Abstract
Auto-encoders and their variational counterparts form a family of (deep) neural networks that serve a wide range of applications in medical research and clinical practice. In this chapter we provide a comprehensive overview of how auto-encoders work and how they can be used to improve medical research. We elaborate on various topics such as dimension reduction, denoising auto-encoders, auto-encoders used for anomaly detection and the applications of representations of data created using auto-encoders. Secondly, we touch upon the subject of variational auto-encoders, explaining their design and training process. We end the chapter with small scale examples of auto-encoders applied to the MNIST dataset and a recent example of an application of a (disentangled) variational auto-encoder applied to ECG-data.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Auto-encoder
- Variational auto-encoder
- Deep learning
- Dimension reduction
- Denoising
- Anomaly detection
- Disentanglement
- Explainable AI
- ECG
1 Introduction
In this chapter the workings of auto-encoders are explained in a way that is understandable for medical researchers and clinicians who have little or no prior training in the field of artificial intelligence (AI). For the more experienced reader we provide several technical intermezzos that contain a more in depth and mathematical explanation of the subject. Furthermore, we provide several examples that show potential use cases of auto-encoders for medical research, whilst also giving a broad set of guidelines on how auto-encoders can be implemented and used by other researchers in medical AI applications.
Auto-encoders and their variational counterparts form a family of (deep) neural networks that serve a wide range of applications in medical research and clinical practice. Auto-encoders were first contemplated in the late 80s, and their popularity grew with the increase in computing power [1]. Their use cases range anywhere from signal/image denoising and anomaly detection tasks to advanced dimension reduction and complex data generation [2, 3].
Unlike most types of deep neural networks, auto-encoders are generally trained in an ‘unsupervised’ manor, meaning that only raw data, without any labels, are required to train the models. This unsupervised nature and the broad set of possible applications make auto-encoders a popular choice in various fields of medical AI research.
2 The Intuition Behind Auto-encoders
Auto-encoders can be considered a dimension reduction or compression technique. Dimension reduction techniques aim to retain as much information from a raw data input as possible into a compressed vector representation (i.e. a set of numbers). The numbers in this vector, which are often referred to as ‘latent variables’, contain (as much as possible) information about the raw data input. If a dimension reduction technique is for example applied to images of written digits (e.g. the MNIST dataset), the reduced vector form of the images may contain information about what digits the image contained, the orientation of the digit and the stroke width of the drawn digit [4]. The amount of reduction applied to the input data is usually inversely related to the amount of information that is retained in the compressed vector form. For example, if an image is reduced to only 3 numbers, a lot of information is lost, and the original cannot be accurately reconstructed. In contrast, if an image that contained 28 × 28 (= 784) pixels is reduced to a vector of 392 digits, much more information is left, albeit in a reduced form. In this context, “information” is a rather abstract concept, and depends on the goal of the user of the dimension reduction technique. For auto-encoders, the main objective is typically to enable both compression and decompression, or in other words reduce the data to such a form that the original data can be reconstructed from this compressed form. Auto-encoders therefore aim to learn the optimal (de)compression functions.
3 Principal Component Analysis
The general idea of auto-encoders has been around for decades. Traditionally the use of auto-encoders has been centered around dimensionality reduction and feature learning. For these purposes, auto-encoders are closely related to Principal Component Analysis (PCA), a technique commonly used in medical research. Both PCA and auto-encoders transform data into a lower dimensional representation, while retaining the original information as much as possible. PCA is a purely mathematical approach to dimension reduction that involves calculating the Singular Value Decomposition (SVD), and is limited to linear transformations. Conversely, (deep) auto-encoders can learn non-linear transformations. For complex data linear transformations are often insufficient for tasks such as classification and dimension reduction. Because of this (deep) auto-encoders often achieve better results than PCA. In fact, when an auto-encoder without any non-linear activations is used, the auto-encoder is likely to approximate PCA [5].
4 Methodology Behind Auto-encoders
Auto-encoders can reconstruct raw input data from extracted latent variables. We therefore make a distinction between the extraction step (i.e. encoding) and the reconstruction step (i.e. decoding). During the training of the auto-encoder, both these steps are performed in sequence. First the raw data is encoded into a set of latent variables, and then the latent variables are decoded back into the raw data form. This approach is what enables the unsupervised learning of auto-encoders, as the output of the model is effectively an approximation of the input. Meanwhile, the latent representation or compressed form of the input data, can be extracted from the middle of the network (after the encoding step). To train the model, a loss or error function is defined, which captures how well the model is doing in terms of reconstructing the original input. The model is then progressively optimized to reduce this reconstruction error.
While the exact architecture of the model may vary depending on the task and data at hand, all auto-encoder models contain a distinctive ‘bottleneck’ or funnel structure. Here the dimensionality of the data is reduced during the encoding step, and increased again during the decoding step. This bottleneck structure ensures the model is unable to simply copy information from the input to the output. Instead it has to compress the data and reconstruct it. By forcing compression of the data through the bottleneck structure and optimizing the model for accurate reconstructions, auto-encoders learn to perform complex steps that allow it to create a latent representation of the data that contains as much important information as possible. We provide a more formal explanation of this process in the technical intermezzo below.
Technical Intermezzo 1
The auto-encoder neural network is trained to ensure that its output data are the same as the input data, which is done through a funnel represented by the latent space (Fig. 1). Even though an auto-encoder is technically a single model; it is common to define the encoder step and the decoder step separately. The encoder E takes the raw data x as input and outputs a latent representation z (Eq. 1). Subsequently, decoder D takes the latent representation z as input and outputs a reconstruction of x, now called \(\hat{\user2{x}}\) (Eq. 2). The so-called latent vector z has a lower dimensionality (is smaller) than the input x and output \(\hat{\user2{x}}\), that both have the same dimensions. As per the MNIST example above, x and \(\hat{\user2{x}}\) would both be of size 28 × 28 pixels, while z is a vector of arbitrary size that is determined by the design and purpose of the auto-encoder (e.g. 1 × 2 for compression to 2 latent variables per sample or 1 × 32 for 32 latent variables per sample).
