Abstract
RNA modifications contribute to the various functions of RNAs in all living organisms. Some of these modifications are dynamic and contribute to the regulation of gene expression. In bacteria, their roles in stress, environmental adaptation, and in infections caused by pathogens have been recently fully recognized. In this review, we describe several methodologies including mass spectrometry, next-generation RNA sequencing methods, biochemical approaches, and cryo-EM structural analysis that are used to detect and localize the modifications in tRNAs and rRNAs. We illustrate how the combination of methods was necessary to avoid technical biases for a successful mapping of the modifications in tRNAs and rRNAs in Staphylococcus aureus.
R. Bahena-Ceron and J. Jaramillo-Ponce contributed equally in this chapter.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
RNA molecules are chemically modified at different positions of the nitrogenous bases or the ribose cycle. These modifications are common to all domains of life and are present in all RNA classes, including stable non-coding RNAs (tRNAs, rRNAs, and some regulatory RNAs) and generally less stable mRNAs. They involve covalent alterations (i.e., methylations, acetylation, and deamination) or isomerization (i.e., pseudouridines) of nucleotides (Fig. 1a). To date, over 170 modifications have been found in RNAs from different organisms, with the most common modification being methylation (see MODOMICS database) (Boccaletto et al. 2022). Overall, they influence chemical and physical properties of RNAs with consequences on their stability, structure, recognition, transport, and cellular localization (Frye et al. 2016). Specific enzymes are responsible for their deposition and removal, providing a large degree of variability (de Crécy-Lagard and Jaroch 2020; Ishitani et al. 2008). These modulations add a sophisticated regulatory layer in gene expression and often connect external stimuli to various cellular processes. This is particularly the case for bacteria, which need to rapidly adapt their growth to environmental stresses, to the host responses, to nutrient availability, and to antimicrobials or dangerous chemical reactive species.
Ribonucleoside modifications commonly found in bacteria. a Chemical structures of some modified ribonucleosides identified in S. aureus tRNAs using mass spectrometry (Antoine et al. 2019). The modification site is highlighted in red and the numbering systems for the nucleobases and ribose rings are indicated. m1A: N1-methyladenosine, Cm: 2′-O-methylcytidine, ac4C: N4-acetylcytidine, I: Inosine, ψ: Pseudouridine, D: Dihydrouridine. b Modified ribonucleosides in rRNAs and tRNAs from E. coli and B. subtilis identified by deep sequencing methods. The position of modifications are depicted in the tRNA cloverleaf structure, and modifications found in rRNAs are listed. Modified nucleosides are colored according to the RNA-seq protocol used for their detection (see Sect. 5). Modifications in black have been assigned with RNA-seq protocols not described. s4U: 4-thiouridine, Gm: 2′-O-methylguanosine, Um: 2′-O-methyluridine, s2C: 2-thiocytidine, k2C: 2-lysidine, ho5U: 5-hydroxyuridine, U*: hypermodified 2- and 5-substituted U34 residues, galQ: Galactosyl-queuosine, Q: Queuosine, ct6A: cyclic N6-threonylcarbamoyladenosine, m6t6A: N6-methyl-threonyl-N6-threonylcarbamoyladenosine, i6A: N6-isopentyladenosine, ms2i6A: 2-methylthio-N6-isopentyladenosine, t6A: N6-threonylcarbamoyladenosine, m1G: N1-methylguanosine, m6A: N6-methyladenosine, m2A: 2-methyladenosine, m7G: N7-methylguanosine, acp3U: 3-(3-amino-3-carboxypropyl)uridine, m5U: 5-methyluridine, m3U: N3-methyluridine, m3ψ: N3-methylpseudouridine, m4Cm: N4-, 2′-O-dimethylcytidine, m5C: 5-methylcytidine, m2G: N2-methylguanosine
Epitranscriptomic regulation often targets the most modified RNA species, tRNAs and rRNAs, influencing the translation process. Both codon decoding and peptide bond formation are affected by specific modifications on tRNAs and rRNAs, which influence translation rates and accuracy. Consequently, they directly impact protein yields, their proper folding, and their activity (Samatova et al. 2020; Antoine et al. 2021). Translation regulation fine tunes protein synthesis to balance the relative abundance of proteins in the same pathways or to provide the correct stoichiometry of complexes (Li et al. 2014). In addition, translational control is well appropriate when fast adaptation is required (Tollerson and Ibba 2020; Duval et al. 2015). In different stress conditions, modulation of RNA modifications can be rapidly achieved by changing the level of the modification enzymes or by affecting their activity.
Stable non-coding RNAs, rRNAs, and tRNAs, which represent in bacteria >95% of the total RNA fraction are extensively modified during their maturation. Despite decades of studies, the comprehensive and reliable profiling of tRNA and rRNA modifications was accomplished only for a few model species. For bacterial species, such comprehensive profiling was performed for Escherichia coli as a prominent representative of Gram-negative bacteria (see MODOMICS) and rather recently, for Bacillus subtilis (de Crecy-Lagard et al. 2020) and Staphylococcus aureus (Antoine et al. 2019) as representative Gram-positive species. Despite the fact that MODOMICS and tRNAdb (Jühling et al. 2009) databases show that all known bacterial tRNAs have numerous common modifications, they also revealed specificity among bacterial species. Dihydrouridine (D), pseudouridine (ψ), 5-methyluridine (m5U/T), and 7-methylguanosine (m7G) are highly common, while 2′-O-methylations (Nm), 4-thiouridine (s4U), and anticodon loop modifications are rarer, and others are unique for specific tRNA species (Fig. 1b). Although Gram-negative and Gram-positive bacterial species share a number of common tRNA modifications, characteristics were identified in the D-arm, e.g., Gm18 is only found in Gram-negative bacteria, while m1A22 is primarily observed in several Gram-positive bacteria. In contrast to tRNAs, rRNA modification profiles are less diverse among bacteria. However, the exact modification position is not necessarily conserved between different bacterial species but a similar modification is often present in the same rRNA structural domain (Piekna-Przybylska et al. 2008). Hence, it is important to gain more knowledge on the evolution of RNA modifications across bacterial phylogeny and analyze the structural environment of chemical modifications to obtain insights into their role and mode of interaction with neighboring residues.
Accurate detection of RNA modifications is still an extremely difficult task, as today there is still not a unique method that provides the whole pattern of RNA modifications. These approaches include various chromatographies (i.e., TLC, HPLC), reverse transcriptase analyses, affinity gel electrophoresis, and more sophisticated methods including liquid chromatography-tandem mass spectrometry analyses of nucleosides or RNA fragments (LC-MS/MS), and deep sequencing approaches (Schaefer et al. 2017; Wetzel and Limbach 2016; Motorin and Helm 2022; Yoluç et al. 2021). Recent nanopore technologies can potentially provide access to multiple modifications in native RNA molecules (Garalde et al. 2018; Begik et al. 2021). Even if these methods offer an extraordinary amount of information and depth, they nevertheless suffer from limitations and biases.
Here we describe the methodologies that were used to determine the modification patterns of stable RNAs (tRNAs and rRNAs) from Staphylococcus aureus, which is an opportunistic human bacterial pathogen causing a large variety of infections. We will especially underline the power of combining complementary methodologies.
2 RNA Purification
2.1 Isolation of Total RNA from S. Aureus
Sample requirements differ depending on the type of technique used for detecting RNA modifications. High amounts of homogenous RNAs are often required for successful analyses and appropriate methods for RNA purification should be optimized. A minimal number of purification steps following efficient cell lysis and RNA stabilization is crucial for correct RNA yield, purity, and integrity. The main challenge in RNA extraction from S. aureus is the disruption of the thick peptidoglycan cell wall. Successful approaches use enzymatic treatment with lysostaphin (Bastos et al. 2010), mechanical bead-beating, or a combination of both methods in the presence of protein denaturants to inhibit RNase activities and abolish RNA-protein interactions (i.e., phenol, guanidine isothiocyanate) (Atshan et al. 2012; Beltrame et al. 2015; França et al. 2011). S. aureus RNA samples are routinely prepared with the FastRNA® Pro blue kit (MP Biomedical), which uses mechanical bead-beating of bacteria suspended in the commercial solution designed to prevent RNA degradation. For large-scale preparations (up to 500 mL of culture in logarithmic phase), bead-beating lysis of S. aureus, suspended in a 1:1 mixture of aqueous buffer and acidic phenol/chloroform/isoamyl alcohol, is performed. RNA is further purified by chloroform extraction and ethanol precipitation.
2.2 Purification of rRNAs and tRNAs
As total RNA samples contain a mixture of different species (rRNAs, tRNAs, mRNAs, sRNAs, etc.), enrichment of the RNA molecule of interest is often beneficial for the analysis of modifications. Several approaches are based on size selection of RNAs (Poulson 1973). A rapid strategy is the use of specific filters (cutoff 50 kDa) to enrich for rRNAs (retentate) or tRNAs (filtrate) after ultrafiltration. Depending on the specific analysis to be performed on the RNAs, more laborious purification methods might be required. To purify small amount of individual 16S and 23S rRNAs as well as tRNAs, S. aureus RNA species can be separated by polyacrylamide gel electrophoresis (PAGE) and subsequently eluted from the excised bands (electroelution or “crush and soak”) (Meyer and Masquida 2016; Petrov et al. 2013). To get larger amounts of RNAs and to avoid gel impurities, chromatographies performed under native conditions are preferred (Kanwal and Lu 2019). For instance, size-exclusion chromatography can be used to purify the different rRNAs (23S, 16S, and 5S) and the tRNA fractions (Chionh et al. 2013; McKenna et al. 2007). Alternatively, weak anion-exchange chromatography using DEAE (diethylaminoethyl) matrix is widely used to separate tRNAs and rRNAs, which elute at low and high salt concentration, respectively (Easton et al. 2010). Finally, to analyze modification profiles of rRNA incorporated in fully matured and actively translating ribosomes, rRNAs (23S, 16S, and 5S) can be recovered by phenol-chloroform extraction of purified S. aureus total ribosomes (Khusainov et al. 2016) or polysomes samples, obtained from sucrose gradients fractionation (Brielle et al. 2017).
