/.
Tzavlaki K, Moustakas A. TGF-β signaling. Biomol Ther. 2020;10:487. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7175140/.
Massagué J, Chen YG. Controlling TGF-beta signaling. Genes Dev. 2000;14:627–44.
Shi Y, Massagué J. Mechanisms of TGF-beta signaling from cell membrane to the nucleus. Cell. 2003;113:685–700.
Kouskoura T, Fragou N, Alexiou M, John N, Sommer L, Graf D, et al. The genetic basis of craniofacial and dental abnormalities. Schweiz Monatsschrift Zahnmed Rev Mens Suisse Odonto-Stomatol Riv Mens Svizzera Odontol E Stomatol. 2011;121:636–46.
Kipreos ET, Pagano M. The F-box protein family. Genome Biol. 2000;1:REVIEWS3002.
Duan S, Cermak L, Pagan JK, Rossi M, Martinengo C, di Celle PF, et al. FBXO11 targets BCL6 for degradation and is inactivated in diffuse large B-cell lymphomas. Nature. 2012;481:90–3. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3344385/.
Tateossian H, Hardisty-Hughes RE, Morse S, Romero MR, Hilton H, Dean C, et al. Regulation of TGF-β signalling by Fbxo11, the gene mutated in the Jeff otitis media mouse mutant. PathoGenetics. 2009;2:5. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2714483/.
Mitani K. Molecular mechanisms of leukemogenesis by AML1/EVI-1. Oncogene. 2004;23:4263–9.
Bertolino E, Reimund B, Wildt-Perinic D, Clerc RG. A novel homeobox protein which recognizes a TGT core and functionally interferes with a retinoid-responsive motif. J Biol Chem. 1995;270:31178–88.
Massagué J, Wotton D. New EMBO member’s review. EMBO J. 2000;19:1745–54. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC302010/.
Tateossian H, Morse S, Parker A, Mburu P, Warr N, Acevedo-Arozena A, et al. Otitis media in the Tgif knockout mouse implicates TGFβ signalling in chronic middle ear inflammatory disease. Hum Mol Genet. 2013;22:2553–65. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3674796/.
Yao W, Frie M, Pan J, Pak K, Webster N, Wasserman SI, et al. C-Jun N-terminal kinase (JNK) isoforms play differing roles in otitis media. BMC Immunol. 2014;15:46.
Azar A, Piccinelli C, Brown H, Headon D, Cheeseman M. Ectodysplasin signalling deficiency in mouse models of hypohidrotic ectodermal dysplasia leads to middle ear and nasal pathology. Hum Mol Genet. 2016;25:3564–77. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5179950/.
Li X, Xu L, Li J, Li B, Bai X, Strauss JF, et al. Otitis media in sperm-associated antigen 6 (Spag6)-deficient mice. PLoS One. 2014;9:e112879.
Konduru AS, Matsuyama S, Lee B-C, Komatsu K, Li J-D. Curcumin inhibits NTHi-induced MUC5AC mucin overproduction in otitis media via upregulation of MAPK phosphatase MKP-1. Int J Inflamm. 2017;2017:4525309.
Mestas J, Hughes CCW. Of mice and not men: differences between mouse and human immunology. J Immunol Baltim Md 1950. 2004;172:2731–8.
Bartel DP. MicroRNA target recognition and regulatory functions. Cell. 2009;136:215–33. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3794896/.
Doo JG, Kim YI, Shim HS, Kim DJ, Park JW, Dong SH, et al. Expression of C-type lectin receptor mRNA in otitis media with effusion and chronic otitis media with and without cholesteatoma. Auris Nasus Larynx. 2019;46:672–80. https://linkinghub.elsevier.com/retrieve/pii/S0385814618306680.
Jung SY, Kim SS, Kim YI, Kim H-S, Kim SH, Yeo SG. Expression of aquaporins mRNAs in patients with otitis media. Acta Otolaryngol (Stockh). 2018;138:701–7.
Kerschner JE, Tripathi S, Khampang P, Papsin BC. MUC5AC expression in human middle ear epithelium of patients with otitis media. Arch Otolaryngol Head Neck Surg. 2010;136:819–24.