Equation 1 Function that represents the encoder part of an auto-encoder. The latent vector (z) is calculated by the encoder (E) based on the input data x.
Equation 2 Function that represents the decoder part of an auto-encoder. The output (x̂) is calculated by the decoder (D) based on the latent vector (z) that was previously calculated by the encoder.
Using this formalization, we can thus define the auto-encoder as two functions as shown above. The objective of the model is to output a reconstruction \(\hat{\user2{x}}\) that is as similar as possible to the original input x while also generating a latent representation (z) of the data after the encoding step. To enforce this similarity, a so-called loss term (or error term) is used during training of the auto-encoder. This loss term is a measure for the difference between input x and output \(\hat{\user2{x}}\). A relatively simple and commonly used function to calculate the loss is the mean squared error (MSE). The loss calculation of the model can then be represented by the following function:
Equation 3 Function to calculate the mean squared error (MSE) loss of the input data x and output data x̅. N = total number of data point in data, i = ith data point in the dataset.
5 Auto-encoders for Denoising and Anomaly Detection
In this section we will provide some use cases for auto-encoders. The first example of a potential use-case is that of denoising data. In the field of medical imaging, the presence of noise in images may limit resolution or decrease interpretability, thereby hampering it’s use for evaluation or further analysis. Therefore, removing noise (i.e. denoising) is commonly performed as a first step. Conventional methods for denoising (medical) images ranges from spatial filters, such as Gaussian or convolutional filters to wavelet based techniques [6]. As described before, auto-encoders can also be used for denoising images. Recent studies have shown that auto-encoder based denoising methods often outperform conventional methods. Gondara showed that using convolutional layers in an auto-encoder led to efficient denoising of medical images, and maybe more importantly, can be used on smaller datasets [7].
Auto-encoders extract information from the input and reconstruct the input data as good as possible. We can use this characteristic to create an auto-encoder that extracts information from a noisy input and reconstructs the input but without the noise. We do this under the assumption that a noisy image is composed of a clean image with noise added to it. We thus want to train the auto-encoder such that it extracts the important information of the clean image but ignores the noise. In order to do so we start with a, non-noisy, input x and add some random noise λ to it. We thus have a new input for the model, which we will call x*, that is the sum of x and λ (e.g. x* (noisy image) = x (image) + λ (noise)). We pass this noisy input through the network and obtain \(\hat{\user2{x}}\), the reconstructed image, as we did before. Meanwhile, we keep the original MSE loss calculation fixed, so it is still the difference between x and \(\hat{\user2{x}}\), however, \(\hat{\user2{x}}\) is now based on the noisy input x* rather than x. The network will thus have to learn how to remove the noise from \(\hat{\user2{x}}\) in order to make it as similar as possible to x.
Denoising auto-encoders can be a useful tool to clean data that stems from real world observations that tend to be very noisy. Lu et al., for example, use denoising auto-encoders to enhance speech recordings [8]. Jifara et al. take a slightly different approach and design their auto-encoder in such a way that it outputs the estimated noise, instead of a reconstruction of the input image (the noise can be subtracted from the noisy image to create a clean image) [9]. They show that this approach improves upon standard denoising auto-encoders on images obtained using chest radiography. Nawarathne et al. use denoising auto-encoders on spectral images extracted from accelerometric data measured on pregnant mothers’ abdomen, in order to improve the analysis of fetal motion during pregnancy [10].
Auto-encoders can also be used as a fully unsupervised method of anomaly detection. For these applications, it is important to understand that auto-encoders only learn to reconstruct data that they have seen during the training of the network. While a network may learn to handle slight differences, it likely performs worse on samples that are very different from the training data. To illustrate this using the MNIST (a dataset containing images of hand drawn digits) example; if a network is only trained on images of the digit 3, it will fail to properly reconstruct the digit 7. Interestingly, we can use this property to detect anomalies or outliers in the dataset, by purposefully training the network on a dataset of which we are certain does not contain any anomalous samples. If we then apply the network to another dataset that does contain outliers, the outliers are likely to have a significantly larger reconstruction error than the non-anomalous samples. It must be kept in mind that all data that is different from that in the training set is considered anomalous. It may therefore be very hard to distinguish between expected anomalous data, and noise in the observations.
Shvetsova et al. show that this approach can be used to detect tissue with metastases in H&E-stained lymph nodes and abnormal chest x-rays [11]. Wei et al. use a similar method to detect suspicious areas in mammograms showing how auto-encoders can also be used to detect the position of the anomaly in an image while only requiring a set of images obtained from healthy ‘normal’ patients [12].