2.3 Purification of Individual tRNAs
Individual tRNAs can be purified from total tRNA using biotinylated oligo DNA probes which specifically hybridize to the target tRNA molecule. Yokogawa and co-workers (Yokogawa et al. 2010) optimized this method for thermostable tRNAs and we adapted it for the purification of S. aureus tRNAs. The procedure consists of four steps: (i) design and preparation of the DNA probe, (ii) probe-resin immobilization, (iii) tRNA-probe hybridization, and (iv) tRNA isolation. The DNA probe is composed of 35–40 nucleotides and contains biotin at the 5′ or 3′ end. Its sequence is usually complementary to the region between the D-arm and the anticodon loop. The biotinylated probe is immobilized on a streptavidin Sepharose resin using standard procedures. Hybridization of target tRNA and DNA probe requires denaturation of tRNA under conditions that maintain tRNA:DNA interaction. The buffer containing tetra-ethylammonium chloride has been reported as the best to destabilize the tRNA tertiary structure and to enhance the interaction between tRNA and DNA probes (Yokogawa et al. 2010). After hybridization, the remaining non-target tRNAs are removed by washing the resin with a non-denaturing buffer. Then, the desired tRNA is eluted with the non-denaturing buffer pre-heated at 60 °C. The quality of the purification is assessed by denaturing PAGE and the identity of the tRNA is confirmed by mass spectrometry (MS). Depending on the yield of the purification, eventual contaminations can be removed by gel purification or ion-exchange chromatography. The purification method described above is efficient for most tRNAs, but further optimization is often required for tRNA isoacceptors, poorly transcribed tRNAs, and tRNAs with high GC content. Methods such as reciprocal circulating chromatography (RCC) (Miyauchi et al. 2007) and chaplet column chromatography (CCC) (Suzuki and Suzuki 2007) have been optimized to increase the yield of GC-rich tRNAs (Ohira et al. 2022).
3 Mass Spectrometry to Identify and Localize Modifications in RNAs
Liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS) is the most trusted method for direct and unambiguous detection of RNA modifications, as it provides precise measurements of molecular masses (accuracy < 1 Da) even from very low amounts of material (Giessing and Kirpekar 2012; Jora et al. 2019). In this setup, the RNA sample is separated by reverse-phase high-performance liquid chromatography (RP-HPLC), and the eluting species are directly injected into a mass spectrometer equipped with two analyzers (Thomas and Akoulitchev 2006). An electrospray ionization (ESI) source (Kebarle and Tang 1993) evaporates and ionizes the analyte, and the formed (parent) ions are separated in the first analyzer according to their mass-to-charge (m/z) ratio. Ions with a defined m/z are selected for collision-induced dissociation (CID) (Ty et al. 2008) and the resulting fragments (daughter ions) are resolved in the second mass analyzer. The recorded MS and MS/MS spectra (plot of m/z values and their relative abundance) are used to depict mass shifts and fragmentation profiles that evidence the presence of additional chemical groups attached to the canonical nucleotides (Thakur et al. 2021).
Two types of complementary LC-MS/MS analyses are performed for the complete mapping of modifications in S. aureus RNAs. In the first analysis, the RNA of interest is hydrolyzed to its nucleoside (sugar+nucleobase) building blocks to identify all resident modifications (Pomerantz and McCloskey 1990). In the second, the RNA is digested with a nucleobase-specific RNase, and the sequences of the resulting oligonucleotides including modifications are determined and placed within the full-length RNA sequence (available from genomic data) (Kowalak et al. 1993). Both analyses require previous purification of the target RNA to obtain reliable data, limiting this “bottom up” mapping strategy to the analysis of one RNA at a time.
3.1 Analysis of Nucleosides
Samples for nucleoside analysis are prepared through enzymatic treatment of the RNA with nuclease P1, snake venom phosphodiesterase I, and bacterial alkaline phosphatase (Crain 1990). However, the reaction is performed under sub-optimal mildly acidic conditions (pH 5) (Wolff et al. 2020) to avoid alteration of labile modifications at higher pH (Jora et al. 2021; Miyauchi et al. 2013). Removal of phosphate groups is critical to make the nucleosides sufficiently hydrophobic to be separated on a reverse-phase column. Canonical nucleosides are the most abundant species and readily resolve into well-defined peaks, eluting according to their hydrophobicity (C < U < G < A) (Gehrke and Kuo 1990). Modified nucleosides are less abundant and they often appear earlier (e.g., D, ψ) or later (e.g., m2G, t6A) than their unmodified counterparts depending on the chemical nature of the modification (Su et al. 2014). Positional isomers (e.g., m1A, m6A, m2A) have the same mass and cannot be distinguished by tandem MS analysis; however, they exhibit different retention times that facilitate their identification when standards are available (Jora et al. 2018).
RP-HPLC of nucleosides is performed with mobile phases (A: water, B: methanol) containing 0.1% formic acid to facilitate protonation of nucleobases during ESI (Cai et al. 2015). MS analysis is performed in positive mode using a triple quadrupole (QQQ) instrument consisting of two quadrupole mass analyzers separated by a collision cell. In most cases, CID of the nucleoside ion occurs at the N-glycosidic bond, resulting in a nucleobase ion and an uncharged ribose moiety (Ross et al. 2016). The recorded m/z values of parent (MS) and daughter ions (MS/MS) as well as the associated neutral loss of unmodified (132 Da) or 2′-O-methylated (146 Da) ribose enable identification of the modified nucleoside. Pseudouridine does not contain an N-glycosidic bond and shows a different but unique fragmentation profile allowing its identification (Dudley et al. 2005). Two strategies of data acquisition can be employed depending on the aim of the experiment. Dynamic multiple reaction monitoring (DMRM) allows the identification and quantification of a defined set of modifications by restraining detection to only those specific m/z values (Thuring et al. 2016). In contrast, neutral loss scan (NLS) is used to identify the full range of modified nucleosides in the sample including novel modifications and consists in detecting all precursors ions that lose a mass equivalent to unmodified or methylated ribose (Kellner et al. 2014).
3.2 Analysis of Oligonucleotides
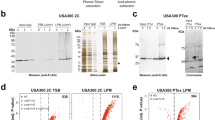
As RNase digestion of tRNA mixtures or long rRNAs often produces fragments with the same sequence, purification of the target molecule is critical to unambiguously map modifications by oligonucleotide analysis. Most of all individual tRNAs of S. aureus can be isolated using two-dimensional polyacrylamide gel electrophoresis (2D-PAGE), in which tRNAs are first separated by their length on a denaturing gel, and then by their conformation on a semi-denaturing gel (Fig. 2a) (Antoine and Wolff 2020; Antoine et al. 2019). For rRNAs, specific sequences can be obtained by RNase H cleavage using DNA probes flanking the region of interest followed by PAGE purification (Hansen et al. 2002; Nakai et al. 1994). In both cases, digestion is performed directly “in-gel” using RNase T1 (cleaving after G, m2G and I leaving a 3’ phosphate end) and the resulting oligonucleotides elute passively from the gel (Taoka et al. 2010). Other RNases, such as RNase A (cleaving after pyrimidines and ψ and leaving a 3’ phosphate end) and RNase U2 (cleaving after purines, preferably A, leaving a 3’ phosphate end), are frequently used in separated analyses to increase the sequence coverage, particularly in G-rich regions (Houser et al. 2015).
Mapping of tRNA modifications by LC-MS/MS analysis of RNase digests. a Isolation of individual tRNAs by 2D-PAGE. Total tRNAs (10 µg) are first separated by size on a denaturing polyacrylamide gel (left). Then, the whole lane is excised and embedded on the top of a semi-denaturing gel. After electrophoretic separation, the different bands migrate into multiple spots, most of which contain individual tRNA species (right). Arrows indicate the direction of migration on the first and second dimensions. Both gels are stained with ethidium bromide. The position of tRNATyr/GUA in the 2D gel is indicated. b LC-MS analysis of in-gel T1 RNase digests of tRNATyr/GUA. The unmodified sequence of the tRNA is shown and the cleavage sites of RNase T1 are labeled with arrowheads. Theoretical masses calculated with MongoOligo (http://rna.rega.kuleuven.be/masspec/mongo.htm) are displayed above the fragments of ≥4 nucleotides. The peak pointed on the elution profile corresponds likely to the oligonucleotide CUAAACG (m = 2267.3 Da) in bold. Indeed, the spacing of the isotopic peaks in its MS spectrum (inset) is 1/2, indicating thus a double-charged anion [M-2H+]2− with m = 2281.3 Da. The discrepancy between the expected and measured mass already evidences the presence of modifications. c Deconvoluted CID spectrum of the parent ion with m/z = 1140.15. Manual interpretation of the MS/MS data yielded the sequence C[D]AA[m1A]CGp containing two modified nucleotides: dihydrouridine and a N1-methyladenosine. Although the position of the methyl group on the adenosine cannot be determined from this analysis, it is known that Gram-positive bacteria display m1A at position 22 of tRNA. The phosphodiester bond between the two unmodified adenosines is shown explicitly to indicate the four possible cleavages during CID. Most abundant breakages occur at the 5′ P-O bond, generating c- and y- ion series (bold) that were used for sequencing
The negatively charged backbone of oligonucleotides raises some challenges for LC-MS/MS analysis. A concern in sample preparation is the presence of alkali cations (Na+, K+), which must be replaced by “volatile” NH4+ cations (Stults et al. 1991). Otherwise, multiple salt adducts are formed upon ESI, leading to peak broadening and interferences that complicate the interpretation of spectra (Bleicher and Bayer 1994). In-gel digests are routinely desalted using “ZipTip” C18 reverse-phase pipette tips (Millipore), but other methods such as RNA precipitation with ethanol and ammonium acetate, cation exchange chromatography, and micro-dialysis can also be used (Castleberry et al. 2008). Due to the high hydrophilicity of phosphate groups, RP-HPLC of oligonucleotides requires a mobile phase containing ion-pairing agents such as triethylamine (TEA) and triethylammonium acetate (TEAA) (Apffel et al. 1997). The ammonium moiety of these molecules masks the charges in the oligonucleotide backbone, while the alkyl group provides hydrophobic surfaces for its interaction with the reverse-phase column. Hexafluoroisopropanol (HFIP) is also included in the mobile phase to improve the ionization of oligonucleotides (McGinnis et al. 2013), which occurs in negative mode (deprotonation). Because TEA/TEAA/HFIP contaminations are difficult to eliminate and interfere with proteomic analyses performed in positive mode (Wetzel and Limbach 2016), it is better to use a dedicated LC-MS/MS system for oligonucleotides analysis.