Jin Shin D, Gan-Undram S, Jin Kim S, Joon Jun Y, Jung Im G, Hyun Jung H. Expression of β-defensins in the tubotympanum of experimental otitis media. Acta Otolaryngol (Stockh). 2006;126:1040–5. https://doi.org/10.1080/00016480600672626.
Trune DR, Kempton B, Hausman FA, Larrain BE, MacArthur CJ. Correlative mRNA and protein expression of middle and inner ear inflammatory cytokines during mouse acute otitis media. Hear Res. 2015;326:49–58. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4492826/.
Kang DW, Dong SH, Kim SH, Kim YI, Park DC, Yeo SG. Expression of endoplasmic reticulum stress-related mRNA in otitis media with effusion. Int J Pediatr Otorhinolaryngol. 2019;121:109–13. https://linkinghub.elsevier.com/retrieve/pii/S0165587619301272.
Hernandez M, Leichtle A, Pak K, Webster NJ, Wasserman SI, Ryan AF. The transcriptome of a complete episode of acute otitis media. BMC Genomics. 2015;16:259.
Ryan AF, Nasamran CA, Pak K, Draf C, Fisch KM, Webster N, et al. Single-cell transcriptomes reveal a complex cellular landscape in the middle ear and differential capacities for acute response to infection. Front Genet. 2020;11:358. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7174727/.
Luo W, Yi H, Taylor J, Li J, Chi F, Todd NW, et al. Cilia distribution and polarity in the epithelial lining of the mouse middle ear cavity. Sci Rep. 2017;7:45870. http://www.nature.com/articles/srep45870.
Tucker AS, Dyer CJ, Romero JMF, Teshima THN, Fuchs JC, Thompson H. Mapping the distribution of stem/progenitor cells across the middle ear during homeostasis and inflammation. Development. 2017;145:dev.154393. https://doi.org/10.1242/dev.154393/264412/Mapping-the-distribution-of-stem-progenitor-cells.
Mulay A, Chowdhury MMK, James CT, Bingle L, Bingle CD. The transcriptional landscape of the cultured murine middle ear epithelium in vitro. Biol Open. 2021;10:bio056564.
Stabenau KA, Zimmermann MT, Mathison A, Zeighami A, Samuels TL, Chun RH, et al. RNA sequencing and pathways analyses of middle ear epithelia from patients with otitis media. Laryngoscope. 2021;131:2590–7. https://doi.org/10.1002/lary.29551.
Liu B, Li J, Cairns MJ. Identifying miRNAs, targets and functions. Brief Bioinform. 2014;15:1–19. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3896928/.
Correia de Sousa M, Gjorgjieva M, Dolicka D, Sobolewski C, Foti M. Deciphering miRNAs’ action through miRNA editing. Int J Mol Sci. 2019;20:6249. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6941098/.
O’Brien J, Hayder H, Zayed Y, Peng C. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol. 2018;9:402.
Place RF, Li L-C, Pookot D, Noonan EJ, Dahiya R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc Natl Acad Sci U S A. 2008;105:1608–13.
Martin HC, Wani S, Steptoe AL, Krishnan K, Nones K, Nourbakhsh E, et al. Imperfect centered miRNA binding sites are common and can mediate repression of target mRNAs. Genome Biol. 2014;15:R51.
Zhang J, Zhou W, Liu Y, Liu T, Li C, Wang L. Oncogenic role of microRNA-532-5p in human colorectal cancer via targeting of the 5′UTR of RUNX3. Oncol Lett. 2018;15:7215. https://doi.org/10.3892/ol.2018.8217.
Dharap A, Pokrzywa C, Murali S, Pandi G, Vemuganti R. MicroRNA miR-324-3p induces promoter-mediated expression of RelA gene. PLoS One. 2013;8:e79467. https://doi.org/10.1371/journal.pone.0079467.
Song J-J, Kwon SK, Cho CG, Park S-W, Chae S-W. Microarray analysis of microRNA expression in LPS induced inflammation of human middle ear epithelial cells (HMEECs). Int J Pediatr Otorhinolaryngol. 2011;75:648–51. https://linkinghub.elsevier.com/retrieve/pii/S0165587611000668.