6 Auto-encoders for Latent Vector and Feature Learning
Perhaps the most interesting applications of auto-encoders are based on the latent vector extracted after the encoding step. The latent vectors contain a condensed form, or a summary, of all the important information in the input data. Exactly what that information is however, is unknown. We only know that the latent vector contains information that the decoder can use to reconstruct the original data. An important aspect of auto-encoders is that they do not guarantee that the latent space is normally distributed. What this means is that we may get unexpected results when we reconstruct samples after manipulating latent representations or when we calculate relationships between latent representations of different samples. For instance, one might expect that two similar looking images yield similar latent vectors when passed through the encoder. However, it is entirely possible that two very different images have a very similar latent vector, while two very similar images have very different latent vectors. An example of this is given in Fig. 4 where we can see that if we look at some MNIST images that are similar in terms of their latent representation, that some of the original non-compressed images are in fact very different. The fact that the latent space of the auto-encoder is not normally distributed also hampers us from directly linking the values in the latent representations to underlying features of the data. In the case of the MNIST example we may for example observe an increase in line-width if we increase the first latent variable of a latent representation by +2 and reconstruct the image. It is however possible that a step of +5 yields a reconstruction in which the digit is rotated instead of a reconstruction where the linewidth is increased further. Variational auto-encoders, discussed later in this chapter, try to enforce a normally distributed latent space which enables a wide range of additional applications.
While the latent representations of auto-encoder are limited by the non-linearity of the latent space they can still be used for a number of applications. The created latent vectors may for example serve as input to other models [13]. If a user has a very large dataset, of which only a small fraction is labeled, it may be beneficial to first train an auto-encoder on the full dataset, and then train a separate classifier on the latent representations of the previously labelled dataset. This approach ensures that sufficient information is extracted from the input data, with less risk of overfitting and unwanted biases.
It is also possible to use the latent vectors as input for another dimension reduction technique that is better at preserving the relationship between samples, but worse at handling large/complex data [14]. It is for example not uncommon to reduce image data to 32 or 64 dimensions using an auto-encoder and then apply t-SNE (or similar dimension reduction techniques) to further reduce the dimension to 2 or 3, so that the data can easily be visualized in a graph [15]. This approach generally performs better than only using an auto-encoder or t-SNE.
7 Variational Auto-encoders
Variational auto-encoders (VAE) are closely related to auto-encoders in terms of their network structure and purposes [16]. The main goal with which they were proposed is however very different from the original ‘vanilla’ auto-encoders. VAEs are a type of generative model, meaning that they can generate (new) data, instead of just compressing and reconstructing existing data. In order to do so, VAEs attempt to learn the distribution (or process) that generated the data on which the model is trained, opposed to simply finding the optimal solution that minimizes reconstruction loss. The latent space variables of regular auto-encoders may have large gaps in their distribution and may be centered around an arbitrary value, while those of VAEs are all normally distributed with a mean of 0 and standard deviation of 1 (stochastic normal distribution). In the case of the latent space of an auto-encoder, there is little relation between values in the latent space and its reconstruction, slightly changing z might lead to completely different reconstructions. With the VAE, there is a very direct relation between the two and slightly changing z will slightly alter the reconstruction while changing z in the opposite direction will have the opposite result. By inserting a latent vector z (with values around zero, and within a few standard deviations) into the decoder of a VAE, one can create ‘new’ data that can usually be considered comparable to the data the VAE was trained on, where a latent vector z containing all zeros approximates the mean of the training data. The general structure of a VAE is visualized in Fig. 2.
General schematic layout of a Variational Auto-encoder neural network. During the training the network latent vector z is sampled from a gaussian distribution parameterized by the outputs of the encoder. These outputs are also used for the calculation of the KL-divergence, which is then combined with the MSE loss (calculated from the original input and the reconstruction) to form the VAE loss function
The training of VAEs is more complex than that of normal auto-encoders, and is described in more detail in the technical intermezzo. It is important to know that VAEs are trained with an additional loss term: the Kullback-Leiber Divergence (KL Divergence). The KL-divergence loss term encourages the latent space of the VAE to have the desired properties described above by enforcing that each individual latent variable follows a unit normal gaussian distribution (with mean = 0 and standard deviation = 1).
Technical Intermezzo 2
VAEs are based on the assumption that all data in the dataset used to train the model was generated from a process involving some unobserved random variable. The data generation process then consists of 2 steps: (1) a value z is generated from some prior distribution \({\varvec{P}}_{{\user2{\theta *}}} \left( {\varvec{z}} \right)\); ; (2) a value x is generated from a conditional distribution \({\varvec{P}}_{{\user2{\theta *}}} ({\varvec{x}}|{\varvec{z}})\). In this process the optimal values of θ(θ*) and z are unknown, and thus need to be calculated from the known values in x. VAEs aim to approximate θ∗ and z even if calculation of the marginal likelihood and true posterior density are intractable. To do so, VAEs use a recognition model \({\varvec{q}}_{\user2{\varphi }} ({\varvec{z}}|{\varvec{x}})\) that approximates the true posterior \({\varvec{P}}_{{\user2{\theta *}}} ({\varvec{x}}|{\varvec{z}})\) and jointly learn the recognition parameter φ together with the generative parameter θ. Using this formalization, we can distinguish between learning a probabilistic encoder \({\varvec{q}}_{\user2{\varphi }} ({\varvec{z}}|{\varvec{x}})\), from which we can sample z when given x and a probabilistic decoder \({\varvec{P}}_{{\varvec{\theta}}} ({\varvec{x}}|{\varvec{z}})\), from which we can sample x when given z. In practice both the probabilistic encoder and decoder are neural networks of which the appropriate architecture can be picked based on the nature of the data in x.
The VAE training objective
To ensure that the approximate distribution \({\varvec{q}}_{\user2{\varphi }} ({\varvec{z}}|{\varvec{x}})\), is close to the real distribution \({\varvec{P}}_{{\varvec{\theta}}} ({\varvec{x}}|{\varvec{z}})\), , we can use the Kullback-Leiber Divergence (KL Divergence) which quantifies the deference between 2 distributions. In the case of VAEs the goal is to minimize this KL Divergence which can be written as follows:
Equation 4 . The Kullback-Leiber Divergence.
Equation 4 can then be rearranged to Eq. 5.