As ESI of oligonucleotides produces several multicharged anions (Potier et al. 1994), data-dependent acquisition (DDA) is used to select the most abundant m/z species for CID, which are typically [M-2H+]−2 and [M-3H+]−3 ions. On high-resolution instruments, as the quadrupole time-of-flight (Q-TOF, accuracy < 0.05 Da), the charge (and thus the mass) of the parent ion can be inferred by inspecting the spacing of isotope peaks in the MS/MS spectra (Polo and Limbach 2001). If the spacing between the selected peak (usually the monoisotopic ion) and its consecutive isotope peak is 1/x, the charge of the parent ion will be -x (Fig. 2b). CID of oligonucleotides occurs along the phosphodiester backbone and generates a ladder of fragments that allow sequence determination (Nakayama et al. 2011; Ni et al. 1996). Among the four possible cleavage sites (Mcluckey et al. 1992), fragmentation of the 5′ P-O bond is the most frequent and produces a series of complementary c- and y-type fragment ions (Fig. 2c). Deconvolution of the multicharge MS/MS spectrum (m/z) to mono-charge spectrum (mass) simplifies the sequencing process, which consists in finding the known terminus (e.g., 3’ Gp end for RNase T1 digests). Then the masses of individual nucleotides are added to match the different peaks until the mass of the parent ion is reached (Fig. 2c). The presence of a modified nucleotide is evidenced by a unique mass shift resulting from the attached chemical group. Only pseudouridine (same mass as U) cannot be mapped with this method. A specific chemical derivatization on pseudouridine with either acrylonitrile (Mengel-Jørgensen and Kirpekar 2002) or N-cyclohexyl-N′-(2-morpholinoehyl)carbodiimide metho-p-toluene-sulfonate (CMCT) (Patteson et al. 2001) produces a detectable mass shift. Although oligonucleotide analysis readily locates single-methylated nucleotides within the RNA sequence, the position of the methyl group on the ribose or the nucleobase cannot be determined, and further analyses are required (Wolff et al. 2020).
4 Biochemical Analyses to Easily Map Specific RNA Modifications
4.1 Primer Extension
RNA modifications induce changes in the physicochemical properties of the nucleobases and ribose rings, including changes in their polarity (Charette and Gray 2000), base stacking (Roost et al. 2015), and coordination with water and ions (Agris 1996). All these events have a direct impact on the activity of RNA-directed DNA polymerases, which are viral enzymes commonly known as reverse transcriptases (RTs) (Potapov et al. 2018). These effects can be exploited to assign modification sites using simple laboratory settings. RNA modifications at the Watson-Crick edge, (e.g., m1A, m1G, m22G) readily interfere with cDNA synthesis, while other modifications (e.g., 2′O-methylation) are less problematic for primer extension, but can be detected with specific protocols. For example, under sub-optimal polymerization conditions, the presence of a modification at the nucleotide base or its ribose in the RNA template induces reverse transcription stop at the +1 position to the modification site (or, rarely, at the neighboring nucleotide) (Fig. 3a, left) (Rebane et al. 2002; Maden et al. 1995). Commonly used conditions include low deoxynucleotides (dNTPs) concentration (Motorin et al. 2007). This stop is readily detected after the separation of the cDNA products by PAGE, along with RNA sequencing profile. Most commonly, primer extension is performed with a radiolabeled DNA oligonucleotide and the radioactive products are detected by autoradiography (Fig. 3a, right). RTs from avian myeloblastosis virus (AMV) and Moloney murine leukemia virus (MMLV) are extensively used. Noteworthy, RT-signatures appear typical for a given modification, but they also depend on the reverse transcriptase used. A recent comparative study, performed on a similar RNA template using several RTs, revealed that RNA modifications can be distinguished by their RT-signature (Werner et al. 2020).
Primer extension analysis to detect several RNA modifications. a On the right, a graphical representation explaining the principles of primer extension. Reverse transcriptase (RT, transparent light blue) stops DNA synthesis (red) at RNA modification sites, in this case at a 2′-O methyl guanosine in RNA (dark blue). On the left, an autoradiography film showing RT stop patterns induced by modifications or stable RNA structures, or unspecific cleavages. Avian myeloblastosis virus (AMV) and Moloney murine leukemia virus (MMLV) RT reactions were used at low nucleotides concentration (0.02 nM), to extend a radiolabeled DNA oligonucleotide complementary to S. aureus 23S rRNA. Using this procedure, the modification Gm 2251 (E. coli numbering) could be confirmed. b Chemical nucleobase derivatizations induced by the presence of pseudouridines and dihydrouridines to be detected by RT stops. Pseudouridines are modified by 1-cyclohexyl-(2-morpholinoethyl)carbodiimide metho-p-toluene sulfonate (CMCT) followed by alkali treatment with Na2CO3, which leaves only the CMC adduct attached to the N3 position. Dihydrouridines subjected to alkaline hydrolysis (pH 10–11) produce ureidopropionic acid after ring opening
Although this technique is simple and inexpensive, it is sometimes difficult to distinguish between modifications and other possible causes of RT stops, such as RNA accidental breaks or presence of stable structures (Stern et al. 1988). To overcome misinterpretations, RT reactions are often conducted in parallel on in vitro transcribed RNAs, devoid of modifications, or on RNA purified from organisms in which specific RNA modification enzymes have been deleted or inactivated (Deryusheva et al. 2012).
Finally, it should be mentioned that many modifications cannot be directly detected by primer extension analysis, like pseudouridines, dihydrouridines, or 5-methylcytosines. For these modifications a specific pre-treatment of the sample prior to the primer extension is necessary (Fig. 3b).
4.2 Nucleotide Derivatization Coupled to Primer Extension Analysis
Given the specific chemical properties that modification brings to nucleotides, diverse options of derivatization inducing RT pauses/stops or nucleotide misincorporation have been developed for RT-silent modified nucleotides (Fig. 3b) (Behm-Ansmant et al. 2011). To map pseudouridines, chemical derivatization with CMCT is commonly used (Bakin and Ofengand 1993). CMCT modifies the N1 and N3 positions in guanosine and uridine, respectively, while pseudouridines are modified both at N1 and N3 positions. Removal of the derivatization from guanosine, uridine, and N1 of ψ is achieved by alkali treatment using Na2CO3 at pH 10 to 11. Only the N3 modification in pseudouridine is maintained due to its specific chemical environment (Ho and Gilham 1971) and can be easily detected by primer extension. Special attention is required at all steps of the labeling procedure to avoid false-positive stops. Therefore, it is important to run, in parallel, samples without treatment, or treated only with CMCT and Na2CO3 separately, to accurately discriminate stops induced by CMCT on pseudouridines from accidental cleavages of the RNA chain during the treatment (Adachi et al. 2019).
The main characteristic of dihydrouridine (D) is the absence of double bond between carbon 5 and 6 and the non-planar and non-aromatic nucleobase ring (Dalluge et al. 1996). The treatment under mild alkaline condition induces the disruption of the saturated nucleobase to produce beta-ureidopropionic acid (Xing et al. 2004) (Fig. 3b). This opened form of the nucleobase induces a specific RT stop during primer extension. Alternatively, the reduction with sodium borohydride (Cerutti and Miller 1967) causes also specific RT stops. Again, to avoid misinterpretation, it is always important to add controls with untreated samples (Igo-Kemenes and Zachau 1969; Xing et al. 2004).
4.3 Affinity Gels
Affinity electrophoretic methods rely on the strong interaction of specific functional groups present on proteins or nucleic acids with a specific reactant contained in the gel matrix (Nakamura and Takeo 1998). The displacement of the sample after gel migration provides signals about the presence or the absence of the RNA modification. For instance, boronate and organomercurial affinity gel electrophoresis have been widely used to identify specific tRNA modifications in various organisms (Tuorto et al. 2018).