Samuels TL, Yan J, Khampang P, MacKinnon A, Hong W, Johnston N, et al. Association of microRNA 146 with middle ear hyperplasia in pediatric otitis media. Int J Pediatr Otorhinolaryngol. 2016;88:104–8.
Janas T, Janas MM, Sapoń K, Janas T. Mechanisms of RNA loading into exosomes. FEBS Lett. 2015;589:1391–8.
Val S, Jeong S, Poley M, Krueger A, Nino G, Brown K, et al. Purification and characterization of microRNAs within middle ear fluid exosomes: implication in otitis media pathophysiology. Pediatr Res. 2017;81:911–8. http://www.nature.com/articles/pr201725.
O’Neill LA, Sheedy FJ, McCoy CE. MicroRNAs: the fine-tuners of Toll-like receptor signalling. Nat Rev Immunol. 2011;11:163–75.
Camussi G, Deregibus MC, Bruno S, Cantaluppi V, Biancone L. Exosomes/microvesicles as a mechanism of cell-to-cell communication. Kidney Int. 2010;78:838–48.
Moon S-K, Moon S-K, Park R, Moon S-K, Park R, Lee H-Y, et al. Spiral ligament fibrocytes release chemokines in response to otitis media pathogens. Acta Otolaryngol (Stockh). 2006;126:564–9. https://doi.org/10.1080/00016480500452525.
Watanabe T, Jono H, Han J, Lim DJ, Li J-D. Synergistic activation of NF-kappaB by nontypeable Haemophilus influenzae and tumor necrosis factor alpha. Proc Natl Acad Sci U S A. 2004;101:3563–8.
Val S, Krueger A, Poley M, Cohen A, Brown K, Panigrahi A, et al. Nontypeable Haemophilus influenzae lysates increase heterogeneous nuclear ribonucleoprotein secretion and exosome release in human middle-ear epithelial cells. FASEB J. 2018;32:1855–67. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5893168/.
Li H-G, Zhao L-H, Bao X-B, Sun P-C, Zhai B-P. Meta-analysis of the differentially expressed colorectal cancer-related microRNA expression profiles. Eur Rev Med Pharmacol Sci. 2014;18:2048–57.
Pellatt DF, Stevens JR, Wolff RK, Mullany LE, Herrick JS, Samowitz W, et al. Expression profiles of miRNA subsets distinguish human colorectal carcinoma and normal colonic mucosa. Clin Transl Gastroenterol. 2016;7:e152. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4822091/.
Lin J, Wang Y, Zou Y-Q, Chen X, Huang B, Liu J, et al. Differential miRNA expression in pleural effusions derived from extracellular vesicles of patients with lung cancer, pulmonary tuberculosis, or pneumonia. Tumour Biol. 2016;37:15835.
Bracken CP, Li X, Wright JA, Lawrence DM, Pillman KA, Salmanidis M, et al. Genome-wide identification of miR-200 targets reveals a regulatory network controlling cell invasion. EMBO J. 2014;33:2040–56. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4195771/.
Inamura K, Ishikawa Y. MicroRNA in lung cancer: novel biomarkers and potential tools for treatment. J Clin Med. 2016;5:36. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4810107/.
Adamczyk P, Narożna B, Szczepankiewicz A, Bręborowicz A, Pucher B, Kotowski M, et al. Decreased miRNA-320e correlates with allergy in children with otitis media with effusion. Auris Nasus Larynx. 2021;48:1061–6.
Gantier MP. The not-so-neutral role of microRNAs in neutrophil biology. J Leukoc Biol. 2013;94:575–83.
Baek D, Villén J, Shin C, Camargo FD, Gygi SP, Bartel DP. The impact of microRNAs on protein output. Nature. 2008;455:64–71.
Hangauer MJ, Vaughn IW, McManus MT. Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS Genet. 2013;9:e1003569.
St Laurent G, Wahlestedt C, Kapranov P. The landscape of long noncoding RNA classification. Trends Genet. 2015;31:239–51.
Arora R, Lee Y, Wischnewski H, Brun CM, Schwarz T, Azzalin CM. RNaseH1 regulates TERRA-telomeric DNA hybrids and telomere maintenance in ALT tumour cells. Nat Commun. 2014;5:5220.
Kleaveland B, Shi CY, Stefano J, Bartel DP. A network of noncoding regulatory RNAs acts in the mammalian brain. Cell. 2018;174:350–362.e17.