The left-hand side of Eq. 5 exactly fits the objective of the VAE: we want to maximize the probability of x from distribution pθ(x) and minimize the difference between the estimated distribution qφ(z|x) and real distribution pθ(z|x). The negation of the right-hand side of the equation gives us the loss which we minimize to find the optimal values for φ and θ.
Equation 6 . The training objective function of the variational auto-encoder.
Equation 6 is known as the Evidence Lower Bound (ELBO) because the KL-divergence is always positive. This means that − \(L_{VAE}\) is the lowest value pθ(x) can take, minimizing \(L_{VAE}\) thus equates to maximizing pθ(x). Even tough Eq. 6 gives a clear definition of a loss term, it cannot directly be used to train a VAE. The expectation term in the loss has to be approximated using a sampling operation, which prevents the flow of gradients during training. To solve this issue, VAEs use the ‘reparameterization trick’ which relies on the assumption that p(z|x) follows a known distribution. This distribution is usually assumed to be a multivariate Gaussian with a diagonal covariance structure (even though the trick works for other distributions as well). Using the parameters of qφ(x|z) and the assumption qφ(x|z) is Gaussian, z can be expressed as a deterministic variable that is produced by some function τφ(x, ε) where ε is sampled form an independent unit normal Gaussian distribution.
Equation 7 . The ‘reparameterization trick’ used to enable the training of variational auto-encoders through backpropagation.
In practice the encoder model of the VAE is constructed so that is outputs a mean (µ) and standard deviation (σ) that parameterize the Gaussian distribution qφ(x|z). Using this set up, the reparameterization trick equates to Eq. 7.
In this chapter we often refer to the embedding or latent representation of data which means the mean µ output of the encoder of the VAE was used and the standard deviation σ was ignored. This can be considered standard practice if a latent representation of input data is desired.
8 Disentanglement and Posterior Collapse
The latent variables of a VAE often encode some underlying characteristics of the data. For images, latent variables can for example encode factors such as the width, height or angle of a shown object [17]. However, different latent variables are often entangled, meaning that multiple variables influence the same characteristic of the data. To improve the explainability of the latent space and better control the generative process of the VAE [18,19,20,21] it can be desirable to disentangle the latent space. Higgins et al. proposed the β-VAE, which adds an additional weight β to the KL-term of the VAE loss, as a very simple but effective way to improve disentanglement [17]. The value of β can be picked based on the desired amount of disentanglement of the latent space. A higher β generally corresponds to better disentanglement. There is however a trade-off between the amount of disentanglement and the reconstruction quality of the VAE, where more disentanglement results in worse reconstructions [22]. VAEs also suffer from a phenomenon called posterior collapse (or KL-vanishing), which causes the model to ignore a subset of the latent variables. Posterior collapse occurs when the uninformative prior distribution matches the variational distribution too closely for a subset of latent variables. This is likely caused by the KL-divergence loss term which encourages the two distributions to be similar [23]. During training, posterior collapse can often be observed when the KL-loss term decreases to (near) zero, which is even more prevalent in VAE variants that add additional weight to the KL-term such as β-VAE [17]. To prevent posterior collapse and improve reconstruction quality of disentangled VAEs, Shao et al. propose the Control-VAE [24]. This method requires a ‘target value’ for the KL-divergence and tunes the weight of the KL-divergence such that it stays close the target value during training.
9 Use Cases for VAEs and Latent Traversals
The generative capabilities and their (disentangled) latent spaces allow for a large number of use-cases of VAEs. VAEs (and VAE based models) can for example be used to improve anomaly detection compared to normal auto-encoders, to create interpretable latent representations that can serve as input for conventional classification models such as logistic regressions, or to perform further analysis of the learned latent variables using techniques such as latent traversals [25, 26].
A latent traversal is a method in which we change one or more latent variables from a sample encoded using the encoder of a VAE, and reconstruct the input sample from these changed latent variables using the decoder. By comparing the original sample and the sample reconstructed from the changed variables one can see which aspects of the data are encoded by these variables. Especially when the latent space is sufficiently disentangled, it is often possible to relate individual latent variables to underlying physiological characteristics of the data.
Latent traversals can be combined with logistic regressions (or other classical statistical models) to infer and visualize relationships between latent variables and the use case (e.g. classification, prediction etc.). We do this by analyzing the weights/coefficients of the logistic regression to see which latent variables have a positive predictive value for a certain class. We can then perform a latent traversal by increasing and decreasing these important latent variables and examining how the reconstructed sample changes. This whole process thus allows us to visualize which features are important for a class. We elaborate on this approach in a practical example applied to electrocardiogram (ECG) data later in this chapter.
10 Auto-encoders Versus Variational Auto-encoders (Summary)
Now that we have discussed both auto-encoders and variational auto-encoders, we can summarize the pros and cons of both model types. An overview of these is given in Table 1. In general, VAEs provide a wider range of applications, while auto-encoders generally produce better reconstructions. We have discussed a similar trade-of regarding the disentanglement of VAEs, where the reconstruction quality of VAEs is inversely related to the amount of disentanglement. These trade-offs lead to the conclusion that it is desirable to use a (disentangled) VAE if a normal auto-encoder is insufficient for the desired use-case.