Boronate affinity electrophoresis is based on the specific interaction between N-acryloyl-3-aminophenylboronic acid (APB) and free cis-hydroxyls of RNA. This is particularly the case of the queuosine (Q) modification, which possesses a cis diol group that delays the migration of the tRNAs during electrophoresis (Igloi and Kössel 1985). On the same principle, an interaction between the sulfur-modified (S) RNA with organomercurials leads to derivatized Hg-S adducts (Igloi 1988). The modified RNA with cis diol or mercury-sulfur can be visualized by different methods. For instance, ethidium bromide staining of the gel is often used for purified tRNA species while Northern blot analyses using 5’end labeled oligonucleotide allows the detection of one tRNA species among total tRNAs samples. Although boronate affinity gels are efficient to separate Q-containing RNA, it is noteworthy that the slower migration can also be observed for RNA species co-transcriptionally modified by NAD (Nübel et al. 2017). Nevertheless, this modification has not yet been observed in tRNAs. Finally, for organomercurial gels, different sensitivity has been reported for specific U-modified RNA, like s4U, and s2U (Zheng et al. 2017).
5 RNA-seq to Map Modifications on Single RNA and Complexed RNA Samples
Historically, profiling of E. coli tRNA modifications was first performed using classical methods of RNA modification analysis (i.e., semi-denaturing 2D RNA gels and RNA fingerprinting), and later combined with LC-MS analysis (Chakraburtty 1980; Kiesewetter et al. 1987; Maden et al. 1995; Mims et al. 1985). However, these analytical approaches are extremely laborious and require highly purified tRNAs, limiting their applicability to low-abundant tRNA species. Similarly, exhaustive profiling of rRNA modifications was hindered by the necessity of targeted rRNA fragmentation into short oligonucleotides of 50–100 nt long for further analysis (Yang et al. 2016). Although CMCT-RT mapping for pseudouridine (ψ) detection in the middle of 1980s (see Sect. 4.3) allowed precise profiling of this abundant modification in tRNAs and rRNAs, other RNA modifications remained understudied.
The development and implementation of deep sequencing mapping approaches as routine analytical protocols for RNA modification mapping was one of the major advances in the field. Major breakthroughs were made for comprehensive mRNA analysis, but also for rRNA and tRNA modification profiling (Motorin and Helm 2019; Motorin and Marchand 2021). Some adaptations had to be introduced for tRNAs characterized by stable tertiary structure and small size. These efforts allowed rapid and simultaneous profiling of different RNAs present in the same biological sample, but most of these methods are still able to detect only one given modification at a time. Analysis of low-abundant RNA species frequently requires enrichment of the RNA of interest, but this limitation does not hinder rRNA and tRNA analysis, since these species are very well represented in total RNA. In practice, fractionation of total RNA to rRNA and tRNA fractions may be beneficial to limit the required sequencing depth, but this step is optional and generally may be omitted. As already mentioned above, the presence of tRNA isodecoders and isoacceptors, which frequently differ only by one of few nucleotides, render the mapping difficult. Several approaches were suggested to alleviate this difficulty. For instance, only unambiguously mapped reads or “modification-aware” alignment algorithms can be retained (Arimbasseri et al. 2015; Behrens et al. 2021). Another possibility is to allow ambiguous or even multiple mapping for the same tRNA sequencing read. Finally, identical and highly similar tRNA sequences can be collapsed into one generic sequence used for reads’ alignment (Pichot et al. 2021). This approach was proven to be efficient to reduce ambiguous mapping but showed modification biases in the case of rare tRNA species.
In addition to single-nucleotide resolution mapping, a comprehensive analysis of RNA modification profiles requires appreciation and/or quantification of the modification stoichiometry site-by-site. This goal can be achieved by LC-MS analysis (with careful calibration of appropriate standards) and by deep sequencing-based methods. Quantification with Ab-based enrichment protocols is relatively imprecise, but many chemicals-based NGS approaches determine modification stoichiometry with ~5–10% error, as it was demonstrated for the RNA Bisulfite sequencing (RBS), RiboMethSeq and HydraPsiSeq protocols. Hence, these methods are extensively applied to monitor RNA modification dynamics.
Considering the tRNA and rRNA modifications commonly found in bacterial species, the most relevant deep sequencing methods for analysis of these RNAs are AlkAnilineSeq to detect m7G, D and ho5C (Marchand et al. 2021a, 2018), HydraPsiSeq for pseudouridine (ψ), some 5-substituted U residues (namely, m5U and wobble base modifications), and lysidine (k2C) (Marchand et al. 2022, 2020) and RiboMethSeq for 2′-O-methylated (Nm) nucleotides (Marchand et al. 2016, 2017; Motorin and Marchand 2018). Other approaches include RNA bisulfite sequencing to identify m5C (and also m4C) and analysis of RT misincorporation signatures for Watson-Crick edge-modified nucleobases (m1A, m1G, etc.). These methods and their applications are discussed below.
5.1 AlkAnilineSeq
AlkAnilineSeq protocol (Marchand et al. 2018) takes advantage of RNA scission at the abasic site created by specific cleavage of the fragile N-glycosidic bond formed by certain RNA modifications. Exposure of RNA containing D, m7G, m3C, and ho5C to elevated pH at high temperature leads to opening and subsequent loss of the modified nucleobase exposing RNA abasic site. Such chemical structures are known to be readily cleaved upon aniline treatment, leading to phosphodiester bond scission and liberating 5′-phosphate group at the N + 1 nucleotide relative to modification. This free 5′-phosphate is used for specific adapter ligation, providing unprecedented specificity of the detection. AlkAnilineSeq was successfully used for mapping all D and m7G residues in tRNAs, and ho5C which is a rather rare rRNA modification found in E. coli and closely related bacterial species. Specificity and sensitivity of AlkAnilineSeq are very high, low background allows detection of even substoichiometric modifications (minimum ~ 0.05 mol of modification/site) (Marchand et al. 2018, 2021a, b).
5.2 HydraPsiSeq
Detection of pseudouridines (ψ) and 5-substituted U residues is based on their resistance to hydrazine cleavage, while unmodified uridines (U) are efficiently cleaved under these conditions (Marchand et al. 2022; Peattie 1979). Thus, protected U residues in RNA may correspond to modified bases and the level of this protection is related to modification stoichiometry. Other bases in RNA (A, C, and G) are insensitive to hydrazine and cleaved only at the background level allowing the normalization of U signals. In addition, some other non-U-derived modified residues show a high sensitivity toward hydrazine cleavage, as shown for lysidine (k2C) present in bacterial tRNAIle (CAU). A similar observation was also made for m7G, but this methylation can be also detected by AlkAnilineSeq. Since protection of the modified nucleotide may also be related to its accessibility/reactivity within the RNA structure, the proportion of false-positive hits is substantially higher for HydraPsiSeq compared to AlkAnilineSeq. Hence, it is highly recommended that pseudouridine (ψ) mapping should be further validated either by conventional CMCT-RT primer extension (Sect. 4.3) or by its high-throughput variant approach PseudoU-Seq (Schwartz et al. 2014). Recently, direct RNA sequencing by Oxford Nanopore Technology (ONT) have been proposed as a new means to detect different modifications including pseudouridine (ψ), which produce characteristic base-calling “error” signatures (Begik et al. 2021; Huang et al. 2021).
5.3 RiboMethSeq
RiboMethSeq protocol is extensively used for mapping and quantification of Nm residues in rRNAs, tRNAs, and mRNAs (Birkedal et al. 2015; Marchand et al. 2016). Detection of Nm residues is based on the resistance of the RNA phosphodiester bond following Nm residue, to cleavage induced by alkaline pH and high temperature. These resistant phosphodiester bonds create “gaps” in random RNA cleavage profile. The depth of the “gap” corresponds to protection, which is related to the level of the 2′-O-methylation, allowing precise quantification.
As HydraPsiSeq, RiboMethSeq is measuring protection against RNA cleavage, the number of false-positive signals is relatively high. Additional validation of the modified candidates can be done using quantitative or semi-quantitative RTL-P or primer extension at low [dNTP] in standard or high-throughput version (2OMe-Seq) (Incarnato et al. 2017). When applied to S. aureus RNA analysis we have observed high rate of false-positive assignation for pseudouridines (ψ) by HydraPsiSeq, while Nm residues have been detected with high confidence.
5.4 RNA BisulfiteSeq
RNA bisulfite sequencing is derived from well-established DNA bisulfite sequencing and insures detection of m5C and m4C (m4Cm in rRNA) in RNAs, together with precise quantification of the modification stoichiometry (Khoddami et al. 2019; Schaefer et al. 2009). Bisulfite treatment at moderate alkaline pH (to reduce RNA degradation) leads to chemical deamination of all C to U, while m5C and m4C are resistant to bisulfite, and thus can be detected as non-deaminated C residues. Application of the method is relatively straightforward for bacterial rRNAs, while m5C was only rarely reported in bacterial tRNAs. Two major limitations have to be considered. First, only A, G, and U residues persist in RNAs after chemical deamination creating highly ambiguous alignment. Indeed, over 50% of RNA-derived reads may show multiple alignments to the reference and thus should be excluded. Second, deamination level of C residues in highly structured RNA regions may be incomplete and these non-deaminated Cs show up as false-positive hits (Squires et al. 2012).