Ahn J-H, Lee H-S, Lee J-S, Lee Y-S, Park J-L, Kim S-Y, et al. nc886 is induced by TGF-β and suppresses the microRNA pathway in ovarian cancer. Nat Commun. 2018;9:1166.
Bridges MC, Daulagala AC, Kourtidis A. LNCcation: lncRNA localization and function. J Cell Biol. 2021;220:e202009045. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7816648/.
Hu Y, Dong H, Huang J, Huang J, Tao D, Huang C, et al. Long non-coding RNA (lncRNA) nuclear enriched abundant transcript 1 (NEAT1) promotes the inflammation and apoptosis of otitis media with effusion through targeting microRNA (miR)-495 and activation of p38 MAPK signaling pathway. Bioengineered. 2021;12:8080–8.
Kraatz J, Clair L, Rodriguez JL, West MA. Macrophage TNF secretion in endotoxin tolerance: role of SAPK, p38, and MAPK. J Surg Res. 1999;83:158–64.
Shames BD, Selzman CH, Pulido EJ, Meng X, Meldrum DR, McIntyre RC, et al. LPS-induced NF-kappaB activation and TNF-alpha release in human monocytes are protein tyrosine kinase dependent and protein kinase C independent. J Surg Res. 1999;83:69–74.
King NE, Zimmermann N, Pope SM, Fulkerson PC, Nikolaidis NM, Mishra A, et al. Expression and regulation of a disintegrin and metalloproteinase (ADAM) 8 in experimental asthma. Am J Respir Cell Mol Biol. 2004;31:257–65.
Knolle MD, Owen CA. ADAM8: a new therapeutic target for asthma. Expert Opin Ther Targets. 2009;13:523–40.
Yamashita M, Fukui H, Sugama K, Horio Y, Ito S, Mizuguchi H, et al. Expression cloning of a cDNA encoding the bovine histamine H1 receptor. Proc Natl Acad Sci U S A. 1991;88:11515–9.
Janssens S, Beyaert R. Functional diversity and regulation of different interleukin-1 receptor-associated kinase (IRAK) family members. Mol Cell. 2003;11:293–302.
Casselbrant ML, Mandel EM, Jung J, Ferrell RE, Tekely K, Szatkiewicz JP, et al. Otitis media: a genome-wide linkage scan with evidence of susceptibility loci within the 17q12 and 10q22.3 regions. BMC Med Genet. 2009;10:85.
Stove V, Van de Walle I, Naessens E, Coene E, Stove C, Plum J, et al. Human immunodeficiency virus Nef induces rapid internalization of the T-cell coreceptor CD8alphabeta. J Virol. 2005;79:11422–33.
Iino Y, Kakizaki K, Katano H, Saigusa H, Kanegasaki S. Eosinophil chemoattractants in the middle ear of patients with eosinophilic otitis media. Clin Exp Allergy. 2005;35:1370–6.
Ingham PW, McMahon AP. Hedgehog signalling: Kif7 is not that fishy after all. Curr Biol. 2009;19:R729–31.
Jia L, Landan G, Pomerantz M, Jaschek R, Herman P, Reich D, et al. Functional enhancers at the gene-poor 8q24 cancer-linked locus. PLoS Genet. 2009;5:e1000597.
Acknowledgments
This work was supported by the National Institutes of Health (NIH)—the National Institute on Deafness and Other Communication Disorders (NIDCD) via grants R01 DC015004 and R01 DC019642 (to RS-C). NKL was supported by the T32 DC012280 grant from NIH-NIDCD (to Sue C. Kinnamon and Herman A. Jenkins). The contents of this chapter are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Lee, N.K., Santos-Cortez, R.L.P. (2023). Genetics and Otitis Media. In: Goycoolea, M.V., Selaimen da Costa, S., de Souza, C., Paparella, M.M. (eds) Textbook of Otitis Media. Springer, Cham. https://doi.org/10.1007/978-3-031-40949-3_10
Download citation
DOI: https://doi.org/10.1007/978-3-031-40949-3_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-40948-6
Online ISBN: 978-3-031-40949-3
eBook Packages: MedicineMedicine (R0)