11 Designing an Auto-encoder and Common Pitfalls
The first step in training an auto-encoder (or any other model) is collecting a representative dataset that can ensure the validity of any findings or insights [27]. As discussed before, auto-encoders only learn to reconstruct data that is similar to the data used during the preceding training phase. It is thus important to collect a heterogeneous dataset that spans the full range of sample variation that will be used for further analysis. The actual type of data can range anywhere from images, to signals to any arbitrary measurement. There is, to the best of our knowledge, no datatype that can inherently not be used to train an auto-encoder. It is however important to remember that more complex data may require a more complex network architecture, or more training data. It is also possible that the standard MSE loss term may not be adequate for certain datatypes where it is important to accurately reconstruct small features, because the MSE loss will deem large features to be more important than small features. An example of this is in the use-case of ECGs, where minor variations in the P-wave can be overshadowed by larger variations in the larger T-wave, and are thus not adequately captured by the auto-encoder.
Both the encoder and decoder part of the auto-encoder consist of a more elaborate neural network. The choice for the network architecture is generally dependent on the data to which the auto-encoder is applied. For simpler data it may be sufficient to use a small number of fully connected linear layers, in combination with non-linear activation functions [28]. For more complex data, such as for example medical images, the encoder network is often composed of several convolutional layers (connected by non-linear activation functions) [7, 9,10,11]. Convolutions are currently the most popular architecture type because they show optimal performance on various types of different data. For signal or timeseries data, 1-dimsional convolutions are a popular choice; for images it is common to use 2-dimensional convolutions [8]. Depending on the number of chosen layers it may also be beneficial to add skip connections (residual connections) to improve the flow of gradients through the network during backpropagation [29]. Various regularization techniques like batch normalization and dropout may also improve performance. It is however generally better to first design a simple network and be certain that these additional tricks improve performance before using them.
In essence, the decoder of the network is often designed to be a mirrored version of the encoder network. Hence, if convolutional layers are used in the encoder, transposed convolutions are used in the decoder [30]. The usage of pooling layers (e.g. min/max-pool, average pool) in the encoder may pose a problem, as no sufficient inverse of these functions exist. In this case it is possible to simply up sample the data in the decoder under the assumption that the model will be expressive enough trough the other layers that do contain weights.
Perhaps the most important design decision is the size of the latent space. Smaller latent vectors generally result in worse reconstructions, conversely larger latent vectors often lead to better reconstructions. The choice of the size of the latent space is thus very dependent on the use case of the auto-encoder. For denoising auto-encoders and anomaly detection tasks, it may be sufficient to reduce the size of the input only slightly during the encoding step. In these cases a very small latent vector is undesirable as it is likely to yield worse reconstructions. By contrast, if the latent representation serves as input for another model, picking the correct size is entirely dependent on the task of the other model. Here a more compressed representation may be desirable as it reduces the amount of information extraction that still must be performed by the other model. If the latent representations are used as input for conventional clustering techniques it is desirable to have an amount of latent variables that is within a reasonable range (e.g. higher than 10 but below 100, dependent on the size of the dataset). When auto-encoders serve as an input to another neural model, the optimal latent space can be selected based on the quality of the reconstructions (if we assume the decoder is functioning perfectly). Simply put, if the reconstructions look decent, there must be enough information in the latent representation to be used in the other model, and the latent space was sufficiently large.
If the goal of the auto-encoder is to create an interpretable latent space, the best choice is likely to use a variational auto-encoder.
12 Examples Using the MNIST Dataset
In this section we perform a number of small-scale experiments to show the how the design of the auto-encoder influences its performance. For this purpose we use the MNIST dataset [4]. This dataset consists of 70,000 grayscale images of handwritten digits and is a popular choice for basic experiments among AI researchers.
We split the dataset into a train, validation and test set (80%, 10%, 10% respectively) and trained 9 neural networks until convergence. As a comparison, we also included 3 examples of the commonly used PCA dimension reduction technique. The tested neural architectures consist of a fully connected (linear) architecture without activation functions, a fully connected architecture with activation functions, and a convolutional neural network. For each architecture we train the network with 3 different latent space sizes (2, 4 and 8 latent variables). We configure the PCA method to also reduce the data to 2, 4 and 8 variables.
All images in the dataset consist of 28 × 28 pixels. Depending on the model architecture we treat the pixel values of the image as either a vector or a matrix. For the fully connected architectures, as well as the commonly used PCA method, we flatten the input 28 × 28 image, resulting in a vector of 1 × 784 pixel values. For a convolutional architecture we keep the image in its original matrix form so that convolutions can better capture the spatial relationships between the pixels in the images, in all directions (i.e. horizontal or vertical).
In Fig. 4 we show the reconstructions of a sample for each of the methods and each tested latent space size. The results clearly show that the reconstruction quality increases as more latent variables are used. We also observe the difference in quality between the different architectures. The fully connected models, without linear activation functions, show the worst results, which are even worse than the PCA method. This is expected, as a linear network is likely to only approximate PCA. The non-linear models, both fully connected and convolutional, show the best results, with the convolution network performing slightly better than the fully connected network. Here the strength of convolutional models becomes clear, as the convolutional networks outperform the fully connected networks while having significantly less parameters (approximately 270,000 for the convolutional networks versus 400,000 for the fully connected networks). We thus see that convolutional neural networks can outperform fully connected neural networks despite having less parameters. This difference in the number of parameters generally causes convolutional networks to be more computationally efficient and converge faster. Additionally, this reduction in computational cost may allow us to further increase the depth/size of the network and potentially improve its performance further (Fig. 3).
In order to highlight the fact that auto-encoders do not preserve the relationship between input samples in the latent space, an additional example is provided. We encode a sample image, as well as the rest of the training dataset, to its latent representation, and look for the images that are closest to the sample image in the latent space. We plot the top 5 closest images in Fig. 5, and observe that images 3, 4 and 5 are not similar to our sample image at all (Fig. 4).