5.5 Analysis of RT Signatures (RT-seq)
Analysis of RT-signatures, e.g., misincorporation/deletion and abortive cDNA synthesis was proposed for tentative mapping of RNA modifications having altered base-pairing properties, like I, m1G, m1A, m3U, m1acp3ψ, and other modifications altering Watson-Crick edge. Since RNA structure or even specific RNA sequences perturb cDNA synthesis by RT, these data can be used only as tentative assignment of potential modifications. More reliable mapping data can be obtained using combinations of several RT enzymes with different properties (Werner et al. 2020), or using specific RT conditions, e.g., in the presence of Mn2+ ions (Kristen et al. 2020). Detection of misincorporation and RT-arrests require adapted protocols, but simple misincorporation/deletion can be directly extracted from RiboMethSeq, HydraPsiSeq, and AlkAnilineSeq raw sequencing data sets. Misincorporation and deletion profiles need to be analyzed in parallel since the signal will appear in one or another scoring system depending on the nature of RNA modification and of the RT enzyme used. For bacterial tRNAs, these protocols allow reliable detection of s4U8, m1A22, I34, m1G37, and of many D and m7G residues.
Combination of the deep sequencing protocols cited above allows, in principle, almost exhaustive mapping of bacterial rRNA and tRNA modifications. Few RNA modifications (i.e., m2G, Q, m6A, i6A, and its derivatives), which are unreactive toward chemical reagents and/or do not lead to particular RT-signature, can be detected by LC-MSMS (Sect. 3) or by specific protocols adapted for them (i.e., NO-Seq (Werner et al. 2021) or specific RT for m6A (Aschenbrenner et al. 2018)).
6 Analysis of rRNA Modifications Using Cryo-EM Structural Studies
In the past decade, cryo electron microscopy (cryo-EM) has emerged as an extraordinary tool to resolve ribosomal structures at high resolution. The tremendous advancements in the development of direct electron detectors and other technical improvements including advanced image processing and structure sorting (Kühlbrandt 2014; Klaholz 2019) have allowed cryo-EM reconstructions to reach resolution levels around 3 Å and better, which permits to visualize post-transcriptional modifications on bacterial and eukaryotic—including human—ribosomes (Cottilli et al. 2022; Natchiar et al. 2017; Stojković et al. 2020; Watson et al. 2020). Nevertheless, the presence of some rRNA elements that, more dynamic than others, limits the assignment to the most stable ribosomal regions. While structures cannot always identify the chemical nature of a modification, they provide key insights into the structural environment of a given chemical modification. For example, 2′-O-methylation can create a more hydrophobic environment that favors Van der Waals contacts with hydrophobic side chains of amino acids, pseudouridines provide an additional possibility for hydrogen bonds through their N1 position, that classical uridines do not have, and some base modifications can increase π-stacking between neighboring nucleotide bases (Natchiar et al. 2018) (Fig. 4). The structural analysis, combined with the various methods for chemical identification discussed above, thus provides important information on the structural role and function of chemical modifications.
rRNA modifications assigned by cryo-EM structure analyses. Nucleotides are numbered according to E. coli rRNA sequences. a S. aureus rRNA methylations can be assigned by cryo-EM looking at extra densities present at specific positions on the nucleotides. For example, an electron density shape difference between the two adenine residues A2496 and A2503 in 23S rRNA has assigned at position 2 a methyl group at the adenine nucleobase A2503 (model: PDB 6YEF; map: EMD-10791 at 3.2 Å, Wang et al. 2021). In this cryo-EM structure, several other S. aureus rRNA methylations have reported (b): m4Cm: 4-methyl 2-O′ methyl cytidine; m2G: 2-methyl guanosine; m3U: 3-methyl uridine; m7G: 7-methyl guanosine; m66A: 6, 6-dimetyl guanosine, Gm: 2-O′ methyl guanosine. c Characteristic bent nucleobase density shape of Dihydrouridine (D) observed on E. coli cryo-EM structure (model: PDB 7K00; map: EMD- 22,586 at1.98 Å, Suzuki and Suzuki 2007). d Interaction between E. coli pseudouridine Ψ2605 with a possible water molecule and comparison with unmodified U2596 from the same 23S rRNA structure (model: PDB 7K00; map: EMD-22,586 at 1.98 Å). W-C edge: Watson-Crick edge; CH edge. e Example of the results obtained by using qptm software (Taoka et al. 2010). Calculated (blue) against difference (green) map from m2A2503 (modified) against A2496 (non-modified)
Structures of several bacterial ribosomes in complex with mRNA and tRNAs have shown the importance of conserved tRNA and rRNA modifications in functional sites of the small (30S) and large ribosomal (50S) subunits. On the 30S ribosomal subunit, where the tRNA anticodon interacts with the mRNA codon in the decoding center, different rRNA modifications have been shown to kinetically regulate the initiation process and to be involved in proofreading (Demirci et al. 2010). For instance, m2G966 and m4Cm1402 (E. coli numbering) both contribute to stabilize the interactions taking place between the tRNA, mRNA, and rRNA in the P-site (Burakovsky et al. 2012; Jenner et al. 2010; Polikanov et al. 2015). On the 50S subunit, several conserved modifications are located at the peptide exit tunnel or are clustered at the Peptidyl Transferase Center (PTC), which may indicate that they assist peptide bond formation. Interestingly, Gm2251 in the P-loop of the 23S rRNA (E. coli numbering) directly interacts with the CCA sequence of the tRNA and thus is expected to contribute to the correct positioning of the tRNA acceptor end (Fischer et al. 2015; Golubev et al. 2020; Polikanov et al. 2015; Watson et al. 2020).
Various strategies were used to analyze modifications from high-resolution structures of ribosomes. With X-ray structures, difference density mapping (between real densities and those artificially obtained from modification-less models) helps to indicate the position of modified nucleotides (Polikanov et al. 2015). With cryo-EM structures, the refinement of specific regions of the ribosomal subunits (focused refinement) allowed reaching better resolution to recognize rRNA modifications in combination with previous annotation using complementary methods (Watson et al. 2020; Golubev et al. 2020). Several cryo-EM structures of S. aureus ribosomes are now available (Belinite et al. 2021; Belousoff et al. 2017; Cimicata et al. 2022; Golubev et al. 2020; Halfon et al. 2019; Khusainov et al. 2020, 2017, 2016; Matzov et al. 2017; Wright et al. 2020), and 10 different methylation sites have been mapped (Golubev et al. 2020) (Fig. 4a, b), but not all modifications could be assigned. Dihydrouridines (D) and pseudouridines (ψ) can be mapped with good confidence at very high resolution, for example on the 2 Å structure of the E. coli ribosome (Watson et al. 2020). Dihydrouridines (D) impose a characteristic bent shape to their pyrimidine ring and can be easily discriminated from uridines (Fig. 4c). Pseudouridines (ψ) can be assigned indirectly, by analyzing in which manner they change the pattern of ions or water interactions. The isomerization of N1 and C5 in ψ maintains the same Watson-Crick base-pairing property (W-C edge) but includes an additional hydrogen bond donor (N1 imino proton) on the C-H edge, which is responsible for water or ions binding (Fig. 4d) (Watson et al. 2020). Assignations of ions, water molecules, and other interactions are still challenging but they will certainly contribute to alleviate better density interpretation (Wang et al. 2021).
To facilitate the analyses of thousands of nucleotides of the ribosome, dedicated software has been recently developed, which automatically calculates difference density maps. Positioning of the RNA modification signals, obtained at specific contour levels, offers now the possibility to restrict the analysis by other methods on only a subset of nucleotides (Fig. 4e) (Stojković et al. 2020). However, efforts are still needed to complete the mapping of all the modifications in S. aureus rRNAs and to gain an evolutionary analysis of the conserved and specific sites by comparing with the extensive analysis done for E. coli (Watson et al. 2020) and T. thermophilus (Polikanov et al. 2015). Structures at higher resolution will help overcoming uncertainty in extra density coming from methylations, to discriminate pseudouridines according to their specific interaction network, and to observe conformational changes induced by modified nucleobases.
7 Discussion
Global analysis of the bacterial epitranscriptome and its variations occurring in response to environmental changes and various stresses is still a major technical challenge. Combination of approaches involving high-resolution structures of RNA machineries, mass spectrometry, and next-generation sequencing is certainly one way to render this objective amenable. As discussed in the review, the choice of the approaches depends on the RNA species to be studied. Recent advances in the Nanopore technology allow now the direct sequencing of full-length native RNA molecules without the need of RT and PCR amplification, and the obtained data normally contain all the information to assign the modifications (reviewed in Begik et al. 2021). However, Nanopore sequencing is still in its infancy and further improvement in the direct RNA sequencing are required for routine application. In most cases, Nanopore data are used only for validation, but not for de novo mapping of RNA modifications. Moreover, application to non-polyadenylated RNA species, like rRNA and tRNA, requires the use of polyadenylation by polyA-polymerase or specific custom primers. The complexity of Nanopore ion current profiles also requires further development of appropriate interpretation informatics pipelines, frequently based on machine learning algorithms or artificial intelligence.
Another major issue is to gain more knowledge on the protein machineries that are required for the modification deposition (writer) or which prevents the modification to take place (eraser). Even if phylogeny can be used to predict the enzymes and pathways required for RNA modifications, little is known in S. aureus. Many questions still remained to be addressed: are modification enzymes regulated upon stress conditions? Are there specific modification enzymes for mRNAs, regulatory RNAs, tRNAs, and rRNAs? What are the enzymes required for biofilm formation and for virulence? Are the modified mRNAs conserved among bacteria? Previous sensitive LC-MS approach has revealed that m6A/A ratio in mRNAs was higher in Gram-negative than in Gram-positive bacteria including S. aureus (Deng et al. 2015). Transcriptomic analysis performed in E. coli and Pseudomonas aeruginosa revealed that m6A was enriched in mRNAs involved in energy metabolism, stress responses, and in several sRNAs (Deng et al. 2015). Due to the rather small genome size, the high diversity of species, and the existing tools to delete genes, the bacteria represent ideal model organisms to decipher the extent of the functions of RNA modifications and of the associated enzyme machineries. No doubt the development of innovative sequencing approaches will be essential to gain full understanding of the bacterial epitranscriptome and to monitor its impact in gene regulation.
References
Adachi H, DeZoysa MD, Yu YT (2019) Detection and Quantification of Pseudouridine in RNA. Methods Mol Biol 1870:219–235
Agris PF (1996) The importance of being modified: roles of modified nucleosides and Mg2+ in RNA structure and function. Prog Nucleic Acid Res Mol Biol 53:79–129
Antoine L, Bahena-Ceron R, Devi Bunwaree H et al (2021) RNA modifications in pathogenic bacteria: impact on host adaptation and virulence. Genes (basel) 12:1125
Antoine L, Wolff P (2020) Mapping of posttranscriptional tRNA modifications by two-dimensional gel electrophoresis mass spectrometry. Methods Mol Biol 2113:101–110
Antoine L, Wolff P, Westhof E et al (2019) Mapping post-transcriptional modifications in Staphylococcus aureus tRNAs by nanoLC/MSMS. Biochimie 164:60–69
Apffel A, Chakel JA, Fischer S et al (1997) Analysis of oligonucleotides by HPLC-electrospray ionization mass spectrometry. Anal Chem 69:1320–1325
Arimbasseri AG, Blewett NH, Iben JR et al (2015) RNA polymerase III output is functionally linked to tRNA dimethyl-G26 modification. PLoS Genet 11(12):e1005671
Aschenbrenner J, Werner S, Marchand V et al (2018) Engineering of a DNA polymerase for direct m(6) a sequencing. Angew Chem Int Ed Engl 57:417–421
Atshan SS, Shamsudin MN, Lung LTT et al (2012) Improved method for the isolation of RNA from bacteria refractory to disruption, including S. aureus producing biofilm. Gene 494:219–224
Bakin A, Ofengand J (1993) Four newly located pseudouridylate residues in Escherichia coli 23S ribosomal RNA are all at the peptidyltransferase center: analysis by the application of a new sequencing technique. Biochemistry 32:9754–9762
Bastos MdCdF, Coutinho BG, Coelho MLV (2010) Lysostaphin: a staphylococcal bacteriolysin with potential clinical applications. Pharmaceuticals 3:1139–1161
Begik O, Lucas MC, Pryszcz LP et al (2021) Quantitative profiling of pseudouridylation dynamics in native RNAs with nanopore sequencing. Nat Biotechnol 39:1278–1291
Behm-Ansmant I, Helm M, Motorin Y (2011) Use of specific chemical reagents for detection of modified nucleotides in RNA. J Nucleic Acids 2011:408053
Behrens A, Rodschinka G, Nedialkova DD (2021) High-resolution quantitative profiling of tRNA abundance and modification status in eukaryotes by mim-tRNAseq. Mol Cell 81:1802–1815
Belinite M, Khusainov I, Soufari H et al (2021) Stabilization of Ribosomal RNA of the Small Subunit by Spermidine in Staphylococcus aureus. Front Mol Biosci 8:738752
Belousoff MJ, Eyal Z, Radjainia M et al (2017) Structural Basis for Linezolid Binding Site Rearrangement in the Staphylococcus aureus Ribosome. mBio 8:e00395-17
Beltrame CO, Côrtes MF, Bandeira PT et al (2015) Optimization of the RNeasy mini kit to obtain high-quality total RNA from sessile cells of Staphylococcus aureus. Braz J Med Biol Res 48:1071–1076
Birkedal U, Christensen-Dalsgaard M, Krogh N et al (2015) Profiling of ribose methylations in RNA by high-throughput sequencing. Angew Chem Int Ed Engl 54:451–455
Bleicher K, Bayer E (1994) Various factors influencing the signal intensity of oligonucleotides in electrospray mass spectrometry. Biol Mass Spectrom 23:320–322
Boccaletto P, Stefaniak F, Ray A et al (2022) MODOMICS: a database of RNA modification pathways. 2021 update. Nucleic Acids Res 50:D231–D235
Brielle R, Pinel-Marie ML, Chat S et al (2017) Purification, identification, and functional analysis of polysomes from the human pathogen Staphylococcus aureus. Methods 117:59–66
Burakovsky DE, Prokhorova IV, Sergiev PV, a. (2012) Impact of methylations of m2G966/m5C967 in 16S rRNA on bacterial fitness and translation initiation. Nucleic Acids Res 40:7885–7895
Cai WM, Chionh YH, Hia F et al (2015) A platform for discovery and quantification of modified ribonucleosides in RNA: application to stress-induced reprogramming of tRNA modifications. Methods Enzymol 560:29–71
Castleberry CM, Rodicio LP, Limbach PA (2008) Electrospray ionization mass spectrometry of oligonucleotides. Curr Protoc Nucleic Acid Chem
Cerutti P, Miller N (1967) Selective reduction of yeast transfer ribonucleic acid with sodium borohydride. J Mol Biol 26:55–66
Chakraburtty K (1980) Recognition of E coli tRNAArg by arginyl tRNA synthetase. Nucleic Acids Res 8:4459–4472
Charette M, Gray MW (2000) Pseudouridine in RNA: what, where, how, and why. IUBMB Life 49:341–351
Chionh YH, Ho CH, Pruksakorn D et al (2013) A multidimensional platform for the purification of non-coding RNA species. Nucleic Acids Res 41:e168
Cimicata G, Fridkin G, Bose T et al (2022) Structural studies reveal the role of helix 68 in the elongation step of protein biosynthesis. mBio 13:e0030622
Cottilli P, Itoh Y, Nobe Y et al (2022) Cryo-EM structure and rRNA modification sites of a plant ribosome. Plant Commun 3:100342
Crain PF (1990) Preparation and enzymatic hydrolysis of DNA and RNA for mass spectrometry. Meth Enzymol 193:782–790
Dalluge JJ, Hashizume T, Sopchik AE, a. (1996) Conformational flexibility in RNA: the role of dihydrouridine. Nucleic Acids Res 24:1073–1079
de Crécy-Lagard V, Jaroch M (2020) Functions of bacterial tRNA modifications: from ubiquity to diversity. Trends Microbiol 29:41–53
de Crecy-Lagard V, Ross RL, Jaroch M et al (2020) Survey and validation of tRNA modifications and their corresponding genes in bacillus subtilis sp subtilis strain 168. Biomolecules 10:977
Demirci H, Ft M, Belardinelli R et al (2010) Modification of 16S ribosomal RNA by the KsgA methyltransferase restructures the 30S subunit to optimize ribosome function. RNA 16(12):2319–2324
Deng X, Chen K, Luo GZ et al (2015) Widespread occurrence of N6-methyladenosine in bacterial mRNA. Nucleic Acids Res 43:6557–6567
Deryusheva S, Choleza M, Barbarossa A et al (2012) Post-transcriptional modification of spliceosomal RNAs is normal in SMN-deficient cells. RNA 18:31–36
Dudley E, Tuytten R, Bond A et al (2005) Study of the mass spectrometric fragmentation of pseudouridine: comparison of fragmentation data obtained by matrix-assisted laser desorption/ionisation post-source decay, electrospray ion trap multistage mass spectrometry, and by a method utilising elect. Rapid Commun Mass Spectrom 19:3075–3085
Duval M, Simonetti A, Caldelari I et al (2015) Multiple ways to regulate translation initiation in bacteria: mechanisms, regulatory circuits, dynamics. Biochimie 114:18–29
Easton LE, Shibata Y, Lukavsky PJ (2010) Rapid, nondenaturing RNA purification using weak anion-exchange fast performance liquid chromatography. RNA 16:647–653
Fischer N, Neumann P, Konevega AL et al (2015) Structure of the E. coli ribosome-EF-Tu complex at <3 a resolution by Cs-corrected cryo-EM. Nature 520:567–570
França A, Melo LD, Cerca N (2011) Comparison of RNA extraction methods from biofilm samples of Staphylococcus epidermidis. BMC Res Notes 4:572
Frye M, Jaffrey SR, Pan T et al (2016) RNA modifications: what have we learned and where are we headed? Nat Rev Genet 17:365–372
Garalde DR, Snell EA, Jachimowicz D et al (2018) Highly parallel direct RNA sequencing on an array of nanopores. Nat Methods 15:201–206
Gehrke CW, Kuo KC (1990) Ribonucleoside analysis by reversed-phase high performance liquid chromatography. J Chromatogr Library 45(Part A):A3–A71
Giessing AMB, Kirpekar F (2012) Mass spectrometry in the biology of RNA and its modifications. J Proteomics 75:3434–3449. https://doi.org/10.1016/j.jprot.2012.01.032
Golubev A, Fatkhullin B, Khusainov I et al (2020) Cryo-EM structure of the ribosome functional complex of the human pathogen Staphylococcus aureus at 3.2 Å resolution. FEBS Lett 594:3551–3567
Halfon Y, Matzov D, Eyal Z et al (2019) Exit tunnel modulation as resistance mechanism of S. aureus erythromycin resistant mutant. Sci Rep 9:11460
Hansen MA, Kirpekar F, Ritterbusch W et al (2002) Posttranscriptional modifications in the A-loop of 23S rRNAs from selected archaea and eubacteria. RNA 8:202–213
Ho NW, Gilham PT (1971) Reaction of pseudouridine and inosine with N-cyclohexyl-N’-beta-(4-methylmorpholinium)ethylcarbodiimide. Biochemistry 10:3651–3657
Houser WM, Butterer A, Addepalli B, a. (2015) Combining recombinant ribonuclease U2 and protein phosphatase for RNA modification mapping by liquid chromatography-mass spectrometry. Anal Biochem 478:52–58
Huang S, Zhang W, Katanski CD et al (2021) Interferon inducible pseudouridine modification in human mRNA by quantitative nanopore profiling. Genome Biol 22:330