We also compare the spread of the values of the latent space of auto-encoders and variational auto-encoders (Fig. 6) to show the differences between both models. To do so we first construct a variational auto-encoder with a latent space of 8 values that uses a similar convolutional architecture as the normal auto-encoder. We than encode all the entries in the training set into their latent representation and create a boxplot for each latent variable. We observe that for the normal auto-encoder the latent variables have mean values that deviate from 0, have larger standard deviations, larger confidence intervals and that the mean value of the variables is often not located at the center of the confidence interval. For the variational auto-encoder we observe that each latent variable does indeed appear to be normally distributed, as was enforced during the training of the VAE.
Illustration of the FactorECG explainable pipeline for ECG interpretation. The VAE consists of three parts, the encoder, the latent space (FactorECG) and the decoder. The model can be made explainable locally (as the individual values of the ECG factors for each ECG are known) and globally (by using factor traversals the influence of individual factors on the ECG morphology can be visualized). Usually, the factors are entered into simple statistical models, such as logistic regression, to perform the task at hand
13 Demonstrator Use Case of an VAE for the Electrocardiogram: The FactorECG
Many studies use deep neural networks to interpret electrocardiograms (ECGs) with high predictive performances, some focusing on tasks known to be associated with the ECG (e.g., rhythm disorders) and others identifying completely novel use cases for the ECG (e.g., reduced ejection fraction) [31,32,33,34]. Most studies do not employ any technique to provide insight into the workings of the algorithm, however, the explainability of neural networks can be considered a essential step towards the applicability of these techniques in clinical practice [35, 36]. In contrast, various studies do use post-hoc explainability techniques, where the ‘decisions’ of the ‘black box’ DNN are visualized after training, usually using heatmaps (e.g.., using Grad-CAM, SHAP or LIME) [37]. In these studies, usually some example ECGs were handpicked, as these heatmap-based techniques only work on single ECGs. Currently employed post-hoc explainability techniques, usually heatmap-based, have limited explainable value as they merely indicate the temporal location of a specific feature in the individual ECG. Moreover, these techniques have been shown to be unreliable, poorly reproducible and suffer from confirmation bias [38, 39].
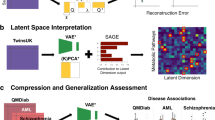
Variational auto-encoders can be used to overcome this by constructing a DNN that is inherently explainable (i.e. explainable by design, instead of investigating post-hoc). One example is the FactorECG, which is part of a pipeline that consists of three components: (1) a variational auto-encoder that learned to encode the ECG into its underlying 21 continuous factors of variation (the FactorECG), (2) a visualization technique to provide insight into these ECG factors, and (3) a common interpretable statistical method to perform diagnosis or prediction using the ECG factors [19]. Model-level explainability is obtained by varying the ECG factors (i.e. latent traversals), while generating and plotting ECGs, which allows for visualization of detailed changes in morphology, that are associated with physiologically valid underlying anatomical and (patho)physiological processes. Moreover, individual patient-level explanations are also possible, as every individual ECG has its representative set of explainable FactorECG values, of which the associations with the outcome are known. When using the explainable pipeline for interpretation of diagnostic ECG statements, detection of reduced ejection fraction and prediction of one-year mortality, it yielded predictive performances similar to state-of-the-art ‘black box’ DNNs. Contrary to the state-of-the-art, our pipeline provided inherent explainability on which ECG features were important for prediction or diagnosis. For example, ST elevation was discovered to be an important predictor for reduced ejection fraction, which is an important finding as it could limit the generalizability of the algorithm to the general population. We have also extended the FactorECG methodology and developed a technique called Query based Latent Space Traversals (qLST) which can be used to relate multiple latent variables to a disease class at once or to explain existing black box classifiers [15].
A longstanding assumption was that the high-dimensional and non-linear ‘black box’ nature of the currently applied ECG-based DNNs was inevitable to gain the impressive performances shown by these algorithms on conventional and novel use cases. Variational auto-encoders allow for reliable clinical interpretation of these models without performance reduction, however, while also broadening their applicability to detect novel features in many other (rare) diseases, as they provide significant dimensionality reduction. The application of such methods will lead to more confidence in DNN-based ECG analysis, which will facilitate the clinical implementation of DNNs in routine clinical practice.
References
Hinton GE, Zemel RS. Autoencoders, minimum description length and Helmholtz free energy. p. 8.
Hinton GE, Salakhutdinov RR. Reducing the dimensionality of data with neural networks. Science. 2006;313(5786):504–7. https://doi.org/10.1126/science.1127647.
Using autoencoders for mammogram compression. PubMed. https://pubmed.ncbi.nlm.nih.gov/20703586/. Accessed 31 Jan 2022.
Lecun Y, Bottou L, Bengio Y, Haffner P. Gradient-based learning applied to document recognition. Proc IEEE. 1998;86(11):2278–324. https://doi.org/10.1109/5.726791.
Baldi P, Hornik K. Neural networks and principal component analysis: learning from examples without local minima. Neural Netw. 1989;2(1):53–8. https://doi.org/10.1016/0893-6080(89)90014-2.
Mohd Sagheer SV, George SN. A review on medical image denoising algorithms. Biomed Signal Process Control 2020;61:102036. https://doi.org/10.1016/j.bspc.2020.102036.
Gondara L. Medical image denoising using convolutional denoising autoencoders. In: 2016 IEEE 16th international conference on data mining workshops (ICDMW), Barcelona, Spain, Dec 2016, pp 241–46. https://doi.org/10.1109/ICDMW.2016.0041
Lu X, Tsao Y, Matsuda S, Hori C. Speech enhancement based on deep denoising auto-encoder. In: Proceedings of interspeech, Jan 2013. p. 436–40.