Igloi GL (1988) Interaction of tRNAs and of phosphorothioate-substituted nucleic acids with an organomercurial. Probing the chemical environment of thiolated residues by affinity electrophoresis. Biochemistry 27(10):3842–3849
Igloi GL, Kössel H (1985) Affinity electrophoresis for monitoring terminal phosphorylation and the presence of queuosine in RNA. Application of polyacrylamide containing a covalently bound boronic acid. Nucleic Acids Res 13:6881–6898
Igo-Kemenes T, Zachau HG (1969) On the specificity of the reduction of transfer ribonucleic acids with sodium borohydride. Eur J Biochem 10:549–556
Incarnato D, Anselmi F, Morandi E et al (2017) High-throughput single-base resolution mapping of RNA 2΄-O-methylated residues. Nucleic Acids Res 45:1433–1441
Ishitani R, Yokoyama S, Nureki O (2008) Structure, dynamics, and function of RNA modification enzymes. Curr Opin Struct Biol 18:330–339
Jenner LB, Demeshkina N, Yusupova G et al (2010) Structural aspects of messenger RNA reading frame maintenance by the ribosome. Nat Struct Mol Biol 17:555–560
Jora M, Borland K, Abernathy S et al (2021) Chemical amination/imination of carbonothiolated nucleosides during RNA hydrolysis. Angew Chem Int Ed Engl 60:3961–3966
Jora M, Burns AP, Ross RL et al (2018) Differentiating positional isomers of nucleoside modifications by higher-energy collisional dissociation mass spectrometry (HCD MS). J Am Soc Mass Spectrom 29:1745–1756
Jora M, Lobue PA, Ross RL et al (2019) Detection of ribonucleoside modifications by liquid chromatography coupled with mass spectrometry. Biochim Biophys Acta Gene Regul Mech 1862:280–290
Jühling F, Mörl M, Hartmann RK et al (2009) tRNAdb 2009: compilation of tRNA sequences and tRNA genes. Nucleic Acids Res 37:D159-162
Kanwal F, Lu C (2019) A review on native and denaturing purification methods for non-coding RNA (ncRNA). J Chromatogr B Analyt Technol Biomed Life Sci 1120:71–79
Kebarle P, Tang L (1993) From ions in solution to ions in the gas phase. Anal Chem 65:972A-986A
Kellner S, Neumann J, Rosenkranz D et al (2014) Profiling of RNA modifications by multiplexed stable isotope labelling. Chem Commun 50:3516–3518
Khoddami V, Yerra A, Mosbruger TL et al (2019) Transcriptome-wide profiling of multiple RNA modifications simultaneously at single-base resolution. Proc Natl Acad Sci USA 116:6784–6789
Khusainov I, Fatkhullin B, Pellegrino S et al (2020) Mechanism of ribosome shutdown by RsfS in Staphylococcus aureus revealed by integrative structural biology approach. Nat Commun 11:1656
Khusainov I, Vicens Q, Ayupov R et al (2017) Structures and dynamics of hibernating ribosomes from Staphylococcus aureus mediated by intermolecular interactions of HPF. EMBO J 36:2073–2087
Khusainov I, Vicens Q, Bochler A et al (2016) Structure of the 70S ribosome from human pathogen Staphylococcus aureus. Nucleic Acids Res 44:10491–10504
Kiesewetter S, Fischer W, Sprinzl M (1987) Sequences of three minor tRNAsArg from E. coli. Nucleic Acids Res 15:3184
Klaholz BP (2019) Deriving and refining atomic models in crystallography and cryo-EM: the latest Phenix tools to facilitate structure analysis. Acta Crystallogr D Struct Biol 75:878–881
Kowalak JA, Pomerantz SC, Crain PF et al (1993) A novel method for the determination of post-transcriptional modification in RNA by mass spectrometry. Nucleic Acids Res 21:4577–4585
Kristen M, Plehn J, Marchand V et al (2020) Manganese Ions Individually Alter the Reverse Transcription Signature of Modified Ribonucleosides. Genes (basel) 11:950
Kühlbrandt W (2014) Biochemistry. The resolution revolution. Science 343:1443–1444
Li GW, Burkhardt D, Gross C et al (2014) Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell 157:624–635
Maden BE, Corbett ME, Heeney PA et al (1995) Classical and novel approaches to the detection and localization of the numerous modified nucleotides in eukaryotic ribosomal RNA. Biochimie 77:22–29
Marchand V, Blanloeil-Oillo F, Helm M et al (2016) Illumina-based RiboMethSeq approach for mapping of 2’-O-Me residues in RNA. Nucleic Acids Res 44:e135
Marchand V, Ayadi L, Ernst FGM et al (2018) AlkAniline-Seq: profiling of m(7) G and m(3) C RNA modifications at single nucleotide resolution. Angew Chem Int Ed Engl 57:16785–16790
Marchand V, Ayadi L, Bourguignon-Igel V et al (2021a) AlkAniline-Seq: a highly sensitive and specific method for simultaneous mapping of 7-methyl-guanosine (m(7)G) and 3-methyl-cytosine (m(3)C) in RNAs by high-throughput sequencing. Meth Mol Biol 2298:77–95
Marchand V, Bourguignon-Igel V, Helm M et al (2021b) Mapping of 7-methylguanosine (m(7)G), 3-methylcytidine (m(3)C), dihydrouridine (D) and 5-hydroxycytidine (ho(5)C) RNA modifications by AlkAniline-Seq. Methods Enzymol 658:25–47
Marchand V, Bourguignon-Igel V, Helm M et al (2022) Analysis of pseudouridines and other RNA modifications using HydraPsiSeq protocol. Methods 203:383–391
Marchand V, Pichot F, Neybecker P et al (2020) HydraPsiSeq: a method for systematic and quantitative mapping of pseudouridines in RNA. Nucleic Acids Res 48:e110
Marchand V, Pichot F, Thuring K et al (2017) Next-generation sequencing-based ribomethseq protocol for analysis of tRNA 2’-O-methylation. Biomolecules 7:13
Matzov D, Aibara S, Basu A et al (2017) The cryo-EM structure of hibernating 100S ribosome dimer from pathogenic Staphylococcus aureus. Nat Commun 8:723
McGinnis AC, Grubb EC, Bartlett MG (2013) Systematic optimization of ion-pairing agents and hexafluoroisopropanol for enhanced electrospray ionization mass spectrometry of oligonucleotides. Rapid Commun Mass Spectrom 27:2655–2664
McKenna SA, Kim I, Puglisi EV et al (2007) Purification and characterization of transcribed RNAs using gel filtration chromatography. Nat Protoc 2:3270–3277
Mcluckey SA, Van Berkel GJ, Glish GL (1992) Tandem mass spectrometry of small, multiply charged oligonucleotides. J Am Soc Mass Spectrom 3:60–70
Mengel-Jørgensen J, Kirpekar F (2002) Detection of pseudouridine and other modifications in tRNA by cyanoethylation and MALDI mass spectrometry. Nucleic Acids Res 30:e135
Meyer M, Masquida B (2016) Polyacrylamide gel electrophoresis for purification of large amounts of RNA. Methods Mol Biol 1320:59–65
Mims BH, Prather NE, Murgola EJ (1985) Isolation and nucleotide sequence analysis of tRNAAlaGGC from Escherichia coli K-12. J Bacteriol 162:837–839
Miyauchi K, Kimura S, Suzuki T (2013) A cyclic form of N 6-threonylcarbamoyladenosine as a widely distributed tRNA hypermodification. Nat Chem Biol 9:105–111
Miyauchi K, Ohara T, Suzuki T (2007) Automated parallel isolation of multiple species of non-coding RNAs by the reciprocal circulating chromatography method. Nucleic Acids Res 35:e24
Motorin Y, Helm M (2019) Methods for RNA modification mapping using deep sequencing: established and new emerging technologies. Genes (basel) 10:35
Motorin Y, Helm M (2022) RNA nucleotide methylation: 2021 update. Wiley Interdiscip Rev RNA 13:e1691
Motorin Y, Marchand V (2018) Detection and analysis of RNA ribose 2’-O-methylations: challenges and solutions. Genes (basel) 9:642
Motorin Y, Marchand V (2021) Analysis of RNA modifications by second- and third-generation deep sequencing: 2020 update. Genes (basel) 12:278
Motorin Y, Muller S, Behm-Ansmant I, a. (2007) Identification of modified residues in RNAs by reverse transcription-based methods. Meth Enzymol 425:21–53
Nakai C, Konishi A, Komatsu Y et al (1994) Sequence-specific cleavage of RNA by a hybrid ribonuclease H. FEBS Lett 339:67–72
Nakamura K, Takeo K (1998) Affinity electrophoresis and its applications to studies of immune response. J Chromatogr B Biomed Sci Appl 715:125–136
Nakayama H, Takahashi N, Isobe T (2011) Informatics for mass spectrometry-based RNA analysis. Mass Spectrom Rev 30:1000–1012
Natchiar SK, Myasnikov AG, Hazemann I et al (2018) Visualizing the role of 2’-OH rRNA methylations in the human ribosome structure. Biomolecules 8:125
Natchiar SK, Myasnikov AG, Kratzat H et al (2017) Visualization of chemical modifications in the human 80S ribosome structure. Nature 551:472–477
Ni J, Pomerantz C, Rozenski J et al (1996) Interpretation of oligonucleotide mass spectra for determination of sequence using electrospray ionization and tandem mass spectrometry. Anal Chem 68:1989–1999
Nübel G, Sorgenfrei FA, Jäschke A (2017) Boronate affinity electrophoresis for the purification and analysis of cofactor-modified RNAs. Methods 117:14–20