Jifara W, Jiang F, Rho S, Cheng M, Liu S. Medical image denoising using convolutional neural network: a residual learning approach. J Supercomput. 2019;75(2):704–18. https://doi.org/10.1007/s11227-017-2080-0.
Nawarathne T et al. Comprehensive study on denoising of medical images utilizing neural network based auto-encoder. Feb 2021. arXiv:2102.01903 [eess]. Accessed 30 Jan 2022. Available http://arxiv.org/abs/2102.01903
Shvetsova N, Bakker B, Fedulova I, Schulz H, Dylov DV. Anomaly detection in medical imaging with deep perceptual autoencoders. IEEE Access. 2021;9:118571–83. https://doi.org/10.1109/ACCESS.2021.3107163.
Wei Q, Shi B, Lo JY, Carin L, Ren Y, Hou, R. Anomaly detection for medical images based on a one-class classification. In: Medical imaging 2018: computer-aided diagnosis, Houston, United States, Feb 2018. p 57. https://doi.org/10.1117/12.2293408
Hinton GE, Krizhevsky A, Wang SD. Transforming auto-encoders. In: Artificial neural networks and machine learning—ICANN 2011, Berlin, Heidelberg, 2011. p. 44–51.
Fabius O, van Amersfoort JR. Variational Recurrent auto-encoders. arXiv:1412.6581 [cs, stat], Jun 2015. Accessed 31 Jan 2022. Available http://arxiv.org/abs/1412.6581
Vessies MB et al. Interpretable ECG classification via a query-based latent space traversal (qLST). arXiv:2111.07386 [cs, eess], Nov 2021, Accessed 31 Jan 2022. Available http://arxiv.org/abs/2111.07386
Kingma DP, Welling M. Auto-encoding variational Bayes. arXiv:1312.6114 [cs, stat], May 2014. Accessed 30 Jan 2022. Available http://arxiv.org/abs/1312.6114
Higgins I et al.: β-VAE: learning basic visual concepts with a constrained variational framework. 2017. p. 22.
Van Steenkiste T, Deschrijver D, Dhaene T. Generating an explainable ECG beat space with variational auto-encoders. arXiv:1911.04898 [cs, eess, stat], Nov 2019. Accessed 30 Jan 2022. Available http://arxiv.org/abs/1911.04898
van de Leur RR et al. Inherently explainable deep neural network-based interpretation of electrocardiograms using variational auto-encoders. Cardiovasc Med. 2022;preprint. https://doi.org/10.1101/2022.01.04.22268759.
Kim J-Y, Cho S. BasisVAE: orthogonal latent space for deep disentangled representation. Sep 2019. Accessed 30 Jan 2022. Available https://openreview.net/forum?id=S1gEFkrtvH
Chen RTQ, Li X, Grosse R, Duvenaud D. Isolating sources of disentanglement in variational autoencoders. arXiv:1802.04942 [cs, stat], Apr 2019. Accessed 30 Jan 30 2022. Available http://arxiv.org/abs/1802.04942
Asperti A, Trentin M. Balancing reconstruction error and Kullback-Leibler divergence in variational autoencoders. arXiv:2002.07514 [cs], Feb 2020. Accessed 30 Jan 2022. Available http://arxiv.org/abs/2002.07514
Lucas J, Tucker G, Grosse R, Norouzi M. Understanding posterior collapse in generative latent variable models. 2019. p. 16.
Shao H et al. Control VAE: controllable variational autoencoder. arXiv:2004.05988 [cs, stat], Jun 2020. Accessed 30 Jan 2022. Available: http://arxiv.org/abs/2004.05988
Guo X, Gichoya JW, Purkayastha S, Banerjee I. CVAD: a generic medical anomaly detector based on Cascade VAE. arXiv:2110.15811 [cs, eess], Jan 2022. Accessed 30 Jan 2022. Available http://arxiv.org/abs/2110.15811
Cakmak AS et al. Using convolutional variational autoencoders to predict post-trauma health outcomes from actigraphy data. arXiv:2011.07406 [cs, eess], Nov 2020. Accessed 25 Jan 2022. Available http://arxiv.org/abs/2011.07406
Ministerie van Volksgezondheid WS. Guideline for high-quality diagnostic and prognostic applications of AI in healthcare—Publicatie - Data voor gezondheid. 28 Dec 2021. https://www.datavoorgezondheid.nl/documenten/publicaties/2021/12/17/guideline-for-high-quality-diagnostic-and-prognostic-applications-of-ai-in-healthcare. Accessed 01 Feb 2022.
Wang Y, Yao H, Zhao S. Auto-encoder based dimensionality reduction. Neurocomputing. 2016;184:232–42. https://doi.org/10.1016/j.neucom.2015.08.104.
He K, Zhang X, Ren S, Sun J. Identity mappings in deep residual networks. In: Computer vision—ECCV, Cham. 2016. p. 630–45. https://doi.org/10.1007/978-3-319-46493-0_38.
Chen H, et al. Low-dose CT with a residual encoder-decoder convolutional neural network. IEEE Trans Med Imaging. 2017;36(12):2524–35. https://doi.org/10.1109/TMI.2017.2715284.
van de Leur RR, et al. Automatic triage of 12-lead ECGs using deep convolutional neural networks. J Am Heart Assoc. 2020;9(10): e015138. https://doi.org/10.1161/JAHA.119.015138.
Attia ZI, et al. Screening for cardiac contractile dysfunction using an artificial intelligence-enabled electrocardiogram. Nat Med. 2019;25(1):70–4. https://doi.org/10.1038/s41591-018-0240-2.
van de Leur RR, et al. Discovering and Visualizing disease-specific electrocardiogram features using deep learning: proof-of-concept in phospholamban gene mutation carriers. Circ Arrhythm Electrophysiol. 2021;14(2): e009056. https://doi.org/10.1161/CIRCEP.120.009056.
Ribeiro AH, et al. Automatic diagnosis of the 12-lead ECG using a deep neural network. Nat Commun. 2020;11(1):1760. https://doi.org/10.1038/s41467-020-15432-4.
Goodman B, Flaxman S. European Union regulations on algorithmic decision-making and a ‘right to explanation.’ AIMag. 2017;38(3):50–7. https://doi.org/10.1609/aimag.v38i3.2741.
Markus AF, Kors JA, Rijnbeek PR. The role of explainability in creating trustworthy artificial intelligence for health care: a comprehensive survey of the terminology, design choices, and evaluation strategies. J Biomed Inform. 2021;113: 103655. https://doi.org/10.1016/j.jbi.2020.103655.
Hughes JW, et al. Performance of a convolutional neural network and explainability technique for 12-lead electrocardiogram interpretation. JAMA Cardiol. 2021;6(11):1285–95. https://doi.org/10.1001/jamacardio.2021.2746.
Hooker S, Erhan D, Kindermans P-J, Kim B. A benchmark for interpretability methods in deep neural networks. arXiv:1806.10758 [cs, stat], Nov 2019. Accessed 31 Jan 2022. Available http://arxiv.org/abs/1806.10758
Adebayo J, Gilmer J, Muelly M, Goodfellow I, Hardt M, Kim B. Sanity checks for saliency maps. arXiv:1810.03292 [cs, stat], Nov 2020, Accessed 31 Jan 2022. Available http://arxiv.org/abs/1810.03292
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Glossary
- Activation function
-
In neural networks, (non-linear) activation functions are used at the output of neurons to convert the input to an ‘active’ or ‘not active’ state. An activation function can be a simple linear or sigmoid function or have more complex arbitrary forms. The Rectified Linear Unit (ReLU) function is currently the most popular choice.
- Back propagation
-
Is a widely used technique in the field of machine learning that is used during the training of a neural network. The technique is used to update the weights of the neural network based on the calculated loss, effectively allowing it to ‘learn’.
- (mini-) Batch
-
A small set of data samples that is fed through the network at once during training. A too small batch size may lead to instability while a too large batch size may lead to depletion of computer resources.
- Convolution
-
Common building block of various neural networks. Convolutional neural networks can be considered the current ‘state of the art’ of neural networks applied to various data sources. Convolutional layers in a neural network a apply a learned filter to the input data which improves the ability of neural networks to comprehend spatial structures. Convolutions can be applied in 1 dimensional (signal/timeseries data) and 2 dimensional (images) forms.
- Decoder
-
Part of the (variational) auto-encoder that decodes the given latent vector into a reconstruction of the original data
- Dimension
-
The dimension of data is the size of the dataset or vector, for a grayscale image this is the height × the width in pixels (e.g. 28 × 28), for an RGB-color image, a third dimension of size 3 is added (e.g. (28 × 28 × 3)
- Encoder
-
Part of the (variational) auto-encoder that encodes the provided data into the latent vector
- Explainability
-
The ability of a (trained) observer to interpret the inner workings of a model. Neural networks are generally considered to be to complex to comprehend by humans and are treated as an ‘unexplainable’ black box. The lack of explainability is a major issue in many of the current clinical applications of neural networks.
- Fullyconnected or linear layer
-
Common building block of neural networks in which every node (or every datapoint) in the input is connected to every node in the output of the layer. Through the weights that are associated with each connection the layer is able perform linear transformations of the input data. Together with non-linear activation functions, fully connected layers make up the most basic forms of neural networks.
- KL Divergence
-
The Kullback-Leiber Divergence is a measure of similarity between two distributions.
- Loss function
-
The loss function of the network defines the training objective of the neural network. The loss, the output of the loss function, is progressively minimized through backpropagation, allowing the network to learn and be optimized for its training objective.
- MNIST
-
A commonly used dataset consisting of image of handwritten digits. MNIST is often used for small scale experiments because of the simplistic nature of the data.
- PCA
-
Principal component analysis. A technique commonly used for dimension reduction. The technique involves the calculation of
- Posterior collapse
-
A phenomenon that can occur during the train of variational autoencoder through which the reconstruction accuracy of the network decreases dramatically if the KL-divergence reduces to much.
- Vector
-
A vector is a single row or column of numbers.
- Matrix
-
A set consisting of multiple rows and columns of numbers.
- Convergence
-
A neural network has reached convergence when further training does no longer improve the model.
- MSE loss
-
Mean Squared Error loss, a measure of difference between two data instances such as images or timeseries. The MSE loss is a common loss function that is used to minimize the reconstruction error in auto-encoders.
- Latent variable
-
A variable that is not directly observed in the data but can be inferred through the usage of a model from other variables that are observed directly. In the case of auto-encoders we refer to the variables in the vector extracted after applying the encoder of the auto-encoder as latent variables.
- Disentanglement
-
The disentanglement of latent variables refers to the process of separating the influence of each latent variable on the reconstructed data.
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Vessies, M., van de Leur, R., Wouters, P., van Es, R. (2023). Deep Learning—Autoencoders. In: Asselbergs, F.W., Denaxas, S., Oberski, D.L., Moore, J.H. (eds) Clinical Applications of Artificial Intelligence in Real-World Data. Springer, Cham. https://doi.org/10.1007/978-3-031-36678-9_13
Download citation
DOI: https://doi.org/10.1007/978-3-031-36678-9_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-36677-2
Online ISBN: 978-3-031-36678-9
eBook Packages: MedicineMedicine (R0)