Ohira T, Minowa K, Sugiyama K et al (2022) Reversible RNA phosphorylation stabilizes tRNA for cellular thermotolerance. Nature 605:372–379
Patteson KG, Rodicio LP, Limbach PA (2001) Identification of the mass-silent post-transcriptionally modified nucleoside pseudouridine in RNA by matrix-assisted laser desorption/ionization mass spectrometry. Nucleic Acids Res 29:49–49
Peattie DA (1979) Direct chemical method for sequencing RNA. Proc Natl Acad Sci USA 76:1760–1764
Petrov A, Wu T, Puglisi EV et al (2013) RNA purification by preparative polyacrylamide gel electrophoresis. Meth Enzymol 530:315–330
Pichot F, Marchand V, Helm M et al (2021) Non-redundant tRNA reference sequences for deep sequencing analysis of tRNA abundance and epitranscriptomic RNA modifications. Genes (basel) 12:81
Piekna-Przybylska D, Decatur WA, Fournier MJ (2008) The 3D rRNA modification maps database: with interactive tools for ribosome analysis. Nucleic Acids Res 36:D178-183
Polikanov YS, Melnikov SV, Soll D et al (2015) Structural insights into the role of rRNA modifications in protein synthesis and ribosome assembly. Nat Struct Mol Biol 22:342–344
Polo LM, Limbach PA (2001) Analysis of Oligonucleotides by Electrospray Ionization Mass Spectrometry. Curr Protoc Nucleic Acid Chem
Pomerantz SC, McCloskey JA (1990) Analysis of RNA hydrolyzates by liquid chromatography-mass spectrometry. Methods Enzymol 193:796–824
Potapov V, Fu X, Dai N et al (2018) Base modifications affecting RNA polymerase and reverse transcriptase fidelity. Nucleic Acids Res 46:5753–5763
Potier N, van Dorsselaer A, Cordier Y et al (1994) Negative electrospray ionization mass spectrometry of synthetic and chemically modified oligonucleotides. Nucleic Acids Res 22:3895–3903
Poulson R (1973) Isolation, Purification and Fractionation of RNA. The ribonucleic acids, pp 243–261
Rebane A, Roomere H, Metspalu A (2002) Locations of several novel 2’-O-methylated nucleotides in human 28S rRNA. BMC Mol Biol 3:1
Roost C, Lynch SR, Batista PJ et al (2015) Structure and thermodynamics of N6-methyladenosine in RNA: a spring-loaded base modification. J Am Chem Soc 137:2107–2115
Ross R, Cao X, Yu N et al (2016) Sequence mapping of transfer RNA chemical modifications by liquid chromatography tandem mass spectrometry. Methods (san Diego, Calif) 107:73–78
Samatova E, Daberger J, Liutkute M et al (2020) Translational control by ribosome pausing in bacteria: how a non-uniform pace of translation affects protein production and folding. Front Microbiol 11:619430
Schaefer M, Kapoor U, Jantsch MF (2017) Understanding RNA modifications: the promises and technological bottlenecks of the ‘epitranscriptome.’ Open Biol 7:1–14
Schaefer M, Pollex T, Hanna K et al (2009) RNA cytosine methylation analysis by bisulfite sequencing. Nucleic Acids Res 37:e12
Schwartz S, Bernstein DA, Mumbach MR et al (2014) Transcriptome-wide mapping reveals widespread dynamic-regulated pseudouridylation of ncRNA and mRNA. Cell 159:148–162
Squires JE, Patel HR, Nousch M et al (2012) Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res 40:5023–5033
Stern S, Moazed D, Noller HF (1988) Structural analysis of RNA using chemical and enzymatic probing monitored by primer extension. Methods Enzymol 164:481–489
Stojković V, Myasnikov AG, Young ID et al (2020) Assessment of the nucleotide modifications in the high-resolution cryo-electron microscopy structure of the Escherichia coli 50S subunit. Nucleic Acids Res 48:2723–2732
Stults JT, Marsters JC, Carr SA (1991) Improved electrospray ionization of synthetic oligodeoxynucleotides. Rapid Commun Mass Spectrom 5:359–363
Su D, Chan CTY, Gu C et al (2014) Quantitative analysis of ribonucleoside modifications in tRNA by HPLC-coupled mass spectrometry. Nat Protoc 9:828–841
Suzuki T, Suzuki T (2007) Chaplet column chromatography: isolation of a large set of individual RNAs in a single step. Meth Enzymol 425:231–239
Taoka M, Ikumi M, Nakayama H et al (2010) In-gel digestion for mass spectrometric characterization of RNA from fluorescently stained polyacrylamide gels. Anal Chem 82:7795–7803
Thakur P, Jora M, Zhao R et al (2021) Mass spectrometry-based methods for characterization of hypomodifications in transfer RNA. Springer. Epitranscriptomics, pp 555–592
Thomas B, Akoulitchev AV (2006) Mass spectrometry of RNA. Trends Biochem Sci 31:173–181
Thuring K, Schmid K, Keller P et al (2016) Analysis of RNA modifications by liquid chromatography-tandem mass spectrometry. Methods 107:48–56
Tollerson R 2nd, Ibba M (2020) Translational regulation of environmental adaptation in bacteria. J Biol Chem 295:10434–10445
Tuorto F, Legrand C, Cirzi C et al (2018) Queuosine-modified tRNAs confer nutritional control of protein translation. EMBO J 37:e99777
Ty H, Kharlamova A, Liu J et al (2008) Ion trap collision-induced dissociation of multiply deprotonated RNA: c/y-Ions versus (a-B)/w-Ions. J Am Soc Mass Spectrom 19:1832–1840
Wang J, Natchiar SK, Moore PB et al (2021) Identification of Mg(2+) ions next to nucleotides in cryo-EM maps using electrostatic potential maps. Acta Crystallogr D Struct Biol 77:534–539
Watson ZL, Ward FR, Méheust R et al (2020) Structure of the bacterial ribosome at 2 Å resolution. Elife 9:e60482
Werner S, Galliot A, Pichot F et al (2021) NOseq: amplicon sequencing evaluation method for RNA m6A sites after chemical deamination. Nucleic Acids Res 49:e23
Werner S, Schmidt L, Marchand V et al (2020) Machine learning of reverse transcription signatures of variegated polymerases allows mapping and discrimination of methylated purines in limited transcriptomes. Nucleic Acids Res 48:3734–3746
Wetzel C, Limbach PA (2016) Mass spectrometry of modified RNAs: recent developments. Analyst 141:16–23
Wolff P, Villette C, Zumsteg J et al (2020) Comparative patterns of modified nucleotides in individual tRNA species from a mesophilic and two thermophilic archaea. RNA 26:1957–1975
Wright A, Deane-Alder K, Marschall E et al (2020) Characterization of the core ribosomal binding region for the oxazolidone family of antibiotics using cryo-EM. ACS Pharmacol Transl Sci 3:425–432
Xing F, Hiley SL, Hughes TR et al (2004) The specificities of four yeast dihydrouridine synthases for cytoplasmic tRNAs. J Biol Chem 279:17850–17860
Yang J, Sharma S, Watzinger P et al (2016) Mapping of complete set of ribose and base modifications of yeast rRNA by RP-HPLC and mung bean nuclease assay. PLoS ONE 11:e0168873
Yokogawa T, Kitamura Y, Nakamura D et al (2010) Optimization of the hybridization-based method for purification of thermostable tRNAs in the presence of tetraalkylammonium salts. Nucleic Acids Res 38:e89
Yoluç Y, Ammann G, Barraud P et al (2021) Instrumental analysis of RNA modifications. Crit Rev Biochem Mol Biol 56:178–204
Zheng C, Black KA, Dos Santos PC (2017) Diverse mechanisms of sulfur decoration in bacterial tRNA and their cellular functions. Biomolecules 7:33
Acknowledgements
This work was supported by the Centre National de la Recherche Scientifique (CNRS), by the French National Research Agency ANR (ANR-21-CE12-0030-01 to [SM]), by the Region Grand Est (N°18P-09227- EpiRNA), and the previous French National Program Investissement d’Avenir (Labex NetRNA) (ANR-10-LABX-0036_NETRNA). This work of the Interdisciplinary Thematic Institute IMCBio, as part of the ITI 2021–2028 program of the University of Strasbourg, CNRS, and Inserm, was supported by IdEx Unistra (ANR-10-IDEX-0002), by SFRI-STRAT’US project (20-SFRI-0012), and EUR IMCBio (IMCBio ANR-17-EURE-0023) under the framework of the France 2030 Program. [HK] is supported by a fellowship from Fondation de la Recherche Médicale (FRM). [BPK] acknowledges support by CNRS, Association pour la Recherche sur le Cancer (ARC), Institut National du Cancer (INCa_16099), the Fondation pour la Recherche Médicale (FRM) and ANR.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bahena-Ceron, R. et al. (2023). Methods to Analyze Post-transcriptional Modifications Applied to Stable RNAs in Staphylococcus aureus. In: Barciszewski, J. (eds) RNA Structure and Function. RNA Technologies, vol 14. Springer, Cham. https://doi.org/10.1007/978-3-031-36390-0_11
Download citation
DOI: https://doi.org/10.1007/978-3-031-36390-0_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-36389-4
Online ISBN: 978-3-031-36390-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)